Abstract
The intestinal immune system has the challenging task of tolerating foreign nutrients and the commensal microbiome, while excluding or eliminating ingested pathogens. Failure in such balance leads to severe diseases such as inflammatory bowel diseases (IBD), food allergies or invasive gastrointestinal infections1. Multiple immune mechanisms are therefore in place to maintain tissue integrity, including balanced generation of effector T (TH) cells and FOXP3+ regulatory T (pTreg) cells, which mediate resistance to pathogens and regulate excessive immune activation, respectively1–4. The gut–draining lymph nodes (gLNs) are critical sites for orchestrating adaptive immunity to luminal perturbations5–7. However, how they manage to simultaneously support tolerogenic and inflammatory reactions is incompletely understood. Here we report that gLNs are immunologically unique according to the functional gut segment they drain. Stromal and dendritic cell gene signatures as well as T cell polarization against the same luminal antigen differed between gLNs, the proximal small intestine–draining gLNs preferentially giving rise to tolerogenic and the distal gLNs to pro-inflammatory T cell responses. This segregation permitted targeting distal gLNs for vaccination and maintenance of duodenal pTreg cell induction during colonic infection. Conversely, the compartmentalized dichotomy was perturbed by surgical removal of select distal gLNs and duodenal infection, impacting both lymphoid organ and tissue immune responses. Our findings reveal that the conflict between tolerogenic and inflammatory intestinal responses is in part resolved by discrete gLN drainage, and encourage gut segment-specific antigen targeting for therapeutic immune modulation.
Appropriate lymphatic trafficking of immune cells to gLNs is essential for intestinal adaptive immunity (Fig.1 a)6,8. Previous studies revealed the drainage map to various gLNs along the murine gut9–12, and described immunological differences between gLNs11,13, but the underlying cellular components and functional consequences of gut segment-specific drainage have not been systematically addressed. We sought to understand how compartmentalized lymphatic drainage of the intestinal milieu contributes to distinct immune responses towards luminal antigens. We first imaged the gut lymphatic system using 3D imaging of solvent–cleared tissue stained with an antibody against the lymphatic endothelial cell (LEC) surface marker LYVE-1. En bloc imaging exposed the lymphatic route of the intestine to gLNs via afferent lymphatics in the mesentery (Fig. 1 a–c, Extended Data Fig. 1a–c, Supplementary Videos 1–4). The size and shape of individual gLNs differs considerably regardless of the microbiota (Extended Data Fig.1d, e). Dye injected into individual gLNs did not spread to other gLNs, suggesting that the lymph remains compartmentalized until it reaches the thoracic duct (Extended Data Fig. 1f–i). Dye injection into the intestinal muscularis confirmed that the gLNs drain different gut segments9–12 despite the network–like lymphatic structure in the gut wall (Extended Data Fig. 1j–q, Supplementary Video 5–7). The progressive shortening of the lymphatic lacteals in the villi along the small intestine is modulated by the microbiome, as germ free (GF) mice displayed lengthened duodenal and shortened ileal lacteals (Extended Data Fig.2a). To explore if the combination of compartmentalized absorption and drainage result in differential nutrient exposure in the gLNs, we tracked uptake of radiolabelled retinol post-feeding, as a lipid–soluble proxy and an immuno-modulatory nutrient, that depends largely on packing into chylomicrons and lymphatic absorption by the upper small intestine (Extended Data Fig. 2b). Indeed, most retinol was absorbed in the duodenum followed by a gradient along the intestine, and this was mirrored in the gLNs (Fig. 1d,e, Extended Data Fig. 2b–g), illustrating that the gLNs are exposed to region–specific lymph composition.
Figure 1. The gLNs are metabolically and immunologically unique according to the gut segment drained.
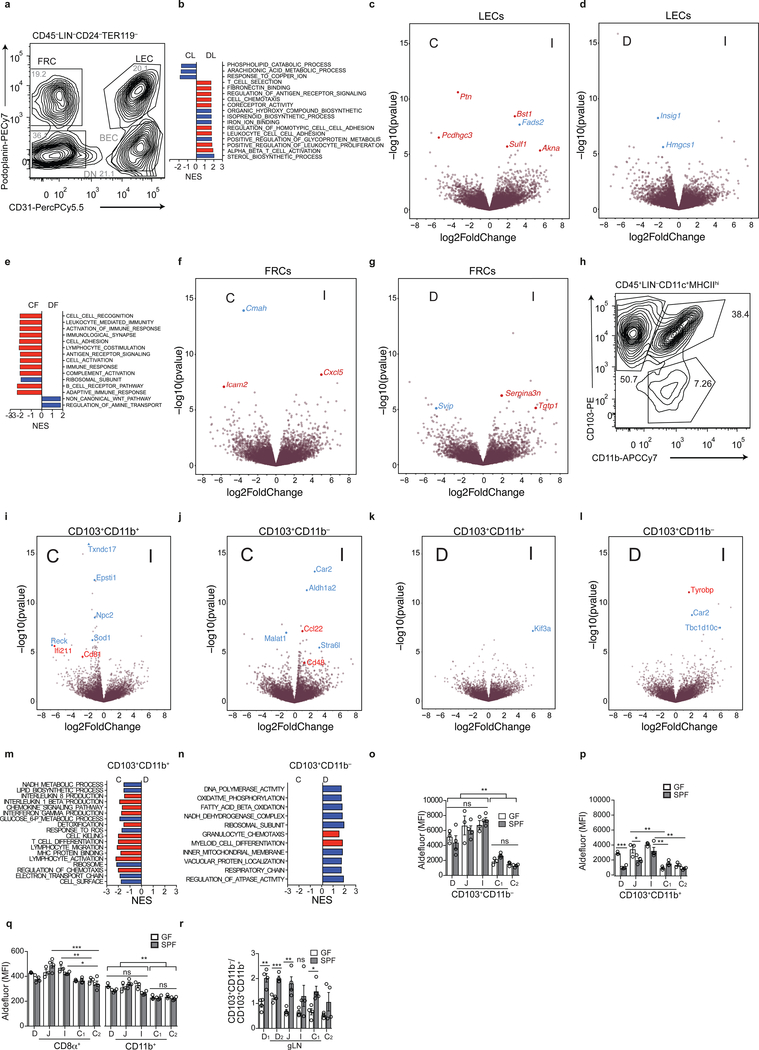
a, Schematic of gLNs. b, c, 3D reconstruction of mouse lymphatics (α-LYVE-1) and position after iDISCO+ (b), co-stained with α-GFP (ItgaxVenus mice) and α-insulin (c). d, e, Percentage of 3H-retinol absorption into indicated gut segment (d) or gLN (e) of mice 3 h after gavage with 1 μCi 3H-retinol in 100 μl olive oil with or without pre-treatment with 5 μl chylomicron formation inhibitor Pluronic L-81 3 h prior to gavage (n=4 per group). f, RNA-seq PCA of FRCs and LECs sorted from D, I, C-gLN from SPF C57BL/6 mice (n=3, pooled from 2 mice each). PC1 accounts for 65.1 %, PC2 for 5.7 % of the variance. g, h, Volcano plot depicting differentially expressed genes among LECs (g) and FRCs (h) from D or C-gLNs. Blue dots indicate metabolic, red dots immunity–related genes. D=duodenum, D1=portal LNs, D2=distal duodenum gLNs, J=jejunum, I=ileum, C(1)=cecal-colonic, C2=ascending colonic, IL/CA=iliac and caudal (both rectal) gLNs, ing=inguinal LN, C=colon. *P < 0.05, **P < 0.01 and ***P < 0.005 (ANOVA).
We asked how compartmentalized drainage impacts gLNs on a cellular level. Stromal cells of pooled gLNs were shown to be tolerance–promoting compared to non–intestinal LNs14,15, therefore we analysed the transcriptome of two major stromal cell populations, lymphatic endothelial cells (LECs) and fibroblastic reticular cells (FRCs), isolated from D-, I and C-gLNs (Extended Data Fig. 3a). Principal component analysis (PCA) indicated LECs and FRCs displayed differential gene expression according to the gut segment they drained (Fig. 1f). LECs from D-gLNs showed a distinct metabolic signature indicative of elevated cholesterol and lipoprotein handling as well as fatty acid utilization, likely driven by the lipid-rich lymph of the proximal small intestine (Fig. 1g, Extended Data Fig. 3b–d, Supplementary Table 1–4). FRC gene expression profiles were affected in a more pronounced manner by location, most evidently in immune cell migration and activation pathways, which were upregulated in the C-gLN (Fig. 1h, Extended Data Fig. 3e, Supplementary Table 5–8). Differences within the small intestine were less extreme, although the I-gLN FRCs also displayed a gene signature associated with leukocyte stimulation (Extended Data Fig. 3f, g).
Because stromal cells may influence antigen–presenting cell function in gLNs14, we probed if gut segment–specific lymphatic drainage influences gene expression profiles of migratory dendritic cells (DCs). The two MHCIIhi migratory DC populations, CD103+CD11b– and CD103+CD11b+ DCs (Extended Data Fig. 3h), are highly represented in gLNs and implicated in tolerogenic and pro-inflammatory responses, respectively16, 17. PCA revealed that DCs segregated between small versus large intestine and less between D- and I-draining gLNs (Fig. 2a). Both DC subsets exhibited immunological and metabolic differences by gLN location (Fig. 2b, c, Extended Data Fig. 3i–n, Supplementary Table 9–16). Duodenal-gLN DCs displayed a less pro-inflammatory signature than their colonic counterparts: D-gLN CD103+CD11b+ DCs expressed lower levels of inflammatory cytokine receptors and pathways such as interferon or IL-1β (Fig. 2b, Extended Data Fig. 3m); D-gLN CD103+CD11b– DCs were specifically enriched for Ccl22, encoding a chemokine associated with migration of Tregs, and Treg–promoting factor Aldh1a2, a rate–limiting enzyme for retinoic acid (RA)11 production from retinol (Fig. 2c, Extended Data Fig. 3j, o–q). Differences in RA production capacity between small and large intestine gLNs were preserved in GF mice (Extended Data Fig. 3o–q), however the ratio of tolerogenic CD103+CD11b– DCs to pro-inflammatory CD103+CD11b+ DCs was inverted in all gLNs analysed (Extended Data Fig. 3r). These data suggest the D-gLNs as tolerogenic environments at least in part due to favourable DC and stromal cell signatures, a phenomenon boosted by the microbiota. Overall, the discrete gLNs appeared poised to mount appropriate immune responses corresponding to the environment of the intestinal region they drain.
Figure 2. Migratory DC profiles and CD4+ T cell polarization differ between gLNs and follow proximal to distal gradients.
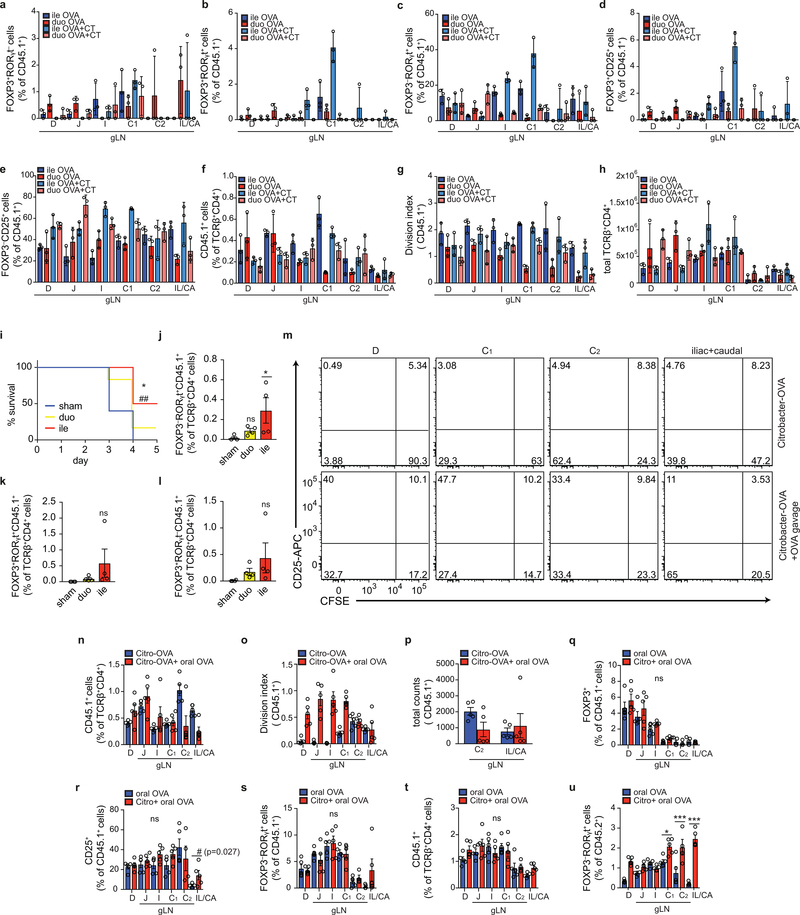
a, RNA-seq PCA of CD103+CD11b+ and CD103+CD11b– DCs sorted from D, I, C-gLN from SPF C57BL/6 mice (n=4, pooled from 2 mice each). PC1 accounts for 52.7%, PC2 for 10% of the variance. b, c, Volcano plot depicting differentially expressed genes among DCs from D or C-gLNs. Blue indicates metabolic, red immunity–related genes. d, Frequency of FOXP3+RORγt+ among TCRβ+CD4+ cells in gLNs from GF or SPF mice (n=3, representing 2 independent experiments). e, Frequency of RORγt+ among TCRβ+CD4+ cells in gLNs from GF mice or GF mice 8 days post-SFB monocolonization (n=3, representing 2 independent experiments). f, Frequency of FOXP3+ among CD45.1+ cells 64 h post-adoptive transfer of 1×106 naïve CD45.1+ OT-II cells into GF and SPF CD45.2 mice (n=4) gavaged OVA 48 h and 24 h pre-analysis. g, Frequency of total FOXP3+ among CD45.1+ cells in LN or spleen 64 h after transfer of 1 × 106 naïve CD45.1+ OT-II cells into SPF CD45.2 mice (n=3) with intravenous injection of OVA 48 h pre-analysis, representative of 2 experiments. h, i, RORγt+ among FOXP3+ CD45.1+ (h) and among CD45.1+ (i) cells in same experiment as f, representing 2 independent experiments. # fewer than 200 cells. LN abbreviations as in Fig. 1; ing=inguinal LN. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA).
We next investigated if CD4+ T cell fates correlated with migratory–DC profiles or the presence of the microbiota. Although GF mice harbour reduced Treg cells (FOXP3+) in the small and large intestines18, we observed no difference in gLN Treg cell frequency between GF and SPF mice (Extended Data Fig. 4a–c). TH17 (RORγt+) effector cells and their corresponding suppressive RORγt+ pTreg cells can be induced by the microbiota1–3, and indeed GF mice displayed a severe reduction in RORγt+ pTreg cells in all tested gLNs, although we did not observe a significant decrease in TH17 cells (Fig. 2d, Extended Data Fig. 4d, e). We then tracked T cell responses to Segmented Filamentous Bacteria (SFB), pathobionts that elicit a strong TH17 response and preferentially colonize the ileum and colon19,20 and found that monocolonization of GF mice with SFB led to the enrichment of RORγt+CD4+ T cells only in the I- and C-gLNs (Fig. 2e). To directly address whether initial CD4+ T cell polarization occurs in a compartmentalized manner, we adoptively transferred naïve Ovalbumin (OVA)-specific CD4+ cells (CD45.1 OT-II cells) and analysed their activation upon OVA gavage17. OT-II cells proliferated and were activated in all gLNs, with the exception of the iliac and caudal gLNs (Extended Data Fig. 4f–i). We then assessed CD4+ T cell fate and its dependence on the microbiota. Amongst the retained OT-II cells (Extended Data Fig. 5a), we observed a gradient of pTreg cell induction that declined in a proximal to distal manner (Fig. 2f). In GF mice, pTreg cell induction was decreased in all gLNs (Fig. 2f, Extended Data Fig. 5b) despite similar antigen access, induced proliferation and increased CD25 expression in the upper small intestine gLNs compared to SPF mice (Extended Fig. 5c–g), suggesting that the less favourable DC composition in GF gLNs contributed to this effect. Limiting antigen availability was also unlikely to explain the gradient of OT-II pTreg cells in the small intestine and C-gLNs of SPF mice, as proliferation and CD25+ frequencies did not differ between these gLNs (Extended Data Fig. 5f, g). Furthermore, intravenous (i.v.) OVA administration also resulted in a decreasing proximal to distal FOXP3 induction gradient, yet similar OT-II cell recovery and proliferation among gLNs (Fig. 2g; Extended Data Fig. 5h–m). In contrast, OT-II RORγt+ TH and OT-II RORγt+ pTreg cells displayed an ascending proximal to distal gradient that was independent of the microbiota (Fig. 2h, i), and correlated with an increase in IL12/23p40+ frequency amongst CD103+CD11b– DCs (Extended Data Fig. 5n). These data indicate that under homeostatic conditions, the D-gLNs are the primary sites for pTreg cell induction by dietary antigen. In contrast, distal gLNs favour TH17 and RORγt+ pTreg cell differentiation, underscoring the immunological distinctness of the proximal and distal intestinal milieus.
We investigated possible consequences of anatomically-segregated T cell fates along the intestine. First, we analysed T cell responses to an antigen delivered to distinct intestinal sites by surgical injection of OVA-containing solution into the duodenum versus ileum. This approach did not elicit significant RORγt– pTreg induction, possibly due to the pro-inflammatory surgical setting and lack of postprandial mechanisms. Ileal injection elicited more pronounced RORγt+ TH and RORγt+ pTreg cell differentiation than intra-duodenum OVA administration in the respective draining gLNs; this response pattern was boosted by co-injection of cholera toxin (CT, Fig. 3a, Extended Data Fig. 6a–h). This result confirms the more pro–inflammatory potential of the I- and C-gLNs when compared to D-gLNs. We next injected CT-OVA into the ileum or duodenum and compared these mice to sham–operated mice in their susceptibility to OVA–expressing Salmonella enterica Typhimurium (Stm-OVA), which infects the ileum and cecum before systemic dissemination. Only mice previously injected with CT-OVA in the ileum displayed a delayed weight loss compared to sham-operated mice (Fig. 3b, c, Extended Data Fig. 6i), which correlated with a reduced Salmonella invasion and an increased frequency of RORγt+ pTreg and TH17 cells in the lamina propria (Extended Data Fig. 6j–l). We then tested whether Citrobacter rodentium infection, anatomically restricted to the cecum and colon, perturbs proximal pTreg induction. Oral OVA-specific pTreg induction in D-gLNs was unaffected by infection with OVA–expressing Citrobacter, which resulted in OVA–specific CD4+ T cell activation in the C-gLNs (Fig. 3.d, e, Extended Data Fig. 6m–p), or upon wild-type Citrobacter (Extended Data Fig. 6q–u).
Figure 3. Compartmentalization of CD4+ T fates permits vaccination via the ileocecal gLNs.
a, Representative flow cytometry plots for FOXP3 and RORγt expression among CD45.1+ OT-II cells in gLN 48 h post duodenal or ileal injection with OVA or OVA and CT, performed 16 h after adoptive transfer of 1 × 106 naïve cells into SPF CD45.2 C57BL/6 mice (n=4). b, c, Percentage of pre-infection weight (b) and splenic Stm-OVA dissemination (c) of mice infected with Stm-OVA 9 days after duodenal or ileal injection with OVA-CT versus sham operation and 10 days post adoptive transfer of 1 × 105 naïve OT-II cells into SPF CD45.2 mice. b, pool of 2 independent experiments, sham (n=11), CT-OVA duo (n=13), CT-OVA ile (n=15), c, single experiment, sham (n=6), CT-OVA duo (n=6), CT-OVA ile (n=7). d, e, Frequency of total FOXP3+ (d) and FOXP3–CD25 + (e) among CD45.1+ in gLNs 64 h post adoptive transfer of 1 × 106 naïve CD45.1+ OT-II cells into SPF CD45.2 mice infected or not with Citrobacter-OVA 9 days before and after gavage of OVA 48 h and 24 h prior to analysis (n=5, represents 2 independent experiments). f, Experimental design for g. g, Frequency of Vβ14+CD45.1+ RORγt+ cells among CD3+CD4+ cells in indicated gLN from mice with sham operation (n=12) or surgical removal of the I- and C1-gLN (ΔicLN, n=14) 64 h (day 3, f) after adoptive transfer of 4 × 105 naïve SFB specific CD45.1+ 7B8tg cells into SPF CD45.2 mice, pooled data from 3 independent experiments with n=4–5 per group each. LN abbreviations as in Fig.1 and 2; duo=duodenum, ile=ileum. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA).
The gLNs are crucial for physiological responses to microbiota, pathogen–resistance mechanisms and oral tolerance7,21,22. We asked whether the I- and C-gLNs are required for the generation of SFB-specific TH17 cells. Upon transfer of naïve SFB–specific CD4+ T cells (7B8tg cells)2 into SPF mice colonized with SFB, we only recovered significant numbers of transferred cells in the I- and C-gLNs, which upregulated RORγt (Fig. 3f, g, Extended Data Fig. 7a–e). Surgical removal of I- and C-gLNs shifted the SFB-specific TH17 response to the gLNs draining gut segments with lower colonization (Fig. 3f,g, Extended Data Fig. 7a–g), which was correlated with an ectopic expansion of 7B8tg cells in the intestine, but only if some SFB was present in the lumen due to recent colonization (Extended data Fig. 7a–c, h–s). It is unlikely that surgery caused lymph rerouting to surrounding gLNs as indicated by absence of dye spreading to adjacent LNs, or SFB expansion in surrounding gut segments as evidenced by SFB–specific 16S qPCR (Extended Data Fig. 8a–c). Hence, these data suggest that individual gLNs also modulate T cell responses in the intestinal tissue in a region–specific manner. If faced with pathogens or pathobionts this regionalization may help to contain tissue damage. Taken together, these results illustrate beneficial effects of discrete gLN drainage, which may avoid immunological conflict between responses induced at different gut sites and help preserve these responses in the tissue.
Viral and bacterial gastrointestinal infections were shown to perturb the generation of pTreg cells recognizing dietary antigen and subsequent oral tolerance6,23. We addressed whether the proximal gLNs alone are necessary or sufficient for oral tolerance, a pTreg cell–dependent process4, by surgical removal of either the D- or the I- and C-gLNs followed by subjecting the mice to an oral antigen-mediated asthma suppression model17. Oral tolerance was intact upon proximal or distal LN removal (Extended Fig. 8d–i), likely because pTregs cells generated in remnant gLNs are sufficient17. To induce an immunological conflict only in the proximal intestine–draining gLNs, we chose the helminth Strongyloides venezuelensis (S.v.) which displays distinct duodenal tropism24. Upon S.v. infection only the D-gLNs were swollen, underwent significant restructuring and displayed increased immune cell counts compared to gLNs of non-infected (N.I.) mice, in correlation with compartmentalized duodenal worm load (Fig. 4a, b, Extended Data Fig. 9a–c, Supplementary Videos 8–17). We also observed a D-gLN selective influx of CD11b+ DCs, previously linked to the induction of Th2 responses25, at the expense of the more tolerogenic CD103+CD11b– and CD8α+ DC subsets, as well as a type 2 immunity signature comprising eosinophils, GATA3+ TH2 cells and GATA3+ Treg cells even after the worm is cleared (Fig. 4 c-h and Extended Data Fig. 9d–l). Upon S.v. infection transferred OT-II cells displayed a significant reduction in upregulation of FOXP3 and CD25 specifically in the D-gLNs upon OVA gavage compared to N.I. mice, despite proportional gLN seeding (Fig. 4i, Extended Data Fig. 10a–e),. Finally we asked if transient infection during the initial antigen exposure impacted tolerance to dietary antigens17 (Fig. 4j). We observed a partial impairment of suppression of eosinophilia and DC infiltration into the bronchial-alveolar fluid (BALF) and lungs, and increased OVA-specific IgG1 levels in the S.v. group compared to N.I. mice (Fig. 4 k-m, Extended Data Fig. 10f, g). Taken together these data show that duodenal infection can perturb local responses as well as impact systemic tolerance to gut antigens.
Figure 4. Duodenal infection leads to a compartmentalized immune conflict in the duodenal gLNs and to compromised oral tolerance.
a, Dissected gLNs from non-infected (N.I.) C57BL/6 mice or mice infected with 700 S. venezuelensis (S.v.) larvae 8 days prior to harvest. b, Total CD45+ cell counts (n=5, represents 3 independent experiments). c-f, Frequency of CD103+CD11b– (c), CD103+CD11b+ (d) and CD103–CD11b+ (e) among MHCIIhiCD11c+ cells and eosinophils among CD45+ cells (f) (n=5, represents 2 independent experiments). g, Representative flow cytometry plot of GATA3+ and FOXP3+ CD4+ T cells. h, Frequency of GATA3+ CD4+ T cells (n=5, represents 3 independent experiments). i, Frequency of total FOXP3+ among CD45.1+ cells in gLNs 64 h post adoptive transfer of 1 × 106 naïve CD45.1+ OT-II cells into CD45.2 mice (n=8, pool of 2 independent experiments) infected with S.v. larvae or N.I. 8 days and gavaged with OVA 48 h and 24 h prior to analysis. j, Scheme of oral tolerance experimental set-up in S.v. infected mice. k-m, Total eosinophils in bronchoalveolar lavage fluid (BALF) (k), frequency of eosinophils among CD45+ cells in lung tissue (l) and OVA-specific IgG1 levels in serum (m) from mice infected with S.v. or N.I. during antigen feeding (+OVA groups) or no feeding (-OVA groups), at 21 d after first immunization with OVA-alum (j) (n=13 for +OVA groups, n=10 for –OVA groups, pool of 2 independent experiments). LN abbreviations as in Fig. 1. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA).
Our data uncover a mechanism by which the intestinal immune system simultaneously handles regulatory versus pro-inflammatory responses: by anatomical segregation of these reactions into functionally distinct gLNs. They also unveil that a cooperation of effector (TH) and corresponding regulatory mechanisms (lineage–defined Treg cells) exist at every site to dampen excessive, gut segment–specific inflammation. It remains to be determined whether specific DC subpopulations with varying tolerogenic or inflammatory roles along gLNs mediate distinct effects towards dietary, microbiota or pathogen-derived antigens3,16. The efficient drainage of dietary antigens into tolerance–promoting lymph nodes associated to the proximal intestine implicate these as sites of natural food allergy prevention, and duodenal infection and dysbiosis as environmental perturbations that can alter the allergic outcome. However, the dietary duodenal route could be exploited therapeutically for inducing tolerance to otherwise inaccessible pro-inflammatory antigen sources, such as dysbiotic bacteria in IBD patients, while selective distal gLN targeting may lead to successful intestinal vaccination.
Methods
Mice
C57BL/6J CD45.1 (B6.SJL-Ptprca Pepcb/BoyJ) were from Jackson laboratories, CD45.2 (C57BL/6) mice were purchased from the Jackson Laboratories or Taconic Farms, and CD45.1 OT-II TCR-transgenic were originally purchased from Taconic Farms and maintained in our facilities. B6.Cg-Tg(Itgax-Venus)1Mnz/J mice were provided by M. Nussenzweig (The Rockefeller University). C57BL/6-Tg (Tcra,Tcrb)2Litt/J (7B8tg) mice were either purchased from Jackson Laboratories or provided by D. Littman (NYU). Mice were used at 7–12 week old, and male mice were used throughout the study. A minimum of biological triplicates was used throughout the study. For in vivo studies, the maximum number of animals available per treatment group were used to counteract biological variation: for all adoptive transfer experiments up to 5 animals per group, for oral tolerance experiments up to 8 animals per group, for infection and surgery experiments up to 8 per group. For surgeries (lymph node removal and intestinal antigen injection) removal surgery and OVA+ cholera toxin surgery group sizes were larger than sham (additional two mice) in the event an animal had to be euthanized during post-operative care due to the higher invasiveness of the procedure. All surviving animals were included in the study. For all studied B6 mice were purchased in batches of 10–20 mice and randomly distributed to cages. Infection/colonization studies: Cages of 5 mice were randomly infected/colonized with SFB or note. Because of the high chance of contamination it was not possible to house infected/colonized and non-infected mice in a single cage. CT-OVA injection: Mice were housed in cages of 4–5 mice and within a cage randomly subjected to sham, duodenal or ileal OVA or OVA-CT injection but ensuring mixed groups were present in each cage (2 vs 2 or 2 vs 3 or 2 vs 2 vs 1). Surgery: Mice were housed as 5 mice/ cage, and each cage housed 2 vs 3 or 3 vs 2 sham operation versus lymph node removal surgery mice, to avoid any potential microbial drift or behavioral changes due to the different treatments affecting the study outcome. Blinding was not possible in this study as the experimenters treating the mice were the same as those analyzing the data. The treatment groups had to be clearly identified throughout the study to prevent cross contamination (gnotobiotic/infection experiments), to be able to draw conclusions (CT-OVA injection in duodenum versus ileum) or were obvious by their anatomy ( lymph nodes were removed). Statistics were applied in an unbiased manner. Mice were maintained at the Rockefeller University animal facilities under specific pathogen-free conditions or germ free conditions. Germ-free C57BL/6 mice were generously provided by S. Mazmanian (Caltech) and imported into germ-free flexible film isolators (Class Biologically Clean Ltd.). Mice monocolonized with SFB were kept in flexible film isolators and originally colonized by gavage with fecal extract from SFB monocolonized mice kept at NYU (Littman lab). SFB colonization was verified by real time PCR using SFB-specific 16S primers; germ-free feces served as a negative, Taconic Farms B6 feces as a positive control. The mice were bred in our germ-free facility for maintenance of the strain and kept on sterilized Autoclavable Mouse Breeder Diet (5021, LabDiet, USA), which was also used for control (ex-germ free) SPF mouse breeding and maintenance.
Rats
6–10 week old female Wistar rats were purchased from Charles River. Animal care and experimentation were consistent with the NIH guidelines and were approved by the Institutional Animal Care and Use Committee at the Rockefeller University.
Reagents
The following reagents were purchased from Sigma: Benzyl ether (108014), Dichloromethane (270997), Fast Green FCF (F7252), Heparin (H3393), Omeprazole (O104), Pluronic L-81 (435430), Pyrantel Pamoate (P6210), Ovalbumin (grade VI, A2512; grade III A5378) and Vancomycin (V2002). LPS-free ovalbumin was from Hyglos, Germany (Cat. no 77161). 3H –retinol and Na125I were from Perkin-Elmer. Cholera toxin was from List Biological Laboratories (Cat. no100B).
Antibodies, staining and flow cytometry
Fluorescent-dye-conjugated antibodies were purchased from BD (USA) (anti CD25, 563061; anti-CD45.1, 563982; anti-CD45.2, 560693; anti-CD31, 557355; anti-CD103, 557495; anti-Ly6C, 560595; anti-NK1.1, 562921; anti-SiglecF, 552126; anti-RORγt, 562894; anti-IL17A, 56022 and 559502; anti Vβ5.1, 553190; anti-Vβ14, 553258), eBioscience (USA) (anti-B220, 48–0452-82; anti-CD3e, 48–0031-82; anti-CD4, 83–0042-42; anti-CD8α, 56–0081-82; anti-CD25, 17–0251-82; anti-CD11b, 47–0112-82; anti-CD11c, 25–0114-82, 17–0114-82 and 56–0114-82; anti-CD24, 48–0242; anti-CD45, 25–0451-82; anti-CD45.1, 25–0453-82 and 45–0454-82; anti-CD69; anti-FOXP3, 12–5773-82; anti-GATA3, 12–9966-42; anti-I-A/I-E (MHCII), 46–5321-82 and 56–5321-82; anti-Ly6G, 48–5931-82; anti-Podoplanin, 25–5381-82; anti-Vα2, 48–5812-82, and Streptavidin, 46–4317-82), or Biolegend (USA) (anti CD3e, 100320; anti-CD8α, 100744; anti-CD11b, 101236; anti-CD64, 139306; anti-TCRβ, 109220; anti-TER119, 116206). Additional antibodies were purchased from BioXCell and labelled in-house (anti CD4, BE0003–1; anti-CD8α, BE0004–1; anti-CD11b, BE007; anti-CD19, BE0150; anti-TCRβ, BE00102). Biotinylated antibodies were purchased from BD Pharmigen (anti-B220, 553086; anti-CD8α, 553029; anti-CD11b, 553309; anti-CD11c, 553800; anti-CD25, 553070; and anti-NK1.1, 553163; anti-TER-119, 553672;) or as follows: anti-IgG1, Bethyl, A90–105B; anti- IgG2c, Bethyl, A90–136B; anti-IgE, eBioscience, 13–5992-82; and anti-Neuropilin, R&D Systems, BAF566. Unconjugated antibodies used were anti-LYVE 1, R and D Systems AF2125; anti-GFP (Aves Labs, GFP-1020); anti-OVA IgG1, Biolegend, 520501; anti-IgE, Invitrogen, RMGE00; IgE isotype control, eBioscience 554118. Horseradish peroxidase conjugated Streptavidin was purchased from Jackson ImmunoResearch Laboratories, Inc. Aqua LIVE/DEAD® Fixable Aqua Dead Cell Stain Kit, L-34965, Cell Trace CFSE and Violet Cell Proliferation kits (C34554 and C34557) were purchased from Life Technologies. For cytokine analysis in T cells, cells were incubated for 3.5 h in RPMI with 10% FBS, Brefeldin A (0.5 μg ml–1, Sigma B7651), ionomycin (0.5 μg ml–1, Sigma I0634), and phorbol 12-myristate 13acetate (100 ng ml–1, Sigma P8139), for cytokines in dendritic cells, cells were incubated for 6h in RPMI with 10% FBS and GolgiPlug (1 to 1000, BD Biosciences 555029). Cell populations were stained with Aqua in PBS, followed by incubation with Fc block and antibodies against the indicated cell surface markers in FACS buffer (PBS, 1% BSA, 10 mM EDTA, 0.02% sodium azide). The cells were analyzed live, or fixed in 1% PFA/PBS. For intracellular staining, cells were first stained for surface epitopes and then fixed, permeabilized and stained according to the manufacturer’s protocol (eBioscience 00–5123-43). Flow cytometry was performed on an LSRII (BD Biosciences) and analysed using FlowJo Software (Tree Star). Cell division index was calculated using the FlowJo formula (http://www.flowjo.com/v765/en/proliferation.html), whereby the index represents the fraction of total cell divisions over the calculated total starting cells.
Tissue clearing, light sheet microscopy and image reconstruction
For tissue clearing and staining, the iDISCO protocol was followed as detailed on the continuously-updated website: http://idisco.info with the following specifics: Tissues were stained in primary and secondary antibodies for 4 days each, and were embedded in 1% agarose-TAE prior to the final dehydration. Blocks were imaged using a LaVision Ultramicroscope II and Software. Images were reconstructed using Imaris 8 Software.
Fast green tracing
Mice were anesthetized with isoflurane and their peritoneal cavity exposed. For mapping intestine to lymph node drainage, 3–5 μl of 10 % Fast Green in PBS were injected into the muscularis of the intestine using a nano-injection device (Nanoject III, Drummond). Spreading of green colour through the lymphatics was followed until it accumulated in the draining lymph node and until it faded again (sham operated mice) and reached the duodenal lumen via the bile duct, no longer than 15 minutes. The same timing was applied in mice in which ileal and cecal mLNs had been removed, even though the dye never accumulated in a lymph node. To trace lymphatic mLN efflux, 1– 3 μl of 10 % Fast Green was injected into the center of the mLN, and efflux monitored for 15 min or until the dye was detected in the thoracic duct.
3H-retinol biodistribution
Mice were fasted for 3h prior to gavage with 150 μl PBS with or without 5 μl chylomicron formation inhibiting Pluronic L-81, followed by 1 μCi 3H –retinol in 100 μl olive oil 30 minutes later, and sampled at indicated time points. Serum was sampled from submandibular vein (systemic) or from the portal vein. Lymph was collected from the thoracic duct under isoflurane anaesthesia using a custom-made glass needle (Micropipette puller, Sutter Instrument). Dissected organs were weighed prior to lysis by mechanical disruption in 0.5 ml hypertonic lysis buffer with 1% Triton-X 100, the lysate mixed with 7 ml Ultima Gold scintillation cocktail (Perkin Elmer) and accumulation of radioactivity measured on a scintillation counter. Input radioactivity was estimated by counting 10 % of the gavaged material.
Segmentation of gut-draining lymph nodes
The murine gLNs consist of one hepatic/celiac LN co-draining the duodenum, pancreatic-duodenal LNs draining the duodenum and separately the ascending and transverse colon, the main mesenteric lymph node (mLN) chain draining the distal duodenum, jejunum, ileum, cecum and proximal ascending colon, and the caudal and iliac lymph nodes draining the descending-distal colon. Mesenteric lymph nodes draining intestinal segments were determined anatomically by following the lymphatic vessels connecting the colon, ileum and jejunum to their lymph nodes. Duodenal lymph nodes were revealed by gavaging 100 μl of olive oil (Sigma) and determining the most stomach-proximal lymph nodes surrounded by chyle, indicative of duodenal drainage, 1h post gavage.
Lymphocyte and APC isolation from lymph nodes
Lymph nodes were dissected into cold HBSS, supplemented with Mg2+ and Ca2+, finely chopped and incubated in 400 U/ml Collagenase D (Roche) in HBSS for 25 min at 37°C, 5% CO2. Collagenase was quenched on ice by addition of final 10% FCS. Single cell suspensions were extracted from connective tissue by taking up and resuspending the digests five times. Erythrocytes were lysed by incubation in erythrocyte lysis buffer (Sigma) for 7 min at RT.
Stromal cell isolation from gLNs
gLNs were dissected into 500 μl cold RPMI, supplemented with 2% FCS and HEPES (dissection medium). Tissues were disrupted using 25 G needles, and Liberase TL (Roche) was added to a final concentration of 0.25 mg/ml, collagenase D (Roche) to a final concentration of 400 U/ml. Tissues were incubated at 37°C on an orbital shaker at 80 rpm for 10 minutes, disrupted at RT by gently pipetting digest up and down 50 times, followed by an additional 20 times. Reaction was stopped by placing digst on ice and adding 14 ml of ice cold dissection medium. Cells were spun down at 700 g for 5 minutes at 4°C, resuspended in erythrocyte lysis buffer (Sigma) topped up with medium again, spun and then subjected to staining.
Lymphocyte and APC isolation from small and large intestine
Intestines were separated from mesentery, and Peyer’s Patches (small intestine) and feces were removed. For segmentation of the small intestine, the upper 25% of the small intestine were taken as duodenum, the next 50% as jejunum and the last 25% as ileum. This division allows for quick processing of multiple specimens, though it can lead to some contamination of the ileum and duodenum with jejunum. The cecum was included in the preparation of the large intestine. Intestines were cut longitudinally and washed twice in PBS. Tissue was cut into 1 cm pieces, mucus was removed by incubating the tissue for 10 min in PBS and 1 μM DTT, and the epithelium removed by two incubations in 25 ml of HBSS + 2% FCS + 30 mM EDTA for 10 min at 37°C at 230 rpm with vigorous shaking after each incubation. Tissues were washed in PBS over a sieve, then finely chopped and digested in 6 ml of RPMI per gut segment (Gibco), 2% FCS, 200 μg/ml DNaseI (Roche) and 2 mg/ml Collagenase 8 (Gibco) for 45 min at 37°C, 5% CO2. Digests were taken up and resuspended 10 times, passed through a sieve and the collagenase quenched by addition of 15 ml of cold RPMI, 2% FCS. Cell pellets were resuspended in 40% Percoll (BD Pharmigen) complemented with RPMI, 2% FCS, passed through a 100 μm mesh and separated by centrifugation in a discontinuous Percoll gradient (80%/40%) at 1000 g for 25 min at room temperature (RT). APCs and lymphocytes were isolated from the interphase, washed, and stained for FACS analysis or subjected to re-stimulation.
Cell isolation for RNA-seq
10–14 week old C57BL/6 males served as gLN donors, and biological triplicates or quadruplicates were collected. Cells were sorted using a FACS Aria cell sorter flow cytometer (Becton Dickinson). MLN dendritic cells were pre-enriched using a Pan Dendritic Cell Isolation Kit (130–100-875, Miltenyi Biotec) and LS MACS Separation Columns (Miltenyi Biotec). Dendritic cells were sorted as Aqua–CD45+Lin–(CD3–B220–NK1.1–CD19–) CD11chi, and the subpopulations further as MHCIIhiCD103+CD11b– and MHCIIhiCD103+CD11b+. Stromal cells were not pre-enriched and sorted as Aqua–CD45–TER119–CD24–TCRβ–B220–CD11c– cells, and the subpopulations further as podoplanin+CD31– (FRCs) and podoplanin+CD31+(LECs). Three hundred cells were sorted directly into 25 μl TCL buffer (Qiagen, 1031576) supplemented with 1% β-mercaptoethanol at single cell precision. Samples were kept at room temperature for 5 min, spun down and kept at −80 °C until further processing.
RNA-seq library preparation and sequencing
RNA was isolated using RNAClean XP beads (Agentcourt, A63987) on a magnetic stand (DynaMag, Invitrogen 12331D). Reverse transcription primers were: P1-RNA-TSO: Biot-rArArUrGrArUrArCrGrGrCrGrArCrCrArCrCrGrArUrNrNrNrNrNrNrGrGrG, P1-T31: Biot-AATGATACGGCGACCACCGATCG31T, P1-PCR: Biot-GAATGATACGGCGACCACCGAT. RNA was eluted for 1 min in RT- cDNA synthesis mix 1 (0. 5 μl P1-T31 (20uM), 0.3 μl RNasin plus (Promega, N2615), 1.5 μl 10 mM dNTP, 3.5 μl 10 mM Tris pH 7.5– 0.5% IGEPAL CA-630 (Sigma) and 1.7 μl RNase free ddH2O) and the beads pipetted up and down ten times. The eluted sample was then incubated for 3 min at 72°C, followed by 1 min on ice, then 7.5 μl of mix 2 was added (3 μl 5X FS Buffer SS, 0.375 μl 100 mM DTT, 0.375 μl RNasin plus, 0.5 μl P1-RNA-TSO (40uM), 0.75 μl Maxima RT Minus H (Thermo Scientific, EP0751), 1.8 μl 5M Betaine (Sigma, B0300), 0.9 μl 50 mM MgCl2 and 0.175 μl RNase free ddH2O. Reverse transcription (RT) occurred during a thermal cycle of one cycle (90 min at 42 °C), 10 cycles (2 min at 50°C, 2 min at 42°C) and one cycle (15 min 70 °C), and the product was kept at 4 °C. The cDNA was then amplified using 15 μl of the RT product, 20 μl 2x KAPA HiFi HS Ready Mix (Kapabiosystems, KK2601), 1.5 μl P1-PCR (10uM), and 3.5 μl RNase free ddH2O. Amplification occurred during following cycle: One cycle of 3 min at 98 °C, 20 cycles (15 sec at 98 °C, 20 sec at 67 °C, 6 min at 72 °C), one cycle (5 min at 72 °C), and the product was kept at 4 °C. 20 μl of PCR product were cleaned up using 16 μl RNAClean XP beads. The cDNA was eluted in 20 μl RNase free ddH2O and kept at −20 °C. Isolated amplicons were confirmed to be 1500–2000 bp long by a High Sensitivity DNA Assay (Bioanalyzer). Concentration of all sample was measured on a Qubit fluorometer (Thermo Fisher), all samples were adjusted to 0.1 ng/μl with ddH2O, and 2.5 μl cDNA were subjected to Nextera XT DNA Library preparation (Illumina) using a Nextera XT Index Kit (Illumina, FC-131–1002) and according to the manufacturer’s protocol, except that all volumes were used at 0.5 x of the indicated volumes. Sample quality was again verified by Bioanalyzer, sample concentrations measured on the Qubit fluorometer and adjusted to a concentration of 4.54 ng/ μl. All samples were pooled at equal contribution and run in multiple lanes. Sequencing was performed using 76 base single-end reading on a NextSeq instrument (Illumina).
RNA-Seq data analysis
Gene expression was quantified using the raw fastq files and the mouse reference transcriptome M18 (Gencode - GRCm38.p6) as input for Kallisto (v.0.43.0). The resultant normalized transcript frequencies were provided to the R package Sleuth for differential analysis (v.0.30.0). Analyses were conducted at the gene level, and the likelihood ratio test was applied to capture gene expression differences among all groups. When testing between group pairs, the Wald test was used to determine differentially expressed genes. Genes with log2 fold changes greater than 1 or less than −1 and a false discovery rate (FDR) of 0.05 were considered significant for downstream studies. Gene set enrichment analysis (GSEA) was performed by ranking genes based on log2 fold changes of paired comparisons and subsequently run as a GSEA Preranked analysis (GSEA v3.0) with the c5.all.v6.1.symbols.gmt (Gene Ontology) gene set database. All pathways with a FDR of 0.25 or less were considered significantly different.
RALDH activity assay
RALDH activity was determined using the Aldefluor kit (STEMCELL™ Technologies) according to the manufacturer’s protocol. Cells were analyzed by flow cytometry 30 min after the addition of substrate.
125I-OVA labelling and biodistribution
Iodination of OVA was performed as described previously18. Mice were fasted for 3 h prior to gavage with 4 × 106 CPM 125I-OVA and 50 mg cold OVA (grade III) in 200 μl PBS, and samples were taken at 1 h and 5 h post gavage. Wet weight of tissues was taken prior to measuring radioactivity on a gamma counter (Packard Cobra). Input radioactivity was estimated by counting 10 % of the gavaged material.
Adoptive T cell transfer
Naïve CD4 T cells from spleen and lymph nodes were isolated by negative selection using biotinylated antibodies against CD8α, CD25, CD11c, CD11b, TER-119, NK1.1, and B220 and anti-biotin MACS beads (Miltenyi Biotec). Purity of transgenic CD4+ T cells was verified by flow cytometry (CD45.1+Vα2+Vβ5+CD25– for OT-II cells, CD45.1 CD45.1+ Vβ14+CD25– for 7B8tg cells, typically >90%). T cells were labeled using the Cell Trace™ Violet or CFSE Cell Proliferation Kit (Life Technologies). For OT-II cells, 1×106 were transferred by retro-orbital injection under isoflurane gas anesthesia. For 7B8tg cells, 4 × 105 cells were transferred for analysis of gLNs 60 h post-transfer and 5000 cells for analysis 7 days or more after transfer in the gut and gLNs.
Oral antigen administration
OVA (grade III, Sigma, A5378) was administered at 50 mg in 200 μl PBS by oral gavage using metal gavage needles. Two doses were given with a 24 h interval, the first dose given 16–24 h post adoptive OT-II cell transfer.
Intravenous antigen administration
Endotoxin low OVA (Worthington) was administered by retro-orbital injection at either 1 mg or 0.1 mg in 100 μl PBS 16 h post adoptive OT-II cell transfer. The high dose corresponds to the maximum of 125I-OVA recovered in blood upon OVA gavage (4% of 50 mg gavage/ ml plasma, see Extended Data Fig. 9d, equivalent to 2% in blood, estimating the blood volume of a 20 g mouse is around 1 ml equating to 1mg); the ten times lower dose is equivalent to doses used previously by others11.
Intestinal CT-OVA injection
Naive OT-II cells were adoptively transferred in to 7 weeks old C57B6 mice 16–24 h before surgery (2 × 106 cells for T cell fate analysis in gLNs, 1 × 105 for subsequent Salmonella typhimurium-OVA infection). Mice were anesthetized subcutaneously with 100 mg/kg ketamine (controlled substance provided by the Rockefeller University animal facility), 10 mg/kg xylazine (Akorn, inc.), and in presence of 5 mg/kg analgesic Meloxicam (Putney, Inc.) in 0.5 ml saline. The abdominal area was then shaved, and sterilized by three cycles of wiping with iodine solution and 70 % ethanol after 20 μl of 0.25% bupivacaine (Hospira, Inc.) was injected intra-dermally at the prospective site of incision. The mouse was placed on a heat mat and covered by a sterile surgical plastic with an opening above the abdomen. All work from here was performed aseptically. Skin was incised in the middle of the abdomen and the peritoneum was cut open. One hundred μl of filter-sterilized PBS solution containing 50 mg of OVA and 5 μg Cholera toxin or not was injected into the duodenum (5 mm after the pylorus) or the ileum (5 mm before the most distal Peyer’s Patch) using a 28 ½ G needle pointing toward the distal intestine. The peritoneal muscles were aligned and sutured using absorbable suture (PDS*II, Ethicon), the skin closed with autoclips (Ken Scientific Corp.), and the wound covered with Triple Antibiotic Ointment (Honeywell Safety Products, USA). Mice were allowed to fully awake in a cage placed on a heat mat. For sham surgery, mice were opened and sutured again. Mice that had undergone different surgeries were pooled in cages. The following day mice were monitored for agility and passing stool. Mice were treated with 0.3 mg/kg of buprinex (controlled substance) and 5 mg/kg Meloxicam 24 h after first Meloxicam and for another 3 days every 24 h.
Salmonella typhimurium-OVA infection
Seven days after CT-OVA injection animals were gavaged with 20 mg Streptomycin in 200 μl PBS. The same day, Salmonella typhimurium-OVA was grown overnight to stationary growth phase in LB broth at 300 rpm, 37 °C. The next day, 100 μl aliquot was sub-cultured in 3 ml LB. After 3.5 h, 50 μl subculture was diluted in 50 ml of PBS, giving rise to a density of 105 CFU per 100 μl, and mice were immediately gavaged with 100 μl of this solution. The infection was timed such that 24 h had elapsed since Streptomycin administration and the mice had been fasted 4 h prior to infection. Initial body weight was taken just prior to gavage. Mice were weighed every 24 h post infection. In the survival study, mice were sacrificed when their weight reached 80 % or less of their starting weight. For analysis of Salmonella typhimurium-OVA dissemination, mice were sacrificed 48 h after infection, and CFU in organs determined by serial dilutions on Salmonella Shigella Agar plates.
Citrobacter rodentium and Citrobater rodentium-OVA infection
Mice were infected with 1–2×109 CFU of Citrobacter rodentium or Citrobater rodentium-OVA. For Citrobater rodentium-OVA infection mice were kept on 100 mg/L Kanamycin in drinking water to ensure the OVA expressing plasmid was not lost. Non- infected control mice were also kept on Kanamycin. Infection was monitored by plating feces onto McConckay agar plates. Nine days post infection 2 × 106 naive OT-II cells were adoptively transferred and mice subjected to OVA gavage as described above.
SFB colonization
SFB was obtained from frozen stocks of cecal contents (kept at −80 °C for less than 6 months) of mice monocolonized with SFB, which were diluted in PBS (2 ml/cecum) and passed through a 70 μm mesh. Mice were colonized by two gavages of 0.4 ml of cecal content preparation, 24 h apart. Colonization was verified by real time PCR of fecal DNA (Quick-DNA Fecal/Soil microbe miniprep kit, Zymo research, Cat. No. D6010). PCR was performed in presence of Power SYBR Green PCR master mix (Applied Biosystems, 4367659) on a Quant Studio 3 PCR machine (Applied Biosystems), using SFB specific 16S primers (fwd: GACGCTGAGGCATGAGAGCAT, rev: GACGGCACGGATTGTTATTCA) The SFB Ct value was normalized by the Ct obtained in the PCR using universal bacterial 16S primers (fwd: ACTCCTACGGGAGGCAGCAGT, rev: ATTACCGCGGCTGCTGGC).
Lymph node surgery
Mice were prepared and sutured and monitored postoperatively as for CT-OVA injection. To expose the ileal and cecal lymph nodes, the cecum was gently pulled out using cotton tipped applicators soaked in saline. To expose the upper duodenal lymph nodes, the duodenum was shifted to the left just under the pylorus; the same loop was shifted to the right to expose the distal duodenal gLNs. Lymph nodes were removed by holding onto the lymph node with tweezers and gently pulling it while slowly and closely cutting around the node with microsurgical scissors, such that no bleeding occurred. For sham operation the cecum was pulled out and the lymph nodes exposed for 2 min. The cecum was then placed back into its original position and the peritoneal cavity filled with 0.5 ml pre-warmed saline. Mice were allowed to recover from surgery for twelve days before colonization with SFB (Jax mice), nineteen days before adoptive transfer of 7B8tg cells, and two weeks before OVA gavage (oral tolerance).
S. venezuelensis passage and infection
S.venezuelensis was maintained in Wistar rats by subcutaneous infection with 30,000 larvae. Day 6–8 p.i. the cecum containing eggs was harvested and spread on Whatman paper which was placed into a beaker with water at 28 °C. The hatching larvae were collected over 4 days and the cycle re-initiated. Mice were infected subcutaneously with 700 larvae/ mouse. Adult worm load was assessed in total epithelial scrapes of the gut.
Alum immunization and airway challenge
Seven days after oral OVA administration, 4 μg of endotoxin-free OVA antigen adsorbed to 40 μl Imject™ Alum Adjuvant (Fisher Scientific) was injected i.p. in a final volume of 400 μl made up with PBS. Immunization was repeated after 7 days. To induce airway inflammation, mice were anesthetized and intranasally administered 10 μg of sterile OVA grade VI in 50 μl PBS (25 μl per nostril) on days 14, 17 and 21 after the first i.p. immunization. Total IgE was measured to confirm previous infection with S. venezuelensis (250–350 ng/ml in plasma compared to 5–10 ng in plasma of uninfected mice) 18.
Bronchoalveolar Lavage (BAL), lung histology and infiltrate analysis by flow cytometry
Mice were anesthetized by i.p. injection of 0.35 ml of 2.5% avertin (Sigma), the trachea was cannulated and lungs were lavaged once with 0.5 ml and then 1.0 ml PBS. Total BAL cells were counted after erythrocyte lysis and stained for FACS analysis. Lungs were perfused via the right ventricle with 10 ml saline to wash out residual blood. One lobe was digested in 400 U/ml collagenase D/HBSS and processed for FACS analysis. Eosinophils were determined as CD45+SSAhiMHCII–CD11b+Ly6GintSiglecF+ and DCs as CD45+MHCII+CD11c+CD64–SiglecF–.
Anti-OVA IgG1 and total IgE ELISAs
ELISAS were performed as described previously18.
Statistical analysis
Statistical analysis was performed in GraphPad Prism 7.0 software. Error bars indicate standard error of the mean (S.E.M.). Multivariate data was analyzed by applying one-way ANOVA and Tukey’s multiple comparison post hoc test, comparison between two treatment conditions by one-tailed unpaired Student’s t-test. Survival curve was analyzed by Mantel-Cox, Gehan-Breslow-Wilcoxon and Logrank. A P value of less than 0.05 was considered significant.
Data availability statement
Source data for all figures are provided with the paper. For RNA-seq experiments the raw data used in this study is available at Sequence Read Archive (SRA) under accession code: SRP166887. The processed data generated here can be obtained at Gene Expression Omnibus database under the accession code: GSE121811.
Extended Data
Extended Data Figure 1. Extended analysis of gLN connectivity and drainage in the peritoneal cavity.
a-e, 3D reconstruction of mouse lymphatics (α-LYVE-1) after solvent clearing (iDISCO+) and light sheet microscopy of the central chain of gLNs connected to the duodenum via afferent vessels (a), the D-gLNs with respect to liver and pancreas (co-stained with α-insulin) (b) and distal intestine gLNs connected to the ileum (c) individually dissected peritoneal gLNs (d) of SPF C57BL/6 mice, or all peritoneal gLNs from GF and SPF C57BL/6 mice (e). D/Duo=duodenum, whereby D1 portal LNs and D2 distal duodenum gLNs, J/Jej=jejunum, I/Ile=ileum, C1=cecal- colonic gLN, C2=ascending colonic gLN. f-i, Fast green spreading upon injection into gLNs draining the duodenum (f), jejunum (g), ileum (h) and cecum-proximal colon from SPF C57BL/6 mice (i). j-m, Fast green spreading upon injection into the lymphatics in the muscularis of the duodenum (j), jejunum (k), ileum (l) and cecum (m) of 10 weeks old mice. Pictures taken up to 15 minutes after dye injection. n-q, 3D reconstruction of mouse lymphatics (α-LYVE-1) after solvent clearing (iDISCO+) and light sheet microscopy in the duodenum (n, o), ileum (p), and colon (q), highlighting the submucosal lymphatic network. The following specifics apply: n, Arrow denotes direction of lymph flow from intestine to gLNs; CD11c was revealed by using ItgaxVenus mice and staining against GFP.
Extended Data Figure 2. Imaging of the intestinal lymphatics and characterization of 3H-retinol absorption along the gut of SPF and GF mice.
(a) 3D reconstruction of mouse lymphatics (α-LYVE-1) after iDISCO+ of villi and submucosa along the intestine of GF and SPF C57BL/6 mice; lymphatics protruding into the villi (lacteals) are pointed out. b, Percentage of 3H-retinol absorption into lymph (L), portal vein serum (PV) or systemic serum (S) of mice 3 h after gavage with 1 μCi 3H-retinol in 100 μl olive oil with or without pre-treatment with 5 μl chylomicron formation inhibitor Pluronic L-81 3 h prior to gavage (n=4 per group). c, d, Percentage of 3H-retinol absorption into duodenum (c), or duodenal gLN (d) of C57BL/6 mice 8 h after gavage with 1 μCi 3H –retinol in 100 μl olive oil with or without pre-treatment with 5 μl chylomicron formation inhibitor Pluronic L-81 3h prior to gavage (n=8 or 9 per group as indicated). Data pooled from two independent experiments with 4 or 5 animals per group each. e-g, Percentage of 3H-retinol absorption into systemic plasma (e), indicated intestinal tissue (f) or gLN (g) from GF or SPF C57BL/6 mice 1 or 5 h after gavage with 1 μCi 3H –retinol in 100 μl olive oil (n= 3 per group). Data representative of two independent experiments. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA). g refers to GF ANOVA, s to SPF ANOVA, $ to 1 h time point, # to 5 h time point.
Extended Data Figure 3. Extended analysis of gLN stromal and dendritic cell differences according to gut segment drained.
a, Flow cytometry plot showing gating of FRCs and LECs for RNA-seq. Pre-gates indicated above. b-g, Bar graph depicting some differentially regulated pathways among LECs sorted from duodenal versus colonic gLN (b), volcano plot depicting differentially expressed genes among LECs sorted from ileal versus colonic gLNs or ileal versus duodenal gLN (c, d), bar graph depicting some differentially regulated pathways among FRCs sorted from duodenal versus colonic gLN (e), volcano plot depicting differentially expressed genes among FRCs sorted from ileal versus colonic gLNs or ileal versus duodenal gLNs (f, g) from SPF C57BL/6 mice identified by RNA-seq. Blue indicates metabolic, red immunity-related pathways or genes. h, Flow cytometry plot showing gating of CD103+CD11b+ and CD103+CD11b– DCs for RNA-seq. Pre-gates indicated above. i-l, Volcano plot depicting differentially expressed genes among CD103+CD11b+ (i, k) and CD103+CD11b– (j, l) DCs sorted from ileal versus colonic gLNs or ileal versus duodenal gLNs from SPF C57BL/6 mice identified by RNA-seq. m, n, Bar graph depicting some differentially regulated pathways among CD103+CD11b+ (m) and CD103+CD11b– (n) sorted from duodenal versus colonic gLN. Blue indicates metabolic, red immunity-related genes and pathways (i-n). o-q, Mean fluorescence intensity (MFI) of fluorescein isothiocyanate–positive boron-dipyrromethene–tagged aminoacetate (Aldefluor) in CD103+CD11b–(o), CD103+CD11b+ (p) CD8α+ and CD11b+ (q) DCs from gLNs from GF and SPF C57BL/6 mice (n=3) assessed by flow cytometry 30 min after the addition of substrate.. r, Ratio of CD103+CD11b+ and CD103+CD11b– DCs in gLNs from SPF and GF mice (n=4, representing 2 independent experiments).. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA). D=duodenum gLNs, J=jejunum, I=ileum, C1=cecal- colonic gLN, C2=ascending colonic gLN, CA/IL=caudal/iliac gLN.
Extended Data Figure 4. Extended analysis of CD4+ T cells in the different gLNs of SPF and GF mice and activation of OT-II CD45.1 cells in all gLNs upon OVA gavage.
a-e, Frequency of TCRβ+CD4+ among CD45+ (a), FOXP3+NRP1+ (b) and FOXP3+NRP1– (c) among TCRβ+CD4+, RORγt+ among FOXP3+NRP1– (d) and FOXP3–RORγt+ among TCRβ+CD4+ (e) cells in indicated gLN from GF or SPF C57BL/6 mice (n=4). Data representative of two independent experiments. f- i, Frequency of CD25+ among CD45.1+ (f) and CD45.1+ among TCRβ+CD4+ cells (g), total CD45.1+ cells (h) and representative CFSE dilution histogram (i) in indicated gLNs 64 h post adoptive transfer of 1 × 106 naïve CD45.1+ OT-II cells into SPF CD45.2 C57BL/6 host mice (n=3) after gavage of OVA 48 h and 24 h prior to analysis. D=duodenum gLNs, J=jejunum, I=ileum, C1=cecal- colonic gLN, C2=ascending colonic gLN, CA/IL=caudal/iliac gLN.
Extended Data Figure 5. Extended analysis of OT-II CD45.1 and dendritic cells upon OVA administration in different gLNs of SPF and GF mice.
a, b, Frequency of CD45.1+ among TCRβ+CD4+ cells (a) and flow cytometry plots for gLN FOXP3 and CD25 expression (b) in indicated gLNs 64 h post adoptive transfer of 1 × 106 naïve CD45.1+ OT-II cells into GF and SPF CD45.2 C57BL/6 host mice (n=4) and after gavage of OVA 48 h and 24 h prior to analysis. c-e, 125I recovery in indicated gLNs (b), intestinal tissue (c) and plasma (d) of GF and SPF C57BL/6 mice (n=3 per time point and group) 1 h or 5 h after gavage with 4 × 106 CPM 125I-OVA in 50 mg cold OVA. f, g, Division index of CD45.1+ (f) and frequency of CD25+ among CD45.1+ (g) cells in indicated gLNs as in a, b. h-m, Frequency of CD45.1+ among TCRβ+CD4+ (h), division index of CD45.1+ (i), total CD45.1+ count (j), frequency of FOXP3hi among CD45.1+(k), representative flow cytometry plot for FOXP3hi and total FOXP3 gating amongst CD3+CD4+ in D-LN (l) and frequency of FOXP3hi among CD45.2+ TCRβ+CD4+ cells (m) in indicated LN or spleen 64 h post adoptive transfer of 1 × 106 naïve CD45.1+ OT-II cells into SPF CD45.2 C57BL/6 host mice (n=3) and after intravenous injection of OVA 48 h prior to analysis. n, Frequency of IL-12/23 p40+ cells among CD103+CD11b– DCs in indicated gLNs of GF and SPF C57BL/6 mice (n=4 per group). # indicates fewer than 200 cells were recovered. D=duodenum, whereby D1 portal LNs and D2 distal duodenum gLNs, J=jejunum, I=ileum, C1=cecal- colonic gLN, C2=ascending colonic gLN, C=colonic, CA/IL=caudal/iliac gLNs, ing=inguinal LN. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA).
Extended Data Figure 6. Extended analysis of OT-II CD45.1 cell seeding and fate in gLNs upon intestinal OVA injection or infection.
a-h, Frequency of FOXP3+RORγt– (a), FOXP3+RORγt+ (b), FOXP3–RORγt+ (c), FOXP3+CD25+ (d), FOXP3–CD25+ (e) among CD45.1+ OT-II cells, frequency of CD45.1+ OT-II among TCRβ+CD4+ cells (f), division index (g) and total TCRβ+CD4+ counts (h) in indicated gLN of mice 48 h post duodenal or ileal injection with OVA or OVA and CT, performed 16 h after adoptive transfer of 1 × 106 naïve OT-II cells into SPF CD45.2 C57BL/6 host mice. n=3. i-l, Survival curve (sham n=5, duo n=6, ile n=8) (i), and frequency in ileal lamina propria of FOXP3–RORγt+CD45.1+ (j) FOXP3–RORγt+CD45.1+ (k) and FOXP3+RORγt–CD45.1+ (l) among TCRβ+CD4+ cells (n=4 per group) of mice infected with Stm-OVA 9 days after duodenal or ileal injection with OVA-CT versus sham operation and 10 days post adoptive transfer of 1 × 105 naïve OT-II cells into SPF CD45.2 C57BL/6 host mice. j-l Intestines harvested 48 h post infection *P < 0.05 (Mantel-Cox and Gehan-Breslow-Wilcoxon, m), ##P < 0.005 (Logrank, a), *P < 0.05 (ANOVA, b-d). m-p, Flow cytometry plot of CD25+ and CFSE (m) frequency of CD45.1+ OT-II among TCRβ+CD4+ cells (n) total CD45.1+ counts (o) and CD45.1+ division index (p) in indicated gLNs 64 h post adoptive transfer of 1 × 106 naïve CD45.1+ OT-II cells into SPF CD45.2 C57BL/6 mice infected with Citrobacter-OVA 9 days before and after gavage of OVA or PBS 48 h and 24 h prior to analysis (n=5). Data representative of two independent experiments. q-u, Frequency of total FOXP3+ (q), FOXP3–CD25+ (r), FOXP3–RORγt+CD45.1+ (s) among CD45.1+, CD45.1+ among TCRβ+CD4+ cells (t) FOXP3–RORγt+CD45.2+ among TCRβ+CD4+ cells (u) in indicated gLNs 64 h post adoptive transfer of 1 × 106 naïve CD45.1+ OT-II cells into SPF CD45.2 C57BL/6 host mice infected or not with Citrobacter 9 days before and after gavage of OVA 48 h and 24 h prior to analysis (n=5). ns=not significant in ANOVA or one tailed t-test comparing gLNs of infected versus non-infected mice. *P < 0.05, **P < 0.01 and ***P < 0.005 (ANOVA), #P < 0.05 (t-test) D=duodengLNs, J=jejunal gLN, I=ileal gLN, C1=cecal- colonic gLN, C2=ascending colonic gLN, IL/CA=iliac/caudal gLNs
Extended Data Figure 7. Extended analysis of CD45.1+ 7B8tg cell fate in gLNs and gut upon distal gLN removal.
a-c, Representative flow cytometry plots of RORγt+ and CD45.1+ frequency among CD3+CD4+ cells in indicated gLN of sham or ΔicLN mice as quantified in Fig. 3g (a), and frequency of Vβ14+CD45.1+ among CD3+CD4+ (b) and RORγt+ among Vβ14+CD45.1+ (c) cells in indicated gLN of mice with sham operation (n=12) or surgical removal of the I- and C1-gLN (ΔicLN, n=14) 64 h (day 3, Fig. 3f) after adoptive transfer of 4 × 105 naïve SFB specific CD45.1+ 7B8tg cells into recently SFB-colonized SPF CD45.2 C57BL/6 Jax mice. Graph represents pooled data from 3 independent experiments with n=4–5 per group each. d, e, Frequency of Vβ14+CD45.1+ among CD3+CD4+ (d) and RORγt+ among Vβ14+CD45.1+ (e) cells in indicated gLN of Taconic sham or ΔicLN mice (n=4) 64 h after adoptive transfer of 4 × 105 naïve SFB specific CD45.1+ cells. Note that SFB is a stable member of the microbial community in Taconic C57BL/6 mice through parental transmission. f, g, Total numbers of CD45+ cells in C2-gLN of recently SFB-colonized SPF mice (f) or Taconic mice (g) at point of harvest, h-n, Frequency of Vβ14+CD45.1+ IL17a+ (h), Vβ14+CD45.1+RORγt+ (i), or Vβ14+CD45.1+ among CD3+CD4+ cells (j), and RORγt+ among Vβ14+CD45.1+ cells (k) or IL17a+ among RORγt+Vβ14+CD45.1+ cells (l) cells in lamina propria of indicated gut segment, and Vβ14+CD45.1+ among CD3+CD4+ (m), RORγt+ among Vβ14+CD45.1+ (n) in gLNs of sham or ΔicLN Jax mice (n=4) 7 days after adoptive transfer of 5000 naïve CD45.1+ 7B8tg cells into recently SFB-colonized SPF CD45.2 C57BL/6 Jax mice. o-s, Frequency of Vβ14+CD45.1+ among CD3+CD4+ (o), RORγt+ among Vβ14+CD45.1+ (p) and IL17a+ among RORγt+Vβ14+CD45.1+ (q) cells in indicated lamina propria, or Vβ14+CD45.1+ among CD3+CD4+ (r), RORγt+ among Vβ14+CD45.1+ (s) in gLNs of sham or ΔicLN Taconic mice 7 days after adoptive transfer of 5000 naïve CD45.1+ 7B8tg cells (n=4). D=duodenum, whereby D1 portal LNs and D2 distal duodenum gLNs, J=jejunum, I=ileum, C1=cecal- colonic gLN, C2=ascending colonic gLN, C=colon. # indicates fewer than 200 cells were recovered. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA).
Extended Data Figure 8. Characterization of effect of selective gLN removal on lymph flow, SFB colonization and oral tolerance.
a, Pictures of Fast Green tracing of ileal lymphatic drainage to the gLNs of SPF C57BL/6 mice, 5 min after injection of 3 μl fast green in sham operated (left column, biological duplicates, see also Extended Data Fig. 2g) or ΔicLN (middle and zoomed in right column, biological triplicates) mice 3 weeks after surgery. Red arrows indicate site of injection, jejunal (J), ileal (I) and cecal-colonic (C1) gLNs are indicated. b, c, Relative quantification of SFB specific 16S in luminal contents of indicated gut segment from recently SFB-colonized SPF C57BL/6 Jax mice (b, n=13 per group from 4 independent experiments) or parentally-colonized Taconic (c, n=7 per group from 4 independent experiments) mice after sham or ΔicLN surgery. *P < 0.05, **P < 0.01 and ***P < 0.005 (ANOVA). d-i, Total eosinophils (d) and DCs (e) in BALF and frequency of eosinophils (f) and DCs (g) among CD45+ cells in lungs, total IgE (h) and anti OVA-IgG1 (i) serum levels in SPF C57BL/6 mice subjected to oral tolerance as in scheme Fig. 4k but without S.v. infection 14 days after sham surgery (n=7), ileal and cecal LN removal (ΔicLN, n=5) or duodenal LN removal (ΔdLN, n=6). Data representative of two independent experiments. Not significant (ns) in ANOVA.
Extended Data Figure 9. Extended analysis of gLN swelling, restructuring, DC and CD4+ T cell subset frequencies upon S. venezuelensis infection.
a, Indicated gLNs positions in non-infected (N.I.) SPF C57BL/6 mice or mice infected with 700 S. venezuelensis (S.v.) larvae 8 days prior to harvest. b, 3D reconstruction of vasculature (α-CD31) after solvent clearing (iDISCO+) and light sheet microscopy of indicated gLNs from non-infected (N.I.) SPF C57BL/6 mice or SPF C57BL/6 mice infected with 700 S. venezuelensis (S.v.) larvae 14 days prior to harvest, bar=500 μm. c, Quantification of mature S.v. worms in gut epithelium 8 days after infection with 700 S.v. larvae. n=8, pooled from two independent experiments. d-g, Frequency of CD11c+ among CD45+ (d), and CD11b+ (e), CD8α+(f) among MHCIIintCD11c+ cells or GATA3+FOXP3+ among TCRβ+CD4+ cells (g) in non-infected (N.I.) SPF C57BL/6 mice or mice infected with 700 S. venezuelensis (S.v.) larvae 8 days prior to harvest. (n=5). Data representative of 3 independent experiments. h-l, frequency of CD103+CD11b+ (h), CD103+CD11b– (i), or CD103–CD11b+ (j) among MHCIIintCD11c+ DCs, and of GATA3+FOXP3– (k) or GATA3+FOXP3+ (l) among TCRβ+CD4+ cells in denoted gLNs of SPF C57BL/6 mice infected with 700 S. venezuelensis (S.v.) larvae 21–50 days prior to harvest, as indicated (n=4 per time point) or non-infected (N.I., harvested day 21) SPF C57BL/6 mice. D=duodenum, whereby D1 portal LNs and D2 distal duodenum gLNs, J=jejunum, I=ileum, C1=cecal- colonic gLN, C2=ascending colonic gLN, C=colon. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA).
Extended Data Figure 10. Extended analysis of OT-II CD45.1 cell seeding, activation and fate and oral tolerance upon S. venezuelensis infection.
a, Representative flow cytometry plot of FOXP3+ and CD25+ CD45. 1 OT-II cells in gLNs 8 days after infection of mice with S.v. larvae or N.I.b-e, Frequency of total FOXP3–CD25+ among CD45.1+ (b), CD45.1+ among TCRβ+CD4+ (c) and total CD45.1+ cells (d) and division index (e) in indicated gLNs 64 h post adoptive transfer of 1 × 106 naïve CD45.1+ OT-II cells into CD45.2 SPF C57BL/6 mice (n=4) infected with S.v. or N.I. 8 days and gavaged with OVA 48 h and 24 h prior to analysis. Data representative of two independent experiments. D=duodenum, whereby D1 portal LNs and D2 distal duodenum gLNs, J=jejunum, I=ileum, C1=cecal- colonic gLN, C2=ascending colonic gLN. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA). f, g, Total DCs in bronchoalveolar lavage fluid (BALF) (f) and frequency of DCs among CD45+ cells in lung tissue (g) from N.I SPF C57BL/6 mice or mice infected with S.v. during antigen feeding (+OVA groups) or no feeding (-OVA groups), at 21 d after first immunization with OVA-alum (n=13 for +OVA groups, n=10 for –OVA groups). Data are pooled from two independent experiments. *P < 0.05, **P < 0.01 and ***P < 0.005 (one-tailed t-test or ANOVA).
Supplementary Material
Acknowledgements
We thank all Mucida Lab members past and present for assistance in experiments, fruitful discussions and critical reading of the manuscript, Bernardo Reis for figure preparations, Fanny Matheis for propagating S. venezuelensis, Aneta Rogoz and Sara Gonzalez for the maintenance of mice, Tomiko Rendon and Beatriz Lopez for genotyping, Kristie Gordon and Kalsang Chosphel for assistance with cell sorting, the Rockefeller University Bio-imaging Research Center for assistance with microscopy and image analysis, the Rockefeller University Genomics Center for RNA sequencing and Rockefeller University employees for continuous assistance. We also thank Michel Nussenzweig, Gabriel Victora (Rockefeller University) and Juan Lafaille (NYU) and their respective lab members for fruitful discussions and suggestions. We are indebted to Dan Littman and Mo Xu (NYU) for generously providing 7B8tg mice and feces from SFB monocolonized mice; and Stephen Galli and Kazufumi Matsushita (Stanford University) for providing S. venezuelensis and guidance on how to maintain it. This work was supported by a Swiss National Science Foundation postdoctoral fellowship and University of Chicago start-up funds (D.E.); a CAPES fellowship (M.C.C.C.); NIH F31 Kirchstein Fellowship, Philip M. Levine Fellowship, and Kavli Neural Systems Institute Graduate Fellowship (P.A.M.); the Leona M. and Harry B. Helmsley Charitable Trust, the Crohn’s & Colitis Foundation of America Senior Research Award, the Burroughs Wellcome Fund PATH Award, National Institute of Health grants R21AI31188, R01DK113375 and R01DK093674 (D.M.).
Footnotes
Competing interests
The authors declare no competing financial interests.
References
- 1.Honda K & Littman DR The microbiota in adaptive immune homeostasis and disease. Nature 535, 75–84, doi: 10.1038/nature18848 (2016). [DOI] [PubMed] [Google Scholar]
- 2.Yang Y et al. Focused specificity of intestinal TH17 cells towards commensal bacterial antigens. Nature 510, 152–156, doi: 10.1038/nature13279 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Xu M et al. c-MAF-dependent regulatory T cells mediate immunological tolerance to a gut pathobiont. Nature 554, 373–377, doi: 10.1038/nature25500 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Mucida D et al. Oral tolerance in the absence of naturally occurring Tregs. The Journal of clinical investigation 115, 1923–1933 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Randolph GJ, Ivanov S, Zinselmeyer BH & Scallan JP The Lymphatic System: Integral Roles in Immunity. Annual review of immunology 35, 31–52, doi: 10.1146/annurev-immunol-041015-055354 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Fonseca DM et al. Microbiota-Dependent Sequelae of Acute Infection Compromise Tissue-Specific Immunity. Cell 163, 354–366, doi: 10.1016/j.cell.2015.08.030 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cording S et al. The intestinal micro-environment imprints stromal cells to promote efficient Treg induction in gut-draining lymph nodes. Mucosal immunology 7, 359–368, doi: 10.1038/mi.2013.54 (2014). [DOI] [PubMed] [Google Scholar]
- 8.Worbs T et al. Oral tolerance originates in the intestinal immune system and relies on antigen carriage by dendritic cells. J Exp Med 203, 519–527, doi: 10.1084/jem.20052016 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Van den Broeck W, Derore A & Simoens P Anatomy and nomenclature of murine lymph nodes: Descriptive study and nomenclatory standardization in BALB/cAnNCrl mice. Journal of immunological methods 312, 12–19, doi: 10.1016/j.jim.2006.01.022 (2006). [DOI] [PubMed] [Google Scholar]
- 10.Carter PB & Collins FM The route of enteric infection in normal mice. J Exp Med 139, 1189–1203 (1974). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Houston SA et al. The lymph nodes draining the small intestine and colon are anatomically separate and immunologically distinct. Mucosal immunology 9, 468–478, doi: 10.1038/mi.2015.77 (2016). [DOI] [PubMed] [Google Scholar]
- 12.Gautreaux MD, Deitch EA & Berg RD Bacterial translocation from the gastrointestinal tract to various segments of the mesenteric lymph node complex. Infection and immunity 62, 2132–2134 (1994). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Veenbergen S et al. Colonic tolerance develops in the iliac lymph nodes and can be established independent of CD103(+) dendritic cells. Mucosal immunology 9, 894–906, doi: 10.1038/mi.2015.118 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Pezoldt J et al. Neonatally imprinted stromal cell subsets induce tolerogenic dendritic cells in mesenteric lymph nodes. Nature communications 9, 3903, doi: 10.1038/s41467-018-06423-7 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hammerschmidt SI et al. Stromal mesenteric lymph node cells are essential for the generation of gut-homing T cells in vivo. J Exp Med 205, 2483–2490, doi:jem.20080039 [pii] 10.1084/jem.20080039 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Durai V & Murphy KM Functions of Murine Dendritic Cells. Immunity 45, 719–736, doi: 10.1016/j.immuni.2016.10.010 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Esterhazy D et al. Classical dendritic cells are required for dietary antigen-mediated induction of peripheral Treg cells and tolerance. Nature immunology 17, 545–555, doi: 10.1038/ni.3408 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Atarashi K et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331, 337–341, doi: 10.1126/science.1198469 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Sano T et al. An IL-23R/IL-22 Circuit Regulates Epithelial Serum Amyloid A to Promote Local Effector Th17 Responses. Cell 163, 381–393, doi: 10.1016/j.cell.2015.08.061 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Atarashi K et al. Th17 Cell Induction by Adhesion of Microbes to Intestinal Epithelial Cells. Cell 163, 367–380, doi: 10.1016/j.cell.2015.08.058 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hadis U et al. Intestinal tolerance requires gut homing and expansion of FoxP3+ regulatory T cells in the lamina propria. Immunity 34, 237–246, doi: 10.1016/j.immuni.2011.01.016 (2011). [DOI] [PubMed] [Google Scholar]
- 22.Balmer ML et al. The liver may act as a firewall mediating mutualism between the host and its gut commensal microbiota. Sci Transl Med 6, 237ra266, doi: 10.1126/scitranslmed.3008618 (2014). [DOI] [PubMed] [Google Scholar]
- 23.Bouziat R et al. Reovirus infection triggers inflammatory responses to dietary antigens and development of celiac disease. Science 356, 44–50, doi: 10.1126/science.aah5298 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mukai K, Karasuyama H, Kabashima K, Kubo M & Galli SJ Differences in the Importance of Mast Cells, Basophils, IgE, and IgG versus That of CD4(+) T Cells and ILC2 Cells in Primary and Secondary Immunity to Strongyloides venezuelensis. Infection and immunity 85, doi: 10.1128/IAI.00053-17 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Tussiwand R et al. Klf4 expression in conventional dendritic cells is required for T helper 2 cell responses. Immunity 42, 916–928, doi: 10.1016/j.immuni.2015.04.017 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Source data for all figures are provided with the paper. For RNA-seq experiments the raw data used in this study is available at Sequence Read Archive (SRA) under accession code: SRP166887. The processed data generated here can be obtained at Gene Expression Omnibus database under the accession code: GSE121811.