Abstract
Mutations in the TP53 tumor suppressor gene are common in many cancer types, including the acute myeloid leukemia (AML) subtype known as complex karyotype (CK) AML. Here, we identify a gain-of-function (GOF) p53 mutation that accelerates CK-AML initiation beyond p53 loss and, surprisingly, is required for disease maintenance. The p53R172H mutation (TP53R175H in humans) exhibits a neomorphic function by promoting aberrant self-renewal in leukemic cells, a phenotype that is present in hematopoietic stem and progenitor cells (HSPCs) even prior to their transformation. We identify the Forkhead box H1 transcription factor (Foxh1) as a critical mediator of mutant p53 function that binds to and regulates stem cell-associated genes and transcriptional programs. Our results identify a context where mutant p53 acts as a bona fide oncogene that contributes to the pathogenesis of CK-AML and suggests a common biological theme for TP53 gain-of-function in cancer.
Keywords: Complex-Karyotype AML, Gain-of-function, enhanced self-renewal, p53, Foxh1
INTRODUCTION
TP53 mutations occur in the majority of human cancers and are often associated with poor outcomes (1,2). TP53 encodes a sequence-specific transcription factor (Kern et al, 1991) that is normally maintained at low levels through strict post-translational control (3). In response to DNA damage, activated oncogenes, or other forms of cellular stress, p53 is stabilized and promotes cell cycle arrest, apoptosis, senescence, or other anti-proliferative programs depending on cellular context (4–7). Most TP53 mutations occur in the DNA binding domain and disrupt its transcriptional activity, thereby preventing these stress responses and enabling aberrant proliferation and survival of mutated cells (8).
Cancer-associated mutations typically inactivate p53 through a two-hit mechanism, whereby one allele acquires a missense mutation and the other undergoes “loss-of-heterozygosity” (LOH) via chromosomal deletion (7). Missense TP53 mutations encode proteins that have attenuated capacity to transactivate wild-type target genes, despite being frequently stabilized owing to reduced interaction with negative regulators (9). These mutant proteins can instill neomorphic gain-of-function (GOF) activities that contribute to cancer phenotypes beyond p53 loss (10). At the organismal level, mice harboring certain germline missense mutations in Trp53 (hereafter referred to as p53) develop an altered tumor spectrum compared to p53 null mice, including a larger fraction of epithelial cancers with increased metastatic potential (11,12). At the cellular level, some GOF p53 mutants promote chemoresistance, invasiveness, and/or an epithelial-to-mesenchymal transition through diverse mechanisms (11–13). Another neomorphic function of mutant p53 involves its ability to facilitate the formation of induced pluripotent stem cells (iPSCs) more so than p53 loss (14,15), though the extent to which this GOF activity is relevant to cancer is poorly understood.
In contrast to their high prevalence in most solid tumors, TP53 mutations occur in around 10% of blood cancers though, when they occur, are associated with poor prognosis (16,17). In acute myeloid leukemia (AML), TP53 mutations are associated with a subtype known as complex karyotype AML (CK-AML), which is defined by the presence of 3 or more cytogenetic abnormalities and a dismal 5-year survival rate of less than 2% (16,17). Functional studies in mice indicate that p53 inactivation in the hematopoietic compartment can produce chemoresistant malignancies with increased leukemia initiating potential, mirroring key features linked to TP53 mutations in AML patients (18–20). Still, whether and how TP53 missense mutations confer GOF activities to p53 in AML is not known.
In this study, we set out to test whether mutant p53 has GOF activity in AML and, if so, to determine the underlying mechanisms behind this effect. We chose to study p53R172H (R175H in humans), a mutant form of p53 that has been shown to confer GOF activity in solid tumors and is the most common allele in AML patients (Dr. Elli Papaemmanuil, personal communication). Several complementary in vitro and in vivo systems were used to compare the biological features of wild-type, p53 null, or p53 mutant alleles, leading us to identify a neomorphic function of mutant p53 in hematopoietic stem and progenitor cells that exerts its effect by enhancing cellular self-renewal beyond that produced by p53 inactivation. We also identify a novel mediator of mutant p53 function, Foxh1, which contributes to the aberrant self-renewal phenotype. As such, suppression of either mutant p53 or Foxh1 ablates this stemness capacity by triggering differentiation. These observations illustrate how mutant p53 can acquire a pro-oncogenic activity that magnifies loss of its tumor suppressive functions and creates a previously unappreciated molecular dependency in AML.
RESULTS
p53R172H accelerates the onset of hematological malignancies beyond effects of p53 deficiency.
We first compared the ability of a mutant or null p53 allele to promote leukemogenesis in a well-defined genetic model. Because of its previously defined GOF activity in other settings, we used a conditional mutant p53 allele harboring a R172H mutation downstream of a “lox-stop-lox” cassette (LSL-p53R172H) (11), which corresponds to TP53R175H, one of the most frequent mutations in clinical cases of AML (17). Mouse cohorts were produced to harbor one p53R172H allele and a p53 floxed allele (p53LSL-R172H/F), or were homozygous for the floxed p53 allele (p53F/F), such that all hematopoietic cells would become p53R172H/Δ and p53Δ/Δ, respectively, in the presence of the hematopoietic-specific Vav1-cre transgene (21). While both the p53R172H/Δ and p53Δ/Δ cohorts eventually succumbed to disease with full penetrance, mice harboring the p53R172H allele became moribund significantly faster (Figure 1A). As commonly observed in p53−/− mice (22), most animals developed thymic lymphoma, although some had leukemias and rarely a mixed disease (leukemia and lymphoma) (Figure 1B).
Figure 1. p53R172H Leads to Accelerated Onset of Hematological Malignancies.
(A) Kaplan-Meier survival curves of native mice with p53Δ/Δ and p53R172H/Δ hematopoietic cells recombined using Vav1-Cre. (B) Spectrum of hematological malignancies arising in p53Δ/Δ (n=15) and p53R172H/Δ (n=11 native mice. (C) Kaplan-Meier survival curves of thymectomized mice transplanted with p53f/f, p53f/f;Mx1-Cre and p53R172H/F;Mx1-Cre bone marrow (BM) (D) Expression of CD11b and Gr1 in the peripheral blood of moribund mice from (C) (E) Representative H&E image of mice with acute myeloid leukemia arising after transplant in thymectomized recipients. ** p <0.005, **** p <0.0001, Log-rank (Mantel-Cox) test.
The above observations are consistent with those previously obtained using mice harboring a humanized p53R248Q allele expressed in the whole body of mice, which also develop T cell lymphoma faster than p53 null mice (23). However, neither model addresses the biology of mutant p53 action in the clinically-relevant context of AML, likely due to the high penetrance of lymphomas that arise in the mouse. Therefore, to bias the model against T cell lymphoma development, we transplanted bone marrow cells from Mx1-Cre;p53R172H/F and Mx1-Cre;p53F/F mice into thymectomized recipients. Three weeks post-transplantation, intraperitoneal injection of PolyI:C was used to activate the interferon-inducible Mx1-Cre, triggering the recombination of floxed alleles in transplanted cells (24). Of note, the use of the inducible Mx1-Cre recombinase enabled transplantation of functionally wild type whole bone marrow (WBM) cells, avoiding biases in engraftment efficiency that might occur upon p53 alteration. As in the autochthonous model, p53R172H/Δ mice succumbed to disease faster than their p53-null counterparts (Figure 1C), though, in this instance, the disease was immunophenotypically and histologically characterized as AML based on the expression of Cd11b and Gr1 on the surface of infiltrating leukemic cells (Figure 1D, E). Thus, p53R172H gives rise to an aggressive leukemia that significantly shortens overall survival as compared to the disease arising from p53 deficient bone marrow cells, providing functional evidence that a p53 mutant allele can contribute to AML beyond effects of p53 loss.
Sustained expression of mutant p53 is required for the maintenance of p53R172H leukemic cells
We next examined whether the p53R172H mutant protein was required for survival of p53 mutant leukemic cells. We utilized previously characterized p53Δ/Δ or p53R172H/Δ and p53R172H/11B3 CK-AML murine cell lines that were created by co-transduction of short hairpin RNAs (shRNAs) targeting Nf1 (GFP-labeled) and Mll3 (mCherry-labeled), hereafter referred to as PNM (19,25). These cell lines were generated to reflect the most common genetic configurations found in patients, including p53 loss alone (p53Δ/Δ), p53R172H over a focal p53 deletion (p53R172H/Δ) or p53R172H over a larger genomic deletion that encompasses p53 (p53R172H/11B3) (25). Control shRNAs against Renilla luciferase (shRen) or p53 (sh.p53) were co-expressed with a blue fluorescent protein (BFP) reporter (Figure S1A) (26). The percentage of BFP+ cells was monitored over time as a measurement of fitness under the two shRNA-containing conditions.
We observed a significant reduction in the relative number of sh.p53 BFP+ cells compared to sh.Ren BFP+ cells over time (Figure 2A light blue and dark blue lines), indicating negative selection against cells in which the mutant protein is depleted. By contrast, sh.p53 had no effect on p53Δ/Δ AML cells, ruling out potential off target effects of the p53 shRNA (Figure 2A black line). p53 depletion in cells harboring p53R172H promoted differentiation and triggered apoptosis as assessed by flow cytometry for CD11b and Annexin V, respectively (Figure 2B,C). Thus, AML cells expressing p53R172H acquire a molecular dependency on mutant p53 that contributes to a differentiation block that sustains leukemogenesis.
Figure 2. Depletion of mutant p53 impairs the growth of p53 mutant leukemias.
(A) The ratio of sh.p53 BFP+ cells compared to sh.Ren BFP+ cells over time in leukemic lines. (B) CD11b expression in the shRNA BFP+ containing cells (n = 3) and (C) Annexin V shown by FACS 7 days after infection. Data presented as mean ± s.e.m (D) Relative expression of TP53 7 days post infection with pRS sh.TP53 in human KY821 cells. (E) CFU in human AML cells with sh.Ren or sh.TP53. Data presented as mean ± s.e.m (n = 3) (F) hCD16 expression by FACS in pRS Ctrl or pRS TP53 cells from growing in CFU assay (G) Percentage of hCD45 in the peripheral blood of mice 5 weeks post transplant in shCntr and shCntr moribund mice (black), shTP53 (blue) and shTP53 moribund mice (orange). Data are represented as mean ± s.e.m (n=5 and n=5) (H) Kaplan-Meier curve from NSG mice transplanted with equal numbers of KY821 cells transduced with wither shCntr or shTP53 (n=5 and n=5). (I) Western blot for human TP53 in cells before injection and in sorted hCD45 cells from the peripheral blood of 3 independent moribund shTP53 mice. *p <0.05, **p <0.005 ***p <0.0005. B. Kruskal-Wallis 1-way ANOVA. C, D, F, J, unpaired t-test.
To determine whether human AML can also depend on sustained expression of mutant p53, we first examined publicly available CRISPR/Cas9 genome-wide screening data (27). In these studies, sgRNAs were introduced into human AML lines expressing Cas9 followed by assessment of their enrichment or depletion over time using next generation sequencing. Interestingly, sgRNAs targeting p53 were invariably enriched in AML lines harboring wild-type TP53 but depleted to varying degrees in those expressing mutant TP53 genes (e.g. R248Q – another hotspot TP53 mutation), suggesting that a subset of the latter group might depend on sustained expression of the mutant protein for their proliferation and/or survival (Figure S1B).
None of the AML cell lines used in the aforementioned CRISPR/Cas9 screen harbored an R175H mutation (corresponding to the mutation we studied in mice). However, we identified a human AML cell line (KY821) that harbored this mutation in the absence of a WT p53 allele (Figure S1C) and used it to functionally test the requirement for this mutant form of TP53 in vitro and in vivo. Cells were transduced with a validated shRNA targeting human p53 (sh.TP53) (28) or empty vector control (Ctrl) and examined for their ability to form colonies in methylcellulose culture in vitro or give rise to leukemia upon transplantation into sub-lethally irradiated NOD-scid IL-2R−/− (NSG) immune-deficient mice. Similar to observations from murine AML, suppression of mutant TP53 in KY821 cells significantly reduced their colony-forming ability (Figure 2D,E) and led to the up-regulation of the myeloid differentiation marker CD16 (Figure 2F), again suggesting that this p53 mutant contributes to a differentiation block.
To test whether TP53R175H mutant cells transduced with sh.TP53 exhibited reduced fitness in vivo, we analyzed peripheral blood for the presence of human CD45-positive leukocytes in recipient mice 5 weeks post-transplantation. Remarkably, this analysis revealed a significant reduction of cells containing sh.TP53 versus control cells (Figure 2G), confirming the suppressive effect of mutant p53 knock-down in p53 mutant-expressing human leukemias. In addition, knockdown of mutant p53 significantly increased the survival of transplanted mice compared to controls (Figure 2H). At the time of death, the resulting leukemias invariably selected against expression of the p53-targeting shRNA and restored the expression of mutant p53, consistent with our observations from murine models (Figure 2I and S1D–F). Therefore, KY821 cells require mutant p53 protein to sustain leukemia. Collectively, these results reveal an oncogenic function for mutant p53 in murine and human AML.
p53R172H-derived leukemia is associated with stem cell transcriptional signatures.
Most studies suggest that the GOF activities of mutant p53 proteins arise from their ability to alter transcription in a manner that is distinct from effects due to p53 loss (29). Therefore, to examine how mutant p53 altered the transcriptome of CK-AML cells, we performed gene expression profiling of PNM p53-null and p53-mutant leukemias via RNA sequencing (RNA-seq) and identified 252 significantly differentially expressed genes (Figure 3A and Table S1). A systematic and unbiased analysis of the data using gene ontology (GO) pathway analysis revealed Tgf-β and stem cell pathways to be positively correlated with the top upregulated genes in our p53R172H murine leukemias (Figure 3B). Notably, these same signatures were enriched in the transcriptional profiles of leukemias obtained from CK-AML patients – the AML subtype in which missense TP53 mutations most commonly occur – compared to other leukemia subtypes or normal hematopoietic cells (www.bloodspot.eu) (31) (Figure 3C). Beyond general stem cell signatures, gene set enrichment analysis indicated that p53 mutant AML was enriched for genes linked to hematopoietic stem cells (HSC) and long term-HSCs (LT-HSCs) (30) (Figure 3D). Therefore, both murine and human mutant p53-expressing (but not p53-deficient) leukemias converge on similar transcriptional pathways, some of which are linked to stemness and inhibition of myeloid differentiation.
Figure 3. Gene expression profile of p53R172H/Δ leukemias.
(A) Volcano plot from RNA-seq comparing p53R172H/Δ leukemias to p53Δ/Δ showing p53 as the most highly expressed gene between the two groups. (B) Gene ontology analysis of pathways associated with genes up-regulated in p53R172H/Δ leukemias. (C) Expression of Gene Ontology-associated pathways in human AML samples and normal human hematopoietic cells (www.bloodspot.eu). (D) Gene Set Enrichment Analysis (GSEA) comparing gene expression between p53R172H/Δ leukemias to p53Δ/Δ and other known signatures related to stem cells and hematopoietic stem cells. Two-tailed t-test * p <0.05, ** p <0.005, *** p <0.0005. HSC: Hematopoietic stem cell, MPP: Multipotential progenitors, CMP: Common myeloid progenitor cell, GMP: Granulocyte monocyte progenitors, MEP: Megakaryocyte-erythroid progenitor cell, early_PM: Early Promyelocyte, late_PM: Late Promyelocyte, MY: Myelocyte, MM: Metamyelocytes, BC: Band cell, PMN: Polymorphonuclear cells, Mono: Monocytes.
p53R172H enhances self-renewal in hematopoietic cells in vitro and in vivo
AML arises from alterations (e.g. mutations) that promote proliferation or enhance the self-renewal of myeloid progenitor cells (32,33). To directly examine the functional consequences of mutant p53 expression in the normal hematopoietic compartment, we isolated adult whole bone marrow (WBM) cells from p53WT/WT, p53Δ/Δ and p53R172H/Δ mice and performed limiting dilution cultures in methylcellulose medium followed by serial replating. Under these cytokine-rich culture conditions, wild-type cells lose their replating capacity after three sequential passages (34). As previously described (35), loss of p53 modestly increased the replating capacity of hematopoietic cells as compared to those expressing wild-type p53 (Figure 4A). In stark contrast, hematopoietic cells expressing p53R172H/Δ not only replated more frequently than p53Δ/Δ and WT controls, but also could replate indefinitely (Figure 4A).
Figure 4. p53R172H induces aberrant self-renewal in vitro and in vivo.
(A) Total number of colony-forming units (CFU) generated by p53WT (black), p53Δ/Δ (red) and p53R172H/Δ (blue) cells. Data are representative from four independent experiments. Error bars correspond to mean ± s.e.m (n=3). (B) Total number of colony-forming units (CFU) generated by p53R172H/Δ cKIT+ cells containing sh.Ren or sh.p53. Data are representative from three independent experiments. Error bars correspond to mean ± s.e.m (n=3). (C) Western-blotting for TP53 and B-ACTIN at passage 4 (P4). (D) Schematic representation of the competitive transplantation protocol. (E) Percentage of CD45.2 cells during the secondary competitive transplant in the PB of mice monitored monthly by bleeding (n=5 per group). (F) Percentage of CD45.2 cells in the PB four months after transplant (n=4–7). (G) Percentage of CD45.2 cells in the BM four months after transplant (n=4–7), (H) Percentage of Megakaryocyte Erythrocyte Progenitors (MEP), (I) Granulocyte Monocyte Progenitors (GMP) and (J) Common Myeloid Progenitors (CMP) in the BM of mice four months after transplant (n=4–7). (K) Schematic representation of the transplant layout. LSKs were sorted from p53R172H/Δ mice and were infected with an sh.Ren or sh.p53 conjugated to BFP fluorescence. Two days after infection equal numbers of cells were transplanted in lethally irradiated CD45.1 mice. (L) BFP fluorescence over time as assessed by monthly bleeds in recipient mice (n=5). (M) Percentage of BFP+ cells in the BM of mice with sh.Ren or sh.p53 four months after transplant (n=3–5). (N) Percentage of BFP+ LSKs in the BM of mice with sh.Ren or sh.p53 four months after transplant (n=3–5). B, C, and D. Two-tailed t-test * p <0.05, ** p <0.005, *** p <0.0005. Error bars correspond to mean ± s.e.m.
This enhanced capability could not be explained by differences in the pre-existing frequency of stem and progenitor cells in the bone marrow, since immunophenotyping of 7 week-old mice (the time of bone marrow collection) did not show an increase in any type of stem or progenitor cell (Figure S2A–K). This increase in self-renewal was observed in p53R172H/Δ but not p53R172H/WT cells, suggesting that loss of the residual WT allele is a prerequisite for this GOF effect (Figure S3A). Furthermore, p53R172H/Δ cells exhibited higher nuclear to cytoplasmic ratio when compared to p53Δ/Δ and WT cells at the same passage (P3) (Figure S3B) and expressed the highest levels of CD34 (Figure S3C), an important marker of long-term repopulation of HSCs (36). The replating phenotype produced by mutant p53 was similar to that produced by disruption of Tet2, a well-characterized leukemia tumor suppressor gene whose loss promotes enhanced self-renewal (Figure S3C) (37,38). Nevertheless, the p53R172H/Δ cells were different than Tet2Δ/Δ cells at the immunophenotypic and morphological levels (Figure S3C, D), suggesting subtle differences between how these alterations act.
To further explore the relationship between mutant p53 and self-renewal potential, we isolated cKit+ bone marrow cells from the p53R172H/Δ mice and transduced them with viral vectors co-expressing an shRNA targeting mutant p53 (sh.p53) or a Renilla control (sh.Ren). These vectors encoded puromycin resistance, allowing us to select and maintain pure shRNA-expressing cells in culture. This isogenic comparison validated our previous observation: p53 knockdown in p53 mutant cells halted replating capacity after the fourth passage (comparable to p53Δ/Δ WBM), whereas sh.Ren cells (which retain expression of mutant p53) were able to replate indefinitely (Figure 4B and 4C). We confirmed that the normalization of self-renewal potential by sh.p53 was not due to an off target effect of RNAi, as the same shRNA had no effect on the fitness of p53Δ/Δ hematopoietic cells (Figure S2L).
Consistent with these findings, competitive transplantation studies using p53WT/WT, p53Δ/Δ and p53R172H/Δ BM (Figure 4D) demonstrated that p53R172H/Δ cells outcompeted wild-type cells to a greater extent than p53Δ/Δ cells (Figure S3E–G), a phenotype that was further exacerbated in secondary transplants (Figure 4E). Analysis of peripheral blood (Figure 4F) and BM (Figure 4G) demonstrated that p53R172H/Δ cells were significantly enriched with respect to p53WT cells. Furthermore, within the BM of p53R172H/Δ mice, HSCs were expanded in relative and absolute number compared to p53Δ/Δ mice (Figure S3H–K). All downstream progenitors, including megakaryocyte/erythroid progenitors (MEPs), granulocyte/macrophage progenitors (GMPs) and common myeloid progenitors (CMPs), were significantly increased in p53R172H/Δ mice (Figure 4H–J).
For further validation, we examined the consequences of inhibiting mutant p53 in normal mouse HSCs. For these experiments, we used the bone marrow of p53R172H/Δ mice to isolate lineage negative (Lin-)/Sca-1+/cKit+ cells (LSKs), a cell population capable of maintaining long-term hematopoiesis in mice (39). To suppress p53, these cells were infected with retroviruses encoding an shRNA (targeting mutant p53 or Renilla control) linked to blue fluorescent protein (BFP) (Figure 4K). Equal numbers of either sh.Ren or sh.p53 cells were mixed with non shRNA expressing p53R172H BFP− cells and transplanted into CD45.1 lethally irradiated mice in an in vivo competition format. The fraction of BFP+ cells in the peripheral blood was monitored for four months. p53R172H/Δ BM cells transduced with the p53 shRNA were progressively depleted from peripheral blood, becoming nearly undetectable by the end of the experiment (Figure 4L).
Analysis of bone marrow at the final time point revealed that fewer cells expressing sh.p53 remained as compared with those expressing control sh.Ren (Figure 4M). Moreover, the proportion of LSKs within the sh.p53 population was lower than in the control sh.Ren population (Figure 4N). Altogether, these observations establish a role for p53R172H in promoting aberrant self-renewal in adult hematopoietic cells in a manner that renders these cells dependent on its action for their maintenance in a pre-malignant setting. Hence, phenotypes linked to p53 mutant GOF can emerge in the hematopoietic system prior to neoplastic transformation.
Foxh1 is required for mutant p53 induced self-renewal in both normal and leukemic cells.
To explore the mechanism by which mutant p53 enhances cellular self-renewal, we surveyed the genes that were expressed at higher levels in p53R172H/Δ leukemias compared to p53Δ/Δ leukemias. One of the most significantly up-regulated genes in our analysis was forkhead box H1 (Foxh1) (Supplementary Table 1), which encodes a key transcription factor that is known to mediate Nodal/Tgf-β signaling during embryogenesis but is not expressed at appreciable levels in the adult hematopoietic compartment or any other mouse tissue (Figure S4A and S4B–C). Like mutant p53 (40), Foxh1 has been linked to EMT (41,42) and can facilitate the reprogramming of fibroblasts into induced pluripotent stem cells (iPSCs) (43). Furthermore, many of the top upregulated genes in p53R172H/Δ leukemias were known Foxh1 targets (44) (Figure 5A). Taken together, these observations led us to hypothesize that Foxh1 might be a key mediator of the p53R172H GOF activity in leukemia.
Figure 5. Identification of Foxh1 as a p53 mutant mediator.
(A) GSEA comparing the expression of genes associated with genomic regions occupied by Foxh1 when compared to the genes differentially expressed in p53R172H and p53Δ/Δ leukemias. (B) Microarray data showing the expression of FoxH1 across normal human hematopoietic cells and different types of leukemias. (C) Overall Survival in patients with AML that have FoxH1 mRNA upregulation or genetic amplification compared to patients without (cbioportal –TCGA). (D) Western-blotting for FOXH1 in HSPCs from p53R172H and p53Δ/Δ cells at Passage 4. (E) Western-Blotting for TP53 and FOXH1 in p53R172H HSPCs transduced with sh.Ren or sh.p53. (F) Western-Blotting for TP53 and FOXH1 in two independent p53R172H murine leukemic cells transduced with sh.Ren or sh.p53. (G) Western-Blotting for TP53 and FOXH1 in p53R175H human leukemic cells transduced with sh.Ren or sh.TP53. * p <0.05, ** p <0.005, *** p <0.0005. Hematopoietic stem cell, MPP: Multipotential progenitors, CMP: Common myeloid progenitor cell, GMP: Granulocyte monocyte progenitors, MEP: Megakaryocyte-erythroid progenitor cell, early_PM: Early Promyelocyte, late_PM: Late Promyelocyte, MY: Myelocyte, MM: Metamyelocytes, BC: Band cell, PMN: Polymorphonuclear cells, Mono: Monocytes.
In agreement, analysis of publicly available expression data from normal human hematopoietic cells and different leukemia subtypes revealed that FOXH1 expression was significantly higher in all AMLs (particularly in CK-AML patients, the majority of which harbor TP53 mutations) compared to any type of normal cell (Figure 5B and S5A–B). In addition, like mutant TP53, FOXH1 upregulation correlated with poor survival in AML patients (45,46) (Figure 5C). Consistent with the RNA-seq results, Foxh1 expression was also higher in p53R172H/Δ hematopoietic stem and progenitor cells (HSPCs) compared to p53Δ/Δ cells growing in colonies (Figure 5D), and knockdown of mutant p53 reduced Foxh1 levels in murine HSPCs (Figure 5E) as well as murine and human leukemic cells (Figures 5F and 5G, respectively). Thus, mutant p53 expression leads to higher levels of Foxh1 in both normal and leukemic cells from mouse and human leukemias.
Foxh1 is a critical effector of p53R172H via its ability to bind and regulate stem cell genes
Since Foxh1 is a transcription factor and its role in cells of hematopoietic origin is not known, we performed chromatin immunoprecipitation with an anti-Foxh1 antibody followed by high-throughput sequencing (ChIP-Seq) in order to identify the genome-wide Foxh1 binding sites in p53 mutant murine leukemic cells (Figure 6A). Sites bound by Foxh1 correlated with the presence of canonical Foxh1 binding motifs (p<1e-1915) as well as with other Tgf-β mediators (47,48) (Figure 6B). GO pathway analysis indicated that genes associated with Foxh1 peaks are involved in BMP and Tgf-β signaling pathways, as well as myeloid leukemia (Figure 6C).
Figure 6. Foxh1 directly binds genes involved in leukemia and stem cell properties.
(A) Pie charts depicting the genome-wide distribution of Foxh1 binding in murine leukemic cells. (B) Motif enrichment analysis for the Foxh1 bound genomic regions in leukemic cells. (C) Gene ontology analysis of genes bound by Foxh1 in murine leukemic cells (D) Venn Diagram depicting the overlap between the genes down regulated upon p53 KD (blue circle, Foxh1 KD (green circle) and Foxh1 ChIP (orange circle). (E) GSEA from transcriptional profile of cells upon Foxh1 KD. (F) Gene tracks for Foxh1 ChIP-seq at the Runx2 and Mef2c loci together with RNA-seq tracks from either sh.Ren (Red), sh. Foxh1 (green) or sh.p53 (blue). (G) Mean log2 expression of transcriptional signature composed of significantly up-regulated genes in p53R172H/Δ versus p53Δ/Δ murine leukemias across AML and normal cells, as well as (H) same comparison for genes downregulated following Foxh1 KD. Student’s t-test between CK- AML and other AML types: * p <0.05, ** p <0.005, *** p <0.0005, **** p <0.00005..
To investigate how many of the genes bound by Foxh1 are dynamically regulated, we suppressed Foxh1 and compared the downregulated genes (green circle) to those physically bound by Foxh1 (orange circle) (Figure 6D). Consistent with a molecular relationship between mutant p53 and Foxh1 expression, there was a significant overlap between the genes that were decreased upon knockdown of mutant p53 (blue circle) or Foxh1 (green circle), with over 50% of genes showing evidence of co-regulation (Fisher Exact Test p-value < 2.2e-16). Gene set enrichment analysis (GSEA) confirmed that genes that were downregulated upon Foxh1 knockdown significantly overlapped with those downregulated upon mutant p53 knockdown (NES=−14.5, FDR=0) (Figure 6E). In line with these results, many of the transcriptional signatures related to hematopoietic stem cells and leukemia (previously shown to be up-regulated in p53R172H/Δ cells) were conversely enriched upon suppression of either Foxh1 (Figure 6E) or p53 (Figure S6A). As examples two genes involved in leukemia and hematopoietic cell differentiation, Runx2 (49) and Mef2c (50) are both bound by Foxh1 and their expression is reduced by either Foxh1 or p53 knockdown (Figure 6F). Hence, our data show that mutant p53 induces Foxh1, which can directly bind to and control genes linked to the differentiation status and self-renewal properties of AML.
To confirm that these Foxh1-dependent transcripts were also observed in human AML, we derived a transcriptional signature composed of differentially up-regulated genes in p53R172H/Δ versus p53Δ/Δ murine leukemias and compared it to expression data from different karyotypes of human AML as well as normal hematopoietic cells (31). We observed a significant up-regulation of p53R172H/Δ murine leukemia-derived transcriptional signature in CK-AML patients (dark red) compared to all other leukemia types (light red) or normal hematopoietic cells (black) (Figure 6G), suggesting our model transcriptionally resembles CK-AML harboring TP53 mutations. A similar increase in Foxh1-dependent transcripts (sh.Foxh1 vs sh.Ren) was also observed in CK-AML patients (Figure 6H). These results reveal a FOXH1 signature in human AML harboring TP53 mutations, further supporting the existence of a functional mutant p53-Foxh1 axis in this disease.
Foxh1 is necessary and sufficient for the enhanced self-renewal phenotype produced by mutant p53.
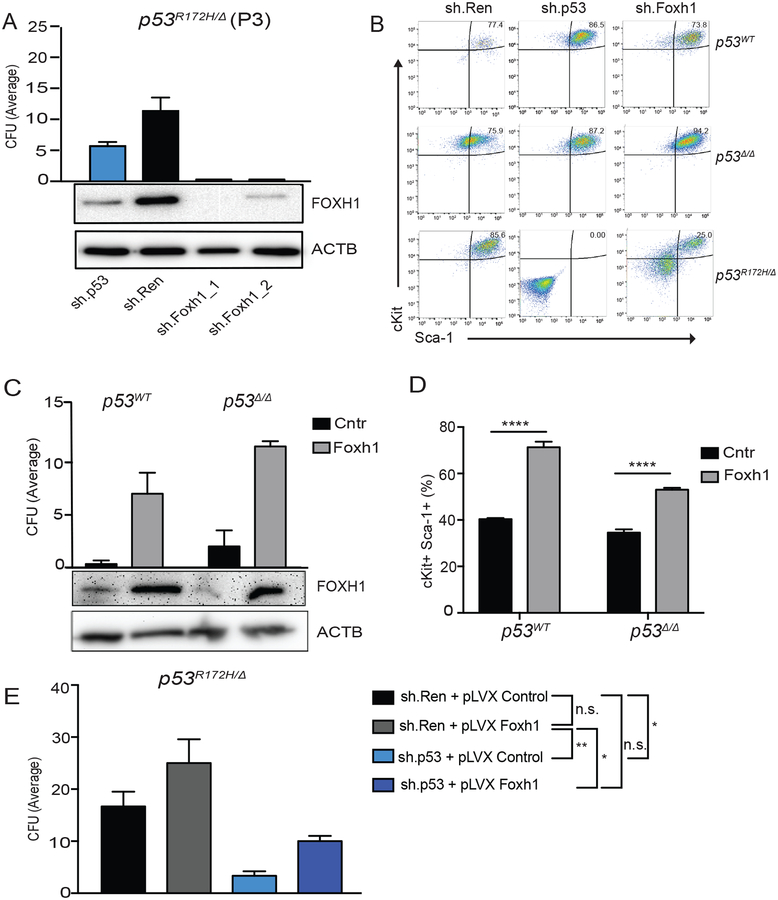
To further explore the links between mutant p53, Foxh1, and aberrant self-renewal, we knocked-down mutant p53 or Foxh1 in p53R172H/Δ HSPCs and performed CFU and immunophenotyping assays. Knockdown of Foxh1 ablated the replating capacity of these cells (Figure 7A), phenocopying in large part the effects observed upon mutant p53 knockdown. Furthermore, similar to mutant p53 ablation, Foxh1 suppression leads to the downregulation of the stem cell factor receptor cKit and the stem cell antigen 1 (Sca-1) in p53R172H/Δ HSPCs (but not in p53 deficient cells), which is functionally accompanied by coarsening of chromatin and nuclear segmentation, morphological changes indicative of differentiation (Figure 7B and Figure S7A and S7B). Conversely, over-expression of Foxh1 in both p53WT/WT and p53Δ/Δ HSPCs endowed these cells with increased colony forming ability (Figure 7C) and also led to the up-regulation of cKit and Sca-1 along with concomitant down-regulation of Gr1/Cd11b (Figure 7D and Figure S7C). Analogous results were also observed in leukemia cells, where suppression of either mutant p53 or Foxh1 in two p53 mutant AML lines reduced cKit expression (Figure S8A–D) and led to cellular differentiation determined by morphological criteria (Figure S8E and S8F) and transcriptional profiling (Figure S8G). Finally, to determine whether Foxh1 could phenocopy mutant p53 effects, we performed an epistasis experiment using p53R172H/Δ HSPCs where mutant p53 was suppressed and cells were co-transduced to overexpress either a Foxh1 cDNA or control vector and examined for self-renewal capacity in serial replating assays. Enforced Foxh1 expression partially reversed the loss of replating potential produced by suppressing mutant p53 (Figure 7E), suggesting that Foxh1 functions downstream of mutant p53 to enforce aberrant self-renewal. Collectively, these data demonstrate that Foxh1 is necessary and sufficient to support an aberrant self-renewal phenotype in both pre-malignant and malignant p53 mutant cells.
Figure 7. Foxh1 is necessary and sufficient for the enhanced self-renewal phenotype produced by mutant p53.
(A) Total number of colony-forming units (CFU) generated by p53R172H/Δ HSPCs containing sh.p53 (blue), sh.Ren (black), shFoxH1.1072 (1) (plum), shFoxH1.1116 (2) (maroon). Data are representative of three experiments. Error bars correspond to mean ± s.e.m (n=3). Lower panels: Western-blotting intensities for FOXH1 for the different conditions. (B) cKit+ and Sca-1+ expression analyzed by FACS in sh.Ren, sh.p53 and sh.FoxH1 containing HSPCs from p53WT, p53Δ/Δ and p53R172H/Δ mice. Data are representative of three experiments. (C) Total number of colony-forming units (CFU) generated by p53Δ/Δ and p53WT HSPCs containing pLVX Control (black) or pLVX Foxh1 cDNA (grey). Data are representative of three experiments. Error bars correspond to mean ± s.e.m (n=3). Lower panel: Western-blotting for FOXH1 for the different conditions. (D) Percentage of cKit+Sca-1+ HSPCs analyzed by FACS in p53Δ/Δ and p53WT cells containing pLVX Control or pLVX Foxh1 cDNA. Data are representative of three experiments. Error bars correspond to mean ± s.e.m (n=3). (E) Total number of colony-forming units (CFU) generated by p53R172H/Δ HSPCs containing sh.Ren + pLVX Control (black), sh.Ren + pLVX Foxh1 cDNA (grey), sh.p53 + pLVX Control (light blue), sh.p53 + pLVX Foxh1 cDNA (dark blue). Data are representative of three experiments. Error bars correspond to mean ± s.e.m (n=3) * p <0.05, ** p <0.005, *** p <0.0005.
Discussion
While mutations in the TP53 tumor suppressor gene ablate its anti-proliferative functions, some mutant p53 alleles also appear to produce “gain-of-function” phenotypes that display more profound phenotypes than those produced by complete p53 loss. Nonetheless, studies investigating the basis for these effects have failed to coalesce around a common biological or molecular mechanism and, indeed, a recent comprehensive analysis of DNA sequence and functional genomics data suggested that the primary selective pressure for the appearance of p53 mutation in cancer arises from their purely inactivating and/or dominant-negative effects (51). Here we studied the action of the most common p53 mutant allele in the initiation and maintenance of acute myeloid leukemia, a well characterized liquid tumor where p53 loss is linked to aggressive and chemoresistant disease and one in which the potential GOF effects of mutant p53 have not been previously examined. We show that this p53 mutant accelerates leukemia onset and, strikingly, is required for leukemia maintenance, such that suppression of mutant p53 eliminates leukemia initiating potential.
The biological basis of AML involves the emergence of leukemic blasts that have acquired mutations that drive increased proliferation of stem and progenitor cells, block their maturation, and promote their aberrant self-renewal (52,53). In our models, the murine p53R172H mutant contributes to the latter activity, such that its elimination in leukemic cells releases the differentiation block. Similar results are observed with human AML cells harboring the equivalent R175H allele and likely explain the apparent requirement of certain other leukemia cells to additional mutant p53 alleles (e.g. R248Q; see Figure S1B – (27)). Taking advantage of well-validated assays of self-renewal in the hematopoietic system, we show that p53R172H acts in non-transformed hematopoietic cells to promote self-renewal to a substantially greater degree than does p53 loss, and that suppression of mutant p53 reverses these effects. Thus, in AML, mutant p53 can produce phenotypes analogous to bone fide oncogenes, such as the MLL-AF9 fusion oncoprotein, which also contributes to leukemia by enforcing aberrant self-renewal and exerts a continuous dependency for disease maintenance (54–56).
While most TP53 mutations disrupt its DNA binding capabilities, the resulting proteins are frequently stable and retain an intact transcriptional activation domain (3,29). Some studies suggest that mutant p53 can be recruited to chromatin to induce gene expression via interaction with other transcriptional regulators, such as NF-Y and ETS2 (29,57–59). Others studies support a model where mutant p53 proteins can directly bind to and perturb the function of other transcription factors, including p63, p73, and SREBP (12,13,57–59). In the AML system studied herein, mutant p53 increases the expression of the Foxh1 transcription factor, leading to its significant upregulation compared to p53Δ/Δ leukemias. Functional studies revealed that p53R172H is necessary and sufficient to induce Foxh1 which, in turn, binds stem cell-associated genes and contributes to enhanced self-renewal. The precise details of how mutant p53 induces Foxh1 remain to be determined and, to date, we have been unable to detect a robust ChIP signal for mutant p53 on DNA or validate an interaction with other p53 family members. Nevertheless, the importance of this molecular axis is supported by the conserved ability of mutant p53 to increase Foxh1 expression cross species and by the significant upregulation Foxh1 expression in CK-AML, a leukemia subtype characterized by a high frequency of TP53 mutations.
Foxh1 is expressed in embryonic stem cells and during early embryonic development. There, it is induced by the TGF-β family member Nodal (60) and acts to promote an epithelial to mesenchymal transition that takes place during gastrulation (42,61). Foxh1 is not normally expressed in the adult hematopoietic system and, as such, mutant p53 appears to hijack Foxh1-driven programs to promote leukemia by enforcing the expression of genes normally involved in promoting stemness and cell plasticity during hematopoiesis. Like the p53R172H allele, p53-deficiency – as produced by two homozygous null p53 alleles – can also accelerate AML development by promoting aberrant self-renewal and enhancing leukemia initiating potential (18). However, the aberrant program produced by the p53 null state is not as potent as that produced by p53R172H and does not depend on elevated levels of Foxh1. Presumably, this distinct rewiring of self-renewal programs explains why cells differentiate and lose leukemia initiating capacity upon mutant p53 elimination to produce a p53 null state, the latter event being known to promote leukemogenesis if present as an initiating event. Thus, in this context, mutant p53 GOF produces a more profound phenotype than p53 loss while concomitantly creating unique molecular dependencies.
Our results may also have relevance to the pathogenesis of human AML. Specifically, beyond the GOF effect of p53R172H in leukemogenesis, this work reveals that mutant p53 can exert its phenotypes even prior to leukemia onset, leading to the continuous self-renewal of HSPCs in vitro and in vivo. Most previous GOF studies of mutant p53 have been in the context of established cancer and, to our knowledge, this is the first study to reveal that a GOF activity can produce molecular dependencies prior to neoplastic transformation. The effect is such that elimination of the mutant protein in pre-malignant cells promotes differentiation, impairs colony-forming ability, and reduces competitive fitness during long-term hematopoiesis in vivo. While TP53 mutations are considered a later event during the progression of many solid tumors (62), they are present as “truncal” mutations with a high variant allele frequency in human AML such that we identified 35 AML cases with the R175H mutation and a median variant allele frequency (VAF) of 36% that showed evidence of clonal or sub-clonal presentation. (63). Indeed, TP53 mutations are often observed in clonal hematopoiesis (CH), a phenomenon where stem cells harboring leukemia-associated mutations expand in the peripheral blood of individuals in more than 0.02 VAF without an overt disease (64–66). We identified 9 cases of CH that harbored TP53 R175H mutations with a high VAF that ranged from 2–9%. Strikingly, 3 out of 9 cases (~33%) only harbored a single mutation - TP53 R175H - strongly suggesting that this is an early event. Of the remaining 6 cases, 2 individuals harbored clonal TP53 R175H mutations while the remaining 4 contained sub-clonal mutations. Furthermore, individuals presenting with clonal hematopoiesis involving a TP53 mutation have the highest risk for progressing to AML (65). Taken together, these observations strongly suggest that TP53R175H mutation is an early, “first-hit” event in the pathogenesis of human AML.
Our data predict that some cancers might depend on continuous expression of mutant TP53 and are consistent with studies on the p53R248Q mutant protein in mouse models of lymphoma, colon cancer, and others (67,68). Nevertheless, it seems unlikely that all mutant TP53 alleles have GOF activity, and even those that do may impinge on distinct molecular mechanisms depending on context. For example, while p53R172H can display GOF effects by promoting invasion and metastasis in a pancreas cancer model, it is not required for the sustained proliferation of pancreatic cancer cells, at least in vitro (12). Moreover, a recent report analyzing broad functional genomic datasets concluded that mutant TP53 was generally dispensable for the proliferation of human cell lines in culture (51). However, this study only assessed the effects of gene disruption on proliferation under in vitro conditions and did not examine signals associated with allele specificity or cellular context that are highly prevalent in the biology of p53 (7), whereas studies in vivo have confirmed a greater advantage of some of the hotspot mutations (69). Indeed, our functional experiments show a requirement for p53R175H in the leukemia initiating potential of a human AML line, which could have been missed by performing a different type of analysis. Additionally, upon detailed inspection of publicly available CRISPR screening data, we note that some TP53 mutant lines appear to depend on mutant TP53 for their survival (Figure S1B) (27). Dissecting the underlying mechanism of this context specificity will be important to determine when and where mutant p53 proteins and their associated molecular programs may represent therapeutic targets.
Still, despite the perplexing range of molecular and biological mechanisms that have been attributed to GOF p53 mutations, our studies reveal convergent features between mutant p53 GOF action in solid and liquid tumors, as well as in experimental models of induced pluripotency (43). In solid tumors, the best characterized phenotype associated with TP53 GOF mutations involves their ability to drive invasion and metastasis (11,12,68,70,71), which is often associated with an EMT and cellular plasticity that enables the acquisition of a more stem cell like state. Consistent with this, the transcriptional profiles of p53 mutant tumors are enriched for gene signatures expressed in embryonic stem cells (72–74). In AML, p53R172H promotes self-renewal, as well as a transcriptional landscape enriched for gene signatures linked to stemness and, intriguingly, EMT. One such gene is Foxh1, which is known to promote EMT during embryonic development, yet we demonstrate here that it can drive aberrant self-renewal in AML. Remarkably, these same players contribute to the generation of iPS cells by the Yamanaka factors: p53R172H enhances iPS cell formation beyond p53 loss (15) and Foxh1 cooperates with p53-deficiency to achieve a similar effect (43). Collectively, these observations point to a convergent action of p53 GOF on cellular plasticity and stemness, which can influence distinct cancer processes depending on context.
METHODS
Animal models
All mouse experiments were approved by the Institutional Animal Care and Use Committee at Memorial-Sloan Kettering Cancer Center. p53LSLR172H and p53LoxP mice were described previously (11). C57BL/6 (B6.SJL-Ptprca Pepcb/BoyJ (CD45.1) mice, thymectomized mice, Vav1Cre and Mx1Cre mice were purchased from Jackson Laboratory and bred in a mouse facility at MSKCC.
Primary cell culture
Total bone marrow cells from p53R172H/F;Vav1Cre, p53F/F;Vav1Cre and p53F/F (WT) mice (6–8 weeks) or purified bone marrow cKit+ cells were transduced as described below with sh.Ren or sh.p53 retrovirus and plated in methylcellulose medium (Methocult M3434, Stem Cell Technologies). Cells were seeded at 10,000 total bone marrow cells or 4,000 purified cKit+ cells per replicate. Colony forming units were enumerated using a Zeiss Axio Observer microscope, and cells were re-plated (4,000 cells/replicate) every 7–10 days. Primary mouse PMN leukemias derived from p53R172H/F;Vav1Cre and p53F/F;Vav1Cre mice were cultured in 1:1 DMEM to IMDM with 10% FBS (Gibco), 100 U/mL penicillin, 100 μg/ml streptomycin, 50 μM 2-mercaptoethanol and recombinant mouse SCF (50 ng/ml), IL3 (10 ng/ml) and IL6 (10 ng/ml) at 37°C in 5% CO2. Puro (puromycin) and Dox (doxycycline) were used at a final concentration of 1 μg/ml for all cell culture assays.
Cell lines
The human leukemia line KY821 (JCRB cell bank) was grown in RPMI 1640 and 20% FBS (Gibco). KL922 (p53Δ/Δ), KL973 (p53R172H/11Β3) and KL974 (p53R172H/Δ) primary mouse cell lines (19,25) were cultured with RPMI, 10% FBS (Gibco) and recombinant mouse SCF (50 ng/ml), IL3 (10 ng/ml) and IL6 (10 ng/ml). PlatinumE (75) cells were a gift from Craig Thomson’s lab and HEK293T were purchased from ATCC (CRL-3216) and were used for retroviral production while maintained in DMEM with 10% FBS. Media were supplemented with 100 U/mL penicillin, 100 μg/ml streptomycin and cultures were incubated at 37°C, 5% CO2. All cells were tested negative for mycoplasma (MP0025–1KT) and were not maintained for longer than 10 passages in culture. For cell line authentication we confirmed TP53 expression by WB and Sanger sequencing confirmed the specific mutation.
Xenotransplantation
Experiments were carried out in accordance with institutional guidelines approved by MSKCC. Immune-deficient NSG mice (6–10 weeks-old) mice were bred under pathogen-free conditions. and were sub-lethally irradiated (200–250 cGy) 3–24 hours before transplantation. KY821 cells were injected by tail-vain injection. CBC analysis was performed on peripheral blood collected from submandibular bleeding using a Drew Hemavet Analyzer (Veterinary Diagnostics).
Histology and microscopy
Tissues were dissected from mice for fixation overnight in 10% formalin (Fisher). Bones were decalcified per manufacturer’s instructions (IDEXX BioResearch), and fixed tissues were dehydrated and embedded in paraffin for sectioning. Paraffin sections (5 μm) were prepared and stained with hematoxylin and eosin (H&E) (Leica Autostainer XL).
Flow cytometry
Single cell suspensions were prepared from bone marrow, spleen or peripheral blood. Red blood cells were lysed with ammonium-chloride-potassium (ACK) buffer, and the remaining cells were resuspended in PBS with 3% FBS. Non-specific antibody binding was blocked by incubation with 20 μg/ml Rat IgG (Sigma-Aldrich) for 15 min, and cells were then incubated with the indicated primary antibodies for 30 min on ice. Stained cells were quantified using a Fortessa analyzer (BD Biosciences) or isolated with a FACSAria II (BD Biosciences). FlowJo software (TreeStar) was used to generate flow cytometry plots. More details on FACS antibodies found in Table S2.
Bone marrow competitive transplantation
Femurs and tibiae were isolated from donor p53F/F (WT), p53F/F;Mx1Cre or p53R172H/F;Mx1Cre (CD45.2), as well as WT (CD45.1) mice. Bones were flushed with PBS and the single-cell suspension was centrifuged (5 min, 0.5g, 4 °C) and treated with red cell ACK lysis buffer, as described above. Total nucleated bone marrow cells were re-suspended in PBS, passed through a 40 μM cell strainer and counted. Donor cells (1 × 106 per genotype, per mouse) were mixed 1:1 with support bone marrow cells (CD 45.1), and transplanted via retro-orbital injection into lethally irradiated (950 Rad) CD45.1 recipient mice. Chimerism was monitored by flow cytometry (anti-CD45.1 and anti-CD45.2, BD Bioscience) of peripheral blood at 4-week intervals post-transplant three weeks after last pIpC injection and for 16 weeks at which time mice were sacrificed and chimerism was assessed in other hematopoietic compartments (bone marrow).
Retroviral transduction
Hematopoietic stem and progenitor cells were isolated by sorting Lin−cKit+Sca1+ cells or enriched using an autoMACS Pro separator (Miltenyi Biotec) using CD117 magnetic microbeads (Miltenyi Biotech, 130–091-224), then cultured in the presence of 50 ng/ml SCF, 10 ng/ml IL-3, and 10 ng/ml IL-6, and infected with concentrated supernatants of LMNe-sh.Ren/sh.p53-BFP or MLPe sh.Ren/sh.p53/sh.FoxH1 retrovirus after 24 and 48 h (more details about the shRNAs in Table S2). Transduction efficiency was determined by reporter fluorescence at 48 h, and 5,000 sorted cells were injected into CD45.1 mice as described above.
Cell viability and DNA synthesis
Apoptosis assays were performed by staining with Annexin V (BD Pharmingen) according to the manufacturer’s instructions, in combination with DAPI. For cell cycle studies, BrdU (BD Pharmingen) was used per manufacturer’s instructions and stained with 2 μg/ml DAPI prior to flow cytometric analysis.
Quantitative RNA expression assays
Total RNA was extracted from cells using the RNeasy Plus Mini Kit (Qiagen). RNA quantity and quality were determined using an Agilent 2100 Bioanalyzer. RNA-seq libraries were prepared from total RNA by polyA selection using oligo-dT beads (Life Technologies) according to the manufacturer’s instructions. The resulting poly-A+ RNA served as input for library construction using standard Illumina protocols. Sequencing was performed on an Illumina HiSeq 2500 sequencer using 50 bp pair-end reads. For mRNA quantification, total RNA was used for cDNA synthesis (Agilent). Real-time PCR reactions were carried out using SYBR Green Master Mix (QuantaBio) and a Via7 (AB Applied biosystems).
RNA sequencing analysis
RNA-Seq library construction and sequencing were performed at the integrated genomics operation (IGO) Core at MSKCC. Poly-A selection was performed. For sequencing, approximately 30 million 50bp paired-end reads were acquired per replicate condition. Resulting RNA-Seq data was analyzed by removing adaptor sequences using Trimmomatic (76). RNA-Seq reads were then aligned to GRCh37.75 (hg19) with STAR (77) and genome-wide transcript counting was performed by HTSeq (78) to generate a matrix of fragments per kilobase of exon per million fragments mapped (RPKM). Gene expressions of RNA-Seq data were managed by hierarchical clustering based on one minus Pearson correlation test using Morpheus (https://software.broadinstitute.org/morpheus/). All results from the RNA-Seq experiments can be found under GSE125097.
Gene set enrichment analysis
Gene set enrichment analysis was performed using gene set as permutation type, 1,000 permutations and log2 ratio of classes, or with gene set and Signal2Noise as metrics for ranking genes. Gene sets used in this study were identified from the Molecular Signatures Database (MSigDB Curated v4.0). Gene pathways and functions were assessed using Ingenuity Pathway Analysis (Qiagen Bioinformatics).
Expression of gene signatures in human AML and normal cells.
Visualization of gene signatures derived from transcriptional data as Log2 expression values for different karyotypes of AML patients described previously in GSE42519 (79). For normal hematopoietic cells values were averaged over all genes in each signature and visualized by using Sinaplot. Student’s t-test was performed using the R programming language (https://www.R-project.org/).
Quantification and Statistical Analysis
All p-values were calculated using unpaired two-tailed Student’s t test with Graphpad Prism software, unless otherwise described in the methods or figure legends. No specific randomization or blinding protocol was used for these analyses. Statistically significant differences are indicated with asterisks in figures with the accompanying p-values in the legend. Error bars in figures indicate standard deviation (SD) or standard error of the mean (SEM) for the number of replicates, as indicated in the figure legend.
Plasmids and viral production
All vectors were derived from the Murine Stem Cell Virus (MSCV, Clontech) retroviral vector backbone. miRE-based shRNAs were designed and cloned as previously described (26) into the LT3GEPIR (TRE3G-GFP-miRE-PGK-PuroR-IRES-rtTA3) vector (80). The mouse Foxh1 cDNA was cloned in a pVXL-hygromycin-mFoxh1 vector (provided by Yilong Zou, Massagué lab, MSKCC). All constructs were verified by sequencing. Lentiviruses were produced by co-transfection of 293T cells with 10 ug LT3GEPIR construct and helper vectors (3.75 ug psPAX2 and 1.25 ugVSV-G). For retroviral infection PlatinumE cells were plated in 15 cm diameter dishes each transfected with 54 ug of MSCV vectors and 2.7ug of VSV-G and 8.1ug of pCl-eco. Transfection of packaging cells was performed using HBS and CaCl2 or Mirus transfection reagent (MIR 2304). Viral supernatants were passed through a 0.45 um filter (Millipore) and concentrated with Centrifugal Filter units (Millipore) to obtain high virus titer. Concentrated virus was supplemented with 8 ug/ml of polybrene (Sigma) before adding to target cells.
Immunoblotting.
For protein lysates, cells were incubated with RIPA buffer supplemented with protease inhibitors (Protease inhibitor tablets, Roche) for 30 min and cleared by centrifugation (15 min 14.000 rpms 4C). Protein was quantified using the DC protein assay (BioRad). The following antibodies were used for immunoblotting: β-ACTIN (ac-15, Sigma), mTP53 (CM5, Leica microsystems), hTP53 (DO-1, Santa Cruz), mFoxh1 (Abcam 49133) and hFOXH1 (Abcam 102590).
Chromatin immunoprecipitation (ChIP).
Chromatin immunoprecipitation was performed as previously described (81). Briefly, KL974 (p53R172H/Δ) cells were fixed with 1% formaldehyde for 15 min, and the cross-linking reaction was stopped by adding 125mM glycine. Cells were washed twice with cold PBS and lysed in buffer (150 mM NaCl, 1%v/v Nonidet P-40, 0.5% w/v deoxycholate, 0.1% w/v SDS, 50 mM Tris pH8, 5m M EDTA) supplemented with protease inhibitors. Cell lysates were sonicated using a Covaris E220 Sonicator to generate fragments less than 400 bp. Sonicated lysates were centrifuged, and incubated overnight at 4°C with specific antibodies (Foxh1 Abcam 49133;). Immunocomplexes were recovered by incubation with 30 ul protein A/G magnetic beads (Thermofisher) for 2h at 4°C. Beads were sequentially washed twice with RIPA buffer, increasing stringency ChIP wash buffers (150 mM NaCl, 250 mM NaCl, 250 mM LiCl) and finally TE buffer. Immunocomplexes were eluted using elution buffer (1% SDS, 100mM NaHCO3), and cross-linking was reverted by addition of 300 mM NaCl and incubation at 65°C overnight. DNA was purified using a PCR purification kit (Qiagen). Input chromatin was used for estimation of relative enrichment. The results for the ChIP-seq experiments can be found under GSE125097.
ChIP-Seq Library Preparation, Illumina Data Analysis and Peak Detection.
ChIP-Seq libraries were prepared at the Center for Epigenetic Research (MSKCC) using the NEBNext® ChIP-Seq Library Prep Master Mix Set for Illumina® (New England BioLabs) following the manufacture’s instructions. Reads were trimmed for quality and Illumina adapter sequences using ‘trim_galore’ before aligning to mouse assembly mm9 with bowtie2 using the default parameters. Aligned reads with the same start site and orientation were removed using the Picard tool MarkDuplicates (http://broadinstitute.github.io/picard/). Density profiles were created by extending each read to the average library fragment size and then computing density using the BEDTools suite (http://bedtools.readthedocs.io). Enriched regions were discovered using MACS2 and scored against matched input libraries (fold change > 2 and p-value < 0.005). Dynamic regions between two conditions were discovered using a similar method, with the second ChIP library replacing input. Peaks were then filtered against genomic ‘blacklisted’ regions (http://mitra.stanford.edu/kundaje/akundaje/release/blacklists/mm9-mouse/mm9blacklist.bed.gz) and those within 500 bp were merged. All genome browser tracks and read density tables were normalized to a sequencing depth of ten million mapped reads. Dynamic peaks were annotated using linear genomic distance and motif signatures were obtained using the ‘de novo’ approach with Homer v4.5 (http://homer.ucsd.edu/).
Supplementary Material
SIGNIFICANCE.
Our study demonstrates how a gain-of-function p53 mutant can hijack an embryonic transcription factor to promote aberrant self-renewal. In this context mutant p53 functions as an oncogene to both initiate and sustain myeloid leukemia and suggests a potential convergent activity of mutant p53 across cancer types.
ACKNOWLEDGMENTS
The authors are grateful to Charles J. Sherr for extensive scientific discussion, suggestions for experiments, and help in formulating and editing the manuscript. We thank Omar Abdel-Wahab Ross L. Levine, and Scott Armstrong for their key input and reagents, and Elli Papaemmanuil for providing analysis of human AML datasets. We also thank members of the Lowe laboratory for comments and discussions, and Yilong Zou and Qiong Wang for providing us with the mFoxh1–expressing vector and Jiajun Zhu for helping optimizing ChIP-seq conditions.
Financial Support: This work was supported by a grant from the National Cancer Institute (R01 CA190261) and the Memorial Sloan Kettering Cancer Center Support Grant (P30 CA008748). E.L. was supported by the Dorris J. Hutchison Student Fellowship (SKD). A.B. was supported by an EMBO long-term fellowship. E.R.K. was supported by an F31 NRSA predoctoral fellowship from the NCI/National Institutes of Health under award number F31CA192835. F.J.S.-R. is an HHMI Hanna Gray Fellow and was partially supported by an MSKCC Translational Research Oncology Training Fellowship (NIH T32-CA160001). S.W.L is an investigator in the Howard Hughes Medical Institute and the Geoffrey Beene Chair of Cancer Biology.
Footnotes
S.W.L. is a founder and member of the scientific advisory board of Mirimus, Inc., Blueprint Medicines, and ORIC Pharmaceuticals; he is also on the scientific advisory board of Constellation Pharmaceuticals and PMV Pharmaceuticals. All of the other authors declare no conflicts of interest.
REFERENCES
- 1.Kandoth C, McLellan MD, Vandin F, Ye K, Niu B, Lu C, et al. Mutational landscape and significance across 12 major cancer types. Nature 2013;502(7471):333–9 doi 10.1038/nature12634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Olivier M, Hollstein M, Hainaut P. TP53 mutations in human cancers: origins, consequences, and clinical use. Cold Spring Harb Perspect Biol 2010;2(1):a001008 doi 10.1101/cshperspect.a001008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Laptenko O, Prives C. Transcriptional regulation by p53: one protein, many possibilities. Cell Death Differ 2006;13(6):951–61 doi 10.1038/sj.cdd.4401916. [DOI] [PubMed] [Google Scholar]
- 4.Serrano M, Lin AW, McCurrach ME, Beach D, Lowe SW. Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 1997;88(5):593–602. [DOI] [PubMed] [Google Scholar]
- 5.Lowe SW, Ruley HE, Jacks T, Housman DE. p53-dependent apoptosis modulates the cytotoxicity of anticancer agents. Cell 1993;74(6):957–67. [DOI] [PubMed] [Google Scholar]
- 6.Kastan MB, Onyekwere O, Sidransky D, Vogelstein B, Craig RW. Participation of p53 protein in the cellular response to DNA damage. Cancer Res 1991;51(23 Pt 1):6304–11. [PubMed] [Google Scholar]
- 7.Kastenhuber ER, Lowe SW. Putting p53 in Context. Cell 2017;170(6):1062–78 doi 10.1016/j.cell.2017.08.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bouaoun L, Sonkin D, Ardin M, Hollstein M, Byrnes G, Zavadil J, et al. TP53 Variations in Human Cancers: New Lessons from the IARC TP53 Database and Genomics Data. Hum Mutat 2016;37(9):865–76 doi 10.1002/humu.23035. [DOI] [PubMed] [Google Scholar]
- 9.Prives C, Hall PA. The p53 pathway. J Pathol 1999;187(1):112–26 doi 10.1002/(SICI)1096–9896(199901)187:1<112::AID-PATH250>3.0.CO;2–3. [DOI] [PubMed] [Google Scholar]
- 10.Muller PA, Vousden KH. Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell 2014;25(3):304–17 doi 10.1016/j.ccr.2014.01.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Olive KP, Tuveson DA, Ruhe ZC, Yin B, Willis NA, Bronson RT, et al. Mutant p53 gain of function in two mouse models of Li-Fraumeni syndrome. Cell 2004;119(6):847–60 doi 10.1016/j.cell.2004.11.004. [DOI] [PubMed] [Google Scholar]
- 12.Weissmueller S, Manchado E, Saborowski M, Morris JPt, Wagenblast E, Davis CA, et al. Mutant p53 drives pancreatic cancer metastasis through cell-autonomous PDGF receptor beta signaling. Cell 2014;157(2):382–94 doi 10.1016/j.cell.2014.01.066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Freed-Pastor WA, Mizuno H, Zhao X, Langerod A, Moon SH, Rodriguez-Barrueco R, et al. Mutant p53 disrupts mammary tissue architecture via the mevalonate pathway. Cell 2012;148(1–2):244–58 doi 10.1016/j.cell.2011.12.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sarig R, Rivlin N, Brosh R, Bornstein C, Kamer I, Ezra O, et al. Mutant p53 facilitates somatic cell reprogramming and augments the malignant potential of reprogrammed cells. J Exp Med 2010;207(10):2127–40 doi 10.1084/jem.20100797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Yi L, Lu C, Hu W, Sun Y, Levine AJ. Multiple roles of p53-related pathways in somatic cell reprogramming and stem cell differentiation. Cancer Res 2012;72(21):5635–45 doi 10.1158/0008-5472.CAN-12-1451. [DOI] [PubMed] [Google Scholar]
- 16.Rucker FG, Schlenk RF, Bullinger L, Kayser S, Teleanu V, Kett H, et al. TP53 alterations in acute myeloid leukemia with complex karyotype correlate with specific copy number alterations, monosomal karyotype, and dismal outcome. Blood 2012;119(9):2114–21 doi 10.1182/blood-2011-08-375758. [DOI] [PubMed] [Google Scholar]
- 17.Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND, et al. Genomic Classification and Prognosis in Acute Myeloid Leukemia. N Engl J Med 2016;374(23):2209–21 doi 10.1056/NEJMoa1516192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhao Z, Zuber J, Diaz-Flores E, Lintault L, Kogan SC, Shannon K, et al. p53 loss promotes acute myeloid leukemia by enabling aberrant self-renewal. Genes Dev 2010;24(13):1389–402 doi 10.1101/gad.1940710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chen C, Liu Y, Rappaport AR, Kitzing T, Schultz N, Zhao Z, et al. MLL3 is a haploinsufficient 7q tumor suppressor in acute myeloid leukemia. Cancer Cell 2014;25(5):652–65 doi 10.1016/j.ccr.2014.03.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zuber J, Radtke I, Pardee TS, Zhao Z, Rappaport AR, Luo W, et al. Mouse models of human AML accurately predict chemotherapy response. Genes Dev 2009;23(7):877–89 doi 10.1101/gad.1771409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ogilvy S, Elefanty AG, Visvader J, Bath ML, Harris AW, Adams JM. Transcriptional regulation of vav, a gene expressed throughout the hematopoietic compartment. Blood 1998;91(2):419–30. [PubMed] [Google Scholar]
- 22.Donehower LA, Harvey M, Vogel H, McArthur MJ, Montgomery CA Jr., Park SH, et al. Effects of genetic background on tumorigenesis in p53-deficient mice. Mol Carcinog 1995;14(1):16–22. [DOI] [PubMed] [Google Scholar]
- 23.Hanel W, Marchenko N, Xu S, Yu SX, Weng W, Moll U. Two hot spot mutant p53 mouse models display differential gain of function in tumorigenesis. Cell Death Differ 2013;20(7):898–909 doi 10.1038/cdd.2013.17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Kuhn R, Schwenk F, Aguet M, Rajewsky K. Inducible gene targeting in mice. Science 1995;269(5229):1427–9. [DOI] [PubMed] [Google Scholar]
- 25.Liu Y, Chen C, Xu Z, Scuoppo C, Rillahan CD, Gao J, et al. Deletions linked to TP53 loss drive cancer through p53-independent mechanisms. Nature 2016;531(7595):471–5 doi 10.1038/nature17157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zuber J, McJunkin K, Fellmann C, Dow LE, Taylor MJ, Hannon GJ, et al. Toolkit for evaluating genes required for proliferation and survival using tetracycline-regulated RNAi. Nat Biotechnol 2011;29(1):79–83 doi 10.1038/nbt.1720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wang T, Yu H, Hughes NW, Liu B, Kendirli A, Klein K, et al. Gene Essentiality Profiling Reveals Gene Networks and Synthetic Lethal Interactions with Oncogenic Ras. Cell 2017;168(5):890–903 e15 doi 10.1016/j.cell.2017.01.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Brummelkamp TR, Bernards R, Agami R. A system for stable expression of short interfering RNAs in mammalian cells. Science 2002;296(5567):550–3 doi 10.1126/science.1068999. [DOI] [PubMed] [Google Scholar]
- 29.Weisz L, Oren M, Rotter V. Transcription regulation by mutant p53. Oncogene 2007;26(15):2202–11 doi 10.1038/sj.onc.1210294. [DOI] [PubMed] [Google Scholar]
- 30.Pietras EM, Reynaud D, Kang YA, Carlin D, Calero-Nieto FJ, Leavitt AD, et al. Functionally Distinct Subsets of Lineage-Biased Multipotent Progenitors Control Blood Production in Normal and Regenerative Conditions. Cell Stem Cell 2015;17(1):35–46 doi 10.1016/j.stem.2015.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bagger FO, Kinalis S, Rapin N. BloodSpot: a database of healthy and malignant haematopoiesis updated with purified and single cell mRNA sequencing profiles. Nucleic Acids Res 2019;47(D1):D881–D5 doi 10.1093/nar/gky1076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Passegue E, Jamieson CH, Ailles LE, Weissman IL. Normal and leukemic hematopoiesis: are leukemias a stem cell disorder or a reacquisition of stem cell characteristics? Proc Natl Acad Sci U S A 2003;100 Suppl 1:11842–9 doi 10.1073/pnas.2034201100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Nowak D, Stewart D, Koeffler HP. Differentiation therapy of leukemia: 3 decades of development. Blood 2009;113(16):3655–65 doi 10.1182/blood-2009-01-198911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nakahata T, Ogawa M. Identification in culture of a class of hemopoietic colony-forming units with extensive capability to self-renew and generate multipotential hemopoietic colonies. Proc Natl Acad Sci U S A 1982;79(12):3843–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Liu Y, Elf SE, Miyata Y, Sashida G, Liu Y, Huang G, et al. p53 regulates hematopoietic stem cell quiescence. Cell Stem Cell 2009;4(1):37–48 doi 10.1016/j.stem.2008.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Matsuoka S, Ebihara Y, Xu M, Ishii T, Sugiyama D, Yoshino H, et al. CD34 expression on long-term repopulating hematopoietic stem cells changes during developmental stages. Blood 2001;97(2):419–25. [DOI] [PubMed] [Google Scholar]
- 37.Moran-Crusio K, Reavie L, Shih A, Abdel-Wahab O, Ndiaye-Lobry D, Lobry C, et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 2011;20(1):11–24 doi 10.1016/j.ccr.2011.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Challen GA, Sun D, Jeong M, Luo M, Jelinek J, Berg JS, et al. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat Genet 2011;44(1):23–31 doi 10.1038/ng.1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Seita J, Weissman IL. Hematopoietic stem cell: self-renewal versus differentiation. Wiley Interdiscip Rev Syst Biol Med 2010;2(6):640–53 doi 10.1002/wsbm.86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Muller PA, Vousden KH. p53 mutations in cancer. Nat Cell Biol 2013;15(1):2–8 doi 10.1038/ncb2641. [DOI] [PubMed] [Google Scholar]
- 41.Chiu WT, Charney Le R, Blitz IL, Fish MB, Li Y, Biesinger J, et al. Genome-wide view of TGFbeta/Foxh1 regulation of the early mesendoderm program. Development 2014;141(23):4537–47 doi 10.1242/dev.107227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Attisano L, Silvestri C, Izzi L, Labbe E. The transcriptional role of Smads and FAST (FoxH1) in TGFbeta and activin signalling. Mol Cell Endocrinol 2001;180(1–2):3–11. [DOI] [PubMed] [Google Scholar]
- 43.Takahashi K, Tanabe K, Ohnuki M, Narita M, Sasaki A, Yamamoto M, et al. Induction of pluripotency in human somatic cells via a transient state resembling primitive streak-like mesendoderm. Nat Commun 2014;5:3678 doi 10.1038/ncomms4678. [DOI] [PubMed] [Google Scholar]
- 44.Charney RM, Forouzmand E, Cho JS, Cheung J, Paraiso KD, Yasuoka Y, et al. Foxh1 Occupies cis-Regulatory Modules Prior to Dynamic Transcription Factor Interactions Controlling the Mesendoderm Gene Program. Dev Cell 2017;40(6):595–607 e4 doi 10.1016/j.devcel.2017.02.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2012;2(5):401–4 doi 10.1158/2159-8290.CD-12-0095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 2013;6(269):pl1 doi 10.1126/scisignal.2004088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Spittau B, Krieglstein K. Klf10 and Klf11 as mediators of TGF-beta superfamily signaling. Cell Tissue Res 2012;347(1):65–72 doi 10.1007/s00441-011-1186-6. [DOI] [PubMed] [Google Scholar]
- 48.Lee KS, Kim HJ, Li QL, Chi XZ, Ueta C, Komori T, et al. Runx2 is a common target of transforming growth factor beta1 and bone morphogenetic protein 2, and cooperation between Runx2 and Smad5 induces osteoblast-specific gene expression in the pluripotent mesenchymal precursor cell line C2C12. Mol Cell Biol 2000;20(23):8783–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Kuo YH, Zaidi SK, Gornostaeva S, Komori T, Stein GS, Castilla LH. Runx2 induces acute myeloid leukemia in cooperation with Cbfbeta-SMMHC in mice. Blood 2009;113(14):3323–32 doi 10.1182/blood-2008-06-162248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Schuler A, Schwieger M, Engelmann A, Weber K, Horn S, Muller U, et al. The MADS transcription factor Mef2c is a pivotal modulator of myeloid cell fate. Blood 2008;111(9):4532–41 doi 10.1182/blood-2007-10-116343. [DOI] [PubMed] [Google Scholar]
- 51.Giacomelli AO, Yang X, Lintner RE, McFarland JM, Duby M, Kim J, et al. Mutational processes shape the landscape of TP53 mutations in human cancer. Nat Genet 2018. doi 10.1038/s41588-018-0204-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Kelly LM, Gilliland DG. Genetics of myeloid leukemias. Annu Rev Genomics Hum Genet 2002;3:179–98 doi 10.1146/annurev.genom.3.032802.115046. [DOI] [PubMed] [Google Scholar]
- 53.Takahashi S Current findings for recurring mutations in acute myeloid leukemia. J Hematol Oncol 2011;4:36 doi 10.1186/1756-8722-4-36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Armstrong SA, Staunton JE, Silverman LB, Pieters R, den Boer ML, Minden MD, et al. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat Genet 2002;30(1):41–7 doi 10.1038/ng765. [DOI] [PubMed] [Google Scholar]
- 55.Somervaille TC, Cleary ML. Identification and characterization of leukemia stem cells in murine MLL-AF9 acute myeloid leukemia. Cancer Cell 2006;10(4):257–68 doi 10.1016/j.ccr.2006.08.020. [DOI] [PubMed] [Google Scholar]
- 56.Zuber J, Rappaport AR, Luo W, Wang E, Chen C, Vaseva AV, et al. An integrated approach to dissecting oncogene addiction implicates a Myb-coordinated self-renewal program as essential for leukemia maintenance. Genes Dev 2011;25(15):1628–40 doi 10.1101/gad.17269211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Zhu J, Sammons MA, Donahue G, Dou Z, Vedadi M, Getlik M, et al. Gain-of-function p53 mutants co-opt chromatin pathways to drive cancer growth. Nature 2015;525(7568):206–11 doi 10.1038/nature15251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Di Agostino S, Strano S, Emiliozzi V, Zerbini V, Mottolese M, Sacchi A, et al. Gain of function of mutant p53: the mutant p53/NF-Y protein complex reveals an aberrant transcriptional mechanism of cell cycle regulation. Cancer Cell 2006;10(3):191–202 doi 10.1016/j.ccr.2006.08.013. [DOI] [PubMed] [Google Scholar]
- 59.Do PM, Varanasi L, Fan S, Li C, Kubacka I, Newman V, et al. Mutant p53 cooperates with ETS2 to promote etoposide resistance. Genes Dev 2012;26(8):830–45 doi 10.1101/gad.181685.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Brennan J, Lu CC, Norris DP, Rodriguez TA, Beddington RS, Robertson EJ. Nodal signalling in the epiblast patterns the early mouse embryo. Nature 2001;411(6840):965–9 doi 10.1038/35082103. [DOI] [PubMed] [Google Scholar]
- 61.Yamamoto M, Meno C, Sakai Y, Shiratori H, Mochida K, Ikawa Y, et al. The transcription factor FoxH1 (FAST) mediates Nodal signaling during anterior-posterior patterning and node formation in the mouse. Genes Dev 2001;15(10):1242–56 doi 10.1101/gad.883901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr., Kinzler KW. Cancer genome landscapes. Science 2013;339(6127):1546–58 doi 10.1126/science.1235122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Wong TN, Ramsingh G, Young AL, Miller CA, Touma W, Welch JS, et al. Role of TP53 mutations in the origin and evolution of therapy-related acute myeloid leukaemia. Nature 2015;518(7540):552–5 doi 10.1038/nature13968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Xie M, Lu C, Wang J, McLellan MD, Johnson KJ, Wendl MC, et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med 2014;20(12):1472–8 doi 10.1038/nm.3733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Abelson S, Collord G, Ng SWK, Weissbrod O, Mendelson Cohen N, Niemeyer E, et al. Prediction of acute myeloid leukaemia risk in healthy individuals. Nature 2018;559(7714):400–4 doi 10.1038/s41586-018-0317-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Steensma DP, Bejar R, Jaiswal S, Lindsley RC, Sekeres MA, Hasserjian RP, et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 2015;126(1):9–16 doi 10.1182/blood-2015-03-631747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Alexandrova EM, Yallowitz AR, Li D, Xu S, Schulz R, Proia DA, et al. Improving survival by exploiting tumour dependence on stabilized mutant p53 for treatment. Nature 2015;523(7560):352–6 doi 10.1038/nature14430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Schulz-Heddergott R, Stark N, Edmunds SJ, Li J, Conradi LC, Bohnenberger H, et al. Therapeutic Ablation of Gain-of-Function Mutant p53 in Colorectal Cancer Inhibits Stat3-Mediated Tumor Growth and Invasion. Cancer Cell 2018;34(2):298–314 e7 doi 10.1016/j.ccell.2018.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Kotler E, Shani O, Goldfeld G, Lotan-Pompan M, Tarcic O, Gershoni A, et al. A Systematic p53 Mutation Library Links Differential Functional Impact to Cancer Mutation Pattern and Evolutionary Conservation. Mol Cell 2018;71(1):178–90 e8 doi 10.1016/j.molcel.2018.06.012. [DOI] [PubMed] [Google Scholar]
- 70.Morton JP, Timpson P, Karim SA, Ridgway RA, Athineos D, Doyle B, et al. Mutant p53 drives metastasis and overcomes growth arrest/senescence in pancreatic cancer. Proc Natl Acad Sci U S A 2010;107(1):246–51 doi 10.1073/pnas.0908428107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Lang GA, Iwakuma T, Suh YA, Liu G, Rao VA, Parant JM, et al. Gain of function of a p53 hot spot mutation in a mouse model of Li-Fraumeni syndrome. Cell 2004;119(6):861–72 doi 10.1016/j.cell.2004.11.006. [DOI] [PubMed] [Google Scholar]
- 72.Shibue T, Weinberg RA. EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat Rev Clin Oncol 2017;14(10):611–29 doi 10.1038/nrclinonc.2017.44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Mizuno H, Spike BT, Wahl GM, Levine AJ. Inactivation of p53 in breast cancers correlates with stem cell transcriptional signatures. Proc Natl Acad Sci U S A 2010;107(52):22745–50 doi 10.1073/pnas.1017001108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Ben-Porath I, Thomson MW, Carey VJ, Ge R, Bell GW, Regev A, et al. An embryonic stem cell-like gene expression signature in poorly differentiated aggressive human tumors. Nat Genet 2008;40(5):499–507 doi 10.1038/ng.127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Morita S, Kojima T, Kitamura T. Plat-E: an efficient and stable system for transient packaging of retroviruses. Gene Ther 2000;7(12):1063–6 doi 10.1038/sj.gt.3301206. [DOI] [PubMed] [Google Scholar]
- 76.Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 2014;30(15):2114–20 doi 10.1093/bioinformatics/btu170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013;29(1):15–21 doi 10.1093/bioinformatics/bts635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Anders S, Pyl PT, Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics 2015;31(2):166–9 doi 10.1093/bioinformatics/btu638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Rapin N, Bagger FO, Jendholm J, Mora-Jensen H, Krogh A, Kohlmann A, et al. Comparing cancer vs normal gene expression profiles identifies new disease entities and common transcriptional programs in AML patients. Blood 2014;123(6):894–904 doi 10.1182/blood-2013-02-485771. [DOI] [PubMed] [Google Scholar]
- 80.Fellmann C, Hoffmann T, Sridhar V, Hopfgartner B, Muhar M, Roth M, et al. An optimized microRNA backbone for effective single-copy RNAi. Cell Rep 2013;5(6):1704–13 doi 10.1016/j.celrep.2013.11.020. [DOI] [PubMed] [Google Scholar]
- 81.Banito A, Li X, Laporte AN, Roe JS, Sanchez-Vega F, Huang CH, et al. The SS18-SSX Oncoprotein Hijacks KDM2B-PRC1.1 to Drive Synovial Sarcoma. Cancer Cell 2018;33(3):527–41 e8 doi 10.1016/j.ccell.2018.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.