Abstract
Background
Filamentous fungi secret hydrolytic enzymes like cellulase and hemicellulase outside the cells, serving as important scavengers of plant biomass in nature and workhorses in the enzyme industry. Unlike the extensive study on the mechanism of cellulase production in fungi, research on spatiotemporal distribution and secretion of cellulase in fungi is lacking, retarding the deeper understanding of the molecular mechanism behind the fungal cellulase production.
Result
Recombinant Trichoderma reesei strains RBGL, RCBH, and RCMC were successfully constructed from T. reesei RUT-C30, expressing red fluorescent protein DsRed-tagged versions of β-glucosidase (BGL), cellobiohydrolase (CBH), and endoglucanase (CMC), respectively. With the assistance of these strains, we found that all three cellulase components BGL, CBH, and CMC diffused throughout the whole fungal mycelium with major accumulation at the hyphal apexes. These enzymes located in ER, Golgi, vacuoles and cell membrane/wall, but not septum, and secreted abundantly into the culture medium. Moreover, the major secretion of CBH and CMC started more early than that of BGL. Brefeldin A (BFA) completely blocked cellulase expression and secretion in T. reesei.
Conclusion
Based on recombinant T. reesei RBGL, RCBH, and RCMC expressing DsRed-fused versions of BGL, CBH, and CMC, respectively, the distribution and secretion of cellulase production in T. reesei were first visualized directly in a dynamic way, preliminarily mapping the location and secretion of T. reesei cellulase and providing evidence for revealing the secretion pathways of cellulase in T. reesei. The obtained results suggest that cellulase excretion majorly occurs via the conventional ER–Golgi secretory pathway, and might be assisted through unconventional protein secretion pathways.
Electronic supplementary material
The online version of this article (10.1186/s13068-019-1538-0) contains supplementary material, which is available to authorized users.
Keywords: Protein secretion, Cellulolytic enzymes, Unconventional secretion pathway, Subcellular localization
Background
Filamentous fungi are evolved to possess efficient protein secretion and are widely used for industrial enzyme production [1]. For example, filamentous fungi like Trichoderma, Penicillium, and Aspergillus are extraordinary in their capability of cellulase/hemicellulase secretion for the efficient deconstruction of recalcitrant polymers in plant biomass including cellulose, hemicellulose, pectin, and lignin [2–4]. Nevertheless, the molecular mechanism remains unknown about the secretory pathways for extracellular enzymes in filamentous fungi. It would benefit the basic understanding of fungal cell biology and improvement of production strains if the sorting, targeting, and release of extracellular proteins of fungi are fully investigated.
Protein secretion in filamentous fungi can occur either by the conventional pathway traversing the endomembrane system or via other alternative routes termed as unconventional secretions [5]. Like other eukaryotic cells, the classical secretory pathway in filamentous fungi is the well-established endoplasmic reticulum (ER)–Golgi pathway. Proteins first enter the ER for folding, modification by glycosylation, disulfide bridge formation, phosphorylation, and subunit assembly before being packaged into transport vesicles for delivery to Golgi compartments. Once in Golgi, the proteins undergo further glycosylation by the addition and/or removal of specific sugar resides, and then are again packed in secretory vesicles and directed to the endosomal/vacuolar system, the Spitzenkörper, or the plasma membrane for exocytosis. In filamentous fungi, exocytosis majorly occurs at growing hyphal apexes where Golgi and ER compartments are accumulated abundantly [5]. Nevertheless, exocytosis at other sites of fungal cells, such as septa [6] and lateral plasma membrane [7, 8], has been observed. Besides the conventional pathway, the unconventional traffic pathways for protein secretion in fungi have also been reported, including Golgi-independent secretion which was BFA-insensitive [9], secretion independent of both ER and Golgi apparatus [10], and extracellular vesicle secretion [11]. For example, Stock et al. reported that the secretion of endochitinase in Ustilago maydis is through the unconventional secretion pathway, independent of the ER and the Golgi-like apparatus [10]. However, the mechanism for this unconventional pathway was not very clear.
Trichoderma reesei is of particular interest because of its remarkable cellulase secretion capability with the extracellular cellulase reaching up to 100 g/L [12]. Meanwhile, it is a model organism for studying the mechanism of cellulase production and secretion, as well as an attractive host for the production of heterologous proteins. Electron microscopic analysis of ER structure in T. reesei RUT-C30 and QM6a is probably the earliest study on the protein secretory pathway of T. reesei [13]. Since then, efforts have been put on imaging and kinetics measurement of the secretion process [13–16], molecular characterization of the secretory pathway components [8, 17–22], and investigation of the relevant transcriptional regulation [23, 24]. The localization and secretion of cellulase has been studied by measurement of cellulase activity in subcellular fractionation [25], electron microscopy [13], and immunoelectron microscopy [26]. Nevertheless, fluorescence microscopy combined with fluorescence protein fusion [8] has yet to be exploited for investigating the spatiotemporal distribution and secretion of cellulase, a powerful strategy capable of providing direct and clear visualization.
In this study, three major cellulase components BGL, CBH, and CMC fused to red fluorescence protein DsRed, respectively, were successfully expressed in T. reesei RUT-C30, resulting in recombinant strains RBGL, RCBH, and RCMC. This allowed the visualization of spatiotemporal distribution and secretion of cellulase in living T. reesei by confocal laser scanning microscopy (CLSM) and fluorescence microplate reader. Moreover, the kinetics of cellulase production and the effect of BFA on cellulase production were investigated in these T. reesei mutants.
Materials and methods
Strains, plasmids, and culture conditions
Escherichia coli DH5α was used for plasmid construction and propagation. T. reesei RUT-C30 (CICC 13052) was utilized for RNA and DNA extraction, and plasmid transformation. E. coli DH5α and Agrobacterium tumefaciens AGL-1 were grown in LB medium with 220 rpm at 37 °C and 28 °C, respectively. T. reesei was grown at 28 °C with 200 rpm on potato dextrose agar (PDA) plate for conidial culture. Plasmid pDht/sk was given as a gift from Professor Zhihua Zhou from Key Laboratory of Synthetic Biology, Shanghai [27]. The primers used in this study are listed in Additional file 1: Table S1. All chemicals used in this study were purchased from Sigma-Aldrich (Sigma-Aldrich, USA).
Construction of recombinant T. reesei strains
The total RNA of T. reesei RUT-C30 was extracted with the RNA extraction Kit from Omega Bio-Tek (GA, USA) following the manufacturer’s instruction. The first-strand cDNA was synthesized from RNA of RUT-C30 using HiScript 1st Strand cDNA Synthesis Kit (Vazyme, China). Then, genes cel3a (BGL, M419DRAFT_136547/TRIREDRAFT_76672), cel7a (CBH, M419DRAFT_125125/TRIREDRAFT_123989), and cel7b (CMC, M419DRAFT_5304/TRIREDRAFT_122081) were amplified using cDNA of T. reesei RUT-C30 as template with primers listed in Additional file 1: Table S1, and cloned into the backbone of plasmid p-DsRed at XbaI loci using the ClonExpress II One-Step Cloning Kit (Vazyme, China) leading to the plasmid pBGL-DsRed, pCBH-DsRed, and pCMC-DsRed (Additional file 1: Fig. S1). Plasmid p-DsRed was constructed by inserting gene DsRed into the plasmid pDht/sk at XbaI loci. The XbaI loci at the end of gene DsRed on plasmid p-DsRed was mutated, leaving single XbaI for further cloning. These four plasmids were transformed into T. reesei RUT-C30 by A. tumefaciens-mediated transformation (AMT) method [28] using hygromycin B as a marker, resulting in recombinant strains RBGL, RCBH, RCMC, and RD, respectively. After transferring and selection, single spore colonies of these mutants were isolated for further study. The insertion of targeted genes into T. reesei genome was confirmed by PCR amplification using primers shown in Additional file 1: Table S2 and the results are shown in Additional file 1: Fig. S2.
Shake flask cultivation
For cellulase production, the conidial suspension (0.5 mL, 107/mL) was inoculated into a 50 mL Erlenmeyer flask-containing 10 mL sabouraud dextrose broth (SDB) and incubated for 48 h at 28 °C with 200 rpm. The culture was then transferred into a 250 mL flask-containing 50 mL TMM media (pH 6.0) [29] with 2% (w/t) cellulose or other carbon sources as indicated in the context for cellulase production, and cultivated at 28 °C with 200 rpm. 0.5 mL cell culture was sampled at different time points as indicated in the text, and centrifuged at 14,000g for 10 min at 4 °C. The supernatants were filtrated with 0.45 µm filter membrane and stored at − 80 °C for cellulase activity assays and fluorescence intensity measurement. The TMM medium was prepared as followed (all concentrations in g/L unless otherwise noted): urea, 1.00; (NH4)2SO4, 4.00; KH2PO4, 6.59; FeSO4 * 7H2O, 0.005; MnSO4 * H2O, 0.0016; ZnSO4 * 7H2O, 0.0014; CoCl2 * 6H2O, 0.002; MgSO4, 0.60; CaCl2, 0.60; Tween-80, 0.0186% (v/v); tryptone, 0.75; yeast extract, 0.25; maleic acid, 11.6; cellulose 2% (w/v), or other carbon sources as indicated in the context. The pH of TMM was adjusted to 6.0 by NaOH.
For BFA treatment, the cultures grown in TMM + 2% cellulose for 4 days were further diluted 1/10 into fresh TMM + 2% cellulose + 50 μg/mL BFA. The diluted cell cultures were cultivated at 28 °C with 200 rpm. Samples were taken at different time points as indicated in the text, and centrifuged at 8000 rpm for 30 min for the separation of the mycelia and the supernatant. The obtained mycelia were used for confocal observation, while the supernatants were utilized for fluorescence intensity detection and cellulase activity assays.
Analysis methods
Cellulase activity assays were carried out by following our previous studies [30–32]. Fluorescence intensity measurement of the filtered cell culture supernatant was performed using a Hitachi 2000 fluorescence spectrophotometer (Hitachi Ltd., Japan) with a 650 nm emission filter. The excitation wavelength was 540 nm.
Confocal imaging of fungal mycelia
Mycelia of T. reesei RUT-C30 and its derivatives were loaded onto a glass slide, covered with a cover glass, and observed on an inverted confocal laser scanning microscope SP8 (Leica, Germany) with a 100 × 1.4 NA oil-immersion objective. The red fluorescence protein DsRed was imaged with the excitation wavelength of 552 nm and the emission wavelength of 570–700 nm. The green fluorescence of the cell membrane/wall dye GC-Chol-PEG-FITC developed in our lab [33] was observed at 500–550 nm with the excitation length of 488 nm.
Preparing protoplasts of T. reesei RBGL for confocal imaging
The preparation of T. reesei protoplast performed according to the published method [34]. Briefly, 5 mL cell cultures of T. reesei RBGL grown in TMM + 2% cellulose for 72 h were centrifuged for 20 min at 8000 rpm to obtain the fungal mycelium, which was resuspended with 6 mL of lysing enzymes from Trichoderma harzianum (L1412, Sigma-Aldrich, USA). 675 mg lysing enzyme was dissolved in 15 mL 1.2 M MgSO4/10 mM sodium phosphate, pH5.8. The mixture was incubated at 25 °C with 90 rpm for 3–4 h. The release of protoplasts during the incubation was monitored on a microscope by counting the protoplasts using a counting chamber, while the fluorescence of the protoplast sample was detected on an inverted confocal laser scanning microscope SP8 with a 100× oil-immersion objective.
Confocal imaging of fungal ER and Golgi
ER-Green (DIOC6(3)) (KeyGEN BioTECH Co. Ltd., China) was prepared as a stock solution of 1 mM in DMSO and stored at − 20 °C in dark. Fungal mycelia were washed with Hank’s balanced salt solution (HBSS) buffer for three times and resuspended in 1 mL HBSS buffer. The pre-warmed ER-Green stock solution at 37 °C was added into the suspension at the ratio of 1:1000 and incubated at 37 °C for 20 min under dark condition. Then, the culture was washed with HBSS for two times and resuspended in HBSS buffer before being detected on an inverted confocal laser scanning microscope SP8 with a 100× oil-immersion objective. The excitation wavelength was 488 nm, and the emission was checked in the range of 498–600 nm.
Golgi-Green (C6-NBD-CERAMIDE) (KeyGEN BioTECH Co. Ltd., China) was used to stain the Golgi. Cell culture was washed with HBSS buffer for three times before being resuspended in HBSS buffer. Golgi-Green (5 µM) and BSA (5 µM) were added into the cell suspension followed by incubation at 4 °C for 30 min. The cell suspension was washed with pre-cooled HBSS and incubated at 37 °C for 30 min. Then, the samples were washed with HBSS for two times and observed by fluorescence microscopy as performed for ER staining.
Results
Cellulase is located throughout the whole hyphal cell with tip-high gradient, but not septum
Expression of cellulases BGL, CBH, and CMC as a fusion protein with red fluorescence protein DsRed under a modified CBH1 promoter (Fig. 1a) in T. reesei RUT-C30 was carried out, leading to recombinant strains RBGL, RCBH, and RCMC, respectively. Expression of DsRed alone in strain RUT-C30 was also performed as a control, which was designated as strain RD. We obtained 5, 2, 10, and 2 transformants for RBGL, RCBH, RCMC, and RD, respectively. The cellulase activity of these mutant strains was measured on day 5 using cellulose as the only carbon source (Additional file 1: Fig. S3), while confocal laser scanning microscopy (CLSM) was applied to check whether there was red fluorescence inside the cells. RBGL-1, RCBH-6, RCMC-8, and RD-1 were chosen for further study according to two rules: their cellulase activities were not affected notably in comparison with the parent strain RUT-C30 (Fig. 1b and Additional file 1: Fig. S3) and red fluorescence was observed in the fungal cells under CLSM (Fig. 1c and Additional file 1: Fig. S4). From here, strains RBGL-1, RCBH-6, RCMC-8, and RD-1 were named as RBGL, RCBH, RCMC, and RD, respectively.
Fig. 1.
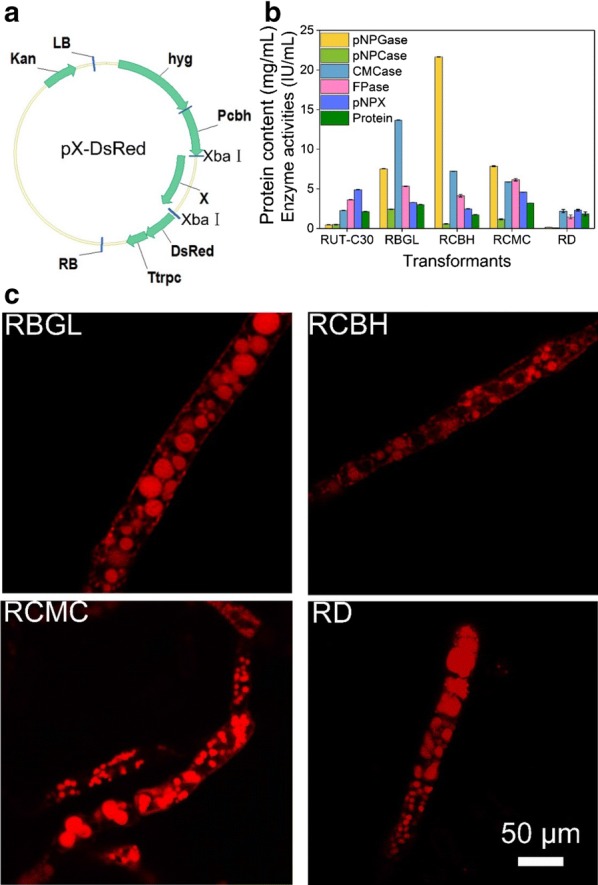
a Schematic illustration of the plasmid pX-DsRed (X = CEL3A, CEL7A, or CEL7B). Kan: kanamycin resistance; LB, left border of the binary vector; RB, right border of the binary vector; Pcbh, a modified CBH1 promoter; Ttrpc, Aspergillus nidulans trpC terminator; hyg, hygromycin B phosphotransferase gene. b Cellulase activities of T. reesei RUT-C30, RBGL, RCBH, RCMC, and RD grown on TMM + 2% cellulose for 5 days. pNPGase: the β-glucosidase activity; pNPCase: the CBH activity; CMCase: the CMC activity; FPase: the filter paper activity. Data are represented as the mean of three independent experiments and error bars express the standard deviations. c Observation of the localization of BGL-DsRed, CBH-DsRed, CMC-DsRed, and DsRed in T. reesei strains RBGL, RCBH, RCMC, and RD, respectively, grown on cellulose for 5 days by STED
The cellular distribution of the three major cellulases was investigated by stimulated emission depletion microscopy (STED) using recombinant strains cultured in TMM containing 2% cellulose for 5 days (Fig. 1c). Significant red fluorescent punctuates were observed throughout the cytoplasm of strains RBGL, RCBH, and RCMC, showing that all three cellulases BGL CBH, and CMC were present abundantly in secretory vesicles/vacuoles of the cytoplasm (Fig. 1c). Interestingly, we also observed red fluorescence on the cell membrane/wall of the three recombinant strains RBGL, RCBH, and RCMC, while no fluorescence was found on the cell membrane of the control strain RD. In line with this, all the hyphae of stains RBGL, RCBH, and RCMC displayed red fluorescence under low magnification due to the distribution of the fusion proteins BGL-DsRed, CBH-DsRed, and CMC-DsRed on cell membrane/wall (Additional file 1: Fig. S4), while RD did not as the fluorescence protein DsRed alone cannot located on the cell membrane/wall. It seems that the three major cellulases BGL, CBH, and CMC were all located on the cell membrane/wall, which is consistent with previous findings [14, 35–37].
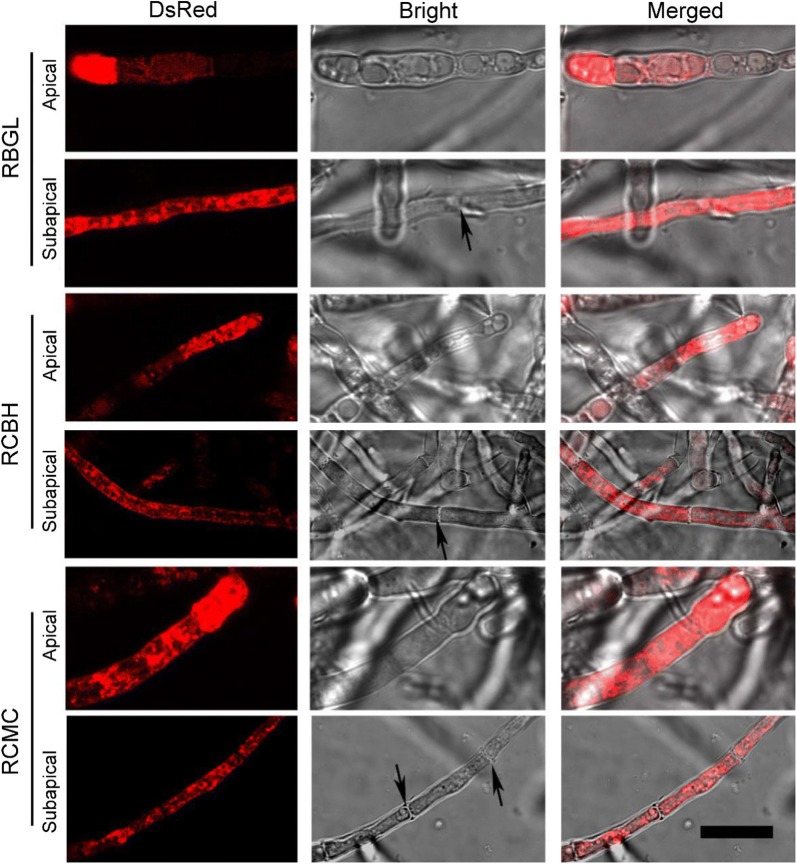
In all three recombinant strains, strong fluorescence was found both at the hyphal tip and distal hyphae regions with brighter fluorescence at the hyphal apexes (Fig. 2). It is probably that all three cellulases accumulated abundantly at the Spitzenkorper of the hyphal tips, indicating that the bulk flow accompanied with polarized apical growth of hyphae is main secretory pathway for cellulase. However, no red fluorescence was obtained at the septa of the three recombinant strains, suggesting that cellulase was not located on the hyphae septa. That all three tested cellulases were distributed throughout the whole hyphae except septum would enable their effective excretion from the fungal cells. This may partially explain the high extracellular cellulase production of RUT-C30. It seems that a complementary mechanism for protein secretion to the bulk flow accompanied with polarized apical growth of hyphae is adapted by T. reesei to release cellulase into the culture medium.
Fig. 2.
Cellular localization of BGL-DsRed, CBH-DsRed, and CMC-DsRed at the apical and subapical regions of the recombinant strains RBGL, RCBH, and RCMC, respectively, grown in TMM + 2% cellulose for 4 days. The black arrowheads indicate the septum (Scale bar = 15 μm)
BGL is located on cell membrane but not cell wall
To further confirm that BGL is distributed on cell membrane/wall, cholesterol-PEG-FITC, a cell membrane/wall dye synthesized in our lab [33], was employed to stain the cell membrane of T. reesei RUT-C30, RBGL and RD (Fig. 3a). The red fluorescent signal on cell membrane/wall of strain RBGL colocalized well with green fluorescence of cholesterol-PEG-FITC, resulting in yellow fluorescence on the cell membrane/wall of strain RBGL as observed in the merged figure of strain RBGL. In contrast, no yellow fluorescence was observed for strain RD, although green fluorescence was found on cell membrane/wall and red fluorescence was obtained in the cytoplasm. This further evidenced that protein BGL was localized on the cell membrane/wall. Nevertheless, whether BGL is on cell membrane or on cell wall is still unknown, as it is so difficult to differentiate cell membrane from cell wall. Therefore, protoplast of strain RBGL cultivated in TMM + 2% cellulose for 3 days was obtained and stained with cholesterol-PEG-FITC (Fig. 3b). In the plasmolysis cells of strain RBGL (Fig. 3b), the overlap of cholesterol-PEG-FITC with BGL-DsRed was reduced notably with only a few regions displaying yellow fluorescence as compared to the untreated mycelium cells of strain RBGL (Fig. 3a). In the enlarged image of one plasmolysis cell which was undergoing plasmolysis, the red fluorescence was being separated from green fluorescence along with the dividing of the cell membrane from the cell wall, while yellow fluorescence was still observed on the regions where plasmolysis did not occur. This result strongly supports that protein BGL is distributed on cell membrane rather than cell wall.
Fig. 3.
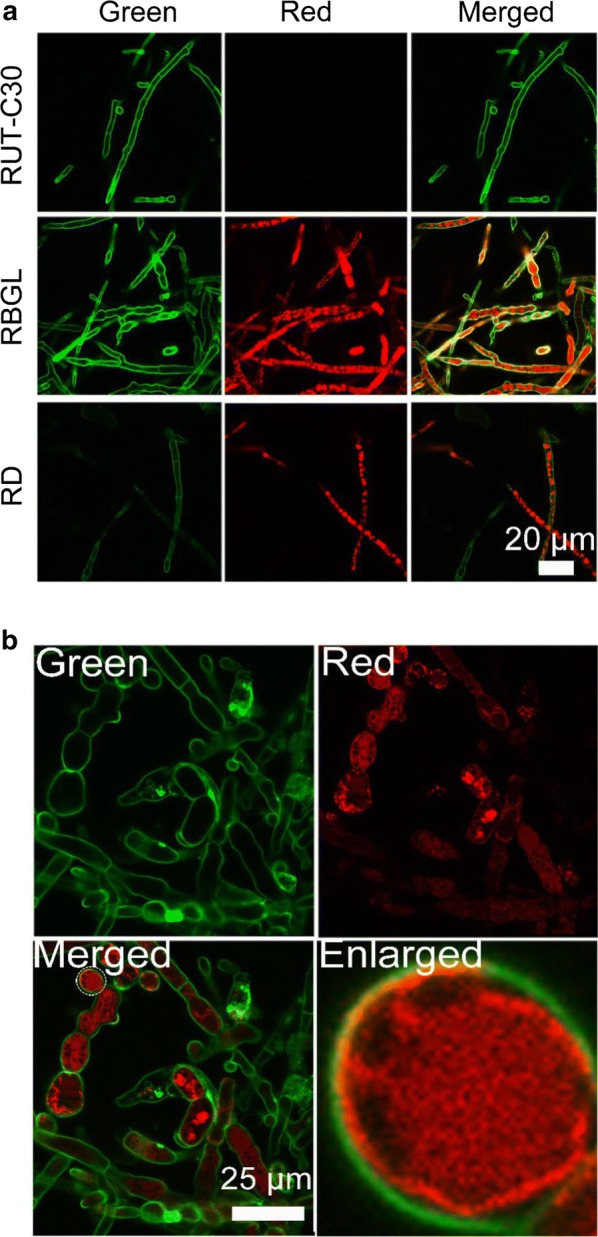
a Confocal images of strains RUT-C30, RBGL, and RD (Red) cultured for 5 days and stained with cholesterol-PEG-FITC (Green). b Confocal images of the protoplasts of strain RBGL (Red) stained with cholesterol-PEG-FITC (Green)
The cellular distribution of cellulase
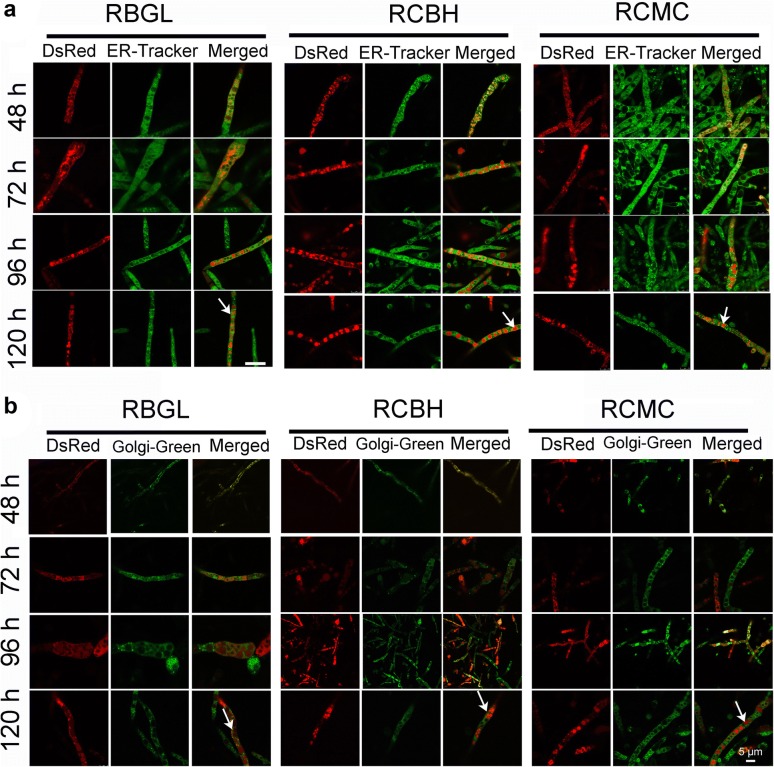
To know exactly the subcellular distribution of cellulases BGL, CMC, and CBH inside the hyphae cells, recombinant strains RBGL, RCBH, and RCMC grown at different time points during the cellulase production was treated with ER-tracker and Golgi-Green to stain ER (Fig. 4a) and Golgi (Fig. 4b), respectively. Green fluorescence from ER-tracker was observed throughout the whole hyphae, suggesting that T. reesei has well-developed ER network, which is consistent with the previous finding that the growing mycelia of T. reesei RUT-C30 were richly endowed with ER [38]. On the other hand, green fluorescence was observed when the three recombinant strains were exposed to Golgi-Green for Golgi-staining, demonstrating that T. reesei has Golgi structure as found by the previous study [39]. Nevertheless, this result contradicts with previous reports that no Golgi-like bodies were found in T. reesei RUT-C30 [26, 38]. Partial overlap of all three tested cellulase components BGL-DsRed, CBH-DsRed, and CMC-DsRed with ER or Golgi was found, as indicated by the observation of yellow fluorescence (Fig. 4a, b). However, the co-localization of BGL-DsRed, CBH-DsRed, and CMC-DsRed with the Golgi structure was poor (Fig. 4b), which coincided with the early finding that the labeling of endoglucanase I in Golgi-like structure was bad [39]. Interestingly, BGL-DsRed, CBH-DsRed, and CMC-DsRed accumulated largely in the lumen of globular vacuoles in addition to ER and Golgi (Fig. 4), which was in line with the result obtained by STED (Fig. 1c). Both xylanase II (XYN II) [26] and endoglucanase I [39] have been found in vacuoles. It is supposed that the cellulase secretion by T. reesei might involve vacuoles [26, 38]. Another hypothesis is that these hydrolytic enzymes are stored in the vacuoles in a poorly active form to avoid the deleterious effect of the hyperproduction of cellulase in T. reesei cells [38]. However, more evidences from experiments are required to support these views.
Fig. 4.
Confcal images of strains RBGL, RCBH, and RCMC stained with ER-Tracker (a) and Golgi-Green (b), respectively. The white arrowheads point out the vacuoles
Monitoring kinetic of cellulase production
Using the recombinant strains RBGL, RCMC, and RCBH, real-time monitor of cellulase production in live T. reesei was performed inside the cells by CLSM (Fig. 5a) and in the supernatant via spectroscopic analysis (Fig. 5b). At 48 h, red fluorescence was clearly observed in the cytoplasm of the hyphae cells of the three mutants, suggesting that BGL-DsRed, CBH-DsRed, and CMC-DsRed were expressed (Fig. 5a). The red fluorescence was strong in the cytoplasm of the three mutant hyphae cells at different time points during the whole fermentation process (from 48 to 168 h) with an increased fluorescence intensity observed from 96 to 144 h (Fig. 5a). In agreement with this finding, the fluorescence intensity in the supernatant of strain RCMC and RCBH remained almost unchanged from 48 to 72 h, followed by a notable increase from 72 to 144 h, while the fluorescence intensity in the supernatant of strain RBGL was not remarkably increased until 96 h. No fluorescence was detected in the supernatant of strain RD during the whole process. Obviously, there is good correlation between the cellulase expression in the hyphae and the cellulase secretion into the culture medium. The effective secretion of CMC and CBH began in the midterm of the fermentation, while the efficient release of BGL started in the late stage.
Fig. 5.
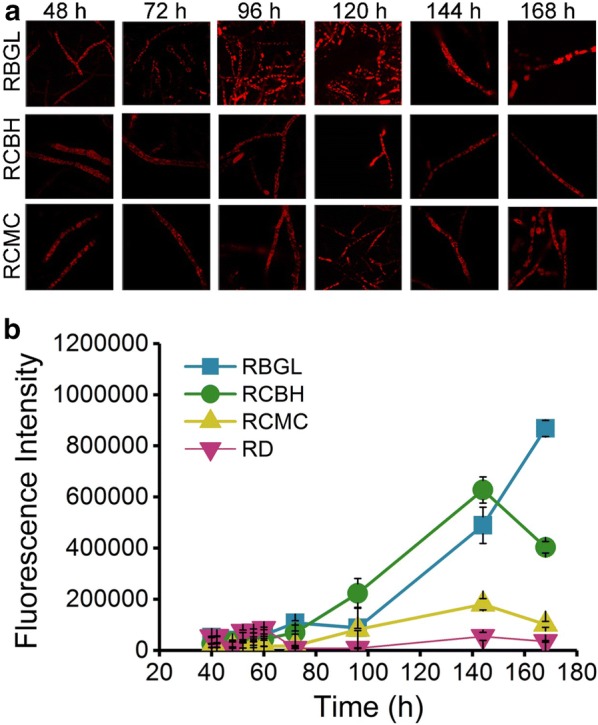
Confocal images of mycelia (a) and fluorescence intensity of the supernatants (b) from the recombinant strains RBGL, RCBH, and RCMC during the cellulase fermentation. The excitation wavelength was 552 nm
BFA inhibits secretion of total cellulase in T. reesei
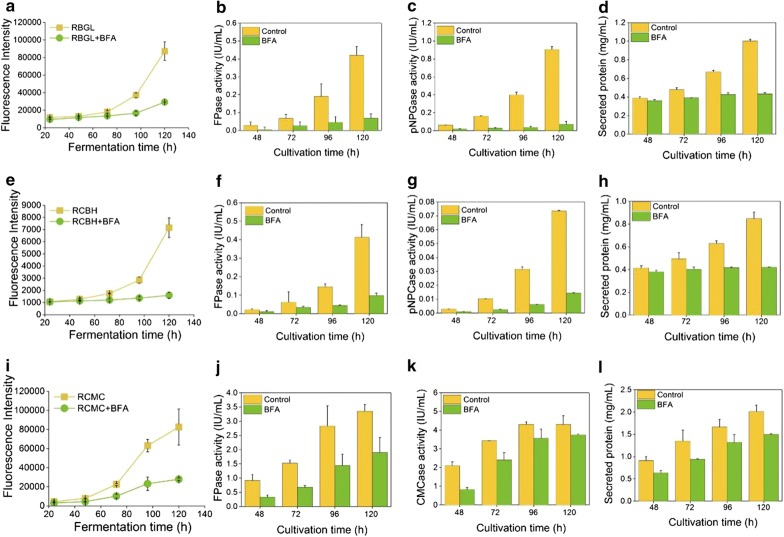
Recombinant strains RBGL, RCMC, and RCBH were exposed to 10 and 50 μg/mL Brefeldin A (BFA), a compound primarily inhibiting vesicle trafficking between ER and Golgi apparatus in fungi [40]. In the presence of 10 μg/mL BFA, red fluorescence in the mycelia of all three recombinant strains was almost unchanged as compared to the untreated samples (Fig. 6 and Additional file 1: Fig. S5), indicating that the cellulase expression was not affected by 10 μg/mL BFA. Nevertheless, the red fluorescence in the corresponding supernatant samples was neglected and remained almost unchanged throughout the whole cellulase production process (Fig. 7a, e, i), suggesting that cellulase secretion was hampered. Accordingly, the cellulase activities and protein concentration in the supernatant of the three recombinant strains were reduced sharply with the treatment of 10 μg/mL BFA (Fig. 7). When the concentration of BFA was increased to 50 μg/mL, red fluorescence in all three mutants was almost abolished, implying that the cellulase expression was inhibited significantly (Fig. 6). Meanwhile, the red fluorescence in the supernatant of the 50 μg/mL BFA-treated strains RBGL, RCMC, and RCBH stayed unchanged during the cellulase production process (Additional file 1: Fig. S6), suggesting that the cellulase secretion was also retarded by BFA. Taken together, BFA inhibited notably cellulase secretion in T. reesei, and even the cellulase expression at high concentration, both leading to notably reduced cellulase production.
Fig. 6.
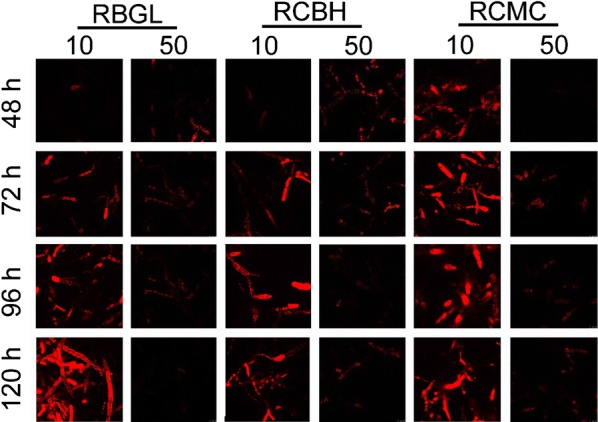
Confocal images of recombinant strains RBGL, RCBH and RCMC cultured in TMM + 2% cellulose + 10 or 50 μg/mL BFA for 48, 72, 96 and 120 h
Fig. 7.
Fluorescence intensity (a, e, i), FPase (b, f, j), pNPGase activity (c), pNPCase activity (g), CMCase activity (k), and secreted protein concentration (d, h, l) in the culture supernatants of recombinant strains RBGL, RCBH, and RCMC cultured in TMM + 2% cellulose + 10 μg/mL BFA for 48, 72, 96, and 120 h
Discussion
Using the industrial strain T. reesei RUT-C30 as the parent strain, we have successfully constructed recombinant strains RBGL, RCBH, and RCMC in which cellulase BGL, CBH, and CMC were fused to red fluorescence protein DsRed. In this way, the localization and secretion of cellulase in living T. reesei were first investigated directly in a dynamic manner by fluorescence imaging method using CLSM and fluorescence microplate reader. Although it presents a well-established, potent way to analyze the spatiotemporal distribution of protein in microorganism study, fluorescence microscopy combined with fluorescence protein fusion is not fully implicated in the study of T. reesei cellulase. Traditionally, the localization and secretion of cellulase has been studied by cellulase activity measurement in subcellular fractionation [23], electron microscopy [13, 38], and immunoelectron microscopy [14, 26, 37]. This is probably due to that the genetic engineering platform of T. reesei is not as well developed as the model microorganisms like Escherichia coli and Saccharomyces cerevisiae. Therefore, it is complicated and cumbersome to perform the strain construction required for fluorescence imaging. It is worth noting that several newly minted of genetic tools for T. reesei have been developed in recent years, including CRISPR/Cas9 [41] and I-SceI-mediated genetic box [42, 43]. With these advances of genetic tools in T. reesei, the fluorescence imaging can definitely become a powerful and common way to investigate spatiotemporal localization and secretion of T. reesei, as demonstrated in this study.
Although the mechanism of cellulase production in fungi has been extensively studied [44], few research on spatiotemporal distribution of cellulase in fungi has been reported, retarding the deeper understanding of the molecular mechanism behind the fungal cellulase production. Here, we found that three major cellulase components BGL, CMC, and CBH were distributed throughout the whole fungal hypha except septa, which is consistent with previous findings [14, 35–38, 45, 46]. By indirect immunofluorescence microscopy and labeled cryosections, protein CBHI was found throughout the whole mycelium including vesicles and hyphal walls, but not on septa [14]. Endoglucanases were found in both hyphal tips and hyphal walls as found by immunoelectron microscopy [37]. Accumulation of CMC in cytosol, vacuoles, and ER of T. reesei was observed in another study using electron microscopy [38]. BGLs have been reported to be extracellular [45], plasma-membrane-bound [36], and intracellular [46]. However, these early studies are based on processed fungal cell samples without studying the localization of cellulase in living T. reesei in real time and in a dynamic way, which instead was achieved in this study. Moreover, for the first time, we demonstrated that BGL1 was located on cell membrane but not cell wall. Owing to the inefficient separation of cell wall and cell membrane by the used techniques, the three major cellulases CMC, CBHI, and BGLI were generally reported to be cell-wall-bounded [14, 35–37].
BFA is an inhibitor of the traditional ER–Golgi secretory pathway, generally utilized to prevent vesicle budding from the Golgi. In many types of cells, the structure of the Golgi compartment was damaged in the presence of BFA [47]. The reported targets of BFA include the guanine nucleotide exchange factors of ARF proteins required for transport vesicle coat formation [48–50], and the BARS proteins that are linked to the integrity of Golgi structure [51]. Our finding that BFA totally blocked the expression and secretion of cellulase in T. reesei is in accordance with the early results. The short-term effect of BFA on the cellulase CBHI synthesis and transport in T. reesei was investigated at both protein and mRNA level [52]. The half-life of the cbh1 mRNA, the rate of CBHI synthesis, and CBHI transport were reduced significantly in the BFA-treated cultures, leading to that only a minute amount of the labeled CBHI with [35S] methionine was detectable in the culture medium at the late stages of the labeling experiment.
The tip-high gradient of intracellular cellulase (Fig. 2), the co-localization of intracellular cellulase with ER and Golgi (Fig. 4), and the complete inhibition of cellulase production by BFA (Fig. 7) all support the proposition that the cellulase secretion is majorly via the classical ER–Golgi secretory pathway. Meanwhile, the localization of cellulase throughout the whole fungal mycelia (Fig. 5) except septa, and the abundant accumulation of cellulase in vacuoles (Figs. 1c, 4) demonstrate that there are probably other unconventional protein secretion pathways for cellulase secretion. Recent studies unraveled several alternative protein secretory pathways to the conventional secretory pathway in fungi, such as Golgi-independent secretion which was BFA-insensitive [9], secretion independent of both ER and Golgi apparatus [10], and extracellular vesicle secretion [11]. Nevertheless, whether and how these unconventional pathways are involved in cellulase requires further extensive study.
Conclusion
For the first time, the spatiotemporally localization and secretion of cellulase was monitored directly in live T. reesei. From T. reesei RUT-C30, we have successfully constructed recombinant T. reesei RBGL, RCBH, and RCMC, which expressed DsRED-fused versions of BGL, CBH, and CMC, namely BGL-DsRed, CBH-DsRed, and CMC-DsRed. In these recombinant strains, BGL, CBH, and CMC was localized throughout the whole fungal mycelia with enormous accumulation at the growing apices, including ER, Golgi, vacuoles, and cell membrane/wall but not septa. In addition, the major secretion of CBH and CMC begun much earlier than that of BGL. In the presence of BFA, the cellulase expression and secretion was severely inhibited in T. reesei, resulting in neglectable fluorescence intensity and cellulase activities. All these results further support the view that cellulase excretion mainly occurs via the conventional ER–Golgi secretory pathway, as evidenced by the abundant accumulation of intracellular cellulase at the advancing tips (Fig. 2), the localization of intracellular cellulase in ER and Golgi (Fig. 4), and the inhibition of cellulase secretion by BFA (Figs. 6, 7), but might be assisted through unconventional protein secretion pathways as it was found that cellulase distributed throughout the whole fungal mycelia (Fig. 5a) except septa with the notable accumulation in vacuoles (Figs. 2, 4).
Additional file
Additional file 1: Fig. S1. Sequences of plasmid pX-DsRed. Pcbh, Ttrpc, hyg, linker, and XbaI were represented in light green, pink, dark green, blue and light red respectively; CEL3A, CEL7A, CEL7B and DsRed were marked in purple, yellow, dark red and grey, respectively; The LB and RB of the binary vector were underlined and in bold. Fig. S2. PCR conformation results for RUT-C30 and the transformants RCMC, RCBH and RBGL. Fig. S3. Cellulase activities of recombinant T. reesei strains RBGL (A), RCBH (B), RCMC (C), and RD (D). The mutants were grown in TMM + 2% cellulose for 5 days. pNPGase: the β-glucosidase activity; pNPCase: the CBH activity; CMCase: the CMC activity; FPase: the filter paper activity. Data are represented as the mean of three independent experiments and error bars expressed the standard deviations. Fig. S4. CLSM imaging of RBGL, RCBH, RCMC, and RD cultivated in TMM + 2% cellulose for 5 days using 10 × objective. Fig. S5. Confocal images of recombinant strains RBGL, RCBH and RCMC cultured in TMM + 2% cellulose + 100 μL or 500 μL DMSO for 48, 72, 96 and 120 h. Fig. S6. Fluorescence intensity (A, C, and E), and secreted protein concentration (B, D, and F) in the culture supernatants of recombinant strains RBGL, RCBH and RCMC cultured in TMM + 2% cellulose + 50 μg/mL BFA for 48, 72, 96 and 120 h. Table S1. PCR primers for plasmid construction. Table S2. PCR primers for confirmation of transformants.
Acknowledgements
We thank the National Natural Science Foundation of China (31700040) and the Fundamental Research Funds for the Central Universities.
Abbreviations
- pNPGase
the β-glucosidase activity
- pNPCase
the CBH activity
- CMCase
the CMC activity
- FPase
the filter paper activity
- pNPXase
the β-xylosidase activity
- AMT
Agrobacterium tumefaciens-mediated transformation
- PDA
potato dextrose agar
- SDB
sabouraud dextrose broth
- TMM
Trichoderma minimal medium
Authors’ contributions
CL and FL conceived and designed the study. CL carried out the majority of the experiments. AP, HY, and RL conducted part of enzyme activity measurement experiments and the detection of fluorescence intensity. ZZ provided the plasmid pXBthg and helped us to design and construct the plasmids used in this study. FW helped to analyze the data. CL and FL analyzed the data and drafted the manuscript. All authors read and approved the final manuscript.
Funding
This study is supported by the National Natural Science Foundation of China (31700040), and the Fundamental Research Funds for the Central Universities.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Chengcheng Li, Email: lichengcheng1162@126.com.
Ai-Ping Pang, Email: applepangtju@126.com.
Hang Yang, Email: 1019604761@qq.com.
Roujing Lv, Email: 2920925948@qq.com.
Zhihua Zhou, Email: zhouzhihua@sippe.ac.cn.
Fu-Gen Wu, Email: wufg@seu.edu.cn.
Fengming Lin, Phone: +86-83791810, Email: linfengming@seu.edu.cn.
References
- 1.Punt PJ, Nick VB, Ana C, Alwin A, Jeroen M, Cees VDH. Filamentous fungi as cell factories for heterologous protein production. Trends Biotechnol. 2002;20:200–206. doi: 10.1016/S0167-7799(02)01933-9. [DOI] [PubMed] [Google Scholar]
- 2.Martinez D, Berka RM, Henrissat B, Saloheimo M, Arvas M, Baker SE, Chapman J, Chertkov O, Coutinho PM, Cullen D, et al. Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn Hypocrea jecorina) Nat Biotech. 2008;26:553–560. doi: 10.1038/nbt1403. [DOI] [PubMed] [Google Scholar]
- 3.Chundawat SP, Beckham GT, Himmel ME, Dale BE. Deconstruction of lignocellulosic biomass to fuels and chemicals. Annu Rev Chem Biomol Eng. 2011;2:121–145. doi: 10.1146/annurev-chembioeng-061010-114205. [DOI] [PubMed] [Google Scholar]
- 4.Nakazawa H, Kawai T, Ida N, Shida Y, Kobayashi Y, Okada H, Tani S, Sumitani J, Kawaguchi T, Morikawa Y, et al. Construction of a recombinant Trichoderma reesei strain expressing Aspergillus aculeatus beta-glucosidase 1 for efficient biomass conversion. Biotechnol Bioeng. 2012;109:92–99. doi: 10.1002/bit.23296. [DOI] [PubMed] [Google Scholar]
- 5.Shoji J-Y, Kikuma T, Kitamoto K. Vesicle trafficking, organelle functions, and unconventional secretion in fungal physiology and pathogenicity. Curr Opin Microbiol. 2014;20:1–9. doi: 10.1016/j.mib.2014.03.002. [DOI] [PubMed] [Google Scholar]
- 6.Hayakawa Y, Ishikawa E, Shoji J-Y, Nakano H, Kitamoto K. Septum-directed secretion in the filamentous fungus Aspergillus oryzae. Mol Microbiol. 2011;81:40–55. doi: 10.1111/j.1365-2958.2011.07700.x. [DOI] [PubMed] [Google Scholar]
- 7.Shoji JY, Arioka M, Kitamoto K. Dissecting cellular components of the secretory pathway in filamentous fungi: insights into their application for protein production. Biotechnol Lett. 2008;30:7–14. doi: 10.1007/s10529-007-9516-1. [DOI] [PubMed] [Google Scholar]
- 8.Valkonen M, Kalkman ER, Saloheimo M, Penttilä M, Read ND, Duncan RR. Spatially segregated SNARE protein interactions in living fungal cells. J Biol Chem. 2007;282:22775–22785. doi: 10.1074/jbc.M700916200. [DOI] [PubMed] [Google Scholar]
- 9.Giraldo MC, Dagdas YF, Gupta YK, Mentlak TA, Yi M, Martinezrocha AL, Saitoh H, Terauchi R, Talbot NJ, Valent B. Two distinct secretion systems facilitate tissue invasion by the rice blast fungus Magnaporthe oryzae. Nat Commun. 2013;4:1996. doi: 10.1038/ncomms2996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Stock J, Sarkari P, Kreibich S, Brefort T, Feldbrugge M, Schipper K. Applying unconventional secretion of the endochitinase Cts1 to export heterologous proteins in Ustilago maydis. J Biotechnol. 2012;161:80–91. doi: 10.1016/j.jbiotec.2012.03.004. [DOI] [PubMed] [Google Scholar]
- 11.Oliveira DL, Juliana R, Joffe LS, Godinho RMC, Rodrigues ML. Where do they come from and where do they go: candidates for regulating extracellular vesicle formation in fungi. Int J Mol Sci. 2013;14:9581–9603. doi: 10.3390/ijms14059581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cherry JR, Fidantsef AL. Directed evolution of industrial enzymes: an update. Curr Opin Biotechnol. 2003;14:438–443. doi: 10.1016/S0958-1669(03)00099-5. [DOI] [PubMed] [Google Scholar]
- 13.Ghosh A, Al-Rabiai S, Ghosh BK, Trimiño-Vazquez H, Eveleigh DE, Montenecourt BS. Increased endoplasmic reticulum content of a mutant of Trichoderma reesei (RUT-C30) in relation to cellulase synthesis. Enzyme Microb Technol. 1982;4:110–113. doi: 10.1016/0141-0229(82)90093-X. [DOI] [Google Scholar]
- 14.Nykanen M, Saarelainen R, Raudaskoski M, Nevalainen K, Mikkonen A. Expression and secretion of barley cysteine endopeptidase B and cellobiohydrolase I in Trichoderma reesei. Appl Environ Microbiol. 1997;63:4929. doi: 10.1128/aem.63.12.4929-4937.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Pakula TM, Katri S, Jaana U, Merja P. The effect of specific growth rate on protein synthesis and secretion in the filamentous fungus Trichoderma reesei. Microbiology. 2005;151:135–143. doi: 10.1099/mic.0.27458-0. [DOI] [PubMed] [Google Scholar]
- 16.Pakula TM, Uusitalo J, Saloheimo M, Salonen K, Aarts RJ, Penttilä M. Monitoring the kinetics of glycoprotein synthesis and secretion in the filamentous fungus Trichoderma reesei: cellobiohydrolase I (CBHI) as a model protein. Microbiology. 1999;146(Pt 1):223–232. doi: 10.1099/00221287-146-1-223. [DOI] [PubMed] [Google Scholar]
- 17.Saloheimo M, Lund M, Penttilä ME. The protein disulphide isomerase gene of the fungus Trichoderma reesei is induced by endoplasmic reticulum stress and regulated by the carbon source. Mol Gen Genet. 1999;262:35–45. doi: 10.1007/s004380051057. [DOI] [PubMed] [Google Scholar]
- 18.Veldhuisen G, Saloheimo M, Fiers MA, Punt PJ, Contreras R, Penttilä M, Hondel CA. Isolation and analysis of functional homologues of the secretion-related SAR1 gene of Saccharomyces cerevisiae from Aspergillus niger and Trichoderma reesei. Mol Gen Genet. 1997;256:446–455. doi: 10.1007/pl00008613. [DOI] [PubMed] [Google Scholar]
- 19.Saloheimo M, Wang H, Valkonen M, Vasara T, Huuskonen A, Riikonen M, Pakula T, Ward M, Penttila M. Characterization of secretory genes ypt1/yptA and nsf1/nsfA from two filamentous fungi: induction of secretory pathway genes of Trichoderma reesei under secretion stress conditions. Appl Environ Microbiol. 2004;70:459–467. doi: 10.1128/AEM.70.1.459-467.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Vasara T, Keranen S, Penttila M, Saloheimo M. Characterisation of two 14-3-3 genes from Trichoderma reesei: interactions with yeast secretory pathway components. Biochim Biophys Acta Mol Cell Res. 2002;1590:27–40. doi: 10.1016/S0167-4889(02)00197-0. [DOI] [PubMed] [Google Scholar]
- 21.Vasara T, Saloheimo M, Keranen S, Penttila M. Trichoderma reesei rho3 a homologue of yeast RH03 suppresses the growth defect of yeast sec15-1 mutation. Curr Genet. 2001;40:119–127. doi: 10.1007/s002940100245. [DOI] [PubMed] [Google Scholar]
- 22.Vasara T, Salusjarvi L, Raudaskoski M, Keranen S, Penttila M, Saloheimo M. Interactions of the Trichoderma reesei rho3 with the secretory pathway in yeast and T. reesei. Mol Microbiol. 2010;42:1349–1361. doi: 10.1046/j.1365-2958.2001.02716.x. [DOI] [PubMed] [Google Scholar]
- 23.Saloheimo M, Valkonen M, Penttila M. Activation mechanisms of the HAC1-mediated unfolded protein response in filamentous fungi. Mol Microbiol. 2003;47:1149–1161. doi: 10.1046/j.1365-2958.2003.03363.x. [DOI] [PubMed] [Google Scholar]
- 24.Valkonen M, Penttila M, Saloheimo M. The ire1 and ptc2 genes involved in the unfolded protein response pathway in the filamentous fungus Trichoderma reesei. Mol Genet Genomics. 2004;272:443–451. doi: 10.1007/s00438-004-1070-0. [DOI] [PubMed] [Google Scholar]
- 25.Kubicek CP. Release of carboxymethyl-cellulase and β-glucosidase from cell walls of Trichoderma reesei. Eur J Appl Microbiol Biotechnol. 1981;13:226–231. doi: 10.1007/BF00500103. [DOI] [Google Scholar]
- 26.Kurzątkowski W, Solecka J, Filipek J, Rozbicka B, Messner R, Kubicek CP. Ultrastructural localization of cellular compartments involved in secretion of the low molecular weight, alkaline xylanase by Trichoderma reesei. Arch Microbiol. 1993;159:417–422. doi: 10.1007/BF00288587. [DOI] [Google Scholar]
- 27.Zou G, Shi SH, Jiang YP, van den Brink J, de Vries RP, Chen L, Zhang J, Ma L, Wang CS, Zhou ZH. Construction of a cellulase hyper-expression system in Trichoderma reesei by promoter and enzyme engineering. Microb Cell Fact. 2012;11:21. doi: 10.1186/1475-2859-11-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhong YH, Wang XL, Wang TH, Jiang Q. Agrobacterium-mediated transformation (AMT) of Trichoderma reesei as an efficient tool for random insertional mutagenesis. Appl Microbiol Biotechnol. 2007;73:1348–1354. doi: 10.1007/s00253-006-0603-3. [DOI] [PubMed] [Google Scholar]
- 29.Minty JJ, Singer ME, Scholz SA, Bae C-H, Ahn J-H, Foster CE, Liao JC, Lin XN. Design and characterization of synthetic fungal–bacterial consortia for direct production of isobutanol from cellulosic biomass. Proc Natl Acad Sci USA. 2013;110:14592–14597. doi: 10.1073/pnas.1218447110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Li C, Lin F, Li Y, Wei W, Wang H, Qin L, Zhou Z, Li B, Wu F, Chen Z. A β-glucosidase hyper-production Trichoderma reesei mutant reveals a potential role of cel3D in cellulase production. Microb Cell Fact. 2016;15:151. doi: 10.1186/s12934-016-0550-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Li C, Lin F, Zhou L, Qin L, Li B, Zhou Z, Jin M, Chen Z. Cellulase hyper-production by Trichoderma reesei mutant SEU-7 on lactose. Biotechnol Biofuels. 2017;10:228. doi: 10.1186/s13068-017-0915-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Li C, Lin F, Sun W, Yuan S, Zhou Z, Wu FG, Chen Z. Constitutive hyperproduction of sorbicillinoids in Trichoderma reesei ZC121. Biotechnol Biofuels. 2018;11:291. doi: 10.1186/s13068-018-1296-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang H-Y, Hua X-W, Jia H-R, Li C, Lin F, Chen Z, Wu F-G. Universal cell surface imaging for mammalian, fungal, and bacterial cells. ACS Biomater Sci Eng. 2016;2:987–997. doi: 10.1021/acsbiomaterials.6b00130. [DOI] [PubMed] [Google Scholar]
- 34.Laurila HO, Nevalainen H, Mäkinen V. Production of protoplasts from the fungi Curvularia inaequalis and Trichoderma reesei. Appl Microbiol Biotechnol. 1985;21:210–212. doi: 10.1007/BF00295124. [DOI] [Google Scholar]
- 35.Messner R, Kubicek CP. Evidence for a single, specific β-glucosidase in cell walls from Trichoderma reesei QM9414. Enzyme Microb Technol. 1990;12:685–690. doi: 10.1016/0141-0229(90)90008-E. [DOI] [Google Scholar]
- 36.Umile C, Kubicek CP. A constitutive, plasma-membrane bound β-glucosidase in Trichoderma reesei. FEMS Microbiol Lett. 1986;34:291–295. [Google Scholar]
- 37.Sprey B. Cellular and extracellular localization of endocellulase in Trichoderma reesei. FEMS Microbiol Lett. 2010;55:283–294. doi: 10.1111/j.1574-6968.1988.tb02816.x. [DOI] [Google Scholar]
- 38.Glenn M, Ghosh A, Ghosh BK. Subcellular fractionation of a hypercellulolytic mutant, Trichoderma reesei Rut-C30: localization of endoglucanase in microsomal fraction. Appl Environ Microbiol. 1985;50:1137–1143. doi: 10.1128/aem.50.5.1137-1143.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ghosh BK, Ganguly T, Ghosh A. Ultrastructural aspects of cellulase biosynthesis in Trichoderma reesei. In: Kubicek CP, Eveleigh DE, Esterbauer H, Steiner W, KubicekPranz EM, editors. Trichoderma cellulases: biochemistry, physiology, genetics and applications. Cambridge: Royal Social Chemistry Press; 1990. pp. 115–138. [Google Scholar]
- 40.Cole L, Davies D, Hyde GJ, Ashford AE. Brefeldin A affects growth, endoplasmic reticulum, Golgi bodies, tubular vacuole system, and secretory pathway in Pisolithus tinctorius. Fungal Genet Biol. 2000;29:95–106. doi: 10.1006/fgbi.2000.1190. [DOI] [PubMed] [Google Scholar]
- 41.Liu R, Chen L, Jiang Y, Zhou Z, Zou G. Efficient genome editing in filamentous fungus Trichoderma reesei using the CRISPR/Cas9 system. Cell Discovery. 2015;1:15007. doi: 10.1038/celldisc.2015.7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Jean Paul O, Mark A, Igor N, Sharief B, Ram AFJ. I-SceI-mediated double-strand DNA breaks stimulate efficient gene targeting in the industrial fungus Trichoderma reesei. Appl Microbiol Biotechnol. 2015;99:10083–10095. doi: 10.1007/s00253-015-6829-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ouedraogo JP, Arentshorst M, Nikolaev I, Barends S, Ram AFJ. I-SceI enzyme mediated integration (SEMI) for fast and efficient gene targeting in Trichoderma reesei. J Biotechnol. 2016;222:25–28. doi: 10.1016/j.jbiotec.2016.02.012. [DOI] [PubMed] [Google Scholar]
- 44.Payne CM, Knott BC, Mayes HB, Hansson H, Himmel ME, Sandgren M, Ståhlberg J, Beckham GT. Fungal cellulases. Chem Rev. 2015;115:1308–1448. doi: 10.1021/cr500351c. [DOI] [PubMed] [Google Scholar]
- 45.Chen H, Hayn M, Esterbauer H. Purification and characterization of two extracellular β-glucosidases from Trichoderma reesei. Biochim Biophys Acta. 1992;1121:54–60. doi: 10.1016/0167-4838(92)90336-C. [DOI] [PubMed] [Google Scholar]
- 46.Inglin M, Feinberg BA, Loewenberg JR. Partial purification and characterization of a new intracellular β-glucosidase of Trichoderma reesei. Biochem. J. 1980;185:515–519. doi: 10.1042/bj1850515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Richard D, Klausner JGD, Jennifer L-S. Brefeldin A: insights into the control of membrane traffic and organelle structure. Int J Biochem Cell Biol. 1992;116:1071–1080. doi: 10.1083/jcb.116.5.1071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chardin P, McCormick F. Brefeldin A: the advantage of being uncompetitive. Cell. 1999;97:153–155. doi: 10.1016/S0092-8674(00)80724-2. [DOI] [PubMed] [Google Scholar]
- 49.Morinaga N, Adamik R, Moss J, Vaughan M. Brefeldin A Inhibited Activity of the Sec7 Domain of p200, a mammalian guanine nucleotide-exchange protein for ADP-ribosylation factors. J Biol Chem. 1999;274:17417–17423. doi: 10.1074/jbc.274.25.17417. [DOI] [PubMed] [Google Scholar]
- 50.Peyroche A, Antonny B, Robineau S, Acker J, Cherfils J, Jackson CL. Brefeldin A acts to stabilize an abortive ARF-GDP-Sec7 domain protein complex: involvement of specific residues of the Sec7 domain. Mol Cell. 1999;3:275–285. doi: 10.1016/S1097-2765(00)80455-4. [DOI] [PubMed] [Google Scholar]
- 51.Spanfò S, Silletta MG, Colanzi A, Alberti S, Fiucci G, Valente C, Fusella A, Salmona M, Mironov A, Luini A, et al. Molecular cloning and functional characterization of brefeldin A-ADP-ribosylated substrate: a novel protein involved in the maintenance of the Golgi structure. J Biol Chem. 1999;274:17705–17710. doi: 10.1074/jbc.274.25.17705. [DOI] [PubMed] [Google Scholar]
- 52.Pakula TM, Laxell M, Huuskonen A, Uusitalo J, Saloheimo M, Penttila M. The effects of drugs inhibiting protein secretion in the filamentous fungus Trichoderma reesei. Evidence for down-regulation of genes that encode secreted proteins in the stressed cells. J Biol Chem. 2003;278:45011–45020. doi: 10.1074/jbc.M302372200. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Fig. S1. Sequences of plasmid pX-DsRed. Pcbh, Ttrpc, hyg, linker, and XbaI were represented in light green, pink, dark green, blue and light red respectively; CEL3A, CEL7A, CEL7B and DsRed were marked in purple, yellow, dark red and grey, respectively; The LB and RB of the binary vector were underlined and in bold. Fig. S2. PCR conformation results for RUT-C30 and the transformants RCMC, RCBH and RBGL. Fig. S3. Cellulase activities of recombinant T. reesei strains RBGL (A), RCBH (B), RCMC (C), and RD (D). The mutants were grown in TMM + 2% cellulose for 5 days. pNPGase: the β-glucosidase activity; pNPCase: the CBH activity; CMCase: the CMC activity; FPase: the filter paper activity. Data are represented as the mean of three independent experiments and error bars expressed the standard deviations. Fig. S4. CLSM imaging of RBGL, RCBH, RCMC, and RD cultivated in TMM + 2% cellulose for 5 days using 10 × objective. Fig. S5. Confocal images of recombinant strains RBGL, RCBH and RCMC cultured in TMM + 2% cellulose + 100 μL or 500 μL DMSO for 48, 72, 96 and 120 h. Fig. S6. Fluorescence intensity (A, C, and E), and secreted protein concentration (B, D, and F) in the culture supernatants of recombinant strains RBGL, RCBH and RCMC cultured in TMM + 2% cellulose + 50 μg/mL BFA for 48, 72, 96 and 120 h. Table S1. PCR primers for plasmid construction. Table S2. PCR primers for confirmation of transformants.