Abstract
Background
4-Thiazolidinone ring is reported to have almost all types of biological activities. Also, it present in many marketed drugs.
Results
Ethyl acetoacetate reacted with phenyl isothiocyanate and ethyl chloroacetate in presence of K2CO3 and DMF to afford the thiazolidinone derivative 5. Thiazolidinone 5 reacted with dimethylformamide-dimethylacetal to afford (Z)-ethyl 2-((Z)-5-((dimethylamino) methylene)-4-oxo-3-phenylthiazolidin-2-ylidene)acetate (6). The structure of thiazolidinone 6 was elucidated from its spectral analysis and X-ray crystallography and was optimized using B3LYP/6-31G(d,p) method. The geometric parameters and NMR spectra were discussed both experimentally and theoretically. Also, the natural charges at the different atomic sites were predicted. The synthesized compounds had moderate cytotoxic activity.
Conclusions
An unexpected synthesis of (Z)-ethyl 2-((Z)-5-((dimethylamino)methylene)-4-oxo-3-phenylthiazolidin-2-ylidene)acetate via deacetylation mechanism. The structure was established using X-ray and spectral analysis. The geometric parameters, and NMR spectra were discussed. The synthesized compounds showed moderate anticancer activity. 
Keywords: Thiazolidinone, X-ray crystallography, Computational studies, DMF-DMA, Cytotoxic activity
Introduction
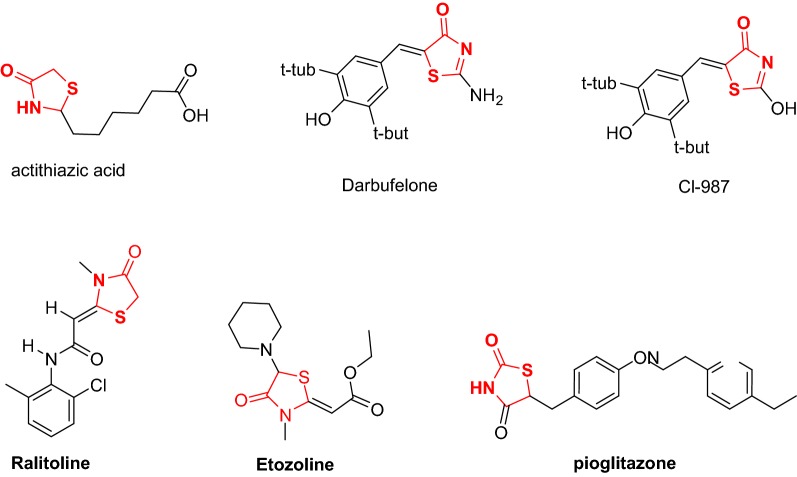
The thiazolidinone ring had diverse biological activities such as antimycobacterial [1], antimicrobial [2], anticancer [3], anticonvulsant [4], antiparasitic [5], antidiabetic [6], and antihypertensive [7]. Also, many clinically used drugs contain thiazolidinone ring in their skeletons such as antibiotic actithiazic acid [8], dual COX/LOX inhibitors (Darbufelone and CI-987) [9], Ralitoline, Etozoline, and Pioglitazone (Fig. 1).
Fig. 1.
Examples of some drugs containing 4-thiazolidinone ring
Due to this diversity in the biological activities of 4-thiazolidinones, there are several procedures have been reported for their synthesis [10–14]. In this research, we synthesize new 4-thiazolidinone derivative 6 in a pure stat, also, we compare cytotoxic activity of synthesized compounds with standard anticancer drug Vinblastine against the colon carcinoma (HCT-116) cell line using MTT assay.
Results and discussion
Chemistry
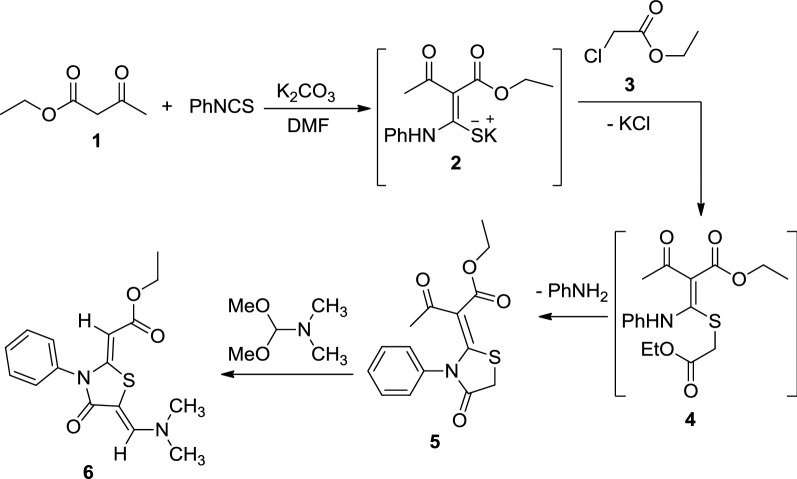
The thiazolidinone derivative 5 was prepared according to the reported method [15]. Refluxing of the thiazolidinone 5 with dimethylformamide–dimethylacetal in DMF gave only one isolable product, the product of this reaction is (Z)-ethyl 2-((Z)-5-((dimethylamino) methylene)-4-oxo-3-phenylthiazolidin-2-ylidene)acetate (6) (Scheme 1). Spectral data (IR, NMR, and X-ray were in a complete agreement with the proposed structure.
Scheme 1.
Synthesis of thiazolidinones 5 and 6
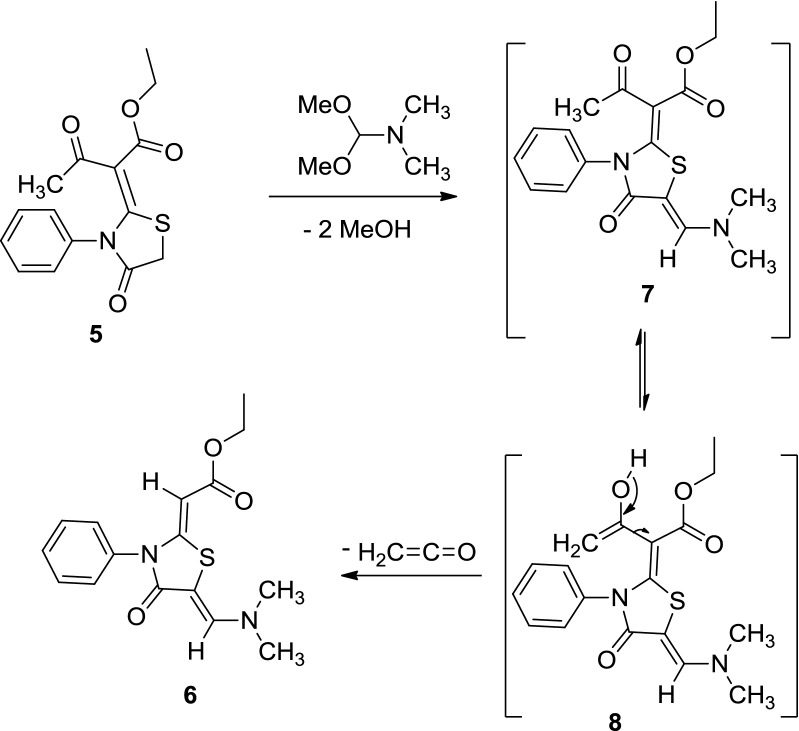
The following reaction mechanism was suggested for the formation of thiazolidinone derivative 6 (Scheme 2). We assumed that the reaction was started between thiazolidinone 5 and dimethylformamide-dimethylacetal to produce an intermediate 7, which underwent enolization of the carbonyl of the acetyl group, followed by elimination of ketene to give the thiazolidinone derivative 6 (Scheme 2). The configuration of thiazolidinone 6 was confirmed using X-ray analysis (Fig. 2) [16].
Scheme 2.
The suggested mechanism for the synthesis of thiazolidinone derivative 6
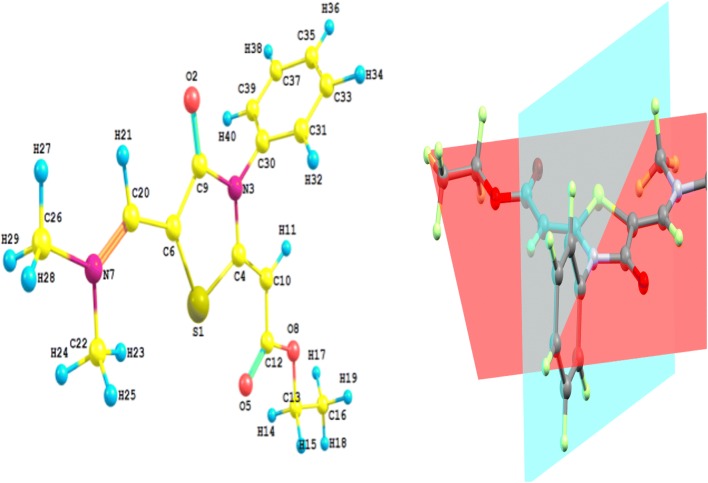
Fig. 2.
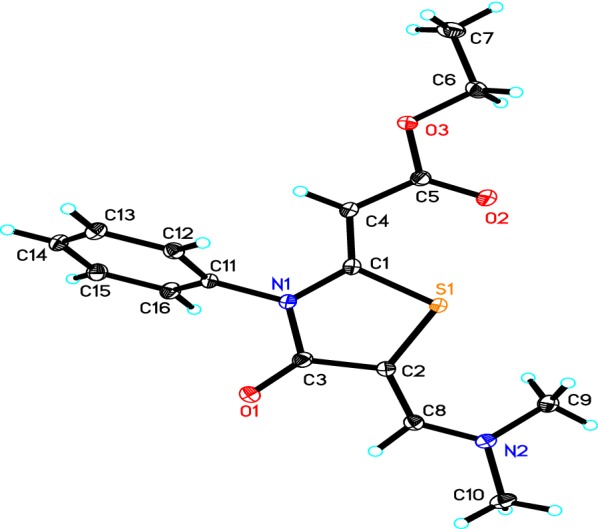
The ORTEP diagram of the final X-ray model of thiazolidinone 6 with displacement ellipsoids drawn at 50% probability level. H-atoms were placed and not included in refinement
Crystal structure determination
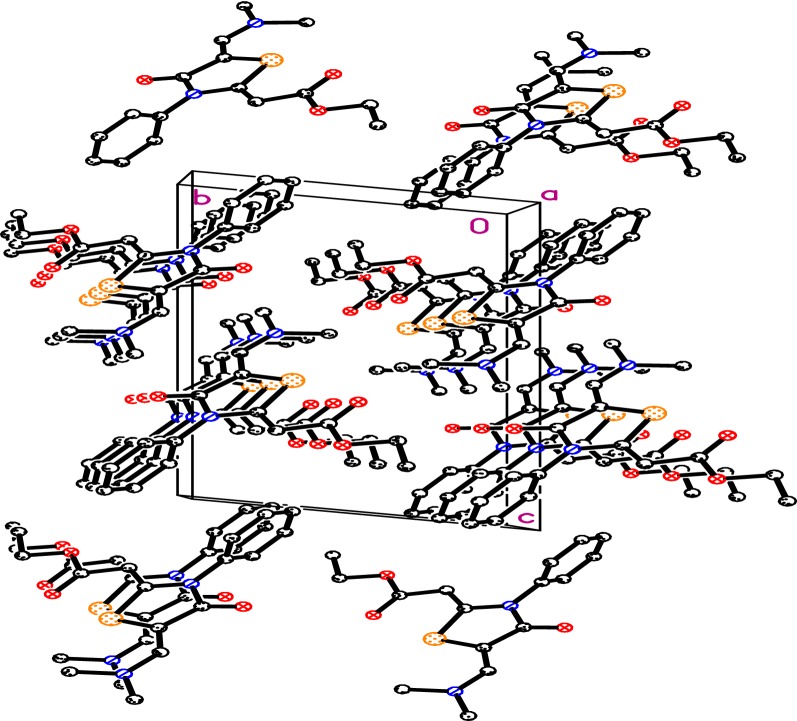
A crystal of dimensions 0.47 × 0.26 × 0.14 mm was selected for X-ray diffraction analysis. Data were collected on a Bruker APEX-II diffractometer equipped with CCD detector and graphite monochromatic Mo Kα radiation (λ = 71,073 Å) at 100 °K. SHELXS-97 [17, 18] was used to solve structure (Table 1). Cell refinement and data reduction were carried out by Bruker SAINT [19]. The final refinement was carried out by full-matrix least-squares techniques with anisotropic thermal data for nonhydrogen atoms on F2. All the hydrogen atoms were placed in calculated positions. The crystal of thiazolidinone 6 (Fig. 2) was finally refined with R factor of 4.46% for 4622 unique reflections. Molecules were found to be packed in crystal lattice through intermolecular hydrogen bonding (Fig. 3). Table 2 summarized some selected geometric parameters for thiazolidinone 6.
Table 1.
The crystal and experimental data of thiazolidinone 6
| Parameters | |
|---|---|
| Empirical formula | C16H18N2O3S |
| Formula weight | 318.38 |
| Temperature | 100 °K |
| Wave length | 0.71073 Å |
| Crystal system | Triclinic, |
| space group | P-1 |
| Unit cell dimensions | a = 5.6502 (6) Å |
| b = 9.1968 (9) Å | |
| c = 14.9469 (16) Å | |
| α = 98.992 (5)° | |
| β = 91.848 (7)° | |
| γ = 96.184 (5)° | |
| Volume | 761.73 (14) Å3 |
| Z | 2 |
| Calculated density | 1.388 Mg m−3 |
| Absorption coefficient | 0.23 mm−1 |
| F(000) | 336 |
| Crystal size | 0.47 × 0.26 × 0.14 mm |
| Theta range for data collection | 1.4° to 30.7° |
| Limiting indices | − 8 ≤h ≤ 8, − 11 ≤k ≤ 13, − 21 ≤l ≤ 21 |
| Reflections collected/unique | 16,661/4622 [R(int) = 0.045] |
| Completeness to theta | 30.7°–98.1% |
| Absorption correction | Semi-empirical from equivalents |
| Max. and min. transmission | 0.7318 and 0.7106 |
| Refinement method | Full-matrix least-squares on F-2 |
| Data/restraints/parameters | 4622/0/203 |
| Goodness-of-fit on F-2 | 1.04 |
| Final R indices [I > 2sigma(I)] | R1 = 0.0344, wR2 = 0.088 |
| R indices (all data) | R1 = 0.0446, wR2 = 0.092 |
| Largest diff. peak and hole | 0.45 and − 0.32 e.Å−3 |
Fig. 3.
Molecular packing of the thiazolidinone 6
Table 2.
Selected geometric parameters (Å, °) for thiazolidinone 6
| S1—C1 | 1.7461 (11) | N1—C1 | 1.3854 (14) |
| S1—C2 | 1.7634 (11) | N1—C3 | 1.4115 (14) |
| O1—C3 | 1.2261 (13) | N1—C11 | 1.4364 (13) |
| O2—C5 | 1.2219 (13) | N2—C8 | 1.3300 (14) |
| O3—C5 | 1.3560 (13) | N2—C9 | 1.4543 (15) |
| O3—C6 | 1.4535 (14) | N2—C10 | 1.4539 (15) |
| C1—S1—C2 | 91.59 (5) | S1—C2—C8 | 129.00 (8) |
| C5—O3—C6 | 114.88 (8) | O1—C3—N1 | 122.06 (9) |
| C1—N1—C3 | 116.76 (9) | O1—C3—C2 | 128.67 (10) |
| C1—N1—C11 | 122.46 (9) | N1—C3—C2 | 109.27 (9) |
| C3—N1—C11 | 120.76 (8) | O2—C5—O3 | 122.19 (10) |
| C8—N2—C9 | 122.19 (9) | O2—C5—C4 | 125.53 (10) |
| C8—N2—C10 | 121.08 (9) | O3—C5—C4 | 112.28 (9) |
| C9—N2—C10 | 116.31 (9) | O3—C6—C7 | 107.39 (9) |
| S1—C1—N1 | 110.58 (8) | N2—C8—C2 | 130.35 (10) |
| S1—C1—C4 | 124.14 (8) | N1—C11—C12 | 120.10 (9) |
| N1—C1—C4 | 125.27 (10) | N1—C11—C16 | 118.75 (9) |
| S1—C2—C3 | 111.73 (8) |
Geometry optimization
The optimized molecular geometry of thiazolidinone 6 is shown in Fig. 4. Table 3 listed the hydrogen bonding data for thiazolidinone 6, and the results of the calculated bond distances and angles are given in Table 4. It is clear that the calculated bond distances and angles agree very well with the experimental results. The bond distances and angles deviated only by 0.027 Å (C20–N7) and 1.6° (C20–C6–S1). The correlation coefficients given in the same table indicated the good agreement between the calculated structure and the X-ray results. Interestingly, the thiazole and phenyl ring planes are not coplanar and are twisted from one another by an angle of 77.1° (exp. 66.5°).
Fig. 4.
The optimized structure of thiazolidinone 6
Table 3.
Hydrogen bonding data for thiazolidinone 6
| D—H···A | D—H | H···A | D···A | D—H···A |
|---|---|---|---|---|
| C8—H8···O1 | 0.95 | 2.53 | 2.8888 (13) | 102 |
| C9—H9A···S1 | 0.98 | 2.77 | 3.1190 (12) | 102 |
| C9—H9B···O2i | 0.98 | 2.33 | 3.3018 (15) | 171 |
| C12—H12···O1ii | 0.95 | 2.37 | 3.3042 (14) | 168 |
Symmetry codes: (i) − x + 1, − y + 1, − z + 1; (ii) x − 1, y, z
Table 4.
The experimental and calculated geometric parameters of thiazolidinone 6
| Bond distances | Calc. | X-ray | Bond angles | Calc. | X-ray |
|---|---|---|---|---|---|
| R(1–4) | 1.773 | 1.746 | A(4-1-6) | 91.0 | 91.6 |
| R(1–6) | 1.786 | 1.763 | A(1-4-3) | 111.0 | 110.6 |
| R(2–9) | 1.223 | 1.226 | A(1-4-10) | 123.9 | 124.1 |
| R(3–4) | 1.387 | 1.385 | A(1-6-9) | 111.6 | 111.7 |
| R(3–9) | 1.417 | 1.412 | A(1-6-20) | 131.0 | 129.0 |
| R(3–30) | 1.435 | 1.436 | A(2-9-3) | 122.8 | 122.1 |
| R(4–10) | 1.362 | 1.361 | A(2-9-6) | 127.8 | 128.7 |
| R(5–12) | 1.227 | 1.222 | A(4-3-9) | 116.9 | 116.8 |
| R(6–9) | 1.463 | 1.444 | A(4-3-30) | 122.8 | 122.5 |
| R(6–20) | 1.363 | 1.369 | A(3-4-10) | 125.2 | 125.3 |
| R(7–20) | 1.357 | 1.330 | A(9-3-30) | 120.2 | 120.8 |
| R(7–22) | 1.456 | 1.454 | A(3-9-6) | 109.5 | 109.3 |
| R(7–26) | 1.456 | 1.454 | A(3-30-31) | 119.9 | 120.1 |
| R(8–12) | 1.360 | 1.356 | A(3-30-39) | 119.6 | 118.7 |
| R(8–13) | 1.441 | 1.453 | A(4-10-12) | 121.0 | 119.8 |
| R(10–12) | 1.448 | 1.444 | A(5-12-8) | 122.6 | 122.2 |
| R(13–16) | 1.517 | 1.506 | A(5-12-10) | 125.7 | 125.5 |
| R(30–31) | 1.397 | 1.391 | A(9-6-20) | 117.4 | 119.2 |
| R(30–39) | 1.396 | 1.388 | A(6-20-7) | 132.2 | 130.3 |
| R(31–33) | 1.394 | 1.391 | A(20-7-22) | 123.4 | 122.2 |
| R(33–35) | 1.396 | 1.389 | A(20-7-26) | 119.3 | 121.1 |
| R(35–37) | 1.396 | 1.390 | A(22-7-26) | 115.6 | 116.3 |
| R(37–39) | 1.394 | 1.390 | A(7-22-23) | 110.6 | 109.5 |
| A(7-22-24) | 108.9 | 109.5 | |||
| A(12-8-13) | 115.6 | 114.9 | |||
| A(8-12-10) | 111.8 | 112.3 | |||
| A(8-13-16) | 107.4 | 107.4 | |||
| A(11-10-12) | 118.8 | 120.1 | |||
| A(31-30-39) | 120.5 | 121.2 | |||
| A(30-31-33) | 119.6 | 118.9 | |||
| A(30-39-37) | 119.6 | 119.4 | |||
| A(32-31-33) | 120.8 | 120.6 | |||
| A(31-33-35) | 120.1 | 120.5 | |||
| A(33-35-37) | 119.9 | 120.0 | |||
| A(35-37-39) | 120.2 | 120.1 | |||
| R2 | 0.9979 | 0.9936 | |||
R2: correlation coefficient
Charge population analysis
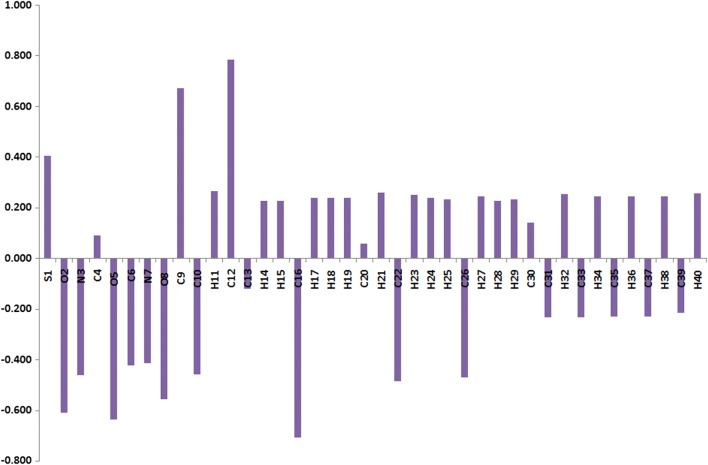
The natural population analysis is performed to predict the natural charges (NC) at the different atomic sites (Fig. 5). It is clear that the sulfur atom is electropositive. The C9 and C12 are the most electropositive C-atoms as these atomic sites are bonded to two strong electronegative atoms. In contrast, the O and N-atoms have electronegative nature where the ring N-atom (N3) is more electronegative than the imine one (N7). Among the O-atoms in the studied molecule, the carbonyl O-atoms are the most electronegative where the O-atom (O5) from the carbonyl group in the ester moiety is more negative than that of the cyclic ketone (O2). As revealed from the molecular electrostatic potential (MEP) map shown in Fig. 6, the most negative regions are located over the O-atoms while the positive region is mainly located over the protons of the two methyl groups.
Fig. 5.
The natural charges at the different atomic sites of thiazolidinone 6
Fig. 6.
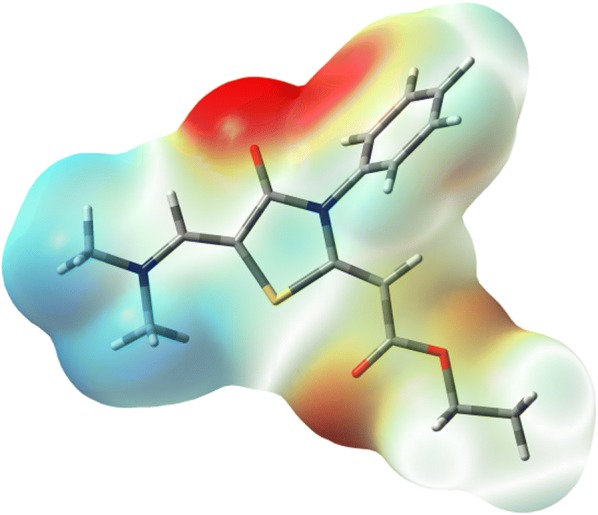
The MEP figure of thiazolidinone 6
Frontier molecular orbitals
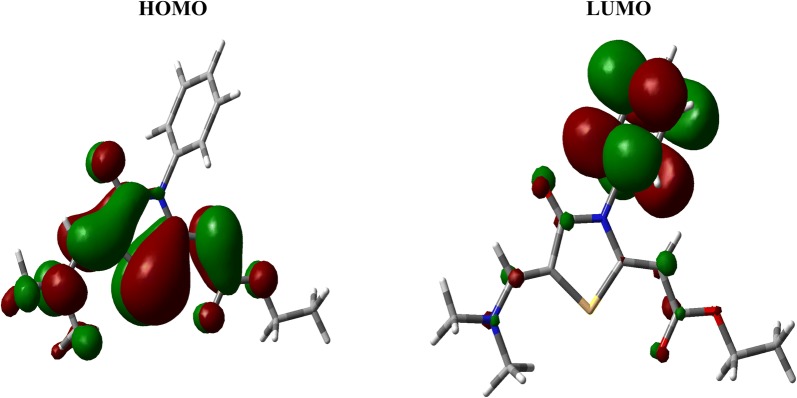
The HOMO and LUMO levels of the thiazolidinone 6 are shown in Fig. 7. The HOMO and LUMO energies are − 5.0584 and − 0.9870 eV, respectively. As a result, the HOMO–LUMO energy gap is calculated to be 4.0714 eV. The HOMO and LUMO are mainly localized over the thiazole and phenyl rings, respectively. Since the HOMO and LUMO levels are mainly located over the π-system of the studied compound 6 so the HOMO–LUMO intramolecular charge transfer is mainly a π–π* transition.
Fig. 7.
The frontier molecular orbitals calculated at the B3LYP/6-311G(d,p) level
NMR spectra
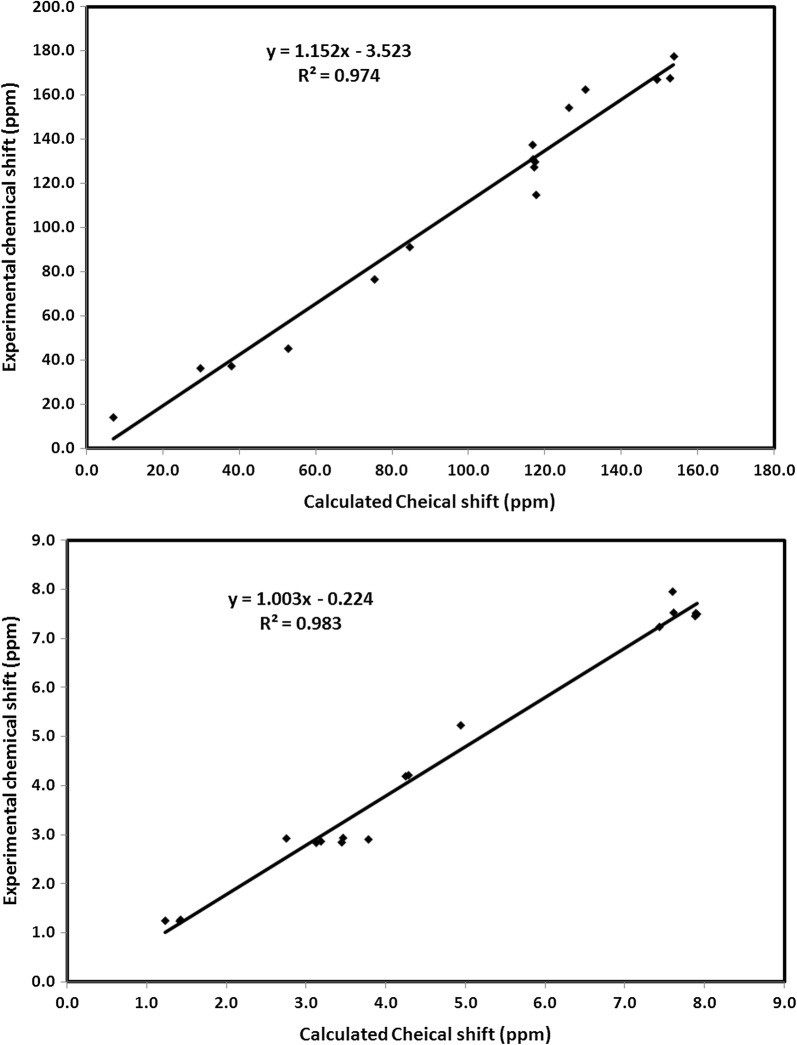
The Gauge-independent atomic orbital (GIAO) calculations were used for accurate prediction of the 1H and 13C isotropic chemical shifts (C.S). The isotropic chemical shifts were used for the identification of organic compounds. The theoretical and experimental chemical shifts are presented in Table 5. Correlation graphs between the experimental and theoretical NMR chemical shifts are shown in Fig. 8. The correlations equations shown in this figure have high R2 values (0.974–0.983) indicating the good agreement between the theoretical and experimental data. The 1H-chemical shifts of the aromatic ring usually appear in the region of 7–8 ppm. In the present case, the aromatic protons were detected at 7.46–7.96 ppm which is in good agreement with B3LYP theoretical values (7.60–7.89 ppm). The aliphatic protons have lower chemical shifts than the aromatic ones (Table 5).
Table 5.
The calculated and experimental 1H and 13C NMR chemical shifts of thiazolidinone 6
| Atom | C. Scalc. | C. Sexp. | Atom | C. Scalc. | C. Sexp. |
|---|---|---|---|---|---|
| C4 | 149.2 | 166.9 | H11 | 4.94 | 5.23 |
| C6 | 84.5 | 91.2 | H14 | 4.25 | 4.19 |
| C9 | 152.6 | 167.5 | H15 | 4.29 | 4.21 |
| C10 | 75.3 | 67.5 | H17 | 1.41 | 1.23 |
| C12 | 153.7 | 177.4 | H18 | 1.23 | 1.24 |
| C13 | 52.7 | 60.9 | H19 | 1.42 | 1.26 |
| C16 | 6.9 | 14.3 | H21 | 7.44 | 7.24 |
| C20 | 130.5 | 162.4 | H23 | 3.78 | 2.90 |
| C22 | 29.7 | 36.4 | H24 | 2.75 | 2.92 |
| C26 | 37.8 | 37.4 | H25 | 3.46 | 2.93 |
| C30 | 126.2 | 154.2 | H27 | 3.13 | 2.83 |
| C31 | 116.8 | 130.9 | H28 | 3.45 | 2.84 |
| C33 | 117.1 | 127.3 | H29 | 3.19 | 2.86 |
| C35 | 116.7 | 137.5 | H32 | 7.61 | 7.53 |
| C37 | 117.3 | 129.7 | H34 | 7.89 | 7.52 |
| C39 | 117.6 | 114.8 | H36 | 7.88 | 7.46 |
| H38 | 7.91 | 7.50 | |||
| H40 | 7.60 | 7.96 |
Fig. 8.
Correlation graphs between the calculated and experimental 1H and 13C NMR chemical shifts
The chemical shifts of the aromatic carbons usually appear in the overlapped region of the spectrum between 100 and 200 ppm [20]. The atom C30 has higher chemical shift than the other aromatic carbons (Table 5). The high chemical shift of C30 is due to the deshielding effect of the electronegative N-atom. Since the oxygen and nitrogen atoms are more electronegative than carbons so the C-atoms (C4, C9, C12, and C20) attached to these electronegative sites were detected at high chemical shifts (162.4–177.4 ppm) compared to the rest of C-atoms.
Cytotoxic activity
The anti-cancer activity of the thiazolidine derivatives 5 and 6 was determined against the colon carcinoma (HCT-116) cell line in comparison with the anticancer drug vinblastine, using MTT assay [21]. The results of the cytotoxic activity were expressed as the mean IC50 of three independent experiments (Table 6). The results revealed that thiazolidinone derivatives 5 and 6 had moderate anticancer activity against colon carcinoma (HCT-116).
Table 6.
Viability values and IC50 of thiazolidinone derivatives 5 and 6 against HCT-116 Cell Line
| S. No | Sample concentration (μg/mL) viability % | |||||||
|---|---|---|---|---|---|---|---|---|
| 50 | 25 | 12.5 | 6.25 | 3.125 | 1.56 | 0 | IC50 (μg) | |
| Vinblastine | 23.08 | 27.35 | 43.59 | 53.85 | 69.23 | 82.54 | 100 | 9.8 |
| 5 | 42.51 | 76.82 | 84.19 | 93.72 | 98.56 | 100 | 100 | 44.5 |
| 6 | 39.43 | 58.15 | 79.51 | 86.42 | 92.63 | 96.47 | 100 | 35.9 |
Experimental section
Chemistry
General
All the melting points were measured on a Gallenkamp apparatus in open glass capillaries and are uncorrected. The IR Spectra were recorded using Nicolet 6700 FT-IR spectrophotometer. 1H- and 13C-NMR spectra were recorded on a JEOL ECP 500 NMR spectrometer operating at 500 MHz. 1H spectra were run at 500 MHz and 13C spectra were run at 125 MHz in deuterated chloroform (CDCl3). Chemical shifts were related to that of the solvent. Chemical shifts δ are expressed in ppm units. Elemental analysis were carried out on a 2400 CHN Elemental Analyzer. The single-crystal X-ray diffraction measurements were accomplished on a Bruker SMART APEX II CCD diffractometer. The biological evaluations of the products were carried out in the Medical Mycology Laboratory of the Regional Center for Mycology and Biotechnology of Al-Azhar University, Cairo, Egypt. The thiazolidinone derivative 5 was prepared as described in the literature [15].
Synthesis of (Z)-ethyl 2-((Z)-5-((dimethylamino)methylene)- 4-oxo-3-phenylthiazolidin- 2-ylidene)acetate (6)
A mixture of thiazolidinone 5 (3.05 g, 10 mmol) and DMF-DMA (1.19 g, 1.33 mL, 10 mmol) in DMF (3 mL) was heated under reflux for 3 h, then left to cool to room temperature. The precipitated solid was filtered off, washed with EtOH and recrystallized from DMF to afford the thiazolidinone derivative 6 in 20% yield, m.p. 227 °C; IR (KBr) v max 1715 (C=O), 1639 (C=O) cm−1; 1H NMR (500 MHz, CDCl3): δ 1.24 (t, 3H. CH3, J = 7.5 Hz), 2.83 (s, 3H, CH3), 2.91 (s, 3H, CH3), 4.20 (q, 2H, CH2, J = 7.5 Hz), 5.23 (s, 1H, CH), 7.24 (s, 1H, CH), 7.38–7.96 (m, 5H, Ar–H); 13C NMR (125 MHz, CDCl3): δ 14.3 (CH3), 36.4, 37.4 (2CH3), 60.9 (CH2), 67.5, 91.2, 114.8, 127.3, 129.7, 130.9, 137.5, 154.2, 162.4, 166.9, 167.5 (C=O), 177.4 (C=O); MS, m/z (%) 304 (95), 258 (55), 243 (100), 215 (25), 77 (Ph, 18). calcd. for C16H18N2O3S: C, 60.36; H, 5.70; N, 8.80. Found: C, 60.28; H, 5.66; N, 8.85.
X-ray analysis
The thiazolidinone 6 was obtained as single crystals by slow evaporation from DMF solution of the pure compound at room temperature. The crystallographic data for thiazolidinone 6 (CCDC 1551169) can be obtained on request from the director, Cambridge Crystallographic Data Center, 12 Union Road, Cambridge CB2 1EW, UK http://www.ccdc.cam.ac.uk/data_request/cif.
Computational details
The X-ray structure coordinates of thiazolidinone 6 were used for geometry optimization followed by frequency calculations. For this task, we used Gaussian 03 software [22] and B3LYP/6‒31G(d,p) method. All the obtained frequencies are positive and no imaginary modes were detected. GaussView4.1 [23] and Chemcraft [24] programs have been used to extract the calculation results and to visualize the optimized structures.
Cytotoxic activity
The cytotoxic activity of the synthesized compounds was carried out at the Regional Center for Mycology and Biotechnology at Al-Azhar University, Cairo, Egypt according to the reported method [21].
Conclusions
A novel synthesis and DFT studies of (Z)-ethyl 2-((Z)-5-((dimethylamino) methylene)-4-oxo-3-phenylthiazolidin-2-ylidene)acetate are presented. The NMR chemical shifts were described based on the GIAO calculations. The calculated results showed good correlations with the experimental data. The anticancer activity of the synthesized compounds against the colon carcinoma (HCT-116) cell line was tested and showed moderate activity.
Authors’ contributions
YNM designed research; YNM, MMA and WF performed research, analyzed the data, YNM, MMA, SSA, SMS, AA, ABM, HA, and NAK wrote the paper. All authors read and approved the final manuscript.
Acknowledgements
The authors extend their appreciation to the Deanship of Scientific Research at King Khalid University for its funding this prolific research group no. (R. G. P. 2/23/40/2019).
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
The crystallographic data for this compound can be obtained free of charge from the Cambridge Crystallographic Data Centre via http://www.ccdc.cam.ac.uk/data_request/cif.
Funding
The authors extend their appreciation to the Deanship of Scientific Research at King Khalid University for its funding this prolific research Group No. (R. G. P. 2/23/40/2019).
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Yahia N. Mabkhot, Phone: +966-11-467-589, Email: ygaber@kku.edu.sa
Mohammed M. Alharbi, Email: Abomashal3003@hotmail.com
Salim. S. Al-Showiman, Email: Abomashal3003@hotmail.com
Saied M. Soliman, Email: saied1soliman@yahoo.com
Nabila A. Kheder, Email: nabila.abdelshafy@gmail.com
Wolfgang Frey, Email: wolfgang.frey@oc.uni-stuttgart.de.
Abdulrhman Asayari, Email: alsayari@kku.edu.sa.
Abdullatif Bin Muhsinah, Email: ajmohsnah@kku.edu.sa.
H. Algarni, Email: halgarni@kku.edu.sa
References
- 1.Babaoglu K, Page MA, Jones VC, McNeil MR, Dong C, Naismith JH, Lee RE. Novel inhibitors of an emerging target in Mycobacterium tuberculosis; substituted thiazolidinones as inhibitors of dTDP-rhamnose synthesis. Bioorg Med Chem Lett. 2003;13:3227–3230. doi: 10.1016/S0960-894X(03)00673-5. [DOI] [PubMed] [Google Scholar]
- 2.Vicini P, Geronikaki A, Anastasia K, Incerti M, Zani F. Synthesis and antimicrobial activity of novel 2-thiazolylimino-5-arylidene-4-thiazolidinones. Bioorg Med Chem. 2006;14:3859–3864. doi: 10.1016/j.bmc.2006.01.043. [DOI] [PubMed] [Google Scholar]
- 3.Jackson JR, Patrick DR, Dar MM, Huang PS. Targeted anti-mitotic therapies: can we improve on tubulin agents? Nat Rev Cancer. 2007;7:107–117. doi: 10.1038/nrc2049. [DOI] [PubMed] [Google Scholar]
- 4.Kwan P, Sills GJ, Brodie MJ. The mechanisms of action of commonly used antiepileptic drugs. Pharmacol Ther. 2001;90:21–34. doi: 10.1016/S0163-7258(01)00122-X. [DOI] [PubMed] [Google Scholar]
- 5.Pink R, Hudson A, Mouriès M-A, Bendig M. Opportunities and challenges in antiparasitic drug discovery. Nat RevDrug Discov. 2005;4:727–740. doi: 10.1038/nrd1824. [DOI] [PubMed] [Google Scholar]
- 6.Fitzgerald SM, Kemp-Harper BK, Parkington HC, Head GA, Evans RG. Endothelial dysfunction and arterial pressure regulation during early diabetes in mice: roles for nitric oxide and endothelium-derived hyperpolarizing factor. Am J Physiol RegulIntegr Comp Physiol. 2007;293:R707–R713. doi: 10.1152/ajpregu.00807.2006. [DOI] [PubMed] [Google Scholar]
- 7.Brizzi V, Francioli M, Brufani M, Filocamo L, Bruni G, Massarelli P. Synthesis, binding affinity and selectivity of new β1- and β2-adrenoceptor blockers. Farmaco. 1999;54:713–720. doi: 10.1016/S0014-827X(99)00077-4. [DOI] [PubMed] [Google Scholar]
- 8.Schenck JR, De Rose AF. Actithiazic acid. II. Isolation and characterization. Arch Biochem Biophys. 1952;40:263–269. doi: 10.1016/0003-9861(52)90110-0. [DOI] [PubMed] [Google Scholar]
- 9.Abdelall E, Kamel G. Synthesis of new thiazolo-celecoxib analogues as dual cyclooxygenase-2/15-lipoxygenase inhibitors: determination of regio-specific different pyrazole cyclization by 2D NMR. Eur J Med Chem. 2016;118:250–258. doi: 10.1016/j.ejmech.2016.04.049. [DOI] [PubMed] [Google Scholar]
- 10.Erlenmeyer H, Oberlin V. Zur Kenntnis der thiazolidone-(4) Helv. 1947;30:1329–1335. doi: 10.1002/hlca.19470300526. [DOI] [PubMed] [Google Scholar]
- 11.Surrey R. The preparation of 4-thiazolidones by the reaction of thioglycolic acid with Schiff bases. J Am Chem Soc. 1947;69:2911. doi: 10.1021/ja01203a507. [DOI] [PubMed] [Google Scholar]
- 12.Troutman HD, Long LM. The synthesis of 2,3-disubstituted-4-thiazolidinones. J Am Chem Soc. 1948;70:3436–3439. doi: 10.1021/ja01190a064. [DOI] [PubMed] [Google Scholar]
- 13.Mclamore WM, Celmer WD, Bogert VV, Pennington FC, Sobin BA, Solomons IA. Structure and synthesis of a new, thiazolidone antibiotic. J Am Chem Soc. 1953;75:105–108. doi: 10.1021/ja01097a030. [DOI] [Google Scholar]
- 14.Vinay V, Lashika K. A review on antimicrobial activity of 4-thiazolidinone derivatives. Int J Res Pharm Sci. 2011;1(1):17–27. [Google Scholar]
- 15.Farhat MF, El-Saghier AMM, Makhlouf MA, Kreddan KM, Elmezoughi AB, Ketene S. S-acetals in heterocyclic synthesis: part 1: synthesis of N-phenyl-2-ylidene and 2,5-diylidene-4-thiazolidinone derivatives. J Sulfur Chem. 2007;28:563–572. doi: 10.1080/17415990701586823. [DOI] [Google Scholar]
- 16.The crystallographic data for thiazolidinone 6 (CCDC 1551169) can be obtained on request from the director, Cambridge Crystallographic Data Center, 12 Union Road, Cambridge CB2 1EW, UK http://www.ccdc.cam.ac.uk/data_request/cif
- 17.Sheldrick GM. A short history of SHELX. Acta Cryst. A. 2008;64:112–122. doi: 10.1107/S0108767307043930. [DOI] [PubMed] [Google Scholar]
- 18.Spek AL. Structure validation in chemical crystallography. Acta Cryst D. 2009;65:148–155. doi: 10.1107/S090744490804362X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Bruker . APEX2 and SAINT. Madison: Bruker AXS Inc.; 2009. [Google Scholar]
- 20.Pihlaja K, Koleinpeter E, editors. Carbon-13 chemical shifts in structural and stereochemical analysis. Deerfield Beach: VCH Publishers; 1994. [Google Scholar]
- 21.Mosmann T. Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods. 1983;65:55–63. doi: 10.1016/0022-1759(83)90303-4. [DOI] [PubMed] [Google Scholar]
- 22.Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JAJ, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Peters-son GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Cliford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA. Gaussian 03, Revision C.02. Wallingford: Gaussian Inc; 2004. [Google Scholar]
- 23.GaussView, Version 4.1, Dennington II, Roy; Keith, Todd; and Millam, John; Semichem, Inc., Shawnee Mission, KS, 2007
- 24.Zhurko GA, Zhurko DA (2005) Chemcraft. Lite Version Build 08. http://www.chemcraftprog.com/. Accessed 1 Apr 2005
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The crystallographic data for this compound can be obtained free of charge from the Cambridge Crystallographic Data Centre via http://www.ccdc.cam.ac.uk/data_request/cif.