Abstract
Endothelin receptors (ETA and ETB) are G-protein-coupled receptors activated by endothelin-1 and are involved in blood pressure regulation. IRL2500 is a peptide-mimetic of the C-terminal tripeptide of endothelin-1, and has been characterized as a potent ETB-selective antagonist, which has preventive effects against brain edema. Here, we report the crystal structure of the human ETB receptor in complex with IRL2500 at 2.7 Å-resolution. The structure revealed the different binding modes between IRL2500 and endothelin-1, and provides structural insights into its ETB-selectivity. Notably, the biphenyl group of IRL2500 penetrates into the transmembrane core proximal to D2.50, thus stabilizing the inactive conformation. Using the newly-established constitutively active mutant, we clearly demonstrate that IRL2500 functions as an inverse agonist for the ETB receptor. The current findings will expand the chemical space of ETR antagonists and facilitate the design of inverse agonists for other class A GPCRs.
Subject terms: X-ray crystallography, Cardiovascular biology
Chisae Nagiri and Wataru Shihoya report the crystal structure of the human endothelin receptor ETB in complex with an ETB antagonist IRL2500, suggesting different binding modes between IRL2500 and endothelin-1, ETB’s endogenous ligand. This study provides structural insights into the specificity of ETB interaction with its ligands.
Introduction
Endothelin receptors (ETRs) are G-protein-coupled receptors (GPCR) activated by vasoactive peptide, endothelins1. Two ETR subtypes (ETA and ETB) are widely expressed in the vascular endothelium, brain, and other circulatory organs2,3. Endothelin-1 (ET-1) activates the both ETRs with sub-nanomolar affinities. The activation of the ETA receptor leads to potent and long-lasting vasoconstriction, whereas that of the ETB receptor induces nitric oxide-mediated vasorelaxation. Therefore, the up-regulation of ET-1 is related to circulatory-system diseases, including pulmonary arterial hypertension (PAH)4–7. Moreover, the autocrine and paracrine signaling functions of ET-1 through the ETA receptor play a critical role in tumor growth and survival8. Thus, ETR antagonists have been developed for the treatment of circulatory-system diseases and cancers6,7. Bosentan is the first orally-active ETR antagonist9,10, and is used to treat PAH. The ETB receptor is the prominent ETR subtype in the brain, with high expression levels in astrocytes11. Stimulation of the ETB receptor modulates astrocytic responses, indicating its important roles in regulating astrocytic functions12. The up-regulation of the astrocytic ETB receptor by ET-1 increases the vascular permeability and reduces the AQP4 levels, thereby aggravating vasogenic brain edema11. The application of ETB-selective antagonists may provide preventive effects against brain edema in the acute phase of brain insults13–16.
To date, most ETR antagonists have been developed based on bosentan17,18. The ETR antagonists that have been developed till now are mostly N-heterocyclic sulfonamides with similar structures and molecular weights, and non-sulfonamide antagonists (atrasentan, ambrisentan, darusentan, and enrasentan) still retain high similarities with each other and with the sulfonamides6. Since the ETR antagonists are chemically very similar7, and the expanded chemical space should be exploited. IRL2500 is a peptide-mimetic ETR antagonist developed based on the partial region of ET-119, not on bosentan. IRL2500 has been characterized as an ETB-selective antagonist with an IC50 value of 1.2 nM20, which shows higher affinity than that of bosentan. In an animal model, the intracerebroventricular administration of IRL2500 attenuated the cold injury-mediated brain edema and disruption of the blood–brain barrier, indicating the neuroprotective effect of IRL250014,15. Clarification of the IRL2500 binding mode would facilitate the expansion of the chemical space of ET agents.
We previously reported the crystal structures of the ETB receptor bound to ET-121 and bosentan22; however, both the binding mode and ETB-selectivity of IRL2500 remained to be elucidated. Here, we present the crystal structure of the ETB receptor in complex with IRL2500. This structure revealed the unique binding mode of IRL2500, which differs from those of ET-1 and bosentan. Structure-guided functional analyses clearly demonstrate that IRL2500 functions as an inverse agonist for the ETB receptor, and thus will provide the basis for the design of inverse agonists for other class A GPCRs.
Results
Overall structure
For crystallization, we used the previously established, thermostabilized ETB receptor (ETB-Y4)22,23. The IC50 value of IRL2500 for ETB-Y4 was similar to that for the wild type receptor in the TGFα shedding assay24 (Fig. 1a), suggesting that the thermostabilizing mutations minimally affect the IRL2500 binding. In contrast, the IC50 value of IRL2500 for the ETA receptor is over 3 μM (Fig. 1a), indicating that IRL2500 has over 100-fold ETB-selectivity, consistent with the previous pharmacological analysis20. To facilitate crystallization, we replaced the third intracellular loop (ICL3) of the receptor with minimal T4 Lysozyme25 (ETB-Y4-mT4L). Using in meso crystallization26, we obtained crystals of ETB-Y4-mT4L in complex with IRL2500 (Supplementary Fig. 1a, b). In total, 58 datasets were collected and merged by the data processing system KAMO27. Eventually, we determined the ETB structure in complex with IRL2500 at 2.7 Å resolution, by molecular replacement using the antagonist-bound ETB structure (PDB code: 5X93) (Table 1).
Fig. 1.
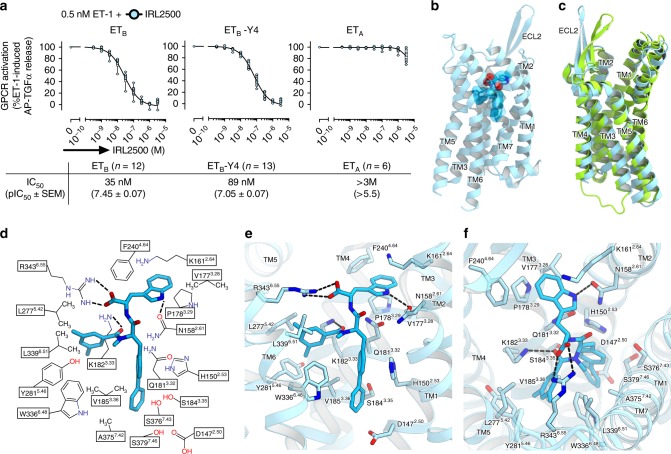
Overall structure of the IRL2500-bound ETB receptor. a The effect of IRL2500 on the ET-1 (0.5 nM)-induced release of AP-TGFα in HEK293 cells expressing the endothelin receptors. For each experiment, the AP-TGFα release response in the absence of IRL2500 is set at 100%. Concentration-response data are displayed as means ± SEM (standard error of the mean) from six to thirteen independent experiments and the pIC50 values are from the indicated number of independent experiments. b The overall structure of the IRL2500-bound ETB receptor. The receptor is shown as a sky blue ribbon model. IRL2500 is shown as a deep sky blue stick model with a transparent surface model. c Superimposition of the IRL2500-bound and ligand-free ETB structures (PDB code: 5GLI), colored sky blue and light green, respectively. d Schematic representation of the interactions between ETB and IRL2500 within 4.5 Å. The dashed lines show hydrogen bonds. e, f Binding pocket for IRL2500, viewed from the membrane plane (e) and the extracellular side (f). The receptor is shown in a sky blue ribbon representation. IRL2500 and receptor residues involved in ligand binding are shown as sticks, colored deep sky blue and sky blue, respectively. The dashed lines show hydrogen bonds
Table 1.
Data collection and refinement statistics
| IRL2500-ETB | |
|---|---|
| Data collection | |
| Space group | I422 |
| Cell dimensions | |
| a, b, c (Å) | 110.0, 110.0, 291.7 |
| α, β, γ (°) | 90, 90, 90 |
| Resolution (Å)a | 43.88–2.70 (2.797–2.70) |
| R meas a | 0.295 (3.946) |
| 〈I/σ(I)〉a | 10.74 (1.08) |
| CC1/2a | 0.995 (0.798) |
| Completeness (%)a | 99.66 (99.35) |
| Redundancya | 19.3 (19.6) |
| Refinement | |
| Resolution (Å) | 43.88–2.70 |
| No. reflections | 25,069 |
| Rwork/Rfree | 0.2216/0.2653 |
| No. atoms | |
| Protein | 3258 |
| Ligand | 43 |
| Water/ion/lipid | 240 |
| Averaged B-factors (Å2) | |
| Protein | 83.41 |
| Ligand | 51.4 |
| Water/ion/lipid | 94.45 |
| R.M.S. deviations from ideal | |
| Bond lengths (Å) | 0.003 |
| Bond angles (°) | 0.65 |
| Ramachandran plot | |
| Favored (%) | 99.26 |
| Allowed (%) | 0.74 |
| Outlier (%) | 0 |
aValues in parentheses are for highest-resolution shell
The overall structure consists of the canonical 7 transmembrane helices (TM), the amphipathic helix 8 at the C-terminus (H8), and two antiparallel β-strands in the extracellular loop 2 (ECL2), as in the previously determined ETB structures (Fig. 1b). The IRL2500-bound structure is similar to the bosentan-bound structure, rather than the ET-1-bound structure (R.M.S.D. values for Cα atoms = 1.34 and 1.95 Å, respectively), reflecting the inactive conformation. We observed a remarkable difference in the conformation of ECL2. The β strands are opened up by 9 Å, as compared with those in the ligand-free structure (Fig. 1c and Supplementary Fig. 2a), and are the widest among the peptide-activated class A GPCRs (Supplementary Fig. 2b). This conformation is facilitated by the crystal packing (Supplementary Fig. 2c), and is not a consequence of IRL2500 binding. This structural feature indicates the ECL2 is highly flexible in the inactive conformation of the ETB receptor, to capture the large peptide ligand endothelin.
IRL2500 binding site
We first describe the IRL2500 binding mode. IRL2500 consists of a tryptophan, a biphenyl group and a 3,5-dimethylbenzoyl group19. The biphenyl group forms a peptide bond with the tryptophan, and a peptoid bond with the dimethylbenzoyl group (Fig. 1d). IRL2500 binds to the transmembrane binding cleft exposed to the extracellular side, with a clear electron density (Supplementary Fig. 3a, b). The carboxylate group of the tryptophan moiety in IRL2500 forms salt bridges with R3436.55 (superscripts indicate Ballesteros–Weinstein numbers28) (Fig. 1e, f). The tryptophan side chain of IRL2500 forms a hydrogen bond with the carbonyl group of the N1582.61 side chain and a cation–π interaction with the K1612.64 side chain, The tryptophan also forms extensive van der Waals interactions with V1773.28, P1783.29, and F2404.64. The dimethylbenzoyl group of IRL2500 forms van der Waals interactions with the hydrophobic pocket, and is surrounded by V1853.36, L2775.42, Y2815.46, W3366.48 and L3396.51. The biphenyl group penetrates deeply into the receptor core proximal to D1472.50, and forms van der Waals interactions with D1472.50, H1502.53, W3366.48, and S3767.43. Overall, the carboxylate of IRL2500 is specifically recognized by the positively charged residues of the ETB receptor, and the other moieties fill the space within the transmembrane binding pocket.
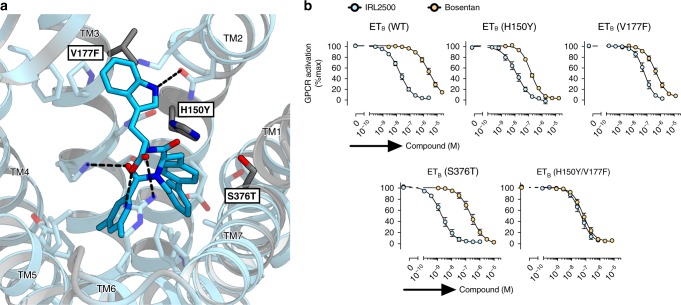
To elucidate the structural basis for the ETB-selectivity of IRL2500, we compared the residues constituting the IRL2500 binding site between the ETB and ETA receptors (Fig. 2a and Supplementary Fig. 4). Most of the residues are conserved, while H1502.53, V1773.28, and S3767.43 are substituted for the bulkier residues Y1292.53, F1613.28, and T3597.43 in the ETA receptor, respectively. These substitutions may cause steric clashes with the aromatic groups of IRL2500 and reduce its affinity. To investigate this hypothesis, we measured the IC50 values of IRL2500 for the H150Y, V177F, and S376T ETB receptor mutants. These mutants showed similar responses for ET-1 in the TGFα shedding assay (Supplementary Fig. 5a). The H150Y mutant showed a similar response for IRL2500, and the S376T mutant showed a 3-fold increased potency (that is, 3-fold smaller IC50value) of IRL2500. Only V177F showed a 6-fold decreased potency of IRL2500 (Fig. 2b and Table 2), suggesting that the F1613.28 in the ETA receptor sterically clashes with the tryptophan moiety of IRL2500. Moreover, the H150Y/V177F double mutant showed the further decreased potency of IRL2500 by about 1.5-fold as compared with that of the V177F mutant only (Fig. 2b and Table 2), suggesting that the two residues have some cooperativity in the ETA receptor. Notably, these mutations did not reduce (actually increased) the potency of the non-selective antagonist bosentan, showing that the two residues (H150Y and V177F) were selective to the IRL2500 inhibition. Taken together, Y1292.53 and F1613.28 in the ETA receptor may specifically clash with IRL2500, partially accounting for its ETB-selectivity. This is consistent with the results of the previous study, in which the replacement of the tryptophan moiety with the smaller moiety valine weakened its ETB-selectivity29.
Fig. 2.
Conservation of the IRL2500 binding site. a Sequence conservation of the IRL2500 binding site between ETA and ETB, mapped onto the IRL2500-bound structure. Conserved and non-conserved residues are colored sky blue and gray, respectively. The receptor residues involved in IRL2500 binding are shown as sticks. The dashed lines show hydrogen bonds. b Effects of IRL2500 and bosentan on the ET-1 (0.5 nM)-induced release of AP-TGFα in HEK293 cells expressing the mutant ETB receptors. For each experiment, the AP-TGFα release response in the absence of IRL2500 is set at 100%. Data are displayed as means ± SEM (standard error of the mean) from four to six independent experiments
Table 2.
Pharmacological characterization of mutant ETB receptors
| ETB--WT | H150Y | V177F | S376T | H150/V177F | ||
|---|---|---|---|---|---|---|
| n | 6 | 4 | 4 | 4 | 4 | |
| ET-1 | EC50 (nM) | 0.14 | 0.090 | 0.093 | 0.16 | 0.11 |
| pEC50 (mean ± SEM) | 9.85 ± 0.07 | 10.05 ± 0.10 | 10.03 ± 0.11 | 9.79 ± 0.04 | 9.97 ± 0.08 | |
| IRL2500 | IC50 (nM) | 21 | 18 | 130 | 6.1 | 180 |
| pIC50 (mean ± SEM) | 7.67 ± 0.09 | 7.74 ± 0.15 | 6.87 ± 0.13 | 8.22 ± 0.15 | 6.75 ± 0.15 | |
| KB (nM) | 2.4 | 0.78 | 7.8 | 0.63 | 12 | |
| pKB (mean ± SEM) | 8.62 ± 0.07 | 9.11 ± 0.12 | 8.11 ± 0.07 | 9.20 ± 0.08 | 7.93 ± 0.14 | |
| ∆KB | 1 | 3.0 | 0.30 | 4.0 | 0.20 | |
| ∆pKB (mean ± SEM) | 0 | 0.47 ± 0.11 | −0.53 ± 0.06 | 0.60 ± 0.03 | -0.70 ± 0.14 | |
| Bosentan | IC50 (nM) | 3,300 | 270 | 1100 | 680 | 270 |
| pIC50 (mean ± SEM) | 5.48 ± 0.11 | 6.57 ± 0.05 | 5.97 ± 0.11 | 6.17 ± 0.16 | 6.56 ± 0.11 | |
| KB (nM) | 400 | 12 | 70 | 71 | 17 | |
| pKB (mean ± SEM) | 6.39 ± 0.09 | 7.92 ± 0.13 | 7.16 ± 0.05 | 7.15 ± 0.10 | 7.78 ± 0.02 | |
| ∆KB | 1 | 35 | 6.1 | 5.8 | 26 | |
| ∆pKB (mean ± SEM) | 0 | 1.55 ± 0.12 | 0.79 ± 0.08 | 0.76 ± 0.11 | 1.41 ± 0.06 |
Comparison of the binding modes of IRL2500, ET-1, and bosentan
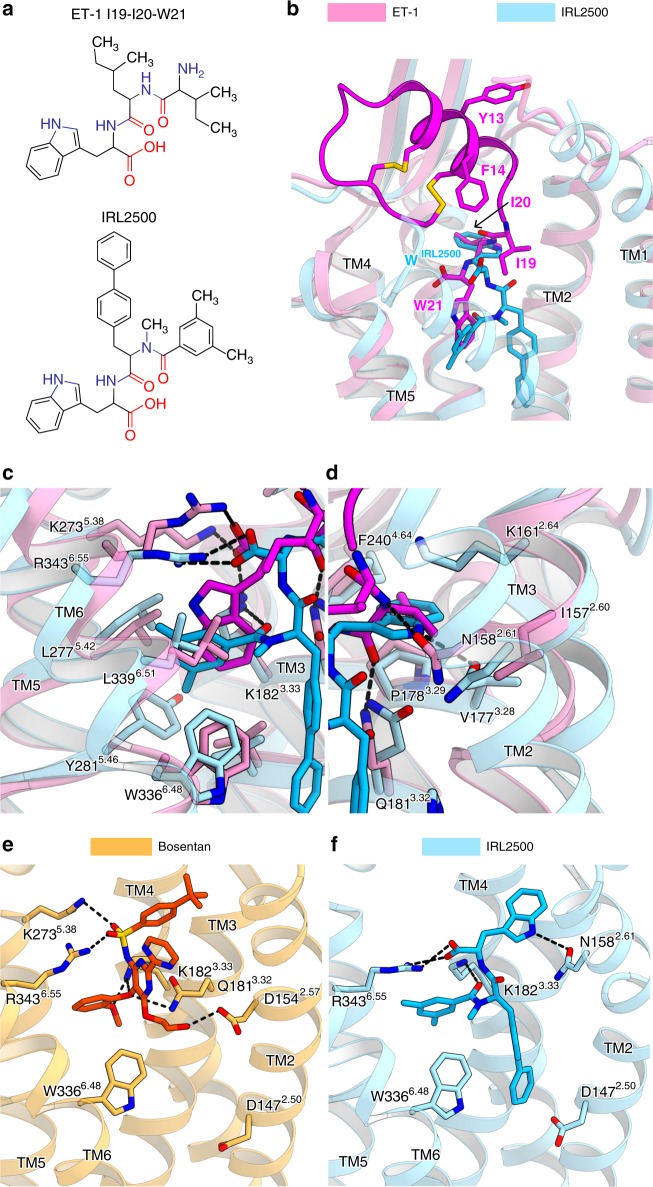
IRL2500 is designed to mimic the F14, I19, I20, and W21 residues in ET-1, which play critical roles in ligand binding to the ETB receptor19. The tryptophan and dimethylbenzoyl groups of IRL2500 seem to be equivalent to W21 and I19 in ET-1, respectively, while the biphenyl group of IRL2500 seems to be equivalent to F14 and I20 of ET-1 (Fig. 3a). However, a comparison between the IRL2500 and ET-1 binding revealed an unexpected difference in their binding interactions (Fig. 3b). The carboxylate of the tryptophan in IRL2500 superimposes well with that of W21 in ET-1, and is coordinated by similar positively charged residues (Fig. 3c). The tryptophan of IRL2500 does not superimpose with the W21 of ET-1, but do with the I20. Instead, the dimethylbenzoyl group of IRL2500 superimposes with the W21. Nevertheless, these hydrophobic moieties of IRL2500 and ET-1 form comparable hydrophobic interactions (Fig. 3c, d). In contrast, the biphenyl group of IRL2500 penetrates into the receptor core, in an opposite manner to the F14 and I20 of ET-1 (Fig. 3b). Overall, while the electrostatic interactions between the carboxylates and the positively charged residues are conserved in IRL2500 and ET-1 binding, the other moieties form distinct interactions with the receptor. The volume of the ligand binding pocket in the ligand-free structure is large, thereby allowing the aromatic moieties of IRL2500 to flip (Fig. 3b).
Fig. 3.
Comparison of binding modes of IRL2500, ET-1, and bosentan. a Chemical structures of IRL2500 and the C-terminal tripeptide of ET-1. b Superimposition of the IRL2500- and ET-1-bound ETB receptors (PDB code: 5GLH). The ET-1- and IRL2500-bound receptors are shown as pink and sky blue ribbons, respectively. IRL2500 is shown as a stick model. ET-1 is shown as a magenta ribbon with stick models of the peptide residues (Y13, F14, I19, I20, and W21). c, d The residues interacting with both IRL2500 and ET-1 are shown as sticks. e, f Binding pockets for bosentan (e) and IRL2500 (f), viewed from the membrane plane. The bosentan-bound receptor (PDB code: 5XPR) is shown as a thin orange ribbon model. The residues involved in bosentan binding (D1542.57, Q1813.32, K1823.33, K2735.38, W3366.48 and R3436.55) and D1472.50 are shown as sticks. Bosentan is shown as an orange stick model. IRL2500 and the IRL2500-bound receptor are colored as in panel (a). The residues involved in IRL2500 binding (N1582.61, K1823.33, R3436.55, D1472.50 and W3366.48) are shown as sticks
IRL2500 has distinct chemical moieties as compared with bosentan, because IRL2500 was not developed based on bosentan. To reveal the similarities and differences in their binding modes, we compared the binding modes of IRL2500 and bosentan in detail (Fig. 3e, f). The carboxylate of IRL2500 and the sulfonamide of bosentan are similarly coordinated by the positively charged residue R3436.55, suggesting that this electrostatic interaction is a common feature of the antagonist binding to the ETB receptor. In addition, like bosentan, the aromatic moieties of IRL2500 fit within the local hydrophobic pockets in the ETB receptor. Overall, IRL2500 has moieties that form similar binding interactions to those of bosentan. However, bosentan lacks the moiety corresponding to the biphenyl group of IRL2500, which deeply penetrates into the receptor core (Fig. 3f). Thus, IRL2500 fits into the pocket more tightly as compared with bosentan, contributing to its higher affinity for the ETB receptor.
IRL2500 functions as an inverse agonist
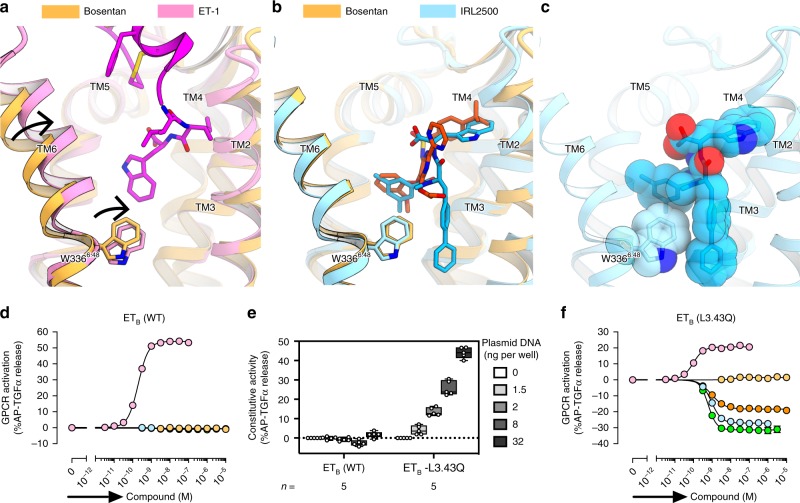
To obtain mechanistic insights into the receptor inactivation by IRL2500, we compared the ETB structures bound to ET-1, bosentan, and IRL2500. Previous structural studies showed that ET-1 binding induces the inward motion of the extracellular portion of TM6 including W3366.48, leading to receptor activation on the intracellular side21 (Fig. 4a). Bosentan binding sterically prevents the inward motion of W3366.48 with its 2-methoxyphenoxy group, and thus functions as an antagonist22 (Fig. 4b). The dimethylbenzoyl group of IRL2500 superimposes well with the 2-methoxyphenoxy group of bosentan and similarly prevents the inward motion (Fig. 4b). Moreover, the dimethylbenzoyl and biphenyl groups of IRL2500 sandwich the W3366.48 side chain, tightly preventing its inward rotation (Fig. 4c). These observations suggest that IRL2500 tightly prevents the transition to the active state, as compared with bosentan, thereby possibly working as an inverse agonist that reduces the basal activity.
Fig. 4.
Inverse agonist activity of IRL2500. a, b Structural changes upon ET-1 and IRL2500 binding, as compared with the bosentan-bound structure, colored as in Fig. 3. Black arrows indicate the inward movements of TM6 and W3366.48. c CPK representations of IRL2500 and the W3366.48 side chain. d Effects of ET-1 and the ETB antagonists (bosentan, IRL2500, K-8794, and BQ-788) on the AP-TGFα release for the ETB receptor. For each experiment, the spontaneous AP-TGFα release response in the absence of the compound is set at the baseline. Data are displayed as means ± SEM from six to eleven independent experiments. e Constitutive activity of the L1923.43Q-mutant ETB receptor (ETB-L3.43Q). HEK293 cells were transfected with titrated volumes of a plasmid encoding the wild-type ETB (ETB-WT) or the L3.43Q-mutant ETB (ETB-L3.43Q) and accumulated AP-TGFα release during 24 h after transfection was measured. The AP-TGFα release signal in the 0 ng receptor plasmid was set as the baseline. Data are displayed as a box-and-whisker plot from five independent experiments with each performed in 4–6 replicates. f Effects of ET-1 and the ETB antagonists (bosentan, IRL2500, K-8794 and BQ-788) on the AP-TGFα release for the constitutively active ETB receptor (ETB-L3.43Q). For each experiment, the spontaneous AP-TGFα release response in the absence of the compound is set at the baseline. Data are displayed as means ± SEM from seven to twelve independent experiments. In the inverse agonist experiments (d, f), the cells were incubated with a test compound for 4 h
To investigate the inverse agonist activity of IRL2500 for the ETB receptor, we first measured the ligand-induced AP-TGFα release responses. In the wild-type ETB, treatment with IRL2500 or the antagonist bosentan with the cells for 4 h-incubation did not change the receptor activation level (Fig. 4d and Table 3). These data suggested that IRL2500 either does not have the inverse agonist activity or that the assay is not sensitive enough to detect the inverse agonist activity. Indeed, we observed that the basal activity of the ETB receptor was very low in the assay and thus we could not distinguish whether IRL2500 functions as an antagonist or an inverse agonist by this assay.
Table 3.
Characterization of inverse agonist activities of endothelin receptor antagonists
| ETB-WT | ETB-L1923.43Q | |
|---|---|---|
| ET-1 | ||
| EC50 (nM) | 0.19 | 0.094 |
| pEC50 (mean ± SEM) | 9.72 ± 0.03 | 10.03 ± 0.04 |
| Emax ± SEM | 53.6 ± 1.7 | 20.5 ± 1.3 |
| n | 10 | 12 |
| Bosentan | ||
| EC50 (nM) | NA | NA |
| pEC50 (mean ± SEM) | NA | NA |
| Emax ± SEM | <2% | <2% |
| n | 11 | 12 |
| IRL2500 | ||
| EC50 (nM) | NA | 0.92 |
| pEC50 (mean ± SEM) | NA | 9.03 ± 0.05 |
| Emax ± SEM | <2% | −27.0 ± 1.1 |
| n | 11 | 12 |
| K-8794 | ||
| EC50 (nM) | NA | 0.61 |
| pEC50 (mean ± SEM) | NA | 9.21 ± 0.04 |
| Emax ± SEM | <2% | −31.5 ± 2.0 |
| n | 6 | 7 |
| BQ-788 | ||
| EC50 (nM) | NA | 0.96 |
| pEC50 (mean ± SEM) | NA | 9.02 ± 0.03 |
| Emax ± SEM | <2% | −18.5 ± 0.7 |
| n | 6 | 7 |
Therefore, we tried the same assay by utilizing a mutation to facilitate the constitutive activity of the ETB receptor. Constitutively active mutant GPCRs have been employed in pharmacological characterizations of inverse agonists30, because such mutant GPCRs allow the signal measurement in a larger detection window. The substitution of the highly conserved L3.43 to glutamine has been identified as a causative activating mutation in the TSHR31 and CYSLTR232 genes, which are related to hyperthyroidism and uveal melanoma, respectively. Therefore, we transferred the L3.43Q mutation into the ETB receptor (ETB-L1923.43Q) and examined its constitutive activity. We found that ETB-L3.43Q induced spontaneous AP-TGFα release (Fig. 4e), indicating that L3.43Q functions as a constitutively active mutation in the ETB receptor. We confirmed that this mutation also increased the constitutive activity in the ETA receptor (Supplementary Fig. 6a). The L3.43Q-mutant ETB and ETA increased the potency of both ET-1 (EC50) and the antagonists (IC50) by approximately 5-fold and 2-fold, respectively (Supplementary Fig. 5b–d). We evaluated the inverse agonist activities of these compounds, using the constitutively active mutant, ETB-L3.43Q (Fig. 4f and Table 3). As expected, the antagonist bosentan did not change the receptor activation from the baseline level. Conversely, IRL2500 reduced the basal activity of the ETB-L3.43Q mutant (EC50 = 0.92 nM). Both bosentan and IRL2500 did not change the basal activity of the ETA-L3.43Q (Supplementary Fig. 6b, c). These data indicate that IRL2500 functions as a potent inverse agonist for the ETB receptor, consistent with the structural observations. The biphenyl group of IRL2500 prevents the inward motion of W3366.48 to stabilize the inactive conformation, and thus IRL2500 functions as an inverse agonist.
In addition to IRL2500, K-8794 and BQ-788 have been characterized as potent ETB-selective antagonists. K-8794 is a high-affinity analog of bosentan22, whereas BQ-788 is a peptide analog with distinct chemical moieties, as compared with those of IRL2500 and bosentan33. We investigated the inverse agonist activities of K-8794 and BQ-788, using the constitutively active mutants of the ETRs (Fig. 4f and Table 3). K-8794 and BQ-788 reduced the basal activity of the ETB-L3.43Q with EC50 values of 0.61 and 0.96 nM, respectively, indicating that they also function as inverse agonists. K-8794 and BQ-788 showed higher and lower efficacies (Emax) of the inverse agonist activity than that of IRL2500, respectively. The K-8794-bound ETB structure has been determined and thus we compare the ETB structures bound to K-8794, bosentan, and IRL2500. The chemical structure of K-8794 is similar to that of bosentan, but K-8794 has a dimethylphenyl group linked to the peptide bond. This modification slightly displaces the overall position of K-8794, thereby moving W3366.48 outward by the interactions with the 6-methoxyphenoxy and the alkyl groups (Supplementary Fig. 7a). Therefore, K-8794 tightly prevents the inward rotation of W3366.48, in a similar manner to IRL2500 (Supplementary Fig. 7b). We thus suggest that this structural feature likely contributes to the inverse agonist activity of K-8794.
Discussion
We have determined the crystal structure of the ETB receptor in complex with the peptide-mimetic drug IRL2500, and thus elucidated the detailed receptor interactions and the structural basis for its ETB selectivity. Although IRL2500 is designed to mimic the partial region of ET-1, its binding mode is quite different. Moreover, using the constitutively active mutant ETB-L3.43Q, we revealed that IRL2500 together with K-8794 and BQ-788, but not bosentan, function as a potent inverse agonist for the ETB receptor, and provided the structural basis for their inverse agonist activities. Our study sheds light on the new aspects of the ETB-selective antagonists, and deepens our understanding of ETR pharmacology.
Although small-molecule ETR antagonists have been developed over the years; however, most ETR antagonists have been designed based on bosentan. Thus, the presently available ETR antagonists are chemically very similar. IRL2500 was developed based on ET-1 and has totally distinct chemical moieties, as compared with those of bosentan. However, the comparison of the IRL2500 and bosentan binding modes revealed the unexpected similarity in their binding interactions. This observation suggests that the charge-complementary interactions in the center of the pocket form the core of the receptor–antagonist interactions, and the other aromatic moieties fit the local hydrophobic pocket. The ligand binding pocket in the inactive ETB structures is larger than those in other GPCR structures, and thus aromatic moieties may be necessary to fit well within the pocket.
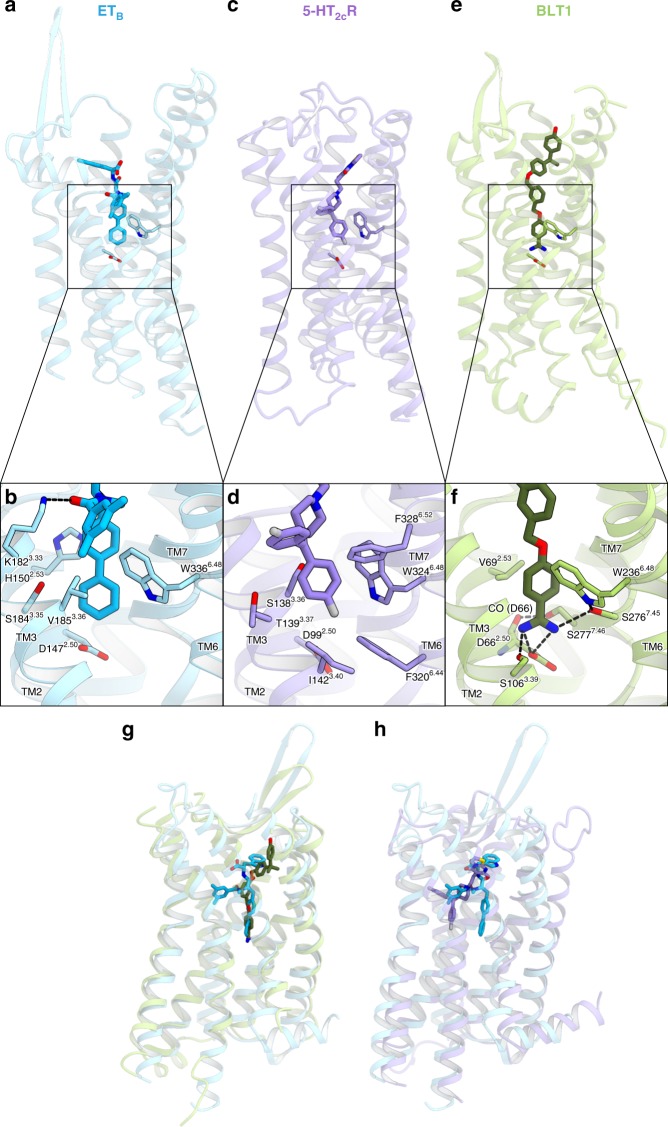
Our study revealed that the biphenyl group of IRL2500 penetrates deeply into the receptor core proximal to D1472.50, preventing the inward motion of W3366.48 in TM6, and thus IRL2500 functions as an inverse agonist (Fig. 5a, b). The deep binding modes of the inverse agonists are also observed in the 5-HT2CR and BLT1 structures (Fig. 5c–f). In the 5-HT2CR structure, the 4-fluorophenyl group of the inverse agonist ritanserin interacts with F3206.44 and W3246.48, the purported “toggle switch” important for GPCR activation34 (Fig. 5c, d). In the BLT1 structure, the benzamidine group of the inverse agonist BIIL260 fits into a sodium binding pocket around D1472.50 (Fig. 5e, f), which is highly conserved among the class A GPCRs35. Sodium selectively competes with agonist binding in most class A GPCRs by stabilizing the inactive conformations, and the sodium binding site is thus an important pocket targeted in the design of negative allosteric modulators and inverse agonists. Instead of the sodium, the benzamidine group of BIIL260 directly hydrogen bonds with D2.50, similarly stabilizing the inactive conformation36 (Fig. 5f). The binding mode of IRL2500 is similar to that of BIIL260 in BLT1, rather than that of ritanserin in the 5-HT2CR (Fig. 5g, h). Although the biphenyl group of IRL2500 does not form any hydrogen-bonding interactions with the receptor, it prevents the conformational change around the D2.50 in a similar manner to the benzamidine moiety of BIIL260 (Fig. 5b, f). For the design of effective inverse agonists, the biphenyl moiety would also be useful as a modulation part along with another moiety that exerts specific and tight binding to the orthosteric site, as well as a benzamidine group.
Fig. 5.
Structural comparison with the inverse agonist-bound GPCR structures. a–f The structures of the IRL2500-bound ETB receptor (a, b), Ritanserin-bound 5-HT2CR (PDB code: 6BQH) (c, d), and the BIIL260-bound BLT1 (PDB code: 5X33) (e, f), shown as sky blue, purple and light green ribbons, respectively. The inverse agonists IRL2500, ritanserin, and BIIL260 are shown as sticks. The binding interactions around the sodium binding site are shown in (b), (d), and (f). The residues involved in ligand binding are represented with sticks. Hydrogen bonds are indicated by black dashed lines. In the IRL2500-bound ETB structure, the biphenyl group of IRL2500 forms van der Waals interactions with the receptor, and does not form any hydrogen-binding interactions. In the BIIL260-bound BLT1 structure, BIIL260 forms hydrogen bonds with D662.50, S1063.39 and S2767.45. g, h Superimposition of the IRL2500-bound ETB structure with the BLT1 (g) and 5-HT2CR (h) structures
Methods
Expression and purification
The haemagglutinin signal peptide, followed by the Flag epitope tag (DYKDDDDK) and a nine-amino-acid linker, was added to the N-terminus of the receptor, and a tobacco etch virus (TEV) protease recognition sequence was introduced between G57 and L66, to remove the disordered N-terminus during the purification process. The C-terminus was truncated after S407, and three cysteine residues were mutated to alanine (C396A, C400A, and C405A) to avoid heterogeneous palmitoylation. To improve crystallogenesis, we introduced four thermostabilizing mutations (R124Y1.55, K270A5.35, S342A6.54, and I381A7.48) and inserted minimal T4 lysozyme25 into intracellular loop 3, between K3035.68 and L3116.23 (ETB-Y4-mT4L22).
The thermostabilized construct ETB-Y4-mT4L was subcloned into a modified pFastBac vector, with the resulting construct encoding a TEV cleavage site followed by a GFP-His10 tag at the C-terminus. The recombinant baculovirus was prepared using the Bac-to-Bac baculovirus expression system (Invitrogen). Sf9 insect cells were infected with the virus at a cell density of 4.0 × 106 cells per ml in Sf900 II medium, and grown for 48 h at 27 °C. The harvested cells were disrupted by sonication, in buffer containing 20 mM Tris–HCl, pH 7.5, and 20% glycerol. The crude membrane fraction was collected by ultracentrifugation at 180,000 g for 1 h. The membrane fraction was solubilized in buffer, containing 20 mM Tris–HCl, pH 7.5, 200 mM NaCl, 1% DDM, 0.2% cholesterol hemisuccinate, 10 μM IRL2500, and 2 mg ml−1 iodoacetamide, for 1 h at 4 °C. The supernatant was separated from the insoluble material by ultracentrifugation at 180,000 g for 20 min, and incubated with TALON resin (Clontech) for 30 min. The resin was washed with ten column volumes of buffer, containing 20 mM Tris–HCl, pH 7.5, 500 mM NaCl, 0.1% LMNG, 0.01% CHS, 10 μM IRL2500, and 15 mM imidazole. The receptor was eluted in buffer, containing 20 mM Tris–HCl, pH 7.5, 500 mM NaCl, 0.01% LMNG, 0.001% CHS, 10 μM IRL2500, and 200 mM imidazole. The eluate was treated with TEV protease and dialyzed against buffer (20 mM Tris–HCl, pH 7.5, 500 mM NaCl, and 10 μM IRL2500). The cleaved GFP-His10 tag and the TEV protease were removed with Co2+-NTA resin. The receptor was concentrated and loaded onto a Superdex200 10/300 Increase size-exclusion column, equilibrated in buffer containing 20 mM Tris–HCl, pH 7.5, 150 mM NaCl, 0.01% LMNG, 0.001% CHS, and 10 μM IRL2500. Peak fractions were pooled, concentrated to 40 mg ml−1 using a centrifugal filter device (Millipore 50 kDa MW cutoff), and frozen until crystallization. During the concentration, IRL2500 was added to a final concentration of 100 μM.
Crystallization
The purified receptor was reconstituted into molten lipid (monoolein and cholesterol 10:1 by mass) at a weight ratio of 1:1.5 (protein:lipid)37. The protein-laden mesophase was dispensed into 96-well glass plates in 30 nl drops and overlaid with 800 nl precipitant solution by a Gryphon LCP robot (Art Robbins Instruments)26. Crystals of ETB-Y4-mT4L bound to IRL2500 were grown at 20 °C in precipitant conditions containing 30% PEG300, 100 mM Bis–tris, pH 7.5, 150 mM sodium phosphate monobasic, and 10 mM TCEP hydrochloride. The crystals were harvested directly from the LCP using micromounts (MiTeGen) or LithoLoops (Protein Wave) and frozen in liquid nitrogen, without adding any extra cryoprotectant.
Data collection and structure determination
X-ray diffraction data were collected at the SPring-8 beamline BL32XU, with 10 × 15 μm2 (width × height) micro-focused beams and an EIGER X 9M detector (Dectris). Various wedge data sets (10°) per crystal were mainly collected with the ZOO system38, an automatic data-collection system developed at SPring-8. The loop-harvested microcrystals were identified by raster scanning and subsequently analyzed by SHIKA39. The collected images were automatically processed with KAMO40 (https://github.com/keitaroyam/yamtbx). Each data set was indexed and integrated with XDS41, and the datasets were hierarchically clustered by using the correlation coefficients of the intensities between datasets41. After the rejection of outliers, 58 data sets were finally merged with XSCALE41. The IRL2500-bound structure was determined by molecular replacement with PHASER42, using the K-8794-bound ETB structure (PDB code: 5X93). Subsequently, the model was rebuilt and refined using COOT43 and PHENIX44, respectively. The final model of IRL2500-bound ETB-Y4-T4L contained residues 91-207, 214-303, and 311-403 of ETB, 1-11 and 19-117 of mT4L, IRL2500, 8 monoolein molecules, 4 phosphoric acids, and 34 water molecules. The model quality was assessed by MolProbity45. Figures were prepared using CueMol (http://www.cuemol.org/ja/).
TGFα shedding assay
The TGFα shedding assay, which measures the activation of Gq/11 and G12/13 signaling24, was performed as described previously22. Briefly, a plasmid encoding an ETB construct with an internal FLAG epitope tag or an ETA construct was transfected, together with a plasmid encoding alkaline phosphatase (AP)-tagged TGFα (AP-TGFα), into HEK293A cells by using a polyethylenimine (PEI) transfection reagent (1 µg ETR plasmid, 2.5 µg AP-TGFα plasmid, and 25 µl of 1 mg per ml PEI solution per 10-cm culture dish). After a 1-day culture, the transfected cells were harvested by trypsinization, washed, and resuspended in 30 ml of Hank’s Balanced Salt Solution (HBSS) containing 5 mM HEPES (pH 7.4). The cell suspension was seeded in a 96-well plate (cell plate) at a volume of 80 μl per well and incubated for 30 min in a CO2 incubator. For the measurement of antagonist activity, IRL2500 was diluted in 0.01% bovine serum albumin (BSA) and HEPES-containing HBSS (assay buffer) and added to the cell plate at a volume of 10 µl per well. After 5 min, ET-1, at a final concentration of 0.5 nM, was added to the cell plate at a volume of 10 µl per well. For the measurement of agonistic activity, after adding 10 µl of the assay buffer, serially diluted ET-1 was mixed with the cells at a volume of 10 µl per well. After a 1 h incubation in the CO2 incubator, aliquots of the conditioned media (80 μl) were transferred to an empty 96-well plate (conditioned media (CM) plate). Similarly, for the measurement of inverse agonist activity, the cells were mixed with 10 µl of the assay buffer, followed by the addition of serially diluted IRL2500, and incubated for 4 h before the transfer of the conditioned media. The AP reaction solution (10 mM p-nitrophenylphosphate (p-NPP), 120 mM Tris–HCl (pH 9.5), 40 mM NaCl, and 10 mM MgCl2) was dispensed into the cell plates and the CM plates (80 µl per well). The absorbance at 405 nm (Abs405) of the plates was measured, using a microplate reader (SpectraMax 340 PC384, Molecular Devices), before and after a 1 h incubation at room temperature. AP-TGFα release was calculated as described previously22. The AP-TGFα release signals were fitted to a four-parameter sigmoidal concentration-response curve, using the Prism 7 software (GraphPad Prism), and the pEC50 (equal to −log10 EC50) and Emax values were obtained.
To obtain equilibrium dissociation constant (KB), for each experiment performed in parallel, IC50 values (IRL2500, K-8794, BQ-788), an EC50 value (ET-1), a Hill slope (K, ET-1), and tested concentration of ET-1 (A; 0.5 nM) were processed as follows46:
The resulting KB values were logarithmically transformed and their negative values (pKB) were used to calculate a difference in pKB value (∆pKB) for a mutant (MT) receptor as follows by using pKB value for WT receptor performed in parallel:
The pKB and the ∆pKB values were used to calculate mean and SEM.
To measure the constitutive activity in a plasmid volume-dependent manner, HEK293 cells were seeded in a 96-well plate at a concentration of 4 × 105 cells per ml in Opti-MEM I Reduced Serum Media (Thermo Fisher Scientific), in a volume of 80 μl per well. A transfection mixture was prepared by mixing the PEI transfection reagent (0.2 µl per well) and plasmids (20 ng AP-TGFα plasmid, titrated ETR plasmid, and an empty vector to balance the total plasmid volume) in Opti-MEM I Reduced Serum Media (20 µl). The mixture was added to the cells, which were then incubated for 24 h before the transfer of the conditioned media. After adding the AP reaction solution, the absorbances of the cells and the CM plates were measured at 20 min intervals. The AP-TGFα release signals were calculated as described above, and the signal in the mock-transfected conditions was set at the baseline. A glutamine mutation was introduced into the internal FLAG epitope-tagged ETB and the ETA constructs at L1923.43 and L1763.43, respectively.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Supplementary information
Acknowledgements
The diffraction experiments were performed at SPring-8 BL32XU (proposal 2017A2527 and 2017B2578). The authors thank the beamline staff at BL32XU of SPring-8 (Sayo, Japan) for technical assistance during data collection. The authors also thank Kouki Kawakami, Takeaki Shibata, and Ayumi Inoue (Tohoku University, Japan) for technical assistance in the characterization of the L3.43Q-mutant ETB and ETA receptors. The authors also thank K. Yamashita for the assistance of the PDB deposition and uploading the raw data. This work was supported by grants from the Platform for Drug Discovery, Informatics and Structural Life Science by the Ministry of Education, Culture, Sports, Science and Technology (MEXT), JSPS KAKENHI grants 16H06294 (O.N.), 17J30010 (W.S.), 30809421 (W.S.), 17K08264 (A.I.), and the Japan Agency for Medical Research and Development (AMED) grants: the PRIME JP17gm5910013 (A.I.) and the LEAP JP17gm0010004 (A.I. and J.A.), and the National Institute of Biomedical Innovation.
Author contributions
C.N. expressed, purified, and crystallized the IRL2500-bound ETB receptor, collected data, and refined the structures. W.S. designed all of the experiments, initially crystallized the receptor, and refined the structure. A.I., F.M.N.K. and J.A. performed and oversaw the cell-based assays. The manuscript was prepared by C.N., W.S., A.I., and O.N. W.S. and O.N. supervised the research.
Data availability
Coordinates and structure factors have been deposited in the Protein Data Bank, under the accession number 6K1Q for the IRL2500-bound structure. The raw X-ray diffraction images are also available at Zenodo (https://zenodo.org/record/2803553). All other data are available from the authors upon reasonable request.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Chisae Nagiri, Wataru Shihoya.
Contributor Information
Wataru Shihoya, Email: wtrshh9@gmail.com.
Osamu Nureki, Email: nureki@bs.s.u-tokyo.ac.jp.
Supplementary information
Supplementary information accompanies this paper at 10.1038/s42003-019-0482-7.
References
- 1.Yanagisawa M, et al. A novel potent vasoconstrictor peptide produced by vascular endothelial cells. Nature. 1988;332:411–415. doi: 10.1038/332411a0. [DOI] [PubMed] [Google Scholar]
- 2.Arai H, et al. Cloning and expression of a cDNA encoding an endothelin receptor. Nature. 1990;348:730–732. doi: 10.1038/348730a0. [DOI] [PubMed] [Google Scholar]
- 3.Sakurai T, et al. Cloning of a cDNA encoding a non-isopeptide-selective subtype of the endothelin receptor. Nature. 1990;348:732–735. doi: 10.1038/348732a0. [DOI] [PubMed] [Google Scholar]
- 4.Channick RN, et al. Effects of the dual endothelin-receptor antagonist bosentan in patients with pulmonary hypertension: a randomised placebo-controlled study. Lancet. 2001;358:1119–1123. doi: 10.1016/S0140-6736(01)06250-X. [DOI] [PubMed] [Google Scholar]
- 5.Rubin LJ, et al. Bosentan therapy for pulmonary arterial hypertension. N. Engl. J. Med. 2002;346:896–903. doi: 10.1056/NEJMoa012212. [DOI] [PubMed] [Google Scholar]
- 6.Maguire JJ, Davenport AP. Endothelin@25—new agonists, antagonists, inhibitors and emerging research frontiers: IUPHAR Review 12. Br. J. Pharmacol. 2014;171:5555–5572. doi: 10.1111/bph.12874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Davenport AP, et al. Endothelin. Pharmacol. Rev. 2016;68:357–418. doi: 10.1124/pr.115.011833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Rosanò L, Spinella F, Bagnato A. Endothelin 1 in cancer: biological implications and therapeutic opportunities. Nat. Rev. Cancer. 2013;13:637–651. doi: 10.1038/nrc3546. [DOI] [PubMed] [Google Scholar]
- 9.Clozel M, et al. Pathophysiological role of endothelin revealed by the first orally active endothelin receptor antagonist. Nature. 1993;365:759–761. doi: 10.1038/365759a0. [DOI] [PubMed] [Google Scholar]
- 10.Clozel M, et al. Pharmacological characterization of bosentan, a new potent orally active nonpeptide endothelin receptor antagonist. J. Pharmacol. Exp. Ther. 1994;270:228–235. [PubMed] [Google Scholar]
- 11.Koyama Y, Michinaga S. Regulations of astrocytic functions by endothelins: roles in the pathophysiological responses of damaged brains. J. Pharmacol. Sci. 2012;118:401–407. doi: 10.1254/jphs.11R13CP. [DOI] [PubMed] [Google Scholar]
- 12.Hammond TR, et al. Endothelin-B receptor activation in astrocytes regulates the rate of oligodendrocyte regeneration during remyelination. Cell Rep. 2015;13:2090–2097. doi: 10.1016/j.celrep.2015.11.002. [DOI] [PubMed] [Google Scholar]
- 13.Moldes O, et al. Neuroprotection afforded by antagonists of endothelin-1 receptors in experimental stroke. Neuropharmacology. 2012;63:1279–1285. doi: 10.1016/j.neuropharm.2012.08.019. [DOI] [PubMed] [Google Scholar]
- 14.Michinaga S, et al. Amelioration of cold injury-induced cortical brain edema formation by selective endothelin ETB receptor antagonists in mice. PLoS One. 2014;9:e102009. doi: 10.1371/journal.pone.0102009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Michinaga S, et al. Improvement of cold injury-induced mouse brain edema by endothelin ETB antagonists is accompanied by decreases in matrixmetalloproteinase 9 and vascular endothelial growth factor-A. Eur. J. Neurosci. 2015;42:2356–2370. doi: 10.1111/ejn.13020. [DOI] [PubMed] [Google Scholar]
- 16.Michinaga S, et al. Delayed administration of BQ788, an ETB antagonist, after experimental traumatic brain injury promotes recovery of blood–brain barrier function and a reduction of cerebral edema in mice. J. Neurotrauma. 2018;35:1481–1494. doi: 10.1089/neu.2017.5421. [DOI] [PubMed] [Google Scholar]
- 17.Palmer MJ. 5 Endothelin receptor antagonists: status and learning 20 years on. Prog. Med. Chem. 2009;47:203–237. doi: 10.1016/S0079-6468(08)00205-1. [DOI] [PubMed] [Google Scholar]
- 18.Mucke HAM. Small-molecule endothelin receptor antagonists: a review of patenting activity across therapeutic areas. IDrugs. 2009;12:366–375. [PubMed] [Google Scholar]
- 19.Früh Th., Saika H., Svensson L., Pitterna Th., Sakaki J., Okada T., Urade Y., Oda K., Fujitani Y., Takimoto M., Yamamura T., Inui T., Makatani M., Umemura I., Teno N., Toh H., Hayakawa K., Murata T. IRL 2500: A potent ETB selective endothelin antagonist. Bioorganic & Medicinal Chemistry Letters. 1996;6(19):2323–2328. doi: 10.1016/0960-894X(96)00421-0. [DOI] [Google Scholar]
- 20.Balwierczak JL, et al. Characterization of a potent and selective endothelin-B receptor antagonist, IRL 2500. J. Cardiovasc. Pharmacol. 1995;26:S393–S396. doi: 10.1097/00005344-199526003-00116. [DOI] [PubMed] [Google Scholar]
- 21.Shihoya W, et al. Activation mechanism of endothelin ETB receptor by endothelin-1. Nature. 2016;537:363–368. doi: 10.1038/nature19319. [DOI] [PubMed] [Google Scholar]
- 22.Shihoya W, et al. X-ray structures of endothelin ETB receptor bound to clinical antagonist bosentan and its analog. Nat. Struct. Mol. Biol. 2017;24:758–764. doi: 10.1038/nsmb.3450. [DOI] [PubMed] [Google Scholar]
- 23.Okuta A, Tani K, Nishimura S, Fujiyoshi Y, Doi T. Thermostabilization of the human endothelin type B receptor. J. Mol. Biol. 2016;428:2265–2274. doi: 10.1016/j.jmb.2016.03.024. [DOI] [PubMed] [Google Scholar]
- 24.Inoue A, et al. TGFα shedding assay: an accurate and versatile method for detecting GPCR activation. Nat. Methods. 2012;9:1021–1029. doi: 10.1038/nmeth.2172. [DOI] [PubMed] [Google Scholar]
- 25.Thorsen TS, Matt R, Weis WI, Kobilka BK. Modified T4 lysozyme fusion proteins facilitate G protein-coupled receptor crystallogenesis. Structure. 2014;22:1657–1664. doi: 10.1016/j.str.2014.08.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Caffrey M, Cherezov V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protoc. 2009;4:706–731. doi: 10.1038/nprot.2009.31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Yamashita K, Hirata K, Yamamoto M. KAMO: towards automated data processing for microcrystals. Acta Crystallogr. D Struct. Biol. 2018;74:441–449. doi: 10.1107/S2059798318004576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ballesteros JA, Weinstein H. [19] Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 1995;25:366–428. doi: 10.1016/S1043-9471(05)80049-7. [DOI] [Google Scholar]
- 29.Sakaki J, et al. Discovery of IRL 3461: a novel and potent endothelin antagonist with balanced ETA/ETB affinity. Bioorg. Med. Chem. Lett. 1998;8:2241–2246. doi: 10.1016/S0960-894X(98)00387-4. [DOI] [PubMed] [Google Scholar]
- 30.Soudijn W, van Wijngaarden I, Ijzerman AP. Structure–activity relationships of inverse agonists for G-protein-coupled receptors. Med. Res. Rev. 2005;25:398–426. doi: 10.1002/med.20031. [DOI] [PubMed] [Google Scholar]
- 31.Trülzsch B, et al. Detection of thyroid-stimulating hormone receptor and Gsalpha mutations: in 75 toxic thyroid nodules by denaturing gradient gel electrophoresis. J. Mol. Med. 2001;78:684–691. doi: 10.1007/s001090000170. [DOI] [PubMed] [Google Scholar]
- 32.Moore AR, et al. Recurrent activating mutations of G-protein-coupled receptor CYSLTR2 in uveal melanoma. Nat. Genet. 2016;48:675–680. doi: 10.1038/ng.3549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ishikawa K, et al. Biochemical and pharmacological profile of a potent and selective endothelin B-receptor antagonist, BQ-788. Proc. Natl Acad. Sci. USA. 1994;91:4892–4896. doi: 10.1073/pnas.91.11.4892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Peng Y, et al. 5-HT2C receptor structures reveal the structural basis of GPCR polypharmacology. Cell. 2018;172:719–730. doi: 10.1016/j.cell.2018.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Katritch V, et al. Allosteric sodium in class A GPCR signaling. Trends Biochem. Sci. 2014;39:233–244. doi: 10.1016/j.tibs.2014.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Hori T, et al. Na+-mimicking ligands stabilize the inactive state of leukotriene B4 receptor BLT1. Nat. Chem. Biol. 2018;14:262–269. doi: 10.1038/nchembio.2547. [DOI] [PubMed] [Google Scholar]
- 37.Shihoya W, et al. Crystal structures of human ETB receptor provide mechanistic insight into receptor activation and partial activation. Nat. Commun. 2018;9:4711. doi: 10.1038/s41467-018-07094-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hirata K, et al. ZOO: an automatic data-collection system for high-throughput structure analysis in protein microcrystallography. Acta Crystallogr. D Biol. Crystallogr. 2019;75:138–150. doi: 10.1107/S2059798318017795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ueno, G. et al. Remote access and automation of SPring-8 MX beamlines. AIP Conf. Proc. 1741, 050021 (2016).
- 40.Yamashita K, Hirata K, Yamamoto M. KAMO: towards automated data processing for microcrystals. Acta Crystallogr. D Biol. Crystallogr. 2018;74:441–449. doi: 10.1107/S2059798318004576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kabsch W. XDS. Acta Crystallogr. D Biol. Crystallogr. 2010;66:125–132. doi: 10.1107/S0907444909047337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.McCoy AJ, et al. Phaser crystallographic software. J. Appl. Crystallogr. 2007;40:658–674. doi: 10.1107/S0021889807021206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Emsley P, Lohkamp B, Scott WG, Cowtan K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 2010;66:486–501. doi: 10.1107/S0907444910007493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Adams PD, et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 2010;66:213–221. doi: 10.1107/S0907444909052925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chen VB, et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 2010;66:12–21. doi: 10.1107/S0907444909042073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Cheng, H. C. The power issue: determination of K-B or K-i from IC50—a closer look at the Cheng-Prusoff equation, the Schild plot and related power equations. J. Pharmacol. Toxicol. Methods46, 61–71 (2001). [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Coordinates and structure factors have been deposited in the Protein Data Bank, under the accession number 6K1Q for the IRL2500-bound structure. The raw X-ray diffraction images are also available at Zenodo (https://zenodo.org/record/2803553). All other data are available from the authors upon reasonable request.