Abstract
Background:
Thoracic aortic dissection is an emergent life-threatening condition. Routine screening for genetic variants causing thoracic aortic dissection is not currently performed for patients or family members.
Methods:
We performed whole exome sequencing of 240 patients with thoracic aortic dissection (n=235) or rupture (n=5) and 258 controls matched for age, sex, and ancestry. Blinded to case-control status, we annotated variants in 11 genes for pathogenicity.
Results:
Twenty-four pathogenic variants in 6 genes (COL3A1, FBN1, LOX, PRKG1, SMAD3, TGFBR2) were identified in 26 individuals, representing 10.8% of aortic cases and 0% of controls. Among dissection cases, we compared those with pathogenic variants to those without and found that pathogenic variant carriers had significantly earlier onset of dissection (41 vs. 57 years), higher rates of root aneurysm (54% vs. 30%), less hypertension (15% vs. 57%), lower rates of smoking (19% vs. 45%), and greater incidence of aortic disease in family members. Multivariable logistic regression showed that pathogenic variant carrier status was significantly associated with age <50 [odds ratio (OR) = 5.5; 95% CI: 1.6-19.7], no history of hypertension (OR=5.6; 95% CI: 1.4-22.3) and family history of aortic disease (mother: OR=5.7; 95% CI: 1.4-22.3, siblings: OR=5.1; 95% CI 1.1-23.9, children: OR=6.0; 95% CI: 1.4-26.7).
Conclusions:
Clinical genetic testing of known hereditary thoracic aortic dissection genes should be considered in patients with a thoracic aortic dissection, followed by cascade screening of family members, especially in patients with age-of-onset <50 years, family history of thoracic aortic disease, and no history of hypertension.
Journal Subject Terms: Aortic Dissection, Genetics
Keywords: aortic dissection, rupture, genetics, diagnostics, pathogenic variants, gene sequencing
Introduction
Thoracic aortic dissection is a life-threatening condition, responsible for 15,000 deaths a year in the United States.1, 2 Approximately 30% of patients presenting with a thoracic aortic aneurysm and dissection have an underlying genetic predisposition,3 which can be associated with syndromic features, such as Marfan syndrome or Loeys-Dietz syndrome, or not associated with syndromic features, as with ACTA2, MYLK, and MYH11 mutations.4 Variants in many genes, including FBN1, SMAD3, and ACTA2, among others, can lead to either syndromic or non-syndromic thoracic aortic aneurysm and dissection.4-6 Recent advances in the field have shown definitive and strong evidence to support the role of pathogenic variants in ACTA2, COL3A1, FBN1, MYH11, SMAD3, TGFB2, TGFBR1, TGFBR2, MYLK, LOX, and PRKG1 as predisposing to hereditary thoracic aortic disease.7
These genetic findings play a critical role for the patient and family members, helping to guide clinical decision-making to prevent or lessen the likelihood of a catastrophic event. Aortic diameter is a central criterion when deciding prophylactic surgical intervention, and the recommended aortic diameter for surgical intervention differs for those with and without an underlying genetic predisposition. The American Heart Association/American College of Cardiology (AHA/ACC) guidelines8 recommend that patients with genetically mediated aneurysms undergo elective surgical repair at an ascending or aortic root diameter of 4.0–5.0 cm, depending on the condition. Whereas patients without a known genetic mutation may undergo elective surgical repair when the ascending or aortic root diameter is ≥ 5.5 cm, there are also established risk factors, such as an aortic diameter growth rate between > 3 to 5mm/year8, 9 that may drive early surgical intervention. Recent work shows that different genes predisposing to hereditary thoracic aortic dissection have varying presentations and courses.10, 11 For instance, patients with ACTA2 mutations more often present with acute aortic dissections whereas patients with Marfan syndrome often present with skeletal and ocular features before thoracic aortic dilation is discovered.12
Despite the potential clinical impact of genetic findings, clinicians are usually not aware that a patient has an underlying pathogenic variant upon initial presentation with a thoracic aortic dissection. The identification of variants known to predispose to thoracic aortic dissection has the potential to improve clinical management and guide treatment strategies for patients and family members. The objective of this study was to evaluate trends in pathogenic variants carriers with a history of thoracic aortic dissection or thoracic aortic rupture, as well as to identify which patients and corresponding family members may benefit from clinic genetic testing. We examined all genes in the genome and none reached exome-wide significance for single variant tests or gene-based burden tests.
Methods
The full methods for this manuscript are available as supplemental material. In accordancy with the Transparency and Openness Promotion (TOP) guidelines the data that support the findings of this study are available from the corresponding author (CJW) upon reasonable request and approval from the institutional internal review board. The study was approved by the institutional review board at the University of Michigan and all subjects provided informed consent.
Results
Annotation of variants from research-level whole exome sequencing identifies pathogenic variants
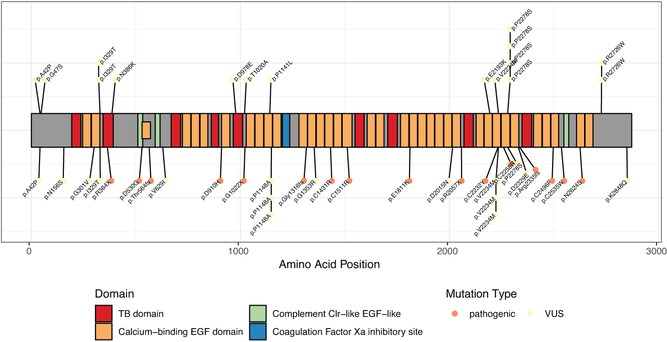
A total of 240 cases with a clinical diagnosis of thoracic aortic dissection (type A or type B) or rupture with or without aortic aneurysm and 258 age-, sex-, and ancestry-matched controls had whole exome sequences available following quality control (see Methods for quality control failures). For the 498 samples passing quality control, 248 variants were annotated blind to the variant carrier’s case or control status. 24 pathogenic variants in 6 genes (COL3A1, FBN1, LOX, PRKG1, SMAD3, TGFBR2) were identified, found exclusively in 26 cases (Table 1), representing 10.8% of cases and 0% of controls. Two variants were seen twice in cases who were first degree relatives. There is a significant burden of pathogenic variants in FBN1 in cases compared to controls (Ncases=18, Ncontrols=0; p-value=2.5×10−5, Supplementary Table 1). These variants are predominantly found in calcium-binding epidermal growth factor domains of FBN1 (Figure 1). We examined the proportion of pathogenic variants that were present in commonly used databases and found that of the 24, 11 were present in dbSNP13, 8 were listed as pathogenic in ClinVar14, and 2 were present in gnomAD15 (see Supplementary Note).
Table 1.
Classification of 24 Pathogenic Variants
| Chromosome:position | Reference Allele | Alternate Allele |
Mutation type | Gene | HGVS protein notation |
ClinVar 9/30/18 | rsID dbSNP 151 |
|---|---|---|---|---|---|---|---|
| 2:189858169 | G | A | Nonsynonymous | COL3A1 | p.G378D | NA | |
| 3:30732950 | G | A | Stop Gain | TGFBR2 | p.W521* | VUS for non-aortic phenotype | |
| 5:121412592 | CCAGA | C | Frameshift | LOX | p.Cys244fs | NA | rs779512296† |
| 10:53227579 | G | A | Nonsynonymous | PRKG1 | p.R177Q | Pathogenic | rs397515330 |
| 15:48707913 | T | C | Nonsynonymous | FBN1 | p.N2624S | VUS | |
| 15:48713849 | G | C | Nonsynonymous | FBN1 | p.C2535W | Pathogenic | |
| 15:48714232 | C | A | Nonsynonymous | FBN1 | p.C2496F | Likely pathogenic | |
| 15:48719947 | TGAAGCAGTACCCTTCCC | T | Frameshift | FBN1 | p.R2335fs | NA | |
| 15:48722967 | A | G | Nonsynonymous | FBN1 | p.C2258R | Pathogenic | rs1057520617 |
| 15:48725107 | C | T | Nonsynonymous | FBN1 | p.C2232Y | Pathogenic | rs1060501054 |
| 15:48730109 | G | A | Stop Gain | FBN1 | p.R2057* | Pathogenic | rs763091520 |
| 15:48744873 | C | T | Nonsynonymous | FBN1 | p.E1811K | Conflicting interpretation of pathogenicity | rs761857514† |
| 15:48760660 | A | G | Nonsynonymous | FBN1 | p.C1511R | Likely pathogenic | rs397515811 |
| 15:48764793 | A | G | Nonsynonymous | FBN1 | p.C1431R | NA | |
| 15:48773870 | C | CT | Frameshift | FBN1 | p.G1316fs | Likely pathogenic | |
| 15:48782066 | C | A | Stop Gain | FBN1 | p.G1022* | NA | rs794728171 |
| 15:48786401 | C | G | Nonsynonymous | FBN1 | p.D910H | NA | |
| 15:48802264 | G | GT | Frameshift | FBN1 | p.Thr564fs | Likely pathogenic | |
| 15:48802366 | T | C | Nonsynonymous | FBN1 | p.D530G | VUS | |
| 15:48808561 | T | C | Essential Splice Site | FBN1 | . | Pathogenic | rs397515756 |
| 15:48812913 | G | A | Stop Gain | FBN1 | p.R364* | Pathogenic | rs794728165 |
| 15:48888576 | C | T | Essential Splice Site | FBN1 | . | Likely pathogenic | |
| 15:67457370 | TGAA | T | In frame deletion | SMAD3 | p.K116del | NA | |
| 15:67462935 | TA | T | Frameshift | SMAD3 | p.Asn218fs | Pathogenic | rs587776881 |
Also present in gnomAD version 2.1
Figure 1.
Distribution of pathogenic and variants of unknown significance in Fibrillin 1 (FBN1). Each point is a sample, with controls above the protein diagram and cases below.
Validation targeted sequencing
Molecular Inversion Probe Sequencing was utilized to ensure the highest level of confidence in the whole exome sequencing variant calls and to protect against potential sample swaps. The carrier status of pathogenic variants identified through whole exome sequencing was verified by Molecular Inversion Probe Sequencing in all 26 samples (Supplementary Table 2).
Research-level whole exome sequencing and implications for precision health
For 17 of the 26 pathogenic variant carriers (hereon pathogenic carriers), the whole exome sequencing results aligned with the current clinical diagnoses in the electronic medical record, including 5 patients (5/17) in which clinical genetic testing previously identified the same pathogenic variant as whole exome sequencing (Table 2 and Supplementary Table 3). Whole exome sequencing results provided validation for 12 pathogenic carriers with a clinical diagnosis of Marfan syndrome based on the Revised Ghent Nosology.16 There were no genetic testing results for the above 12 patients other than the whole exome sequencing results from this study. For the 9 remaining pathogenic carriers, whole exome sequencing and annotation of pathogenic variants added diagnostic precision to the clinical diagnosis (Table 2). Specifically, 8 of these pathogenic carriers (8/9) lacked a specific clinical diagnosis, but whole exome sequencing and history of thoracic aortic dissection shifted the clinical diagnosis per guidelines to Marfan syndrome16 (FBN1, n=4), vascular Ehlers-Danlos syndrome8 (COL3A1, n=1), or familial thoracic aortic disease (LOX, PRKG1, and SMAD3, n=3). For 1 pathogenic carrier (1/9) there was an incorrect diagnosis of Marfan syndrome, which was amended to Loeys-Dietz syndrome based on a pathogenic variant identified in TGFBR2 and history of an acute Type A aortic dissection. In addition, the whole exome sequencing results provide a basis for cascade screening for the family members of all 26 cases per AHA guidelines.8
Table 2.
Comparison Between Clinical Diagnosis and Pathogenic Variants Identified With Whole Exome Sequencing
| Clinical Diagnosis Matched |
Clinical Diagnosis Changed |
Diagnostic improvement and implications for clinical care |
||||
|---|---|---|---|---|---|---|
| Number of Variants |
Genes | Number of Variants |
Genes | Number of Variants |
Genes | |
| Clinical genetic testing previously performed | 5 | FBN1*, SMAD3*, PRKG1* | 0 | 0 | 0 | - |
| No prior clinical genetic testing | 12 | FBN1† | 1 | TGFBR2‡ | 8 | FBN1§, SMAD3§, LOX§, COL3A1§ |
Clinical diagnois and clinical genetic testing were consistent with the whole exome sequencing results
Clinical diagnosis based on the Revised Ghent Nosology without clinical genetic testing was consistent with whole exome sequencing results.
Clinical diagnosis without clinical genetic testing was inconsistent with whole exome sequencing results
Clinical diagnosis without clinical genetic testing would be improved by the whole exome result.
Variants of unknown significance
86 of the 248 annotated variants in aortopathy genes were annotated as VUS. After excluding one of each first degree relative pair (see Methods) and cases with pathogenic variants, 58 of 213 cases (27.2%) and 51 of 258 controls (19.8%) had at least one VUS identified from whole exome sequencing, which was not significant (p-value=0.072, Supplementary Table 4). There is, however, a significant association between pathogenic variants and cases (p-value=2.8×10−7, Supplementary Table 4). None of the 11 genes demonstrated association between carrier status for VUS and thoracic aortic dissection or rupture case/control status (Supplementary Table 1).
Clinical characteristics between pathogenic variant and non-pathogenic variant carriers
The pathogenic carriers were significantly younger with a median of 41 years (age range 18–61 years) versus 57 years (age range 17–89 years) of age. 77% of pathogenic carriers were < 50 years old while 72% of non-pathogenic carriers were >50 years old. Pathogenic carriers also had significantly more root aneurysms (54% vs. 30%), less hypertension (15% vs. 57%), and less history of smoking (19% vs. 45%) compared to the non-pathogenic carriers. Moreover, the pathogenic carriers had a greater incidence of thoracic aortic disease in parents, siblings, and children (all p-values<0.05) (Table 3). Pathogenic carriers presented with more type A than type B dissections although this comparison was not significant (69.2% vs. 58.9%, p=0.421). One pathogenic carrier had a bicuspid aortic valve compared to 17 non-pathogenic variant carriers with biscuspid aortic valves. Multivariable logistic regression showed that pathogenic carriers were significantly more likely to have dissection age < 50 years old, family history of thoracic aortic disease, and no history of hypertension (Table 4).
Table 3.
Demographic and Clinical Characteristics at the Time of Dissection
| Variables | All Patients N= 240 |
Non-Pathogenic N=214 |
Pathogenic N=26 |
P |
|---|---|---|---|---|
| Age of onset, years | 56 (45, 66) | 57 (47, 67) | 38 (26, 48) | <.001 |
| Age of dissection, years | 56 (45, 67) | 57 (47, 67) | 41 (29, 50) | <.001 |
| Male | 159 (66) | 146 (68) | 13 (50) | 0.102 |
| Race (% Caucasian) | 212 (88) | 190 (89) | 22 (85) | 0.76 |
| Ethnicity (% non-Hispanic) | 224 (93) | 198 (93) | 26 (100) | 0.30 |
| Thoracic aortic indications | ||||
| Root aneurysm | 78 (33) | 64 (30) | 14 (54) | 0.025 |
| Ascending aneurysm | 119 (50) | 107 (50) | 12 (46) | 0.87 |
| Arch aneurysm | 59 (25) | 55 (26) | 4 (15) | 0.34 |
| Descending aneurysm | 71 (30) | 66 (31) | 5 (19) | 0.32 |
| Max aneurysmal diameter, mm | 48 (42, 57) | 47 (42, 55) | 57 (48, 71) | 0.03 |
| Type A aortic dissection | 144 (60) | 126 (59) | 18 (69) | 0.42 |
| Type B aortic dissection | 91 (38) | 84 (39) | 7 (27) | 0.31 |
| Rupture | 5 (2.1) | 4 (1.9) | 1 (3.8) | 0.441 |
| Risk Factors | ||||
| HTN | 126 (53) | 122 (57) | 4 (15) | <.001 |
| Dyslipidemia | 42 (18) | 40 (19) | 2 (7.7) | 0.27 |
| Smoking history (former/current) | 102 (43) | 97 (45) | 5 (19) | 0.02 |
| Type 2 diabetes mellitus | 6 (2.5) | 6 (2.8) | 0 (0) | 1.00 |
| Medications | ||||
| ACE-I | 29 (12) | 27 (13) | 2 (7.7) | 0.75 |
| Calcium channel blocker | 11 (4.6) | 11 (5.1) | 0 (0) | 0.61 |
| ARB | 14 (5.8) | 13 (6.1) | 1 (3.8) | 1.00 |
| Βeta-Blocker | 68 (28) | 62 (29) | 6 (23) | 0.69 |
| Anti-HTN medications (% yes) | 83 (35) | 77 (36) | 6 (23) | 0.28 |
| Number of HTN medications | 0.40 | |||
| 0 | 157 (65) | 137 (64) | 20 (77) | |
| 1 | 50 (21) | 46 (21) | 4 (15) | |
| 2 | 27 (11) | 26 (12) | 1 (3.8) | |
| 3 | 6 (2.5) | 5 (2.3) | 1 (3.8) | |
| Family history, first-degree relative | ||||
| Mother | 41 (17) | 31 (15) | 10 (50) | 0.008 |
| Father | 47 (20) | 39 (18) | 8 (31) | 0.22 |
| Sibling, at least one known | 42 (18) | 30 (14) | 12 (46) | <.001 |
| Child, at least one known | 18 (7.5) | 10 (5) | 8 (31) | <.001 |
| CLIA genetic testing (% yes) | 20 (8.0) | 15 (7.0) | 5 (19.2) | 0.05 |
| Pathogenic variant | 5 (2.0) | 0 (0) | 5 (19.2) | <.001 |
| Likely pathogenic or VUS | 8 (3.8) | 8 (3.8) | 0 (0) | 0.604 |
| No variant identified | 7 (3.3) | 7 (3.3) | 0 (0) | 1.0 |
Values are median (IQR) or n (%).
Correction for multiple statistical tests was not performed.
ACE-I=angiotensin converting enzyme inhibitor; ARB=Angiotensin II receptor blocker; CLIA: Clinical Laboratory Improvement Amendments; HTN=hypertension
Table 4.
Risk factors for cases with a pathogenic variant
| Variables | OR | 95% Wald Confidence Limits | P-value | |
|---|---|---|---|---|
| Lower | Upper | |||
| Age ≤ 50 vs > 50 | 5.5 | 1.6 | 19.7 | 0.008 |
| Sex (female vs male) | 1.1 | 0.3 | 3.8 | 0.84 |
| Caucasian | 0.7 | 0.1 | 3.1 | 0.60 |
| Root aneurysm | 1.7 | 0.6 | 5.2 | 0.34 |
| Hypertension | 5.6 | 1.4 | 22.3 | 0.015 |
| Smoking history | 2.6 | 0.7 | 9.9 | 0.16 |
| Family history | ||||
| Mother | 5.7 | 1.4 | 22.3 | 0.013 |
| Father | 0.3 | 0.1 | 1.6 | 0.17 |
| Siblings | 5.1 | 1.1 | 23.9 | 0.04 |
| Children | 6.0 | 1.4 | 26.7 | 0.017 |
Hypertension was defined as no hypertension versus had a diagnosis of hypertension. Smoking history was defined as no smoking history versus had a smoking history. Family history was defined as aortic disease noted within a first-degree relative.
Discussion
The current study reports our initial experience with research-level whole exome sequencing in patients with thoracic aortic dissection or rupture with or without aneurysm. We tested 240 cases and 258 controls for pathogenic variants in 11 genes known to cause aortic dissection.7 By whole exome sequencing and validation targeted sequencing, we found pathogenic variants in 10.8% of cases and 0% of controls. 58 (27.2%) cases and 51 (19.8%) controls were identified as carriers of variants of unknown significance.
In the general population, the incidence of pathogenic variants in our 11 genes of interest is very low (1 × 10−7 %, see Supplementary Note). Our diagnostic yield of 10.8% parallels the 9.3% in previous work, which identified pathogenic variants in the same 11 genes based on research-level whole exome sequencing of 355 patients with sporadic aortic dissection and early onset (≤56 years of age).17 In contrast, the yield of whole exome sequencing in 102 thoracic aortic aneurysm and dissection patients was much lower, with only 3.9% of cases carrying a pathogenic variant in one of 21 genes of interest.18 Similarly, Weerakkody et al.19 performed targeted genetic analysis of 15 genes in a mixed cohort of 967 familial and sporadic thoracic aortic aneurysm or dissection cases, and identified 49 pathogenic or likely pathogenic variants in 47 patients which represents a diagnostic yield of 4.9%. We report a two-fold increased proportion of pathogenic variant carriers (10.8%) in a cohort with a more severe phenotype consisting of only thoracic aortic dissection or rupture cases, suggesting the utility of pursuing a clinical genetic diagnosis in this patient group specifically. The 89% of dissection cases that do not have a pathogenic variant may be due to a pathogenic variant currently annotated as a VUS, a pathogenic variant in a gene not yet identified, a high polygenic risk of aortopathy, and/or environmental risk factors. Additional studies of dissection cases may help identify novel genes underlying risk in remaining cases. Notably, the incidence of BAV in non-pathogenic variant carriers (17/216) is higher than that of the general population similar to other studies20, indicating that BAV is a risk factor for aortic dissection even in the absence of a known pathogenic variant.
The significant risk factors for a pathogenic variant in patients with thoracic aortic dissection or rupture were young age (<50 years old), no history of hypertension, but strong family history of thoracic aortic aneurysm, dissection or rupture (Table 4). This is in agreement with a recent study in familial and sporadic cases of aneurysm or dissection of the thoracic aorta which demonstrated a significantly increased probability of harboring a pathogenic or likely pathogenic variant in cases which were syndromic, young (age <50), or with a known or probable family history.19 Patients with pathogenic variants in TGFBR1/2 (Loeys-Dietz syndrome), FBN1 (Marfan syndrome), and MYH11 have a higher risk of aortic dissection and suffer more complications from aortic dissection, including death. Therefore AHA/ACC guideline recommends early and aggressive prophylactic operation to resect the abnormal thoracic aorta in patients with pathogenic variants.8 Our results support the clinical importance of obtaining clinical genetic testing of known hereditary thoracic aortic dissection genes for thoracic aortic dissection and rupture patients, especially those with onset < 50 years old, no history of hypertension, and a positive family history of thoracic aortic disease.
It is important to clarify that other circumstances may exist that would warrant similar or different recommendations based on our findings. For instance, if a patient had a positive family history of thoracic aortic disease, clinical genetic testing for the patient and family members especially the offspring would be recommended despite the patient’s age at the time of dissection (< or > 50 years of age). If a patient had a negative family history and was < 50 years of age, clinical genetic testing for the patient would be recommended, but cascade screening for family members would only be recommended if a pathogenic variant was identified in the patient. Beyond clinical genetic testing, screening with a CT angiogram (CTA) or MRI would be recommended to rule out thoracic aortic disease among the patient’s family members. If a patient had a negative family history and was > 50 years of age, clinical genetic testing for the patient or family members would not be recommened, although screening with a CTA or MRI would be recommended to rule out thoracic aortic disease among the patient’s family members. Routine surveillance should be performed for all patients surviving a thoracic aortic dissection. Less frequent surveillance utilizing a CTA or MRI is recommended for family members without thoracic aortic diease at initial CTA or MRI since family members may have a higher risk of thoracic aortic dissection compared to the normal population.
We did not find a difference in the percentage of VUS in 11 dissection genes among cases compared to controls (p = 0.07). In contrast, a previous study found a significantly increased burden of VUS in hereditary thoracic aortic dissection genes in dissection cases < 56 years of age compared with public controls (p = 2 × 10−8).17 However, several differences in the two studies may contribute to the varied results. Whereas the sample size of the previous study’s control group was substantially higher, we analyzed cases and controls from the same batch and performed all quality control and variant annotation blinded to case or control status. Additionally, a focus on younger onset dissection cases may identify higher rates of VUS that may actually be pathogenic.21 Although the 2015 American College of Medical Genetics guidelines22 state that “a variant of uncertain significance should not be used in clinical decision-making,” we found evidence that VUS from clinical genetic testing resulted in the introduction of syndromic labels and diagnoses into the electronic medical record. Specifically, a VUS in TGFBR2 was subsequently described as a “novel change likely causing Loeys-Dietz syndrome.” The statistically similar rate of VUS in cases and controls demonstrates the need for a greater understanding of the high frequency of VUS in controls (15% in Guo et al17 and 20% in this study) and careful interpretation of VUS in clinical practice.
To address the limitation that our sample processing and whole exome sequencing was not performed in a CLIA-certified laboratory, we verified pathogenic variants using MIPS. Furthermore, we performed expert-annotation of variant pathogenicity blinded to case or control status. This, coupled with the absence of pathogenic variants in controls, provides increased confidence in the results. These precautions lend additional evidence that the research-level whole exome sequencing results are of high enough quality to return findings to patients, which will trigger verification by clinical genetic testing performed in a CLIA-certified laboratory and cascade screening for the same pathogenic variant in family members. Electronic medical record review of the cases with a pathogenic variant suggested an average of 4 (3.88) first degree relatives per patient that would now be candidates for cascade screening. We are also limited by the 1) retrospective review, 2) possibility of incomplete electronic medical records, especially if a patient was seen at an outside institution, and 3) potential for limited family history knowledge.
In conclusion, this work provides evidence that whole exome sequencing and annotation can accurately identify pathogenic variants in established genes for hereditary thoracic aortic dissection in patients with a thoracic aortic dissection or rupture. Moreover, the results highlight meaningful implications for precision health by providing clinical guidance on how to manage both patients and family members. We recommend clinical genetic testing of hereditary thoracic aortic dissection genes in patients who have suffered a thoracic aortic dissection, especially for those with an onset prior to 50 years old, a family history of thoracic aortic disease, and no history of hypertension. Clinical genetic testing may help to prevent catastrophic events, such as thoracic aortic dissections and death, for family members of pathogenic variant carriers who have a high risk but have yet to develop the phenotype.
Supplementary Material
Acknowledgments:
Sequencing/Genotyping services were provided through the RS&G Service by the Northwest Genomics Center at the University of Washington, Department of Genome Sciences, under U.S. Federal Government contract number HHSN268201100037C from the National Heart, Lung, and Blood Institute. The authors acknowledge the University of Michigan Medical School Central Biorepository for providing biospecimen storage, management, and distribution services in support of the research reported in this publication. We acknowledge the University of Michigan DNA Sequencing Core. We thank the clinicians, staff, and study participants from the CHIP Biorepository and Michigan Genomics Initiative.
Sources of Funding: National Institutes of Health (R01-HL127564, R35-HL135824, and R01-HL142023 to C.J.W., K08HL130614 and R01HL141891 to B.Y., R01HL109942 to D.M.M., and R01HL122684 and R01HL139672 to S.K.G.). National Science Foundation (DGE 1256260) to B.N.W. The Phil Jenkins and Darlene & Stephen J. Szatmari Funds to B.Y. The Joe D. Morris Collegiate Professorship, the David Hamilton Fund, and the Phil Jenkins Breakthrough Fund in Cardiac Surgery to H.P.
Footnotes
Disclosures: Himanshu J. Patel (hjpatel@med.umich.edu) is a consultant for WL gore Edwards and Medtronic, and these efforts are “Modest”. Cristen J Willer’s spouse works at Regeneron Pharmaceuticals. Other Disclosures: none
References:
- 1.Kent KC, et al. Screening for abdominal aortic aneurysm: a consensus statement. J Vasc Surg. 2004;39:267–269. [DOI] [PubMed] [Google Scholar]
- 2.Clouse WD, et al. Acute aortic dissection: population-based incidence compared with degenerative aortic aneurysm rupture. Mayo Clin Proc. 2004;79:176–180. [DOI] [PubMed] [Google Scholar]
- 3.Milewicz DM, et al. Heritable Thoracic Aortic Disease Overview In: Adam MP, Ardinger HH, Pagon RA, Wallace SE, Bean LJH, Stephens K and Amemiya A, eds. GeneReviews((R)) Seattle (WA): University of Washington, Seattle University of Washington, Seattle; GeneReviews is a registered trademark of the University of Washington, Seattle. All rights reserved.; 1993. [PubMed] [Google Scholar]
- 4.Pomianowski P, et al. The genetics and genomics of thoracic aortic disease. Ann Cardiothorac Surg. 2013;2:271–279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Brownstein AJ, et al. Genes Associated with Thoracic Aortic Aneurysm and Dissection: An Update and Clinical Implications. Aorta (Stamford). 2017;5:11–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Brownstein AJ, et al. Genes Associated with Thoracic Aortic Aneurysm and Dissection: 2018 Update and Clinical Implications. Aorta (Stamford). 2018;6:13–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Renard M, et al. Clinical Validity of Genes for Heritable Thoracic Aortic Aneurysm and Dissection. J Am Coll Cardiol. 2018;72:605–615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hiratzka LF, et al. 2010 ACCF/AHA/AATS/ACR/ASA/SCA/SCAI/SIR/STS/SVM guidelines for the diagnosis and management of patients with thoracic aortic disease. Circulation. 2010;121:e266–369. [DOI] [PubMed] [Google Scholar]
- 9.Erbel R, et al. 2014 ESC Guidelines on the diagnosis and treatment of aortic diseases: Document covering acute and chronic aortic diseases of the thoracic and abdominal aorta of the adult. The Task Force for the Diagnosis and Treatment of Aortic Diseases of the European Society of Cardiology (ESC). Eur Heart J. 2014;35:2873–2926. [DOI] [PubMed] [Google Scholar]
- 10.Wallace SE, et al. MYLK pathogenic variants aortic disease presentation, pregnancy risk, and characterization of pathogenic missense variants. Genet Med. 2019;21:144–151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bradley TJ, et al. The Expanding Clinical Spectrum of Extracardiovascular and Cardiovascular Manifestations of Heritable Thoracic Aortic Aneurysm and Dissection. Can J Cardiol. 2016;32:86–99. [DOI] [PubMed] [Google Scholar]
- 12.Regalado ES, et al. Aortic Disease Presentation and Outcome Associated With ACTA2 Mutations. Circ Cardiovasc Genet. 2015;8:457–464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Sherry ST, et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 2001;29:308–311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Landrum MJ, et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014;42:D980–985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Lek M, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Loeys BL, et al. The revised Ghent nosology for the Marfan syndrome. J Med Genet. 2010;47:476–485. [DOI] [PubMed] [Google Scholar]
- 17.Guo DC, et al. Heritable Thoracic Aortic Disease Genes in Sporadic Aortic Dissection. J Am Coll Cardiol. 2017;70:2728–2730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ziganshin BA, et al. Routine Genetic Testing for Thoracic Aortic Aneurysm and Dissection in a Clinical Setting. Ann Thorac Surg. 2015;100:1604–1611. [DOI] [PubMed] [Google Scholar]
- 19.Weerakkody R, et al. Targeted genetic analysis in a large cohort of familial and sporadic cases of aneurysm or dissection of the thoracic aorta. Genet Med. 2018;20:1414–1422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Michelena HI, et al. Natural history of asymptomatic patients with normally functioning or minimally dysfunctional bicuspid aortic valve in the community. Circulation. 2008;117:2776–2784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kwartler CS, et al. Variants of Unknown Significance in Genes Associated with Heritable Thoracic Aortic Disease Can Be Low Penetrant “Risk Variants”. Am J Hum Genet. 2018;103:138–143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Richards S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.