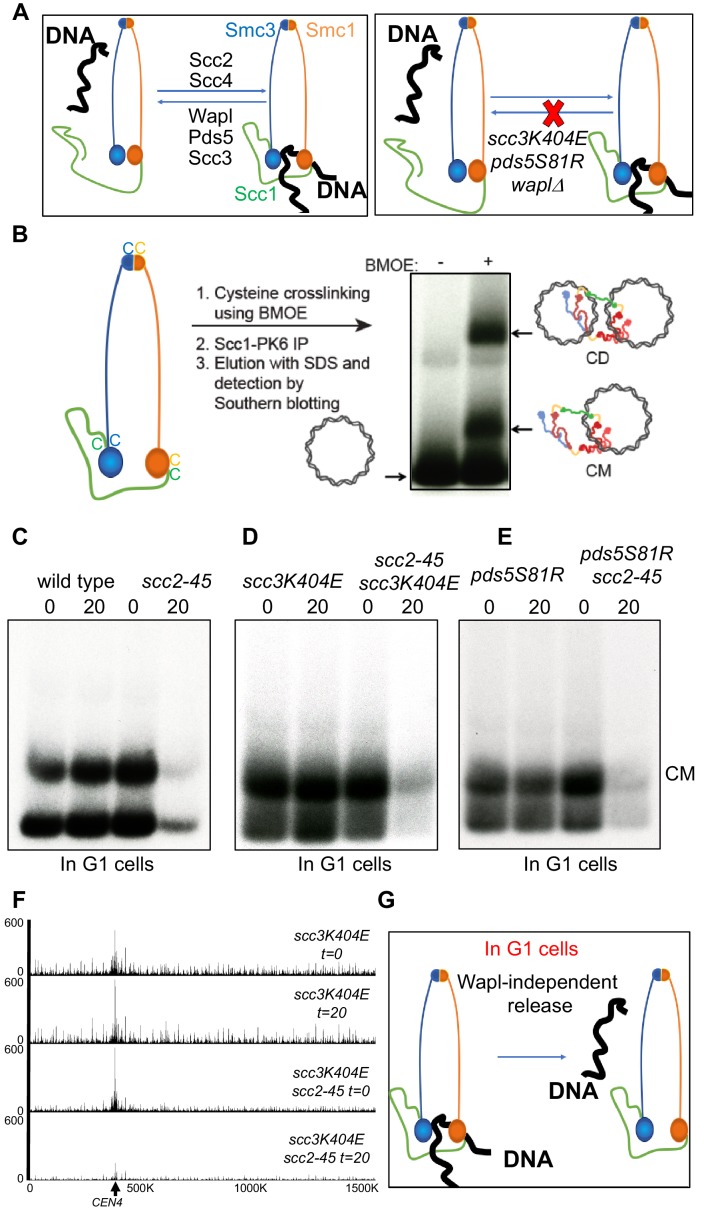
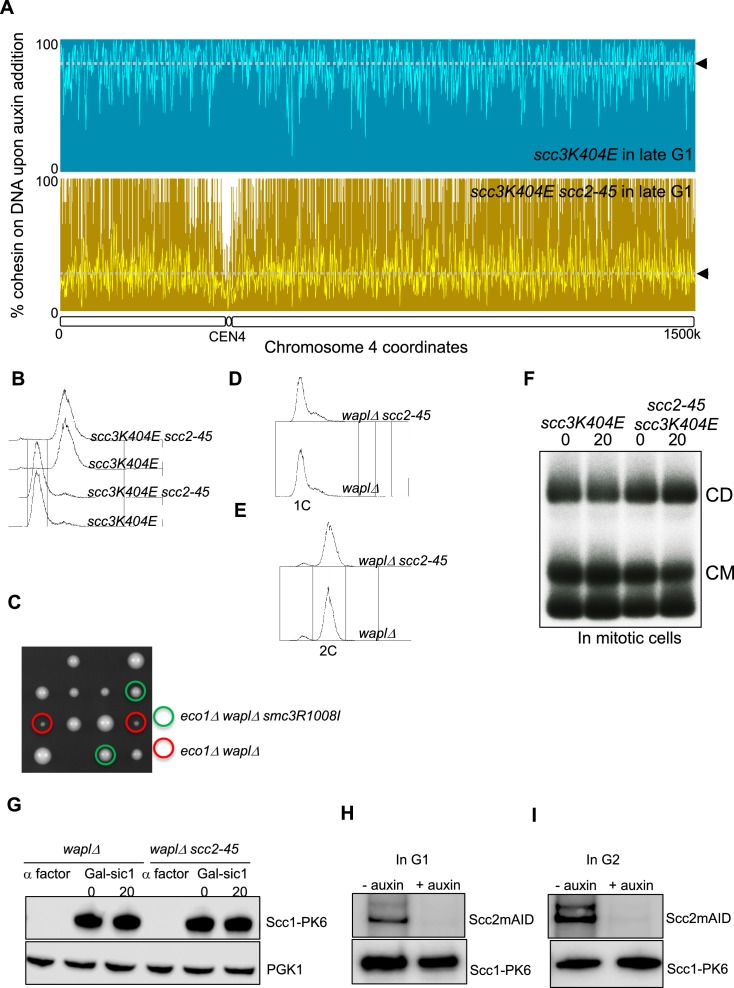
Figure 1. A Wapl-independent activity releases cohesin from chromosomes in G1 cells.
(A) Cohesin’s association with DNA is regulated by two opposing activities: Scc2-Scc4 complex loads cohesin onto DNA, while Pds5, Scc3 and Wapl constitute the releasing activity that releases cohesin from DNA by opening the Smc3-Scc1 interface. Mutations in Scc3 (scc3K404E) Pds5 (pds5S81R) and deletion of the WAPL gene abrogate Wapl mediated releasing activity and lead to cohesin’s stable association with DNA. (B) Schematic of the mini-chromosome IP assay: 6C strain (K23889) with cysteine pairs at all three ring subunit interfaces (2C Smc3: E570C S1043C, 2C Smc1: G22C K639C and 2C Scc1 C56 A547C) and carrying a 2.3 kb circular mini-chromosome was subjected to in vivo crosslinking with BMOE. DNAs associated with cohesin immune-precipitates (Scc1-PK6) were denatured with SDS and separated by agarose gel electrophoresis. Southern blotting reveals two forms of DNA unique to cells treated with BMOE: CMs (cohesin entrapping individual mini-chromosomes) and CDs (cohesin entrapping a pair of sister mini-chromosomes). (C) WT (K23972) and scc2-45 (K25238) 6C strains were arrested in late G1 by overexpression of nondegradable Sic1 at 25°C as described in Materials and Methods. The cultures were shifted to 37°C for 20 min, aliquots drawn before (0) and after (20) temperature shift (to inactivate Scc2) were subjected to mini-chromosome IP. (D) scc3K404E (K25313) and scc3K404E scc2-45 (K25316) 6C strains were arrested in late G1. The cultures were shifted to 37°C for 20 min, aliquots drawn before (0) and after (20) temperature shift (to inactivate Scc2) were subjected to mini-chromosome IP. Also see S1B. (E) pds5S81R (K25311) and pds5S81R scc2-45 (K25312) 6C strains were arrested in late G1. The cultures were shifted to 37°C for 20 min, aliquots drawn before (0) and after (20) temperature shift (to inactivate Scc2) were subjected to mini-chromosome IP. (F) scc3K404E (K25313) and scc3K404E scc2-45 (K25316) strains were arrested in late G1. The cultures were shifted to 37°C for 20 min, aliquots drawn before (0) and after (20) temperature shift (to inactivate Scc2) were analysed by Calibrated-ChIP- sequencing (Scc1-PK). Cohesin profile along chromosome four is shown. Also see Figure 1—figure supplement 1. (G) Even in cells that lack Wapl mediated releasing activity, inactivation of Scc2 in G1 cells leads to release of DNA entrapped within cohesin rings. This suggests that an activity that is Wapl-independent is capable of releasing cohesin from DNA.