Abstract
Stenotrophomonas maltophilia DF07 is a gram negative bacterium isolated from polluted San Jacinto River sediment near Moncrief Park in Channelview, Texas. The genome of strain DF07 (chromosome and plasmid) was compiled at the scaffold level and can be accessed through the National Center for Biotechnology Information database under accession NZ_NJGC00000000. The DF07 genome consists of a total of 4,801,842 bp encoding for approximately 4,351 functional proteins. Approximately 86 proteins are associated with broad-spectrum antibiotic resistance, 11 are associated with bacteriocin production, and a total of 17 proteins encode for an assortment of Mycobacterium-like virulence and invasion operons. S. maltophilia DF07 is genetically similar to the nosocomial S. maltophilia strain AU12-09, but also harbors an unusually large plasmid that encodes for over 150 proteins of unknown function. Taken together, this strain is potentially an important antibiotic reservoir and its origin within a recreational park merits further study of the area.
Keywords: Antibiotic resistance, Genome, Plasmid, San Jacinto River, Stenotrophomonas maltophilia
Specifications Table
| Subject area | Microbiology |
| More specific subject area | Microbial genetics |
| Type of data | Figures, DNA sequence |
| How data was acquired | DNA sequencing: Illumina Miseq Bioinformatics: NCBI Prokaryotic Genomes Automatic Annotation Pipeline (PGAAP), the RAST web server |
| Data format | Raw and analyzed |
| Experimental factors | Genomic DNA from pure microbial culture |
| Experimental features | Microbial sample was isolated from polluted sediment along San Jacinto River and whole genome sequenced using Illumina MiSeq technology |
| Data source location | Polluted sediment/soil USA: Texas, Channelview, Moncrief Park (29.805,619, −95.095,543) |
| Data accessibility | Strain data is uploaded to National Center for Biotechnology Information database under accession NZ_NJGC00000000. Direct link to data: https://www.ncbi.nlm.nih.gov/Traces/wgs/NJGC01?display=proteins&page=1 |
| Related research article | L. Zhang, M. Morrison, P.O. Cuív, P. Evans, C.M. Rickard. Genome sequence of Stenotrophomonas maltophilia strain AU12-09, isolated from an intravascular catheter. Genome Announc., 1 (2013), e00195-13 [1]. |
Value of the data
|
1. Data
Sequence analysis identifies DF07 as a novel strain of Stenotrophomonas maltophilia, a member of the Xanthomonadaceae family from the Gammaprotebacteria class. This Gram-negative bacterium is ubiquitously distributed throughout both soil and aquatic environments. S. maltophilia is known to be an opportunistic human pathogen. According to MASH genome analysis, the closest relative of S. maltophilia DF07 is S. maltophilia AU12-09, a nosocomial isolate collected from a hospital intravascular catheter (Fig. 1) [1]. Genome annotation reveals that S. maltophilia DF07 possesses many of the antibiotic resistance determinants identified in strain AU12-09. These determinants include a complement of 12 β-lactamase enzymes and associated proteins, aminoglycoside inactivation enzymes, fluoroquinolones resistance proteins as well as 5 tripartite and 27 multidrug pump proteins related to antibiotic efflux (Fig. 2). Of particular note is the large bacterial plasmid found within the DF07 strain. This plasmid has a length of 209,390 bp and encodes for approximately 179 genes, many of which have unknown functions. S. maltophilia DF07 also encodes for several chromosomal and plasmid based Mycobacterium-like virulence and invasion operons (Fig. 3).
Fig. 1.
UPGMA dendogram of S. maltophilia DF07 and the top five similar bacterial genomes.
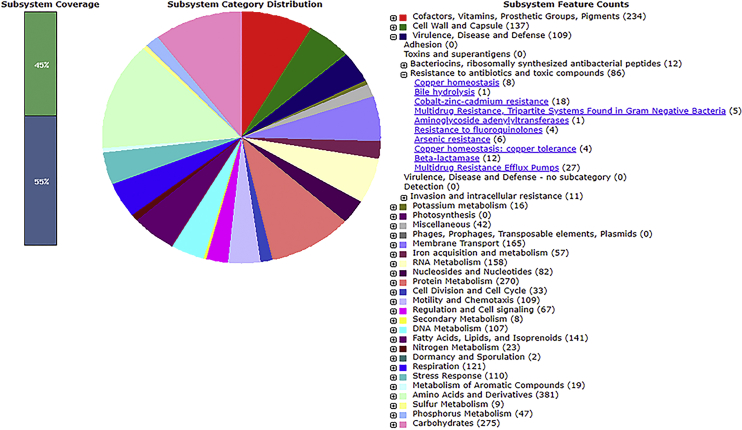
Fig. 2.
Subsystem category distribution of major protein coding genes on the chromosome of S. maltophilia DF07 as annotated by the RAST annotation server. The bar chart shows the subsystem coverage in percentage (the green bar corresponds to percentage of proteins identified in one of the listed subsystems). The pie chart shows the distribution of the 25 most abundant subsystem categories.
Fig. 3.
Subsystem category distribution of major protein coding genes on the plasmid of S. maltophilia DF07 as annotated by the RAST annotation server. The bar chart shows the subsystem coverage in percentage (the green bar corresponds to percentage of proteins identified in one of the listed subsystems). The pie chart shows the distribution of the 25 most abundant subsystem categories.
2. Experimental design, materials, and methods
2.1. Sample collection
Sediment was collected from the bottom of a 12 inch hole dug by the bank of the San Jacinto River alongside Moncrief Park in northern Channelview, Texas. Moncrief Park lies west of the now enclosed San Jacinto River Waste Pits, a submerged Superfund site once used for dumping of paper mill waste.
2.2. Sample screening
Carbon selective media was prepared as previously described in Iyer et al., 2016 [2]. A total of 5 mL of carbon selective media was used for initial sample inoculation. Dibenzofuran added at a final concentration of 100 μg/mL was used as a screening agent and potential carbon source. Subcultures were performed over five weeks before plating onto minimal agar plates supplemented with dibenzofuran.
2.3. Genomic DNA preparation
Plated colonies, yellow in coloration, were revitalized in 5 mL Luria-Bertani medium and grown overnight. Total cellular DNA of the overnight culture was then extracted using a Qiagen DNeasy Blood and Tissue kit.
2.4. Whole genome sequencing
Prepared sample DNA was shipped to Genewiz (South Plainfield, NJ) who performed Illumina MiSeq paired-end sequencing (Table 1).
Table 1.
S. maltophilia DF07 genome statistics.
| Assembly statistics | |
| platform | Illumina MiSeq (2*250) paired end |
| genome size(bp) | 4,801,355 |
| number of contigs | 154 (chromosome) + 6 (plasmid) |
| average coverage |
223.26x |
| Annotation statistics | |
| GC content | 66.40 (chromosome) + 62.20 (plasmid) |
| total genes | 4,522 |
| coding genes | 4,351 |
| RNAs | 87 |
2.5. Genome annotation
Raw sequence data was first quality checked in Fastqc [3] and poor reads filtered out using BBTools [4]. Good sequence reads were then assembled with the Spades 3.10 program [5]. Annotation was performed both through the NCBI Prokaryotic Genomes Automatic Annotation Pipeline (http://www.ncbi.nlm.nih.gov/genomes/static/Pipeline.html) and RAST server (See Table 1) [6], [7].
2.6. Phylogeny analysis
The Mash program was used first for species identification, then to map the five closest bacterial hits based on their Mash distances to the sample strain using the Mash sketch database for RefSeq release 70 (k-mer size = 21, sketch size = 1000) [8]. The file was then imported into R and the Ggdendrogram [9], [10] package used to create a phylogenetic tree (See Fig. 1).
Acknowledgments
Funding for whole genome sequencing of S. maltophilia DF07 was provided by the National Institute of Standards and Technology (NIST) (G110008 58,106).
Author contributions
RI was the principal investigator of this research and all research work was conducted in her laboratory space. RI also submitted compiled sequence data to NCBI and proofread the manuscript. AD carried out genome assembly and phylogenetic analysis. BI performed RAST analysis and helped with drafting the manuscript. All authors read and approved the final text.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
- 1.Zhang L., Morrison M., Cuív P.O., Evans P., Rickard C.M. Genome sequence of Stenotrophomonas maltophilia strain AU12-09, isolated from an intravascular catheter. Genome Announc. 2013;1 doi: 10.1128/genomeA.00195-13. e00195-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Iyer R., Aggarwal J., Iken B. Genome of Pseudomonas nitroreducens DF05 from dioxin contaminated sediment downstream of the San Jacinto River waste pits reveals a broad array of aromatic degradation gene determinants. Genomics data. 2017;14:40–43. doi: 10.1016/j.gdata.2017.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bioinformatics B. Babraham Institute; Cambridge, United Kingdom: 2011. FastQC a Quality Control Tool for High Throughput Sequence Data.http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ [Google Scholar]
- 4.Bushnell B. 2016. BBMap Short Read Aligner.http://sourceforge.net/projects/bbmap [Google Scholar]
- 5.Bankevich A., Nurk S., Antipov D., Gurevich A.A., Dvorkin M., Kulikov A.S., Lesin V.M., Nikolenko S.I., Pham S., Prjibelski A.D., Pyshkin A.V., Sirotkin A.V., Vyahhi N., Tesler G., Alekseyev M.A., Pevzner P.A. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012;19:455–477. doi: 10.1089/cmb.2012.0021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Tatusova T., DiCuccio M., Badretdin A., Chetvernin V., Ciufo S., Li W. second ed. National Center for Biotechnology Information; Bethesda, MD: 2013. Prokaryotic Genome Annotation Pipeline. The NCBI Handbook. [Google Scholar]
- 7.Aziz R.K., Bartels D., Best A.A., DeJongh M., Disz T., Edwards R.A., Formsma K., Gerdes S., Glass E.M., Kubal M., Meyer F. The RAST server: rapid annotations using subsystems technology. BMC Genomics. 2008;9:75. doi: 10.1186/1471-2164-9-75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ondov B.D., Treangen T.J., Melsted P., Mallonee A.B., Bergman N.H., Koren S., Phillippy A.M. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 2016;17(1):132. doi: 10.1186/s13059-016-0997-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wickham H., Hester J., Francois R. 2016. Readr: Read Tabular Data.https://CRAN.Rproject.org/package=readr [Google Scholar]
- 10.de Vries A., Ripley B.D. 2016. Ggdendro: Create Dendrograms and Tree Diagrams Using ‘ggplot2’. R Package Version 0.1–20.https://CRAN.R-project.org/package= ggdendro [Google Scholar]