Abstract
Gastroesophageal reflux disease (GERD) is the main risk factor for Barrett’s tumorigenesis. In this study, we investigated the role of NRF2 in response to exposure acidic bile salts (ABS), in conditions that mimic GERD, using Barrett’s esophagus cell models. We detected an increase in NRF2 protein levels, following exposure to ABS. We found oxidization of cysteines (cysteines with oxidized thiol groups) in KEAP1 protein with a weaker interaction between NRF2 and KEAP1, following ABS exposure. Treatment with bile salts increased nuclear NRF2 levels, enhancing its transcription activity, as measured by an ARE (antioxidant response element) luciferase reporter assay. The mRNA expression levels of NRF2 target genes, HO-1 and GR, was increased in response to ABS exposure. Using genetic overexpression and knockdown of NRF2, we found that NRF2 has a critical role in suppressing ABS-induced ROS levels, oxidative DNA damage, DNA double strand breaks, and apoptosis. Collectively, our results suggest that transient induction of NRF2 in response to ABS, plays a pivotal role in protecting esophageal cells by maintaining the levels of oxidative stress and DNA damage below lethal levels under GERD conditions.
Keywords: Barrett’s, oxidative stress, DNA damage
Introduction
The incidence of esophageal adenocarcinoma (EAC) has been rising rapidly during the past few decades in the United States and other Western countries[1–3]. The development of Barrett’s esophagus (BE), where the normal squamous epithelium is replaced by intestinal glandular metaplastic epithelium, due to chronic gastroesophageal reflux disease (GERD), is the main risk factor for EAC [4–6]. The abnormal reflux of acidic bile salts (ABS) into the lower esophagus generates high levels of reactive oxygen species (ROS) and oxidative stress that induce oxidative DNA damage. Chronic oxidative stress and DNA damage are high risk factors associated with tumor initiation and progression of EAC[7–9]. The GERD-associated genotoxic environment can lead to cell death, if oxidative stress and DNA damage lesions are uncorrected, reaching lethal levels. Normal cells have adaptive intact anti-oxidative response systems that counteract the toxic reactive species, such as ROS, hydrogen peroxide (H2O2), reactive nitrogen species (RNS), and nitric oxide (NO)[10, 11]. While low concentrations of these potentially toxic molecules are used for adaptive intracellular signaling, their physiological concentrations must be tightly regulated.
Nuclear factor erythroid 2-related factor 2 (NRF2) is a transcription factor which belongs to the Cap ‘N’ Collar (CNC) family that contains a conserved basic leucine zipper (bZIP) structure that regulates the expression of antioxidant proteins that protect against oxidative damage triggered by injury and inflammation. Under normal conditions, NRF2 binds to its physiological inhibitor KEAP1 (Kelch-like erythroid cell-derived protein with CNC homology [ECH]-associated protein 1). KEAP1 is a thiol-rich protein that contains multiple reactive cysteines. Under basal conditions, KEAP1 functions as an adaptor protein in the Cul3-based E3 ligase complex which mediates constant degradation of NRF2 through the ubiquitin-proteasome pathway [12–14]. The half-life of the NRF2 protein is usually short under normal conditions[15], which maintains cellular NRF2 at a low level to prevent constitutive activation of antioxidant response. Upon exposure to stresses, KEAP1 is inactivated, mainly by direct modification of cysteine thiol residues, resulting in decreased NRF2 ubiquitination and degradation[15, 16]. Accumulation of NRF2 is followed by its nuclear localization to activate transcription of numerous target genes that contain the antioxidant response element (ARE)/electrophile response element (EpRE), in cell and context dependent manners [17]. NRF2 target genes encode detoxification enzymes, such as NAD(P)H, quinone oxidoreductase 1 (NQO1) and glutathione S-transferases (GSTs); antioxidant enzymes like heme oxygenase-1 (HO-1); glutathione reductase (GR) and peroxiredoxins (PRDX), and several other enzymes involved in glutathione (GSH) synthesis and metabolism, such as glutamate–cysteine ligase (GCL) subunits.
In the present study, we investigated the role of NRF2 in protecting BE epithelial cells from abnormal exposure to ABS in conditions that mimic GERD. We demonstrated that NRF2 is transiently induced in response to ABS, a step that is critical in reducing ABS-induced oxidative stress, DNA damage, and cell death.
Materials and Methods
Cell lines
Three immortalized cell lines originating from non-neoplastic Barrett’s esophagus, BAR-10T (kindly provided by Dr. Rhonda Souza), CPA and CPB (purchased from American Type Culture Collection, ATCC, Manassas, VA) were used in this study. Cells were cultured at 37°C with 5% CO2 in DMEM/F12 (1:1) medium supplemented with 0.4 μg/ml hydrocortisone, 20 mg/L adenine, 140 μg/ml bovine pituitary extract, 20 ng/ml recombinant human epidermal growth factor and 0.1% ITS (Insulin-transferrin-sodium selenite), and 5% FBS.
Bile salts
A bile salts cocktail, consisting of an equal molar mixture of sodium salts of glycocholic acid, taurocholic acid, glycodeoxycholic acid, deoxycholic acid (CALBIOCHEM, La Jolla, CA), and glycochenodeoxycholic acid (Sigma-Aldrich, Saint Louis, MO), was prepared to reflect the mixture of bile salts in the distal esophagus during GERD, as previously reported [9, 18, 19]. To mimic pathophysiology conditions of GERD, we used the bile salts cocktail in acidic medium (pH4, ABS) to treat Barrett’s epithelial cells.
Quantitative real time RT-PCR (qRT-PCR)
Total RNA was isolated using the Direct-Zol RNA kit (Zymo Research, Irvine, CA). Single-stranded complementary DNA was subsequently synthesized from RNA using the iScript cDNA synthesis Kit (Bio-Rad, Hercules, CA). Primers for NRF2 target genes were designed using Primer 3 on line tool (http://bioinfo.ut.ee/primer3–0.4.0/primer3/). The primers were obtained from Integrated DNA Technologies (Coralville, Iowa). The quantitative real-time polymerase chain reaction (qRT-PCR) was performed using an iCycler (Bio-Rad) with the threshold cycle number determined by the use of iCycler software version 3.0. Reactions were performed in triplicate, the threshold numbers were averaged, and fold expression was calculated as previously reported [20]. The mRNA expression results were normalized to the average CT of HPRT1.
Cloning and construction of NRF2 pTetOne inducible expression system
A full length of NRF2 coding sequence [21] with Flag-tag was amplified from normal cDNA by PCR using Platinum PCR SuperMix High Fidelity (ThermoFisher Scientific, Waltham, MA) and cloned into the pTetOne vector (Takara Bio USA, Mountain View, CA). The pTetOne-NRF2 plasmid was co-transfected with linear hygromycin marker (Takara Bio USA) at 20:1 ratio, following supplier instructions. 72 h after transfection, the cells were treated with hygromycin at 100 μg/ml (for CPA cells) to select cells stably expressing the vector. Doxycycline (DOX, 100 ng/ml) was added to cells for 72 h to induce NRF2 expression. Induction of NRF2 was validated using qRT-PCR and Western blotting.
Construction of NRF2 knockdown cell model
NFE2L2 (NRF2) sgRNA crispr/cas9 all-in-one lentivector was purchased from Applied Biological Materials (Abm, Richmond, BC, Canada) and co-transfected with second generation packaging mix (Abm) into 293LTV cells (Cell Biolabs, San Diego, CA) following supplier’s protocol. Cells were infected with NRF2 lentiviral particles with polybrene at a concentration of 8 μg/ml. Cells were selected with puromycin and single clones were isolated. NRF2 mRNA and protein levels were examined using qRT-PCR and Western blotting.
Detection of oxidized cysteine sulfates of KEAP1 protein
CPB cells were treated with ABS (pH4, 200 μM) for 20 min, then cells were cultured with regular medium with 5 μM Dimedone for 1 h. Cells were harvested and co-immunoprecipitation (co-IP) was performed with antibody against KEAP1 (Proteintech, 10503–2-AP, Rosemont, IL) using Dynabeads protein A for immunoprecipitation (Thermofisher). The input and IP proteins were loaded and separated by sodium dodecyl sulphate polyacrylamide gel electrophoresis and transferred to nitrocellulose membranes. We used a rabbit antibody which recognizes all forms of cysteine with an oxidized thiol group (sulfenic RSOH, sulfinic RSO2H and sulfonic RSO3H) (Enzo Life Science, Farmingdale, NY)[22]. To avoid heavy chain background, a second mouse antibody against rabbit IgG light chain was applied (Cell Signaling, Danvers, MA), followed by anti-mouse IgG with HRP (Cell Signaling). Immuno-reactive protein bands were visualized by enhanced chemiluminescence (Thermofisher) and images were captured by a ChemiDoc XRST Image System (Bio-Rad).
Detection of NRF2 and KEAP1 interaction by co-immunoprecipitation assay
Co-immunoprecipitation assay was performed using protein-G magnetic beads (Millipore, Billerica, MA, USA) and the primary antibodies against NRF2 (Abcam, ab62352) and KEAP1 (Proteintech, 10503–2-AP), according to the supplier’s instruction. IgG from rabbit (Cell signaling) was used as the negative control. The same amount of protein from each group was incubated with antibody-bound beads, continuously mixing at 4°C overnight. The input and IP proteins were denatured at 90°C for 10 min. Western blot analysis was performed following standard protocols using the primary antibodies against NRF2 (Abcam, ab62352) and KEAP1 (Proteintech, 10503–2-AP). For KEAP1 Western blotting, we used a mouse-anti-rabbit IgG light chain as the secondary antibody followed by anti-mouse IgG antibody with HRP (Cell signaling) to reduce the background of heavy chain.
3D organotypic culture and bile acids treatment
3D organotypic reconstruct cultures were established as previously described[21, 23]. Briefly, human esophageal fibroblasts (ScienCell, Carlsbad, CA) were seeded into a 3D matrix (75,000 cells/well) containing collagen I (High concentration rat tail collagen, Corning, Corning, NY) and Matrigel (BD Biosciences, Franklin Lakes, NJ) and incubated for 7 days at 37° C. Then, CPB cells were seeded (500,000 cells/well) on top of the fibroblast matrix. Cultures were incubated for an additional 14 days. Cells were treated with ABS (pH4, 200 μM) by adding the ABS cocktail into the chambers covering the epithelial cells for 30 min. The culture medium was replaced by regular medium for 2 h. The 3D cultures were harvested, fixed in 4% paraformaldehyde for 1 h, and then transferred to 70% ethanol for overnight, followed by embedding paraffin blocks. Blocks were cut into glass slides and processed for HE staining and immunocytochemistry.
Detection of NRF2 nuclear translocation by immunofluorescence
Cells were seeded in 8-chamber culture slides. On the second day, cells were treated with ABS (pH4, 200 μM) for 20 min, followed by replacement with regular culture medium for 2 h. Cells were fixed with 4% paraformaldehyde in PBS at room temperature for 45 min, followed by permeabilization with 0.5% Triton X-100 in PBS for 2 min on ice. After blocking, cells were incubated with primary antibody against NRF2 (ABE413, Millipore Sigma, Burlington, MA) overnight at 4° C. Cells were incubated with anti-rabbit secondary antibody labelled with Fluor-488 for 1 h at room temperature. After washing, cells were covered with Vectashield mounting medium with DAPI (Vector Laboratories, Burlingame, CA).
Detection of NRF2 ARE luciferase reporter activity
BE cells were seeded into 12-well plates. On the second day, cells were co-transfected with PGL4.37[luc2P/ARE/Hygro] vector (Promega, Madison, WI) and pCMV-beta-gal vector (Promega). 48 h after transfection, cells were treated with bile salts cocktail (BS) at a concentration of 200 μM for 6 h, together with control treatment of PBS (we did not use BS in acidic medium, because we found that acidic conditions significantly impairs luciferase activity). Cells were harvested and lysed with 1x luciferase lysis buffer. Lysates were divided into two parts: for measurement of luminescence of ARE luciferase activity and for measurement of absorbance of β-galactosidase using a FLUOstar OPTIMA microplate reader (BMG LABTECH, Cary, NC). ARE luciferase activity results were normalized to β-galactosidase expression levels of each sample. The experiments were set in triplicate.
Immunofluorescence of 3D organotypic cell culture
We carried out immunofluorescence staining in paraffin-embedded 3D culture slides as previously described[23, 24]. Primary antibodies (anti-NRF2, Millipore; anti-HO-1, Proteintech) were incubated over night at 4° C. Secondary anti-mouse or anti-rabbit antibody with Fluor-488 or Fluor-586 labeling were applied for 1 h at room temperature, protected from light. Slides were then mounted with Vectashield mounting medium with DAPI (Vector Laboratories). Images were examined and documented using an Olympus BX41 fluorescence microscope.
Detection of oxidative DNA damage and DNA double strand breaks
CPA cells expressing the NRF2-pTetOne Inducible System were seeded into 8-chamber culture slides and treated with doxycycline (100 ng/ml) for 48 h. CPB cells with or without NRF2 knockdown were seeded into 8-chamber culture slides. On the second day, cells were treated with ABS (pH4, 200 μM) for 20 min, then cells were returned to regular culture medium for 2 h. Cells were fixed with 4% paraformaldehyde in PBS at room temperature for 45 min and immunofluorescence assay was performed as described above. Primary antibodies against p-H2A.X (ser139) (80312, Cell Signaling) and 8-Oxyguanine (MAB3560, Millipore, Sigma) were applied. p-H2A.X and 8-oxoGuanine nuclear staining positive cells were counted. We counted at least 400 cells from four different areas using the ImageJ software. The percentage of nuclear positive cells versus total number of cells was calculated and statistically analyzed using the GraphPad Prism software.
Western blot analysis
Cells were lysed using RIPA buffer in the presence of proteinase inhibitor cocktail and phosphatase inhibitor (Santa Cruz Biotechnology, Dallas, TX). Protein concentration was determined by a BCA Protein Assay (Thermo Scientific) using a FLUOstar OPTIMA microplate reader (BMG). Equal amounts of proteins were loaded and separated by sodium dodecyl sulphate polyacrylamide gel electrophoresis and transferred to nitrocellulose membranes, followed by Western blot analysis using standard protocols. The primary antibodies were: anti-NRF2 antibody (Millipore), Anti-KEAP1 antibody (Proteintech), anti-cleaved caspase 3 and caspase 3 antibodies (Cell Signaling), anti-cleaved PARP and PARP antibodies (Cell Signaling), anti-histone H3 antibody, anti-β-tubulin antibody (Cell Signaling), and anti-β-actin antibody (Sigma). Horseradish peroxidase-conjugated anti-mouse and anti-rabbit secondary antibodies were purchased from Cell Signaling Technology. Immunoreactive protein bands were visualized by enhanced chemiluminescence and images were captured by a ChemiDoc XRST Image System (Bio-Rad).
Statistical Analysis
Data were expressed as the mean ± standard error of the mean for parametric data. Statistical analysis was performed using unpaired Student t test for independent two samples. 1-way analysis of variance (ANOVA) with post-test comparison was used to analyze the independent samples of 3 or more. All statistical analyses were done using the GraphPad Prism software. For all analyses, p<0.05 was considered statistically significant.
Results
NRF2 protein level is induced following a short exposure to acidic bile salts
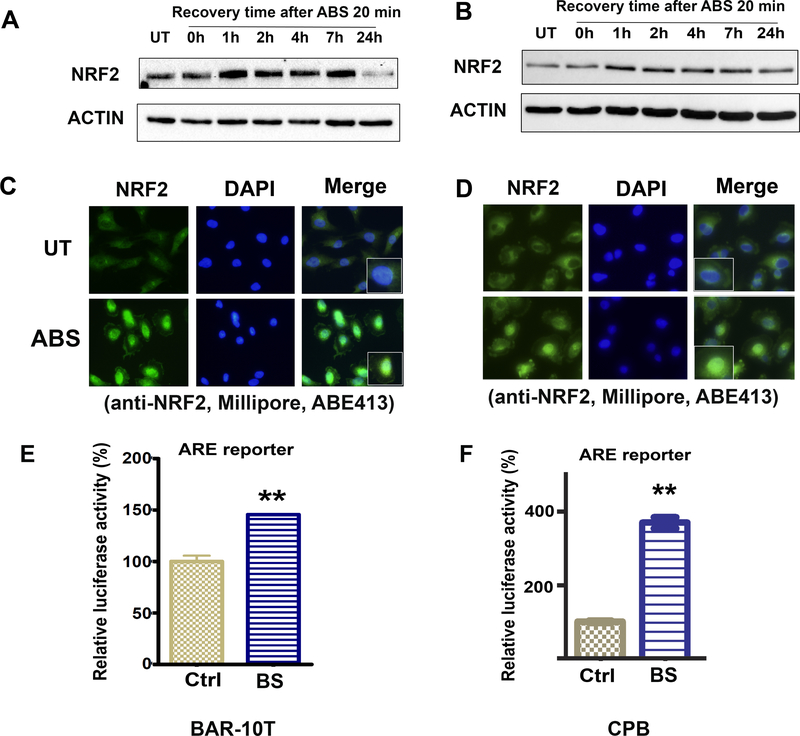
To mimic the pathophysiological condition of a gastro-esophageal reflux episode, we treated BAR-10T, and CPB cells with ABS cocktail (200 μM, pH4) for 20 min, followed by replacement with a regular culture medium. Cells were allowed to recover for 1, 2, 4, 7, and 24 h. We detected induction of the NRF2 protein level at the 1 h recovery time point, that returned back to baseline after 24 h (Figure 1A and 1B). Next, we performed an immunofluorescence assay to determine nuclear localization of NRF2. As shown in Figure 1 C (BAR-10T) and D (CPB), the majority of induced NRF2 protein was localized to the nucleus, following exposure to ABS. The nuclear accumulation of NRF2 protein was further confirmed by Western blot of cytosolic and nuclear fraction of cell lysates (Supplementary Figure 1, A and B). Using a luciferase reporter assay with a luc2P reporter that contains 4 copies of antioxidant response elements (ARE), as a measure of NRF2 activity, we found that exposure of cells to bile salts (BS) can significantly induce the reporter activity in BAR-10T and CPB cells (Figure 1 E and F). Similar results were obtained in another BE cell line, CPA cells (Supplementary Figure 2).
Figure 1. NRF2 protein is transiently induced and activated by short exposure to acidic bile salts.
BAR-10T (A) and CPB (B) cells were exposed to acidic bile salts (ABS, pH4, 200 μM) for 20 min, then recovered in regular culture media for the designated time points. Western blot analysis of NRF2 protein was carried out. C and D, immunofluorescence staining of NRF2 protein (green) in BAR-10T (C) and CPB (D) cells. Cells were exposed to ABS for 20 min and recovered for 2 h, based on Figure 1 A and B. DAPI (blue) was used to stain nucleus. E and F, luciferase assay to evaluate the ARE (antioxidant response element) reporter activity in BAR-10T (E) and CPB (F) cells after cells were exposed to bile salts cocktails (we used BS instead of ABS in this assay, because we found that acidic conditions significantly impaired luciferase activity). Data presented as relative luciferase activity after normalized to β-galactosidase expression level of the same samples. Ctrl, control with PBS; BS, bile salts.
Up-regulation of heme oxygenase-1 (HO-1) and glutathione reductase (GR) expression levels in response to acidic bile salts
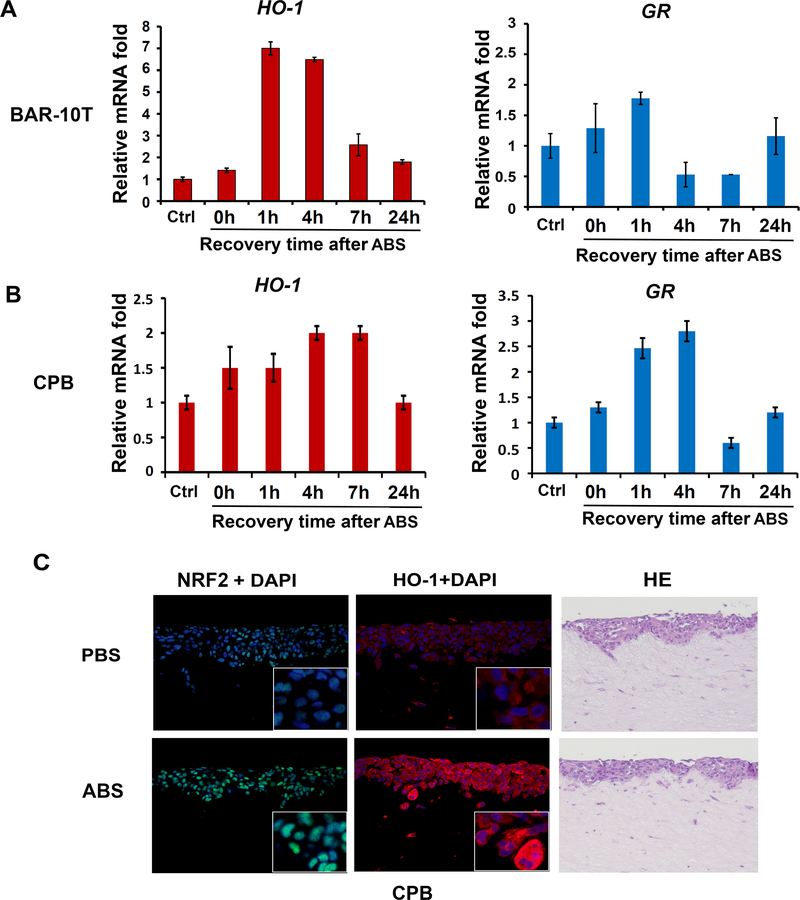
NRF2 plays a pivotal role in regulating antioxidant response through transcriptional regulation of target genes that contain ARE. By using a luciferase reporter assay that measures the ARE activity, the results demonstrated induction of reporter activity in response to bile salts treatment (Figure 1). Therefore, we used qRT-PCR to determine the expression of several known NRF2 target genes that are related to oxidative stress; GR, HO-1, GSTM2, GSTM3, GPX1, GPX3, GPX4, GPX7, NQO1 and GSTP1 (Supplementary Figure 3). Two genes, heme oxygenase-1 (HO-1) and glutathione reductase (GR) were the most significantly and consistently induced in response to bile salts in all the cell lines that we tested (BAR-10T, CPA, CPB), whereas the up-regulation of other genes was limited to one or two cell lines (Supplementary Figure 3). In concordant with the increase NRF2 protein level and activity, the up-regulation of HO-1 and GR expression after ABS exposure was transient too; the levels of HO-1 and GR returned to baseline after 7–24 h (Figure 2, A and B). To mimic GERD in physiological 3D growth pattern of epithelial cells, we established 3D organotypic culture of CPB cells and treated the culture with ABS for 30 min. Immunofluorescence staining of organotypic culture cells confirmed the nuclear translocation of NRF2 protein and induction of HO-1 protein after ABS treatment (Figure 2C).
Figure 2. Transient exposure to acidic bile salts mediates induction of NRF2 target genes.
BAR-10T (A) and CPB (B) cells were exposed to acidic bile salts (ABS, pH4) for 20 min, then returned to regular culture media for the designated time points. qRT-PCR was carried out to quantify the mRNA levels of NRF2 target genes, normalized to that in control cells. Two representative target genes, HO-1 and GR are shown. Results of other NRF2 target genes are shown in Supplementary Figure 3. C. CPB cells were cultured in a 3D organotypic culture model as described in the Methods section. At the end of 3D culture, cells were exposed to ABS for 30 min, recovered for 1 h in regular medium. Immunocytofluorescence assay was applied for NRF2 protein (green) and HO-1 protein (red). DAPI (blue) was used to stain nucleus. HE staining images were used as control.
Acidic bile salts induced oxidation of cysteines in KEAP1
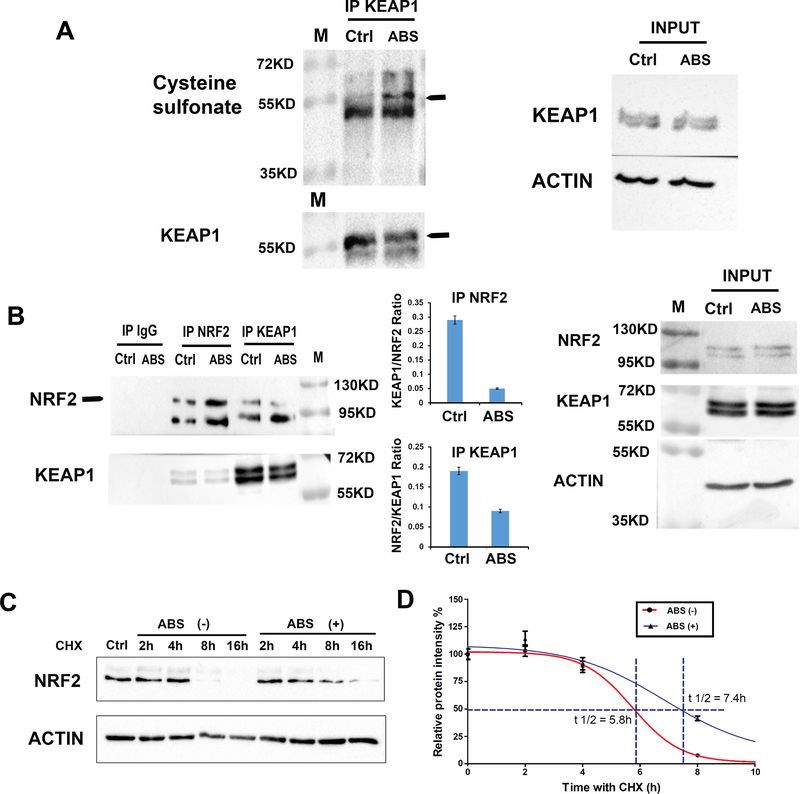
KEAP1 (Kelch-like erythroid cell-derived protein with CNC homology [ECH]-associated protein 1) is a known NRF2 physiological inhibitor[14, 25, 26]. Oxidation of cysteines in KEAP1 results in release of NRF2 from KEAP1, allowing its accumulation. To investigate if ABS treatment can affect cysteine oxidation of KEAP1, CPB cells were treated with ABS followed by incubation with 5 μM dimedone for 1 h. Dimedone is a cell-permeable, nucleophilic reagent that reacts selectively with the electrophilic sulfur atom in sulfenic acid to form a stable thioether product[22]. Immunoprecipitation (IP) was carried out with an anti-KEAP1 antibody (Proteintech), then blotted with an antibody recognizing dimedone-derivatized cysteine sulfate. As shown in Figure 3A, ABS induced more oxidized cysteine sulfate of KEAP1 (arrowed bands). Concordant with this finding, the KEAP1-NRF2 protein-protein interaction was reduced after ABS exposure, as compared to control cells, with an overall increase of NRF2 level after ABS (Figure 3B). Next, we examined the NRF2 protein stability using cycloheximide (CHX, a protein synthesis inhibitor) with/without ABS treatment. Data revealed that ABS treatment significantly prolonged NRF2 half-life time (Figure 3, C and D).
Figure 3. Acidic bile salts induce cysteine oxidation of KEAP1 and enhanced NRF2 stability in esophageal epithelial cells.
A, CPB cells were treated with acidic bile salts (ABS) or PBS (Ctrl), then returned to regular media with dimedone for 1 h. Western blot analysis, using a specific antibody recognizing dimedone-derivatized cysteine sulfates, was applied to co-IP proteins using antibody against KEAP1. Bands with arrowhead display the KEAP1. B, CPB cells were treated with acidic bile salts (ABS) or PBS (Ctrl), then returned to regular media for 1 h. co-IP using antibodies against NRF2 and KEAP1 were performed and Western blot analysis was used to detect the binding between NRF2 and KEAP1. Bands with arrowhead (upper bands) indicates the NRF2. Quantification of bands intensity was performed with ImageJ, the KEAP1/NRF2 ratio (when IP NRF2) and NRF2/KEAP1 ratio (when IP KEAP1) were shown. C, CPB cells were treated with ABS or PBS, then returned to regular medium with existence of cycloheximide (CHX, 100 μg/ml). Cells from different time points were collected and subjected to Western blot analysis. The bands’ intensities were quantified with ImageJ. The NRF2 half-life time was plotted in Panel D.
NRF2 protects esophageal cells from acidic bile salts-induced oxidative stress, oxidative DNA damage, and double strand breaks
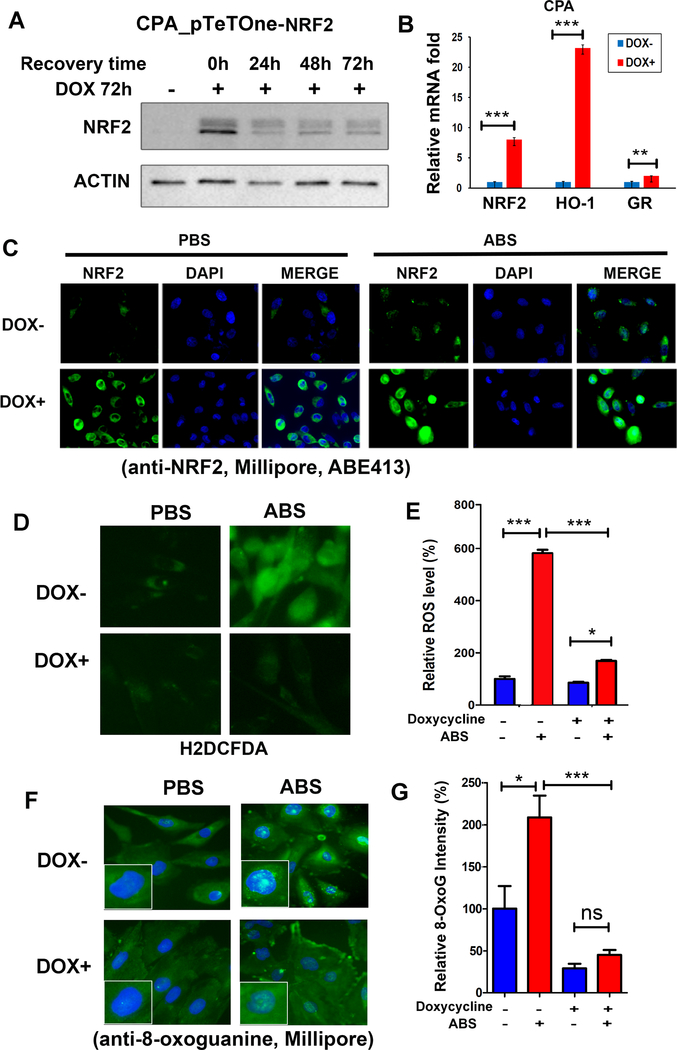
To examine the role of NRF2 in esophageal epithelial cells, we generated a doxycycline inducible NRF2 system in CPA cells using the pTetOne-NRF2 expression plasmid. Western blot analysis and immunofluorescence assay confirmed that NRF2 protein is inducible in CPA cells, following doxycycline treatment (Figure 4 A and C) and NRF2 target genes were induced (Figure 4 B). Treatment of CPA cells with ABS induced significant intracellular ROS, however, this increase of ROS was diminished in doxycycline-treated pTetOne-NRF2 cells (Figure 4 D and E). Similarly ABS treatment induced a significant increase of oxidative DNA damage, as measured by positive nuclear staining of 8-oxoguanine (Figure 4 F and G), an oxidative DNA damage marker [27]. However, induction of NRF2 by doxycycline-treatment of pTetOne-NRF2 cells, significantly reduced levels of 8-oxoguanine lesions by ABS (Figure 4 F and G). In addition, we also measured levels of double strand breaks using p-H2A.X (ser 139) immunofluorescence. The levels of p-H2A.X foci, following exposure to ABS, were significantly lower in doxycycline-treated pTetOne-NRF2 cells, as compared to controls (Figure 5 A and B). In addition, we also transfected CPA cells with a NRF2-expressing plasmid with EGFP-tag. Figure 5, panels C and D, confirms overexpression of NRF2 with induction of NRF2 target genes in these cells. Exposure to acidic bile salts induced a significant increase in double stand breaks foci, as measured by p-H2AX (ser 139) immunofluorescence. However, cells expressing NRF2-EGFP (green) exhibited absent to low levels of double stand break foci (Figure 5E).
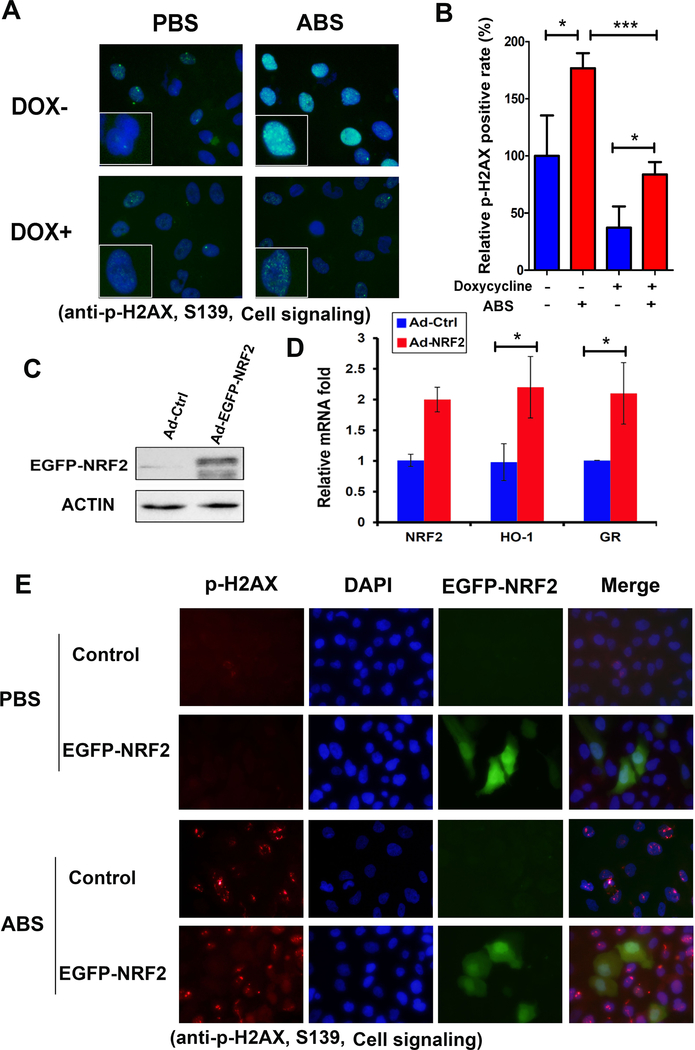
Figure 4. Reconstitution of NRF2 protects esophageal cells from acidic bile salts-induced oxidative stress and oxidative DNA damage.
A, NRF2 protein is induced by doxycycline (DOX) in CPA cells stably transfected with the pTetOne Inducible System. B, qRT-PCR confirmed that NRF2 target genes were induced upon DOX treatment. C, immunofluorescence assay to detect NRF2 protein (green) in cells with/without DOX and exposure to ABS. D, acidic bile salts (ABS, pH4) induced a significant generation of intracellular ROS indicated by green fluorescence (H2DCFDA). However, the generation of ROS was significantly suppressed in the presence of DOX-induced NRF2. E, similar results were obtained with flow cytometry assay to quantitate the intracellular ROS. F, immunofluorescence assay for 8-oxoguanine (8-OxoG), an oxidative DNA damage marker. ABS induced marked oxidative DNA damage (green color, merged with DAPI), located predominantly in nucleus. However, in the presence of DOX-induced NRF2, followed by ABS, the level of oxidative DNA damage was significantly inhibited. G, quantification data of oxidative DNA damage by using ImageJ. *, p<0.05; ***, p<0.001, ns, not significance.
Figure 5. Exogenous expression of NRF2 protects esophageal epithelial cells from acidic bile salts-induced double strand breaks.
A, immunofluorescence assay for p-H2AX (ser 139) protein, a DNA double strand break marker in CPA cells stably transfected with the pTetOne Inducible System. ABS induced high levels of DNA double strand breaks (green color, merged with DAPI) in cells without DOX. However, in the presence of DOX-induced NRF2, ABS-induced DNA double strand break was significantly inhibited. B, quantitative data of DNA double stand breaks in panel A, using ImageJ. *, p<0.05; ***, p<0.001. C, CPA cells were infected with NRF2-EGFP adenoviral particle, Western blot shows that NRF2 protein is enforced expressed. D, qRT-PCR shows significant induction of HO-1 and GR, two representative NRF2 target genes, following enforced NRF2 expression. E, CPA cells were expressed with NRF2-EGFP followed by ABS treatment. Immunofluorescence assay was used to identify the DNA double stand breaks foci, as indicated by red fluorescence of p-H2AX (ser 139) protein. ABS exposure induced DNA double strand breaks in cells without NRF2-EGFP expression; however, cells with ectopic expression of NRF2 (green) were devoid of DNA double strand breaks.
To confirm the above results, stable knockdown of NRF2 in CPB cells, where endogenous NRF2 levels are relatively higher than CPA, was established using crispr/cas9. As shown in Figure 6A, the crispr/cas9 cells (NRF2 KD) expressed significantly lower NRF2 levels, as compared to control cells. Knockdown of NRF2 led to a significant decrease in NRF2 target genes, HO-1 and GR (Figure 6B). Treatment of these cells with ABS induced significantly higher intracellular ROS levels than that in control cells, indicating that knockdown of NRF2 sensitized cells to oxidative stress (Figure 6C). Not surprisingly, the oxidative DNA damage level, as measured by positive 8-oxoguanine lesions, was significantly higher in NRF2 knockdown (NRF2 KD) cells, following exposure to ABS, as compared to control cells (Figure 6 D and E). Similarly, knockdown of NRF2 sensitized CPB cells to ABS-induced double strand breaks (Figure 6 F and G). To further confirm the protective role of NRF2, we reconstituted NRF2 in CPB cells with stable crispr/cas9 NRF2 knockdown. The reconstitution of NRF2 reduced ABS-induced oxidative DNA damage lesions (Supplementary Figure 4). Collectively, our results demonstrate a protective role of NRF2 against oxidative DNA lesions and double strand breaks in conditions of acidic bile salts exposure that mimic GERD.
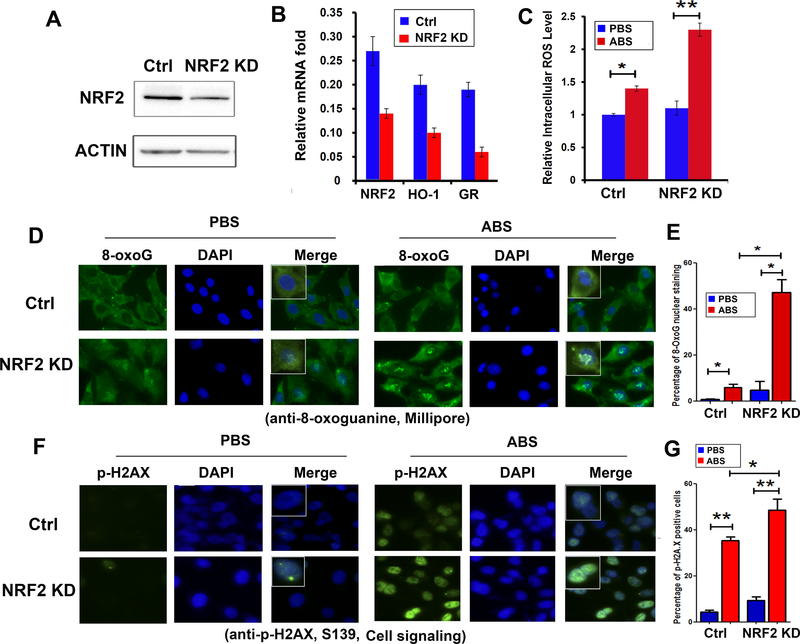
Figure 6. Knockdown of NRF2 sensitizes esophageal cells to acidic bile salts-induced oxidative stress, oxidative DNA damage and DNA double strand breaks.
A, Western blot analysis of NRF2 protein after knockdown of NRF2 (NRF2 KD) in CPB cells. B, qRT-PCR shows significant down-regulation of HO-1 and GR, two representative NRF2 target genes, following NRF2 knockdown. C, intracellular levels of ROS were enhanced after NRF2 knockdown, following exposure to ABS. D, immunofluorescence staining of 8-oxoguanine (8-oxoG) in the cells with or without NRF2 knockdown (NRF2 KD), exposed to either PBS or ABS. 8-oxoG is stained in green and merged with DAPI (blue). E, quantitative data of 8-oxoG in D, using ImageJ. F, immunofluorescence staining of p-H2AX (ser 139) in cells with or without NRF2 knockdown (NRF2 KD), exposed to either PBS or ABS. p-H2AX is stained green and merged with DAPI (blue). G, quantitative data of p-H2AX in F, using ImageJ. *, p<0.05; **, p<0.01.
NRF2 protects esophageal cells against acidic bile slats-induced apoptosis
Uncontrolled accumulation of unrepaired DNA damage lesions is a lethal condition. The above results suggest that NRF2 may play a cyto-protective role in esophageal cells, in conditions where cells are abnormally exposed to acidic bile acids, similar to GERD. We, therefore, measured the levels of apoptosis, using Annexin V assay, following exposure to ABS. As shown in Figure 7 A–C, the number of apoptotic cells was significantly increased in the absence of NRF2 (NRF2 KD) as compared to control cells, after exposure to ABS. In agreement with these results, Western blot analysis showed a significant increase in the levels of cleaved caspase 3 and cleaved PARP in NRF2 KD cells, as compared to control cells after exposure to ABS (Figure 7D).
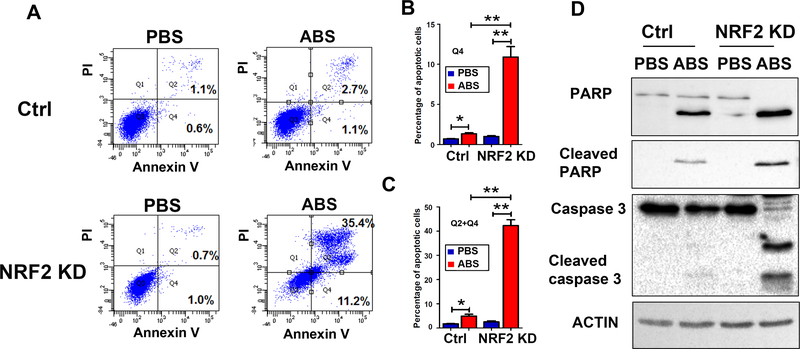
Figure 7. Knockdown of NRF2 promotes acidic bile salts-induced apoptosis in esophageal cells.
A, CPB cells with NRF2 knockdown (NRF2 KD) and control cells (Ctrl) were treated with acidic bile salts (ABS, pH4). Flow cytometry was used to detect annexin V and propidium iodide (PI) staining in cells. Representative flow cytometry profiles are shown. B, Bar graph shows the percentage of annexin V positive cells in NRF2 KD cells, as compared to control cells. C, Bar graph shows the percentage of positive cells with both annexin V and PI staining in NRF2 knockdown (NRF2 KD) cells, as compared to control cells. D, Western blot analysis of cleaved PARP and cleaved caspase 3, two representative apoptosis markers, following knockdown NRF2 and exposure to ABS. *, p<0.05; **, p<0.01.
Discussion
The incidence of GERD and its associated BE is increasing at alarming levels in the United States and other Western countries [4, 6, 28]. Although BE can be viewed as a protective adaptive mechanism in response to chronic GERD, it is also a precancerous condition and a major risk factor for esophageal adenocarcinoma (EAC) [3, 5, 29]. Exposure to acidic bile salts in GERD conditions induces high levels of ROS and DNA damage in esophageal cells [7, 9, 30], events that are linked to tumorigenesis of EAC.
In this study, we used a cocktail of five bile salts, common components of reflux in lower esophagus [18, 19]. We demonstrated that NRF2 was transiently induced following a single 20 min exposure that mimicked a GERD episode. NRF2 is a transcriptional factor that plays a pivotal role in cellular anti-oxidative function through mediating expression of antioxidant genes[12, 31]. Our analyses indicated that NRF2 mediated reducing ROS levels in BE cells. We detected nuclear translocation of NRF2 with activation of its transcription activity, in response to exposure to acidic bile salts. Upon oxidative stress, ROS can oxidize cysteine in KEAP1 molecule and leads to release of NRF2 from KEAP1-NRF2-Cul3 complex, free of ubiquitination degradation[32]. Accordingly, we detected weaker binding of KEAP1 with NRF2, along with increased oxidized cysteine in KEAP1 molecule upon ABS exposure. Although we found that the increase in NRF2 protein level is due to increase in its half-life along with oxidation of KEAP1, the possibility of other additional mechanisms cannot be definitively ruled out. For example, phosphorylation[33] and acetylation[34] of NRF2, interaction with other molecules such as p21[35], and autophagy[36] have been described to regulate NRF2 in the context of cellular stress (reviewed in [37, 38]).
We investigated the expression of several targets of NRF2. We detected a consistent and significant induction of expression of heme oxygenase-1 (HO-1) and glutathione reductase (GR) in several cell models (BAR-10T, CPA, CPB). Of note, NQO1 and GSTP1, reported to be activated by NRF2 in response to some cellular stress stimuli[39], did not show a similar response following exposure to ABS. This data suggests that activation of targets of NRF2 could be cell and context dependent. HO-1 catalyzes the rate-limiting step in heme degradation, leading to the generation of equimolar amounts of iron ions, biliverdin and CO[40, 41]. These molecules contribute to the roles of HO-1 in regulating important biological processes, such as inflammation, apoptosis, and cell proliferation [40–42]. Glutathione reductase (GR), also known as glutathione-disulfide reductase (GSR), is an enzyme that catalyzes the reduction of glutathione disulfide (GSSG) to the sulfhydryl form glutathione (GSH), which is a critical molecule in resisting oxidative stress and maintaining the reducing environment of the cell[43, 44]. Based on our findings, HO-1 and GR appear to be important mediators of response to acidic bile salts in esophageal cells. We acknowledge the need for comprehensive analyses to fully characterize the anti-oxidant targets in esophageal cells in response to the unique oxidative environment created by exposure to acidic bile salts reflux.
We and others have previously reported that acidic bile salts generate high levels of ROS, oxidative stress, oxidative DNA damage and double strand breaks in esophageal cells[7, 9]. Therefore, we investigated whether NRF2 plays a role in regulating these events and maintaining genotoxic damage below lethal levels. Using a number of ectopic expression and knockdown models, our results confirmed that NRF2 was an essential factor in reducing the burden of exposure to ABS by lowering the levels of intracellular ROS, oxidative DNA damage and double strand breaks. In fact, these factors were essential in keeping DNA damage below lethal levels. In absence of NRF2, there was a significant increase in the levels of apoptosis. These data suggest that NRF2 plays a protective role in maintaining cellular homeostasis in oxidative conditions in response to ABS. However, we speculate that the role of NRF2 could be a double sword mechanism where some surviving cells may, over time, accumulate sufficient DNA damage lesions, below lethal levels, that promote esophageal tumorigenesis. This would be consistent with the etiology of EAC where chronic exposure to GERD, over several years or decades, is a risk factor for carcinogenesis.
In summary, our results indicate that transient induction of NRF2, in response to reflux conditions, plays a critical role in regulating oxidative stress and genotoxic events. The antioxidant functions of NRF2 protect esophageal cells and maintain cellular homeostasis in a highly oxidative environment.
Supplementary Material
Highlights.
Gastroesophageal reflux disease (GERD) is the main risk factor for esophageal Barrett’s tumorigenesis.
Using conditions that mimic GERD, we demonstrate transient induction of NRF2 in response to acidic bile salts (ABS).
Acidic bile salts induced oxidation of KEAP1, with release and nuclear accumulation of NRF2.
Activation of NRF2 protected esophageal cells by suppressing ABS-induced ROS levels, oxidative DNA damage, DNA double strand breaks, and apoptosis.
Acknowledgements
This study was supported by development funds from the Sylvester Comprehensive Cancer Center, and grants from the U.S. National Institutes of Health (R01CA206563 and R01CA224366) and the U.S. Department of Veterans Affairs (1IK6BX003787 and I01BX001179). The content of this work is solely the responsibility of the authors and do not necessarily represent the official views of the Department of Veterans Affairs, National Institutes of Health, or the University of Miami.
Footnotes
Conflict of interest: All the authors declare no conflict of interest.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Devesa SS, Blot WJ and Fraumeni JF Jr. Changing patterns in the incidence of esophageal and gastric carcinoma in the United States. Cancer. 1998; 83(10):2049–2053. [PubMed] [Google Scholar]
- 2.Parkin DM, Pisani P and Ferlay J. Global cancer statistics. CA Cancer J Clin. 1999; 49(1):33–64, 31. [DOI] [PubMed] [Google Scholar]
- 3.Conteduca V, Sansonno D, Ingravallo G, Marangi S, Russi S, Lauletta G and Dammacco F. Barrett’s esophagus and esophageal cancer: an overview. Int J Oncol. 2012; 41(2):414–424. [DOI] [PubMed] [Google Scholar]
- 4.di Pietro M Barrett’s esophagus and reflux disease. Endoscopy. 2010; 42(11):910–915. [DOI] [PubMed] [Google Scholar]
- 5.Cossentino MJ and Wong RK. Barrett’s esophagus and risk of esophageal adenocarcinoma. Semin Gastrointest Dis. 2003; 14(3):128–135. [PubMed] [Google Scholar]
- 6.Bonino JA and Sharma P. Barrett’s esophagus. Curr Opin Gastroenterol. 2006; 22(4):406–411. [DOI] [PubMed] [Google Scholar]
- 7.Dvorak K, Payne CM, Chavarria M, Ramsey L, Dvorakova B, Bernstein H, Holubec H, Sampliner RE, Guy N, Condon A, Bernstein C, Green SB, Prasad A and Garewal HS. Bile acids in combination with low pH induce oxidative stress and oxidative DNA damage: relevance to the pathogenesis of Barrett’s oesophagus. Gut. 2007; 56(6):763–771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lee JS, Oh TY, Ahn BO, Cho H, Kim WB, Kim YB, Surh YJ, Kim HJ and Hahm KB. Involvement of oxidative stress in experimentally induced reflux esophagitis and Barrett’s esophagus: clue for the chemoprevention of esophageal carcinoma by antioxidants. Mutat Res. 2001; 480–481:189–200. [DOI] [PubMed] [Google Scholar]
- 9.Peng D, Belkhiri A, Hu T, Chaturvedi R, Asim M, Wilson KT, Zaika A and El-Rifai W. Glutathione peroxidase 7 protects against oxidative DNA damage in oesophageal cells. Gut. 2012; 61(9):1250–1260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Halliwell B Antioxidant defence mechanisms: from the beginning to the end (of the beginning). Free Radic Res. 1999; 31(4):261–272. [DOI] [PubMed] [Google Scholar]
- 11.Sharma R, Yang Y, Sharma A, Awasthi S and Awasthi YC. Antioxidant role of glutathione Stransferases: protection against oxidant toxicity and regulation of stress-mediated apoptosis. Antioxid Redox Signal. 2004; 6(2):289–300. [DOI] [PubMed] [Google Scholar]
- 12.Motohashi H and Yamamoto M. Nrf2-Keap1 defines a physiologically important stress response mechanism. Trends Mol Med. 2004; 10(11):549–557. [DOI] [PubMed] [Google Scholar]
- 13.Itoh K, Wakabayashi N, Katoh Y, Ishii T, Igarashi K, Engel JD and Yamamoto M. Keap1 represses nuclear activation of antioxidant responsive elements by Nrf2 through binding to the amino-terminal Neh2 domain. Genes & development. 1999; 13(1):76–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kobayashi A, Kang MI, Okawa H, Ohtsuji M, Zenke Y, Chiba T, Igarashi K and Yamamoto M. Oxidative stress sensor Keap1 functions as an adaptor for Cul3-based E3 ligase to regulate proteasomal degradation of Nrf2. Mol Cell Biol. 2004; 24(16):7130–7139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Itoh K, Wakabayashi N, Katoh Y, Ishii T, O’Connor T and Yamamoto M. Keap1 regulates both cytoplasmic-nuclear shuttling and degradation of Nrf2 in response to electrophiles. Genes Cells. 2003; 8(4):379–391. [DOI] [PubMed] [Google Scholar]
- 16.Kobayashi M, Itoh K, Suzuki T, Osanai H, Nishikawa K, Katoh Y, Takagi Y and Yamamoto M. Identification of the interactive interface and phylogenic conservation of the Nrf2-Keap1 system. Genes Cells. 2002; 7(8):807–820. [DOI] [PubMed] [Google Scholar]
- 17.Itoh K, Tong KI and Yamamoto M. Molecular mechanism activating Nrf2-Keap1 pathway in regulation of adaptive response to electrophiles. Free Radic Biol Med. 2004; 36(10):1208–1213. [DOI] [PubMed] [Google Scholar]
- 18.Nehra D, Howell P, Williams CP, Pye JK and Beynon J. Toxic bile acids in gastro-oesophageal reflux disease: influence of gastric acidity. Gut. 1999; 44(5):598–602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kauer WK, Peters JH, DeMeester TR, Feussner H, Ireland AP, Stein HJ and Siewert RJ. Composition and concentration of bile acid reflux into the esophagus of patients with gastroesophageal reflux disease. Surgery. 1997; 122(5):874–881. [DOI] [PubMed] [Google Scholar]
- 20.Peng DF, Razvi M, Chen H, Washington K, Roessner A, Schneider-Stock R and El-Rifai W. DNA hypermethylation regulates the expression of members of the Mu-class glutathione S-transferases and glutathione peroxidases in Barrett’s adenocarcinoma. Gut. 2009; 58(1):5–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Chen Z, Hu T, Zhu S, Mukaisho K, El-Rifai W and Peng DF. Glutathione peroxidase 7 suppresses cancer cell growth and is hypermethylated in gastric cancer. Oncotarget. 2017; 8(33):54345–54356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Seo YH and Carroll KS. Profiling protein thiol oxidation in tumor cells using sulfenic acid-specific antibodies. Proc Natl Acad Sci U S A. 2009; 106(38):16163–16168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Le Bras GF, Loomans HA, Taylor CJ, Revetta FL and Andl CD. Activin A balance regulates epithelial invasiveness and tumorigenesis. Lab Invest. 2014; 94(10):1134–1146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Loomans HA and Andl CD. Intertwining of Activin A and TGFbeta Signaling: Dual Roles in Cancer Progression and Cancer Cell Invasion. Cancers (Basel). 2014; 7(1):70–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Jaramillo MC and Zhang DD. The emerging role of the Nrf2-Keap1 signaling pathway in cancer. Genes & development. 2013; 27(20):2179–2191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kansanen E, Kuosmanen SM, Leinonen H and Levonen AL. The Keap1-Nrf2 pathway: Mechanisms of activation and dysregulation in cancer. Redox Biol. 2013; 1(1):45–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Guetens G, De Boeck G, Highley M, van Oosterom AT and de Bruijn EA. Oxidative DNA damage: biological significance and methods of analysis. Crit Rev Clin Lab Sci. 2002; 39(4–5):331–457. [DOI] [PubMed] [Google Scholar]
- 28.Bernstein IT, Kruse P and Andersen IB. Barrett’s oesophagus. Dig Dis. 1994; 12(2):98–105. [DOI] [PubMed] [Google Scholar]
- 29.Drewitz DJ, Sampliner RE and Garewal HS. The incidence of adenocarcinoma in Barrett’s esophagus: a prospective study of 170 patients followed 4.8 years. Am J Gastroenterol. 1997; 92(2):212–215. [PubMed] [Google Scholar]
- 30.Hong J, Chen Z, Peng D, Zaika A, Revetta F, Washington MK, Belkhiri A and El-Rifai W. APE1-mediated DNA damage repair provides survival advantage for esophageal adenocarcinoma cells in response to acidic bile salts. Oncotarget. 2016; 7(13):16688–16702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kobayashi A, Ohta T and Yamamoto M. Unique function of the Nrf2-Keap1 pathway in the inducible expression of antioxidant and detoxifying enzymes. Methods Enzymol. 2004; 378:273–286. [DOI] [PubMed] [Google Scholar]
- 32.Yamamoto M, Kensler TW and Motohashi H. The KEAP1-NRF2 System: a Thiol-Based Sensor-Effector Apparatus for Maintaining Redox Homeostasis. Physiol Rev. 2018; 98(3):1169–1203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sun Z, Huang Z and Zhang DD. Phosphorylation of Nrf2 at multiple sites by MAP kinases has a limited contribution in modulating the Nrf2-dependent antioxidant response. PLoS One. 2009; 4(8):e6588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sun Z, Chin YE and Zhang DD. Acetylation of Nrf2 by p300/CBP augments promoter-specific DNA binding of Nrf2 during the antioxidant response. Mol Cell Biol. 2009; 29(10):2658–2672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Chen W, Sun Z, Wang XJ, Jiang T, Huang Z, Fang D and Zhang DD. Direct interaction between Nrf2 and p21(Cip1/WAF1) upregulates the Nrf2-mediated antioxidant response. Mol Cell. 2009; 34(6):663–673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Lau A, Wang XJ, Zhao F, Villeneuve NF, Wu T, Jiang T, Sun Z, White E and Zhang DD. A noncanonical mechanism of Nrf2 activation by autophagy deficiency: direct interaction between Keap1 and p62. Mol Cell Biol. 2010; 30(13):3275–3285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Silva-Islas CA and Maldonado PD. Canonical and non-canonical mechanisms of Nrf2 activation. Pharmacol Res. 2018; 134:92–99. [DOI] [PubMed] [Google Scholar]
- 38.Bryan HK, Olayanju A, Goldring CE and Park BK. The Nrf2 cell defence pathway: Keap1dependent and -independent mechanisms of regulation. Biochem Pharmacol. 2013; 85(6):705–717. [DOI] [PubMed] [Google Scholar]
- 39.Lee S, Hur EG, Ryoo IG, Jung KA, Kwak J and Kwak MK. Involvement of the Nrf2-proteasome pathway in the endoplasmic reticulum stress response in pancreatic beta-cells. Toxicol Appl Pharmacol. 2012; 264(3):431–438. [DOI] [PubMed] [Google Scholar]
- 40.Loboda A, Damulewicz M, Pyza E, Jozkowicz A and Dulak J. Role of Nrf2/HO-1 system in development, oxidative stress response and diseases: an evolutionarily conserved mechanism. Cell Mol Life Sci. 2016; 73(17):3221–3247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Nitti M, Piras S, Marinari UM, Moretta L, Pronzato MA and Furfaro AL. HO-1 Induction in Cancer Progression: A Matter of Cell Adaptation. Antioxidants (Basel). 2017; 6(2). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wegiel B, Nemeth Z, Correa-Costa M, Bulmer AC and Otterbein LE. Heme oxygenase-1: a metabolic nike. Antioxid Redox Signal. 2014; 20(11):1709–1722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Deponte M Glutathione catalysis and the reaction mechanisms of glutathione-dependent enzymes. Biochim Biophys Acta. 2013; 1830(5):3217–3266. [DOI] [PubMed] [Google Scholar]
- 44.Couto N, Wood J and Barber J. The role of glutathione reductase and related enzymes on cellular redox homoeostasis network. Free Radic Biol Med. 2016; 95:27–42. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.