Abstract
Background
Analysis of melanomas for actionable mutations has become the standard of care. Recently, a classification scheme has been proposed that categorizes BRAF mutations based on their mechanisms for activation of the MAPK pathway.
Methods
In this analysis BRAF, KIT, NRAS, and PIK3CA mutations were examined by next generation sequencing (NGS) in 446 melanomas in a clinical diagnostic setting. KRAS and HRAS were also analyzed to elucidate coexisting BRAF and RAS mutations. BRAF mutations were categorized into class-1 (kinase-activated, codon 600), class-2 (kinase-activated, non-codon 600) and class-3 (kinase-impaired), based on the newly proposed classification scheme.
Results
NGS demonstrated high analytic sensitivity. Among 355 mutations detected, variant allele frequencies were 2–5% in 21 (5.9%) mutations and 2–10% in 47 (13%) mutations. Mutations were detected in BRAF (42%), NRAS (25%), KIT (4.9%) and PIK3CA (2.7%). The incidence of class-1, class-2 and class-3 mutations were 33% (26% p.V600E and 6.1% p.V600K), 3.1 and 4.9% respectively. With a broader reportable range of NGS, class-1, class-2 and class-3 mutations accounted for 77, 7.4 and 12% of all BRAF mutations. Class-3 mutations, commonly affecting codons 594, 466 and 467, showed a higher incidence of coexisting RAS mutations, consistent with their RAS-dependent signaling. Significant association with old age and primary tumors of head/neck/upper back suggest chronic solar damage as a contributing factor for melanomas harboring BRAF p.V600K or class-3 mutations.
Conclusion
This study categorizes the range, frequency, coexisting driver mutations and clinical characteristics of the three classes of BRAF mutations in a large cohort of melanomas in a clinical diagnostic setting. Further prospective studies are warranted to elucidate the clinical outcomes and benefits of newly developed targeted therapy in melanoma patients carrying each class of BRAF mutation.
Electronic supplementary material
The online version of this article (10.1186/s12885-019-5864-1) contains supplementary material, which is available to authorized users.
Keywords: BRAF, NRAS, Melanoma, Kinase-impaired, Categorization
Background
Integrated multiplatform analyses including whole exon sequencing and whole genome sequencing have uncovered the landscape of genomic alterations and important implications for prognosis and therapy in different subtypes of melanomas [1–4]. Most melanomas had potentially actionable mutations in the mitogen-activated protein kinase (MAPK) pathway and phosphatidylinositol 3-kinase (PI3K/AKT/mTOR) pathway. These included BRAF mutations affecting codon 600, NRAS mutations affecting codon 61 (and less frequently codons 12 and 13), and KIT mutations within exons 9 and 11. Monotherapy with BRAF inhibitors and combined therapy with BRAF inhibitors and MEK inhibitors have been approved by the Food and Drug Administration in the United States of America and other countries worldwide for targeted therapy of metastatic melanoma [5–10]. In NRAS-mutated advanced melanoma, a progression-free survival benefit was observed in patients treated with binimetinib (a MEK inhibitor) compared with dacarbazine [11]. Tyrosine kinase inhibitors (imatinib, nilotinib and dasatinib) have also shown benefits for melanoma patients with activating KIT mutations [12–14]. Mutation detection among these genes is recommended for standard of care targeted therapy in patients with metastatic melanoma.
Cobas 4800, a real-time polymerase chain reaction (PCR) mutational assay, was the first FDA-approved companion diagnostic test to detect the BRAF p.V600E mutation [15]. However, next generation sequencing (NGS) platforms have become popular in the clinical diagnostic setting for simultaneous mutational profiling of a panel of genes for melanomas and other cancers [16, 17]. Using NGS assays with a higher analytic sensitivity and a broader reportable range, a variety of BRAF mutations were detected within exon 11 and 15 [18–20]. We previously categorized these BRAF mutations based on their impact on the kinase activity (activated versus impaired) of the protein, and proposed potential clinical implications [17, 21].
More recently, BRAF mutations are further categorized by Yao et al. based on their mechanisms for activation of the MAPK pathway [22, 23]. This classification includes class-1 (constitutive active RAS-independent monomer with high BRAF kinase activity involving codon 600), class-2 (constitutive active RAS-independent dimers with high or intermediate BRAF kinase activity involving codons outside 600, including BRAF fusion mutants), and class-3 (low or no BRAF kinase activity) mutations. Pre-clinical and clinical studies have demonstrated distinct oncogeneic mechanisms for each class of BRAF mutation, which in turn might predict different therapeutic strategies [17, 23, 24]. Previous studies, prior to the recent categorization of 3 classes of BRAF mutation, have shown impact of non-V600 BRAF mutations on disease characteristics and clinical outcomes in melanomas and colorectal cancers [18, 25–27]. Furthermore, categorization of BRAF mutations in lung cancers, based on the classification scheme proposed by Yao et al. has shown that cancers with class-2 and class-3 mutations have a more aggressive course and less favorable prognosis [28].
In this study BRAF, NRAS, KRAS, HRAS, PIK3CA and KIT mutations were examined in melanomas. We demonstrated coexisting mutations of these driver genes in melanomas, and categorize BRAF mutations based on the new classification system to elucidate their association with clinical characteristics.
Methods
Materials
A total of 502 formalin-fixed paraffin-embedded (FFPE) specimens with a diagnosis of malignant melanoma (n = 500) or melanoma in situ (n = 2) were submitted to the Molecular Diagnostics Laboratory at The Johns Hopkins Hospital between August 2013 and November 2017. The assay failed in 26 specimens (5.2%). Two metastatic specimens with prior BRAF inhibitor treatment were excluded. This include one pleural effusion specimen carrying 49% BRAF p.V600E, 19% NRAS p.Q61K and 3.3% NRAS p.Q61R coexisting mutations. The remaining 474 specimens with NGS results were submitted from 457 tumors of 455 patients (Additional file 1: Table S1). Paired specimens were submitted from 19 patients, including 9 patients with primary and metastatic tumor specimens and 8 patients with 2 metastatic tumor specimens showing the same mutational status, and 2 patients with two primary tumor specimens showing different mutational status. Eleven of 457 tumors with an estimated tumor cellularity of less than 10% and no BRAF, RAS, KIT and PIK3CA mutations detected were excluded for analysis. Among the remaining 446 tumors, there were 20 primary and/or metastatic mucosal melanomas, 2 metastatic uveal melanomas, 286 primary, recurrent or metastatic cutaneous melanomas, and 138 metastatic melanomas of unknown origin or without information of the primary site. Estimated tumor cellularity of these 446 melanoma specimens ranged from less than 10% to nearly 100%. Accompanied hematoxylin and eosin stained slides were reviewed by a pathologist (MTL). FFPE tissues from 1 to 10 unstained, 5 or 10-μm-thick sections were macro-dissected using Pinpoint reagents according the manufacturer’s protocol (ZymoResearch, Orange, CA). DNA was purified using QIAmp DNA kit (Qiagen, Valencia, CA) as described previously [29]. Concentration of DNA was determined by Qubit 2.0 Fluorometer (Life Technologies, Carlsbad, California). The Johns Hopkins Medicine institutional review board granted approval to this study.
Next generation sequencing (NGS)
NGS was conducted using AmpliSeq Cancer Hotspot Panel (v2) (Life Technologies, Carlsbad, California) for targeted multi-gene amplification as described previously [29]. Sequencing data of the targeted genes were analyzed using Torrent Suite (Life Technologies). Mutations were identified and annotated through both Torrent Variant Caller and direct visual inspection of the binary sequence alignment/map (BAM) file on the Broad Institute’s Integrative Genomics Viewer (IGV) [30]. Melanoma specimens were analyzed for BRAF (NM_004333), KIT (NM_000222), NRAS (NM_002524) and PIK3CA (NM_006218) genes for clinical reporting. BRAF mutations of the first 152 melanoma specimens were previously categorized based on their impact on the kinase activity [17]. KRAS (NM_033360) and HRAS (NM_005343) mutations were retrospectively analyzed to elucidate coexistence of RAS and kinase-impaired BRAF mutations. GNAQ (NM_002072) and GNA11 (NM_002067) mutations were retrospectively analyzed for the uveal melanomas. During our validation of this NGS assay, a cutoff of background noise at 2% was chosen for single nucleotide variations [31]. BRAF mutations were categorized according to the recently proposed classification scheme [23]. These included class-1 (high kinase activity involving codon 600), class-2 (high or intermediate kinase activity involving codons outside 600), and class-3 (impaired kinase activity) mutations.
Statistical analysis
Fisher exact test or χ2 test were performed to calculate P values.
Results
Mutational profiling of 446 melanomas
NGS revealed 118 (26%) of 446 melanomas with no mutation, 42% with BRAF mutations, 25% with NRAS mutations, 4.9% with KIT mutations, and 2.0, 2.7 and 2.7% with HRAS, KRAS and PIK3CA mutations, respectively (Table 1). Metastatic melanomas of unknown origin or without information of the primary site showed similar mutation rates to those of cutaneous melanomas, suggesting most of these cases were of cutaneous origin. Mucosal melanomas showed a higher incidence of no mutation (P < .01) and a lower incidence of BRAF mutations (P < .001), as compared to those of cutaneous melanomas. Analysis of the GNA11 and GNAQ genes in two metastatic uveal melanomas revealed GNA11 p.Q209L and GNAQ p.Q209P, respectively.
Table 1.
Mutational profiling of 446 melanomas
| Melanoma | Negative | BRAF | HRAS | KRAS | NRAS | KIT | PIK3CA |
|---|---|---|---|---|---|---|---|
| Uveal (n = 2) | 2 (100%) | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) | 0 (0%) |
| Mucosal (n = 20) | 12 (60%) | 1 (5.0%) | 0 (0%) | 1 (5.0%) | 4 (20%) | 3 (15%) | 1 (5.0%) |
| Cutaneous (n = 286)a | 72 (25%) | 132 (46%) | 9 (3.1%) | 7 (2.4%) | 64 (22%) | 14 (4.9%) | 6 (2.1%) |
| Others (n = 138)b | 32 (23%) | 56 (41%) | 0 (0%) | 4 (2.9%) | 42 (30%) | 5 (3.6%) | 5 (3.6%) |
| Total (n = 446)a | 118 (26%) | 189 (42%) | 9 (2.0%) | 12 (2.7%) | 110 (25%) | 22 (4.9%) | 12 (2.7%) |
a11 specimens with no mutation detected and an estimated tumor cellularity of less than 10% were excluded
bMetastatic melanomas of unknown origin or without information of the primary site
BRAF mutations
A total of 30 unique BRAF mutations were detected in 189 BRAF-mutated melanomas, including one tumor with two BRAF mutations (Table 2). Eighteen (9%) mutations were located within exon 11 and 172 (91%) within exon 15. p.V600E (62%) and p.V600K (14%) were the two most common BRAF mutations (Fig. 1). Mutations occurring outside codon 600 were seen in 43 tumors (23% of BRAF-mutated melanomas or 9.6% of melanomas). There were 147 (77%) class-1 mutations, 14 (7.4%) class-2 or likely class-2 mutations, and 22 (12%) class-3 or likely class-3 mutations (impaired kinase activity) and 7 mutations of unknown class (3.7%) (Table 2 and Fig. 1). The overall incidence among this cohort with 446 melanoma specimens is 33% for class-1 mutation (26% for p.V600E and 6.1% for p.V600K), 3.1% for class-2 mutations, and 4.9% for class-3 mutations (Table 3). Common locations affected by class-3 BRAF mutations were codons 594 (7 tumors), 466 (6 tumors) and 467 (4 tumors).
Table 2.
BRAF mutations in 189 melanomas
| aa change | cDNA change | Exon | Case No.a | Classb |
|---|---|---|---|---|
| p.G464Ic | c.1390_1391delinsAT | 11 | 1 | likely 2 |
| p.G464R | c.1390G > A | 11 | 1 | likely 2 |
| p.G466A | c.1397G > C | 11 | 1 (1) | 3 |
| p.G466E | c.1397G > A | 11 | 2 | 3 |
| p.G466 V | c.1397G > T | 11 | 3 (1) | 3 |
| p.S467 L | c.1400C > T | 11 | 4 (1) | 3 |
| p.G469A | c.1406G > C | 11 | 2 | 2 |
| p.G469E | c.1406G > A | 11 | 2 (2) | 3 |
| p.G469R | c.1405G > A | 11 | 1 | 2 |
| p.G469 V | c.1406G > T | 11 | 1 (1) | 2 |
| p.N581I | c.1742A > T | 15 | 1 | 3 |
| p.N581 T | c.1742A > C | 15 | 1 | likely 3 |
| p.L584F | c.1750C > T | 15 | 1 (1) | UK |
| p.E586K | c.1756G > A | 15 | 1 | UK |
| p.G593D | c.1778G > A | 15 | 1 | UK |
| p.D594E | c.1782 T > A | 15 | 1 (1) | 3 |
| p.D594G | c.1781A > G | 15 | 1 (1) | 3 |
| p.D594N | c.1780G > A | 15 | 5 (2) | 3 |
| p.G596C | c.1786G > T | 15 | 1 | likely 3 |
| p.L597Q | c.1790 T > A | 15 | 2 (1) | 2 |
| p.L597R | c.1790 T > G | 15 | 1 (1) | likely 2 |
| p.L597S | c.1789_1790delinsTC | 15 | 1 | likely 2 |
| p.T599dup | c.1795_1797dup | 15 | 1 | UK |
| p.V600E | c.1799 T > Ad | 15 | 117 (44) | 1 |
| p.V600K | c.1798_1799delinsAA | 15 | 27 (7) | 1 |
| p.V600R | c.1798_1799delinsAG | 15 | 3 (1) | 1 |
| p.K601E | c.1801A > G | 15 | 4 (2) | 2 |
| p.R603* | c.1807C > T | 15 | 1 | UK |
| p.S605Ic | c.1814G > T | 15 | 1 (1) | UK |
| p.S607F | c.1820C > T | 15 | 1 | UK |
aa change: amino acid change; Case no.: case number. UK: unknown class
aNumber in the parenthesis indicates case number of BRAF mutations previously reported [17]
bLikely 2 or 3: other mutations involving the same codon have been categorized as 2 or 3
cNot reported in the COSMIC database
dTwo with c.1799_1800delinsAA
Fig. 1.
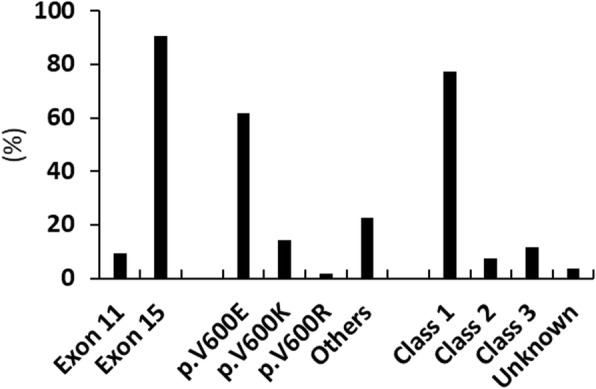
Distribution of BRAF mutations. BRAF mutations are categorized into class-1 (high kinase activity involving codon 600), class-2 (high or intermediate kinase activity involving codons outside 600), and class-3 (impaired BRAF kinase activity). [23]
Table 3.
Incidences of class-1, class-2 and class-3 BRAF mutations
| BRAF | Class-1 | p.V600E | p.V600K | Class-2 | Class-3 | Unclassified | |
|---|---|---|---|---|---|---|---|
| Uveal (n = 2) | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Mucosal (n = 20) | 1 | 0 | 0 | 0 | 0 | 1 | 0 |
| Cutaneous (n = 286) | 132 (46%) | 105 (37%)a | 82 (29%) | 20 (7.0%) | 10 (3.5%) | 14 (4.9%) | 3 |
| H/N/UB (n = 98) | 44 (45%) | 28 (29%) | 14 (14%) | 14 (14%) | 5 (5.1%) | 10 (10%) | 1 |
| non-H/N/UB (n = 188) | 88 (47%) | 77 (41%)a | 68 (36%) | 6 (3.2%) | 5 (2.7%) | 4 (2.1%) | 2 |
| Others (n = 138)b | 56 (41%) | 42 (30%) | 35 (25%) | 7 (5.1%) | 4 (2.9%) | 7 (5.1%) | 4 |
| Total (n = 446) | 189 (42%) | 147 (33%) a | 117 (26%) | 27 (6.1%) | 14 (3.1%) | 22 (4.9%) | 7 (1.6%) |
H/N/UB: Primary tumors at head, neck or upper back
aIncluding 3 tumors with p.V600R
bMetastatic melanomas of unknown origin or without information of the primary site, including one tumor with p.S605I and p.V600E within the same allele
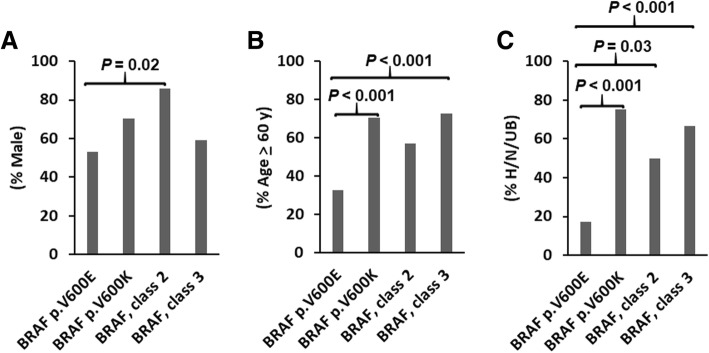
We and others have previously shown different clinical characteristics between patients with BRAF p.V600E and p.V600K mutation [32–34]. Therefore, BRAF class-1 p.V600E and p.V600K mutations were separated for comparison with the class-2 and class-3 mutations. The p.V600K, class-2 and class-3 mutations were associated with male gender and/or old age (> 60 years), as compared to the p.V600E mutation (Fig. 2a and b). In both male and female patient populations with BRAF mutations, old age was significantly associated with a higher incidence of p.V600K, class-2 or class-3 mutations and a lower incidence of p.V600E mutation (Additional file 1: Table S2). In old patient and young patient populations, there was no significant association of different BRAF mutations with gender, although male patients showed a trend of higher incidence of p.V600K, class-2 or class-3 mutations (P = 0.17) and a lower incidence of p.V600E mutation (P = 0.07) in the old patient population. The p.V600K, class-2 and class-3 mutations showed a significantly higher incidence of primary tumors located at head, neck and upper back as compared to the p.V600E mutation (75, 50 and 67% vs. 17%, Fig. 2c).
Fig. 2.
Association of p.V600E (n = 117), p.V600K (n = 27), class-2 (n = 14) and class-3 (n = 22) BRAF mutations with gender (a), age (b) and locations of the primary tumor at head, neck and upper back (H/N/UB) (c). The primary tumor site was known in 82, 20, 10 and 15 for melanomas with p.V600E, p.V600K, class-2 and class-3 mutations, respectively
Association of primary tumors located within the sun exposure areas with the BRAF p.V600K mutation and class 3 BRAF mutations
The incidence of BRAF mutation was similar in cutaneous melanomas with the primary tumor located within or outside the head, neck and upper back areas (45% vs. 47%). However, cutaneous melanomas with the primary tumor located within the head, neck or upper back areas showed a higher incidence of p.V600K mutation (14% vs. 3.2%, P = 0.001), class-2 mutations (5.1% vs. 2.7%, P = 0.32) and class-3 mutations (10% vs. 2.1%, P < 0.01), but a lower incidence of class-1 mutations (29% vs. 41%, P = 0.04) and p.V600E mutation (14% vs. 36%, P < 0.001), as compared to cutaneous melanomas with the primary tumor located outside the head, neck and upper back areas (Table 3). There was no difference in the incidence of each mutation category between the cutaneous melanomas and metastatic melanomas of unknown origin or without information of the primary site.
NRAS, KRAS, HRAS and PIK3CA mutations
A total of 18 unique NRAS mutations were detected in 110 NRAS-mutated melanomas, including 2 tumors with 2 NRAS mutations. There were 92 (82%) NRAS mutations involving codon 61, and 16 (14%) mutations involving codon 12 or codon 13. p.Q61R (38%) and p.Q61K (31%) were two most common NRAS mutations. KRAS mutations were detected in 12 melanomas. There were 8 mutations involving codon 12 or 13 and one mutation involving codon 61. HRAS mutations were detected in 9 melanomas including 2 with p.G13 N (c.37_38delinsAA) resulting from a CC > TT (or GG > AA) alteration, a signature of UV-damage. There were 4 mutations involving codon 12 or codon 13 and 4 mutations involving codon 61. PIK3CA mutations were detected in 12 melanomas, including 1 tumor with two PIK3CA mutations. p.E545K (38%) was the most common PIK3CA mutation. Mutations affecting the 3 most common codons (p.E542, p.E545 and p.H1047) account for only 54% of PIK3CA mutations.
KIT mutations
A total of 17 unique KIT mutations were detected in 22 melanomas, include one with two KIT mutations and one with 3 KIT mutations (Additional file 1: Table S3). There were 16 (64%) exon 11 mutations which might be sensitive to imatinib and 5 (20%) exon 17 mutations which are resistant (codon 816) or intermediately responsive (codon 822) to imatinib. p.L576P (28%) was the most common KIT mutation.
Variant allele frequency (VAF)
NGS detected 355 mutations among the 474 specimens. The VAF was 2–5% in 21 (5.9%) of 355 mutations, 2–10% in 47 (13%) mutations, and 2–20% in 87 (25%) mutations (Additional file 1: Table S4). VAF was below the limit of detection of pyrosequencing (approximately 5% VAF) in 4.6 and 4.1% of BRAF and NRAS mutations, respectively. VAF was below the limit of detection of Sanger sequencing (approximately 10–20% VAF) in 13–28% of BRAF mutations, 9–15% of NRAS mutations, and 15–38% of KIT mutations.
Mutational profiling of paired specimens
Paired specimens were submitted from 19 patients. Mutational profile was concordant in 17 pairs and discordant in 2 pairs (Table 4). In pair 18, a biopsy specimen taken from the left lower lip showed malignant melanoma with a mitosis rate at 10/mm2 and a BRAF p.V600E mutation at 11% VAF, consistent with an estimated tumor cellularity at 20–40%. However, the same BRAF mutation was not detected in a biopsy specimen with a 60–80% estimated tumor cellularity of malignant melanoma taken from the lower lip one month later. This specimen showed a mitosis rate at 0/mm2. In pair 19, no mutations were detected in a resection specimen of right vulva with a diagnosis of melanoma in situ and 20–40% estimated tumor cellularity. However, a resection specimen take from right vulva 39 months later showed malignant melanoma harboring a KIT p.N655K mutation at 3.7% VAF in a context of 30–50% estimated tumor cellularity.
Table 4.
Mutational profiling of 19 patients with paired specimens
| Cases | Specimens | Mutationsa |
|---|---|---|
| Same mutation | ||
| Pair 01 | Skin, neck (Re) | NRAS p.Q61L |
| Lymph node, neck (Re) | NRAS p.Q61L | |
| Pair 02 | Skin, thigh (Re) | NRAS p.Q61R |
| Soft tissue, thigh (Re) | NRAS p.Q61R | |
| Pair 03 | Skin, upper arm (Bx) | NRAS p.Q61R |
| Kidney (FNA) | NRAS p.Q61R | |
| Pair 04 | Soft tissue, axillar (Re) | NRAS p.G13R |
| Lymph node, axillar (Bx) | NRAS p.G13R | |
| Pair 05 | Lymph node, neck (Re) | BRAF p.V600K |
| Lung (Re) | BRAF p.V600K | |
| Pair 06 | Soft tissue, upper arm (Re) | NRAS p.Q61R |
| Soft tissue, scalp (Bx) | NRAS p.Q61R | |
| Pair 07 | Lymph node, axillar (Re) | BRAF p.V600E |
| Soft tissue, chest (Re) | BRAF p.V600E | |
| Pair 08 | Lymph node, groin (Bx) | BRAF p.V600E |
| Lymph node, neck (FNA) | BRAF p.V600E | |
| Pair 09 | Skin, scalp (Bx) | No mutation |
| Lymph node, neck (FNA) | No mutation | |
| Pair 10 | Skin, thigh (Re) | NRAS p.Q61R |
| Lymph node, iliac (Bx) | NRAS p.Q61R | |
| Pair 11 | Soft tissue, axillar (Re) | NRAS p.Q61R |
| Lymph node, axillar (Re) | NRAS p.Q61R | |
| Pair 12 | Skin, lower leg (Re) | NRAS p.Q61L |
| Lymoh node, groin (Re) | NRAS p.Q61L | |
| Pair 13 | Skin, upper back (Bx) | BRAF p.V600E |
| Soft tissue, upper back (Bx) | BRAF p.V600E | |
| Pair 14 | Brain (Re) | NRAS p.Q61R |
| Lymph node, neck (Re) | NRAS p.Q61R | |
| Pair 15 | Skin, chest (Re) | No mutation |
| Lymph node, axillar (Bx) | No mutation | |
| Pair 16 | Skin, nose (Re) | KIT p.L576 |
| Liver (Bx) | KIT p.L576 | |
| Pair 17 | Lung (Re) | No mutation |
| Lung (Re) | No mutation | |
| Different mutations | ||
| Pair 18 | Left lower lip (Bx) | BRAF p.V600E |
| Lower lip (Bx) | No mutation | |
| Pair 19 | Skin, right vulva (Re) | No mutation |
| Skin, right vulva (Re) | KIT p.N655K | |
Bx: biopsy; FNA: fine needle aspiration; Re: resection or excision
aMutations in the BRAF, HRAS, KIT, KRAS, NRAS and PIK3CA genes
Coexisting mutations
Coexisting mutations within BRAF, RAS, PIK3CA and KIT genes were observed in 31 (9.5%) of 328 tumors with mutations (Table 5). There were 6 tumors with coexisting mutations of the same gene (1 with 2 BRAF, 2 with 2 NRAS, 1 with 2 PIK3CA, 1 with 2 KIT, and 1 with 3 KIT mutations), and 26 tumors with coexisting mutations of different genes, including a tumor with 2 NRAS and one PIK3CA mutation. BRAF and NRAS-mutated melanomas showed a significantly lower incidence of coexisting mutations of different genes (10 and 8.1%, respectively) as compared to PIK3CA, HRAS and KIT-mutated melanomas (75, 67 and 36%; all P < .001).
Table 5.
Melanomas with coexisting mutations in 31 melanomasa
| BRAF | HRAS | KRAS | NRAS | PIK3CA | KIT | CM (%)b | |
|---|---|---|---|---|---|---|---|
| BRAF | 1c | 3 | 1 | 4 | 5 | 5 | 18 (10%) |
| HRAS | 3 | 0 | 1 | 0 | 1 | 1 | 6 (67%) |
| KRAS | 1 | 1 | 0 | 1 | 0 | 0 | 3 (25%) |
| NRAS | 4 | 0 | 1 | 2d | 2 | 1 | 9 (8.1%) |
| PIK3CA | 5 | 1 | 0 | 2 | 1 | 1 | 9 (75%) |
| KIT | 5 | 1 | 0 | 1 | 1 | 2e | 8 (36%) |
aIncluding 29 tumors with 2 mutations and 2 tumors with 3 mutations
bCoexisting mutation (CM) within different genes
cp.S605I with unknown kinase activity and p.V600E within the same allele
dOne with NRAS p.Q61L and p.Y64D of unknown activating status, and one with NRAS p.G13D, p.Q61K and PIK3CA p.E542G
eOne with p.V560A and p.N822Y, and one with p.P573S, p.F681I and p.N822I
Coexisting BRAF and activating PIK3CA or RAS mutations
Coexisting BRAF and activating RAS mutations were observed in none of 147 melanomas with a class-1 BRAF mutation, 2 (14%) of 14 melanomas with a class-2 BRAF mutation, and 5 (23%) of 22 melanomas with a class-3 BRAF mutation (Table 6). The incidence of coexisting activating RAS mutations was significantly higher in the class-2 and class-3 BRAF mutations (P < 0.01 and P < .001, respectively), as compared to the class-1 BRAF mutations. The VAF of BRAF mutations were relatively concordant with that of RAS mutations, except for one tumor with coexistence of class-2 BRAF mutation and NRAS mutation. Activating PIK3CA mutations affecting codons 542, 545 or 1047 were observed in 2 melanomas with a class-1 BRAF mutation and 1 melanoma with a class-3 BRAF mutation.
Table 6.
Coexisting BRAF and activating RAS or PIK3CA mutations
| BRAF kinase activity | BRAF a | RAS or PIK3CAa |
|---|---|---|
| Coexisting BRAF and RASb | ||
| Class-2 BRAF mutation | p.K601E (35%) | NRAS p.G13 N (34%) |
| p.G464R (5.5%) | NRAS p.Q61R (55%) | |
| Class-3 BRAF mutation | p.G466A (30%) | KRAS p.G12D (49%) |
| p.G466 V (19%) | HRAS p.G13 N (26%) | |
| p.S467 L (26%) | NRAS p.Q61K (24%) | |
| p.D594N (47%) | NRAS p.G12S (42%) | |
| p.D594N (59%) | HRAS p.Q61K (46%) | |
| Coexisting BRAF and PIK3CAc | ||
| Class-1 BRAF mutation | p.V600E (11%) | PIK3CA p.545 K (7.5%) |
| p.V600K (56%) | PIK3CA p.545 K (24%) | |
| Class-3 BRAF mutation | p.G466E (17%) | PIK3CA p.545 K (3.1%) |
aPercentage in the parenthesis indicates mutant allele frequency
bA case with BRAF p.V600E and HRAS p.Q25* was not included
cIncluding only PIK3CA mutations affecting codons 542, 545 or 1047
Discussion
NGS has been clinically validated for mutational profiling of melanomas [18, 19, 35]. We have previously shown a test feasibility of 95% among the first 165 melanoma specimens submitted for NGS testing [36]. This retrospective analysis of 502 melanoma specimens for quality assessment reaffirms the strength of NGS. NGS demonstrates a great analytic sensitivity and broad reportable ranges in clinical mutational profiling. With an analytic sensitivity of 10–20% VAF, Sanger sequencing would have missed 13% or 28% of BRAF mutations with less than 10% or 20% VAFs. The analytic sensitivity can be improved to approximately 5% VAF by mutation-specific real time PCR assays, such as cobas BRAF mutation test, which was designed to detect only hot spot codon 600 mutations [15, 37]. Eight p.V600E or p.V600K mutations with less than 5% VAF and all non-codon 600 mutations in this series would have been missed by the cobas BRAF mutation test.
NGS detected a variety of BRAF mutations located outside the reportable ranges of cobas BRAF mutation test. These included 43 (9.6%) non-codon 600 BRAF mutations detected from 446 melanoma specimens. We have previously categorized these non-codon 600 mutation based on their kinase activity and proposed potential treatment strategy [17]. More recently, BRAF mutations are further categorized into 3 classes based on their distinct mechanisms to activate MAPK pathway [22, 23]. BRAF kinase activity are high for class-1 mutants, high to intermediate for class-2 mutants, and impaired or dead for class-3 mutants. Both class-1 and class-2 mutants are RAS-independent, and signal as monomers and dimers, respectively. Class-3 mutants amplify ERK signaling in a RAS-dependent fashion. Therefore, they require coexisting mechanisms to maintain activation of RAS. This is supported by the observation of a significantly higher incidence of coexisting KRAS/NRAS/HRAS mutations in melanomas with class-3 mutations [17, 18, 21, 23]. In this study, coexisting RAS mutation was observed in 23% of class-3 mutants, but none of the 147 class-1 mutants when specimens with prior treatment of BRAF inhibitor were excluded. Among the 502 submitted specimens, the only tumor carrying coexisting class-1 BRAF mutation and activating NRAS mutations was seen in a pleural effusion specimen taken 3 months after combined therapy with BRAF inhibitor and MEK inhibitor. Detection of BRAF p.V600E (VAF of 49%), NRAS p.Q61K (VAF of 19%) and NRAS p.Q61 (VAF of 3.3%) in this pleural effusion specimen are consistent with two NRAS-mutated resistant subclones emerging following targeted therapy [38]. Since NF1 gene is not included in the AmpliSeq Cancer Hotspot Panel (v2), the incidence of coexistence of NF1 and class-3 BRAF mutations is not known in this series [23].
In addition to select patients with melanomas for targeted therapy, mutational status, such as BRAF vs. NRAS mutation or p.V600E vs. p.V600K BRAF mutations may affect clinicopathological characteristics and outcomes [32–34, 39]. p.V600K mutation has been associated with male gender, old age and/or head/neck primary tumor location [32–34]. In a Chinese population examined by Sanger sequencing of exon 15, kinase impaired BRAF mutations involving codons 594 and 596 were observed in 7 (3.4%) of 208 mucosal melanomas and 6 (1.1%) of 544 non-mucosal melanomas. Codons 594/596 mutation predicted a good prognosis in this study with a small cohort of kinase impaired BRAF mutations [26]. NGS examination using AmpliSeq sequencing panel among 699 advanced melanomas revealed 6% non-codon 600 BRAF mutations [18]. Non-V600 mutations are more common in primary tumors of the head and neck.
Categorization in a large cohort of BRAF-mutated non-small cell lung cancers based on the new classification system have showed less favorable outcome in patient with class-2 or class-3 BRAF mutations [28]. In this study, we applied this new classification system to a cohort of 446 melanoma specimens submitted to a clinical diagnostics laboratory. NGS identified 33% class-1 BRAF mutations and 9.6% non-codon 600 mutation, including 3.1% class-2 mutation, 4.9% class-3 mutations and 1.6% unclassified mutations. However, referring of specimens with wild type BRAF codon 600 to the laboratory for NGS may have led to a bias toward a lower incidence of class-1 mutation. The current study confirms a previous consensus that p.V600K is more prevalent in old male and primary melanoma of head/neck. This is also true for non-V600 BRAF mutations, especially the class-3 mutations. Our findings support that duration of sun damage is a significant contributory factor for both p.V600K and class-3 mutations. A larger cohort of class-2 mutations is needed for further clarification.
Classification of the BRAF mutations according to their mechanisms to induce activation of the MAPK pathway also provides rationales for therapeutic strategy in the future [23, 24]. Class-2 and class-3 mutants function as RAS-dependent dimers and RAS-independent dimers respectively [22, 23]. Therefore, melanomas with class-2 or class-3 mutations are not expected to respond to current FDA-approved BRAF inhibitors, which are monomer selective. Case studies have shown benefit of MEK inhibitors in melanoma patients with class-2 mutation [40–42]. Combined therapy with BRAF and MEK inhibitors have shown improved efficacy in patients class-1 BRAF mutations, [9, 10] as well as inhibitory effects in cell lines harboring class-2 or class-3 BRAF mutations [43, 44]. Recently developed dimer inhibitors, such as αC-IN inhibitor and RAF inhibitor PLX8394 have therapeutic potential against the class-2 and class-3 BRAF mutants [45, 46].
Tumor heterogeneity may have important clinical implications. Multiregional analyses to include the precursor lesions have shown that BRAF and NRAS mutations are early drivers for melanoma tumorigenesis [47]. Therefore, they are expected to be present in the primary tumor and the metastatic tumor. However, evidences of intra-tumor and inter-tumor heterogeneity have been repeatedly reported. Although a high discordance rate of the BRAF p.V600E mutation has been reported in 8 (44%) of 18 paired primary and metastatic melanoma specimens, [48] BRAF and NRAS mutations are generally highly concordant with variation of discordance rates depending on the metastatic sites in larger cohort studies [49, 50]. In this study for quality assessment, discrepancy of BRAF and NRAS mutations is not identified in 9 pairs of primary and metastatic specimens and 8 pairs of metastatic specimens. Discordance of BRAF or KIT mutation was seen in two pairs of primary tumors. Tissue identify was confirmed by analysis of 17 single nucleotide polymorphisms within the NGS panel, according to an operation procedure proposed for validation of discordant trunk drivers in patients with multiple lung cancer specimens [51]. Our findings are consistent with a higher discordance rate in patients with multiple primary melanomas or multiple lung nodules [52, 53].
Conclusion
In this retrospective study for quality assessment, NGS demonstrates a high analytic sensitivity and broad reportable ranges. A variety of recurrent BRAF mutations were detected in melanomas in a clinical diagnostics setting. Categorization of 3 classes of BRAF mutations according to their mechanisms of signal transduction to activate MAPK pathway showed sun damage could contribute to tumorigenesis of melanomas carrying the BRAF p.V600K mutation or class-3 BRAF mutations. Further prospective studies are warranted to elucidate the clinical outcomes and benefits of newly developed targeted therapy in melanoma patients carrying each class of BRAF mutation.
Additional file
Table S1. 474 specimens from 457 tumors of 455 patients with melanoma. Table S2. Patients stratified according to gender and age. Table S3. KIT mutations in 22 melanomas. Table S4. Variant allele frequency (VAF) detected by next generation sequencing. (DOCX 22 kb)
Acknowledgements
Not applicable.
Abbreviations
- BAM
Binary sequence alignment/map
- FFPE
Formalin-fixed paraffin-embedded
- IGV
Integrative Genomics Viewer
- MAPK
Mitogen-activated protein kinase
- NGS
Next-generation sequencing
- PCR
Polymerase chain reaction
- VAF
Variant allele frequency
Authors’ contributions
PML, LHT and MTL analyzed the data and drafted the manuscript; AP developed the bioinformatics pipeline and assisted in data gathering and analysis; ER and GZ participated in the data generation and analysis; CDG and JRE participated in the design of the study, data generation and analysis. All authors have read and approved the final manuscript.
Funding
This study was supported by 1UM1CA186691–01 from the NIH-National Cancer Institute. The funding agency played no role in study design, data analysis, data interpretation and in the writing of this manuscript.
Availability of data and materials
Data will be made available upon email request to MTL.
Ethics approval and consent to participate
The Johns Hopkins Medicine institutional review board granted approval to this study. The IRB has determined that informed consent from individual patients was not necessary for this study.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Parvez M. Lokhandwala and Li-Hui Tseng contributed equally to this work.
Contributor Information
Parvez M. Lokhandwala, Email: plokhan1@jhmi.edu
Li-Hui Tseng, Email: lhtseng@ntu.edu.tw.
Erika Rodriguez, Email: erodri17@jhmi.edu.
Gang Zheng, Email: gzheng5@jhmi.edu.
Aparna Pallavajjalla, Email: apallav2@jhmi.edu.
Christopher D. Gocke, Email: cgocke1@jhmi.edu
James R. Eshleman, Email: jeshlem@jhmi.edu
Ming-Tseh Lin, Email: mlin36@jhmi.edu.
References
- 1.Hodis E, Watson IR, Kryukov GV, Arold ST, Imielinski M, Theurillat JP, Nickerson E, Auclair D, Li L, Place C, et al. A landscape of driver mutations in melanoma. Cell. 2012;150(2):251–263. doi: 10.1016/j.cell.2012.06.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Cancer Genome Atlas N Genomic Classification of Cutaneous Melanoma. Cell. 2015;161(7):1681–1696. doi: 10.1016/j.cell.2015.05.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hayward NK, Wilmott JS, Waddell N, Johansson PA, Field MA, Nones K, Patch AM, Kakavand H, Alexandrov LB, Burke H, et al. Whole-genome landscapes of major melanoma subtypes. Nature. 2017;545(7653):175–180. doi: 10.1038/nature22071. [DOI] [PubMed] [Google Scholar]
- 4.Robertson AG, Shih J, Yau C, Gibb EA, Oba J, Mungall KL, Hess JM, Uzunangelov V, Walter V, Danilova L, et al. Integrative analysis identifies four molecular and clinical subsets in Uveal Melanoma. Cancer Cell. 2017;32(2):204–220 e215. doi: 10.1016/j.ccell.2017.07.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sosman JA, Kim KB, Schuchter L, Gonzalez R, Pavlick AC, Weber JS, McArthur GA, Hutson TE, Moschos SJ, Flaherty KT, et al. Survival in BRAF V600-mutant advanced melanoma treated with vemurafenib. N Engl J Med. 2012;366(8):707–714. doi: 10.1056/NEJMoa1112302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.McArthur GA, Chapman PB, Robert C, Larkin J, Haanen JB, Dummer R, Ribas A, Hogg D, Hamid O, Ascierto PA, et al. Safety and efficacy of vemurafenib in BRAF(V600E) and BRAF(V600K) mutation-positive melanoma (BRIM-3): extended follow-up of a phase 3, randomised, open-label study. Lancet Oncol. 2014;15(3):323–332. doi: 10.1016/S1470-2045(14)70012-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Flaherty KT, Robert C, Hersey P, Nathan P, Garbe C, Milhem M, Demidov LV, Hassel JC, Rutkowski P, Mohr P, et al. Improved survival with MEK inhibition in BRAF-mutated melanoma. N Engl J Med. 2012;367(2):107–114. doi: 10.1056/NEJMoa1203421. [DOI] [PubMed] [Google Scholar]
- 8.Hauschild A, Grob JJ, Demidov LV, Jouary T, Gutzmer R, Millward M, Rutkowski P, Blank CU, Miller WH, Jr, Kaempgen E, et al. Dabrafenib in BRAF-mutated metastatic melanoma: a multicentre, open-label, phase 3 randomised controlled trial. Lancet. 2012;380(9839):358–365. doi: 10.1016/S0140-6736(12)60868-X. [DOI] [PubMed] [Google Scholar]
- 9.Dummer R, Ascierto PA, Gogas HJ, Arance A, Mandala M, Liszkay G, Garbe C, Schadendorf D, Krajsova I, Gutzmer R, et al. Encorafenib plus binimetinib versus vemurafenib or encorafenib in patients with BRAF-mutant melanoma (COLUMBUS): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2018;19(5):603–615. doi: 10.1016/S1470-2045(18)30142-6. [DOI] [PubMed] [Google Scholar]
- 10.Dummer R, Ascierto PA, Gogas HJ, Arance A, Mandala M, Liszkay G, Garbe C, Schadendorf D, Krajsova I, Gutzmer R, et al. Overall survival in patients with BRAF-mutant melanoma receiving encorafenib plus binimetinib versus vemurafenib or encorafenib (COLUMBUS): a multicentre, open-label, randomised, phase 3 trial. Lancet Oncol. 2018;19(10):1315–1327. doi: 10.1016/S1470-2045(18)30497-2. [DOI] [PubMed] [Google Scholar]
- 11.Dummer R, Schadendorf D, Ascierto PA, Arance A, Dutriaux C, Di Giacomo AM, Rutkowski P, Del Vecchio M, Gutzmer R, Mandala M, et al. Binimetinib versus dacarbazine in patients with advanced NRAS-mutant melanoma (NEMO): a multicentre, open-label, randomised, phase 3 trial. Lancet Oncol. 2017;18(4):435–445. doi: 10.1016/S1470-2045(17)30180-8. [DOI] [PubMed] [Google Scholar]
- 12.Carvajal RD, Antonescu CR, Wolchok JD, Chapman PB, Roman RA, Teitcher J, Panageas KS, Busam KJ, Chmielowski B, Lutzky J, et al. KIT as a therapeutic target in metastatic melanoma. JAMA. 2011;305(22):2327–2334. doi: 10.1001/jama.2011.746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Guo J, Carvajal RD, Dummer R, Hauschild A, Daud A, Bastian BC, Markovic SN, Queirolo P, Arance A, Berking C, et al. Efficacy and safety of nilotinib in patients with KIT-mutated metastatic or inoperable melanoma: final results from the global, single-arm, phase II TEAM trial. Ann Oncol. 2017;28(6):1380–1387. doi: 10.1093/annonc/mdx079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kalinsky K, Lee S, Rubin KM, Lawrence DP, Iafrarte AJ, Borger DR, Margolin KA, Leitao MM, Jr, Tarhini AA, Koon HB, et al. A phase 2 trial of dasatinib in patients with locally advanced or stage IV mucosal, acral, or vulvovaginal melanoma: a trial of the ECOG-ACRIN Cancer research group (E2607) Cancer. 2017;123(14):2688–2697. doi: 10.1002/cncr.30663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Anderson S, Bloom KJ, Vallera DU, Rueschoff J, Meldrum C, Schilling R, Kovach B, Lee JR, Ochoa P, Langland R, et al. Multisite analytic performance studies of a real-time polymerase chain reaction assay for the detection of BRAF V600E mutations in formalin-fixed, paraffin-embedded tissue specimens of malignant melanoma. Arch Pathol Lab Med. 2012;136(11):1385–1391. doi: 10.5858/arpa.2011-0505-OA. [DOI] [PubMed] [Google Scholar]
- 16.Frampton GM, Fichtenholtz A, Otto GA, Wang K, Downing SR, He J, Schnall-Levin M, White J, Sanford EM, An P, et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol. 2013;31(11):1023–1031. doi: 10.1038/nbt.2696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zheng G, Tseng LH, Chen G, Haley L, Illei P, Gocke CD, Eshleman JR, Lin MT. Clinical detection and categorization of uncommon and concomitant mutations involving BRAF. BMC Cancer. 2015;15:779. doi: 10.1186/s12885-015-1811-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Siroy AE, Boland GM, Milton DR, Roszik J, Frankian S, Malke J, Haydu L, Prieto VG, Tetzlaff M, Ivan D, et al. Beyond BRAF(V600): clinical mutation panel testing by next-generation sequencing in advanced melanoma. J Invest Dermatol. 2015;135(2):508–515. doi: 10.1038/jid.2014.366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Carter J, Tseng LH, Zheng G, Dudley J, Illei P, Gocke CD, Eshleman JR, Lin MT. Non-p.V600E BRAF mutations are common using a more sensitive and broad detection tool. Am J Clin Pathol. 2015;144(4):620–628. doi: 10.1309/AJCP85ATMJOZOUDJ. [DOI] [PubMed] [Google Scholar]
- 20.Leichsenring J, Stogbauer F, Volckmar AL, Buchhalter I, Oliveira C, Kirchner M, Frohling S, Hassel J, Enk A, Schirmacher P, et al. Genetic profiling of melanoma in routine diagnostics: assay performance and molecular characteristics in a consecutive series of 274 cases. Pathology. 2018;50(7):703–710. doi: 10.1016/j.pathol.2018.08.004. [DOI] [PubMed] [Google Scholar]
- 21.Wan PT, Garnett MJ, Roe SM, Lee S, Niculescu-Duvaz D, Good VM, Jones CM, Marshall CJ, Springer CJ, Barford D, et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell. 2004;116(6):855–867. doi: 10.1016/S0092-8674(04)00215-6. [DOI] [PubMed] [Google Scholar]
- 22.Yao Z, Torres NM, Tao A, Gao Y, Luo L, Li Q, de Stanchina E, Abdel-Wahab O, Solit DB, Poulikakos PI, et al. BRAF mutants evade ERK-dependent feedback by different mechanisms that determine their sensitivity to pharmacologic inhibition. Cancer Cell. 2015;28(3):370–383. doi: 10.1016/j.ccell.2015.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yao Z, Yaeger R, Rodrik-Outmezguine VS, Tao A, Torres NM, Chang MT, Drosten M, Zhao H, Cecchi F, Hembrough T, et al. Tumours with class 3 BRAF mutants are sensitive to the inhibition of activated RAS. Nature. 2017;548(7666):234–238. doi: 10.1038/nature23291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dankner M, Rose AAN, Rajkumar S, Siegel PM, Watson IR. Classifying BRAF alterations in cancer: new rational therapeutic strategies for actionable mutations. Oncogene. 2018;37(24):3183–3199. doi: 10.1038/s41388-018-0171-x. [DOI] [PubMed] [Google Scholar]
- 25.Kim DW, Haydu LE, Joon AY, Bassett RL, Jr, Siroy AE, Tetzlaff MT, Routbort MJ, Amaria RN, Wargo JA, McQuade JL, et al. Clinicopathological features and clinical outcomes associated with TP53 and BRAF(N)(on-)(V)(600) mutations in cutaneous melanoma patients. Cancer. 2017;123(8):1372–1381. doi: 10.1002/cncr.30463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wu X, Yan J, Dai J, Ma M, Tang H, Yu J, Xu T, Yu H, Si L, Chi Z, et al. Mutations in BRAF codons 594 and 596 predict good prognosis in melanoma. Oncol Lett. 2017;14(3):3601–3605. doi: 10.3892/ol.2017.6608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Jones JC, Renfro LA, Al-Shamsi HO, Schrock AB, Rankin A, Zhang BY, Kasi PM, Voss JS, Leal AD, Sun J, et al. (Non-V600) BRAF mutations define a clinically distinct molecular subtype of metastatic colorectal Cancer. J Clin Oncol. 2017;35(23):2624–2630. doi: 10.1200/JCO.2016.71.4394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Dagogo-Jack I, Martinez P, Yeap BY, Ambrogio C, Ferris LA, Lydon C, Nguyen T, Jessop NA, Iafrate AJ, Johnson BE, et al. Impact of BRAF mutation class on disease characteristics and clinical outcomes in BRAF-mutant lung Cancer. Clin Cancer Res. 2019;25(1):158–165. doi: 10.1158/1078-0432.CCR-18-2062. [DOI] [PubMed] [Google Scholar]
- 29.Zheng G, Lin MT, Lokhandwala PM, Beierl K, Netto GJ, Gocke CD, Eshleman JR, McCarthy E, Illei PB. Clinical mutational profiling of bone metastases of lung and colon carcinoma and malignant melanoma using next-generation sequencing. Cancer Cytopathol. 2016;124(10):744–753. doi: 10.1002/cncy.21743. [DOI] [PubMed] [Google Scholar]
- 30.Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP. Integrative genomics viewer. Nat Biotechnol. 2011;29(1):24–26. doi: 10.1038/nbt.1754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lin MT, Mosier SL, Thiess M, Beierl KF, Debeljak M, Tseng LH, Chen G, Yegnasubramanian S, Ho H, Cope L, et al. Clinical validation of KRAS, BRAF, and EGFR mutation detection using next-generation sequencing. Am J Clin Pathol. 2014;141(6):856–866. doi: 10.1309/AJCPMWGWGO34EGOD. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Menzies AM, Haydu LE, Visintin L, Carlino MS, Howle JR, Thompson JF, Kefford RF, Scolyer RA, Long GV. Distinguishing clinicopathologic features of patients with V600E and V600K BRAF-mutant metastatic melanoma. Clin Cancer Res. 2012;18(12):3242–3249. doi: 10.1158/1078-0432.CCR-12-0052. [DOI] [PubMed] [Google Scholar]
- 33.Bucheit AD, Syklawer E, Jakob JA, Bassett RL, Jr, Curry JL, Gershenwald JE, Kim KB, Hwu P, Lazar AJ, Davies MA. Clinical characteristics and outcomes with specific BRAF and NRAS mutations in patients with metastatic melanoma. Cancer. 2013;119(21):3821–3829. doi: 10.1002/cncr.28306. [DOI] [PubMed] [Google Scholar]
- 34.Chen G, Dudley J, Tseng LH, Smith K, Gurda GT, Gocke CD, Eshleman JR, Lin MT. Lymph node metastases of melanoma: challenges for BRAF mutation detection. Hum Pathol. 2015;46(1):113–119. doi: 10.1016/j.humpath.2014.09.014. [DOI] [PubMed] [Google Scholar]
- 35.Reiman A, Kikuchi H, Scocchia D, Smith P, Tsang YW, Snead D, Cree IA. Validation of an NGS mutation detection panel for melanoma. BMC Cancer. 2017;17(1):150. doi: 10.1186/s12885-017-3149-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zheng G, Tsai H, Tseng LH, Illei P, Gocke CD, Eshleman JR, Netto G, Lin MT. Test feasibility of next-generation sequencing assays in clinical mutation detection of small biopsy and fine needle aspiration specimens. Am J Clin Pathol. 2016;145(5):696–702. doi: 10.1093/ajcp/aqw043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Halait H, Demartin K, Shah S, Soviero S, Langland R, Cheng S, Hillman G, Wu L, Lawrence HJ. Analytical performance of a real-time PCR-based assay for V600 mutations in the BRAF gene, used as the companion diagnostic test for the novel BRAF inhibitor vemurafenib in metastatic melanoma. Diagn Mol Pathol. 2012;21(1):1–8. doi: 10.1097/PDM.0b013e31823b216f. [DOI] [PubMed] [Google Scholar]
- 38.Shi H, Hugo W, Kong X, Hong A, Koya RC, Moriceau G, Chodon T, Guo R, Johnson DB, Dahlman KB, et al. Acquired resistance and clonal evolution in melanoma during BRAF inhibitor therapy. Cancer Discov. 2014;4(1):80–93. doi: 10.1158/2159-8290.CD-13-0642. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Thomas NE, Edmiston SN, Alexander A, Groben PA, Parrish E, Kricker A, Armstrong BK, Anton-Culver H, Gruber SB, From L, et al. Association between NRAS and BRAF mutational status and Melanoma-specific survival among patients with higher-risk primary Melanoma. JAMA Oncol. 2015;1(3):359–368. doi: 10.1001/jamaoncol.2015.0493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Dahlman KB, Xia J, Hutchinson K, Ng C, Hucks D, Jia P, Atefi M, Su Z, Branch S, Lyle PL, et al. BRAF(L597) mutations in melanoma are associated with sensitivity to MEK inhibitors. Cancer Discov. 2012;2(9):791–797. doi: 10.1158/2159-8290.CD-12-0097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Bowyer SE, Rao AD, Lyle M, Sandhu S, Long GV, McArthur GA, Raleigh JM, Hicks RJ, Millward M. Activity of trametinib in K601E and L597Q BRAF mutation-positive metastatic melanoma. Melanoma Res. 2014;24(5):504–508. doi: 10.1097/CMR.0000000000000099. [DOI] [PubMed] [Google Scholar]
- 42.Marconcini R, Galli L, Antonuzzo A, Bursi S, Roncella C, Fontanini G, Sensi E, Falcone A. Metastatic BRAF K601E-mutated melanoma reaches complete response to MEK inhibitor trametinib administered for over 36 months. Exp Hematol Oncol. 2017;6:6. doi: 10.1186/s40164-017-0067-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Noeparast A, Giron P, De Brakeleer S, Eggermont C, De Ridder U, Teugels E, De Greve J. Type II RAF inhibitor causes superior ERK pathway suppression compared to type I RAF inhibitor in cells expressing different BRAF mutant types recurrently found in lung cancer. Oncotarget. 2018;9(22):16110–16123. doi: 10.18632/oncotarget.24576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Noeparast A, Teugels E, Giron P, Verschelden G, De Brakeleer S, Decoster L, De Greve J. Non-V600 BRAF mutations recurrently found in lung cancer predict sensitivity to the combination of Trametinib and Dabrafenib. Oncotarget. 2017;8(36):60094–60108. doi: 10.18632/oncotarget.11635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Karoulia Z, Wu Y, Ahmed TA, Xin Q, Bollard J, Krepler C, Wu X, Zhang C, Bollag G, Herlyn M, et al. An integrated model of RAF inhibitor action predicts inhibitor activity against oncogenic BRAF signaling. Cancer Cell. 2016;30(3):485–498. doi: 10.1016/j.ccell.2016.06.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yao Z, Gao Y, Su W, Yaeger R, Tao J, Na N, Zhang Y, Zhang C, Rymar A, Tao A, et al. RAF inhibitor PLX8394 selectively disrupts BRAF dimers and RAS-independent BRAF-mutant-driven signaling. Nat Med. 2019;25(2):284–291. doi: 10.1038/s41591-018-0274-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Shain AH, Yeh I, Kovalyshyn I, Sriharan A, Talevich E, Gagnon A, Dummer R, North J, Pincus L, Ruben B, et al. The genetic evolution of Melanoma from precursor lesions. N Engl J Med. 2015;373(20):1926–1936. doi: 10.1056/NEJMoa1502583. [DOI] [PubMed] [Google Scholar]
- 48.Yancovitz Molly, Litterman Adam, Yoon Joanne, Ng Elise, Shapiro Richard L., Berman Russell S., Pavlick Anna C., Darvishian Farbod, Christos Paul, Mazumdar Madhu, Osman Iman, Polsky David. Intra- and Inter-Tumor Heterogeneity of BRAFV600EMutations in Primary and Metastatic Melanoma. PLoS ONE. 2012;7(1):e29336. doi: 10.1371/journal.pone.0029336. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Colombino M, Capone M, Lissia A, Cossu A, Rubino C, De Giorgi V, Massi D, Fonsatti E, Staibano S, Nappi O, et al. BRAF/NRAS mutation frequencies among primary tumors and metastases in patients with melanoma. J Clin Oncol. 2012;30(20):2522–2529. doi: 10.1200/JCO.2011.41.2452. [DOI] [PubMed] [Google Scholar]
- 50.Casula M, Colombino M, Manca A, Caraco C, Botti G, Ascierto PA, Lissia A, Cossu A, Palmieri G, Italian Melanoma I. Low levels of genetic heterogeneity in matched lymph node metastases from patients with Melanoma. J Invest Dermatol. 2016;136(9):1917–1920. doi: 10.1016/j.jid.2016.05.103. [DOI] [PubMed] [Google Scholar]
- 51.Tseng LH, De Marchi F, Pallavajjalla A, Rodriguez E, Xian R, Belchis D, Gocke CD, Eshleman JR, Illei P, Lin MT. Clinical validation of discordant trunk driver mutations in paired primary and metastatic lung cancer specimens. Am J Clin Pathol. 2019; Accepted. [DOI] [PMC free article] [PubMed]
- 52.Colombino M, Sini M, Lissia A, De Giorgi V, Stanganelli I, Ayala F, Massi D, Rubino C, Manca A, Paliogiannis P, et al. Discrepant alterations in main candidate genes among multiple primary melanomas. J Transl Med. 2014;12:117. doi: 10.1186/1479-5876-12-117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Roepman P, Ten Heuvel A, Scheidel KC, Sprong T, Heideman DAM, Seldenrijk KA, Herder GJM, Kummer JA. Added value of 50-gene panel sequencing to distinguish multiple primary lung cancers from pulmonary metastases: a systematic investigation. J Mol Diagn. 2018;20(4):436–445. doi: 10.1016/j.jmoldx.2018.02.007. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1. 474 specimens from 457 tumors of 455 patients with melanoma. Table S2. Patients stratified according to gender and age. Table S3. KIT mutations in 22 melanomas. Table S4. Variant allele frequency (VAF) detected by next generation sequencing. (DOCX 22 kb)
Data Availability Statement
Data will be made available upon email request to MTL.