SUMMARY
Quorum sensing molecules (QSMs) are secreted by bacteria to signal population density. Upon reaching a critical concentration, QSMs induce transcriptional alterations in bacteria that enable virulence factor expression and biofilm formation. It is unclear whether mammalian hosts can recognize QSMs to trigger responsive antibacterial immunity. We report that mouse mast cell-specific G protein-coupled-receptor Mrgprb2, and its human homologue MRGPRX2, are receptors for Gram-positive QSMs, including competence-stimulating peptide (CSP)-1. CSP-1 activates Mrgprb2/MRGPRX2, triggering mast cell degranulation which inhibits bacterial growth and prevents biofilm formation. Such antibacterial functions are reduced in Mrgprb2-deficient mast cells, while wildtype mast cells fail to inhibit the growth of bacterial strains lacking CSP-1. Mrgprb2-knockout mice exhibit reduced bacterial clearance, while pharmacologically activating Mrgprb2 in vivo eliminates bacteria and improves disease score. These findings identify a host defense mechanism that uses QSMs as an “Achilles heel” and suggest MRGPRX2 as a potential therapeutic target for controlling bacterial infections.
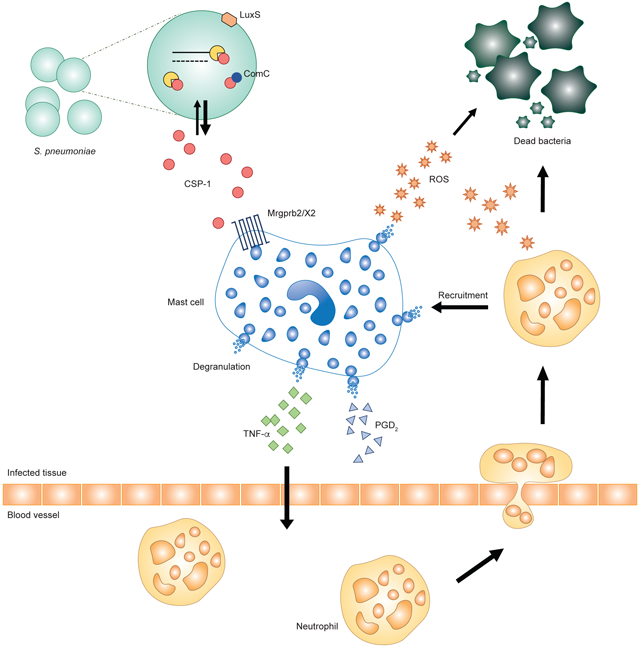
GRAPHICAL ABSTRACT
eTOC blurb
Bacteria use quorum sensing signaling for cross-species communication. Pundir et al. report that host mast cells detect Gram-positive bacteria-derived quorum sensing molecules via the Mrgpr receptors. Mrgpr activation triggers antibacterial activity and immune cell recruitment to efficiently clear bacteria, while animals deficient of Mrgpr are hypersusceptible to bacterial infection.
INTRODUCTION
Mast cells localize at areas where the host interfaces with the external environment and; therefore, are first to encounter pathogens (Marshall, 2004). Mast cells interact with pathogens through surface and intracellular receptors (Urb and Sheppard, 2012), including pattern-recognition receptors (Sandig and Bulfone-Paus, 2012), receptors for bacterial toxins (Saito et al., 1987), antimicrobial peptides (Pundir and Kulka, 2010) and complement proteins (Nilsson et al., 1996), and Fc receptors (Qiao et al., 2006). Upon stimulation, mast cells undergo degranulation, releasing preformed mediators such as histamine, proteases, and tumor necrosis factor (TNF)-α. In addition, mast cell activation induces de novo synthesis of lipid mediators and transcription and release of cytokines and chemokines (Dawicki and Marshall, 2007). These mediators have potent antimicrobial and immunomodulatory properties.
MAS-related G-protein coupled receptors (Mrgprs) are a family of GPCRs first identified in sensory neurons (Dong et al., 2001). We recently reported that mouse Mrgprb2, an ortholog of human MRGPRX2, is specifically expressed in connective tissue mast cells and is the sole receptor for numerous cationic substances including inflammatory peptides and peptidergic drugs associated with pseudoallergic reactions (McNeil et al., 2015). Considering that most pathogens secrete peptides, we hypothesized that Mrgprs can be activated by these peptides and play a role in the immune response to bacterial infections.
Quorum sensing is a cell-to-cell communication mechanism used by bacteria to orchestrate collective behaviors governing antibiotic resistance, biofilm formation, and virulence. The process is conducted by quorum sensing molecules (QSMs); therefore, QSMs present a promising target through which to control infections (Waters and Bassler, 2005). QSMs modulate immunological responses and directly impact the host (Rutherford and Bassler, 2012). If and how the host immune system detects and responds to interbacterial communication is unknown.
Here, we show that Mrgprb2 and MRGPRX2 can detect QSMs produced by Gram-positive bacteria. Upon Mrgprb2/X2 activation, mast cells degranulate and release antibacterial mediators which are pivotal to controlling bacterial infection. Removing Mrgprb2 causes significant deficits in bacterial clearance within animals while, conversely, pharmacological activation of Mrgprb2 can enhance antibacterial immunity. We have identified an innate immune mechanism that detects critical bacterial masses and initiates a timely immune response to combat the infection.
RESULTS
Mrgprb2 and MRGPRX2 are mast cell receptors for QSMs
We screened a number of bacteria-derived compounds and found that bacterial peptides with a net positive charge >2.5 activated HEK cells expressing MRGPRX2 and Mrgprb2 (Figures S1A and S1B, Tables S1 and S2). MRGPRX2- and Mrgprb2-HEK cells did not respond to weakly positive or negatively charged peptides (Table S1). MRGPRX2/b2 were activated by peptides at nano to micromolar concentrations (Table S3), a physiologically relevant range (Mashburn-Warren et al., 2012, Zhu and Lau, 2011). Interestingly, all the cationic peptides that activated Mrgprs were known QSMs derived from Gram-positive bacteria (e.g. competence stimulating peptide (CSP) from Streptococcus pneumoniae, Entf-metabolite from Enterococcus faecium, and Streptin-1 from S. pyogenes).
We determined whether QSMs activate mast cells. While a substantial percentage of wildtype (WT) peritoneal mast cells were activated by peptide QSMs, the effect was completely abolished in Mrgprb2MUT mast cells (Figure 1A and B). Peptide QSMs also induced histamine release from WT peritoneal mast cells, an effect that was abrogated in Mrgprb2MUT mast cells (Figure S1C). Furthermore, peptide QSMs evoked a dose-dependent release of β-hexosaminidase from control siRNA-treated LAD2 human mast cells (LAD2-Cntr). The effect was diminished in MRGPRX2-siRNA treated LAD2 cells (LAD2-X2kd; Figure S1D).
Figure 1. Quorum sensing peptides activate MRGPRX2 and Mrgprb2.
(A) Changes in intracellular calcium levels in peritoneal mast cells isolated from WT or Mrgprb2MUT mice in response to CSP-1 (50 μM), CSP-2 (75 μM), CSP (75 μM), Entf (100 μM), and Streptin-1 (50 μM). The black line on the graphs indicates the application of QSM. (n=3).
(B) The percentage of WT and Mrgprb2MUT peritoneal mast cells responding to QSMs. (n=3).
(C-E) Transformed binding isotherms CSP-1 to MRGPRX2 (C), Mrgprb2 (D), and Mrgprc11 (E). (n=3).
(F) Changes in ear thickness after CSP-1 or vehicle injections. (n = 660 total mast cells analyzed per group).
(G-J) Representative 3D confocal images of Av.SRho signal (white) in whole mount ears (G and I) and associated quantification of the percentage of mast cell degranulation per field of view (H and J) upon injection of vehicle (G and H) or CSP-1 (I and J). Yellow arrows point exteriorized Av.SRho+ granule structures. Scale bar, 50 μm. (n=2, 4 mice per group).
Mean ± SEM. two-tailed unpaired Student’s t-test. n.s., not significant. See also Figure S1 and Tables S1-S3.
CSP-1 was the most potent Mrgprb2/X2-activating peptide with the lowest EC50 value (Table S3) and was, therefore, used as a representative ligand for further experiments. To be certain that CSP-1 directly binds the identified receptors, we assayed thermophoresis of each receptor in the presence and absence of CSP-1. CSP-1 bound MRGPRX2 with a KD of 0.46 ± 1.7 μM (Figure 1C) and Mrgprb2 with a KD of 70.1 ± 3.2 μM (Figure 1D). CSP-1 exhibited negligible affinity for the closely related Mrgprc11 (Figure 1E).
We confirmed if CSP-1 interacts with Mrgprb2 in vivo to trigger mast cell degranulation. Injections of vehicle induced comparable swelling in WT and Mrgprb2MUT mice (Figure 1F). Injecting CSP-1 induced pronounced ear swelling in WT mice than in Mrgprb2MUT mice (Figure 1F). Mice injected with vehicle displayed minimal mast cell activation in either genotype (Figure 1G). Compared to Mrgprb2MUT mice, CSP-1 injection caused significant mast cell degranulation in WT mice (Figure 1I). Less than 22% of mast cells were degranulated at vehicle-injected sites in both genotypes (Figure 1H). CSP-1 triggered net 53% of mast cells to degranulate in WT, versus net 0% in Mrgprb2MUT mice (Figure 1J).
Mast cell activation via MRGPRX2/b2 facilitates bacterial killing
We explored whether Mrgprs activation of mast cells controls bacterial burden. Most clinical isolates of S. pneumoniae produce two different QSMs, CSP-1 and CSP-2 (Havarstein et al., 1997). We used strain D39 which produces CSP-1, the most potent Mrgprb2/X2-activating QSM. A significant suppression of the rate of D39 growth was observed at 6 h when co-cultured with LAD2 cells (Figure S2A). LAD2 reduced the viability of bacteria at multiplicities of infection (MOI) of 10 or 100 colony-forming units (CFUs) per mast cell (Figure S2B and S2C). D39, however, did not affect mast cell viability (data not shown). Quorum sensing is a cell density-dependent determinant of bacterial physiology (Davies et al., 1998); therefore, MOI 100 was chosen for subsequent experiments as it reflects a more physiologically relevant dose. LAD2-Cntr inhibited the growth of D39 by 35%. LAD2-X2kd exhibited decreased ability to suppress D39 proliferation, with up to 96% bacteria remaining viable (Figure 2A). Similarly, D39 grew more rapidly with Mrgprb2MUT peritoneal mast cells than with WT (88 vs. 50% viability, respectively; Figure 2B).
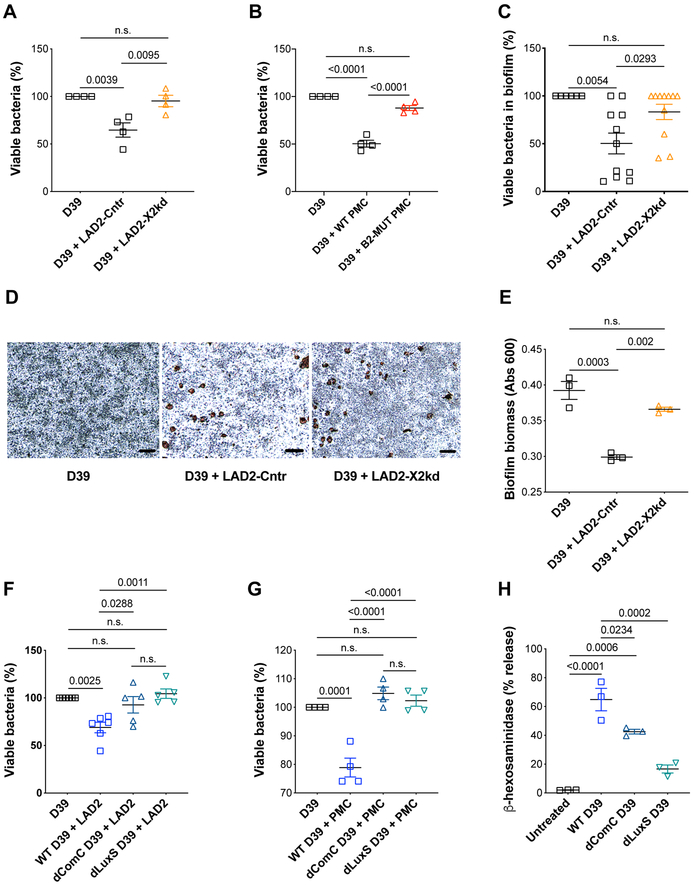
Figure 2. MRGPRX2 and Mrgprb2 restrict bacterial growth in a quorum sensing signaling-dependent manner.
(A) Viability of D39 in co-cultures with medium or LAD2 cells transfected with control (LAD2- Cntr) and MRGPRX2 siRNA (LAD2-X2kd) at MOI 100 (n=4).
(B) Viability of D39 in co-cultures with medium, WT, and Mrgprb2MUT peritoneal mast cells at MOI 100 (n=5).
(C) Viability of D39 biofilm in the presence of medium (n=6), LAD2-Cntr (n=11), and LAD2-X2kd (n=11).
(D) Representative image of D39 biofilms stained with crystal violet after incubation with medium or LAD2 cells. Mast cells were counter-stained with safranin (brown; n=3).
(E) Quantification of the total biofilm mass, as determined in Fig. 2D (n=3).
(F-G) Viability of WT, dComC, or dLuxS D39 in cultures with LAD2 (F; n=5) and WT peritoneal mast cells (G; n=4).
(H) Release of β-hexosaminidase from LAD2 cells in response to supernatants from WT, dComC, and dLuxS D39 (n=3).
Mean ± SEM. One-way ANOVA with Tukey’s multiple comparison test. Scale bar, 50 mm. n.s., not significant; PMC, peritoneal mast cells. See also Figure S2.
Inside the host, bacteria are found as biofilms which allow them to subvert immune responses and establish chronic infections (Lewis, 2007). Knowing if Mrgprs are able to disrupt biofilms would be an important finding. Pneumococci were allowed to form a biofilm on abiotic surfaces (Figure S2D). Mast cells reduced biofilm viability, with a maximum reduction observed at 6 h (Figure S2E). Biofilm viability was significantly higher in the presence of LAD2-X2kd (83%) compared with LAD2-Cntr (50%; Figure 2C).
We assessed biofilm integrity using crystal violet to stain the biomass and the extracellular matrix (Skogman et al., 2012). Biofilm alone was uniformly dark. Biofilm treated with LAD2-Cntr was more porous, with clear white spaces surrounding mast cells, indicating decreased integrity. This effect was mitigated in biofilm treated with LAD2-X2kd, as displayed by a medium intensity of staining (Figure 2D). LAD2-Cntr significantly reduced biofilm biomass compared with LAD2-X2kd (Figure 2E).
Inhibition of bacterial growth is quorum sensing dependent
In Gram-positive bacteria, the gene comC encodes the CSP-1 precursor (Cvitkovitch et al., 2003). The autoinducer-2 (AI-2) quorum sensing pathway, encoded by the luxS gene, operates universally in both Gram-positive and Gram-negative bacteria (Stroeher et al., 2003). We used D39-isogenic quorum sensing mutant strains to test whether the disruption of quorum sensing pathways would affect Mrgprs’ ability to limit bacterial growth. As expected, the viability of WT D39 was significantly reduced in cultures with human (Figure 2F) and mouse (Figure 2G) mast cells. Mast cells failed to effectively reduce the viability of dComC and dLuxS D39 (Figure 2F and G). The phenotype observed was not attributable to differences in growth rate, as each strain showed similar growth rates (Figure S2F). Correspondingly, supernatants from WT D39 induced LAD2 degranulation while supernatants from dComC and dLuxS D39 displayed an abated activity (Figure 2H). Thus, disrupting the quorum sensor machinery reduces Mrgprs’ ability to limit the bacterial growth, perhaps due to an insufficient release of antibacterial mediators following mast cell degranulation. Interestingly, only supernatants harvested from WT D39 at 6 and 8 h caused mast cell degranulation (Figure S2G). This is consistent with the observed antimicrobial activity of Mrgprs at 6 h. CSP-1 triggers competence during the exponential growth phase, suggesting that the effect is seen at 6 h due to CSP-1 accumulation to the critical levels needed to signal quorum sensing (Vidal et al., 2013).
Mrgprs elicit mediator release and recruitment of immune cells
We characterized the mechanisms underlying Mrgprs’ antibacterial activity. High levels of bactericidal reactive oxygen species (ROS) were detected in LAD2-Cntr after treatment with CSP-1 that were reduced in LAD2-X2kd (Figure S3A). Furthermore, WT D39 supernatants evoked an increase in ROS in LAD2; the effect was diminished with dComC supernatants (Figure S3B). CSP-1 induced TNF production from LAD2-Cntr, whereas the response was blunted in LAD2-X2kd (Figure S3C). LAD2-Cntr released prostaglandin D2 (PGD2; Figure S3D), but not cysteinyl leukotrienes (CysLT; Figure S3E), in response to CSP-1. The release of PGD2 was reduced from CSP-1-treated LAD2-X2kd (Figure S3D). The bactericidal activity of Mrgprs did not involve the formation of mast cell extracellular traps (Figure S3F). In addition, unlike peritoneal macrophages, peritoneal mast cells were incapable of phagocytosing D39 (Figure S3G).
To study the roles of Mrgprb2+ mast cells in the upper respiratory tract, we developed a nasopharyngeal murine model where colonization of the nasopharynx by S. pneumoniae was examined. The number of mast cells in the nasal epithelium between vehicle-treated WT and Mrgprb2MUT groups did not differ (Figure 3A and B). After infection with D39, there was a significant increase in the number of mast cells in WT mice; however, Mrgprb2MUT mice had no such increase (Figure 3A and B). This mast cell increase paralleled higher TNF levels in the nasal lavage fluid (NLF) samples from WT mice than from colonized Mrgprb2MUT animals (Figure 3C). Furthermore, WT mice infected with WT D39 had more TNF in their NLF than WT mice infected with dComC D39 (Figure 3D).
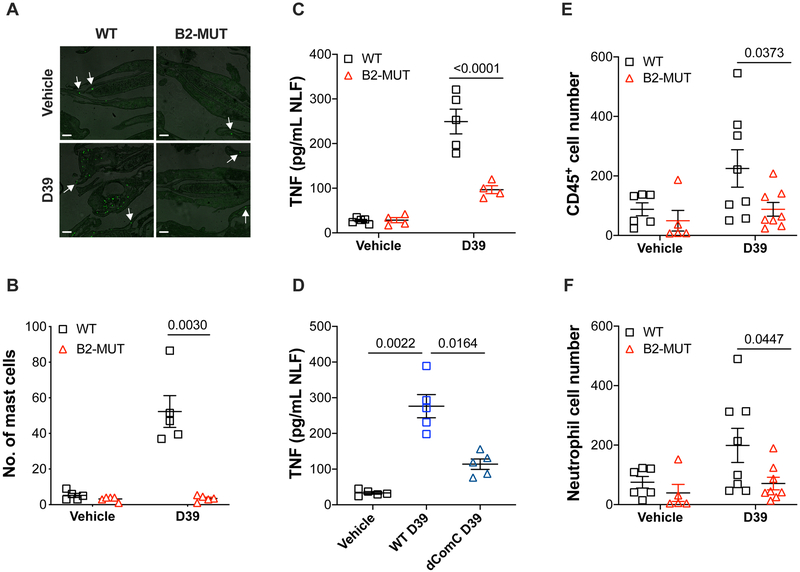
Figure 3. Mrgprb2 activation elicits the release of antibacterial mediators and promotes immune cell recruitment.
(A) Representative images showing recruitment of avidin+ mast cells (green) into the nasal epithelium of WT and Mrgprb2MUT mice after infection with vehicle or D39. Scale bar, 50 μm.
(B) Quantification of mast cells in the nasal epithelium from WT and Mrgprb2MUT mice after infection with vehicle or D39 (n=5).
(C-D) TNF levels in the NLF after nasopharyngeal colonization of (C) WT (n=5) and Mrgprb2MUT mice with WT D39 or (D) WT mice with WT and dComC D39 (n=4).
(E-F) Flow cytometric analysis of (E) CD45 for all hematopoietic cells and (F) CD11b, Ly6G, and Ly6C for neutrophils in the NLF from WT and Mrgprb2MUT mice after infection with vehicle (n=6 WT, n=5 MUT) or D39 (n=8).
Mean ± SEM. Two-way ANOVA test. See also Figure S3.
We asked if Mrgprb2 recruits other immune cells to the site of infection to further aid in the clearance of bacteria. Flow cytometric analysis (Figure S3H) demonstrated an increased number of CD45+ cells in WT NLF after infection with D39, but there were significantly less CD45+ cells in NLF from Mrgprb2MUT mice (Figure 3E). Furthermore, CD11b+Ly6G+ neutrophils infiltrated the nasopharynx in WT mice in higher numbers compared to Mrgprb2MUT mice (Figure 3F).
Mrgprb2 deficient mice are more susceptible to bacterial infections
We infected Mrgprb2MUT mice with Gram-positive bacteria to assess the role of Mrgprb2 in controlling bacterial infection in vivo. We targeted three different sites – nasopharynx, peritoneum, and skin – as they are major portals of entry for bacteria. These surfaces are enriched in connective tissue mast cells expressing Mrgprb2 (Figure 4A). In our pneumococcal nasopharyngeal colonization model, Mrgprb2MUT mice exhibited impaired bacterial clearance, as demonstrated by higher bacterial loads in their NLF compared to WT mice (Figure 4B). Consistently, infected Mrgprb2MUT mice gained less body weight than WT mice (Figure 4C). In a peritoneal infection model, Mrgprb2MUT mice displayed higher bacterial loads in peritoneal cavities after infection with vancomycin-resistant E. faecium (VRE) than WT mice (Figure 4D).
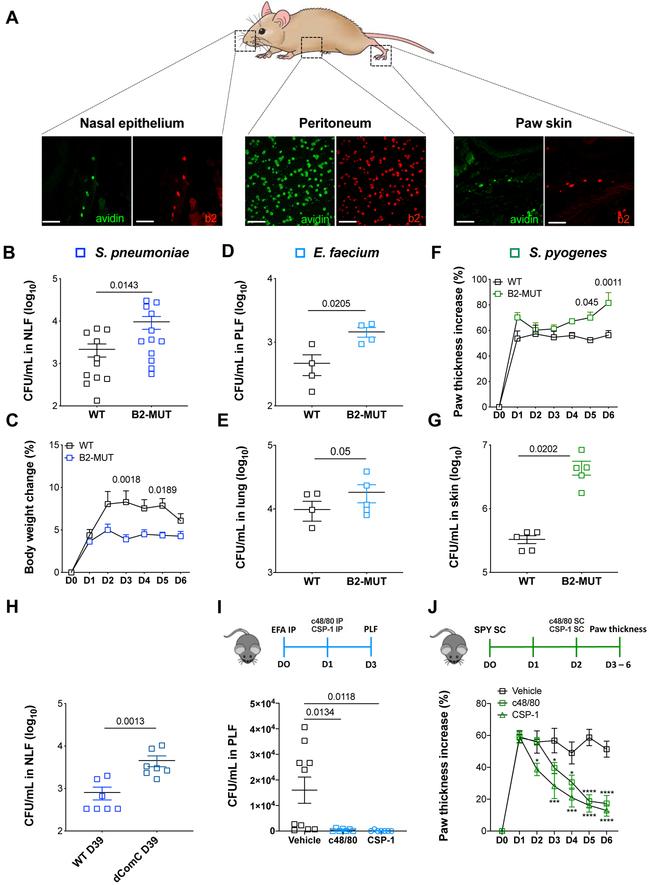
Figure 4. Mrgprb2 confers protective immunity against bacteria.
(A) Representative confocal images of sections of nasopharynx, cytospins of PLF, and sections of glabrous skin from BAC transgenic mice in which tdTomato expression is controlled by eGFP-Cre expression from the Mrgprb2 locus (red). Mast cells (green) were identified by staining for avidin (n=3). Scale bar, 50 μm.
(B) Bacterial loads in the NLF from WT and Mrgprb2MUT mice post-infection with D39 (n=12).
(C) Changes in body weights of WT and Mrgprb2MUT mice infected with D39 (n=10).
(D-E) Bacterial loads in the peritoneal cavities (D, n=4) and lung homogenates (E, n=5) from WT and Mrgprb2MUT mice infected with VRE.
(F-G) Changes in paw thickness (F) and bacterial loads in the footpads (G) of WT and Mrgprb2MUT mice infected with S. pyogenes (n=5).
(H) Bacterial clearance from the nasopharynx of WT mice infected with WT and dComC D39 (n=7).
(I) Bacterial loads in the PLF from WT mice treated with vehicle (n=9) or c48/80 and CSP-1 (n=10) after infection with VRE.
(J) Changes in paw thickness of WT mice treated with vehicle (n=9) or c48/80 and CSP-1 (n=9) after infection with S. pyogenes.
*p < 0.05, ***p < 0.001, ****p < 0.0001. Mean ± SEM. Two-tailed unpaired Student’s t-test or Two-way ANOVA test. EFA, E. faecium; PLF, peritoneal lavage fluid; SPY, S. pyogenes. See also Figure S4.
Since intraperitoneal (i.p.) colonization with VRE results in systemic infection (Leendertse et al., 2008), enterococci were also isolated from lungs. Mrgprb2MUT mice had higher bacterial loads in lung homogenates than WT mice (Figure 4E). In a dermal infection model, subcutaneous (s.c.) infection with S. pyogenes resulted in the development of footpad edema that was significantly larger and displayed a prolonged progression in Mrgprb2MUT mice compared with WT (Figure 4F). Mrgprb2MUT mice also showed impaired clearance of S. pyogenes from hind paws compared with WT mice (Figure 4G). Surprisingly, the antibacterial activity of Mrgprb2 was not limited to Gram-positive bacteria. Subcutaneous injection of Gram-negative Pseudomonas aeruginosa into the hind paws of Mrgprb2MUT mice resulted in higher edema than in WT mice (Figure S4A).
We tested whether Mrgprb2 control of bacterial infection was quorum-sensing dependent. Significantly higher bacteria were recovered in NLF from WT mice that were colonized with dComC strain. Conversely, WT mice colonized with WT D39 had lower bacteria in NLF (Figure 4H). We then performed these experiments in Mrgprb2MUT mice. As expected, mutant mice colonized with WT D39 had more bacteria in their NLF. Mutant mice infected with dComC had reduced bacteria, likely due to impaired ability of CSP-1 mutants in colonizing the nasopharynx (Figure S4B).
During nasopharyngeal carriage, pneumococci acquires genes for resistance to antibiotics such as trimethoprim (Tmp) via transformation, a process regulated by quorum sensing network (Lattar et al., 2018). We investigated whether Mrgpr binding of CSP-1 affects pneumococcal transformation frequency (tF). In the absence of mast cells, the tF of D39 was 4.65 × 10−6. When cultured with LAD2 cells, D39 displayed a statistically significant reduction in their ability to transform, with a tF of 2.17 × 10−6 (Figure S4C).
During an infection, the host produce cationic peptides (Hancock and Diamond, 2000). We used our infection models to investigate whether endogenously-produced peptides could be activating Mrgprb2, as opposed to the bacterial QSM. Subcutaneous injection of supernatants from WT D39 into WT mice hind paws led to an upregulation in the transcription of mouse defensin (Def)b3 and Defb4. Injecting supernatant from the dComC D39 induced comparable expression of Defb3 and Defb4 (Figure S4D and S4E). It is likely defensin upregulation occurred via activation of toll-like receptors in the epithelial cells. In the pneumococcal model, nasopharyngeal colonization with WT and dComC D39 failed to induce any changes in the expression of Defb1, Defb2, Defb3, Defb4, Defb14, and cathelicidin Camp in the nasal epithelium (Figure S4F). These results suggest a direct interaction between QSMs and Mrgprb2.
Pharmacological activation of Mrgprb2 sufficiently eliminates bacteria and reduces disease score
We asked whether direct stimulation of Mrgpr could induce protective immunity against bacteria. We injected WT mice with either compound 48/80, an Mrgprb2-specific agonist (McNeil et al., 2015), or CSP-1. The compounds were injected locally at the site of bacterial infection to specifically activate Mrgprb2-expressing mast cells. Mice that received 48/80 and CSP-1 i.p. after infection with VRE displayed strikingly lower bacterial counts in their peritoneal lavages compared with saline-treated animals (Figure 4I). Similarly, s.c. injection of 48/80 and CSP-1 into the hind paws colonized with S. pyogenes resulted in a remarkable reduction in footpad edema, with paw size returning to a normal thickness within 96 h post-treatment (Figure 4J).
DISCUSSION
In rodents, there are at least two subsets of mast cell population: the connective tissue type (CTMCs) and the mucosal type (MMCs) (Bienenstock et al., 1982, Enerback, 1966). While CTMCs are present in connective tissues, MMCs are prominent within the gastrointestinal mucosa, where they are key modulators of barrier function and homeostasis (Kurashima and Kiyono, 2014). CTMCs respond well to basic secretagogues, whereas MMCs respond poorly, likely due to their lack of expression of Mrgprs (Seifert, 2015). We demonstrate that CTMC-expressing Mrgprb2/X2 recognize and are activated by bacterial cationic QSMs. Upon exposure to CSP-1 or S. pneumoniae, mast cells degranulate and release ROS, TNF-α, and PGD2 in Mrgprb2/X2, as well as quorum sensing signaling, dependent manner. Mast cell-derived cytokines and lipid mediators contribute to the recruitment of immune cells, such as neutrophils, and clearance of infections (Galli et al., 2005).
In the current study, we focused on select peptides produced by human pathogen Gram-positive Streptococcaceae but it is likely that the list of cationic QSMs that activate Mrgprs is greater than explored here. Mrgpr activation may affect disease outcome with other Grampositive bacteria such as Staphylococcaceae, as they also exhibit quorum sensing and are able to form biofilms (Yarwood and Schlievert, 2003). Mrgprb2 also elicited an antimicrobial activity towards P. aeruginosa. Even though Gram-negative bacteria do not use peptide signals for quorum sensing, there could be unknown downstream cationic products released during quorum sensing signaling that could potentially activate Mrgprb2. There could also be peptides, bacteria- or host-derived, released in response to an infection with P. aeruginosa independent of quorum sensing networks that could lead to the phenotype observed (Kariminik et al., 2017)
To date, studies of the bacterial quorum sensing systems have focused on Gram-negative bacteria, their effect on secondary immune targets, and treatment strategies based on the manipulation of bacterial genetics. P. aeruginosa is one of the best characterized examples. P. aeruginosa quorum sensing network plays a dual role in the modulation of the immune system. P. aeruginosa QSMs are able to either stimulate or suppress immune responses against the bacteria. (Pritchard, 2006, Kariminik et al., 2017). The responsiveness of immune cells, epithelial cells, and tissues to QSMs has been traditionally measured as changes in intracellular calcium, cyclic AMP, nitric oxide, cytokine/chemokine levels in heterologous cell systems and primary cell lines (Freund et al., 2018, Verbeurgt et al., 2017). The effect of such interactions between ligand and secondary immune targets on defense against pathogens has not been explored. Here, we reveal that mast cell receptors Mrgprb2 and MRGPRX2 directly bind peptide QSM. This interaction provides an insight into the fundamental mechanism by which the innate immune system can detect Gram-positive bacteria.
The finding that dComC bacteria deficient in CSP-1 cannot be recognized and killed by mast cells in vitro, or effectively cleared by host in vivo, further reinforces that Mrgprs’ recognition of the quorum sensing network is key to a successful immune response. Results from dLuxS strain were an interesting finding as AI-2 is not a peptide. It did not activate Ca2+ mobilization in MRGPRX2-HEK and Mrgprb2-HEK cells (Table S1) or β-hexosaminidase release from LAD2 cells (Figure S2H). As the transformation is driven by CSP (the product of comC) in Streptococci, reduced transformation efficiency in luxS mutants suggests that LuxS/AI-2 might regulate the competence pathway upstream (and, therefore, modulate the production of CSP) (Vidal et al., 2013).
We show that in the presence of Mrgprs, pneumococci transform at a lower frequency as the quorum sensing signal is sequestered towards mast cell function. Our data may have implications for pneumococcal evolution and acquisition of antimicrobial resistance in the upper airways but also it appears to have a profound effect on pneumococcal adaptation occurring during colonization of nasopharynx where pneumococci encounter mast cells. As Com is required for virulence during nasopharyngeal colonization (Lattar et al., 2018), scavenging CSP by its eukaryotic receptor not just triggers an immune response but also impairs Com-activated virulence. Overall, our research has uncovered MRGPRX2 as a specific therapeutic target that can be used to harness the immune system to combat bacterial infections.
STAR METHODS
CONTACT FOR REAGENT AND RESOURCE SHARING
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, Xinzhong Dong (xdong2@jhmi.edu).
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Mice
C57Bl/6J, genetically Mrgprb2-deficient (Mrgprb2MUT, B2-MUT), and Mrgprb2-Cre mice were bred under specific pathogen-free conditions in the Johns Hopkins University School of Medicine Animal Facility and weaned at 3-3.5 weeks of age. Mrgprb2-Cre; Rosa26-lsl-tdTomato BAC transgenic reporter mice were obtained by crossing Mrgprb2-Cre animals with Rosa26- lLoxP-STOP-LoxP (lsl)-tdTomato animals (Jackson labs). Mice were kept in community cages (4-5 mice per cage) at light periods of 12 h and fed water and mouse chow ad libitum. Male and female mice were used for experiments at 8-10 weeks of age. All animal procedures were conducted in accordance with institutional guidelines and with approval from the Animal Care and Use Committee at the Johns Hopkins University School of Medicine.
Cell lines
Laboratory of Allergic Disease (LAD2) mast cells derived from male human mast cell leukemia (Kirshenbaum et al., 2003) were obtained from Drs. A. S. Kirshenbaum and D. Metcalfe (National Institutes of Health). LAD2 cell line has been shown to express MRGPRX2 endogenously (Subramanian et al., 2013). Cells were cultured in StemPro-34 medium (Life Technologies) supplemented with 2 mM L-glutamine, 100 /mL penicillin, 50 μg/mL streptomycin and 100 ng/mL recombinant human stem cell factor (SCF; Peprotech), maintained at 37°C and 5% CO2, and split by hemidepletion every 7 days. Female human embryonic kidney (HEK)293 cells were maintained in DMEM supplemented with 10% fetal bovine serum (FBS), penicillin-streptomycin and L-glutamine at 37°C and 5% CO2. All cell lines were tested to be negative for mycoplasma. Cell lines were authenticated by flow cytometry for cell surface markers, or obtained from authentic stocks (ATCC).
Bacterial Strains
Streptococcus pneumoniae strains D39 WT (Avery strain, clinical isolate capsular serotype 2), dComC (D39-derivative comC null mutant), dLuxS (D39-dervative luxS null mutant) (Vidal et al., 2013), and SPJV29 Tmpr (Lattar et al., 2018) were previously prepared. The Com or LuxS quorum sensor machinery was disrupted by knocking out the genes encoding for the secreted molecule that activates quorum sensing signaling, either comC (for CSP; dComC D39) or luxS (for AI-2; dLuxS D39), respectively (Vidal et al., 2013). Enterococcus faecium strain VRE and Streptococcus pyogenes strain SF370; M1 GAS were procured from ATCC. Stock cultures were maintained in 20% glycerol at −80°C. All strains were cultured on tryptic soy agar with 5% sheep blood (TSA). Pseudomonas aeruginosa strain 19660 (Shao et al., 2012) was cultured on cetrimide agar plates.
METHOD DETAILS
Experimental design
No sample size calculation was performed. Sample size was determined based on previous experiments and significance levels determined as described in the individual Figure legends.
No data were excluded. All attempts at replication were successful. Samples were allocated into experimental groups at the start of each individual experiments. In case of animals, genetically identical animals were age and sex-matched and allocated to experimental groups at the start of the experiments. Investigators were blind to group allocation (both genotype and treatment) during in vivo data collection and assessment.
Molecules and preparation
Various bacteria-derived molecules and peptides utilized in this study are listed in Table S1. Sequences of quorum sensing peptides are listed in Table S2. Compounds were dissolved either in saline or DMSO and stored at −80°C before thawing on ice and diluting in appropriate assay buffer.
Mouse genotyping
Primers and annealing temperatures were as follows:
Wildtype mice – ggttcctgggcatccgtat, cttccgcctgaaccttcggt, 64.5°C
Mrgprb2MUT – gttcctgggcatccgcac, cttccgcctgaaccttcggt, 61.8°C
Mrgprb2-Cre – tatatcatggccgacaagca, cagaccgcgcgcctgaaga, 62°C
tdTomato – aagggagctgcagtggagta, ccgaaaatctgtgggaagtc, ggcattaaagcagcgtatcc, ctgttcctgtacggcatgg, 61°C.
Murine peritoneal mast cell isolation and calcium imaging
Adult male or female animals were euthanized by CO2 asphyxiation and peritoneal mast cells were collected through a lavage of the abdomen, as previously described (McNeil et al., 2015). Briefly, 2 × 6 mL of ice-cold mast cell dissociation medium (MCDM; HBSS with 3% FBS and 10 mM HEPES, pH 7.2) was injected into the peritoneal cavity and the abdomen was massaged for 60 s. The medium with cell suspension was aspirated and centrifuged at 200 g for 5 min at RT. The pellets were resuspended in 2 mL MCDM, layered over 4 mL of an isotonic 70% Percoll solution, and centrifuged at 500 g for 20 min at 4°C. Mast cells recovered in the pellet were resuspended in DMEM with 10% FBS, 100 U/mL penicillin, 50 μg/ml streptomycin and 25 ng/ml recombinant mouse stem cell factor (mSCF; Peprotech), and plated onto glass coverslips coated with 30 μg/ml fibronectin (Sigma). After 2 h of incubation at 37°C and 5% CO 2, mast cells were loaded with 1:2000 Fluo-4, acetomethoxy ester (Molecular Probes) along with 0.02% Pluronic F-127 (1:1000; Molecular Probes) in calcium imaging buffer (CIB: NaCl 125 mM, KCl 3 mM, CaCl2 2.5 mM, MgCl2 0.6 mM, HEPES 10 mM, glucose 20 mM, NaHCO3 1.2 mM, sucrose 20 mM, pH 7.4) for 30 min at 37°C, followed by wash ing once in CIB. Fluorescence measurements were performed at 488 nm excitation. QSMs were tested at a concentration approximately 2X the EC50 as calculated by the values reported in Table S3.
Calcium imaging with HEK293 cells
Calcium imaging was performed as previously described (McNeil et al., 2015). Briefly, HEK293 cells stably expressing MRGPRX2 or Mrgprb2 receptors along with the Gα15 protein were cultured on sterile 50 μg/ml poly-D-lysine coated glass coverslips overnight. Next day, cells were loaded with 1:1000 fura-2, acetomethoxy ester (Molecular Probes) along with 1:1000 pluronic acid for 30 min in the dark at 37°C and 5% CO2. After washing twice with CIB, cells were imaged during 340- and 380-nm excitation to detect intracellular free calcium.
EC50 determination
HEK293 cells stably expressing Gα15 and either MRGPRX2 or Mrgprb2 were plated at 4 × 104 cells per well in 96-well plates and incubated overnight. The next day, media was removed and replaced with imaging solution from the FLIPR Calcium 5 assay kit (Molecular Devices), diluted according to manufacturer's suggestions in Hank's Balanced Salt Solution (HBSS) with 20 mM HEPES, pH 7.4. Cells were incubated at 37°C f or 60 min, and allowed to recover for 15 min at RT before imaging in a Flexstation 3 (Molecular Devices). Wells were imaged according to manufacturer's specifications for 120 s, with 50 μl of QSM at 3X concentration added 30 s after the imaging began. Responses were determined by subtracting the minimum signal from the maximum signal. QSMs were tested in duplicate wells, the signals were averaged, and EC50s were determined for each trial by normalizing to the peak response to the substance in that trial. All peptides were dissolved in HBSS+HEPES solution.
Histamine release assay
Peritoneal mast cells were isolated as described and after 2 h of recovery in DMEM + mSCF, resuspended in CIB and counted. Cells were seeded on to 20 μg/ml fibronectin-coated 96-well plates for 45 min at 37°C and 5% CO 2 in 75μL volume at 1000 cells/well. For the assay, all compounds were dissolved in CIB and then added at 2X the test concentration (final volume 150 μL). After 15 min, 100 μl of supernatant was harvested and frozen at −80°C until assay. Histamine levels were determined with a histamine assay kit (Enzo) according to the manufacturer’s instructions.
siRNA transfection
ON-TARGET plus MRGPRX2 SMARTpool siRNA and control siRNA (GE Dharmacon) were transfected into LAD2 cells with Lipofectamine 3000 (Life Technologies) following the manufacturer’s instructions. Briefly, a total of 1 × 106 cells in antibiotic-free medium were plated in 6-well plates and transfected with 100 nM siRNA. At 48 h, knockdown was confirmed by qPCR. Knockdown efficiency was 86.41 ± 4.84%.
Measurement of histamine release from human mast cells
LAD2 cells were washed, resuspended in BSA-free HEPES at 0.1 × 106 per well, and incubated with QSMs for 30 min at 37°C and 5% CO 2. A histamine (Sigma-Aldrich) stock solution of 100 μg/ml was prepared and stored at −20°C. The working standards of 4000 to 7.8 ng/ml were freshly prepared using 2-fold serial dilution. O-phthalaldehyde (OPT; Sigma-Aldrich) was dissolved in acetone-free methanol (10 mg/ml) and kept in dark at 4°C. Histamine standards and cell-free supernatants (60 μl were transferred to a black 96-well flat-bottom microplate and mixed with 12 μl 1M NaOH and 3 μl OPT. After 4 min at room temperature, 6 μl 3M HCl was added to stop the histamine-OPT reaction. Fluorescence intensity was measured at 460 nm (355 nm excitation).
Mast cell degranulation assay
LAD2 cells washed, resuspended in 0.4% BSA-HEPES buffer at 0.5 × 106 cells/mL, and treated with compounds for 30 min at 37°C and 5% CO 2. β-hexosaminidase in the supernatants and cell lysates solubilized with 0.1% Triton-X-100 were quantified by hydrolysis of p-nitrophenyl N-acetyl-β-D-glucosamide (pNAG; Sigma-Aldrich) in 0.1 M sodium citrate buffer (pH 4.5) for 90 min at 37°C. The percentage of β-hexosaminidase release was calculated as a percent of total content.
Generation of cells stably expressing GFP-tagged Mrgprs
Mrgprb2 and MRGPRX2 were inserted into pEGFP-N1 by restriction cloning and transfected into HEK293 cells using Lipofectamine 3000 as per the manufacturer’s instructions. After 48 h, cells were then selected using 0.5 mg/mL G418 for three weeks, after which GFP-positive cells were sorted by fluorescence-activated cell sorting (FACS) and monoclonally expanded. Two lines expressing similar levels of Mrgprb2 and MRGPRX2, as measured by GFP fluorescence, were selected for the study.
Microscale thermophoresis binding
Binding isotherms for Mrgprb2, MRGPRX2, and Mrgrpc11 towards CSP-1 was determined by microscale thermophoresis with the NanoTemper monolith NT.115 instrument. Thermophoresis of a molecule is affected by physical parameters such as size, charge, and solvation. By extension, the thermophoresis of one molecule is altered when it interacts with another; and therefore, it can be used to measure interactions between molecules (Duhr and Braun, 2006). Receptors were crudely purified as a membrane fraction from cells stably independently expressing GFP-tagged MRGPRX2, Mrgprb2, or Mrgprc11 (Vasavda et al., 2017). Lyophilized CSP-1 was dissolved in binding buffer and subsequently incubated with each receptor for 5 minutes at room temperature in binding buffer. Mrgprb2 was assayed in a buffer of 20 mM HEPES, 125 mM NaCl, 2 mM MgCl2, 4.5 mM KCl, and 2 mM CaCl2 at pH 7.4. MRGPRX2 was assayed in a buffer of 10 mM Na2HPO4, 1.8 mM KH2PO4, 137 mM NaCl, and 2.7 mM KCl at pH 7.4. Microscale thermophoretic experiments were executed using 20% LED power and 40% MST power. KDs were calculated using the law of mass action by evaluating the change in normalized fluorescence from three independent experiments. Samples with large changes in initial fluorescence (> +/− 10% initial capillary fluorescence) were omitted from analysis.
Ear pinnae mast cell staining and ear thickness
Eight μg of avidin-sulfhorhodamine 101 (Av.SRho) in 20 μl of PBS were injected intradermally (i.d.) into both ear pinnae of WT and Mrgprb2MUT mice, as previously described (Reber et al., 2017). Seven days later, ear thickness was measured and left ear pinnae were injected i.d. with 50 μM of CSP-1 in a final volume of 20 μl and the right ear pinnae with 20 μl of vehicle control. Ears thickness was measured again 45 min later and mice were euthanized before ear excision. Whole ears were fixed in 4% PFA overnight, then mounted on microscopy slides and placed under a LSM710 Meta inverted confocal laser-scanning microscope. High resolution Z-stack images (1024 × 1024 pixels) of Av.SRho fluorescent signal were acquired using a 20x objective in each whole mount ear pinna. Images were then processed using Image J software.
Preparation of bacterial inoculum
An overnight agar culture was used to prepare a cell suspension in Todd-Hewitt broth containing 0.5% (w/v) yeast extract (THY; for S. pneumoniae and S. pyogenes) or BHI broth (for E. faecium) to an OD600 of 0.1. This suspension was placed in an Anaeropack™-CO2 generator jar (Thermo Fisher Scientific) and incubated in a 37°C incubator until the culture reached an OD600 of 0.45 (mid-log phase). E. faecium was propagated in a shaking incubator at 37°C in atmospheric conditions (i.e. CO2 levels were not adjusted) to achieve an OD600 of 1.0. Bacteria were harvested, centrifuged for 1 min at 14,000 g, washed with sterile PBS, and adjusted to the required concentrations.
Bacterial supernatant collection
Bacterial strains were inoculated overnight on TSA plates at 37°C and 5% CO2. Next day, 25 mL of THY was inoculated with bacteria and OD600 adjusted between 0.08 – 0.1. The bacterial suspensions were incubated at 37°C and 5% CO2 for 2, 6, 4, and 8 h. At each time points, samples were collected and centrifuged at 4000 rpm for 10 min. The supernatants were filtered through a 0.45 μm filter, aliquoted, and immediately stored at −80°C. Samples were thawed on ice prior to use.
Bacterial killing
Overnight cultures of D39 WT, dComC or dLuxS were used to inoculate 5 mL of THY at OD600 = 0.1 and cultured at 37°C and 5% CO2. until the OD600 reached 0.45. Peritoneal mast cells were isolated as described. Instead of plating on cover slips, cells were allowed to recover for 2 h in antibiotic-free medium. After determining cell counts, the densities of LAD2 and peritoneal mast cells were adjusted at 104 cells/0.9 mL and the cell suspensions were seeded in 24-well plates (900 μL/well). After reaching an OD600 = 0.45, 100 μL of bacterial suspension was spun at 14,000 g for 1 min. The pellet was resuspended in 1 mL sterile PBS, and 100 μL (MOI 100) was added to the wells containing cells. The plate was then incubated at 37°C and 5% CO2 for 6 h, after which 100 μL of culture medium was removed for serial dilution and CFU enumeration by agar plating.
Biofilm formation
Briefly, bacteria were diluted from overnight culture (1:100) in THY and 0.1 ml of the suspension was added to each well of a 96-well polystyrene treated plate (O'Toole, 2011). Plates were incubated for 2 h at 37°C in 5% CO 2. Planktonic cells were removed and washed once with PBS before mast cells were added into each well at a MOI of 100. Biofilms were grown 37°C in 5% CO2 until the indicated time points. Planktonic cells were removed and the biofilms were washed with PBS before being resuspended in THY for colony-forming units (CFUs) enumeration by agar plating. The percentage of viable cells was calculated by normalizing the CFUs of the treated biofilms to the untreated, medium only control at the same time point, which contains 100% viable cells.
Determination of biofilm mass
To stain biofilms, planktonic cells were first removed. Crystal violet (1%) was added to sample and stained for 60 seconds and then washed three times with PBS. To develop a counterstain for the mast cells, the samples were then stained with iodine for 60 seconds, ethanol (95%) for 5 seconds, and safranin (1%) for 60 seconds, subsequently in the specific order, where washes (3X) with PBS were performed after each staining agent. To quantify the amount of biofilm biomass stained with crystal violet, 100% ethanol was used to solubilize the biofilm. After 15 minutes, absorbance (OD600) was read on the dissolved biofilms.
EIA
LAD2 cells were treated with 10 mM CSP-1 for 4 h at 37°C and 5% CO 2. Cell-free supernatants were collected and assayed using human TNF ELISA set (BD Biosciences), Cysteinyl Leukotriene ELISA kit and Prostaglandin D2 ELISA kit (Cayman Chemicals). For in vivo TNF assays, WT and Mrgprb2MUT animals were colonized with S. pneumoniae and NLF was collected after 6 h, as described. TNF levels were determined in cell-free NLF using mouse TNF-alpha Quantikine ELISA kit (R&D Systems).
ROS generation assay
LAD2 cells were treated with 10 μM CSP-1 or 20% v/v supernatants from WT, dComC, and dLuxS D39 strains for 4 h at 37°C and 5% CO 2 and assayed using DCFDA cellular ROS detection assay kit (Abcam).
Murine model of pneumococcal colonization
Unanesthetized male mice were colonized intranasally with 10 μL of S. pneumoniae suspension containing ~107 CFUs. The inoculum was plated immediately on TSA plates to determine viable counts. At the time indicated, mice were euthanized by CO2 asphyxiation, the trachea was exposed and a small incision was made. A 26-gauge, 1 mL insulin syringe connected to a 4-cm PE-20 polyethylene tube was inserted into the trachea, tied off using a silk suture, and 1 mL sterile PBS was instilled. Lavage fluid (NLF) was collected from the nares, serially diluted in PBS and plated onto TSA plates. Colonies were counted after overnight incubation at 37°C.
Murine model of peritonitis
Female mice were injected intraperitoneally with ~108 CFUs of E. faecium in 100 μL saline. The inoculum was plated immediately on TSA plates to determine viable counts. At the times indicated, mice were euthanized by CO2 asphyxiation, a peritoneal lavage was performed with 5 mL sterile PBS using a 26-guage needle and peritoneal lavage fluid (PLF) was collected in 15 mL polypropylene tubes. After the collection of PLF, the abdomen was opened, the lungs were harvested and homogenized in 1 mL sterile PBS. CFUs were determined from serial dilutions of PLF and lung homogenates by plating onto TSA plates and overnight incubation at 37°C.
Murine model of subcutaneous infection
The right hind footpads of male mice were subcutaneously infected with 5 μL of S. pyogenes or P. aeruginosa suspension containing ~107 and ~106 CFUs, respectively. The inoculum was plated immediately on TSA plates to determine viable counts. The left hind footpad received 5 μL sterile PBS. After infection, the thickness of paws was measured using a digital Vernier calipers every 24 h. At the time indicated, mice were euthanized by CO2 asphyxiation. Footpad skin was depilated with Nair and harvested using a 6-mm biopsy punch. Skin was homogenized in 1 mL sterile PBS and serial dilutions were plated onto TSA plates. After overnight incubation at 37°C, the CFUs were counted.
Transformation reactions and calculation of transformation frequency
S. pneumoniae D39 WT strain was made competent using standard procedures and then transformed with 500 ng of pure DNA containing 100 ng CSP-1 in a reaction volume of 200 μL (Havarstein et al., 1995, Lattar et al., 2018). The density of parent strain was counted on TSA plates supplemented with 3% sheep blood and trimethoprim (Tmp, 10 μg/mL). The transformation frequency (tF) was calculated as number of transformants relative to the total pneumococci recovered in the transformation reaction.
Avidin staining of mast cells
Animals were anesthetized with chloral hydrate and transcardially perfused with PBS and ice-cold 4% paraformaldehyde (PFA). The whole nasal cavity with attached cribiform plate and footpad skin sections were dissected. Nasal tissue was decalcified in Shandon™ TBD-2™ (Fisher Scientific) on ice for 5 h with gentle rotation. After being washed in PBS, tissues were equilibrated sequentially in 15% and 30% sucrose, embedded in optimum cutting temperature compound, and sectioned (20 μm width) with a cryostat. Frozen nasal sections were processed from cribiform plate side. The sections on slides were dried at 37°C for 1 h, and fixed with 4% PFA at RT for 10 min. The slides were pre-incubated in blocking buffer (10% normal goat serum, 0.2% Triton-X-100 in PBS, pH 7.4) for 1 h at RT, then incubated with 1:250 Avidin-FITC (Sigma-Aldrich) for 1 h. Slides were then washed five times with PBS and mounted with Fluoromount G mounting media (eBioscience).
Peritoneal mast cells were isolated as described except that plating on fibronectin-coated coverslips, cells were allowed to recover for 2 h in DMEM at 37°C and 5% CO2. Cells were then spun at 1000 rpm for 5 min at 4°C on a Cytospin (Thermo Scientific), fixed with 4% PFA at RT for 10 min, stained with 1:1000 Avidin-FITC for 30 min at RT, and washed with PBS before immediate imaging.
Real-time qPCR
Total RNA was extracted from skin and airway tissues using Direct-zol RNA kit (Zymo Research). Reverse transcription was carried out using iScript cDNA synthesis kit (BioRad). Gene expression was analyzed by real-time qPCR on a StepOnePlus system (Applied Biosystems). For each qPCR assay, a total of 4 ng of cDNA was used, and all reactions were performed in duplicated for 40 cycles as per the manufacturer’s recommendation. Taqman gene expression primers and gene expression master mix were used. All data were normalized to Hprt internal control and are reported as ratio of Hprt expression.
In vivo phagocytosis assay
S. pneumoniae was grown to 0.5 OD600 in 5 mL THY culture at 37°C, pelleted by centrifugation at 4000 g for 5 min, and resuspended in 2 mL of 10 μM CFSE in PBS. After 1 h of incubation at RT in dark, the culture was centrifuged and the pellet was washed three times with 15 mL PBS. After final wash, the pellet was fully resuspended with 5 mL PBS and OD600 was adjusted to 0.45. Mice were injected i.p. with 100 μL (1 × 106 CFU) of S. pneumoniae. After 6 h, mice were sacrificed and PLF was collected as described. Samples were then processed for flow cytometry.
Quantification of mast cell extracellular trap DNA (MCETs)
LAD2 cells were plated in a black 96-well plate at a density of 1 × 105 cells per well, and activated with 100 nM PMA, 10 μM CSP-1 or 1% Triton-X-100 (to determine total DNA content). After 2 hours, 2 U DNase was added to appropriate wells. After a further 45 min, 5 μM Sytox Green dye was added to each well and fluorescence was measured 15 min later at emission/excitation = 504/523 nm. The amount of extracellular DNA was calculated as:
Flow cytometry
Peritoneal and nasal lavage fluid samples were collected from animals and spun at 200 g for 5 min at 4°C. The cell pellets were resuspended in 1 mL PBS and stained with 1 μL Live/Dead Fixable Aqua Dead cell stain for 30 min on ice. The samples were centrifuged at 200 g for 5 min at 4°C, resuspended in 100 μL ice-cold FACs buffer (1X PBS-2% FBS) and then Fc blocked for 10 min on ice. The cells were stained for 30 min with indicated antibodies, spun at 200 g for 5 min at 4°C and washed twice with 1 mL ice-cold FACs buffer. Final cell pellets were suspended in 300 μL FACs buffer, run on cytoFLEX (Beckman Coulter) and analyzed using FlowJo v.10 software. The following antibodies from Biolegend were used in flow analysis: rat anti-mouse CD45 (clone 30-F11) FITC, rat anti-mouse CD11-b (M1/70) PE/Dazzle, rat anti-mouse Ly6G (1A8) BV421, rat anti-mouse Ly6C (HK1.4) AF647 and APC, rat anti-mouse CD117 (ACK2) PerCP/Cy5.5, hamster anti-mouse FcεRI (MAR-1) PE/Cy7, rat anti-mouse F4/80 (BM8) PE, and hamster anti-mouse CD11c (N418) BV605. The cells were initially gated for single cells based on forward and side scatter (FSC-A/SSC-A) followed by two double exclusion gates (SSC-A/SSC-H and FSC-H/FSC-W). Then, dead cells were excluded based on their positivity for Live/Dead Fixable Aqua Dead cell staining. Live cells were then gated for CD45+ cells. Mast cells in peritoneal lavage fluid were identified as CD117+FcεRI+. Peritoneal macrophages were identified as F4/80+Ly6C−. Neutrophils in nasal lavage fluid were identified as CD11b+Ly6G+Ly6C−.
QUANTIFICATION AND STATISTICAL ANALYSIS
Data are presented as mean ± standard error of mean (SEM), and were analyzed by 1) two-tailed unpaired Student’s t-test, 2) one-way ANOVA with Tukey’s multiple comparison test, and 3) two-way ANOVA with Sidak’s multiple comparison test, using Prism v7.0d (GraphPad). Statistical significance was defined as P<0.05. All the figures have data that are representative of at least two independent in vivo experiments, or at least 3 independent in vitro experiments.
DATA AVAILABILITY
All data generated or analyzed during this study are included in this article.
Supplementary Material
KEY RESOURCES TABLE
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| Rat anti-mouse CD45 (clone 30-F11) | BioLegend | Cat # 103108, RRID:AB_312973 |
| Rat anti-mouse/human CD11b (clone M1/70) | BioLegend | Cat# 101256, RRID:AB_2563648 |
| Rat anti-mouse Ly-6G (clone 1A8) | BioLegend | Cat# 127628, RRID:AB_2562567 |
| Rat anti-mouse Ly-6C (clone HK1.4) | BioLegend | Cat# 128010, RRID:AB_1236550 |
| Rat anti-mouse Ly-6C (clone HK1.4) | BioLegend | Cat# 128016, RRID:AB_1732076 |
| Rat anti-mouse CD117 (c-kit) (clone ACK2) | BioLegend | Cat# 135134, RRID:AB_2629635 |
| Hamster anti-mouse FcεRIa(clone MAR-1) | BioLegend | Cat# 134318, RRID:AB_10640122 |
| Rat anti-mouse F4/80 (clone BM8) | BioLegend | Cat# 123110, RRID:AB_893486 |
| Hamster anti-mouse CD11c (clone N418) | BioLegend | Cat#117334, RRID:AB_2562415 |
| Avidin-FITC | Sigma-Aldrich | Cat# A2901 |
| Bacterial and Virus Strains | ||
| Streptococcus pneumoniae: Clinical isolate capsular serotype 2: Avery strain D39 | (Vidal et al., 2013) | N/A |
| Streptococcus pneumoniae: dComC (D39-derivative comC null mutant) | (Vidal et al., 2013) | N/A |
| Streptococcus pneumoniae: dLuxS (D39-derivative luxSnull mutant) | Vidal et al., 2013) | N/A |
| Streptococcus pneumoniae: SPJV29 D39 Tmpr (Trimethoprim resistant) | (Lattar et al., 2018) | N/A |
| Enterococcus faecium: strain VRE | ATCC | Cat# 700221 |
| Streptococcus pyogenes: M1 serotype strain: strain SF370; M1 GAS | ATCC | Cat# 700294 |
| Pseudomonas aeruginosa: Strain 19660 | (Shao et al., 2012) | N/A |
| Chemicals, Peptides, and Recombinant Proteins | ||
| CSP-1 | AnaSpec | Cat# AS-63779 |
| CSP-2 | AnaSpec | Cat# AS-63877 |
| Delta toxin (1-26) | AnaSpec | Cat# AS-62496 |
| Listeriolysin; LLO (91-99) | AnaSpec | Cat# AS-64870 |
| Staphylococcal enterotoxin B domain (144-153) | AnaSpec | Cat# AS-63835 |
| CFP10 (71-85) | AnaSpec | Cat# AS-61689 |
| C2-HSL | Cayman Chemicals | Cat# 23241 |
| 3-oxo-C8-HSL | Cayman Chemicals | Cat# 10011206 |
| Gallidermin | Enzo Life Sciences | Cat# ALX-380-072 |
| PQS | Sigma-Aldrich | Cat# 94398 |
| HHQ | Sigma-Aldrich | Cat# SML0747 |
| DHQ | Sigma-Aldrich | Cat# 144789 |
| Pyocyanin | Sigma-Aldrich | Cat# P0046 |
| Nisin | Sigma-Aldrich | Cat# N5764 |
| BIP | EZBiolab | Custom synthesis |
| BIP2 | EZBiolab | Custom synthesis |
| CSFphrC1 | EZBiolab | Custom synthesis |
| PaEDF3 | EZBiolab | Custom synthesis |
| CSP | GenScript | Custom synthesis |
| SilCR | GenScript | Custom synthesis |
| Streptin-1 | GenScript | Custom synthesis |
| Entf | GenScript | Custom synthesis |
| Autoinducer 2 | OMM Scientific | N/A |
| StemPro-34 SFM (1x) | Thermo Fisher Scientific | Cat# 10639011 |
| DMEM, high glucose, GlutaMAX supplement | Thermo Fisher Scientific | Cat# 10569044 |
| FBS, HI | Thermo Fisher Scientific | Cat# 16140071 |
| Recombinant human stem cell factor | Peprotech | Cat# 300-07 |
| Recombinant murine stem cell factor | Peprotech | Cat# 250-03 |
| G418 Sulfate | Thermo Fisher Scientific | Cat# 30-234-CR |
| Poly-D-lysine hydrobromide | Sigma-Aldrich | Cat# P7280 |
| Fibronectin | Sigma-Aldrich | Cat# F0895 |
| Fluo-4, AM | Thermo Fisher Scientific | Cat# F14201 |
| Fura-2, AM | Thermo Fisher Scientific | Cat# F1225 |
| Pluronic; F-127 | Sigma-Aldrich | Cat# P3000MP |
| Lipofectamine 3000 | Thermo Fisher Scientific | Cat# L3000015 |
| CFSE | Sigma-Aldrich | Cat# 21888 |
| Avidin sulforhodamine 101 conjugate | Marker Gene Tech | Cat# M1124 |
| Sytox Green | Thermo Fisher Scientific | Cat# S7020 |
| Crystal Violet | Sigma-Aldrich | Cat# C0775 |
| Safranin O | Sigma-Aldrich | Cat# S2255 |
| Fluoromount-G | eBioscience | Cat# 00-4958-02 |
| Shandon TBD-2 Decalcifier | Thermo Fisher Scientific | Cat# 6764003 |
| DMSO | Sigma-Aldrich | Cat# D2650 |
| Percoll | Sigma-Aldrich | Cat# P4937 |
| Histamine | Sigma-Aldrich | Cat# H7250 |
| O-phthalaldehyde | Sigma-Aldrich | Cat# P0657 |
| 4-Nitrophenyl N-acetyl-β-D-glucosaminide | Sigma-Aldrich | Cat# N9376 |
| Triton X-100 | Sigma-Aldrich | Cat# X-100 |
| Brain Heart Infusion broth | BD | Cat# 221812 |
| Todd Hewitt Broth | Sigma-Aldrich | Cat# T1438 |
| Tryptic Soy Agar | Sigma-Aldrich | Cat# 22901 |
| Sheep Blood | Thermo Fisher Scientific | Cat# R54004 |
| Blood agar (TSA with 5% sheep blood) | Thermo Fisher Scientific | Cat# R01201 |
| ON-TARGETplus Non-targeting Pool | Dharmacon | Cat# D-001810-10 |
| SMARTpool: ON-TARGETplus MRGPRX2 siRNA | Dharmacon | Cat# L-005666 |
| Trimethoprim | Sigma-Aldrich | Cat# T7883 |
| Critical Commercial Assays | ||
| Histamine ELISA kit | Enzo Life Sciences | Cat# ENZ-KIT140-0001 |
| Mouse TNF-alpha Quantikine ELISA kit | R&D Systems | Cat# MTA00B |
| Human TNF ELISA set | BD Biosciences | Cat# 555212 |
| DCFDA – Cellular ROS detection assay | Abcam | Cat# ab113851 |
| Cysteinyl Leukotriene ELISA Kit | Cayman Chemicals | Cat#500390 |
| Prostaglandin D2ELISA Kit | Cayman Chemicals | Cat# 512031 |
| FLIPR Calcium 5 Assay Kit | Molecular Devices | Cat# R8186 |
| Direct-Zol RNA Miniprep Plus Kit | Zymo Research | Cat# R2071 |
| iSCRIPT cDNA Synthesis Kit | Bio-Rad | Cat# 1708891 |
| Experimental Models: Cell Lines | ||
| Human: LAD2 cells | Laboratory of Allergic Diseases, NIH | N/A |
| Human: HEK 293 cells | ATCC | Cat# ATCC CRL-1573 |
| Experimental Models: Organisms/Strains | ||
| Mouse: C57Bl/6J | The Jackson Lab | Cat#000664 |
| Mouse: Mrgprb2−/−;C57Bl/6J Mrgprb2 knockout | This lab | N/A |
| Mouse: Mrgprb2-Cre; C57Bl/6J Cre-recombinase under Mrgprb2 promoter | This lab | N/A |
| Mouse: tdTomato (tdT); B6; 129S - Gt(ROSA) 26Sortm14(CAG-tdTomato)Hze/J | The Jackson Lab | Cat# 007908 |
| Oligonucleotides | ||
| Primer: WT mice Forward: GGTTCCTGGGCATCCGTAT |
(McNeil et al., 2015) | N/A |
| Primer: WT mice Reverse: CTTCCGCCTGAACCTTCGGT |
(McNeil et al., 2015) | N/A |
| Primer: Mrgprb2MUT Forward: GTTCCTGGGCATCCGCAC |
(McNeil et al., 2015) | N/A |
| Primer: Mrgprb2MUT Reverse: CTTCCGCCTGAACCTTCGGT |
(McNeil et al., 2015) | N/A |
| Primer: Mrgprb2-Cre Forward: TATATCATGGCCGACAAGCA |
(McNeil et al., 2015) | N/A |
| Primer: Mrgprb2MUT Reverse: CAGACCGCGCGCCTGAAGA |
(McNeil et al., 2015) | N/A |
| Primer: tdTomato WT
Forward: AAGGGAGCTGCAGTGGAGTA |
(McNeil et al., 2015) | N/A |
| Primer: tdTomato WT
Reverse: CCGAAAATCTGTGGGAAGTC |
(McNeil et al., 2015) | N/A |
| Primers: tdTomato Mutant
Reverse: GGCATTAAAGCAGCGTATCC |
(McNeil et al., 2015) | N/A |
| Primer: tdTomato Mutant
Forward: CTGTTCCTGTACGGCATGG |
(McNeil et al., 2015) | N/A |
| Defb1 | Thermo Fischer Scientific | Cat#4331182, Assay ID: Mm00432803_m1 |
| Defb2 | Thermo Fischer Scientific | Cat#4331182, Assay ID: Mm00657074_m1 |
| Defb3 | Thermo Fischer Scientific | Cat#4331182, Assay ID: Mm01614469_m1 |
| Defb4 | Thermo Fischer Scientific | Cat#4331182, Assay ID: Mm00731768 m1 |
| Defb14 | Thermo Fischer Scientific | Cat#4331182, Assay ID: Mm00806979_m1 |
| Camp | Thermo Fischer Scientific | Cat#4331182, Assay ID: Mm00438285_m1 |
| PENK | Thermo Fischer Scientific | Cat#4331182, Assay ID: Mm01212875_m1 |
| Hprt | Thermo Fisher Scientific | Cat#4331182, Assay ID: Mm03024075_m1 |
| Recombinant DNA | ||
| Plasmid: EGFP-N1 | Clontech | Cat#6085-1 |
| Software and Algorithms | ||
| Image J | NIH | https://imagej.nih.gov/ij/? |
| MO.Affinity Analysis v2.3 | NanoTemper Tech | https://nanotempertech.com/monolith-mo-control-software/ |
| FlowJo v.10 | FlowJo, LLC | https://www.flowjo.com |
| Prism 7.0 | GraphPad Software | https://www.graphpad.com/scientific-software/prism/ |
| Illustrator | Adobe | https://www.adobe.com/products/illustrator.html |
Highlights.
The mammalian receptor Mrgprb2/X2 can detect bacterial quorum sensing molecules (QSMs)
QSM detection by Mrgprb2/X2 in mast cells elicits antibacterial mediator release
Mrgprb2 recognition of QSMs is critical for an effective immune response to bacteria
Pharmacologic activation of Mrgprb2/X2 enhances bacterial clearance
ACKNOWLEDGMENTS
We acknowledge valuable discussions with Drs. Benjamin McNeil, Shukti Chakravarti, and Mengfei Chen. The work was supported by grants from the NIH to X.D. (NS054791 and AI135186), J.E.V. (R21AI112768-01A1), S.H.S (MH18501), and C.V. (T32GM73009). N.G. is supported by the Société Française de Dermatologie, the Société Française d’Allergologie, the Marie Skłodowska-Curie Individual Fellowship (H2020-MSCA-IF-2016 #749629), the European Research Council (ERC-2018-STG #802041), and the INSERM. P.P. received a Canadian Institutes of Health Research Fellowship.
Footnotes
DECLARATION OF INTERESTS
The authors declare no competing interests.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES:
- BIENENSTOCK J, BEFUS AD, PEARCE F, DENBURG J & GOODACRE R 1982. Mast cell heterogeneity: derivation and function, with emphasis on the intestine. J Allergy Clin Immunol, 70, 407–12. [DOI] [PubMed] [Google Scholar]
- CVITKOVITCH DG, LI YH & ELLEN RP 2003. Quorum sensing and biofilm formation in Streptococcal infections. J Clin Invest, 112, 1626–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DAVIES DG, PARSEK MR, PEARSON JP, IGLEWSKI BH, COSTERTON JW & GREENBERG EP 1998. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science, 280, 295–8. [DOI] [PubMed] [Google Scholar]
- DAWICKI W & MARSHALL JS 2007. New and emerging roles for mast cells in host defence. Curr Opin Immunol, 19, 31–8. [DOI] [PubMed] [Google Scholar]
- DONG X, HAN S, ZYLKA MJ, SIMON MI & ANDERSON DJ 2001. A diverse family of GPCRs expressed in specific subsets of nociceptive sensory neurons. Cell, 106, 619–32. [DOI] [PubMed] [Google Scholar]
- DUHR S & BRAUN D 2006. Why molecules move along a temperature gradient. Proc Natl Acad Sci U S A, 103, 19678–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ENERBACK L 1966. Mast cells in rat gastrointestinal mucosa. 2. Dye-binding and metachromatic properties. Acta Pathol Microbiol Scand, 66, 303–12. [DOI] [PubMed] [Google Scholar]
- FREUND JR, MANSFIELD CJ, DOGHRAMJI LJ, ADAPPA ND, PALMER JN, KENNEDY DW, REED DR, JIANG P & LEE RJ 2018. Activation of airway epithelial bitter taste receptors by Pseudomonas aeruginosa quinolones modulates calcium, cyclic-AMP, and nitric oxide signaling. J Biol Chem, 293, 9824–9840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- GALLI SJ, NAKAE S & TSAI M 2005. Mast cells in the development of adaptive immune responses. Nat Immunol, 6, 135–42. [DOI] [PubMed] [Google Scholar]
- HANCOCK RE & DIAMOND G 2000. The role of cationic antimicrobial peptides in innate host defences. Trends Microbiol, 8, 402–10. [DOI] [PubMed] [Google Scholar]
- HAVARSTEIN LS, COOMARASWAMY G & MORRISON DA 1995. An unmodified heptadecapeptide pheromone induces competence for genetic transformation in Streptococcus pneumoniae. Proc Natl Acad Sci U S A, 92, 11140–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- HAVARSTEIN LS, HAKENBECK R & GAUSTAD P 1997. Natural competence in the genus Streptococcus: evidence that streptococci can change pherotype by interspecies recombinational exchanges. J Bacteriol, 179, 6589–94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- KARIMINIK A, BASERI-SALEHI M & KHEIRKHAH B 2017. Pseudomonas aeruginosa quorum sensing modulates immune responses: An updated review article. Immunol Lett, 190, 1–6. [DOI] [PubMed] [Google Scholar]
- KIRSHENBAUM AS, AKIN C, WU Y, ROTTEM M, GOFF JP, BEAVEN MA, RAO VK & METCALFE DD 2003. Characterization of novel stem cell factor responsive human mast cell lines LAD 1 and 2 established from a patient with mast cell sarcoma/leukemia; activation following aggregation of FcepsilonRI or FcgammaRI. Leuk Res, 27, 677–82. [DOI] [PubMed] [Google Scholar]
- KURASHIMA Y & KIYONO H 2014. New era for mucosal mast cells: their roles in inflammation, allergic immune responses and adjuvant development. Exp Mol Med, 46, e83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LATTAR SM, WU X, BROPHY J, SAKAI F, KLUGMAN KP & VIDAL JE 2018. A Mechanism of Unidirectional Transformation, Leading to Antibiotic Resistance, Occurs within Nasopharyngeal Pneumococcal Biofilm Consortia. MBio, 9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LEENDERTSE M, WILLEMS RJ, GIEBELEN IA, VAN DEN PANGAART PS, WIERSINGA WJ, DE VOS AF, FLORQUIN S, BONTEN MJ & VAN DER POLL T 2008. TLR2-dependent MyD88 signaling contributes to early host defense in murine Enterococcus faecium peritonitis. J Immunol, 180, 4865–74. [DOI] [PubMed] [Google Scholar]
- LEWIS K 2007. Persister cells, dormancy and infectious disease. Nat Rev Microbiol, 5, 48–56. [DOI] [PubMed] [Google Scholar]
- MARSHALL JS 2004. Mast-cell responses to pathogens. Nat Rev Immunol, 4, 787–99. [DOI] [PubMed] [Google Scholar]
- MASHBURN-WARREN L, MORRISON DA & FEDERLE MJ 2012. The cryptic competence pathway in Streptococcus pyogenes is controlled by a peptide pheromone. J Bacteriol, 194, 4589–600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MCNEIL BD, PUNDIR P, MEEKER S, HAN L, UNDEM BJ, KULKA M & DONG X 2015. Identification of a mast-cell-specific receptor crucial for pseudo-allergic drug reactions. Nature, 519, 237–41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- NILSSON G, JOHNELL M, HAMMER CH, TIFFANY HL, NILSSON K, METCALFE DD, SIEGBAHN A & MURPHY PM 1996. C3a and C5a are chemotaxins for human mast cells and act through distinct receptors via a pertussis toxin-sensitive signal transduction pathway. J Immunol, 157, 1693–8. [PubMed] [Google Scholar]
- O'TOOLE GA 2011. Microtiter dish biofilm formation assay. J Vis Exp. [DOI] [PMC free article] [PubMed] [Google Scholar]
- PRITCHARD DI 2006. Immune modulation by Pseudomonas aeruginosa quorum-sensing signal molecules. Int J Med Microbiol, 296, 111–6. [DOI] [PubMed] [Google Scholar]
- PUNDIR P & KULKA M 2010. The role of G protein-coupled receptors in mast cell activation by antimicrobial peptides: is there a connection? Immunol Cell Biol, 88, 632–40. [DOI] [PubMed] [Google Scholar]
- QIAO H, ANDRADE MV, LISBOA FA, MORGAN K & BEAVEN MA 2006. FcepsilonR1 and toll-like receptors mediate synergistic signals to markedly augment production of inflammatory cytokines in murine mast cells. Blood, 107, 610–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- REBER LL, SIBILANO R, STARKL P, ROERS A, GRIMBALDESTON MA, TSAI M, GAUDENZIO N & GALLI SJ 2017. Imaging protective mast cells in living mice during severe contact hypersensitivity. JCI Insight, 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- RUTHERFORD ST & BASSLER BL 2012. Bacterial quorum sensing: its role in virulence and possibilities for its control. Cold Spring Harb Perspect Med, 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- SAITO H, OKAJIMA F, MOLSKI TF, SHA'AFI RI, UI M & ISHIZAKA T 1987. Effects of ADP-ribosylation of GTP-binding protein by pertussis toxin on immunoglobulin E-dependent and -independent histamine release from mast cells and basophils. J Immunol, 138, 3927–34. [PubMed] [Google Scholar]
- SANDIG H & BULFONE-PAUS S 2012. TLR signaling in mast cells: common and unique features. Front Immunol, 3, 185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- SEIFERT R 2015. How do basic secretagogues activate mast cells? Naunyn Schmiedebergs Arch Pharmacol, 388, 279–81. [DOI] [PubMed] [Google Scholar]
- SHAO H, LEE S, GAE-SCOTT S, NAKATA C, CHEN S, HAMAD AR & CHAKRAVARTI S 2012. Extracellular matrix lumican promotes bacterial phagocytosis, and Lum−/− mice show increased Pseudomonas aeruginosa lung infection severity. J Biol Chem, 287, 35860–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- SKOGMAN ME, VUORELA PM & FALLARERO A 2012. Combining biofilm matrix measurements with biomass and viability assays in susceptibility assessments of antimicrobials against Staphylococcus aureus biofilms. The Journal Of Antibiotics, 65, 453. [DOI] [PubMed] [Google Scholar]
- STROEHER UH, PATON AW, OGUNNIYI AD & PATON JC 2003. Mutation of luxS of Streptococcus pneumoniae affects virulence in a mouse model. Infect Immun, 71, 3206–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- SUBRAMANIAN H, GUPTA K, LEE D, BAYIR AK, AHN H & ALI H 2013. beta-Defensins activate human mast cells via Mas-related gene X2. J Immunol, 191, 345–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- URB M & SHEPPARD DC 2012. The role of mast cells in the defence against pathogens. PLoS Pathog, 8, e1002619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- VERBEURGT C, VEITHEN A, CARLOT S, TARABICHI M, DUMONT JE, HASSID S & CHATELAIN P 2017. The human bitter taste receptor T2R38 is broadly tuned for bacterial compounds. PLoS One, 12, e0181302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- VIDAL JE, HOWERY KE, LUDEWICK HP, NAVA P & KLUGMAN KP 2013. Quorum-sensing systems LuxS/autoinducer 2 and Com regulate Streptococcus pneumoniae biofilms in a bioreactor with living cultures of human respiratory cells. Infect Immun, 81, 1341–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- WATERS CM & BASSLER BL 2005. Quorum sensing: cell-to-cell communication in bacteria. Annu Rev Cell Dev Biol, 21, 319–46. [DOI] [PubMed] [Google Scholar]
- YARWOOD JM & SCHLIEVERT PM 2003. Quorum sensing in Staphylococcus infections. J Clin Invest, 112, 1620–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ZHU L & LAU GW 2011. Inhibition of competence development, horizontal gene transfer and virulence in Streptococcus pneumoniae by a modified competence stimulating peptide. PLoS Pathog, 7, e1002241. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data generated or analyzed during this study are included in this article.