Abstract
The proteasome multicatalytic proteinase (MCP) is a 20S complex that plays a major role in nonlysosomal pathways of intracellular protein degradation. A polyclonal antibody against rat liver MCP was used to investigate the distribution of MCP in the CNS of the rat and its subcellular localization within the neurons. As expected, MCP immunoreactivity (MCP-IR) was distributed ubiquitously in the rat CNS but not homogeneously. The most intensely stained neurons were the pyramidal cortical neurons of layer 5 and the motor neurons of the ventral horn in the spinal cord, which show an intense nuclear and cytoplasmatic MCP-IR and clearly stained processes. Additionally, some populations of large neurons in the mesencephalon and brainstem also displayed a moderate MCP-IR in their perikarya. The vast majority of neurons in the remaining structures did not show a strong cytoplasmatic MCP-IR, but their nuclei displayed an intense MCP-IR. The subcellular localization also was studied by immunoelectron microscopy. MCP-IR was intense in the neuronal nuclei, and significant staining also was found in the cytoplasm, dendritic, and axonic processes (including some myelinated axons) and in synaptic boutons, as illustrated in the cerebellar cortex. The distribution of MCP in the rat CNS and its subcellular localization are discussed in relation to (1) the distribution of calpain, the other major nonlysosomal cellular protease, and (2) the possible role of MCP in the degradation of regulatory proteins and key transcription factors that are essential in many neuronal responses.
Keywords: multicatalytic proteinase, proteasome, prosome, immunohistochemistry, immunoelectron microscopy, CNS, rat
The multicatalytic proteinase (MCP), proteasome or prosome, is the major nonlysosomal proteinase accounting for the intracellular protein degradation via ubiquitin-dependent and ubiquitin-independent pathways (Rivett, 1993; Ciechanover, 1994;Hochstrasser, 1995). MCP is present in all natural kingdoms. Its structure, in all cases, seems to be a heterodimer, α7β7β7α7composed of (1) a single type of α subunit and β subunit in archebacteria (Lowe et al., 1995), (2) two kinds of α and β subunits in eubacteria (Tamura et al., 1995), and (3) no less than seven different types of α and β subunits in eukaryotes (Tanaka et al., 1992; Heinemeyer et al., 1994). The eukaryotic 20S proteasome appears as the core catalytic component of a 26S complex that is competent for the degradation of both polyubiquitinylated and some nonubiquitinylated proteins (Ciechanover, 1994; Hochstrasser, 1995). The proteasome appears to be responsible for the bulk of protein turnover (Rock et al., 1994) as well as for the degradation of regulatory short half-life proteins such as transcription factors, key-regulatory metabolic enzymes, and cyclins (Hochstrasser, 1995) and for the production of antigenic peptides presented by the class I major histocompatibility complex (Gaczynska et al., 1993; Rock et al., 1994). At the subcellular level, the proteasome has been localized (for review, see Rivett and Knecht, 1993) both in the nucleus and in the cytoplasm. In the latter, it can be found either free or in association with the endoplasmic reticulum (Rivett and Knecht, 1993) and with cytoskeletal components (Olink-Coux et al., 1994).
The importance of MCP in neuronal function is stressed by a recent report that the specific proteasome inhibitor lactacystin (Fenteany et al., 1995) promotes neurite outgrowth of N2A neuroblastoma cell line in culture (Fenteany et al., 1994), indicating that the MCP is implicated in the promotion or maintenance of the differentiated neuronal phenotype apart from its role in cell cycle progression (Katagiri et al., 1995). These results, together with the implication of the proteasome in the degradation of key transcription factors (Hochstrasser, 1995), suggest that it may be playing an important role in the nervous system. However, information concerning the detailed distribution and localization of the MCP complex in the CNS is needed to understand the function of the proteasome in the brain. To this date, a single immunohistochemical study has been reported on the MCP localization in the mammalian brain (Kamakura et al., 1988), apart from a study on the localization of the proteasome in Lewy bodies (Kwak et al., 1991; Masaki et al., 1994). Thus, a detailed study of the distribution and subcellular localization of the proteasome in the rat CNS has been carried out by means of immunohistochemical techniques, both at the light and electron microscopic levels. MCP immunoreactivity (MCP-IR) was found to be distributed heterogenously in the rat CNS, primarily localized in the nucleus, and consistent staining also is found in the cytoplasm, dendritic, and axonic processes (including some myelinated axons) as well as in synaptic boutons.
MATERIALS AND METHODS
MCP purification and antibody characterization. Rat liver and brain MCP were purified as described elsewhere (Arribas and Castaño, 1990). Antibody production against the native rat liver MCP complex was made in rabbits as described previously (Arribas et al., 1991). The polyclonal antibody used in the present studies immunoprecipitates the native MCP complex (Arribas et al., 1991). To verify the specificity of the polyclonal antibody against rat brain, purified rat brain MCP (5 μg) and a total extract of rat brain (100 μg of total protein) were separated on 14% SDS-PAGE, transferred to nitrocellulose, immunoblotted with anti-MCP antibody at 1:200 dilution, and developed with peroxidase-labeled goat anti-rabbit antibody at 1:1000 dilution as described elsewhere (Arribas et al., 1991).
Brain immunohistochemical procedures. Sprague Dawley rats (210–300 gm) were used, fed on stock diet and water supplied ad libitum, and housed under controlled conditions providing light from 7:00 A.M. to 7:00 P.M. The animals were anesthetized with nembutal (sodium pentobarbital, 33 mg/kg, i.p.) and then perfused transcardially with 50 ml of 0.9% saline followed by 500 ml of a fixative solution containing 4% paraformaldehyde in PBS (20 mm NaPi, 100 mm NaCl, pH 7.4) or 4% paraformaldehyde, 0.1% glutaraldehyde in phosphate buffer (PB) (0.1 m NaPi, pH 7.4) for light and electron microscopic studies, respectively. The brains then were removed, blocked, and post-fixed for 3 hr in 4% paraformaldehyde at room temperature and then cryoprotected by overnight immersion in 0.1% PB containing 30% sucrose at 4°C.
For light microscopic studies, consecutive sections (40 μm) were cut through the coronal plane on a freezing microtome. In some cases, cresyl violet stain and acetylcholinesterase histochemistry were developed additionally in adjacent series of sections after a slight modification to Geneser-Jensen and Blackstad method (1971). The sections were washed in TBS (25 mm Tris-Cl, pH 7.5, 0.15 m NaCl) and then preincubated for 1 hr at room temperature in a blocking solution containing TBS, 3% bovine serum albumin (BSA), 10% normal goat serum (NGS), and 0.1% Triton X-100. After blocking, the slices were incubated overnight at 4°C, with the primary antibody at a dilution of 1:200 in TBS containing 3% BSA and 2% NGS. After three washes of the sections at room temperature (10 min each) in TBS, they were incubated for 1 hr at room temperature with a 1:1000 dilution of the secondary antibody (peroxidase-labeled goat anti-rabbit) (Bio-Rad, Hercules, CA) prepared in TBS containing 3% BSA and 2% NGS. Sections were washed again as described above and developed under visual control in TBS containing 0.05% of 3,3′-diaminobenzidine (DAB) (Sigma, St. Louis, MO) and 0.01% H2O2; on one occasion, peroxidase-enhanced developing with 0.01% CoCl2and 0.01% NiS04 was used. The sections then were dehydrated, cleared in xylene and mounted on gelatin-coated slides, air-dried, and coverslipped. Controls with preimmune serum or the secondary antibody alone developed under the same conditions were all negative. The atlas of Paxinos and Watson (1986) was used for the abbreviations and the localization of the brain structures.
Electron microscopy. Fixed and cryoprotected blocks of tissue were frozen rapidly in liquid nitrogen and thawed in cold 0.1 m PB to improve antibody penetration. Coronal sections (40 μm) were cut on a vibratome, and immunostaining was performed as used for the light microscopy except that Triton X-100 was omitted from the incubation solutions. The immunocytochemical reaction was developed under visual control by incubating the tissue sections in PBS containing 0.006% DAB for 10 min, and then 0.003% H2O2 was added to the same solution. The reaction was stopped with 0.1 m PB. Subsequently, the sections were washed in 0.1 MPB, post-fixed in 1% osmium tetroxide in 0.1 m PB for 1 hr, dehydrated in aqueous ethanol of increasing concentrations, and block-stained in uranyl acetate (1% in 70% ethanol) in the dark for 30 min at room temperature. The sections then were mounted on resin slides (Durcupan, Fluka, Ronkonkoma, NY) under a plastic coverslip and incubated for 48 hr at 56°C. Selected areas of the rat brain were dissected out and reembedded in Durcupan. Ultrathin sections were cut, mounted on Formvar-coated copper grids, and studied with a Jeol 1200 EX electron microscope.
RESULTS
Characterization of anti-MCP polyclonal antibody
Figure 1 shows an immunoblot analysis of purified rat brain MCP and total brain homogenate with the anti-MCP polyclonal antibody at dilution 1:200 (the titer of this antibody shows positive reaction up to 1:1000–1:2000 dilution). The polyclonal antibody detected primarily three polypeptide bands of MCP, either in the purified MCP complex or in the total rat brain homogenate, a clear indication of its specificity. Furthermore, this antiserum was able to immunoprecipitate the native MCP complex (Arribas et al., 1991), and similar immunoblot results were obtained with anti-MCP antibody affinity-purified against purified rat liver MCP (data not shown). Preimmune serum from the same rabbit was negative (data not shown). These data clearly show the specificity of the polyclonal anti-MCP antibody used for the immunolocalization studies presented below.
Fig. 1.
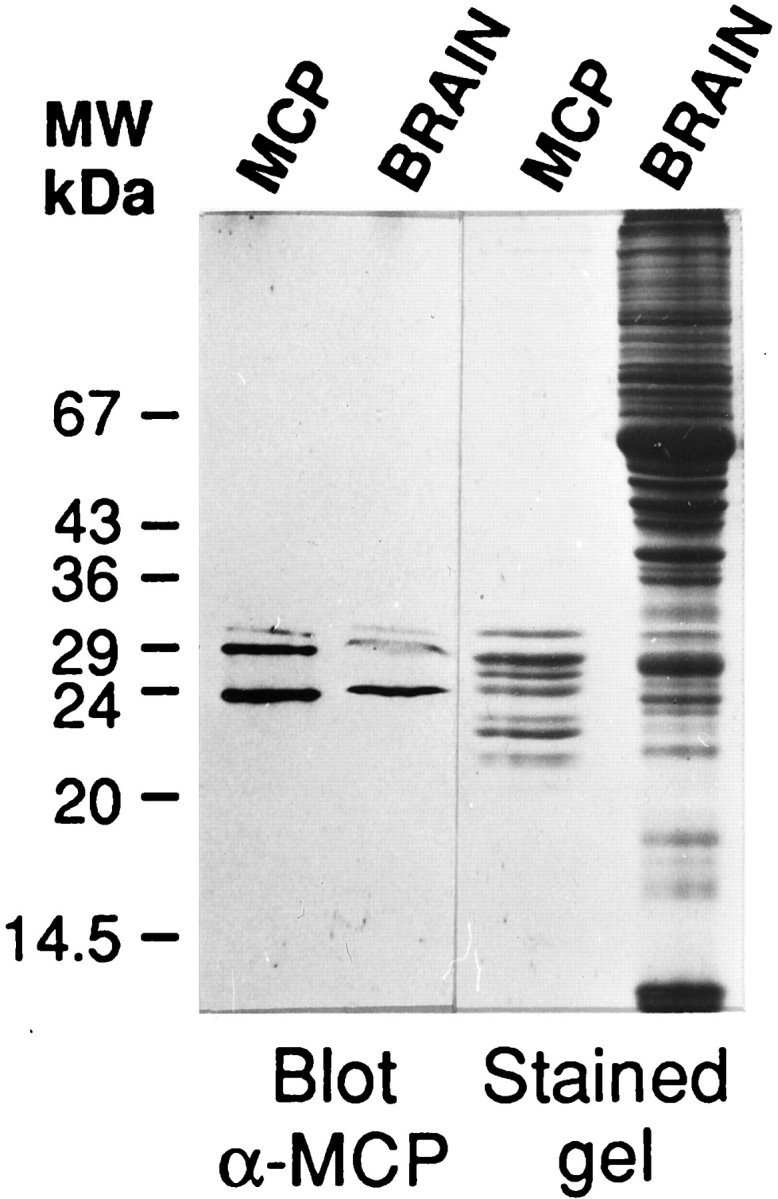
Immunoblot analysis of purified MCP and total brain homogenate with anti-MCP polyclonal antibody. Proteins, purified rat brain MCP (MCP), and total brain homogenate (BRAIN) were separated in 14% SDS-polyacrylamide gels and either stained with Coomassie blue R-250 (stained gel) or blotted to nitrocellulose and probed with the anti-MCP polyclonal antibody at 1:200 dilution (Blotα-MCP). A drawing of the complete gel with the migration of appropriate molecular weight standards is shown.
Distribution of MCP-IR in the rat CNS
The immunolocalization of the MCP complex in the rat CNS was performed with the anti-MCP antibody, the specificity of which is shown in Figure 1, at 1:200 dilution, and no difference in distribution was observed using dilutions that ranged from 1:100 to 1:800 of the primary antibody. As expected, the distribution of MCP-IR in the CNS was found to be ubiquitous (Kamakura et al., 1988); the vast majority of neurons in every structure of the rat CNS displayed MCP-IR and primarily in the cell nucleus (see below). Similar results were obtained when anti-C2 (Arribas et al., 1994) subunit-specific antibodies were used (data not shown).
Cerebral cortex
The great majority of neurons in the neocortex, regardless of their laminar or areal localization, displayed MCP-IR (Fig.2A). However, a remarkably intense immunoreactivity was detected in the pyramidal neurons of layer 5, because of the fact that in addition to the nuclear MCP-IR, immunostaining also was present in the cytoplasm (Fig.3A,A′). Thus, their cell bodies were clearly outlined, and their apical dendrites could be traced ascending toward the superficial layers, in contrast to the rest of cortical neurons. This pattern of immunoreactivity was constant in all neocortical areas. In addition, in both cingular and retrosplenial cortices, the cells in layer 2 were more intensely stained than the neurons in other layers (Fig. 2A).
Fig. 2.

Coronal sections showing MCP-IR in cortical and prosencephalic structures of the rat CNS. A, MCP-IR in the cingular cortex. The six cortical layers are defined at theleft, and the asterisk marks the white matter; the arrowheads point to the pial surface.B, Section through the rostroventral portions of the rat’s brain showing intense MCP-IR in the piriform cortex and the olfactory tubercle (Tu). The asteriskindicates the heterogeneous distribution of MCP-IR in the Tu. Thearrowheads point to the pyramidal layer of the piriform cortex. C, MCP-IR in the hippocampal formation. Note the dense MCP-IR in the granule cells of the dentate gyrus and in the pyramidal cells in fields CA1–3 of Ammon’s horn. The area inbrackets in CA1 is shown at a higher magnification inD. E, Hippocampal formation in a control section incubated with preimmune serum as primary antibody.F, Intense MCP-IR in the bed nucleus of the stria terminalis. Abbreviations for Figures 2, 3, 4, 5, 6, 7: ac, anterior commissure; Ax, axon; b, terminal bouton;BST, bed nucleus of the stria terminalis;CA1, CA2, CA3, fields CA1–3 of Ammon’s horn; CC, central canal;cc, corpus callosum; Cg, cingular cortex;CPu, caudate–putamen complex; cu, cuneate fasciculus; d, dendrite; ER, endoplasmic reticulum; Gi, gigantocellular reticular nucleus; GL, granule cell layer of the cerebellum;gr, gracile fasciculus; icp, inferior cerebellar peduncle; Int, interposed cerebellar nucleus;IOD, dorsal nucleus of the inferior olive;lfu, lateral funiculus of the spinal cord;Lat, lateral cerebellar nucleus; LV, lateral ventricle; LVe, lateral vestibular nucleus;MG, medial geniculate nucleus; ML, molecular layer of the cerebellum; mt, mossy fiber terminal; MVe, medial vestibular nucleus;N, neuron; n, nucleus;Pir, piriform cortex; RSG, retrosplenial granular cortex; S, septum; Sol M, medial nucleus of the solitary tract; vfu, ventral funiculus of the spinal cord; 1–6, spinal cord layers; IV, fourth ventricle; 12, hippoglossal nucleus. Scale bar (shown in D):A–C, E, 1 mm;D, F, 250 μm.
Fig. 3.

Detailed MCP-IR localization in selected cortical and hippocampal regions. A, MCP-IR in the pyramidal cells from layer 5 of the retrosplenial granular cortex. Note the MCP-IR displayed by the majority of the cortical neurons and by the fine apical dendrites ascending toward the pial surface.A′, The same cortical field as in A, shown at a higher magnification. Note the nonimmunoreactive nucleoli within several of the immunoreactive nuclei and the MCP-IR of the perikarya in the larger neurons. The arrowheads point to MCP-IR in the initial segments of apical dendrites; the small arrows mark an apical dendrite traversing the cortical field.B, MCP-IR in the pyramidal neurons of the CA3 hippocampal region. B′, The same field as inB, shown at a higher magnification. MCP-IR is localized primarily in the nuclei of the pyramidal neurons; again, negative nucleoli are clearly distinguishable in some of these neurons. MCP-IR also is seen in the apical dendrites within the stratum radiatum. Scale bar (shown in B): A, B: 60 μm; A′, B′, 100 μm.
Regarding the paleocortex, cells in layer 2 of the piriform cortex were prominently MCP-positive (Fig. 2B), clearly standing out from the surrounding structures. Medial to it, the superficial cell layer of the olfactory tubercle also displayed an intense MCP-IR. Also in the olfactory tubercle, the islands of Calleja could be identified as circular or oval areas showing a moderately stained neuropil in which MCP-positive neurons were immersed (data not shown).
Hippocampus
In Ammon’s horn, the pyramidal cells from CA1 to CA3 were intensely immunoreactive (Fig. 2C,D). MCP-IR was localized primarily in the nuclei, whereas the perikarya apparently were devoid of immunoreactivity. However, within the stratum radiatum, several MCP-positive structures were visible, corresponding to the proximal portions of the apical dendrites of the pyramidal cells (Fig. 3B,B′). Sparse cells, moderately to intensely stained, also were visible in the stratum radiatum and in the stratum lacunosum-moleculare. The granule cells in the dentate gyrus also were very intensely immunoreactive, although no reaction product could be detected in the cell processes. The hilar region also contained abundant MCP-positive cells.
Basal prosencephalon
A dense MCP-IR was found in the septal nuclei, in the amygdalar complex, and especially in the neurons of the bed nucleus of the stria terminalis (Fig. 2F). In addition, it also is interesting to note that MCP-IR varied among the different components of the basal ganglia. Thus, the caudate–putamen complex displayed a light MCP-IR, whereas the neurons in the globus pallidus, entopeduncular nucleus, and subthalamic nucleus were more intensely immunoreactive.
Diencephalon, mesencephalon, and brainstem
The neurons in the paraventricular thalamic nuclei showed the heaviest immunoreactivity within the thalamus, being furthermore immersed in an MCP-positive neuropil. The reticular nucleus also presented an intense MCP-IR, whereas the rest of the thalamic nuclei displayed a moderate immunostaining. The hypothalamus also was strongly immunoreactive, especially in its anterior and ventral areas.
Both at the mesencephalic and brainstem levels, intensely MCP-immunoreactive axons were visible in the cranial nerves (data not shown). The most notable finding was the presence of a moderate MCP-IR within the perikarya in several populations of large neurons, in addition to the nuclear immunostaining, allowing the delineation of the cellular contours of these large neurons and, in most cases, the proximal portions of their neuronal processes also were clearly visible. These neuronal populations comprised the motor nuclei (Fig.4C), the cells of the intermediate gray layer of the superior colliculus, the magnocellular portion of the red nucleus, the lateral vestibular nucleus (Fig.4A,B), and the reticular formation, especially its gigantocellular part (Fig.4D). In the remaining structures, a clear perikaryal immunostaining could not be detected. However, there were other nuclei displaying a strong nuclear MCP-IR, such as the interpeduncular nucleus, which also had a heavy MCP-positive neuropil, the substantia nigra, the pontine nuclei, the mesencephalic nucleus of the trigeminal nerve, and the nuclei of the solitary tract.
Fig. 4.
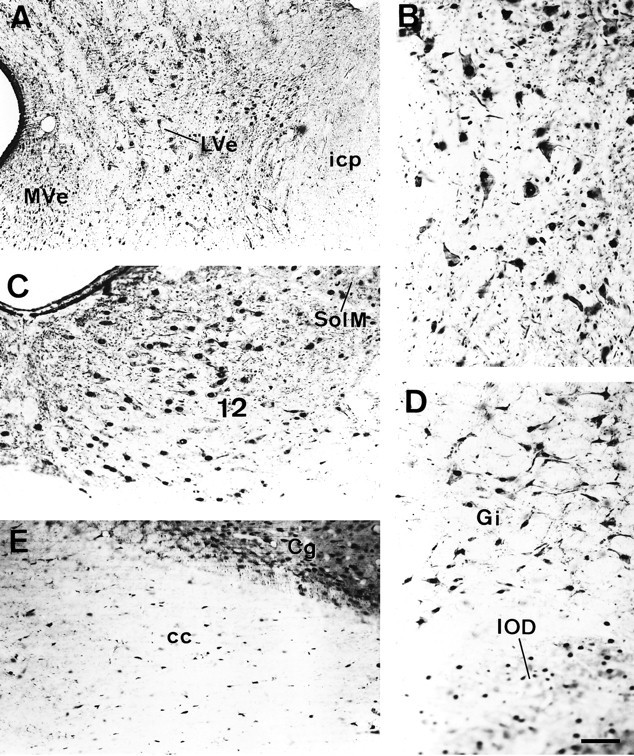
MCP-IR in different areas of the brainstem.A, Coronal section through the lateral vestibular nucleus. B, The same field as in A, shown at a higher magnification. Note the intense MCP-IR displayed by these large neurons in both the nucleus and cytoplasm and, occasionally, in the neuronal processes. C, MCP-IR in the motor nucleus of the 12th cranial nerve and in some neurons of the medial nucleus of the solitary tract. D, Neurons of the gigantocellular reticular formation. The cellular contours are clearly depicted because of the dense MCP-IR in the cytoplasm. The neurons, most of which appear out of focus at the lower portion of the micrograph, correspond to the dorsal nucleus of the inferior olive. E, Coronal section through the corpus callosum. Note the abundant MCP-IR of the cells in the white matter that mostly correspond to glial cells. Scale bar (shown in D): 120 μm.
Cerebellum
The most intense MCP-IR corresponded to the layer of the Purkinje cells (Fig. 5A). Thus, the nuclei of these cells were heavily immunoreactive, whereas their perikarya were faintly stained. However, their cell contours appeared quite neatly outlined, apparently because of MCP-positive structures bordering the neuronal somata or apposed to them (Fig. 5A′, arrowheads). The apical dendrites of the Purkinje cells also were heavily stained and could be seen traversing the molecular layer. In addition, a moderately intense MCP-positive neuropil was present in the molecular layer in which sparse MCP-IR cells were immersed (Fig.5A,A′). In contrast, the granule cell layer displayed a light although variable MCP-IR, ranging from a faint to moderate immunoreactivity (Fig.5A,A′). Also, intensely immunoreactive cells of a larger size appeared sparsely distributed within this cell layer (Fig. 5A,A′). The neurons in the deep cerebellar nuclei were intensely immunoreactive (Fig.5B,B′). These large neurons displayed an intense MCP-IR, both in the nuclei and in the perikarya, so that their somata appeared well-delineated in most cases (Fig. 5B′), and occasionally, some neuronal processes also were visible.
Fig. 5.
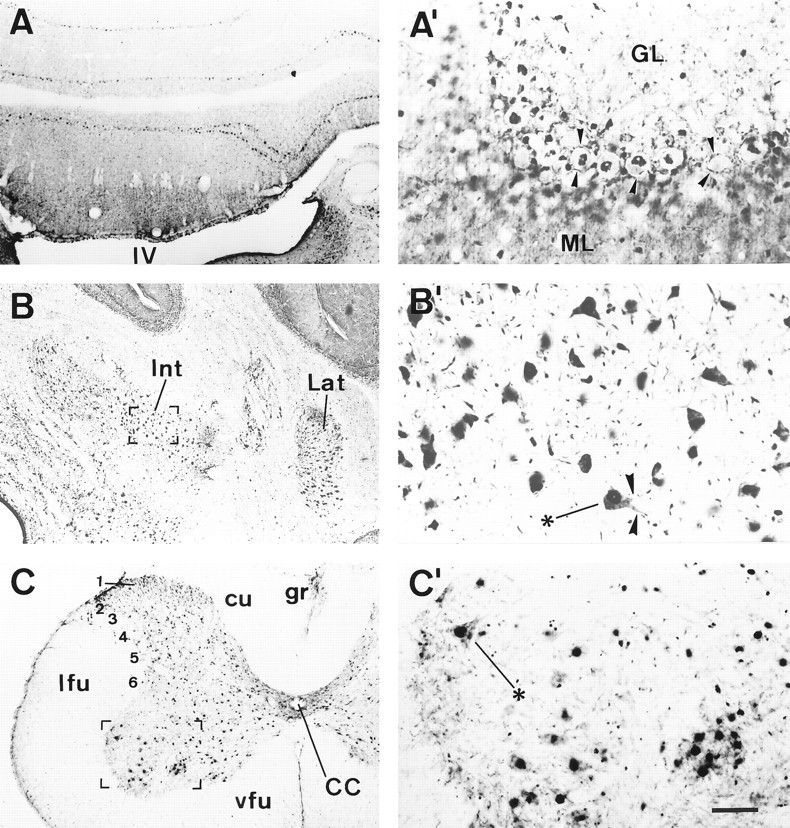
MCP-IR in the cerebellum and spinal cord.Left, Coronal sections showing MCP-IR in different structures at low magnification. The areas in bracketsin B and C are shown in B′and C′, respectively, shown at a higher magnification.A, Cerebellar cortex; note the dense MCP-IR displayed by the Purkinje cells. A′, Cerebellar cortex shown at a higher magnification. Several Purkinje cells displaying intense MCP-IR in the nucleus are shown in the center. Note the nonimmunoreactive nucleoli as well as the lightly labeled perikarya. The arrowheads point to immunoreactive structures outlining the Purkinje cell somata, and below them, another row of Purkinje cells appears out of focus. The molecular layer displays a moderate MCP-IR in which scattered MCP-positive cell nuclei are prominent. The granule cell layer is faintly immunoreactive, whereas intensely immunoreactive cells, likely Golgi cells, are scattered in this layer. B, Deep cerebellar nuclei;B′, higher magnification of neurons in the interposed cerebellar nucleus with intense MCP-IR in their perikarya and, in some cases, in the neuronal processes (arrowheads).C, Spinal cord; C′, neurons from the anterior horn shown at a higher magnification. Scale bar (shown inC′): A–C, 1 mm;A′–C′, 250 μm.
Spinal cord
The immunostaining pattern was similar to the one observed in the cerebral cortex. Thus, most neurons in all layers of the spinal cord were heavily stained (Fig. 5C). Within periependymal lamina 10, MCP-positive neuropil also was present. However, the motor neurons in the ventral horn displayed the densest MCP-IR, and their cell bodies were clearly delineated by means of MCP-IR within the perikarya and the proximal processes (Fig. 5C′).
White matter
MCP-IR also was detected in the white matter (for example, see Fig. 5B). Thus, most of the main fiber tracts in the brain displayed at least a light immunoreactivity. In addition, abundant MCP-positive cell bodies corresponding to glial cells were visible both within the white matter (Fig. 4E) and in the close vicinity of neuronal somata, within the gray matter (Figs.4B, 5C′).
Subcellular localization of the MCP complex in neurons
At the subcellular level, MCP-IR appeared primarily located within the nuclei of the neurons. In contrast, most neurons show only a slight immunostaining in the cytoplasm except in some neural structures containing rather large neurons. The perikarya of these large neurons displayed a moderate MCP-IR that appeared distributed homogeneously within the cytoplasm, in most cases, filling the proximal portions of the dendrites. However, in some cases, reaction product aggregates were visible within the perikaryon (Fig. 5B′,C′,asterisks). This subcellular distribution of the MCP complex in nervous tissue also was verified in primary cultures of neurons and glial cells by indirect immunofluorescence showing that MCP-IR is primarily located in the nuclei of both types of cells (data not shown). To confirm the findings observed under light microscopy and to refine the localization of MCP at the subcellular level, several brain areas were studied with immunoelectron microscopy. In full agreement with the observations at the light microscopic level, MCP-IR was primarily localized in the nuclei of the neurons from cortical, hippocampal, cerebellar, or spinal cord tissue; electron-dense DAB reaction product appeared densely clustered within the nucleoplasm, clearly delineating the nonimmunoreactive nucleoli (data not shown). Regarding the perikarya, much less reaction product was found consistently in the cytoplasm than in the nuclei of the neurons of the different brain areas examined. MCP-IR in the cytoplasm appeared in association with the endoplasmic reticulum or formed sparse clusters within the cytoplasmic matrix (Fig.6B,D); the Golgi apparatus and the mitochondria were devoid of labeling.
Fig. 6.
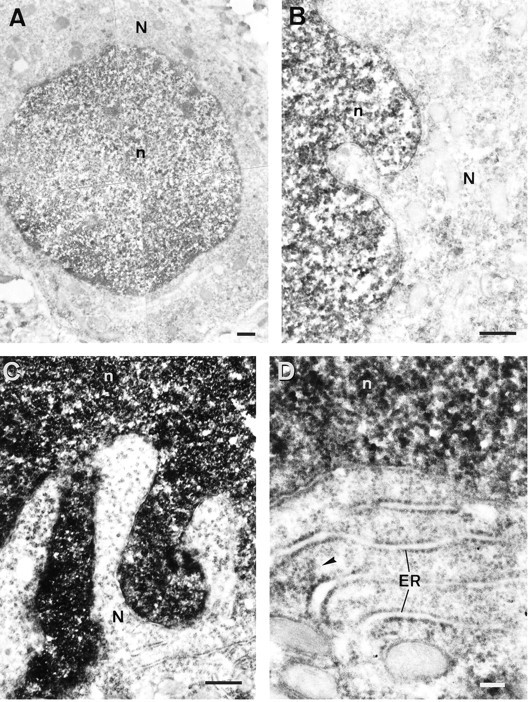
Electron micrographs showing the subcellular distribution of MCP-IR. A, Medium-size neuron from the parietal cortex displaying an intense MCP-IR in the nucleus and a light immunoreactivity in the cytoplasm. B, Detail of another cortical neuron showing the dense clusters of reaction product in the nucleus. The external nuclear membrane is almost free of labeling, whereas a light and heterogeneously distributed immunoreactivity is visible in the cytoplasm. Mitochondria in the perikarya appear devoid of MCP-IR. C, Purkinje cell from the cerebellum. Again, note the dense MCP-IR in the nucleus in contrast to the low immunoreactivity in the perikarya. D, Another Purkinje cell shown at a higher magnification. Note how MCP-IR in the cytoplasm is closely associated with the endoplasmic reticulum. Thearrowhead points to a cluster of reaction product between two cisternae of endoplasmic reticulum. Scale bars:A–C, 500 nm; D, 100 nm.
To study the subcellular localization in neuronal processes, we focused on the cerebellum because of its regularly arrayed structure. As described above, the Purkinje cells displayed an intense nuclear MCP-IR and a light immunoreactivity in the cytoplasm. In contrast, the apical dendrites of the Purkinje cells showed an intense MCP-IR that is clearly visible (in longitudinal sections) as they emerge from the Purkinje cells or traverse the molecular layer toward the pial surface. Also in the molecular layer, numerous MCP-positive neuronal processes were visible (Fig. 7A–C), probably corresponding to dendritic branches of the Purkinje cells. In some cases (Fig. 7A,B), nonimmunoreactive terminal boutons were making asymmetric synaptic contacts with these dendritic profiles. In the Purkinje cell layer, abundant MCP-immunoreactive terminal boutons, corresponding to nerve endings of the basket cells, were making symmetric axosomatic contacts with the Purkinje cell somata (Fig.7D,E). The granule cells showed low MCP-IR both at the light (see above) and electron microscopic levels. In contrast, presynaptic terminals with intense MCP-IR were observed in the granule cell layer. These MCP-positive large terminal boutons, filled with abundant mitochondria and making asymmetric synaptic contacts with nonreactive dentritic processes, were identified as mossy fiber terminals contacting granule cell dendrites (Fig.7F,G). Finally, myelinated axons within the granule cell layer also displayed an intense MCP-IR (Fig.7I). In addition, myelinated axons displaying MCP-IR frequently were observed in other brain areas, as illustrated in Figure7H by an axon from the cerebral cortex.
Fig. 7.
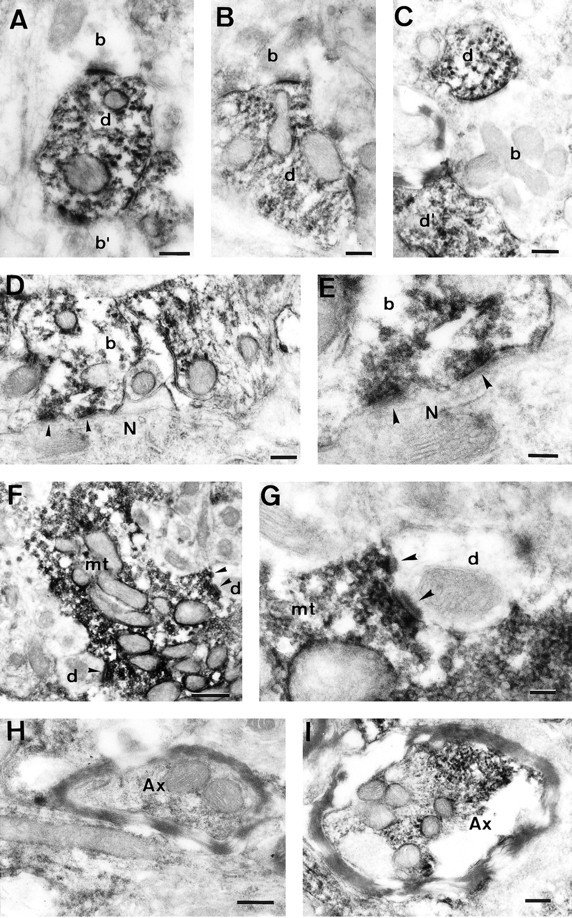
Electron micrographs showing MCP-IR within neuronal processes. A–C, MCP-IR in several dendritic processes from the molecular layer of the cerebellum. Nonimmunoreactive terminal bouton profiles (A, B) are making asymmetric synaptic contacts with the dendritic processes. C shows a nonimmunoreactive preterminal nerve fiber closely related to immunoreactive dendritic processes. D, MCP-IR in a basket cell terminal bouton making a symmetric synaptic contact with the soma of a Purkinje cell. The arrowheads mark the points of synaptic contact. E, The same synaptic contacts as in D, but at a higher magnification.F, MCP-IR in a mossy terminal bouton within the granule cell layer of the cerebellum making asymmetric synaptic contacts with several nonimmunoreactive dendritic processes. Thearrowheads mark the points of synaptic contact.G, Higher magnification of two of the synaptic contacts shown in F. H and Iillustrate MCP-IR in two myelinated axons from the cerebral cortex and the granule cell layer of the cerebellum, respectively. Scale bars:A–D, H, I, 200 nm; E, G, 100 nm; F, 500 nm.
DISCUSSION
A detailed study of the distribution of MCP-IR was carried out in the rat CNS using a polyclonal antibody against rat liver MCP. Our main findings are as follows. (1) MCP-IR is found ubiquitously in the rat CNS. (2) Despite this, MCP-IR is distributed heterogenously within the CNS. (3) MCP is localized primarily in the nuclei of neurons. These results, obtained at the light microscopic level, were confirmed by electron microscopic studies. The higher resolution of electron microscopy allowed us to define further the subcellular localization of MCP within neuronal somata and processes. MCP-IR was found in the cytoplasm, either free or in association with the endoplasmic reticulum. MCP-IR also was present in dendritic and axonic processes as well as in synaptic terminals.
As reported previously (Kamakura et al., 1988), MCP-IR is found ubiquitously along the rat CNS. In fact, the great majority of neurons in almost every structure of the encephalon and spinal cord displayed MCP-IR. However, MCP-IR at the subcellular level is localized primarily in the nuclei of neurons and glial cells, both in nervous tissue and in culture. This finding is in clear contrast to the previous report of Kamakura and colleagues (1988), who localized MCP-IR primarily in the cytoplasm of neurons, whereas only a faint staining was occasionally found within the nucleus. Nevertheless, our findings are in perfect agreement with the subcellular distribution of MCP in other tissues (Arizti el al., 1993; Rivett and Knecht, 1993). At least partially, this striking difference could be explained by the differences in the fixation procedures. The fixation solution used in the previous report was acetone at −20°C. However, during our study, we observed that in some cases, when the fixation had been carried out either with ethanol, methanol, or acetone at −20°C, MCP-IR was localized exclusively in the cytoplasm, even in cultured cells. Actually, we have observed invariably, by means of standard DNA stainings such as 4,6-diamidino-2-phenylindole (DAPI) or cresyl violet, that when our results showed MCP-IR only in the cytoplasm, the nuclear staining was either poor or negative. Therefore, DAPI staining (immunofluorescence) (Arizti et al., 1993) or cresyl violet staining (immunohistochemical procedure) always was included in our studies to check that the nucleus of neurons and glial cells were accessible. Similar results regarding the effect of the fixative have been reported recently (Machiels et al., 1995).
Another major finding of our study was that the distribution of MCP-IR in the rat CNS varied depending on the neural structures studied. The most intensely stained neurons were the pyramidal cortical neurons of layer 5 and the motor neurons of the ventral horn in the spinal cord. These cells displayed nuclear and cytoplasmatic MCP-IR and clearly stained processes. In addition, some populations of large neurons in the mesencephalon and brainstem also displayed a moderate MCP-IR in their perikarya, delineating the cell contours. Representative examples of this last group include the magnocellular portion of the red nucleus, the intermediate gray layer of the superior colliculus, the lateral vestibular nucleus, the deep cerebellar nuclei, the motor nuclei of the brainstem, and the reticular formation, especially its gigantocellular part. It is important to note, however, that the vast majority of neurons in the remaining structures showed only a slight cytoplasmatic MCP-IR, but their nuclei displayed an intense MCP-IR. Prominent examples of this group were the cortical neurons of layers 2–4 and layer 6, the piriform cortex and the olfactory tubercle, the amygdalar complex, the hippocampal formation, the nucleus of the bed stria terminalis, the paraventricular thalamic nucleus, and the Purkinje cells in the cerebellum.
A relevant point is the comparison of the distribution and subcellular localization for MCP and for calpain, the other major nonlysosomal protease of the cell, in the rat CNS. Calpain I or μ-calpain (Hamakubo et al., 1986; Perlmutter et al., 1990) as well as MCP also is distributed ubiquitously in the rat CNS, but unlike MCP, it is localized in the cytoplasm and neural processes, whereas the cell nucleus is devoid of calpain I. However, in spite of this, the regional distribution of MCP-IR coincides with the distribution of calpain I in the different structures of the rat CNS (Hamakubo et al., 1986;Perlmutter et al., 1990). In contrast, the distribution of MCP-IR differs greatly from the distribution of calpain II (Hamakubo et al., 1986). This is particularly clear in the hippocampus, in which μ-calpain is present in almost all pyramidal and granule cells as well as in some other neurons of the archicortex (Fukuda et al., 1990). Calpain has been implicated in the damage of neurons induced by ischemia. The activation of calpain by ischemia-induced calcium release and its contribution to neuronal cell damage have been based on studies demonstrating the protective role of certain proteolytic inhibitors on postischemic neuronal death, as confirmed by the presence of normal long-term potentiation (LTP) in those hippocampal neurons after the ischemic procedure (Lee et al., 1991). These studies have used leupeptin and the specific calpain inhibitor IN-acetyl-leu-leu-nor-leucinal. Interestingly, both of these inhibitors also are potent inhibitors of the MCP activity (Orlowski, 1990; Rivett, 1993), and the doses of calpain inhibitor I used in those studies clearly will suppress both MCP and calpain activity (Figueiredo-Pereira et al., 1994; Rock et al., 1994). Therefore, MCP proteolytic activity may be relevant in pathological damage of neurons during hypoxia.
Finally, the present demonstration of the localization of the proteasome in synaptic boutons and in the neuronal cell nucleus might have additional functional significance. The proteasome has been found by immunoblot in synaptosomes and other subcellular fractions fromAplysia neurons (Chain et al., 1995), and ubiquitin-dependent degradation of the regulatory subunit of the cAMP-dependent protein kinase was correlated with long-term presynaptic facilitation in this system (Hegde et al., 1993). Those results, together with the results presented in this study, and the fact that we also found MCP by immunoblot analysis in synaptosomal preparations from rat brain strongly support the hypothesis that the proteasome may participate in the mechanism of maintenance of LTP. In a similar context, it has been described recently in Aplysia,Drosophila, and mice that the cAMP-dependent CREB family of transcription factors is involved in LTP as a consequence of the activation of the cAMP-dependent protein kinase (for review, see Frank and Greenberg, 1994). Therefore, LTP appears to be a neurobiological process dependent on a sophisticated network of short- and long-term changes in transcription factors. Thus, we may hypothesize that the proteasome, which clearly is involved in the degradation of some inducible transcription factors such as NFKB/IKB (Palombella et al., 1994; Chen et al., 1995), fos (Tsurumi et al., 1995), jun (Treier et al., 1994; Jariel-Encontre et al., 1995), and p53 (Scheffner et al., 1990; Ciechanover et al., 1994), also might be implicated in the degradation of these transcription factors modulating LTP, via ubiquitin-dependent or ubiquitin-independent pathways.
Footnotes
This work was supported by Fundación Ramón Areces and Comisión Interministerial de Ciencia y Tecnología (SAF94-0685) to J.G.C. and Fondo de Investigaciones Sanitarias de la Seguridad Social (FIS; FIS 93/0337, FIS 96/0488) to J.M.G.-A. E.M. is the recipient of a postdoctoral fellowship supported by FIS 93/0337, and P.A. was supported by a grant from Comunidad Autónoma de Madrid. We thank Dr. R. Martinez-Murillo for his help and comments, E. Sánchez for technical assistance, and A. Fernández for the photographic work on this paper.
Correspondence should be addressed to Dr. José G. Castaño, Departamento de Bioquímica e Instituto de Investigaciones Biomédicas, Facultad de Medicina, Universidad Autónoma de Madrid, 28029 Madrid, Spain.
REFERENCES
- 1.Arribas J, Arizti P, Castaño JG. Antibodies against the C2 COOH-terminal region discriminate the active and latent forms of the multicatalytic proteinase complex. J Biol Chem. 1994;269:12858–12864. [PubMed] [Google Scholar]
- 2.Arribas J, Castaño JG. Kinetic studies of the differential effect of detergents on the peptidase activities of the multicatalytic proteinase from rat liver. J Biol Chem. 1990;265:13969–13973. [PubMed] [Google Scholar]
- 3.Arribas J, Luz-Rodriguez M, Alvarez-Do-Forno R, Castaño JG. Autoantibodies against the multicatalytic proteinase in patients with systemic lupus erythematosus. J Exp Med. 1991;173:423–427. doi: 10.1084/jem.173.2.423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Arizti P, Arribas J, Castaño J G. Modulation of the multicatalytic proteinase complex by lipids, interconversion and proteolytic processing. Enzyme Protein. 1993;47:285–295. doi: 10.1159/000468686. [DOI] [PubMed] [Google Scholar]
- 5.Chain DG, Hegde AN, Yamamoto N, Liu-Marsh B, Schwartz JH. Persistent activation of cAMP dependent protein kinase by regulated proteolysis suggests a neuron-specific function of the ubiquitin system in Aplysia. J Neurosci. 1995;15:7592–7603. doi: 10.1523/JNEUROSCI.15-11-07592.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chen Z, Hagler J, Palombella VJ, Melandri F, Scherer D, Ballard D, Maniatis T. Signal-induced site-specific phosphorylation targets I kappa B alpha to the ubiquitin-proteasome pathway. Genes Dev. 1995;9:1586–1597. doi: 10.1101/gad.9.13.1586. [DOI] [PubMed] [Google Scholar]
- 7.Ciechanover A. The ubiquitin-proteasome proteolytic pathway. Cell. 1994;79:13–21. doi: 10.1016/0092-8674(94)90396-4. [DOI] [PubMed] [Google Scholar]
- 8.Ciechanover A, Shkedy D, Oren M, Bercovich Degradation of the tumor suppressor protein p53 by the ubiquitin-mediated proteolytic system requires a novel species of ubiquitin-carrier protein, E2. J Biol Chem. 1994;269:9582–9589. [PubMed] [Google Scholar]
- 9.Fenteany G, Standaert RF, Lane WS, Choi S, Corey EJ, Schreiber SL. Inhibition of proteasome activities and subunit-specific amino-terminal threonine modification by lactacystin. Science. 1995;268:726–731. doi: 10.1126/science.7732382. [DOI] [PubMed] [Google Scholar]
- 10.Fenteany G, Standaert RF, Reichard GA, Corey EJ, Schreiber SL. A beta-lactone related to lactacystin induces neurite outgrowth in a neuroblastoma cell line and inhibits cell cycle progression in an osteosarcoma cell line. Proc Natl Acad Sci USA. 1994;91:3358–3362. doi: 10.1073/pnas.91.8.3358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Figueiredo-Pereira ME, Banik N, Wilk S. Comparison of the effect of calpain inhibitors on two extralysosomal proteinases: the multicatalytic proteinase complex and m-calpain. J Neurochem. 1994;62:1989–1994. doi: 10.1046/j.1471-4159.1994.62051989.x. [DOI] [PubMed] [Google Scholar]
- 12.Fukuda T, Adachi E, Kawashima S, Yoshiya I, Hashimoto PH. Immunohistochemical distribution of calcium-activated neutral proteinases and endogeneous CANP inhibitor in the rabbit hippocampus. J Comp Neurol. 1990;302:100–109. doi: 10.1002/cne.903020108. [DOI] [PubMed] [Google Scholar]
- 13.Frank DA, Greenberg ME. CREB: a mediator of long-term memory from mollusks to mammals. Cell. 1994;79:5–8. doi: 10.1016/0092-8674(94)90394-8. [DOI] [PubMed] [Google Scholar]
- 14.Gaczynska M, Rock KL, Goldberg AL. Role of proteasomes in antigen presentation. Enzyme Protein. 1993;47:354–369. doi: 10.1159/000468693. [DOI] [PubMed] [Google Scholar]
- 15.Geneser-Jensen FA, Blackstad TW. Distribution of acetyl cholinesterase in the hippocampal region of the guinea pig. I. Entorhinal area, parasubiculum and presubiculum. Z Zellforsch Mikrosk Anat. 1971;114:460–481. doi: 10.1007/BF00325634. [DOI] [PubMed] [Google Scholar]
- 16.Hamakubo T, Kannagi R, Murachi T, Matus A. Distribution of calpains I and II in rat brain. J Neurosci. 1986;6:3103–3111. doi: 10.1523/JNEUROSCI.06-11-03103.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hegde AN, Goldberg A, Schwartz JH. Regulatory subunits of cAMP-dependent protein kinase are degraded after conjugation to ubiquitin: a molecular mechanism underlying long-term synaptic plasticity. Proc Natl Acad Sci USA. 1993;90:7436–7440. doi: 10.1073/pnas.90.16.7436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Heinemeyer W, Trondle N, Albrecht G, Wolf DH. PRE5 and PRE6, the last missing genes encoding 20S proteasome subunits from yeast? Indication for a set of 14 different subunits in the eukaryotic proteasome core. Biochemistry. 1994;33:12229–12237. doi: 10.1021/bi00206a028. [DOI] [PubMed] [Google Scholar]
- 19.Hochstrasser M. Ubiquitin, proteasomes, and the regulation of intracellular protein degradation. Curr Opin Cell Biol. 1995;7:215–223. doi: 10.1016/0955-0674(95)80031-x. [DOI] [PubMed] [Google Scholar]
- 20.Jariel-Encontre I, Pariat M, Martin F, Carillo S, Salvat C, Piechaczyk M. Ubiquitinylation is not an absolute requirement for degradation of c-Jun protein by the 26 S proteasome. J Biol Chem. 1995;270:11623–11627. doi: 10.1074/jbc.270.19.11623. [DOI] [PubMed] [Google Scholar]
- 21.Kamakura K, Ishiura S, Nonaka I, Sugita H. Localization of ingensin in rat central nervous system and skeletal muscle. J Neurosci Res. 1988;20:473–478. doi: 10.1002/jnr.490200410. [DOI] [PubMed] [Google Scholar]
- 22.Katagiri M, Hayashi M, Matsuzaki K, Tanaka H, Omura S. The neuritogenesis inducer lactacystin arrests cell cycle at both G0/G1 and G2 phases in neuro 2a cells. J Antibiot (Tokyo) 1995;48:344–346. doi: 10.7164/antibiotics.48.344. [DOI] [PubMed] [Google Scholar]
- 23.Kwak S, Masaki T, Ishiura S, Sugita H. Multicatalytic proteinase is present in Lewy bodies and neurofibrillary tangles in diffuse Lewy body disease brains. Neurosci Lett. 1991;128:21–24. doi: 10.1016/0304-3940(91)90751-e. [DOI] [PubMed] [Google Scholar]
- 24.Lee KS, Frank S, Vanderklish P, Arai A, Lynch G. Inhibition of proteolysis protects hippocampal neurons from ischemia. Proc Natl Acad Sci USA. 1991;88:7233–7237. doi: 10.1073/pnas.88.16.7233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lowe J, Stock D, Jap B, Zwickl P, Baumeister W, Huber R. Crystal structure of the 20S proteasome from the archaeon T. acidophilum at 3.4 A resolution [see comments]. Science. 1995;268:533–539. doi: 10.1126/science.7725097. [DOI] [PubMed] [Google Scholar]
- 26.Machiels BM, Henfling ME, Broers JL, Hendil KB, Ramekers FC. Changes in immunocytochemical detectability of proteasome epitopes depending on cell growth and fixation conditions of lung cancer cell lines. Eur J Cell Biol. 1995;66:282–292. [PubMed] [Google Scholar]
- 27.Masaki T, Ishiura S, Sugita H, Kwak S. Multicatalytic proteinase is associated with characteristic oval structures in cortical Lewy bodies: an immunocytochemical study with light and electron microscopy. J Neurol Sci. 1994;122:127–134. doi: 10.1016/0022-510x(94)90288-7. [DOI] [PubMed] [Google Scholar]
- 28.Olink-Coux M, Arcangeletti C, Pinardi F, Minisini R, Huesca M, Chezzi C, Scherrer K. Cytolocation of prosome antigens on intermediate filament subnetworks of cytokeratin, vimentin and desmin type. J Cell Sci. 1994;107:353–366. [PubMed] [Google Scholar]
- 29.Orlowski M. The multicatalytic proteinase complex, a major extralysosomal proteolytic system. Biochemistry. 1990;29:10289–10297. doi: 10.1021/bi00497a001. [DOI] [PubMed] [Google Scholar]
- 30.Palombella VJ, Rando OJ, Goldberg AL, Maniatis T. The ubiquitin-proteasome pathway is required for processing the NFkappa B1 precursor protein and the activation of NF-kappa B. Cell. 1994;78:773–785. doi: 10.1016/s0092-8674(94)90482-0. [DOI] [PubMed] [Google Scholar]
- 31.Paxinos G, Watson C. Academic; New York: 1986. The rat brain in stereotaxic coordinates. . [DOI] [PubMed] [Google Scholar]
- 32.Perlmutter LS, Gall C, Baudry M, Lynch G. Distribution of calcium-activated protease calpain in the rat brain. J Comp Neurol. 1990;296:269–276. doi: 10.1002/cne.902960207. [DOI] [PubMed] [Google Scholar]
- 33.Rivett JA. Proteasomes: multicatalytic proteinase complexes. Biochem J. 1993;291:1–10. doi: 10.1042/bj2910001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Rivett JA, Knecht E. Proteasome location. Curr Biol. 1993;3:127–129. doi: 10.1016/0960-9822(93)90173-l. [DOI] [PubMed] [Google Scholar]
- 35.Rock KL, Gramm C, Rothstein L, Clark K, Stein R, Dick L, Hwang D, Goldberg AL. Inhibitors of the proteasome block the degradation of most cell proteins and the generation of peptides presented on MHC class I molecules. Cell. 1994;78:761–771. doi: 10.1016/s0092-8674(94)90462-6. [DOI] [PubMed] [Google Scholar]
- 36.Scheffner M, Werness BA, Huibregtse JM, Levine AJ, Howley PM. The E6 oncoprotein encoded by human papillomavirus types 16 and 18 promotes the degradation of p53. Cell. 1990;63:1129–1136. doi: 10.1016/0092-8674(90)90409-8. [DOI] [PubMed] [Google Scholar]
- 37.Tamura T, Nagy I, Lupas A, Lottspeich F, Schoofs G, Tanaka K, De Mot R, Baumeister W. The first characterization of a eubacterial proteasome: the 20S complex of Rhodococcus. Curr Biol. 1995;5:766–774. doi: 10.1016/s0960-9822(95)00153-9. [DOI] [PubMed] [Google Scholar]
- 38.Tanaka K, Tamura T, Yoshimura T, Ichihara A. Proteasomes: protein and gene structures. New Biol. 1992;4:173–187. [PubMed] [Google Scholar]
- 39.Treier M, Staszewski LM, Bohmann D. Ubiquitin-dependent c-Jun degradation in vivo is mediated by the delta domain. Cell. 1994;78:787–798. doi: 10.1016/s0092-8674(94)90502-9. [DOI] [PubMed] [Google Scholar]
- 40.Tsurumi C, Ishida N, Tamura T, Kakizuka A, Nishida E, Okumura E, Kishimoto T, Inagaki M, Okazaki K, Sagata N, Ichihara A, Tanaka K. Degradation of c-Fos by the 26S proteasome is accelerated by c-Jun and multiple protein kinases. Mol Cell Biol. 1995;15:5682–5687. doi: 10.1128/mcb.15.10.5682. [DOI] [PMC free article] [PubMed] [Google Scholar]


