SUMMARY
The plant pathogenic basidiomycete fungi Phakopsora pachyrhizi and Phakopsora meibomiae cause rust disease in soybean plants. Phakopsora pachyrhizi originated in Asia–Australia, whereas the less aggressive P. meibomiae originated in Latin America. In the New World, P. pachyrhizi was first reported in the 1990s to have spread to Hawaii and, since 2001, it has been found in South America. In 2004, the pathogen entered continental USA. This review provides detailed information on the taxonomy and molecular biology of the pathogen, and summarizes strategies to combat the threat of this devastating disease.
Taxonomy: Phakopsora pachyrhizi Syd. & P. Syd; uredial anamorph: Malupa sojae (syn. Uredo sojae); Domain Eukaryota; Kingdom Fungi; Phylum Basidiomycota; Order Uredinales; Class Urediniomycetes; Family Phakopsoraceae; Genus Phakopsora (http://www.indexfungorum.org). The nomenclature of rust spores and spore‐producing structures used within this review follows Agrios GN (2005) Plant Pathology, 5th edn. London: Elsevier/Academic Press.
Host range: In the field, P. pachyrhizi infects leaf tissue from a broad range (at least 31 species in 17 genera) of leguminous plants. Infection of an additional 60 species in other genera has been achieved under laboratory conditions.
Disease symptoms: At the beginning of the disease, small, tan‐coloured lesions, restricted by leaf veins, can be observed on infected soybean leaves. Lesions enlarge and, 5–8 days after initial infection, rust pustules (uredia, syn. uredinia) become visible. Uredia develop more frequently in lesions on the lower surface of the leaf than on the upper surface. The uredia open with a round ostiole through which uredospores are released.
INTRODUCTION
Plant pathogens that cause serious epidemics on crop plants have the power to strongly influence human communities (Stukenbrock and McDonald, 2008). One recent example of such a scenario is the invasion of South and North America by Phakopsora pachyrhizi, the causal agent of Asian soybean rust, from Asia into the Western Hemisphere. Although soybean rust caused by the less aggressive species P. meibomiae has been present in Central and South America since the 1970s, the disease became a major problem only after the invasion of P. pachyrhizi (Yorinori et al., 2005). Currently, P. pachyrhizi is one of the most important economical threats for soybean growers in South America. In 2002–2003, the disease spread through Brazil and caused losses estimated at US$2 billion for 2003 (Yorinori et al., 2005). The disease reached continental USA in 2004 (Schneider et al., 2005). Despite prevalent concerns, the expected high yield losses did not occur in the USA, probably because of unfavourable weather conditions for pathogen spread and/or pathogen populations not fully adapted to the new habitat (Christiano and Scherm, 2007). As the uredospores of P. pachyrhizi are not frost tolerant, the pathogen would need to overwinter in the southern areas of the USA on alternative host plants, such as kudzu (Pueraria lobata). When the temperatures rise in early summer, the pathogen can spread out, but it does not always reach the soybean‐growing areas further north (Hartman and Haudenshield, 2009). Continuous adaptation of the pathogen to novel habitats, such as the continental USA, should be expected. Therefore, new strategies are needed to combat this growing threat.
The pathogen is able to defoliate soybean fields within a few days and may lead to complete crop failure. As soybean varieties with resistance to all isolates of the pathogen population are not yet available, and fungicidal treatments are expensive, soybean cultivation in many areas invaded by P. pachyrhizi is endangered. At present, Asian soybean rust can be controlled by the use of several fungicides in different classes with different modes of action (Miles et al., 2007). However, the level of control depends on disease pressure and the accurate timing of application (Mueller et al., 2009). Furthermore, it has been demonstrated that farnesyl acetate, a naturally occurring plant volatile emitted by rust‐infected plants, negatively affects haustorial development. Thus, this compound might be used to control soybean rust in an environmentally compatible manner (Mendgen et al., 2006). Biological control of soybean rust has not been researched in detail. Fungal hyperparasites growing as a cottony mycelium on pustules of soybean rust in Taiwan and Thailand have been isolated and identified as species of Tuberculina, Verticillium and Trichothecium (Saksirirat and Hoppe, 1990, 1991a). In laboratory and growth chamber experiments, V. psalliotae was found to be particularly effective in colonizing soybean rust pustules on detached leaves and on potted plants, but field tests have not yet been performed (Saksirirat and Hoppe, 1990). It would, however, be overoptimistic to expect that biological control could replace or significantly complement chemical rust control, mainly because of its dependence on high humidity.
This pathogen profile builds on the information presented in the last major review on P. pachyrhizi (Bromfield, 1984) and clarifies the sometimes confusing nomenclature and taxonomy of the fungal pathogen species causing soybean rust, as well as giving an historical overview of the pathogen's global movement from the East to the West. Furthermore, we present knowledge on the host range and infection process. We also describe the current strategies employing molecular genetic tools which are being pursued to establish durable resistance in soybean plants against P. pachyrhizi.
TAXONOMY
Traditionally, the taxonomy of fungi was based on differences in morphological and developmental criteria, such as the type of reproductive organs, spore size or sporogenesis. During recent decades, classifications have been revised using DNA sequence data from different genes which, in some cases, has led to the splitting of an ostensibly uniform species into several new species (de Gruyter et al., 2009; James et al., 2006). In particular, for the rust fungi, morphological and phenotypical characteristics are often unsuitable for an accurate classification because of their very complex life cycles, involving up to six different types of spore, and alterations between different host plants which inherently cause morphological differences (Kirk et al., 2001). Therefore, work is in progress to resolve the relationships amongst rust fungi at the family level by examination of gene sequences (Aime, 2006). According to this study, the family Phakopsoraceae Cummins & Hirats. f. contains a morphologically diverse group of 12–13 different teleomorphic genera and 10 different anamorphic‐form genera. Furthermore, both the family Phakopsoraceae and the genus Phakopsora Dietel itself must be divided into two monophyletic, unrelated lineages. At the species level, the genus Phakopsora contains at least 90 morphologically variable species with an unsatisfactory taxonomic classification (Ono et al., 1992).
HISTORICAL ASPECTS OF NOMENCLATURE
The agent of Asian soybean rust was first recorded in 1902 in Japan by Hennings, and referred to as Uredo sojae Henn. (Hennings, 1903). In 1914, Hans and Paul Sydow described the fungus under its current name, Phakopsora pachyrhizi Syd. & P. Syd, based on an isolate obtained from the leguminous host plant Pachyrhizus erosus (L.) Urb. (=Pachyrhizus angulatus) in Taiwan (Sydow and Sydow, 1914). Since then, phakopsoroid fungi on legumes have been reported several times under confusing taxonomic names, most probably because of a lack of comparative morphological studies (Bromfield, 1984). Generally, such studies are difficult to perform when full life cycle information, including all spore types of a rust fungus, is not available. Today, it is clear that soybean rust is caused by at least one other Phakopsora species, Phakopsora meibomiae (Arthur) Arthur, which is a minor and less aggressive pathogen on soybean (Bonde et al., 2006; Ono et al., 1992). However, historically, taxonomists had made no distinction between the Asian–Australian isolate of the fungus (now referred to as Phakopsora pachyrhizi) and the Latin American isolate (now named Phakopsora meibomiae), and both isolates were seen as belonging to the same species. Based on differences in virulence on soybean, it was proposed that the American isolate may represent a forma specialis which is less well adapted to soybean (Bromfield, 1984). Indications that this interpretation was false came from a comparative study on isoenzymes, which revealed great polymorphisms (Bonde, 1988). The picture of two independent species was further confirmed by morphological differences and the analysis of nucleotide sequences, which were found to be only 80% similar within the ribosomal internal transcribed spacer region of P. pachyrhizi and P. meibomiae (Frederick et al., 2002; Ono et al., 1992).
HISTORY OF DISTRIBUTION
Soybean [Glycine max (L.) Merrill], in the family Fabaceae, is the most economically important species affected by Asian soybean rust (Bromfield, 1984). The cultivation of G. max is thought to have started in the eastern half of north China in the 11th century bc and, from there, it was propagated in the following centuries all over the Eastern Hemisphere (Bromfield, 1984). Unfortunately, the climatic conditions which support soybean cultivation also favour the development of soybean rust disease. Hence, it was not surprising that the disease was first reported from Japan and Taiwan (see above). In 1934, it appeared for the first time in Australia, where soybean has been cultivated since 1920. The first occurrence of soybean rust in Puerto Rico was reported in 1976 (Vakili and Bromfield, 1976) and, since then, it has been observed in Brazil, Colombia and Costa Rica (Bromfield, 1984). It is now clear that these early outbreaks of the disease in Central and South America and the Caribbean Islands were caused by P. meibomiae, the less aggressive Latin American species (Bonde et al., 2006).
The more aggressive Asian–Australian species, P. pachyrhizi, was not reported in the New World until 1994, when it was discovered on soybean in Hawaii (Killgore et al., 1994). From 1997 to 2001, P. pachyrhizi was reported from several southern and central African countries (Levy, 2005). According to Freire et al. (2008), the American continent was free of P. pachyrhizi until 2001. Then, P. pachyrhizi was reported in Paraguay, and became established in Bolivia, Brazil and Argentina in 2002 and 2003 (Freire et al., 2008). In November 2004, Asian soybean rust was reported for the first time in the continental USA (Schneider et al., 2005; Stokstad, 2004). The long‐distance dispersal of the spores from South America to North America was attributed to wind storms and, together with increasing human mobility, possibly was even spread via clothing (Hartman and Haudenshield, 2009; Isard et al., 2005). The spread of P. pachyrhizi within North America, however, has so far been limited, presumably because of a low inoculum available in 2005 and the dry weather conditions in 2006 (Christiano and Scherm, 2007). Teliospore formation by P. pachyrhizi has so far not been observed in North America, and the role of teliospores in the life cycle of the fungus is anyway unclear (see below). In the USA, during the soybean‐free time of the year, P. pachyrhizi overwinters in the south on alternative hosts, such as the broadly distributed immigrant legume kudzu [Pueraria montana (Lour.) Merr. var. lobata (Willd.) Maesen & S.M. Almeida] (Christiano and Scherm, 2007).
HOST RANGE
Unlike the highly specialized grass‐infecting rust fungi, P. pachyrhizi and P. meibomiae have wide host ranges and are able to sporulate on 31 species in 17 genera and 42 species in 19 genera of leguminous plants, respectively (Ono et al., 1992). Remarkably, little overlap was found among legumes infected by both rusts, with the exception of five cultivated crops, i.e. Crotalaria micans Link, G. max, Pa. erosus, Phaseolus lunatus L. and Vigna unguiculata (L.) Walp. (Ono et al., 1992). In addition to the host plants from which the pathogens were isolated in nature, both species are able to infect and sporulate on a range of other plants when artificially inoculated. In this respect, P. pachyrhizi has a broader host range (60 additional species in 26 genera) than P. meibomiae (18 species in 12 genera) (Ono et al., 1992). However, the longer time span needed to sporulate and the less frequently found uredia illustrate the lower degree of compatibility between both Phakopsora species and these additional hosts in comparison with their soybean host.
LIFE CYCLE
The life cycles of rust fungi tend to be complex, involving different types of spores with specialization to particular hosts. Sexual reproduction of the dikaryotic rusts involves the formation of teliospores, karyogamy and reduction division, ending with the formation of four haploid basidiospores which infect the host on which they were formed or, in the case of the heteroecious rusts, an alternative host species. For P. pachyrhizi, there is a lack of information on sexual reproduction. Although teliospores were observed in Asia on several hosts, including soybean, their germination has never been reported from nature (Bromfield, 1984). However, teliospore germination and basidiospore formation can be induced under laboratory conditions (Saksirirat and Hoppe, 1991b). Nevertheless, nothing is known about the basidial infection process or whether P. pachyrhizi is autoecious or heteroecious. Spermatia and aeciospores, rust spore types which are typically formed together with host alternation among heteroecious rusts, are also unknown for this fungus (Bromfield, 1984).
Phakopsora pachyrhizi forms asexual uredospores on short stalks within a uredium 5–8 days after inoculation on colonized leaves (1, 2). Uredospores are released from uredia through an ostiole and dispersed by wind. Under appropriate conditions, uredospores germinate with a single germ tube. Interestingly, unilateral illumination leads to germ tube emergence on the shaded side and growth away from the light (Koch and Hoppe, 1987). Germ tube growth is terminated by the formation of a globose, nonornamented appressorium, approximately the same size as a uredospore (Fig. 2b). As appressoria were also formed on artificial membranes, such as cellulose nitrate membranes, thigmo‐ rather than chemo‐differentiation seems to be involved (Koch and Hoppe, 1988). In contrast with uredospore‐derived infections by most other rust fungi, P. pachyrhizi penetrates leaves directly rather than using stomatal openings (Fig. 3). However, penetration by P. pachyrhizi through stomata has been observed occasionally on nonhost plants, such as barley, and, in this case, the fungus was able to colonize the mesophyll, but never sporulated (M. Loehrer and U. Schaffrath, unpublished work). However, infection by basidiospores in most rust species employs the direct mode of cuticle penetration (Heath, 1997; Mellersh and Heath, 2001). Penetration by P. pachyrhizi starts with the formation of a funnel‐shaped structure, termed the appressorial cone, within the appressorium (Fig. 3) (Koch et al., 1983). This cone is contiguous with the cell wall of the penetration hypha, which is also referred to as the transepidermal vesicle (Bromfield, 1984 and references therein). On penetration, the epidermal cells collapse (Fig. 4), become disorganized and show signs of cell death, as evidenced by trypan blue staining (2, 3) (Keogh et al., 1980). After penetration, the hypha grows through the epidermal cell and reaches the intercellular space; a septum is formed which separates it from the emerging primary hypha (Koch et al., 1983), sometimes also termed the invading hypha (Bromfield, 1984). The primary hypha may branch to form secondary hyphae and, finally, haustorium mother cells differentiate in close contact with mesophyll cells (1, 2) (Koch et al., 1983). The formation of the first haustoria was observed at 24–48 h after initial contact between uredospores and a susceptible host (Fig. 2c). Thereafter, intense colonization of the mesophyll tissue occurs and the intercellular spaces of the leaves become filled with fungal mycelium (Fig. 1). The asexual reproduction of P. pachyrhizi is completed when newly developed uredia originate from uredial primordia (Koch et al., 1983).
Figure 1.
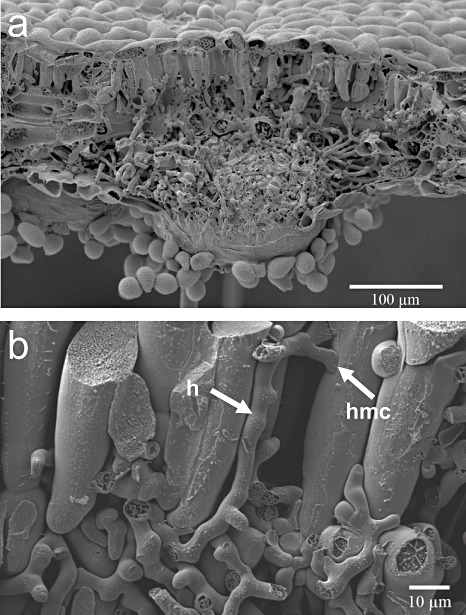
Scanning electron micrographs showing soybean leaves colonized by P. pachyrhizi. a) Section through a leaf blade with an uredium. Intercellular spaces are heavily colonized by P. pachyrhizi mycelium and dispersed uredospores are present at the lower leaf surface. b) Branched hyphae (h) are formed in the intercellular space of soybean leaves. Haustorial mother cells (hmc) get in close contact with particular mesophyll cells which are subsequently invaded and mature haustoria are formed. (Permission and copyrights of figures are owned by Bayer CropSience AG, Monheim Germany.)
Figure 2.
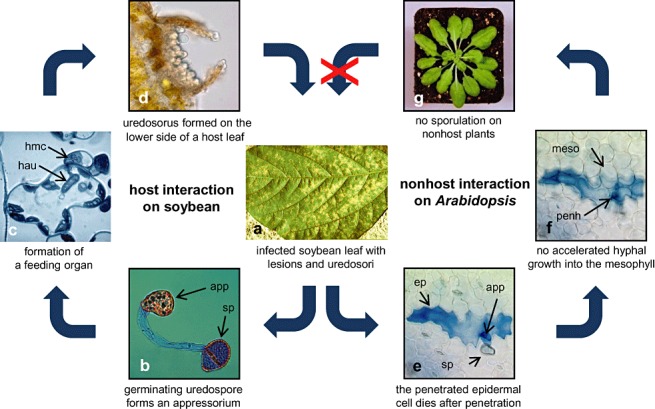

Interaction of P. pachyrhizi with its host soybean and the nonhost plant Arabidopsis. a) Upon infection P. pachyrhizi forms uredia which are located mainly on the lower side of the leaf. Newly formed uredospores are dispersed by wind. b) Spores landing on leaves germinate and form an appressoria as depicted in the interference contrast micrograph. c) Intercellular hyphae form haustorial mother cells (hmc) from which haustoria (hau) develop inside mesophyll cells. d) The life cycle of P. pachyrhizi is completed with the formation of uredospores in uredia. e) In the nonhost interaction between the fungus and Arabidopsis uredospores germinate, form appressoria and penetrate epidermal cells as known from the host type of interaction. Similarly penetrated epidermal cells of host and nonhost plants die, as indicated after trypan‐blue staining. f) Fungal growth is restricted at the mesophyll boundary. Pictures shown in e and f are optical sections from the same infection site focused either on the epidermal or mesophyll layer. g) P. pachyrhizi is unable to complete its life cycle and does not sporulate on wild‐type Arabidopsis plants. sp, uredospore; app, appressorium; epi, epidermal cell; hau, haustorium; hmc, haustorial mother cell; penh, penetration hypha; meso, mesophyll cell. (Fig. 2c reprinted from Koch et al., 1983).
Figure 3.
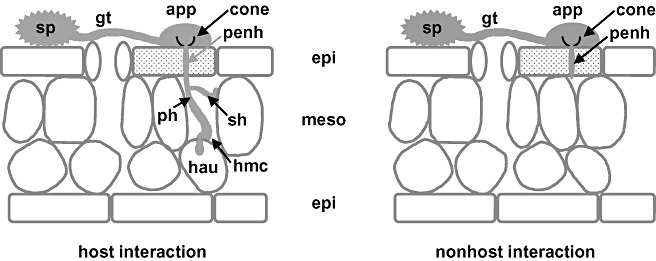
Initial developmental stages during interactions of P. pachyrhizi with host and nonhost plants (schematic). Germinating uredospores (sp) produce a single germ tube (gt) terminated with an appressorium (app). A funnel‐like structure, the appressorial cone (cone), is built inside the appresorium and continues into a penetration hypha (penh) which traverses epidermal cells (epi). Penetrated cells, in consequence, die (punctated). In the intercellular space of the mesophyll a primary hypha (ph) is separated from the penetration hypha by a septum and branches into several secondary hyphae (sh). Finally , a haustorial mother cell (hmc) is formed from which the pathogen invades a mesophyll cell (meso) forming the first haustorium (hau) are established in the host interaction (modified from Koch et al. 1983). Left side: host interaction showing all developmental stages of infection; right side: nonhost interaction with e.g. wild‐type Arabidopsis plants. Fungal invasion is stopped the latest after transversion of the epidermal cell at the border to the mesophyll.
Figure 4.
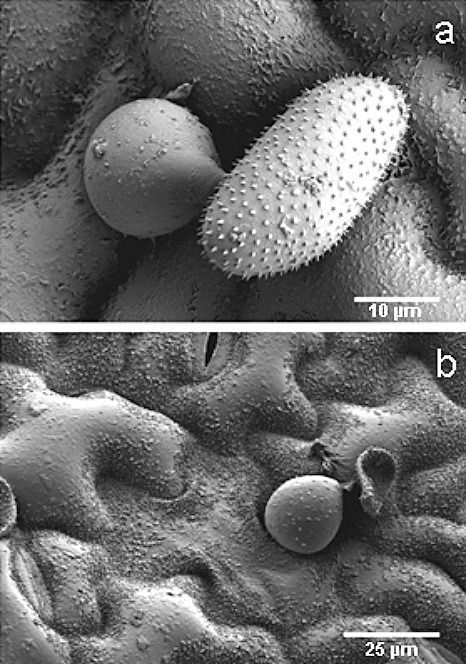
Infection structures of P. pachyrhizi (race Thai 1) on the adaxial leaf surface of soybean (cv. Erin). a) Early stage of appressorium formation with a short germ tube, six hours post inoculation (h p.i.). b) Subsequent invasion of the fungus into the epidemis causes cell collapse and death of penetrated cells as observed at 24 h p.i. Low temperature cryo scanning electron microscopy by Prof. Kurt Mendgen, University of Konstanz, Germany.
RESISTANCE AGAINST P. PACHYRHIZI IN SOYBEAN
Breeding for resistance to P. pachyrhizi has been conducted by classical germplasm screens based on three infection phenotypes: ‘Tan’, ‘RB’ and ‘immune’. Susceptible interactions (Tan) are characterized by tan‐coloured lesions with sporulating uredia, whereas resistant cultivars generally develop reddish‐brown lesions with little or no sporulation (RB). The number of uredia on the lower leaf surface is generally two to five per Tan lesion and zero to two per RB lesion. The so‐called ‘immune’ phenotype is an incompatible interaction without any visible disease symptoms on host leaves (Bromfield, 1984; Pham et al., 2009). None of the soybean cultivars in present commercial production is resistant to all P. pachyhrizi isolates (Hartman et al., 2005). However, five accessions have been identified which each show resistance to certain isolates of P. pachyrhizi. On this basis, five hypothetical major resistance genes (R genes), Rpp1, Rpp2, Rpp3, Rpp4 and Rpp5, have been predicted (Bromfield,, 1984; Bromfield and Hartwig, 1980; Hartwig and Bromfield, 1983; Morceli et al., 2008). For example, the soybean accession PI 230970, which contains the soybean resistance gene Rpp2, produces an RB‐type reaction in response to inoculation with P. pachyrhizi isolate Taiwan 72‐1, whereas inoculation with other isolates of P. pachryhizi, including Taiwan 80‐2, results in a fully susceptible Tan reaction (Hartman et al., 2005).
Although all Rpp genes have been mapped to particular linkage groups (LGs), Rpp1 on LG‐G, Rpp2 on LG‐J, Rpp3 on LG‐C2, Rpp4 on LG‐G and Rpp5 on LG‐N, so far none of the genes have been cloned (Bromfield,, 1984; Bromfield and Hartwig, 1980; Hartwig and Bromfield, 1983; Hyten et al., 2007; Morceli et al., 2008). Most progress has been made for Rpp4‐mediated resistance by identifying a single candidate gene within a 2 cM region on chromosome 18, which shows sequence similarity to the lettuce RGC2 family of so‐called coiled‐coil nucleotide‐binding site leucine‐rich repeat (NBS‐LRR) resistance genes (Meyer et al., 2009). Although Rpp2‐ and Rpp4‐mediated resistances have been stable for 6 years in Brazil and more than 20 years in Asia, respectively (Hartman et al., 2005), single, dominantly inherited R gene‐mediated resistance against P. pachyrhizi has been overcome in nature several times because of the great capacity of the fungus to develop new races (Bromfield, 1984). Generally, this scenario of the breakdown of R gene‐mediated resistance is known as the ‘boom and bust’ syndrome. In addition to pyramiding known Rpp resistance genes into modern cultivars to create a more durable and broad‐spectrum disease resistance, the recruitment of novel sources of resistance to P. pachyrhizi is desirable. Thus, the analysis of more than 16 000 soybean accessions has led to the identification of at least 22 additional accessions carrying dominant resistance that is not governed by Rpp2 or Rpp4 (Garcia et al., 2008; Hartman et al., 2005; Monteros et al., 2007; Oloka et al., 2008; Pham et al., 2009; Pierozzi et al., 2008). Furthermore, at least two different recessive sources of resistance are known (Calvo et al., 2008; Pierozzi et al., 2008). Interestingly, one of these loci was identified as a putative recessive allele of Rpp2 (Garcia et al., 2008). Recessively inherited types of resistance seem to be mechanistically different from R gene‐mediated resistance and might be based on a lack of compatibility (Panstruga, 2003). Thus, the identification of the corresponding pathogen genes might also provide new insights into the requirements for the establishment of the biotrophic interaction between P. pachyrhizi and soybean.
A prerequisite for the introduction of known Rpp genes into commercially deployable, high‐yielding soybean cultivars is the association of phenotypic features with molecular markers. So far, simple sequence repeat (SSR) markers have been identified for Rpp1, Rpp2, Rpp4, Rpp5 and the novel ‘Hyuuga’ red–brown lesion resistance gene, which can now be used for marker‐assisted selection (Garcia et al., 2008; Hyten et al., 2007; Monteros et al., 2007; Silva et al., 2008). In addition, screening for resistance to P. pachyrhizi has been expanded to close relatives of soybean, such as G. soja and other perennial Glycine ssp. (Hartman, 1992; Hartman et al., 2005). Once identified, these traits can be introduced into high‐yielding soybean lines by crossing. Alternative strategies include breeding for tolerance, i.e. the selection of plants which, under field conditions, react to infection with P. pachyrhizi with less yield loss (Hartman et al., 2005).
MOLECULAR ASPECTS OF RESISTANCE TO P. PACHYRHIZI
A large‐scale transcript profiling approach conducted with Rpp1 soybean plants has revealed an up‐regulation in the expression of genes for lipoxygenase and peroxidase in an incompatible interaction, suggesting an important function for these genes in Rpp1‐mediated resistance (Choi et al., 2008). Biphasic gene expression in response to P. pachyrhizi infection has been found in soybean plants with Rpp2‐mediated resistance (van de Mortel et al., 2007). Differences in gene expression between compatible and incompatible interactions peaked at 12 h post‐inoculation (hpi) and returned to almost basal levels by 24 hpi. At later stages, starting at 72 hpi, a second, smaller wave of defence gene expression could be observed, which was significantly earlier in the resistant than in the susceptible interaction. The early transcriptional response observed in susceptible and resistant plants might represent a general response of soybean to the nonspecific recognition of any pathogen, presumably by interaction with microbe‐associated or microbe‐induced molecular patterns (MAMPs and MIMPs) (Mackey and McFall, 2006). By contrast, the induction of genes involved in the phenylpropanoid pathway was observed during the second wave of gene expression at least 1 day earlier in the resistant than in the susceptible soybean variety. A similar up‐regulation of genes involved in this pathway was also found in Rpp1‐carrying plants after infection with compatible or incompatible P. pachyrhizi isolates (Choi et al., 2008). It remains to be seen whether Rpp1‐ and Rpp2‐mediated resistances employ similar defence mechanisms. Based on whole‐genome expression analyses, it seems that timely induction rather than the involvement of specific genes determines the outcome of the interaction between soybean and P. pachyrhizi (van de Mortel et al., 2007). In addition, even the growth stage of soybean at the time of infection is a critical parameter which needs to be considered when assaying the gene induction pattern (Panthee et al., 2009).
Complementary to transcriptional analyses in the host, gene transcript profiling has also been performed with P. pachyrhizi (Posada‐Buitrago and Frederick, 2005; Tremblay et al., 2009). As it is not trivial to separate infection structures of the biotrophic pathogen from host tissue, analysis was performed, for example, with uredospores germinated in water (Posada‐Buitrago and Frederick, 2005). In this study, 499 unique expressed sequence tags (ESTs) were identified, comprising approximately 4%–5% of the totally expressed genes which could be estimated for filamentous fungi. Recently, a cDNA library was constructed from uredinia separated from host tissue by laser‐captured microdissection (Tremblay et al., 2009). About 80% of genes identified in this study shared no homology to previously described Phakopsora genes. This result demonstrates stage‐specific gene expression in the development of uredinia.
ALTERNATIVE SOURCES FOR RESISTANCE TO P. PACHYRHIZI
Most recently, a study was performed to investigate the nonhost resistance of Arabidopsis against P. pachyrhizi (Loehrer et al., 2008). Nonhost resistance is defined as the resistance of all genotypes of a given plant species against all genotypes of a fungal species for which it is not considered to be a host (Heath, 2000). Similar to the interactions of P. pachyrhizi with other nonhost plants (Hoppe and Koch, 1989), on wild‐type Arabidopsis germination, appressorium formation and transit of the fungus through penetrated epidermal cells were not altered phenotypically in comparison with the soybean host. The predominant involvement of jasmonic acid‐related defences during the initial stages of infection raised the interesting question as to whether P. pachyrhizi mimics a necrotroph to circumvent salicylic acid‐related defences, which potentially might be more effective against this biotrophic pathogen (Loehrer et al., 2008). Additional evidence that the success of invasion by P. pachyrhizi depends on the killing of penetrated epidermal cells comes from the observation that barley plants expressing the cell death BAX inhibitor‐1 protein are less frequently invaded by the pathogen than are nontransformed plants (Hoefle et al., 2009). Certainly, more research is needed to identify potential candidate genes that could be used to engineer sustainable resistance into soybean against P. pachyrhizi.
CONCLUDING REMARKS AND FURTHER PERSPECTIVES
Until now, research on Asian soybean rust has mainly focused on host range studies, epidemiology and the evaluation of yield loss and control measures. These studies have mainly centred on the diseased plant. However, as soybean cultivars with resistance to all known races of P. pachyrhizi are not yet available, and because it is questionable whether they will ever exist, more basic research is also needed on the pathogen itself. Crucial elements in the infection process of the fungus and possibilities of interference to adapt novel plant protection strategies need to be identified. One important step forward in this direction is the assessment of fungal gene expression during distinct phases of the plant–pathogen interaction. So far, Posada‐Buitrago and Frederick (2005) have reported the EST clones characteristic for the germination stage of P. pachyrhizi on soybean. However, more information is required on gene transcripts which are specifically up‐regulated during appressorium formation, epidermal penetration, invasive growth and, in particular, haustorium formation to obtain a complete picture on the requirements for the pathogen's asexual life cycle. Experimental access towards this goal has been provided by Voegele and co‐workers who have analysed the interaction between Uromyces fabae and Vicia faba (Kemen et al., 2005; Link and Voegele, 2008). A methodological bottle‐neck, however, is the inability to transform rust fungi and to utilize established tools of reverse and forward genetics. The application of these molecular technologies, however, will probably determine whether the threat of Asian soybean rust will successfully be averted in the major areas of soybean cultivation.
ACKNOWLEDGEMENTS
We gratefully acknowledge comments on the manuscript by Brian Deverall and Alan J. Slusarenko. Electron micrographs were kindly provided by Kurt Mendgen and Bayer CropScience AG. KG and work on soybean rust in the Department of Plant Physiology is supported by BASF Plant Science Company GmbH. ML is supported by a grant from RWTH Aachen University.
REFERENCES
- Aime, M.C. (2006) Toward resolving family‐level relationships in rust fungi (Uredinales). Mycoscience, 47, 112–122. [Google Scholar]
- Bonde, M.R. (1988) A comparison of isoenzymes of Phakopsora pachyrhizi from the eastern hemisphere and the new world. Phytopathology, 78, 1491–1494. [Google Scholar]
- Bonde, M.R. , Nester, S.E. , Austin, C.N. , Stone, C.L. , Frederick, R.D. , Hartman, G.L. and Miles, M.R. (2006) Evaluation of virulence of Phakopsora pachyrhizi and P. meibomiae isolates. Plant Dis. 90, 708–716. [DOI] [PubMed] [Google Scholar]
- Bromfield, K.R. (1984) Soybean Rust. St. Paul, MN: American Phytopathological Society. [Google Scholar]
- Bromfield, K.R. and Hartwig, E.E. (1980) Resistance to soybean rust and mode of inheritance. Crop. Sci. 20, 254–255. [Google Scholar]
- Calvo, E.S. , Kiihl, R.A.S. , Garcia, A. , Harada, A. and Hiromoto, D.M. (2008) Two major recessive soybean genes conferring soybean rust resistance. Crop. Sci. 48, 1350–1354. [Google Scholar]
- Choi, J. , Alkharouf, N. , Schneider, K. , Matthews, B. and Frederick, R. (2008) Expression patterns in soybean resistant to Phakopsora pachyrhizi reveal the importance of peroxidases and lipoxygenases Funct. Integr. Genomics, 8,314–359. [DOI] [PubMed] [Google Scholar]
- Christiano, R.S.C. and Scherm, H. (2007) Quantitative aspects of the spread of Asian soybean rust in the southeastern United States, 2005–2006. Phytopathology, 97, 1428–1433. [DOI] [PubMed] [Google Scholar]
- Frederick, R.D. , Snyder, C.L. , Peterson, G.L. and Bonde, M.R. (2002) Polymerase chain reaction assays for the detection and discrimination of the soybean rust pathogens Phakopsora pachyrhizi and P. meibomiae . Phytopathology, 92, 217–227. [DOI] [PubMed] [Google Scholar]
- Freire, M.C.M. , De Oliveira, L.O. , De Almeida, A.M.R. , Schuster, I. , Moreira, M.A. , Liebenberg, M.M. and Mienie, C.M.S. (2008) Evolutionary history of Phakopsora pachyrhizi (the Asian soybean rust) in Brazil based on nucleotide sequences of the internal transcribed spacer region of the nuclear ribosomal DNA. Genet. Mol. Biol. 31, 920–931. [Google Scholar]
- Garcia, A. , Calvo, É. , Souza Kiihl, R. , Harada, A. , Hiromoto, D. and Vieira, L. (2008) Molecular mapping of soybean rust (Phakopsora pachyrhizi) resistance genes: discovery of a novel locus and alleles. Theor. Appl. Genet. 117, 545–553. [DOI] [PubMed] [Google Scholar]
- De Gruyter, J. , Aveskamp, M.M. , Woudenberg, J.H.C. , Verkley, G.J.M. , Groenewald, J.Z. and Crous, P.W. (2009) Molecular phylogeny of Phoma and allied anamorph genera: towards a reclassification of the Phoma complex. Mycological. Res. 113, 508–519. [DOI] [PubMed] [Google Scholar]
- Hartman, G.L. (1992) Sources of resistance to soybean rust in perennial Glycine species. Plant Dis. 76, 396–399. [Google Scholar]
- Hartman, G.L. and Haudenshield, J.S. (2009) Movement of Phakopsora pachyrhizi (soybean rust) spores by non‐conventional means. Eur. J. Plant Pathol. 123, 225–228. [Google Scholar]
- Hartman, G.L. , Miles, M.R. and Frederick, R.D. (2005) Breeding for resistance to soybean rust. Plant Dis. 89, 664–666. [DOI] [PubMed] [Google Scholar]
- Hartwig, E.E. and Bromfield, K.R. (1983) Relationships among 3 genes conferring specific resistance to rust in soybeans. Crop. Sci. 23, 237–239. [Google Scholar]
- Heath, M.C. (1997) Signalling between pathogenic rust fungi and resistant or susceptible host plants. Ann. Bot. 80, 713–720. [Google Scholar]
- Heath, M.C. (2000) Nonhost resistance and nonspecific plant defenses. Curr. Opin. Plant Biol. 3, 315–319. [DOI] [PubMed] [Google Scholar]
- Hennings, P. (1903) Einige neue japanische Uredinales (in German). Hedwigia, IV (Suppl.), 107–108. [Google Scholar]
- Hoefle, C. , Loehrer, M. , Schaffrath, U. , Frank, M. , Schultheiss, H. and Hückelhoven, R. (2009) Transgenic suppression of cell death limits penetration success of the soybean rust fungus Phakopsora pachyrhizi into epidermal cells of barley. Phytopathology, 99, 220–226. [DOI] [PubMed] [Google Scholar]
- Hoppe, H.H. and Koch, E. (1989) Defense reactions in host and nonhost plants against the soybean rust fungus (Phakopsora pachyrhizi Syd). J. Phytopathol. 125, 77–88. [Google Scholar]
- Hyten, D.L. , Hartman, G.L. , Nelson, R.L. , Frederick, R.D. , Concibido, V.C. , Narvel, J.M. and Cregan, P.B. (2007) Map location of the Rpp1 locus that confers resistance to soybean rust in soybean. Crop. Sci. 47, 837–838. [Google Scholar]
- Isard, S.A. , Gage, S.H. , Comtois, P. and Russo, J.M. (2005) Principles of the atmospheric pathway for invasive species applied to soybean rust. Bioscience, 55, 851–861. [Google Scholar]
- James, T.Y. , Kauff, F. , Schoch, C.L. , Matheny, P.B. , Hofstetter, V. , Cox, C.J. , Celio, G. , Gueidan, C. , Fraker, E. , Miadlikowska, J. , Lumbsch, H.T. , Rauhut, A. , Reeb, V. , Arnold, A.E. , Amtoft, A. , Stajich, J.E. , Hosaka, K. , Sung, G‐H , Johnson, D. , O'Rourke, B. , Crockett, M. , Binder, M. , Curtis, J.M. , Slot, J.C. , Wang, Z. , Wilson, A.W. , Schuszler, A. , Longcore, J.E. , O'Donnell, K. , Mozley‐Standridge, S. , Porter, D. , Letcher, P.M. , Powell, M.J. , Taylor, J.W. , White, M.M. , Griffith, G.W. , Davies, D.R. , Humber, R.A. , Morton, J.B. , Sugiyama, J. , Rossman, A.Y. , Rogers, J.D. , Pfister, D.H. , Hewitt, D. , Hansen, K. , Hambleton, S. , Shoemaker, R.A. , Kohlmeyer, J. , Volkmann‐Kohlmeyer, B. , Spotts, R.A. , Serdani, M. , Crous, P.W. , Hughes, K.W. , Matsuura, K. , Langer, E. , Langer, G. , Untereiner, W.A. , Lucking, R. , Budel, B. , Geiser, D.M. , Aptroot, A. , Diederich, P. , Schmitt, I. , Schultz, M. , Yahr, R. , Hibbett, D.S. , Lutzoni, F. , McLaughlin, D.J. , Spatafora, J.W. and Vilgalys, R. (2006) Reconstructing the early evolution of Fungi using a six‐gene phylogeny. Nature, 443, 818–822. [DOI] [PubMed] [Google Scholar]
- Kemen, E. , Kemen, A.C. , Rafiqi, M. , Hempel, U. , Mendgen, K. , Hahn, M. and Voegele, R.T. (2005) Identification of a protein from rust fungi transferred from haustoria into infected plant cells. Mol. Plant–Microbe Interact. 18, 1130–1139. [DOI] [PubMed] [Google Scholar]
- Keogh, R.C. , Deverall, B.J. and Mcleod, S. (1980) Comparison of histological and physiological responses to Phakopsora pachyrhizi in resistant and susceptible soybean. Trans. Br. Mycol. Soc. 74, 329–333. [Google Scholar]
- Killgore, E. , Heu, R. and Gardner, D.E. (1994) First report of soybean rust in Hawaii. Plant Dis. 78, 1216–1216. [Google Scholar]
- Kirk, P. , Cannon, P. , David, J. and Stalpers, J. (2001) Dictionary of the Fungi, 9th edn. Wallingford: CABI. [Google Scholar]
- Koch, E. and Hoppe, H.H. (1987) Effect of light on uredospore germination and germ tube growth of soybean rust (Phakopsora pachyrhizi Syd). J. Phytopathol. Phytopathol. Z. 119, 64–74. [Google Scholar]
- Koch, E. and Hoppe, H.H. (1988) Development of infection structures by the direct‐penetrating soybean rust fungus (Phakopsora pachyrhizi Syd) on artificial membranes. J. Phytopathol. Phytopathol. Z. 122, 232–244. [Google Scholar]
- Koch, E. , Ebrahimnesbat, F. and Hoppe, H.H. (1983) Light and electron‐microscopic studies on the development of soybean rust (Phakopsora pachyrhizi Syd) in susceptible soybean leaves. Phytopathol. Z. 106, 302–320. [Google Scholar]
- Levy, C. (2005) Epidemiology and chemical control of soybean rust in southern Africa. Plant Dis. 89, 669–674. [DOI] [PubMed] [Google Scholar]
- Link, T.I. and Voegele, R.T. (2008) Secreted proteins of Uromyces fabae: similarities and stage specificity. Mol. Plant Pathol. 9, 59–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loehrer, M. , Langenbach, C. , Goellner, K. , Conrath, U. and Schaffrath, U. (2008) Characterization of nonhost resistance of Arabidopsis to the Asian soybean rust. Mol. Plant–Microbe Interact. 21, 1421–1430. [DOI] [PubMed] [Google Scholar]
- Mackey, D. and McFall, A.J. (2006) MAMPs and MIMPs: proposed classifications for inducers of innate immunity. Mol. Microbiol. 61, 1365–1371. [DOI] [PubMed] [Google Scholar]
- Mellersh, D.G. and Heath, M.C. (2001) Plasma membrane‐cell wall adhesion is required for expression of plant defense responses during fungal penetration. Plant Cell, 13, 413–424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mendgen, K. , Wirsel, S.G.R. , Jux, A. , HoVmann, J. and Boland, W. (2006) Volatiles modulate the development of plant pathogenic rust fungi. Planta, 224, 1353–1361. [DOI] [PubMed] [Google Scholar]
- Meyer, J.D.F. , Silva, D.C.G. , Yang, C. , Pedley, K.F. , Zhang, C. , Van De Mortel, M. , Hill, J.H. , Shoemaker, R.C. , Abdelnoor, R.V. , Whitham, S.A. and Graham, M.A. (2009) Identification and analyses of candidate genes for Rpp4‐mediated resistance to Asian soybean rust in Soybean. Plant Physiol. 150, 295–307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miles, M.R. , Levy, C. , Morel, W. , Mueller, T. , Steinlage, T. , Rij, N. , Frederick, R.D. and Hartman, G.L. (2007) International fungicide efficacy trials for the management of soybean rust. Plant Dis. 91, 1450–1458. [DOI] [PubMed] [Google Scholar]
- Monteros, M.J. , Missaoui, A.M. , Phillips, D.V. , Walker, D.R. and Boerma, H.R. (2007) Mapping and confirmation of the ‘Hyuuga’ red–brown lesion resistance gene for Asian soybean rust. Crop. Sci. 47, 829–836. [Google Scholar]
- Morceli, T.G.S. , Trevisoli, S.H.U. , Morceli, J.A.A. , Kiih, E.S. , Di Mauro, A.O. and Garcia, A. (2008) Identification and validation of microsatellite markers linked to the Rpp5 gene conferring resistance to Asian soybean rust. Pesq. Agropec. Bras. 43, 1533–1541. [Google Scholar]
- Van De Mortel, M. , Recknor, J.C. , Graham, M.A. , Nettleton, D. , Dittman, J.D. , Nelson, R.T. , Godoy, C.V. , Abdelnoor, R.V. , Almeida, A.M.R. , Baum, T.J. and Whitham, S.A. (2007) Distinct biphasic mRNA changes in response to Asian soybean rust infection. Mol. Plant–Microbe Interact. 20, 887–899. [DOI] [PubMed] [Google Scholar]
- Mueller, T.A. , Miles, M.R. , Morel, W. , Marois, J.J. , Wright, D.L. , Kemerait, R.C. , Levy, C. and Hartman, G.L. (2009) Effect of fungicide and timing of application on soybean rust severity and yield. Plant Dis. 93, 243–248. [DOI] [PubMed] [Google Scholar]
- Oloka, H.K. , Tukamuhabwa, P. , Sengooba, T. and Shanmugasundram, S. (2008) Reaction of exotic soybean germplasm to Phakopsora pachyrhizi in Uganda. Plant Dis. 92, 1493–1496. [DOI] [PubMed] [Google Scholar]
- Ono, Y. , Buritica, P. and Hennen, J.F. (1992) Delimitation of Phakopsora, Physopella and Cerotelium and their species on Leguminosae . Mycol. Res. 96, 825–850. [Google Scholar]
- Panstruga, R. (2003) Establishing compatibility between plants and obligate biotrophic pathogens. Curr. Opin. Plant Biol. 6, 320–326. [DOI] [PubMed] [Google Scholar]
- Panthee, D. , Marois, J. , Wright, D. , Narváez, D. , Yuan, J. and Stewart, C. (2009) Differential expression of genes in soybean in response to the causal agent of Asian soybean rust (Phakopsora pachyrhizi Sydow) is soybean growth stage‐specific. Theor. Appl. Genet. 118, 359–370. [DOI] [PubMed] [Google Scholar]
- Pham, T.A. , Miles, M.R. , Frederick, R.D. , Hill, C.B. and Hartman, G.L. (2009) Differential responses of resistant soybean entries to isolates of Phakopsora pachyrhizi . Plant Dis. 93, 224–228. [DOI] [PubMed] [Google Scholar]
- Pierozzi, P.H.B. , Ribeiro, A.S. , Moreira, J.U.V. , Laperuta, L.C. , Rachid, B.F. , Lima, W.F. , Arias, C.A.A. , Olivera, M.F. and Toledo, J.F.F. (2008) New soybean (Glycine max, Fabales, Fabaceae) sources of qualitative genetic resistance to Asian soybean rust caused by Phakopsora pachyrhizi (Uredinales, Phakopsoraceae). Genet. Mol. Biol. 31, 505–511. [Google Scholar]
- Posada‐Buitrago, M.L. and Frederick, R.D. (2005) Expressed sequence tag analysis of the soybean rust pathogen Phakopsora pachyrhizi . Fungal Genet. Biol. 42, 949–962. [DOI] [PubMed] [Google Scholar]
- Saksirirat, W. and Hoppe, H.H. (1990) Verticillium psalliotae, an effective mycoparasite of the soybean rust fungus Phakopsora pachyrhizi Syd. J. Plant Dis. Prot. 97, 622–633. [Google Scholar]
- Saksirirat, W. and Hoppe, H.H. (1991a) Degradation of uredospores of the soybean rust fungus (Phakopsora pachyrhizi Syd) by cell‐free culture filtrates of the mycoparasite Verticillium psalliotae Treschow. J. Phytopathol. 132, 33–45. [Google Scholar]
- Saksirirat, W. and Hoppe, H.H. (1991b) Teliospore germination of soybean rust fungus (Phakopsora pachyrhizi Syd). J. Phytopathol. 132, 339–342. [Google Scholar]
- Schneider, R.W. , Hollier, C.A. , Whitam, H.K. , Palm, M.E. , McKemy, J.M. , Hernández, J.R. , Levy, L. and DeVries‐Paterson, R. (2005) First report of soybean rust caused by Phakopsora pachyrhizi in the continental United States. Plant Dis. 89, 774–774. [DOI] [PubMed] [Google Scholar]
- Silva, D. , Yamanaka, N. , Brogin, R. , Arias, C. , Nepomuceno, A. , Di Mauro, A. , Pereira, S. , Nogueira, L. , Passianotto, A.L. and Abdelnoor, R. (2008) Molecular mapping of two loci that confer resistance to Asian rust in soybean. Theor. Appl. Genet. 117, 57–63. [DOI] [PubMed] [Google Scholar]
- Stokstad, E. (2004) Agriculture—plant pathologists gear up for battle with dread fungus. Science, 306, 1672–1673. [DOI] [PubMed] [Google Scholar]
- Stukenbrock, E.H. and McDonald, B.A. (2008) The origins of plant pathogens in agro‐ecosystems. Annu. Rev. Phytopathol. 46, 75–100. [DOI] [PubMed] [Google Scholar]
- Sydow, H. and Sydow, P. (1914) A contribution to knowledge of the parasitic fungi on the island of Formosa. Ann. Mycol. 12, 105. [Google Scholar]
- Tremblay, A. , Li, S. , Scheffler, B.E. and Matthews, B.F. (2009) Laser capture microdissection and expressed sequence tag analysis of uredinia formed by Phakopsora pachyrhizi, the causal agent of Asian soybean rust. Physiol. Mol. Plant Pathol. 73, 163–174. [Google Scholar]
- Vakili, N.G. and Bromfield, K.R. (1976) Phakopsora rust on soybean and other legumes in Puerto‐Rico. Plant Dis. Rep. 60, 995–999. [Google Scholar]
- Yorinori, J.T. , Paiva, W.M. , Frederick, R.D. , Costamilan, L.M. , Bertagnolli, P.F. , Hartman, G.E. , Godoy, C.V. and Nunes, J. (2005) Epidemics of soybean rust (Phakopsora pachyrhizi) in Brazil and Paraguay from 2001 to 2003. Plant Dis. 89, 675–677. [DOI] [PubMed] [Google Scholar]


