Abstract
Over the last decade, arthropods have been shown to harbour a rich diversity of viruses. Through viral metagenomics a large diversity of single-stranded (ss) DNA viruses have been identified. Here we examine the ssDNA virome of the hematophagous New Zealand blackfly using viral metagenomics. Our investigation reveals a plethora of novel ssDNA viral genomes, some of which cluster in the viral families Genomoviridae (n = 9), Circoviridae (n = 1), and Microviridae (n = 108), others in putative families that, at present, remain unclassified (n = 20) and one DNA molecule that only encodes a replication associated protein. Among these novel viruses, two putative multi-component virus genomes were recovered, and these are most closely related to a Tongan flying fox faeces-associated multi-component virus. Given that the only other known multi-component circular replication-associated (Rep) protein encoding single-stranded (CRESS) DNA viruses infecting plants are in the families Geminiviridae (members of the genus Begomovirus) and Nanoviridae, it appears these are likely a new multi-component virus group which may be associated with animals. This study reiterates the diversity of ssDNA viruses in nature and in particular with the New Zealand blackflies.
Keywords: blackfly, sandfly, Austrosimulium sp., CRESS DNA virus, Circoviridae, Genomoviridae
1. Introduction
Hematophagous insects are responsible for vectoring a wide range of pathogens. Vectors of important mammalian and avian pathogens, such as mosquitoes, are members of the order Diptera (suborder: Nematocera). One group of blood sucking insects, commonly referred to as blackflies or sandflies, are members of the Simuliidae family [1]. Although blackflies are found globally, only some species consume a blood meal. For those species that feed on blood, it is the adult females that do so, whereas the males feed primarily on nectar. Aquatic environments are essential for the life cycle of blackflies, with the egg, larval and pupae stages all occurring in flowing water, followed by the emergence of a winged adult form.
Blackflies are known to transmit a handful of parasites, predominantly protozoa and parasitic worms. In humans, the most important blackfly vectored pathogen is the parasitic nematode, Onchocerca volvulus which causes river blindness. Although rare, globally O. volvulus has a significant impact on human health in Africa, affecting over 18 million people [2,3]. Avian haemazoan parasites, such as those in the genus Leucocytozoon, are also commonly transmitted by blackflies, with different blackfly species showing preference to the bird species they bite and harbouring a variety of parasite linages [4,5]. The most studied arbovirus transmitted by blackflies is vesicular stomatitis virus (family Rhabdoviridae, genus Vesiculovirus) that typically infects livestock, however, zoonotic events have also been reported [6]. Invertebrate iridoviruses (family Iridoviridae), which are double-stranded DNA viruses, have been identified in blackflies across the globe, often causing a covert infection [7,8].
Several species of blackfly are endemic to New Zealand, two of which, the New Zealand blackfly (Austrosimulium australense) and the west coast blackfly (A. ungulatum), consume blood meals [9,10]. They are notorious for their persistence in pursuit of a blood meal and in various locations are present in overwhelming swarms. Despite having some knowledge on the ecology of New Zealand blackflies, we know very little about the viruses that are circulating in these insects. No human pathogens vectored by blackflies in New Zealand have been documented and, with the exception of protozoal transmission in some avian species, little is known about the microorganisms circulating in these insects.
In recent years, studies have used viral metagenomics as a non-biased approach for identifying viruses circulating in hematophagous and phytophagous insects [11,12,13,14,15]. This has resulted in the identification of a vast number of viruses, circulating in the hosts these insects are feeding on, from the surrounding environment, and those that infect the insects themselves. Arthropods have been broadly shown to harbour a wide range of viruses with circular replication-associated (Rep) protein encoding single-stranded (CRESS) DNA genomes [11,13,14,16,17]. At present there are many established CRESS DNA virus families; Bacillidnaviridae, Circoviridae, Geminiviridae, Genomoviridae, Nanoviridae, and Smacoviridae, all of whose members encode Reps that share conserved replication associated motifs, an origin of replication and are usually <6 kb in size [18]. Also, viruses in the family Microviridae which infect bacteria, typically encode a major capsid, minor capsid and a replication initiation protein, and range in size from 4–7 kb [19]. In addition, numerous novel CRESS DNA viruses have been identified in arthropods which are yet to be taxonomically classified [11,20].
Viral metagenomic studies reveal a great deal about viruses circulating in arthropods. For example, a study on mosquitoes [14] identified viral sequences with similarities to animal-infecting ssDNA viruses (families: Anelloviridae, Circoviridae, Parvoviridae), double-stranded (ds) DNA viruses (families: Herpesviridae, Poxviridae and Papillomaviridae), plant-infecting ssDNA viruses (families: Geminiviridae and Nanoviridae), and three bacteria-infecting dsDNA viruses (families: Myoviridae, Podoviridae and Siphoviridae). Viral metagenomic studies allows for a snapshot of the viruses circulating in the vectors hosts, the insect and surrounding ecosystem to be identified. Based on the lack of information on the widespread New Zealand hematophagous insect commonly known as the blackfly, we undertook a metagenomics approach to investigate the associated ssDNA virome.
2. Materials and Methods
2.1. Collection of Blackflies and Isolation of Viral Nucleic Acid
For this project, 40 individual blackflies were collected from North Canterbury, New Zealand in 2015. The 40 individuals were collected from a single site. The sex of the individuals, and whether they had consumed a blood meal, was determined. These samples were pooled and homogenized using a pestle in 2 mL of SM buffer (0.1 M NaCl, 50 mM Tris/HCl-pH 7.4, 10 mM MgSO4). The homogenized sample was centrifuged for 10 min at 10,000 rpm and the resulting supernatant was filtered through a 0.2 µM filter. The viral particles in the filtrate were precipitated overnight at 4 °C with 15% PEG and following this the solution was centrifuged at 14,000 rpm for 10 min and resulting pellet resuspended in 500 µL of SM buffer. Following this, 200 µL of the resuspended material was subsequently used to isolate viral DNA using the High Pure Viral Nucleic Acid Kit (Roche Diagnostics, USA) according to the manufacturer’s specifications. The viral nucleic acid was then used in a rolling circle amplification reaction with the TempliPhi™ kit (GE Healthcare, USA) to preferentially amplify circular DNA molecules.
2.2. High-Throughput Sequencing and Viral Genome Verification
Rolling circle amplified DNA was used to prepare 2 × 150 bp paired-end libraries for sequencing on an Illumina 2500 platform at Macrogen Inc. (Korea). The paired-end reads were de novo assembled using metaSPAdes v 3.12.0 [21] and resulting contigs (>750 nts) were filtered for viral-like sequences using BLASTx [22] against viral protein database generated from the GenBank RefSeq depository. For contigs with similarities to viruses in the Microviridae family, full de novo assembled genomes were confirmed by mapping raw reads using BBMap [23], and deemed credible with a coverage level of greater than 10×. For viral contigs with similarities to other ssDNA viruses, abutting primers (Table S1) were designed to recover the complete genomes by PCR using Kapa HiFi HotStart DNA polymerase (KAPA Biosystems, USA). Amplicons were resolved on a 0.7% agarose gel, the correct size amplicons were excised, gel purified and cloned into pJET1.2 cloning vector (ThermoFisher Scientific, USA). Recombinant plasmids containing the viral genomes were purified from transformed XL blue E. coli competent cells and Sanger sequenced at Macrogen Inc. (Korea) by primer walking. The Sanger sequences were assembled using Geneious software V11.1.5.
2.3. Network Construction, Phylogenetic and Similarity Comparison Analyses
The blackfly viral rep (CRESS DNA viruses) and the major capsid protein (mcp) gene (microviruses) together with those available in GenBank were extracted, translated and used to build a Rep and MCP protein sequence dataset. The Rep of circoviruses, geminiviruses, smacoviruses, genomoviruses and nanoviruses and the MCP of viruses in the subfamily Gokushovirinae (family Microvividae) clustered first with CD-HIT [24] using 0.9 sequence identity cut off and a representative from each cluster was included in the final dataset. The Rep and MCP protein sequences were used separately to build sequence similarity networks (E-value = 1 × 10−5) using the EFI-Enzyme similarity tool server [25,26]. The MCP similarity network was constructed using minimum similarity score 10−175 and the Rep with 10−60. The score is the similarity threshold for connected nodes, i.e., proteins, with each other and thus those with scores below this value are not connected. Protein similarity networks were visualised using Cytoscape V3.7.0 [27].
Based on the network clusters, unique cluster sequences datasets were built for each major cluster (comprised of ten or more members) that contained a blackfly derived sequence. The datasets included genomoviruses, circoviruses and five additional clusters labelled cluster group 1–5 for the unclassified CRESS DNA viruses and two microvirus groups labelled cluster MV group 1 and 2. The sequences in each of these cluster groups were aligned using MUSCLE [28] and maximum likelihood phylogenetic trees inferred using PHYML [29] with the best fit models, determined using ProtTest [30]. The substitution models used are genomoviruses, LG+I+G; circoviruses, rtREV+G+I; CRESS group 1, rtREV+G+I; CRESS group 2, WAG+G+I; CRESS group 3, rtREV+G; CRESS group 4, WAG+G+I; CRESS Group 5, WAG+G+I, MV group 1, rtRev+G+I; MV group 2, rtREV+G+I+F. Branches with aLRT support of <0.8 were collapsed using TreeGraph2 [31]. The Maximum likelihood phylogenetic trees were midpoint rooted with the exception of the genomoviruses which were rooted using the geminivirus Reps and the cycloviruses with circoviruses Reps.
BLASTx [22] comparisons were undertaken for any singletons to determine the most closely related sequences in GenBank. For clusters comprised of less than four sequences an amino acid pairwise comparison using SDT V1.2 [32] was undertaken.
3. Results and Discussion
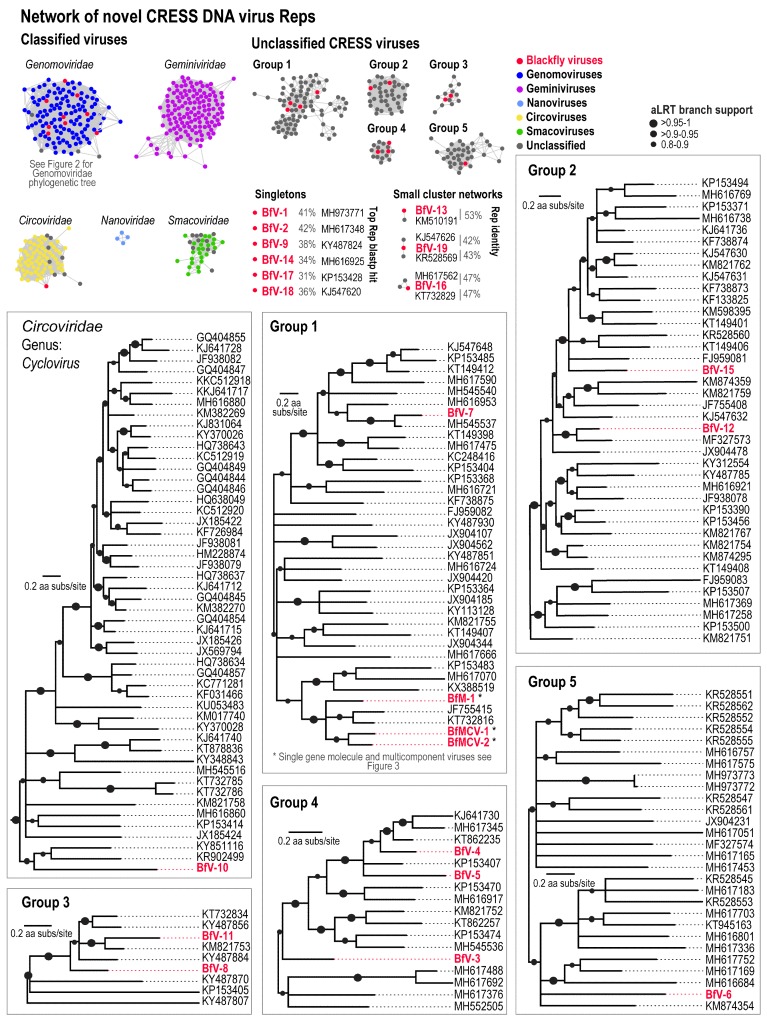
In a pool of 40 individual blackflies, diverse CRESS DNA viruses were identified. Cluster analyses using the Rep protein, which is the most conserved gene among CRESS DNA viruses, provides a broad overview of the extensive range of viruses recovered in the blackfly samples (Table 1 and Figure 1). The network analyses results reveal (Figure 1) that the Reps from the blackfly derived viruses or DNA molecules cluster with those in the Genomoviridae family (n = 9), Circoviridae family (n = 1), unclassified CRESS DNA viruses (n = 15) or as singletons (n = 6).
Table 1.
Summary of the genomes of blackfly associated viruses including genome organisation and GenBank Accession numbers.
| Family/Subfamily | Genus | Virus Name | Accession Number | Size (nts) | ORF Orientation |
|---|---|---|---|---|---|
| Genomoviridae | Gemycircularirus | Blackfly genomovirus-1 SF02 506 | MK433242 | 2290 | Bidirectional |
| Blackfly genomovirus-2 SF02 631 | MK433234 | 2131 | Bidirectional | ||
| Blackfly genomovirus-3 SF02 1766 | MK433235 | 2138 | Bidirectional | ||
| Blackfly genomovirus-4 SF02 836 | MK433236 | 2181 | Bidirectional | ||
| Blackfly genomovirus-5 SF02 599 | MK433237 | 2182 | Bidirectional | ||
| Blackfly genomovirus-6 SF02 459 | MK433238 | 2186 | Bidirectional | ||
| Blackfly genomovirus-7 SF02 767 | MK433239 | 2195 | Bidirectional | ||
| Blackfly genomovirus-9 SF02 507 | MK433241 | 2217 | Bidirectional | ||
| Genomoviridae | Gemyduguvirus | Blackfly genomovirus-8 SF02 579 | MK433240 | 2226 | Bidirectional |
| Unclassified CRESS DNA virus | Unassigned | Blackfly DNA virus-1 SF02 666 | MK433215 | 1805 | Unidirectional |
| Blackfly DNA virus-2 SF02 583 | MK433216 | 1936 | Bidirectional | ||
| Blackfly DNA virus-3 SF02 402 | MK433217 | 2005 | Unidirectional | ||
| Blackfly DNA virus-4 SF02 664 | MK433218 | 2047 | Unidirectional | ||
| Blackfly DNA virus-5 SF02 839 | MK433219 | 2058 | Unidirectional | ||
| Blackfly DNA virus-6 SF01 308 | MK433220 | 2172 | Bidirectional | ||
| Blackfly DNA virus-7 SF02 462 | MK433221 | 2306 | Bidirectional | ||
| Blackfly DNA virus-8 SF02 1137 | MK433222 | 2337 | Bidirectional | ||
| Blackfly DNA virus-9 SF02 881 | MK433223 | 2389 | Bidirectional | ||
| Blackfly DNA virus-10 SF02 899 | MK433224 | 2444 | Bidirectional | ||
| Blackfly DNA virus-11 SF02 963 | MK433225 | 2449 | Bidirectional | ||
| Blackfly DNA virus-12 SF02 422 | MK433226 | 2466 | Unidirectional | ||
| Blackfly DNA virus-13 SF02 413 | MK433227 | 2501 | Bidirectional | ||
| Blackfly DNA virus-14 SF02 295 | MK433228 | 2583 | Bidirectional | ||
| Blackfly DNA virus-15 SF02 403 | MK433229 | 2697 | Bidirectional | ||
| Blackfly DNA virus-16 SF02 377 | MK433230 | 2704 | Bidirectional | ||
| Blackfly DNA virus-17 SF02 1426 | MK433231 | 2835 | Unidirectional | ||
| Blackfly DNA virus-18 SF02 66 | MK433232 | 3003 | Bidirectional | ||
| Blackfly DNA virus-19 SF02 380 | MK433233 | 2002 | Bidirectional | ||
| Unclassified Circular DNA molecules | Unassigned | Blackfly DNA molecule 1 - rep | MK561604 | 1157 | Unidirectional |
| Unclassified Multi-component virus | Unassigned | Blackfly multicomponent virus 1 - rep | MK561605 | 1163 | Unidirectional |
| Blackfly multicomponent virus 1 - cp | MK561606 | 1154 | Unidirectional | ||
| Blackfly multicomponent virus 2 - rep | MK561607 | 1136 | Unidirectional | ||
| Blackfly multicomponent virus 2 - cp | MK561608 | 1133 | Unidirectional | ||
| Microviridae/Gokushovirinae | Unassigned | Blackfly microvirus-1_ SF02_SP_9 | MK249160 | 4873 | Unidirectional |
| Blackfly microvirus-2_ SF02_SP_11 | MK249161 | 4544 | Unidirectional | ||
| Blackfly microvirus-3_ SF02_SP_12 | MK249162 | 4625 | Unidirectional | ||
| Blackfly microvirus-4_ SF02_SP_13 | MK249163 | 4660 | Unidirectional | ||
| Blackfly microvirus-5_ SF02_SP_14 | MK249164 | 4594 | Unidirectional | ||
| Blackfly microvirus-6_ SF02_SP_15 | MK249165 | 4675 | Unidirectional | ||
| Blackfly microvirus-7_ SF02_SP_17 | MK249166 | 4379 | Unidirectional | ||
| Blackfly microvirus-8_ SF02_SP_18 | MK249167 | 4483 | Unidirectional | ||
| Blackfly microvirus-9_ SF02_SP_20 | MK249168 | 4324 | Unidirectional | ||
| Blackfly microvirus-10_ SF02_SP_21 | MK249169 | 4656 | Unidirectional | ||
| Blackfly microvirus-11_ SF02_SP_22 | MK249170 | 4565 | Unidirectional | ||
| Blackfly microvirus-12_ SF02_SP_24 | MK249171 | 4231 | Unidirectional | ||
| Blackfly microvirus-13_ SF02_SP_25 | MK249172 | 4582 | Unidirectional | ||
| Blackfly microvirus-14_ SF02_SP_27 | MK249173 | 4694 | Unidirectional | ||
| Blackfly microvirus-15_ SF02_SP_28 | MK249174 | 4557 | Unidirectional | ||
| Blackfly microvirus-16_ SF02_SP_30 | MK249175 | 4689 | Unidirectional | ||
| Blackfly microvirus-17_ SF02_SP_31 | MK249176 | 4474 | Unidirectional | ||
| Blackfly microvirus-18_ SF02_SP_32 | MK249177 | 4841 | Unidirectional | ||
| Blackfly microvirus-19_ SF02_SP_34 | MK249178 | 4727 | Unidirectional | ||
| Blackfly microvirus-20_ SF02_SP_36 | MK249179 | 4498 | Unidirectional | ||
| Blackfly microvirus-21_ SF02_SP_38 | MK249180 | 4461 | Unidirectional | ||
| Blackfly microvirus-22_ SF02_SP_44 | MK249181 | 6565 | Unidirectional | ||
| Blackfly microvirus-23_ SF02_SP_45 | MK249182 | 4565 | Unidirectional | ||
| Blackfly microvirus-24_ SF02_SP_46 | MK249183 | 4633 | Unidirectional | ||
| Blackfly microvirus-25_ SF02_SP_49 | MK249184 | 4521 | Unidirectional | ||
| Blackfly microvirus-26_ SF02_SP_50 | MK249185 | 4731 | Unidirectional | ||
| Blackfly microvirus-27_ SF02_SP_51 | MK249186 | 4581 | Unidirectional | ||
| Blackfly microvirus-28_ SF02_SP_52 | MK249187 | 4667 | Unidirectional | ||
| Blackfly microvirus-29_ SF02_SP_54 | MK249188 | 4637 | Unidirectional | ||
| Blackfly microvirus-30_ SF02_SP_57 | MK249189 | 4980 | Unidirectional | ||
| Blackfly microvirus-31_ SF02_SP_67 | MK249190 | 4901 | Unidirectional | ||
| Blackfly microvirus-32_ SF02_SP_69 | MK249191 | 4633 | Unidirectional | ||
| Blackfly microvirus-33_ SF02_SP_71 | MK249192 | 4901 | Unidirectional | ||
| Blackfly microvirus-34_ SF02_SP_73 | MK249193 | 4870 | Unidirectional | ||
| Blackfly microvirus-35_ SF02_SP_74 | MK249194 | 4765 | Unidirectional | ||
| Blackfly microvirus-36_ SF02_SP_75 | MK249195 | 4789 | Unidirectional | ||
| Blackfly microvirus-37_ SF02_SP_78 | MK249196 | 4774 | Unidirectional | ||
| Blackfly microvirus-38_ SF02_SP_79 | MK249197 | 4711 | Unidirectional | ||
| Blackfly microvirus-39_ SF02_SP_80 | MK249198 | 4858 | Unidirectional | ||
| Blackfly microvirus-40_ SF02_SP_82 | MK249199 | 4799 | Unidirectional | ||
| Blackfly microvirus-41_ SF02_SP_83 | MK249200 | 4825 | Unidirectional | ||
| Blackfly microvirus-42_ SF02_SP_84 | MK249201 | 4779 | Unidirectional | ||
| Blackfly microvirus-43_ SF02_SP_87 | MK249202 | 4768 | Unidirectional | ||
| Blackfly microvirus-44_ SF02_SP_88 | MK249203 | 4615 | Unidirectional | ||
| Blackfly microvirus-45_ SF02_SP_89 | MK249204 | 4630 | Unidirectional | ||
| Blackfly microvirus-46_ SF02_SP_91 | MK249205 | 4733 | Unidirectional | ||
| Blackfly microvirus-47_ SF02_SP_93 | MK249206 | 4721 | Unidirectional | ||
| Blackfly microvirus-48_ SF02_SP_97 | MK249207 | 4646 | Unidirectional | ||
| Blackfly microvirus-49_ SF02_SP_98 | MK249208 | 4700 | Unidirectional | ||
| Blackfly microvirus-50_ SF02_SP_99 | MK249209 | 4706 | Unidirectional | ||
| Blackfly microvirus-51_ SF02_SP_100 | MK249210 | 4721 | Unidirectional | ||
| Blackfly microvirus-52_ SF02_SP_104 | MK249211 | 4692 | Unidirectional | ||
| Blackfly microvirus-53_ SF02_SP_106 | MK249212 | 4680 | Unidirectional | ||
| Blackfly microvirus-54_ SF02_SP_107 | MK249213 | 4706 | Unidirectional | ||
| Blackfly microvirus-55_ SF02_SP_108 | MK249214 | 4647 | Unidirectional | ||
| Blackfly microvirus-56_ SF02_SP_110 | MK249215 | 4618 | Unidirectional | ||
| Blackfly microvirus-57_ SF02_SP_115 | MK249216 | 4608 | Unidirectional | ||
| Blackfly microvirus-58_ SF02_SP_116 | MK249217 | 4596 | Unidirectional | ||
| Blackfly microvirus-59_ SF02_SP_117 | MK249218 | 4590 | Unidirectional | ||
| Blackfly microvirus-60_ SF02_SP_118 | MK249219 | 4621 | Unidirectional | ||
| Blackfly microvirus-61_ SF02_SP_120 | MK249220 | 4617 | Unidirectional | ||
| Blackfly microvirus-62_ SF02_SP_122 | MK249221 | 4591 | Unidirectional | ||
| Blackfly microvirus-63_ SF02_SP_123 | MK249222 | 4499 | Unidirectional | ||
| Blackfly microvirus-64_ SF02_SP_124 | MK249223 | 4510 | Unidirectional | ||
| Blackfly microvirus-65_ SF02_SP_125 | MK249224 | 4507 | Unidirectional | ||
| Blackfly microvirus-66_ SF02_SP_126 | MK249225 | 4581 | Unidirectional | ||
| Blackfly microvirus-67_ SF02_SP_128 | MK249226 | 4503 | Unidirectional | ||
| Blackfly microvirus-68_ SF02_SP_129 | MK249227 | 4570 | Unidirectional | ||
| Blackfly microvirus-69_ SF02_SP_130 | MK249228 | 4498 | Unidirectional | ||
| Blackfly microvirus-70_ SF02_SP_131 | MK249229 | 4558 | Unidirectional | ||
| Blackfly microvirus-71_ SF02_SP_135 | MK249230 | 4488 | Unidirectional | ||
| Blackfly microvirus-72_ SF02_SP_137 | MK249231 | 4551 | Unidirectional | ||
| Blackfly microvirus-73_ SF02_SP_138 | MK249232 | 4335 | Unidirectional | ||
| Blackfly microvirus-74_ SF02_SP_139 | MK249233 | 4547 | Unidirectional | ||
| Blackfly microvirus-75_ SF02_SP_142 | MK249234 | 4538 | Unidirectional | ||
| Blackfly microvirus-76_ SF02_SP_143 | MK249127 | 4471 | Unidirectional | ||
| Blackfly microvirus-77_ SF02_SP_145 | MK249128 | 4529 | Unidirectional | ||
| Blackfly microvirus-78_ SF02_SP_146 | MK249129 | 4521 | Unidirectional | ||
| Blackfly microvirus-79_ SF02_SP_148 | MK249130 | 4549 | Unidirectional | ||
| Blackfly microvirus-80_ SF02_SP_149 | MK249131 | 4460 | Unidirectional | ||
| Blackfly microvirus-81_ SF02_SP_150 | MK249132 | 4460 | Unidirectional | ||
| Blackfly microvirus-82_ SF02_SP_156 | MK249133 | 4443 | Unidirectional | ||
| Blackfly microvirus-83_ SF02_SP_159 | MK249134 | 4438 | Unidirectional | ||
| Blackfly microvirus-84_ SF02_SP_160 | MK249135 | 4483 | Unidirectional | ||
| Blackfly microvirus-85_ SF02_SP_162 | MK249136 | 4350 | Unidirectional | ||
| Blackfly microvirus-86_ SF02_SP_163 | MK249137 | 4484 | Unidirectional | ||
| Blackfly microvirus-87_ SF02_SP_164 | MK249138 | 4502 | Unidirectional | ||
| Blackfly microvirus-88_ SF02_SP_165 | MK249139 | 4487 | Unidirectional | ||
| Blackfly microvirus-89_ SF02_SP_166 | MK249140 | 4410 | Unidirectional | ||
| Blackfly microvirus-90_ SF02_SP_167 | MK249141 | 4459 | Unidirectional | ||
| Blackfly microvirus-91_ SF02_SP_168 | MK249142 | 4401 | Unidirectional | ||
| Blackfly microvirus-92_ SF02_SP_169 | MK249143 | 4387 | Unidirectional | ||
| Blackfly microvirus-93_ SF02_SP_174 | MK249144 | 4445 | Unidirectional | ||
| Blackfly microvirus-94_ SF02_SP_175 | MK249145 | 4400 | Unidirectional | ||
| Blackfly microvirus-95_ SF02_SP_178 | MK249146 | 4422 | Unidirectional | ||
| Blackfly microvirus-96_ SF02_SP_181 | MK249147 | 4391 | Unidirectional | ||
| Blackfly microvirus-97_ SF02_SP_183 | MK249148 | 4368 | Unidirectional | ||
| Blackfly microvirus-98_ SF02_SP_184 | MK249149 | 4293 | Unidirectional | ||
| Blackfly microvirus-99_ SF02_SP_185 | MK249150 | 4351 | Unidirectional | ||
| Blackfly microvirus-100_ SF02_SP_188 | MK249151 | 4276 | Unidirectional | ||
| Blackfly microvirus-101_ SF02_SP_189 | MK249152 | 4318 | Unidirectional | ||
| Blackfly microvirus-102_ SF02_SP_190 | MK249153 | 4205 | Unidirectional | ||
| Blackfly microvirus-103_ SF02_SP_192 | MK249154 | 4320 | Unidirectional | ||
| Blackfly microvirus-104_ SF02_SP_195 | MK249155 | 4302 | Unidirectional | ||
| Blackfly microvirus-105_ SF02_SP_206 | MK249156 | 4238 | Unidirectional | ||
| Blackfly microvirus-106_ SF02_SP_208 | MK249157 | 4237 | Unidirectional | ||
| Blackfly microvirus-107_ SF02_SP_211 | MK249158 | 4170 | Unidirectional | ||
| Blackfly microvirus-108_ SF02_SP_213 | MK249159 | 4232 | Unidirectional |
Figure 1.
A network and phylogenetic analyses of the circular replication-associated (Rep) protein encoding single-stranded (CRESS) DNA virus replication-associated (Rep) protein sequences. The network analyses shows the clustering of the blackfly CRESS DNA viruses (shown as red circles) within taxonomic family groupings, larger and smaller groupings of unclassified CRESS DNA viruses (only those groups containing CRESS DNA viruses from this study), and singletons from this study that do not cluster with other known Reps. Closest Rep comparisons and percentage similarity are shown for the smaller groups and singletons. Maximum likelihood phylogenetic trees of the Rep sequence of circovirus (showing only the Cyclovirus genus clade) and the five larger unclassified CRESS DNA virus groupings. Blackfly originating sequence names are shown in red. The maximum likelihood phylogenetic tree of the Rep sequence for the genomoviruses is shown in Figure 2.
3.1. Blackfly CRESS DNA Viruses Clustering with Genomoviruses
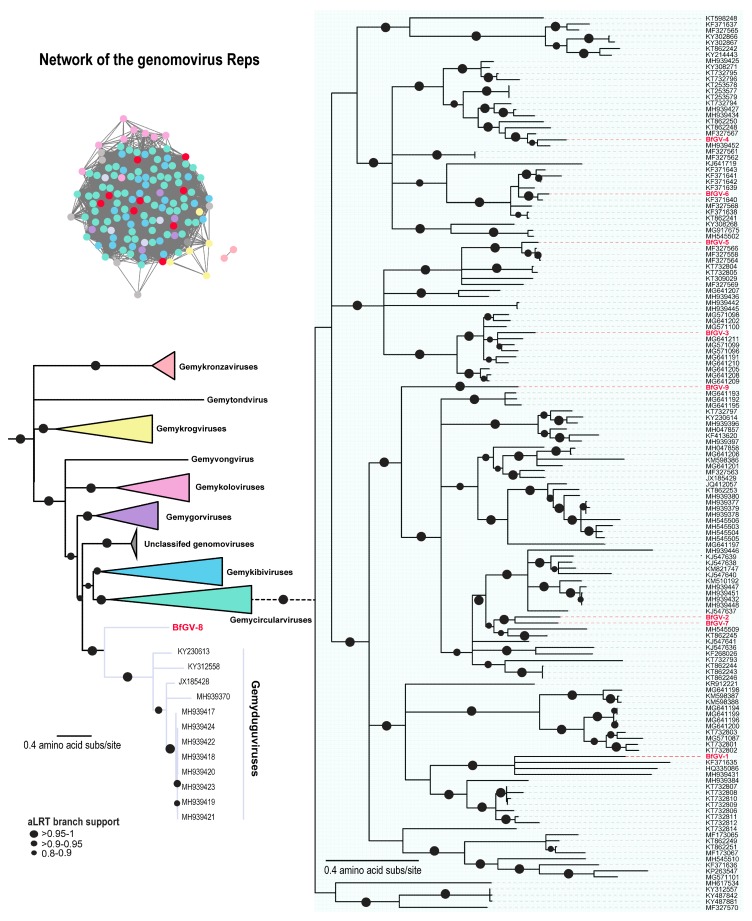
Genomoviruses have been frequently found to be associated with various arthropods [11,14,16,17,20,33,34] in addition to various other samples. Genomoviruses are typically ~2 kb in genome size and likely infect fungi based on a study which showed a genomovirus species, Sclerotinia sclerotiorum hypovirulence virus 1, is able to replicate and cause hypovirulence in Sclerotinia sclerotiorum [35]. Taxonomically, genomoviruses group in nine genera; Gemycircularvirus, Gemyduguivirus, Gemygorvirus, Gemykibivirus, Gemykolovirus, Gemykrogvirus, Gemykronzavirus, Gemytondvirus and Gemyvongvirus. Here we describe nine novel genomoviruses hereby referred to as blackfly genomovirus (BfGV) 1–9 (Table 1, Figure 1 and Figure 2). Eight of these (BfGV 1–7, -9) can be more broadly classified into the Gemycircularvirus genus and BfGV-8 Rep lies basal to that of the members of Gemyduguivirus genus (Figure 2). Based on the 78% full genome species demarcation [36] the seven of the genomoviruses, BfGV-8 whose Rep is basal to those of gemyduguiviruses and six (BfGV-1, -2, -4, -5, -9) assigned to the Gemycircularvirus genus, are new species. The genome of BfGV-3 shares 79% identity with a genomovirus from snowshoe hare faeces (MG611211; [37]) and BfGV-6 shares 81% identity with a genomovirus from porcine faeces (KF371640; [38]. All share a conserved nonanucleotide “TAATATTAT” sequence which is nicked by Rep to initiate replication. The Reps of these genomoviruses share between 33–95% amino acid identities with those of all genomoviruses whereas the capsid proteins (CPs) share 29–79% identity. (Supplementary Data 1). Replication associated motifs in the Rep are conserved throughout the nine genomovirus genomes (Table S2).
Figure 2.
Network and phylogenetic analysis of the Rep sequences of genomovirus, each genus with a corresponding colour and blackfly originating genomoviruses shown in red.
3.2. Divergent CRESS DNA Viruses
A large cohort of Rep-encoding viruses exist that are divergent and do not, at present, fall into classified viral families. These vary in genome organisation but all encode a Rep and putative CP. Several novel CRESS DNA viruses were also discovered in this study, all but one cluster outside major virus family groups and are distributed across five larger network clusters and three smaller clusters contain two to three sequences (Table 1, Figure 1). Nine do not cluster with any other Reps therefore are represented as singletons (Table 1, Figure 1). These range in genome size from ~1.8 to 3 kb and are referred to as blackfly DNA virus (BfV) 1–19 (Table 1). Genome organisation is highly variable, see Table 1 for details on open-reading frame orientations. Interestingly, BfV-18 appears to have an unusual rep which contains two predicted intron regions. All Rep encoding molecules harbour RCR and SF3 helicase motifs, with the exception of BfV-1 and -2 which are apparently missing motif C and RCR motif II, respectively.
The Rep of BfV-7 in group 1 is most closely related to that from a giant house spider (MH545537) [11], sharing 77% Rep identity (Sup data 2). In group 2, the Reps of BfV-12 and -15 share 70% identity with each other, and 68% with giant panda circovirus 1 Rep (MF327573) and 60% with dragonfly larvae-associated circular virus-2 Rep (KF738874) [39], respectively. Group 3 includes Reps of BfV-8 which shares 68% with those of Pacific flying fox faeces-associated circular DNA virus-15 (KT732834) [40] and BfV-11 which shares 70% with sewage-associated circular DNA virus-18 (KM821753) [41]. Reps of BfV-3, -4 and -5 in group 4 share 62% identity with CRESS virus from a minnow (MH617376) [42], and 76% and 68% with llama faeces-associated circular DNA virus-1 (KT862235) [43], respectively. The Rep of BfV-6 in group 5 shares 48% identity with that of Avon-Heathcote estuary-associated circular virus 24 (KM874354) [44].
Several viruses related to those in the family Circoviridae have also been identified in arthropods [11,14,15,45,46,47,48]. Here we identify a blackfly DNA virus (BfV-10) whose Rep clusters with members of the Circoviridae family and it is phylogenetically basal to the cycloviruses (Figure 1). The Rep of BfV-10 share 35–43% amino acid identity to Reps of circoviruses.
Reps of BfV-13, -19 and -16 form small clusters sharing 42–53% identities to their closest related Reps from Bromus-associated circular DNA virus 2 (KM510191) [49], Lytechinus variegatus variable sea urchin-associated circular virus (KR528569) [50], sewage-associated circular DNA virus-2 (KJ547626) [41], CRESS virus sp. isolate ctbg173 (MH617562) [42], Pacific flying fox faeces-associated circular DNA virus-2 (KT732829) [40]. BfV-1, -2, -9, -14, -17 and -18 Reps are singletons sharing 31–42% with Reps of Apis mellifera virus 15 (MH973771) [34], Circoviridae sp. isolate ctbe41 (MH617348) [42], uncultured virus clone CG155 (KY487824) [51], CRESS virus sp. isolate ctdb65 (MH616925) [42], Lake Sarah-associated circular virus-18 (KP153428) [44] and sewage-associated circular DNA virus-1 (KJ547620) [41] (Figure 1).
3.3. Multi-Component Viruses and Circular Rep-Encoding DNA Molecule
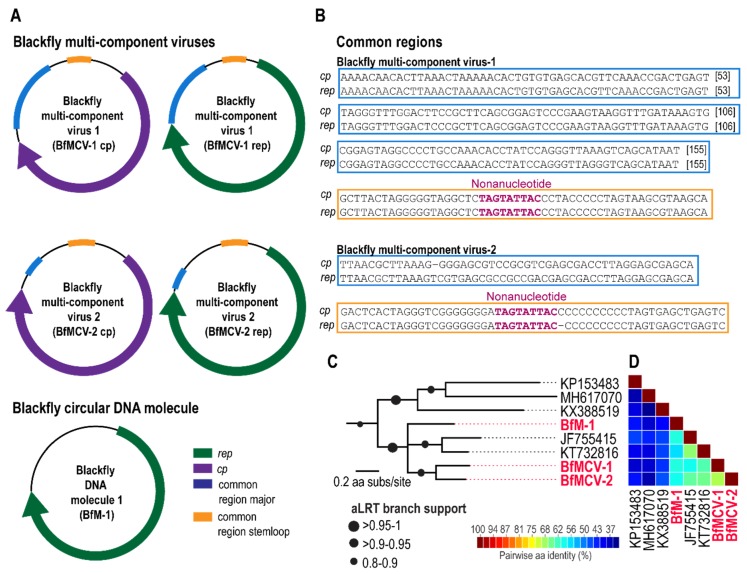
Viruses in the Nanoviridae family infect plants and are comprised of up to eight separate circular DNA components, all encoding a different functional gene including a Rep. Members of another plant infecting virus family, Geminiviridae, are known to have satellite molecules which enhance replication and pathogenicity. More recently a multi-component virus was isolated from faecal samples of Pacific flying fox, this, like other multi-component DNA viruses shares sequence recognition regions in the intergenic regions. In this study, three circular DNA molecules that encode a rep gene were identified. For two of these, cognate molecules encoding a cp gene were identified and therefore these represent two multicomponent viruses (Table 1, Figure 3A–C, Supplementary Data 2). These components have common regions in the intergenic region (Figure 3B), such common regions in multicomponent viruses act as recognition sites for initiation of replication [52]. These, referred to as blackfly multicomponent virus (BfMCV) 1 and 2, have Reps most similar to another multicomponent virus recovered from a Pacific flying fox multicomponent virus (KT732816) [40], sharing 65% and 67% aa similarity, respectively (Figure 3C,D). The cognate BfMCV 1 and 2 CPs share 44% with leaf-footed bug circular genetic element (MH545544) [11] and 43% with Pacific flying fox multicomponent virus (KT732816) [40]. No cognate molecule was identified for the third DNA molecule, referred to as Blackfly DNA molecule 1, however, this molecule is most closely related to the other two multicomponent viruses detected here, sharing 61–62% Rep identity, and to rodent stool associated circular genome virus (JF755415) [53] sharing 62% Rep identity. A cognate molecule encoding a CP may therefore not have been recovered, although no molecule was identified in the high-throughput sequencing data, or this genetic element may represent replication. All molecules have the same nonanucleotide sequence ‘TAGTATTAC’.
Figure 3.
(A) Genome organisation of two blackfly multicomponent DNA viruses (1 and 2) and a DNA molecule indicating open reading frames and common regions (B) Alignment of the common regions from blackfly multicomponent virus 1 and 2 (C) Phylogeny of Rep for the two multicomponent viruses, DNA molecule and closest relatives (D) Pairwise comparison of the Reps show percentage similarity.
3.4. Bacteria-Infecting CRESS DNA Viruses
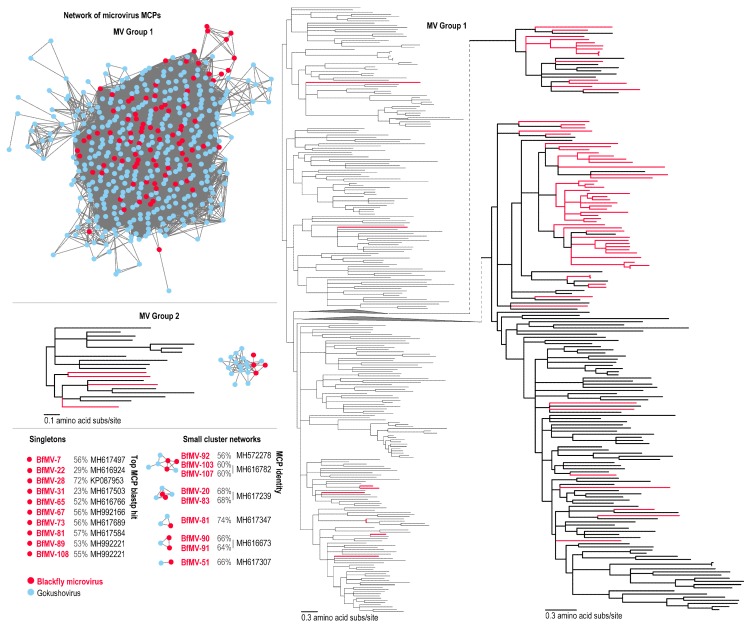
Microviruses, which infect bacteria, are commonly found where their host is present including the microbiome of arthropods [34,54] and harbour unidirectional genomes which encode a replication initiation protein, major and minor capsid proteins and other accessory proteins [19]. Divided into two subfamilies, Gokushovirinae and Bullavirinae (International committee for virus taxonomy: 2018 release; https://talk.ictvonline.org/taxonomy/), microviruses are largely host specific [19]. Here, we identify 108 novel microviruses from blackflies. Using the most well conserved protein, the MCP, we show the wide scope of microvirus diversity present in blackflies (Figure 4). A large proportion of blackfly microviruses (BfMVs) MCPs group with those of gokushoviruses (Figure 4). The MCPs of 88 BfMVs cluster in MV group 1, four in MV group 2 and the remaining are part of small network clusters or are singletons. Although a few BfMVs in the MV group 1 are interspersed throughout the MCP phylogeny, the majority fall in two major clades (Figure 4), sharing between 51–85% MCP identity with other microviruses (Supplementary Data 3). The MCPs of BfMVs in MV group 2, along with the other small network clusters, share 29–74% identity with those of other microviruses (Figure 4 and Supplementary Data 3). The singletons range in similarity to nearest neighbours in the public database sharing between 23–72% MCP protein sequence identity (Figure 4 and Supplementary Data 3).
Figure 4.
Network analyses of the microvirus major capsid protein. Maximum likelihood phylogeny of the MCP sequences of microviruses for the two major groups (MV group 1 and 2) are shown. Smaller networks and singletons are shown with closet Rep comparisons and percentage similarities. Blackfly derived sequence are shown in red.
Many of the newly described microviruses identified in this study group with those isolated from other arthropods such as those from honey bees [34]. Taking into consideration that only 40 blackflies were sampled, the sheer number and diversity of microviruses that have been found here is remarkable. This may, in fact, reflect an equally diverse associated bacterial community in the blackfly and the animals from which they obtain a blood meal. Furthermore, this, taken with the recent recovery of many novel microviruses from arthropods [34,54], indicates the presence of distinct arthropod microvirus populations.
4. Conclusions
The ongoing expansion of our knowledge of the CRESS DNA viruses facilitated by high-throughput sequencing approaches emphasizes the breadth of diversity in various ecosystems and organisms. The diversity and number of CRESS DNA viruses associated with arthropods alone is staggering [11,14,16,20,48]. Here, we report the identification of CRESS DNA viruses associated with blackflies in New Zealand. Nine genomoviruses, 19 unclassified CRESS DNA viruses, two multicomponent viruses, a circular rep-encoding DNA molecule and 108 microviruses. To date, the only multicomponent CRESS viruses in established families are the nanoviruses and geminiviruses which both infect plants. Other than these, our group have identified novel multicomponent viruses previously in faecal matter of Pacific flying foxes [40] and now in blackflies. These appear to be related (i.e., CP and Reps) and thus eludes to the fact that there are other Rep encoding multicomponent viruses [40], given the presence of a completely unique group comprised of the BfMCVs and the pacific flying fox multicomponent virus beyond those that belong to the Geminiviridae and Nanoviridae families. Due to the blood feeding nature of blackflies, these viruses may have originated from the blackflies, the hosts they feed from or surrounding environment. Although further investigation is needed to truly determine the hosts of the CRESS viruses, we know the microviruses infect bacteria and therefore, prominent groups of related microviruses may reflect a specific blackfly bacterial profile. This study provides knowledge on viruses associated with the understudied hematophagous blackfly of New Zealand and further displays the sheer breadth of CRESS DNA viral diversity in arthropods globally.
Supplementary Materials
The following are available online at https://www.mdpi.com/1999-4915/11/6/532/s1, Table S1: Primer details for each recovered blackfly virus genomes and molecules; Table S2: Rolling circle replication and helicase motifs identified in the Reps of the eukaryotic CRESS DNA viruses; Data S1: Pairwise identity comparison of the full genome, Rep and CP sequences of the genomoviruses recovered in this study together with all previously identified.
Author Contributions
Conceptualization, S.K., M.W. and A.V.; methodology, S.K., K.S., R.S.F., M.W. and A.V.; formal analysis, S.K. and A.V.; investigation, S.K., K.S., R.S.F., M.W. and A.V.; resources, A.V.; data curation, S.K. and A.V.; writing—original draft preparation, S.K. and A.V.; writing—review and editing, S.K., K.S., R.S.F., M.W. and A.V.; visualization, S.K. and A.V.; supervision, A.V.; project administration, A.V.; funding acquisition, S.K., M.W. and A.V.
Funding
This research was funded a grant awarded to S.K., M.W. and A.V. from the Brian Mason Scientific and Technical Trust of New Zealand (grant # 2015/09).
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results.
References
- 1.Adler P.H., Cheke R.A., Post R.J. Evolution, epidemiology, and population genetics of black flies (Diptera: Simuliidae) Infect. Genet. Evol. 2010;10:846–865. doi: 10.1016/j.meegid.2010.07.003. [DOI] [PubMed] [Google Scholar]
- 2.Lustigman S., James E.R., Tawe W., Abraham D. Towards a recombinant antigen vaccine against Onchocerca volvulus. Trends Parasitol. 2002;18:135–141. doi: 10.1016/S1471-4922(01)02211-5. [DOI] [PubMed] [Google Scholar]
- 3.Kalluri S., Gilruth P., Rogers D., Szczur M. Surveillance of arthropod vector-borne infectious diseases using remote sensing techniques: A review. PLoS Pathog. 2007;3:e116. doi: 10.1371/journal.ppat.0030116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Hellgren O., Bensch S., Malmqvist B. Bird hosts, blood parasites and their vectors—Associations uncovered by molecular analyses of blackfly blood meals. Mol. Ecol. 2008;17:1605–1613. doi: 10.1111/j.1365-294X.2007.03680.x. [DOI] [PubMed] [Google Scholar]
- 5.Sato Y., Tamada A., Mochizuki Y., Nakamura S., Okano E., Yoshida C., Ejiri H., Omori S., Yukawa M., Murata K. Molecular detection of Leucocytozoon lovati from probable vectors, black flies (Simuliudae) collected in the alpine regions of Japan. Parasitol. Res. 2009;104:251–255. doi: 10.1007/s00436-008-1183-1. [DOI] [PubMed] [Google Scholar]
- 6.Lichty B.D., Power A.T., Stojdl D.F., Bell J.C. Vesicular stomatitis virus: Re-inventing the bullet. Trends Mol. Med. 2004;10:210–216. doi: 10.1016/j.molmed.2004.03.003. [DOI] [PubMed] [Google Scholar]
- 7.Piégu B., Guizard S., Spears T., Cruaud C., Couloux A., Bideshi D.K., Federici B.A., Bigot Y. Complete genome sequence of invertebrate iridovirus IIV-25 isolated from a blackfly larva. Arch. Virol. 2014;159:1181–1185. doi: 10.1007/s00705-013-1918-x. [DOI] [PubMed] [Google Scholar]
- 8.Williams T. Covert iridovirus infection of blackfly larvae. Proc. R. Soc. Lond. Ser. B Biol. Sci. 1993;251:225–230. doi: 10.1098/rspb.1993.0033. [DOI] [Google Scholar]
- 9.Crosby T. Austrosimulium (Austrosimulium) dumbletoni n. sp. from New Zealand (Diptera: Simuliidae) N. Z. J. Zool. 1976;3:17–19. doi: 10.1080/03014223.1976.9517894. [DOI] [Google Scholar]
- 10.Crosskey R.W. Blackflies (Simuliidae) In: Lane R.P., Crosskey R.W., editors. Medical Insects and Arachnids. Springer; Dordrecht, The Netherlands: 1993. pp. 241–287. [DOI] [Google Scholar]
- 11.Rosario K., Mettel K.A., Benner B.E., Johnson R., Scott C., Yusseff-Vanegas S.Z., Baker C.C., Cassill D.L., Storer C., Varsani A., et al. Virus discovery in all three major lineages of terrestrial arthropods highlights the diversity of single-stranded DNA viruses associated with invertebrates. PeerJ. 2018;6:e5761. doi: 10.7717/peerj.5761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Li C.X., Shi M., Tian J.H., Lin X.D., Kang Y.J., Chen L.J., Qin X.C., Xu J., Holmes E.C., Zhang Y.Z. Unprecedented genomic diversity of RNA viruses in arthropods reveals the ancestry of negative-sense RNA viruses. Elife. 2015;4:e05378. doi: 10.7554/eLife.05378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ng T.F., Duffy S., Polston J.E., Bixby E., Vallad G.E., Breitbart M. Exploring the diversity of plant DNA viruses and their satellites using vector-enabled metagenomics on whiteflies. PLoS ONE. 2011;6:e19050. doi: 10.1371/journal.pone.0019050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ng T.F.F., Willner D.L., Lim Y.W., Schmieder R., Chau B., Nilsson C., Anthony S., Ruan Y., Rohwer F., Breitbart M. Broad surveys of DNA viral diversity obtained through viral metagenomics of mosquitoes. PLoS ONE. 2011;6:e20579. doi: 10.1371/journal.pone.0020579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sadeghi M., Altan E., Deng X., Barker C.M., Fang Y., Coffey L.L., Delwart E. Virome of >12 thousand Culex mosquitoes from throughout California. Virology. 2018;523:74–88. doi: 10.1016/j.virol.2018.07.029. [DOI] [PubMed] [Google Scholar]
- 16.Waits K., Edwards M.J., Cobb I.N., Fontenele R.S., Varsani A. Identification of an anellovirus and genomoviruses in ixodid ticks. Vir. Genes. 2018;54:155–159. doi: 10.1007/s11262-017-1520-5. [DOI] [PubMed] [Google Scholar]
- 17.Kraberger S., Hofstetter R.W., Potter K.A., Farkas K., Varsani A. Genomoviruses associated with mountain and western pine beetles. Vir. Res. 2018;256:17–20. doi: 10.1016/j.virusres.2018.07.019. [DOI] [PubMed] [Google Scholar]
- 18.Zhao L., Rosario K., Breitbart M., Duffy S. Eukaryotic circular rep-encoding single-stranded DNA (CRESS DNA) viruses: Ubiquitous viruses with small genomes and a diverse host range. In: Kielian M., Mettenleiter T.C., Roossinck M.J., editors. Advances in Virus Research. Elsevier; Cambridge, MA, USA: 2019. pp. 71–133. [DOI] [PubMed] [Google Scholar]
- 19.Cherwa J.E., Fane B.A. Microviridae: Microviruses and Gokushoviruses. John Wiley & Sons; Hoboken, NJ, USA: 2011. [DOI] [Google Scholar]
- 20.Dayaram A., Potter K.A., Pailes R., Marinov M., Rosenstein D.D., Varsani A. Identification of diverse circular single-stranded DNA viruses in adult dragonflies and damselflies (Insecta: Odonata) of Arizona and Oklahoma, USA. Infect. Genet. Evol. 2015;30:278–287. doi: 10.1016/j.meegid.2014.12.037. [DOI] [PubMed] [Google Scholar]
- 21.Bankevich A., Nurk S., Antipov D., Gurevich A.A., Dvorkin M., Kulikov A.S., Lesin V.M., Nikolenko S.I., Pham S., Prjibelski A.D., et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012;19:455–477. doi: 10.1089/cmb.2012.0021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Altschul S.F., Gish W., Miller W., Myers E.W., Lipman D.J. Basic local alignment search tool. J. Mol. Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 23.Bushnell B. BBMap Short-Read Aligner, and Other Bioinformatics Tools. University of California; Berkeley, CA, USA: 2015. [Google Scholar]
- 24.Ying H., Beifang N., Ying G., Limin F., Weizhong L. CD-HIT Suite: A web server for clustering and comparing biological sequences. Bioinformatics. 2010;26:680–682. doi: 10.1093/bioinformatics/btq003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gerlt J.A., Bouvier J.T., Davidson D.B., Imker H.J., Sadkhin B., Slater D.R., Whalen K.L. Enzyme function initiative-enzyme similarity tool (EFI-EST): A web tool for generating protein sequence similarity networks. Biochim. Biophys. Acta. 2015;1854:1019–1037. doi: 10.1016/j.bbapap.2015.04.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zallot R., Oberg N.O., Gerlt J.A. Democratizedgenomic enzymology web tools for functional assignment. Curr. Opin. Chem. Biol. 2018;47:77–85. doi: 10.1016/j.cbpa.2018.09.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lopes C.T., Franz M., Kazi F., Donaldson S.L., Morris Q., Bader G.D. Cytoscape web: An interactive web-based network browser. Bioinformatics. 2010;26:2347–2348. doi: 10.1093/bioinformatics/btq430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Edgar R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004;32:1792–1797. doi: 10.1093/nar/gkh340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Guindon S., Dufayard J.F., Lefort V., Anisimova M., Hordijk W., Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010;59:307–321. doi: 10.1093/sysbio/syq010. [DOI] [PubMed] [Google Scholar]
- 30.Darriba D., Taboada G.L., Doallo R., Posada D. ProtTest 3: Fast selection of best-fit models of protein evolution. Bioinformatics. 2011;27:1164–1165. doi: 10.1093/bioinformatics/btr088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Stover B.C., Muller K.F. TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinform. 2010;11:7. doi: 10.1186/1471-2105-11-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Muhire B.M., Varsani A., Martin D.P. SDT: A virus classification tool based on pairwise sequence alignment and identity calculation. PLoS ONE. 2014;9:e108277. doi: 10.1371/journal.pone.0108277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kraberger S., Polston J.E., Capobianco H.M., Alcala-Briseno R.I., Fontenele R.S., Varsani A. Genomovirus genomes recovered from Echinothrips americanus sampled in Florida, USA. Genome Announc. 2017;5:e00445-17. doi: 10.1128/genomeA.00445-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kraberger S., Cook C.N., Schmidlin K., Fontenele R.S., Bautista J., Smith B., Varsani A. Diverse single-stranded DNA viruses associated with honey bees (Apis mellifera) Infect. Genet. Evol. 2019;71:179–188. doi: 10.1016/j.meegid.2019.03.024. [DOI] [PubMed] [Google Scholar]
- 35.Yu X., Li B., Fu Y., Jiang D., Ghabrial S.A., Li G., Peng Y., Xie J., Cheng J., Huang J., et al. A geminivirus-related DNA mycovirus that confers hypovirulence to a plant pathogenic fungus. Proc. Natl. Acad. Sci. USA. 2010;107:8387–8392. doi: 10.1073/pnas.0913535107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Varsani A., Krupovic M. Sequence-based taxonomic framework for the classification of uncultured single-stranded DNA viruses of the family Genomoviridae. Vir. Evol. 2017;3 doi: 10.1093/ve/vew037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kraberger S., Waits K., Ivan J., Newkirk E., VandeWoude S., Varsani A. Identification of circular single-stranded DNA viruses in faecal samples of Canada lynx (Lynx canadensis), moose (Alces alces) and snowshoe hare (Lepus americanus) inhabiting the Colorado San Juan Mountains. Infect. Genet. Evol. 2018;64:1–8. doi: 10.1016/j.meegid.2018.06.001. [DOI] [PubMed] [Google Scholar]
- 38.Sikorski A., Massaro M., Kraberger S., Young L.M., Smalley D., Martin D.P., Varsani A. Novel myco-like DNA viruses discovered in the faecal matter of various animals. Vir. Res. 2013;177:209–216. doi: 10.1016/j.virusres.2013.08.008. [DOI] [PubMed] [Google Scholar]
- 39.Dayaram A., Galatowitsch M., Harding J.S., Arguello-Astorga G.R., Varsani A. Novel circular DNA viruses identified in Procordulia grayi and Xanthocnemis zealandica larvae using metagenomic approaches. Infect. Genet. Evol. 2014;22:134–141. doi: 10.1016/j.meegid.2014.01.013. [DOI] [PubMed] [Google Scholar]
- 40.Male M.F., Kraberger S., Stainton D., Kami V., Varsani A. Cycloviruses, gemycircularviruses and other novel replication-associated protein encoding circular viruses in Pacific flying fox (Pteropus tonganus) faeces. Infect. Genet. Evol. 2016;39:279–292. doi: 10.1016/j.meegid.2016.02.009. [DOI] [PubMed] [Google Scholar]
- 41.Kraberger S., Arguello-Astorga G.R., Greenfield L.G., Galilee C., Law D., Martin D.P., Varsani A. Characterisation of a diverse range of circular replication-associated protein encoding DNA viruses recovered from a sewage treatment oxidation pond. Infect. Genet. Evol. 2015;31:73–86. doi: 10.1016/j.meegid.2015.01.001. [DOI] [PubMed] [Google Scholar]
- 42.Tisza M.J., Pastrana D.V., Welch N.L., Stewart B., Peretti A., Starrett G.J., Pang Y.-Y.S., Varsani A., Krishnamurthy S.R., Pesavento P.A. Discovery of several thousand highly diverse circular DNA viruses. BioRxiv. 2019;2019:555375. doi: 10.1101/555375. [DOI] [Google Scholar]
- 43.Steel O., Kraberger S., Sikorski A., Young L.M., Catchpole R.J., Stevens A.J., Ladley J.J., Coray D.S., Stainton D., Dayaram A., et al. Circular replication-associated protein encoding DNA viruses identified in the faecal matter of various animals in New Zealand. Infect. Genet. Evol. 2016;43:151–164. doi: 10.1016/j.meegid.2016.05.008. [DOI] [PubMed] [Google Scholar]
- 44.Dayaram A., Galatowitsch M.L., Arguello-Astorga G.R., van Bysterveldt K., Kraberger S., Stainton D., Harding J.S., Roumagnac P., Martin D.P., Lefeuvre P., et al. Diverse circular replication-associated protein encoding viruses circulating in invertebrates within a lake ecosystem. Infect. Genet. Evol. 2016;39:304–316. doi: 10.1016/j.meegid.2016.02.011. [DOI] [PubMed] [Google Scholar]
- 45.Rosario K., Marinov M., Stainton D., Kraberger S., Wiltshire E.J., Collings D.A., Walters M., Martin D.P., Breitbart M., Varsani A. Dragonfly cyclovirus, a novel single-stranded DNA virus discovered in dragonflies (Odonata: Anisoptera) J. Gen. Virol. 2011;92:1302–1308. doi: 10.1099/vir.0.030338-0. [DOI] [PubMed] [Google Scholar]
- 46.Padilla-Rodriguez M., Rosario K., Breitbart M. Novel cyclovirus discovered in the Florida woods cockroach Eurycotis floridana (Walker) Arch. Virol. 2013;158:1389–1392. doi: 10.1007/s00705-013-1606-x. [DOI] [PubMed] [Google Scholar]
- 47.Wang B., Sun L.D., Liu H.H., Wang Z.D., Zhao Y.K., Wang W., Liu Q. Molecular detection of novel circoviruses in ticks in northeastern China. Ticks Tick Borne Dis. 2018;9:836–839. doi: 10.1016/j.ttbdis.2018.03.017. [DOI] [PubMed] [Google Scholar]
- 48.Dayaram A., Potter K.A., Moline A.B., Rosenstein D.D., Marinov M., Thomas J.E., Breitbart M., Rosario K., Arguello-Astorga G.R., Varsani A. High global diversity of cycloviruses amongst dragonflies. J. Gen. Virol. 2013;94:1827–1840. doi: 10.1099/vir.0.052654-0. [DOI] [PubMed] [Google Scholar]
- 49.Kraberger S., Farkas K., Bernardo P., Booker C., Argüello-Astorga G.R., Mesléard F., Martin D.P., Roumagnac P., Varsani A. Identification of novel Bromus- and Trifolium-associated circular DNA viruses. Arch. Virol. 2015;160:1303–1311. doi: 10.1007/s00705-015-2358-6. [DOI] [PubMed] [Google Scholar]
- 50.Rosario K., Schenck R.O., Harbeitner R.C., Lawler S.N., Breitbart M. Novel circular single-stranded DNA viruses identified in marine invertebrates reveal high sequence diversity and consistent predicted intrinsic disorder patterns within putative structural proteins. Front. Microbiol. 2015;6:696. doi: 10.3389/fmicb.2015.00696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Pearson V.M., Caudle S.B., Rokyta D.R. Viral recombination blurs taxonomic lines: Examination of single-stranded DNA viruses in a wastewater treatment plant. PeerJ. 2016;4:e2585. doi: 10.7717/peerj.2585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Timchenko T., de Kouchkovsky F., Katul L., David C., Vetten H.J., Gronenborn B. A single rep protein initiates replication of multiple genome components of faba bean necrotic yellows virus, a single-stranded DNA virus of plants. J. Virol. 1999;73:10173–10182. doi: 10.1128/jvi.73.12.10173-10182.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Phan T.G., Kapusinszky B., Wang C., Rose R.K., Lipton H.L., Delwart E.L. The fecal viral flora of wild rodents. PLoS Pathog. 2011;7:e1002218. doi: 10.1371/journal.ppat.1002218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Schmidlin K., Kraberger S., Fontenele R.S., De Martini F., Chouvenc T., Gile G.H., Varsani A. Genome sequences of microviruses associated with Coptotermes formosanus. Microbiol. Resour. Announc. 2019;8:e00185-19. doi: 10.1128/MRA.00185-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.