Abstract
Background
Immune infiltration may predict survival and have clinical significance in lung cancer. However, immune signatures derived from immune profiling based on bulk tumor transcriptomes have not been systematically established in lung adenocarcinoma. We aimed to construct an immune cell infiltrating score, using a new algorithm for evaluating immune infiltration, to improve the prognostic model of lung adenocarcinoma.
Methods
Public datasets of lung adenocarcinoma from the Gene Expression Omnibus and The Cancer Genome Atlas were adopted as the training and validation cohorts. Fractions of different immune cell subtypes in each sample were estimated using the CIBERSORT algorithm. The immune infiltrating score was further developed by a least absolute shrinkage and selection operator regression model. The prognostic value and clinical relationship of the model was then further explored.
Results
An immune infiltrating score model was established on the basis of the immune cells in the training cohort. A high score was associated with significantly worse survival in patients with lung adenocarcinoma (P < 0.001). The prognostic value of the score was confirmed in the validation cohort. The immune infiltrating score could improve the accuracy of predictions of survival when combined with the staging system. Furthermore, the score was potentially associated with patient smoking status and histologic subtype of lung adenocarcinoma. Its possible association with the efficacy of adjuvant chemotherapy was not statistically significant.
Conclusion
The immune cell infiltrating score has prognostic significance in predicting overall survival in patients with lung adenocarcinoma.
Electronic supplementary material
The online version of this article (10.1186/s12967-019-1964-6) contains supplementary material, which is available to authorized users.
Keywords: Lung adenocarcinoma, Score, Prognosis, Immune infiltration
Background
Lung cancer remains the leading cause of cancer incidence and death worldwide in 2018, when it was estimated to comprise 2.1 million new cases and cause 1.8 million deaths [1]. Among all patients with lung cancer, lung adenocarcinoma accounts for the largest proportion, and its incidence continues to increase [2]. The literature has shown that phenotypes of cancers are dependent on the tumor microenvironment, especially tumor-infiltrating immune cells [3, 4]. The immune response is characterized by many types of cells, and differences in their potential prognostic value depend on the cancer type [5]. Computational approaches were adopted to characterize immune-interactions in lung adenocarcinoma, mainly focused on B cell and CD8+ T cell [6]. Meta-analysis also showed that CD8+, CD3+ and FOXP3+ T cells infiltration had significantly prognostic value in lung adenocarcinoma [7]. Furthermore, the prognostic significance of integrated immune infiltration in some cancers has been validated, and this has been proposed to supplement the tumor, node, and metastasis (TNM) staging system for predicting patient survival [8–13]. However, very few studies reported integrated immune infiltration in lung adenocarcinoma.
Most previous studies have adopted an immunohistochemistry method to evaluate the landscape of tumor-infiltrating immune cell subtypes. However, a bioinformatics tool, known as the CIBERSORT algorithm, has been previously developed for computational enumeration of immune cell subtypes. The landscape and proportions of tumor-infiltrating immune cells are evaluated on the basis of the changes in expression of immune-relative and other genes in one sample. This meta-gene approach has been well designed and validated in previous studies [9, 14–16].
In the present study, CIBERSORT was used to estimate the fraction of immune cell types based on different lung adenocarcinoma gene expression profiles. Least absolute shrinkage and selection operator (LASSO) Cox regression analysis was utilized to establish an immune cell infiltrating score model in both training and validation cohorts. We believe that the immune infiltrating score could assist in predicting survival, along with clinical factors, in lung adenocarcinoma.
Methods
Study cohort and data processing
We adopted the public datasets from the Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo/) as the training cohort. Four GEO datasets (GSE30219, GSE37745, GSE50081 and GSE68465, the latter also known as the Director’s Challenge Consortium) were included in this study. All of the four datasets were derived from the genechips of Affymetrix® (Santa Clara, California, USA). GSE30219, GSE37745 and GSE50081 were based on the GPL570 genechip, while GSE68465 were from the GPL96 microarray. Then, raw data and genechip files were downloaded directly. A total of 85, 106, 127 and 443 samples of lung adenocarcinoma from GSE30219, GSE37745, GSE50081 and GSE68465 datasets were enrolled. We used a robust multichip average method via RMAExpress for background adjustment, quantile normalization and summary to process the gene profiles [17–19].
Level 3 RNA sequence data from lung adenocarcinoma samples were also downloaded from The Cancer Genome Atlas (TCGA) before June 5, 2018 (https://portal.gdc.cancer.gov/). A total of 594 samples were obtained, comprising 535 adenocarcinoma and 59 normal lung samples. Baseline clinicopathological factors and treatment information were also acquired from TCGA. Primary lung adenocarcinoma samples with complete follow-up and baseline information (age, sex and pathological stage) were included as the validation cohort in this study. Due to the original records of all included datasets, the seventh edition of the TNM staging method was utilized in our study. Information regarding histologic subtypes of lung adenocarcinoma was intergraded with records from the University of California Santa Cruz Xena database (http://xena.ucsc.edu/) and previous literature [20–22]. According to previous studies, tumors were classified into three subgroups (low-, intermediate- and high-grade adenocarcinoma) regarding the survival data for each predominant lung adenocarcinoma subtype [23, 24].
Estimation of tumor-infiltrating immune cell types
The CIBERSORT method was adopted to quantify the proportions of the immune cell in both training and validation cohorts [25, 26]. Common methods, like flow cytometry and immunohistochemistry, rely on few phenotype markers and tissue disaggregation prior to cytometry, may cause damaged or lost cells, altering analyses [26]. The CIBERSORT was developed to accurately evaluate the relative levels of the 22 immune cell phenotypes, especially closely related types, using signature gens within complex expression mixtures [26]. Such mixtures could derive from both patients’ solid tissues and blood profiled by array or RNA-sequencing [26]. The 22 immune cells are mainly composed of B cells, T cells, macrophages, dendritic cells, plasma cells, natural killer cells and mast cells. To process data by the CIBERSORT, the targeted gene profile was uploaded to its website (https://cibersort.stanford.edu). Then, a heat-map table with levels of 22 immune cell subtypes returned as the final results. As previously described, we set the threshold P-value < 0.05 and excluded samples without immune infiltration in training (5 patients) and validation (78 patients) cohorts [25]. The optimal cut-off values of the proportions of different immune cells in the training cohort were calculated on the basis of the prognostic significance using X-Tile software [27]. Each immune cell fraction level of samples from both cohorts was then divided into two groups based on the cut-off point. The immune cell fraction level divided by the cut-off value was valued as 0 or 1 in the subsequent scoring formula.
Statistical analysis
All statistical analyses in this study were performed using R version 3.5.0 (R Foundation for Statistical Computing, Vienna, Austria) and IBM SPSS Statistics 22.0 (IBM, Inc., Armonk, NY, USA). We used the LASSO Cox regression model to select the ideal prognostic model among the estimated immune cell subtypes using the glmnet package in R [28]. The optimal model parameter λ and corresponding coefficients were determined by tenfold cross validations. We chose λ via 1 − SE (standard error) criteria as previously recommended [8, 9]. Based on the final immune infiltrating score model and corresponding coefficients, a formula for the score was constructed using the training cohort. The sensitivity and specificity of the prognosis prediction of the immune cell infiltrating score were evaluated by a time-dependent receiver operating characteristic (ROC) curve in R software. The nearest neighbor estimation method in the ROC curve was adopted with cut-off times of 1, 3 and 5 years. Survival curves were estimated using the Kaplan–Meier method and the log-rank test was utilized for comparing survival curves. Univariate and multivariate Cox proportional hazard models were adopted to identify all significant prognostic factors. The prognostic value of the included variables was evaluated by Harrell’s concordance index (c-index) [29]. The score stratified by subgroups were compared using the one-way ANOVA method, if normal distribution and homogeneity of variance were followed. Correlation analysis was also conducted to explore the potential relationship between the score and the expression of immune checkpoint regulators. In this study, a two-tailed P-value of < 0.05 was considered statistically significant.
Results
Construction of the immune cell infiltrating score model
After the initial selection process, 751 and 418 patients were enrolled as the training and validation cohorts, respectively. The characteristics of the patients are listed in Table 1 and Additional file 1: Table S1. Optimal cut-off values for the fraction of each immune cell type in the training cohort were generated as described in “Methods” section (Additional file 2: Table S2). LASSO Cox regression analysis was performed, based on the fraction of immune cells in the training cohort (Fig. 1a, b). Finally, the formula of the immune cell infiltrating score for lung adenocarcinoma in the training cohort was built (Additional file 3). The prognostic accuracy of the scoring model was evaluated using time-dependent ROC analysis in the training cohort. The predictive time points were set at 1, 3 and 5 years, respectively (Fig. 2a). The area under the ROC curve for the immune infiltrating score at each time point was 0.674, 0.684 and 0.675, respectively.
Table 1.
Baseline characteristics of patients with lung adenocarcinoma in the training cohort
| Variable | Baseline characteristics (n = 751) |
|---|---|
| Affymetrix® platform | |
| HG-U133 Plus 2.0 (GPL570) | 313 (41.7) |
| HG-U133A (GPL96) | 438 (58.3) |
| Age | 64.6 ± 10.0 |
| Sex | |
| Male | 394 (52.5) |
| Female | 357 (47.5) |
| Tumor stage | |
| Stage I | 515 (68.6) |
| Stage II–III | 232 (30.9) |
| Stage IV | 4 (0.5) |
| Smoking status | |
| Current smoker | 67 (8.9) |
| Ex-smoker | 322 (42.9) |
| Non-smoker | 71 (9.5) |
| Unknown | 291 (38.7) |
| Adjuvant chemotherapy | |
| Yes | 102 (13.6) |
| No | 377 (50.2) |
| Unknown | 272 (36.2) |
Continuous data (age) was presented as mean ± standard deviation and categorical data as number (proportion of that the subgroup accounted for the whole group)
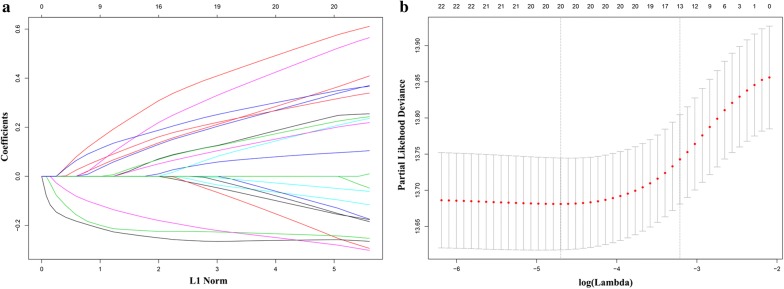
Fig. 1.
a Coefficient profiles of immune cell types in the LASSO Cox regression model. b Tenfold cross-validation for turning parameter selection in the LASSO Cox regression model. λ is the turning parameter. The partial likelihood deviance is plotted in log(λ), in which vertical lines are shown at the optimal values by minimum criteria and 1 − SE criteria
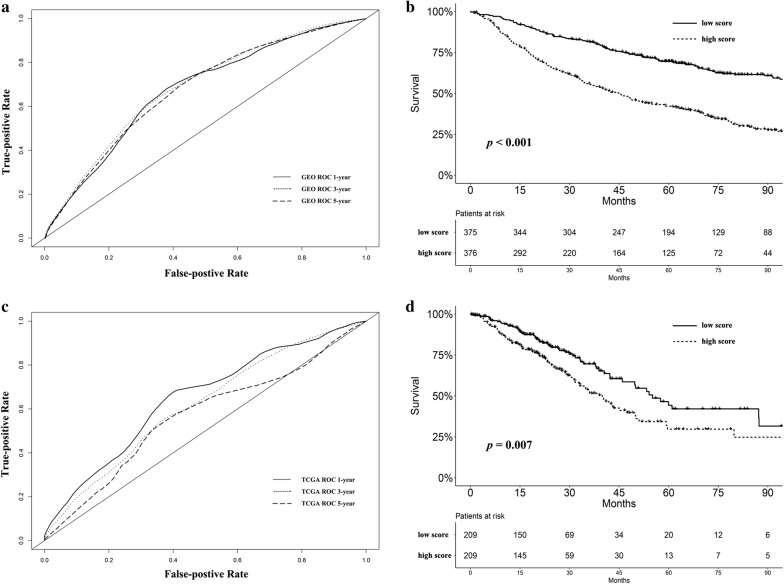
Fig. 2.
a The immune cell infiltrating score evaluated by time-dependent ROC curves at 1, 3 and 5 years in the training cohort. The area under the curve at each time point was 0.674, 0.684 and 0.675, respectively. b Survival analysis stratified by immune cell infiltrating score in the training cohort (P < 0.001). c The immune cell infiltrating score evaluated by time-dependent ROC curves at 1, 3 and 5 years in the validation cohort. The area under the curve at each time point was 0.650, 0.616 and 0.567, respectively. d Survival analysis stratified by immune cell infiltrating score in the validation cohort (P = 0.013)
Survival analyses were performed stratified by the score, in the training cohort (Fig. 2b). Patients were separated into low- and high-score groups by the median score of the cohort. We observed that patients with a high score had significantly worse survival than those with a low score (P < 0.001, Fig. 2b). A high score was also associated with significantly shorter progression-free survival (P < 0.001, Additional file 4: Figure S1).
As a continuous variable, the immune cell infiltrating score was found to have significantly prognostic value using univariate Cox regression analysis (P < 0.001, Table 2). Multivariate analysis confirmed that the immune infiltrating score was an independent prognostic factor in the training cohort, along with the TNM staging system (P < 0.001, Table 2). According to the corresponding c-index, the combination of the scoring model and TNM stage showed a significantly better predictive value than the TNM stage alone in this cohort (P < 0.001, Table 3).
Table 2.
Results of univariate and multivariate Cox regression of overall survival in the training cohort
| Cohort | Training cohort | |||||
|---|---|---|---|---|---|---|
| Variable | Univariate | Multivariate | ||||
| HR | 95% CI | P-value | HR | 95% CI | P-value | |
| Immune infiltrating score | 6.459 | 4.409–9.462 | < 0.001 | 5.322 | 3.574–7.926 | < 0.001 |
| Age | 1.025 | 1.014–1.035 | < 0.001 | 1.024 | 1.014–1.035 | < 0.001 |
| Sex | 0.031 | 0.079 | ||||
| Female | Reference | Reference | ||||
| Male | 1.245 | 1.021–1.518 | 0.031 | 1.196 | 0.979–1.460 | 0.079 |
| TNM stage | < 0.001 | < 0.001 | ||||
| Stage I | Reference | Reference | ||||
| Stage II–III | 2.682 | 2.195–3.278 | < 0.001 | 2.443 | 1.996–2.990 | < 0.001 |
| Stage IV | 2.256 | 0.721–7.053 | 0.162 | 1.835 | 0.586–5.740 | 0.297 |
HR hazard ratio, CI confidence interval
Table 3.
Comparison of the accuracy of survival prediction between the TNM stage with and without the immune infiltrating score
| Cohort | TNM stage | Immune infiltrating score + TNM stage | P-value | ||
|---|---|---|---|---|---|
| c-index | 95% CI | c-index | 95% CI | ||
| Training cohort | 0.619 | 0.594–0.643 | 0.686 | 0.659–0.714 | < 0.001 |
| Validation cohort | 0.661 | 0.621–0.700 | 0.695 | 0.650–0.740 | < 0.001 |
CI confidence interval
Validation of the immune cell infiltrating score model
We adopted a validation group based on the dataset from TCGA to evaluate the prognostic value of the proposed scoring model. The same formula for the immune cell infiltrating score and optimal cut-off point for each immune cells were applied to the validation group. Likewise, ROC analysis was adopted to assess the prognostic value of the scoring model. The area under the curve was 0.650, 0.616 and 0.567 at the predictive time of 1, 3 and 5 years, respectively (Fig. 2c). In the validation group, a high score was associated with significantly worse prognosis (P = 0.013, Fig. 2d).
Consistent with the previous findings, the score was an independent prognostic factor in the validation cohort, based on the univariate and multivariate Cox regression model (P = 0.001 and 0.002, Additional file 5: Table S3). Furthermore, the combination of the scoring model and TNM stage improved the prognostic model of lung adenocarcinoma in this cohort (P < 0.001, Table 3).
Clinical significance and bioinformatics analyses based on the immune cell infiltrating score
In the training group, data regarding adjuvant chemotherapy were documented only in the GSE68465 and GSE37745 datasets. Patients were divided into low and high score groups based on the score median value. Given the differences in staging methods and initial study design, patients with stage I disease were excluded from this analysis. The survival advantage of the low-score group was observed in patients with or without adjuvant chemotherapy (P = 0.004 and 0.002, Additional file 6: Figure S2A, B). Moreover, there was a trend for better prognosis in patients with a low score who received adjuvant chemotherapy, although the difference was not statistically significant (P = 0.909, Fig. 3a). No significant prognostic difference regarding adjuvant chemotherapy was observed in high score group (P = 0.764, Fig. 3b). We also extracted data on adjuvant chemotherapy from the validation group. There was a trend for better survival in low score group who received chemotherapy (P = 0.084, Additional file 6: Figure S2C). No difference was found in patients without chemotherapy (P = 0.761, Additional file 6: Figure S2D). Furthermore, patients with a low score may benefit from adjuvant chemotherapy (P = 0.010, Fig. 3c), while no difference was observed in high-score group (P = 0.213, Fig. 3d).
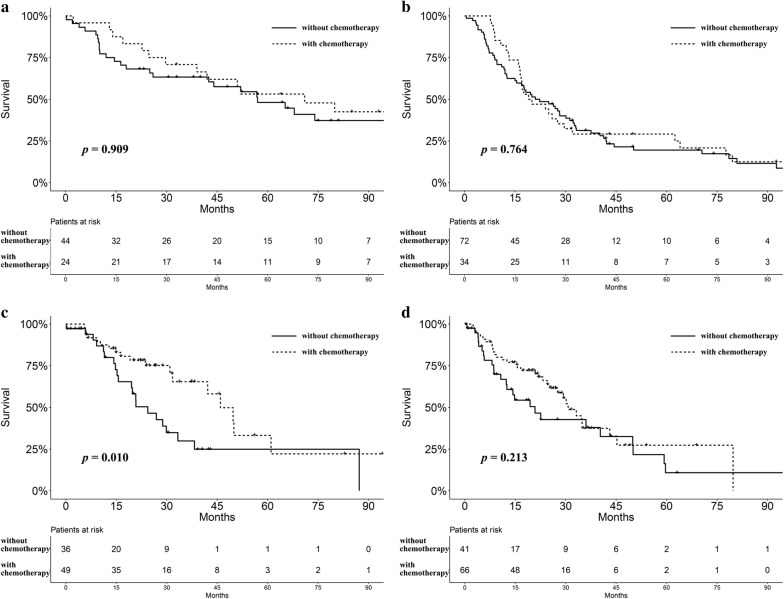
Fig. 3.
a Survival analysis for patients (stage II–IV) with low scores stratified by the receipt of adjuvant chemotherapy in the training cohort (P = 0.909). b Survival analysis for patients (stage II–IV) with high scores stratified by the receipt of adjuvant chemotherapy in the training cohort (P = 0.764). c Survival analysis for patients (stage II–IV) with low scores stratified by the receipt of adjuvant chemotherapy in the validation cohort (P = 0.010). d Survival analysis for patients (stage II–IV) with high scores stratified by the receipt of adjuvant chemotherapy in the validation cohort (P = 0.213)
Smoking status was included in two datasets (GSE50081 and GSE68465) for the training cohort. Patients with unknown smoking status were excluded. We observed that patients who are current smokers had significantly higher score than those who do not smoke (P < 0.001, Fig. 4a). In the validation cohort, patients who are current or reformed smokers (< 15 years) had higher scores than non-smokers or reformed smokers (> 15 years), although the difference was not statistically significant (P = 0.932, Fig. 4b).
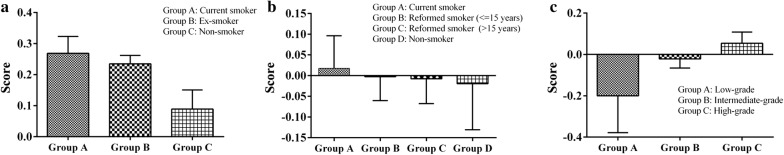
Fig. 4.
a Histograms of the immune cell infiltrating score stratified by the patient smoking status in the training cohort (P < 0.001). b Histograms of the immune cell infiltrating score stratified by the patient smoking status in the validation cohort (P = 0.932). c Histograms of the immune cell infiltrating score stratified by the histologic subtype of lung adenocarcinoma in the validation cohort (P = 0.003). High-grade: micropapillary or solid predominant lung adenocarcinoma; intermediate-grade: acinar or papillary predominant lung adenocarcinoma; low-grade: lepidic predominant lung adenocarcinoma
Furthermore, we explored potential relationships between the immune cell infiltrating score and histologic subtypes of lung adenocarcinoma. In the validation cohort, 186 patients with information on histologic subtypes were included. We found that high-grade adenocarcinoma subtypes were associated with significantly higher score than low-grade tumors (P = 0.003, micropapillary/solid vs acinar/papillary vs lepidic predominant lung adenocarcinoma, Fig. 4c).
Owing to the difference of the microarray platforms used for the training cohort, correlation analyses between the immune infiltrating score and the expression of immune checkpoint regulators or inflammatory mediators were performed in the validation cohort. We found that there was a significant positive correlation between the score and some of the regulators and mediators, including PD-L1 (P = 0.002), IDO1 (P = 0.003) and LAG3 (P = 0.017), IFNB1 (P = 0.002), IL-1A (P = 0.005), TNFA (P = 0.010) and IL-6 (P = 0.007) (Additional file 7: Table S4).
Discussion
In this study, we adopted the CIBERSORT method to calculate the fraction of 22 immune cell in lung adenocarcinoma. This newly developed algorithm can work accurately for bulk tumor samples profiled by microarray or RNA-Sequencing. And unlike flow or mass cytometry-based methods, it is applicable to archived RNA and cellular specimens [26]. Previous studies have validated the efficacy of the CIBERSORT method [14–16]. On the basis of the estimated fractions of signature immune cells, LASSO regression was utilized to construct an immune cell infiltrating score model, of which the predictive accuracy has been demonstrated previously [14, 28, 30, 31]. In both cohorts, the prognostic value of the scoring model was confirmed. A lower score was associated with significantly better prognosis, which may be due to more immunity-activating lymphocyte infiltrations. Further analyses also implied that the score could significantly improve the accuracy of survival prediction when combined with the TNM staging system. Significant correlation between the score and the expression of common immune checkpoint regulators or inflammatory mediators was also observed, including PD-L1, IDO1 and LAG3, which further confirmed its potential values. Previous studies also indicated that the expression of PD-L1 and LAG-3 was related with early recurrence and poor prognosis [32, 33]. In a word, we hope that our work will provide new insights into the construction of staging or prognostic models in lung adenocarcinoma.
The role of different tumor-infiltrating lymphocytes in lung adenocarcinoma has been explored separately [7, 34]. Kinoshita et al. [35] found that CD8+ T-cells accumulation was identified as a negative prognostic factor in lung adenocarcinoma, but not in lung squamous carcinoma. Comprehensive immune profiling of lung adenocarcinoma revealed plasma cell infiltration was related to worse prognosis [36]. Mast cell exosomes were also reported to promote lung adenocarcinoma cell proliferation [37]. The function of mast cells against or in favor of the cancers still requires investigations [38, 39]. Previous studies also observed that the prognostic implication differs according to histological types and smoking status [35, 40]. The potential role of tumor-infiltrating lymphocytes may require exploration comprehensively according to immune microenvironment. The functionality of CD8+ tumor-infiltrating lymphocytes was reported to be affected by competition between antitumor and exhaustion programs [41]. Choi and Na investigated the relationship between the immune landscape and tumor glucose metabolism in lung adenocarcinoma [42]. Varn et al. adopted computational approaches to characterize tumor-immune interactions, mainly focused on six immune cell subtypes [6]. In our study, we adopted the CIBERSORT method with 22 cell phenotypes in lung adenocarcinoma like the study of Gentles et al. [5]. The final formula of the immune cell infiltrating score was composed of 13 types of immune cells using LASSO regression, which may better characterize the potential internal interactions and microenvironment.
Several studies have focused on the relationships between tumor-infiltrating lymphocytes and the efficacy of adjuvant chemotherapy [8, 43–45]. In this study, the potential associations with chemotherapy were not all statistically significant. The weak effect may be due to small sample size in this part of analyses and the difficulty in finding the optimal cut-off value for the score. Previous studies showed that the chemotherapy sensitivity may depend on the lymphocytes infiltrating the tumor [8, 9]. One potential mechanism is that the interferon secreted by lymphocytes could sensitize cells to chemotherapy [46]. The innate and adaptive immune responses may also be activated by immunogenic tumor cell death [47]. More investigations may be required for the underlying mechanisms [48]. Our study also indicated that the score was significantly associated with patient smoking status and histologic subtype, which may help us to better characterize patient subgroups and choose the optimal treatment.
The strength of this study is the large cohort derived from several institutions, which outnumbered than most similar researches of lung adenocarcinoma. We adopted a newly developed CIBERSORT method to estimate the level of tumor-infiltrating lymphocytes, which outperformed other methods (like immunohistochemistry) regarding noise, unknown mixture content and closely related cell types. There are also some limitations that should be noted. First, the public datasets in this study were based on different genechips, but we adopted the RMAExpress method which designed for Affymetrix® microarrays and merged the raw data. All the datasets were collected with different study purposes, which may lead to heterogeneities in baseline features and therapies. Also, only retrospective analyses were made. Future prospective studies will be needed to confirm the findings. Second, some clinicopathologic factors were missing or incomplete, especially histological subtype, smoking status and adjuvant chemotherapy. Further investigations should collect more clinical factors and endpoints, along with possible alternative methods for testing and validation, which may provide more evidence for lung adenocarcinoma [13]. In addition, optimal cut-off value and practical approaches are essential for future clinical application.
Conclusion
An immune cell infiltrating score model was established based on the immune cells in lung adenocarcinoma. A high score was associated with significantly worse survival. The scoring model could improve the accuracy of predictions of survival when combined with the staging system. It was also shown that the score was closely associated with the smoking status, histologic subtype and the expression of some immune checkpoint regulators or inflammatory mediators. This study may provide new insights into the construction of staging or prognostic models in lung adenocarcinoma. The close associations between the immune infiltrating score and clinical factors may help us to better identify patient subgroups and choose the optimal treatment for patients with lung adenocarcinoma.
Additional files
Additional file 1: Table S1. Baseline characteristics of patients with lung adenocarcinoma in the validation cohort. Continuous data (age) was presented as mean ± standard deviation and categorical data as number (proportion of that the subgroup accounted for the whole group).
Additional file 2: Table S2. Cut-off value for immune cell fractions in the training cohort.
Additional file 3. Additional information.
Additional file 4: Figure S1. Survival analysis of the progression-free time stratified by immune cell infiltrating score in the training cohort (P < 0.001).
Additional file 5: Table S3. Results of univariate and multivariate Cox regression of overall survival in the validation cohort. HR = hazard ratio; CI = confidence interval.
Additional file 6: Figure S2. A: Survival analysis for patients (stage II-IV) who received chemotherapy stratified by immune cell infiltrating score in the training cohort (P = 0.004). B: Survival analysis for patients (stage II–IV) who did not receive chemotherapy stratified by immune cell infiltrating score in the training cohort (P = 0.002). C: Survival analysis for patients (stage II–IV) who received chemotherapy stratified by immune cell infiltrating score in the validation cohort (P = 0.084). D: Survival analysis for patients (stage II–IV) who did not receive chemotherapy stratified by immune cell infiltrating score in the validation cohort (P = 0.761).
Additional file 7: Table S4. Correlation between the immune infiltrating score and the expression of immune checkpoint regulators or inflammatory mediators in the validation cohort.
Acknowledgements
We would like to thank International Science Editing Co. for the language editing service.
Authors’ contributions
Study design: XY, YS, WG, CZ, QW. Data acquisition: XY, YS, CZ, ML, TL. Data analysis: XY, YS, JX, WG, CZ. Data interpretation: JX, ZL, WJ, WG, CZ, QW. Drafting of the manuscript: XY, YS, CZ, ML, TL. Revision of the manuscript: JX, WJ, ZL, WG, QW. All authors contributed significantly to this work and agreed to be accountable for the work. All authors read and approved the final manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (Grant Nos. 81672268, 81572295) (https://www.nsfc.gov.cn/) and the Research Development Fund of Zhongshan Hospital, Fudan University (2016ZSFZ56).
Availability of data and materials
All datasets were adopted in this study are available in the GEO (https://www.ncbi.nlm.nih.gov/geo/) and TCGA (https://portal.gdc.cancer.gov/) database.
Ethics approval and consent to participate
Appropriate Ethics Committees of Zhongshan Hospital approved this study to be exempt research.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Xiaodong Yang and Yu Shi contributed equally to this work
Contributor Information
Xiaodong Yang, Email: 10301010022@fudan.edu.cn.
Yu Shi, Email: shi.yu@zs-hospital.sh.cn.
Ming Li, Email: 14301050161@fudan.edu.cn.
Tao Lu, Email: lutao901118@163.com.
Junjie Xi, Email: xi.junjie@zs-hospital.sh.cn.
Zongwu Lin, Email: lin.zongwu@zs-hospital.sh.cn.
Wei Jiang, Email: jiang.wei1@zs-hospital.sh.cn.
Weigang Guo, Email: guo.weigang@zs-hospital.sh.cn.
Cheng Zhan, Email: czhan10@fudan.edu.cn.
Qun Wang, Email: wang.qun@zs-hospital.sh.cn.
References
- 1.Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi: 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- 2.Meza R, Meernik C, Jeon J, Cote ML. Lung cancer incidence trends by gender, race and histology in the United States, 1973–2010. PLoS ONE. 2015;10(3):e0121323. doi: 10.1371/journal.pone.0121323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Mantovani A, Allavena P, Sica A, Balkwill F. Cancer-related inflammation. Nature. 2008;454(7203):436–444. doi: 10.1038/nature07205. [DOI] [PubMed] [Google Scholar]
- 4.Swann JB, Smyth MJ. Immune surveillance of tumors. J Clin Invest. 2007;117(5):1137–1146. doi: 10.1172/JCI31405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Gentles AJ, Newman AM, Liu CL, et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat Med. 2015;21(8):938–945. doi: 10.1038/nm.3909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Varn FS, Tafe LJ, Amos CI, Cheng C. Computational immune profiling in lung adenocarcinoma reveals reproducible prognostic associations with implications for immunotherapy. Oncoimmunology. 2018;7(6):e1431084. doi: 10.1080/2162402X.2018.1431084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Geng Y, Shao Y, He W, et al. Prognostic role of tumor-infiltrating lymphocytes in lung cancer: a meta-analysis. Cell Physiol Biochem. 2015;37(4):1560–1571. doi: 10.1159/000438523. [DOI] [PubMed] [Google Scholar]
- 8.Jiang Y, Zhang Q, Hu Y, et al. ImmunoScore signature: a prognostic and predictive tool in gastric cancer. Ann Surg. 2018;267(3):504–513. doi: 10.1097/SLA.0000000000002116. [DOI] [PubMed] [Google Scholar]
- 9.Zeng D, Zhou R, Yu Y, et al. Gene expression profiles for a prognostic immunoscore in gastric cancer. Br J Surg. 2018;105(10):1338–1348. doi: 10.1002/bjs.10871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Xiong Y, Wang K, Zhou H, et al. Profiles of immune infiltration in colorectal cancer and their clinical significant: a gene expression-based study. Cancer Med. 2018;7(9):4496–4508. doi: 10.1002/cam4.1745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhang XM, Song LJ, Shen J, et al. Prognostic and predictive values of immune infiltrate in patients with head and neck squamous cell carcinoma. Hum Pathol. 2018;82:104–112. doi: 10.1016/j.humpath.2018.07.012. [DOI] [PubMed] [Google Scholar]
- 12.Duan J, Xie Y, Qu L, et al. A nomogram-based immunoprofile predicts overall survival for previously untreated patients with esophageal squamous cell carcinoma after esophagectomy. J Immunother Cancer. 2018;6(1):100. doi: 10.1186/s40425-018-0418-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Pages F, Mlecnik B, Marliot F, et al. International validation of the consensus Immunoscore for the classification of colon cancer: a prognostic and accuracy study. Lancet. 2018;391(10135):2128–2139. doi: 10.1016/S0140-6736(18)30789-X. [DOI] [PubMed] [Google Scholar]
- 14.Fu H, Zhu Y, Wang Y, et al. Identification and validation of stromal immunotype predict survival and benefit from adjuvant chemotherapy in patients with muscle-invasive bladder cancer. Clin Cancer Res. 2018;24(13):3069–3078. doi: 10.1158/1078-0432.CCR-17-2687. [DOI] [PubMed] [Google Scholar]
- 15.Mao Y, Feng Q, Zheng P, et al. Low tumor infiltrating mast cell density confers prognostic benefit and reflects immunoactivation in colorectal cancer. Int J Cancer. 2018;143(9):2271–2280. doi: 10.1002/ijc.31613. [DOI] [PubMed] [Google Scholar]
- 16.Zhou L, Xu L, Chen L, et al. Tumor-infiltrating neutrophils predict benefit from adjuvant chemotherapy in patients with muscle invasive bladder cancer. OncoImmunology. 2017;6(4):e1293211. doi: 10.1080/2162402X.2017.1293211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bolstad BM, Irizarry RA, Astrand M, Speed TP. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2003;19(2):185–193. doi: 10.1093/bioinformatics/19.2.185. [DOI] [PubMed] [Google Scholar]
- 18.Irizarry RA, Bolstad BM, Collin F, et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 2003;31(4):e15. doi: 10.1093/nar/gng015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Irizarry RA, Hobbs B, Collin F, et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4(2):249–264. doi: 10.1093/biostatistics/4.2.249. [DOI] [PubMed] [Google Scholar]
- 20.Cancer Genome Atlas Research Network Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511(7511):543–550. doi: 10.1038/nature13385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Travis WD, Brambilla E, Noguchi M, et al. International association for the study of lung cancer/American Thoracic Society/European Respiratory Society international multidisciplinary classification of lung adenocarcinoma. J Thorac Oncol. 2011;6(2):244–285. doi: 10.1097/JTO.0b013e318206a221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Travis WD, Brambilla E, Nicholson AG, et al. The 2015 World Health Organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol. 2015;10(9):1243–1260. doi: 10.1097/JTO.0000000000000630. [DOI] [PubMed] [Google Scholar]
- 23.Lee HY, Cha MJ, Lee KS, et al. Prognosis in resected invasive mucinous adenocarcinomas of the lung: related factors and comparison with resected nonmucinous adenocarcinomas. J Thorac Oncol. 2016;11(7):1064–1073. doi: 10.1016/j.jtho.2016.03.011. [DOI] [PubMed] [Google Scholar]
- 24.Russell PA, Wainer Z, Wright GM, et al. Does lung adenocarcinoma subtype predict patient survival?: a clinicopathologic study based on the new International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society international multidisciplinary lung adenocarcinoma classification. J Thorac Oncol. 2011;6(9):1496–1504. doi: 10.1097/JTO.0b013e318221f701. [DOI] [PubMed] [Google Scholar]
- 25.Ali HR, Chlon L, Pharoah PD, et al. Patterns of immune infiltration in breast cancer and their clinical implications: a gene-expression-based retrospective study. PLoS Med. 2016;13(12):e1002194. doi: 10.1371/journal.pmed.1002194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Newman AM, Liu CL, Green MR, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12(5):453–457. doi: 10.1038/nmeth.3337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Camp RL, Dolled-Filhart M, Rimm DL. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res. 2004;10(21):7252–7259. doi: 10.1158/1078-0432.CCR-04-0713. [DOI] [PubMed] [Google Scholar]
- 28.Tibshirani R. The lasso method for variable selection in the Cox model. Stat Med. 1997;16(4):385–395. doi: 10.1002/(SICI)1097-0258(19970228)16:4<385::AID-SIM380>3.0.CO;2-3. [DOI] [PubMed] [Google Scholar]
- 29.Harrell FJ, Califf RM, Pryor DB, et al. Evaluating the yield of medical tests. JAMA. 1982;247(18):2543–2546. doi: 10.1001/jama.1982.03320430047030. [DOI] [PubMed] [Google Scholar]
- 30.Qu L, Wang Z, Chen Q, et al. Prognostic value of a long non-coding RNA signature in localized clear cell renal cell carcinoma. Eur Urol. 2018;74(6):756–763. doi: 10.1016/j.eururo.2018.07.032. [DOI] [PubMed] [Google Scholar]
- 31.Dai W, Feng Y, Mo S, et al. Transcriptome profiling reveals an integrated mRNA–lncRNA signature with predictive value of early relapse in colon cancer. Carcinogenesis. 2018;39(10):1235–1244. doi: 10.1093/carcin/bgy087. [DOI] [PubMed] [Google Scholar]
- 32.He Y, Yu H, Rozeboom L, et al. LAG-3 protein expression in non-small cell lung cancer and its relationship with PD-1/PD-L1 and tumor-infiltrating lymphocytes. J Thorac Oncol. 2017;12(5):814–823. doi: 10.1016/j.jtho.2017.01.019. [DOI] [PubMed] [Google Scholar]
- 33.Zhang ML, Kem M, Mooradian MJ, et al. Differential expression of PD-L1 and IDO1 in association with the immune microenvironment in resected lung adenocarcinomas. Mod Pathol. 2019;32(4):511–523. doi: 10.1038/s41379-018-0160-1. [DOI] [PubMed] [Google Scholar]
- 34.Bremnes RM, Busund LT, Kilvaer TL, et al. The role of tumor-infiltrating lymphocytes in development, progression, and prognosis of non-small cell lung cancer. J Thorac Oncol. 2016;11(6):789–800. doi: 10.1016/j.jtho.2016.01.015. [DOI] [PubMed] [Google Scholar]
- 35.Kinoshita T, Kudo-Saito C, Muramatsu R, et al. Determination of poor prognostic immune features of tumour microenvironment in non-smoking patients with lung adenocarcinoma. Eur J Cancer. 2017;86:15–27. doi: 10.1016/j.ejca.2017.08.026. [DOI] [PubMed] [Google Scholar]
- 36.Kurebayashi Y, Emoto K, Hayashi Y, et al. Comprehensive immune profiling of lung adenocarcinomas reveals four immunosubtypes with plasma cell subtype a negative indicator. Cancer Immunol Res. 2016;4(3):234–247. doi: 10.1158/2326-6066.CIR-15-0214. [DOI] [PubMed] [Google Scholar]
- 37.Xiao H, Lasser C, Shelke GV, et al. Mast cell exosomes promote lung adenocarcinoma cell proliferation—role of KIT-stem cell factor signaling. Cell Commun Signal. 2014;12:64. doi: 10.1186/s12964-014-0064-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Derakhshani A, Vahidian F, Alihasanzadeh M, et al. Mast cells: a double-edged sword in cancer. Immunol Lett. 2019;209:28–35. doi: 10.1016/j.imlet.2019.03.011. [DOI] [PubMed] [Google Scholar]
- 39.Reddy SM, Reuben A, Barua S, et al. Poor response to neoadjuvant chemotherapy correlates with mast cell infiltration in inflammatory breast cancer. Cancer Immunol Res. 2019;7(6):1025–1035. doi: 10.1158/2326-6066.CIR-18-0619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kinoshita T, Muramatsu R, Fujita T, et al. Prognostic value of tumor-infiltrating lymphocytes differs depending on histological type and smoking habit in completely resected non-small-cell lung cancer. Ann Oncol. 2016;27(11):2117–2123. doi: 10.1093/annonc/mdw319. [DOI] [PubMed] [Google Scholar]
- 41.O’Brien SM, Klampatsa A, Thompson JC, et al. Function of human tumor-infiltrating lymphocytes in early-stage non-small cell lung cancer. Cancer Immunol Res. 2019;7(6):896–909. doi: 10.1158/2326-6066.CIR-18-0713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Choi H, Na KJ. Integrative analysis of imaging and transcriptomic data of the immune landscape associated with tumor metabolism in lung adenocarcinoma: clinical and prognostic implications. Theranostics. 2018;8(7):1956–1965. doi: 10.7150/thno.23767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Jang N, Kwon HJ, Park MH, et al. Prognostic value of tumor-infiltrating lymphocyte density assessed using a standardized method based on molecular subtypes and adjuvant chemotherapy in invasive breast cancer. Ann Surg Oncol. 2018;25(4):937–946. doi: 10.1245/s10434-017-6332-2. [DOI] [PubMed] [Google Scholar]
- 44.Zhang H, Liu H, Shen Z, et al. Tumor-infiltrating neutrophils is prognostic and predictive for postoperative adjuvant chemotherapy benefit in patients with gastric cancer. Ann Surg. 2018;267(2):311–318. doi: 10.1097/SLA.0000000000002058. [DOI] [PubMed] [Google Scholar]
- 45.Ignatiadis M, Van den Eynden G, Roberto S, et al. Tumor-infiltrating lymphocytes in patients receiving trastuzumab/pertuzumab-based chemotherapy: a TRYPHAENA substudy. J Natl Cancer Inst. 2019;111(1):69–77. doi: 10.1093/jnci/djy076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Galluzzi L, Buqué A, Kepp O, et al. Immunological effects of conventional chemotherapy and targeted anticancer agents. Cancer Cell. 2015;28(6):690–714. doi: 10.1016/j.ccell.2015.10.012. [DOI] [PubMed] [Google Scholar]
- 47.Ghiringhelli F, Apetoh L. The interplay between the immune system and chemotherapy: emerging methods for optimizing therapy. Expert Rev Clin Immunol. 2014;10(1):19–30. doi: 10.1586/1744666X.2014.865520. [DOI] [PubMed] [Google Scholar]
- 48.Zitvogel L, Apetoh L, Ghiringhelli F, et al. The anticancer immune response: indispensable for therapeutic success? J Clin Invest. 2008;118(6):1991–2001. doi: 10.1172/JCI35180. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Table S1. Baseline characteristics of patients with lung adenocarcinoma in the validation cohort. Continuous data (age) was presented as mean ± standard deviation and categorical data as number (proportion of that the subgroup accounted for the whole group).
Additional file 2: Table S2. Cut-off value for immune cell fractions in the training cohort.
Additional file 3. Additional information.
Additional file 4: Figure S1. Survival analysis of the progression-free time stratified by immune cell infiltrating score in the training cohort (P < 0.001).
Additional file 5: Table S3. Results of univariate and multivariate Cox regression of overall survival in the validation cohort. HR = hazard ratio; CI = confidence interval.
Additional file 6: Figure S2. A: Survival analysis for patients (stage II-IV) who received chemotherapy stratified by immune cell infiltrating score in the training cohort (P = 0.004). B: Survival analysis for patients (stage II–IV) who did not receive chemotherapy stratified by immune cell infiltrating score in the training cohort (P = 0.002). C: Survival analysis for patients (stage II–IV) who received chemotherapy stratified by immune cell infiltrating score in the validation cohort (P = 0.084). D: Survival analysis for patients (stage II–IV) who did not receive chemotherapy stratified by immune cell infiltrating score in the validation cohort (P = 0.761).
Additional file 7: Table S4. Correlation between the immune infiltrating score and the expression of immune checkpoint regulators or inflammatory mediators in the validation cohort.
Data Availability Statement
All datasets were adopted in this study are available in the GEO (https://www.ncbi.nlm.nih.gov/geo/) and TCGA (https://portal.gdc.cancer.gov/) database.