Abstract
Viruses encounter many challenges within host cells in order to replicate their nucleic acid. In the case of DNA viruses, one challenge that must be overcome is recognition of viral DNA structures by the host DNA damage response (DDR) machinery. This is accomplished in elegant and unique ways by different viruses as each has specific needs and sensitivities dependent on its life cycle. In this review we focus on three DNA tumor viruses and their interactions with the DDR. The viruses Epstein-Barr virus (EBV), Kaposi’s sarcoma-associated herpesvirus (KSHV), and human papillomavirus (HPV) account for nearly all of the virus-associated human cancers worldwide. These viruses have also been excellent models for the study of oncogenic virus-mediated cell transformation. In this review we will discuss how each of these viruses engage and subvert aspects of the host DDR. The first level of DDR engagement is a result of the genetic linkage between the oncogenic potential of these viruses and their ability to replicate. Namely, the promotion of cells from quiescence into the cell cycle to facilitate virus replication can be sensed through aberrant cellular DNA replication structures which activate the DDR and hinders cell transformation. DNA tumor viruses subvert this growth-suppressive DDR through changes in viral oncoprotein expression which ultimately facilitates virus replication. An additional level of DDR engagement is through direct detection of replicating viral DNA. These interactions parallel that observed in other DNA virus systems in that the need to subvert these intrinsic sensors of aberrant DNA structure in order to replicate must be in place. DNA tumor viruses are no exception. This review will cover the molecular features of DNA tumor virus interactions with the host DDR and the consequences on virus replication.
1. Introduction
This review will focus on the interplay between DNA tumor viruses and the host DNA damage response (DDR). We will first discuss the components of the DDR sensing machinery and signaling pathways, then the interplay between viruses and the DDR. The role of the DDR in regulating replication of each specific virus, EBV, KSHV, and HPV, will then be discussed individually. Finally, we will address emerging themes in the interactions between viral proteins and DNA with the DDR and convergent roles of the DDR in regulating latent versus lytic virus replication.
1.1. The DNA Damage Response
Our DNA needs to be replicated faithfully in the face of continuous damage by both endogenous and exogenous agents. Damaged DNA that is not repaired during the cell cycle has the potential to cause genomic instability which could ultimately result in cancer. The DNA damage response is a signal transduction pathway that senses and recruits repair complexes to the site of damage and can stall cell cycle progression until the repair is complete (reviewed in [1]). The presence of damaged DNA triggers a response mediated by members of the phophinositide-3-like kinase (PIKK) family of serine/threonine kinases: ataxia-telangiectasia mutated (ATM), ataxia-telangiectasia and RAD3-related (ATR) or DNA-dependent protein kinase (DNA-PK) [2]. This leads to the activation via phosphorylation of a number of downstream factors leading to repair or in the face of irreparable damage, senescence or apoptosis.
The most detrimental form of DNA damage is double stranded breaks (DSBs) which are sensed by the ATM arm of the DDR (Figure 1A and reviewed in [3]). In response to DSBs, the Mre11-Rad50-Nbs1 (MRN) complex binds to the site of DNA damage and ATM is recruited to the damaged DNA where it interacts with the MRN complex [3, 4]. ATM then undergoes an activating auto-phosphorylation at serine 1981 which causes it to transition from a dimer to a monomer [5, 6]. ATM is further activated through additional phosphorylations as well as acetylation of lysine 3016 by TIP60 [7]. Recent evidence also implicates the modulation of phosphorylation by protein phosphatase 2A [8] and WIP1 phosphatases in the activation of ATM [9]. ATM then phosphorylates H2AX on S139 (known as γH2AX) [10] which recruits the critical adaptor protein MDC1 [11]. The ubiquitin ligase proteins RNF8 and RNF168 bind to ATM-phosphorylated MDC1 leading to the ubiquitination of γH2AX which scaffolds the interactions with 53BP1 and Brca1 [12–15]. These events nucleate and retain a host of DNA repair factors at sites of DNA damage as well as enable the phosphorylation of downstream targets of ATM including Chk2, p53 and Cdc25 family members that mediate cell cycle arrest and apoptosis [2, 16].
Figure 1. Schematic diagram of the ATM and ATR DNA damage response signaling pathways.
a) ATM activation responding to a DNA double-stranded break (DSB). The initial complex of Mre11, Rad50, and Nbs1 targets the DSB and recruits ATM to activate its kinase domain. This leads to phosphorylation and monomerization, which then triggers a cascade of target phosphorylations. H2AX, the histone H2A variant, is first phosphorylated and provides a scaffold for MDC1, which recruits the ring finger Ub ligases RNF8 and RNF168 to ubiquitinate H2AX. Concomitantly, enzymes such as Tip60 are recruited that can facilitate both ATM activation and chromatin de-compaction through acetylation of ATM directly and histone tails, respectively. The ubiquitination of H2AX further promotes recruitment and retention of factors such as 53BP1 and Brca1 that establish stable DDR foci propagating ATM signaling to effector molecules such as the checkpoint kinase Chk2, the transcription factor p53, and others. b) ATR activation responding to ssDNA exposure following DNA replicative stress. Following replication fork collapse, ssDNA exposure is sensed by the protein RPA that coats these regions and serves as a scaffold to recruit the kinase ATR through its co-factor ATRIP. The critical ATR activating factor TopBP1 is recruited through the Rad9-Rad1-Hus1 (9-1-1) complex at the replication fork. Finally, Claspin recruitment to the complex enables ATR phosphorylation of Chk1 and many other downstream effectors. A common downstream target is p53, which can be phosphorylated by both ATM and ATR leading to phenotypic outcomes ranging from DNA repair to apoptosis and senescence depending on the strength and duration of the DNA damaging signal.
ATR is essential in cells for DNA replication as it responds to single stranded DNA (ssDNA) exposure following replication fork collapse and the uncoupling of the replication complex (Figure 1B and reviewed in [17]). Replication protein A (RPA) coats the ssDNA and recruits ATR through its partner ATR interacting protein (ATRIP) [18]. Topoisomerase-binding protein 1 (TopBP1) is recruited to the ssDNA through the Rad9-Rad1-Hus1 (9-1-1) complex and then binds to both ATR and ATRIP. This stimulates ATR kinase activity leading to the phosphorylation of downstream effectors [16, 19] including the critical mediator Claspin, which recruits Chk1 to ssDNA enabling ATR to phosphorylate and activate its kinase domain to promote checkpoint activation [2, 20, 21].
Chk1 is the primary effector of the intra-S and G2/M checkpoints while Chk2 exerts influence in G1/S and intra-S [22, 23]. Both Chk1 and Chk2 phosphorylate the Cdc25 family of phosphatases which function to activate cyclin-dependent kinases (CDKs) by removing inhibitory phosphorylations [2]. Chk2 also modulates cell cycle progression by phosphorylating E2F and both Chk1 and Chk2 can phosphorylate p53 with downstream consequences on G1/S phase progression and apoptosis. Chk1 and Chk2 also play a role in DNA repair through the phosphorylation and activation of components of the homologous recombination (HR) pathway [23]. Despite the apparent dissection of function between the ATM and ATR pathway signaling to Chk2 and Chk1 respectively, these pathways are often functionally redundant where common targets are temporally offset in being phosphorylated by one kinase or the other [24]. Nevertheless, the engagement of the DDR leads to a rapid accumulation of factors with characteristic activating phosphorylations at sites of DNA damage. These hallmarks are useful measures of DDR activity, but detailed genetic analysis of individual signaling molecules is necessary to get an accurate picture of how these complex signaling networks are built.
1.2. Viruses and the DDR
Viruses face extraordinary challenges upon infection of a host cell. Both RNA and DNA viruses must evade innate sensors of foreign nucleic acids and other pattern recognition receptors to begin protein production, genome replication, and new particle formation. In the case of DNA viruses, a major challenge involves a specific conflict set up between the viral and host DNA genomes. The viral DNA genome may be recognized as damaged DNA and initiate a DNA damage response (DDR) that could have beneficial or catastrophic consequences on virus replication. For instance, adenovirus DNA structures are recognized by the host DDR to prevent cleavage and packaging of progeny DNA. An adenovirus protein E3 ubiquitin ligase complex (E4orf3/E1B55k) specifically degrades the components of the host DNA damage sensing Mre11/Rad50/Nbs1 (MRN) complex to allow faithful DNA processing and prevent concatemerization [25]. This landmark paper highlighted the complexity of the conflict between DNA viruses and the host DDR. Subsequent work in other DNA viruses has indicated both a positive role for the DDR in some aspects of virus replication as well as a limiting role in others [26].
In the specific case of DNA tumor viruses, several aspects of virus replication and virus-induced cell transformation involve the DDR [27, 28]. Viral oncoproteins are required to drive infected cells from quiescence into the cell cycle in order to promote virus replication. The consequence of aberrant viral oncoprotein expression is cellular DNA replicative stress that can activate a growth-suppressive DDR. In the setting of viral latency or during virus replication, viral proteins can directly engage and either activate or inactivate components of the host DDR. This engagement can engender the specificity necessary to either promote certain aspects of the DDR such as repair factor recruitment or negate other activities such as downstream activation of apoptosis. A final mechanism by which DNA tumor viruses can engage the DDR is through direct recognition of aberrant viral DNA structures. Under certain circumstances such as latent DNA replication of large DNA tumor viruses (e.g. gamma-herpesviruses) DDR factor recruitment is important in maintaining episomal DNA. Similarly, during lytic replication of these viruses, DDR factors assemble at sites of viral DNA replication to recruit factors likely important for recombination and repair of viral DNA prior to packaging. However, robust DDR signaling can activate checkpoints and apoptosis that must be quenched downstream in order to allow virus replication to proceed. The coordination of these processes is an area of intense study and DNA tumor viruses provide an excellent model with which to strengthen our understanding of these responses. In this review we will focus on the mechanisms by which human DNA tumor viruses engage and subvert the host DNA damage response. The three representative viruses discussed will be Epstein-Barr virus, Kaposi’s sarcoma-associated herpesvirus, and the human papillomaviruses.
2. Epstein-Barr virus and the DNA damage response
2.1. Epstein-Barr virus
Epstein-Barr virus (EBV) is a human gamma-herpesvirus that is found ubiquitously in the adult population where it establishes a lifelong latency with no discernible clinical manifestations in the majority of those infected [29]. Infection with EBV generally occurs early in life and the virus establishes a life-long latency in B lymphocytes. EBV has been linked to a number of benign and malignant diseases such as infectious mononucleosis, endemic Burkitt’s lymphoma (eBL) and the epithelial cell-derived nasopharyngeal carcinomas (NPC) [29]. EBV is a concern in immuno-compromised patients where it has been linked to a large subset of HIV associated B-cell lymphomas and post-transplant lymphoproliferative disorders [29].
Initial EBV infection of epithelial cells typically results in lytic replication which is followed by infection of resting B cells where long term latency is established. Most EBV induced malignancies are associated with the latent stage of the viral replicative cycle during which only a limited set of viral proteins and small non-coding RNAs are produced. The proteins expressed during latent infection include the EBV nuclear antigens (EBNAs 1, 2, 3A, 3B, 3C and LP) and the latent membrane proteins (LMP1 and LMP2) [30]. Akin to many oncogenic viruses, EBV alters the host cell to ensure long term proliferation and survival. This includes induction of the cell cycle and evasion of innate host defenses that would result in long-term senescence or apoptosis. The first viral proteins expressed in the infected cell are EBNA-LP and EBNA2 which transcriptionally up-regulate the expression of a number of viral and cellular genes inducing a transition of resting B cells into the cell cycle [31, 32]. These genes include the viral EBNA1 and EBNA3A, 3B, and 3C as well as cellular S phase initiating proto-oncogenes such as c-Myc, E2F1, and cyclin D2 [31] (Figure 2A, bottom). The EBNA3 proteins modulate EBNA2 dependent gene expression in addition to regulating a number of other cellular targets [33, 34]. EBNA2 and EBNA3C then co-activate the viral LMP1, a constitutively active mimic of the TNF receptor and B-cell co-stimulatory molecule CD40 [35, 36]. This triggers NFκB, p38 and JNK signaling pathways resulting in the modulation in gene expression patterns to allow for long-term cellular proliferation and survival. Additionally, LMP2A acts a mimic of the B cell receptor leading to constitutive, high level expression of PI3K signaling and pro-survival pathways [37]. EBNA1 is a DNA binding transcription factor that primarily functions to replicate and segregate the EBV episome [30].
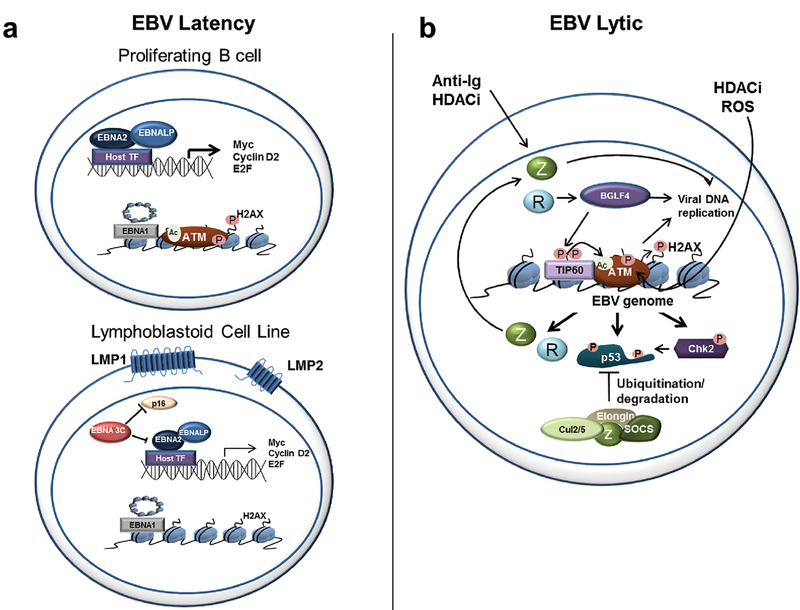
Figure 2. Epstein-Barr virus and the DNA damage response.
a) DDR activation during EBV latency. Top, Early after B cell infection with EBV, cells hyper-proliferate dependent on the expression of the viral EBNA2 and EBNA-LP proteins driving c-Myc and other S phase-promoting genes. This results in an activated DDR including ATM and H2AX phosphorylation. Bottom, in later divisions after infection, and also in immortalized lymphoblastoid cell lines (LCLs), the expression of the EBNA3C protein mitigates the DDR, presumably through attenuating the CDK inhibitor p16. Additional viral proteins are expressed such as LMP1 and LMP2 that contribute to the LCL survival phenotype. B) DDR activation during EBV lytic infection. Reactivation from diverse stimuli converge on expression of the critical viral trans-activator Z. In the case of anti-Ig treatment of latently-infected B cells or pan-HDAC inhibition, these effects are direct on the Z promoter (top left). However, other molecules such as DNA damaging agents like reactive oxygen species (ROS) can directly promote ATM activation. The consequence of this activation is also Z expression through the ATM-dependent regulation of viral lytic promoters and Tip60-mediated acetylation of viral chromatin. Expression of the viral kinase BGLF4 further supports this mechanism through direct activation of Tip60, ATM, and H2AX. Ultimately the downstream signaling from ATM can lead to the deleterious effect of p53-mediated apoptosis. However, the viral Z protein can form an E3 ubiquitin ligase complex to promote p53 degradation at late stages during viral lytic replication.
EBV encodes three distinct latency programs which are tightly linked to the differentiation state of the infected B cell and exhibit distinct patterns of gene expression [38]. Infection of resting primary human peripheral blood B cells with EBV in vitro transforms them into indefinitely proliferating; non-tumorigenic lymphoblastoid cell lines (LCLs) which express the full set of latency genes (latency III). In contrast, the more restricted latency II program is found in normal EBV-infected germinal center B cells, Hodgkin’s lymphomas, and NPCs where only EBNA1, the LMPs and viral non-coding RNAs are detected. A latency I expression profile is found in BL where only the non-coding viral RNAs and EBNA1 is expressed. A more recently described latency type, Wp-restricted, is found in approximately 20% of EBV-positive BLs, is often deleted for EBNA2 and retains all other latent genes [39]. Finally, normal resting peripheral blood memory B cells in asymptomatic individuals display a latency 0 type of gene expression where essentially all genes are silenced, save EBNA1 during cell division and perhaps the viral miRNAs [40].
2.2. The role of the DDR in suppressing EBV-induced cell proliferation
Viral induction of proliferation and survival pathways can result in oncogenic transformation as evidenced by the tight association of latent EBV infection with many types of malignancies. However, these malignancies are rare despite the greater than 90% prevalence of the virus in the adult population. In support of this, the efficiency of B cell transformation is low with only 1% of infected cells becoming immortalized [41, 42]. This implies that the host cell must be able to restrict virally induced proliferation. This was addressed by work by Nikitin, et al. who demonstrated that at least 10% of this restriction is due to the induction of the DDR [43]. Infection of primary B cells with EBV leads to a transient period of hyper-proliferation that peaks 4–6 days after infection and eventually slows to the rate observed in LCLs. This hyper-proliferative phase coincides with the activation of the ATM-dependent DDR indicated by the phosphorylation of H2AX, ATM, Chk2 and the formation of 53BP1 nuclear foci (Figure 2A). These cells were demonstrated to be nearly 100% latently infected and the DDR markers did not co-localize with viral episomes. This indicates that the DDR was activated in response to host DNA damage and not to the replication or presence of the viral DNA. Inhibition of ATM or Chk2 enhanced B-cell transformation implying that the DDR is an innate mechanism that the cell uses to restrict viral-mediated proliferation. The enhanced transformation in the presence of the inhibitors was only effective during the hyper-proliferative period. As the infected B cells transitioned to become immortalized LCLs there was a reduction in both DDR markers and sensitivity to the DDR.
The differences in host gene expression between resting B cells, transformed LCLs and this hyper-proliferative population could reveal how the host cell can limit transformation. Indeed, gene expression microarrays indicated that there are distinct transcriptomic changes between the hyper-proliferating cells in comparison to either LCLs or uninfected B cells [43–45]. These include enrichment for genes that are involved in cell cycle progression and the DDR. This also corresponded to a transition from a period of elevated EBNA-LP and EBNA2 activity to an accumulation of EBNA3 proteins. As previously noted, EBNA-LP and −2 up-regulate expression of a number of genes involved in cell cycle progression and this activity is repressed though the action of the EBNA3 proteins [46]. This could indicate a model in which initial infection causes a transient period of hyper-proliferation induced by EBNA2 regulated genes such as c-Myc, which results in replicative stress and induction of the DDR-mediated growth arrest.
In order to establish latency, the virus must overcome the DDR induced cell-cycle arrest. Loss of EBNA3C prevents EBV mediated B-cell transformation and also results in a dramatic increase in the activation of DDR markers [43]. Previous work has demonstrated that EBNA3C represses the MDM2 antagonist p14ARF and the cyclin-dependent kinase inhibitor p16INK4a through epigenetic silencing of their promoter [47, 48]. Given recent evidence suggesting that persistent DDR signaling at sites of irreparable DNA damage leading to senescence can be exacerbated by over-expression of p16 [49], the role of EBNA3C in suppressing p16 may, in fact, be critical for its ability to attenuate the EBV-induced DDR.
EBNA3C has also been demonstrated to interact with a number of host factors including RBPJ, CtBP, NCoR, HDACs, and others [50–53]. Notably it was shown to directly interact with Chk2 and this interaction relieved a G2/M cell cycle block [54]. EBNA3C has also been linked to S-phase entry through the stabilization of the Cyclin D1/CDK6 complex [55] and recruitment of SCFSkp2 E3 ligase for ubiquitin mediated degradation of pRB [56]. Therefore, EBNA3C is a key viral factor that is likely to regulate the host DDR through multiple mechanisms throughout the cell cycle.
2.3. EBV and genomic instability
DNA damage that is not repaired could ultimately lead to genomic instability and cancer. Unlike the karyotypically stable LCLs [57], chromosomal aberrations have been observed in EBV associated Burkitt’s lymphomas (BLs) and other cancers (reviewed in [58]). A direct link between EBV viral proteins and genomic instability was uncovered by the Masucci group in studies looking at paired EBV negative and positive BL lines. The presence of EBV caused a 3–10 fold increase in chromosomal abnormalities including dicentric chromosomes as well as the induction of a DDR response as indicated by the presence of γH2AX [59]. Further analysis revealed that EBNA1 alone induced the chromosomal dysfunction and up-regulation of the DDR [60]. EBNA1 increased expression of the transcription factor Nox2, which in turn increased levels of reactive oxygen species (ROS) resulting in DNA damage and aberrant chromosomal structures. In a follow up study, the authors found telomere dysfunction in a number of EBV positive BL cell lines while the corresponding EBV negative lines were karyotypically normal [61]. BL cells that express EBNA1 exhibit increased telomere dysfunction such as extra-chromosomal telomere signals, the accumulation of telomere dysfunction induced foci (TIFs), and telomeric-sister chromatid exchange. While EBNA1 had no effect on the overall expression of the shelterin components, it did reduce the association of TRF2 with telomeres. EBNA1 appears to induce this damage through ROS as compounds that act as ROS scavengers were able to inhibit the appearance of DDR markers and reconstitute TRF2-protected telomeres [61].
EBNA1 is not the only viral protein implicating in inducing genomic instability [58]. In an EBV-negative B lymphoma cell line, BJAB, ectopic expression of LMP1 or EBNA3C also promoted genomic instability as detected by chromosomal aberrations and the up-regulation of DDR markers [62]. The three viral proteins appear to induce this instability through unique mechanisms. Studies in epithelial cells initially indicated that LMP1 could inhibit DNA repair through inducing the cytoplasmic accumulation and thus inhibition of Foxo3, a transcription factor implicating in cell cycle arrest and DNA repair mechanisms [63]. Furthermore, LMP1-expressing B cells had a reduced capacity to repair damaged DNA and a decrease in ATM response [62]. In contrast, EBNA3C over-expression promoted aneuploidy through a mitotic spindle checkpoint failure. More specifically, EBNA3C led to a decrease in the expression of the checkpoint protein BubR1 which binds cdc2 and prevents anaphase progression [62]. Additionally, the interaction of EBNA3C with Chk2 and other cell cycle components discussed above could be involved in promoting genomic instability through aberrant mitotic progression [54, 64, 65]. Collectively, these studies indicate that the aberrant expression of EBV latent oncoproteins may contribute to oncogenesis through disrupting normal controls of DNA repair and cell cycle checkpoints.
2.4. Latent EBV DNA replication requires the DDR
During viral latency the EBV episome is replicated once per cell cycle by EBNA1 ([30]; reviewed in [66]). EBNA1 binds to specific regions of the EBV genome, namely oriP, the origin of plasmid replication, and recruits the cellular replication machinery. EBNA1 binds to a family of repeats (FR) and a dyad symmetry (DS) element within oriP [67]. Host proteins including the cellular origin replication complex (ORC) and mini chromosome maintenance (MCM) complex are enriched in association with EBNA1 at the DS elements during G1 and early S phase [68]. Interestingly, there are also repeat elements adjacent to EBNA1 binding sites within the DS that are very similar to telomere repeat elements [69, 70]. In fact, these sites bind the shelterin components TRF1, TRF2 and hRap1. Shelterin components normally function to protect telomeres from DDR sensing (reviewed in [71]). Through these sites, EBNA1 and TRF2 interact and recruit ORC to the viral episome [72, 73]. TRF2 and hRap1 binding are both critical for EBNA1 mediated replication while TRF1 appears to be inhibitory [70]. Thus, EBV co-opts the normal cellular DNA machinery of telomeric DNA repeats to recruit shelterin components to facilitate replication of the viral genome. This is likely important to suppress hyper-activation of the DDR at latent EBV genomes as occurs at protected telomeric ends due to the activity of TRF2 [71].
The firing of oriP is also delayed in early S phase by a host checkpoint regulation that is dependent on the recruitment of HDACs by TRF2 [74]. In early S phase the nucleosomes adjacent to the TRF2 binding sites are de-acetylated. As S phase progresses, the nucleosomes become acetylated and oriP replication ensues [74]. The activity of the DDR protein Chk2 is down-regulated after interaction with TRF2 [75] in a cell cycle dependent manner with enrichment in S phase [76]. Chk2 depletion by siRNA or Chk2 kinase inhibitor inhibits oriP replication and episome maintenance and advances the replication timing of OriP [76]. Mechanistically, the depletion of Chk2 alters histone de-acetylation at DS sites within oriP. In vitro studies indicate that Chk2 phosphorylates TRF2 on its amino terminal domain and a phospho-mimetic TRF2 mutant precludes ORC recruitment to oriP.
In addition to shelterin components, EBNA1 also recruits the DNA damage sensing MRN complex to oriP during S phase [77]. In the absence of Nbs1, similar to loss of Chk2, oriP maintenance and EBV genome maintenance is suppressed indicating an important role for MRN recruitment to the oriP. Therefore, the oriP DNA element serves as a unique scaffold for DDR factors and signaling that is required to orchestrate the faithful replication of EBV DNA during latency as well as presumably to promote the proper resolution of Holliday junction structure formed late during each S phase [77].
2.5. The role of the DDR in facilitating EBV lytic replication
EBV latently infected B cells can switch into a lytic phase of infection generating progeny virions that can infect a new host. Lytic reactivation is a highly regulated process that begins with expression of two immediate early viral genes, BZLF1 and BRLF1 which encode the proteins Z and R, respectively [46]. This expression results in the transactivation of early lytic genes which encode viral proteins responsible for replication. After viral replication, the late genes are transcribed, resulting in the production of the structural proteins needed to produce new virions. EBV, like many viruses, usurps host cell cycle controls in order to provide an S phase like environment for efficient replication.
The best characterized viral protein involved in this process is Z which can interact with and alter components of the cell cycle checkpoint in addition to its role as a transcription factor. The link between Z and the DDR was first observed by Kudoh, et al. using a tetracycline inducible BZLF1 (Z) construct to induce lytic replication [78]. Induction of Z correlated with the phosphorylation and activation of ATM, Chk2, H2AX and p53 as well as the co-localization of ATM-S1981 and the MRN complex with EBV replication centers. The activation of these proteins would normally lead to cell cycle arrest and apoptosis. However, the authors also observed hyper-phosphorylation of Rb and the subsequent release and activation of E2F1 and its downstream targets such as cyclins A and E indicating cell cycle progression.
One of the ways that Z regulates cell cycle progression is through its interaction with p53 [78–80]. Overexpression of Z in B958 cells induces the activation of p53 but without the subsequent increase in protein levels. Additionally, the expression of downstream targets such as MDM2 and p21CIP1 were not elevated [78, 80] although p53 retained its ability to bind to DNA and had increased post translational modification [80]. Subsequent studies revealed that Z can function as an adaptor of the Elongin B/C-cul2/5-SOCS-(ECS) box ubiquitin ligase complex through direct interaction with Cul2 and Cul5 [81]. The targeting and subsequent degradation of p53 by Z was enhanced by phosphorylation of p53 on S15 and S20 as a result of the activation of the ATM pathway.
To further delineate the interplay between the DDR and lytic replication, the Hayward group identified host substrates phosphorylated by a conserved herpesvirus kinase [82]. The EBV orthologue, BGLF4, is a mimic of host cyclin-dependent kinases and plays a key role in the establishment of an S-phase like environment in infected cells. Using a protein microarray to identify BGLF4 substrates, Li, et al. observed enrichment for proteins involved in the ATM arm of the DDR including TIP60, a chromatin remodeling protein that is involved in both DNA repair and transcription [82]. BGLF4 interaction with TIP60 results in its phosphorylation and the activation of its histone and protein acetylation activity. TIP60 has previously been shown to activate ATM in response to DNA damage [7] and its activation by BGLF4 also results in the phosphorylation of ATM and its downstream effectors (Figure 2B). Critically, loss of either TIP60 or ATM suppressed EBV DNA replication after lytic induction [82]. Mechanistically, BGLF4 and TIP60 are recruited to the EBV lytic replication origin (OriLyt) and other viral promoters stimulating both γH2AX and acetylated H2AK5 leading to increased viral lytic gene expression and DNA replication.
The importance of the DDR for lytic replication was further confirmed by the Kenney group exploring the importance of ATM in response to induction by a variety of stimuli [83]. Treatment of Akata BL cells with the ATM inhibitor KU55933 suppressed viral reactivation after induction with paclitaxel, a chemotherapeutic agent; sodium butyrate an HDAC inhibitor; activation of the B cell receptor by anti-IgG treatment; cytokine treatment with TGFβ or treatment with the proteosome inhibitor bortezomib. The importance of ATM for lytic reactivation was also seen in both EBV positive BL lines as well as in epithelial cells. Of note, not all of the BL lines used in this study contain functional p53 indicating that the role of ATM in lytic reactivation is at least partially p53 independent. Since ROS was recently shown to activate ATM kinase activity [84] the authors speculated that it also might lead to lytic reactivation. Indeed, ROS did lead to lytic reactivation which was blocked with subsequent treatment with the ATM inhibitor or by knocking down ATM with shRNAs.
Viruses subvert host ubiquitination pathways in most phases of their life-cycle in order to evade innate immune responses and to degrade proteins that would hinder their replication. Similarly, ubiquitin specific proteases are also targeted by viral proteins such as EBNA1 to alter the turnover of proteins needed for replication [85]. Recent work by the Masucci group has demonstrated that the tegument protein, BPLF1 is a de-neddylase that regulates Cullen-ring ligases (CRL) to alter S phase progression [86]. BPLF1 is able to cleave both ubiquitin adducts as well as NEDD8 conjugates in transfected cells although evidence points to its de-neddylation activity as being more functionally relevant. BPLF1 directly interacts with CRLs and modulates their activity as seen through an accumulation of substrates such as Cdc25A, p21 and Cdt1, an S phase licensing factor. The stabilization of Cdt1 through its de-neddylation promotes aberrant S phase entry and leads to the accumulation of phosphorylated ATM and Chk2. Inhibition of BPLF1 by shRNA reduced EBV replication to a level similar to that seen with inhibition of Z [86].
While S phase entry is a critical aspect of EBV lytic DNA replication, the consequences on cell cycle arrest through aberrant entry via Cdt1 stabilization could be catastrophic for the virus. Interestingly, recent work by the Pagano laboratory indicates that BPLF1 can de-ubiquitinate the major host DNA replication processivity factor PCNA [87]. Since mono-ubiquitinated PCNA (PCNA-Ub) recruits trans-lesion synthesis (TLS) polymerase to sites of DNA damage [88] and replaces the replicative polymerase allowing it to bypass the lesion, the authors speculate that stalling of cellular replication may free cellular factors that are needed for viral replication [87]. Additionally, the downstream signaling consequences of PCNA-Ub such as strong checkpoint activation and apoptosis would be ameliorated by BPLF1. Thus, BPLF1 simultaneously functions to promote pseudo-S phase maintenance through Cdt1 stabilization while cutting off downstream signaling through inhibiting PCNA-Ub.
A model can then be considered in which DNA damage or ROS leads to the activation of ATM (Figure 2B). This leads to the activation of the Z and R promoters and the expression of the immediate early genes. Data from Tsurumi, Kenney, and other groups indicate that p53 may be important at this early stage in lytic replication [89]. The combined activity of Z and R would then lead to the up-regulation of BGLF4 which phosphorylates TIP60 resulting in an increase in its HAT activity. TIP60 then further activates ATM as well as the OriLyt promoter leading to lytic reactivation. Viral replication is aided at this late stage by the inhibition of p53 mediated apoptosis through Z-dependent ubiquitination as well as by the regulation of DDR signaling by BPLF1.
3. Kaposi’s sarcoma-associated herespvirus and the DDR
3.1. Kaposi’s sarcoma-associated herespvirus
Kaposi’s sarcoma-associated herpesvirus (KSHV) is a large, double-stranded DNA virus of the gamma herpesvirus family that was originally identified in the lesions of patients suffering from Kaposi’s sarcoma [90]. KSHV has been found in all forms of KS as well as primary effusion lymphoma (PEL) and multicentric Castleman’s disease (reviewed in [91]). Like all members of the herpesvirus family, KSHV can replicate in both lytic and latent stages. During latency, the virus is maintained as an extra-chromosomal episome that is linked to the host chromosome through the virally encoded protein latency-associated nuclear antigen (LANA) [92].
The major viral transcripts expressed during latency function to keep the cell alive and replicating. The proteins include LANA, viral cyclin (v-cyclin), viral FLICE inhibitory protein (vFLIP), and the kaposins, and KSHV also encoded many miRNAs [91]. LANA plays a role in abrogating host tumor suppressor pathways by inhibiting p53 and Rb-E2F [91, 93–95] in addition to its critical role in episome maintenance [92]. LANA also activates telomerase reverse transcriptase (TERT) expression further promoting infected-cell survival [91, 96].
KSHV v-cyclin is structurally homologous to cellular D-type cyclins and can bind to and activate cellular CDK6 (reviewed in [97, 98]). The v-cyclin/CDK6 complex has a broader substrate range than its cellular counterpart and is resistant to inhibition by CDK inhibitors. vFLIP can activate NFκB by binding to the activating IKK complex which results in the expression of anti-apoptotic genes and the induction of cytokine secretion [91, 99]. Importantly, vFLIP also serves as a key negative regulator of autophagy in latently-infected cells [100].
3.2. The role of the DDR in suppressing KSHV-induced cell proliferation
The first link between latent KSHV infection and the DDR came from the Ojala group exploring the effect of v-cyclin in telomerase immortalized endothelial cells [101] (Figure 3A). Over-expression of v-cyclin decreased cellular proliferation through the induction of an S phase arrest. They also demonstrated that viral protein expression could lead to cellular senescence as has been observed by overexpression of classical cellular oncogenes. Both primary and immortalized endothelial cells had a strong induction of DDR markers such as phosphorylated H2AX, ATM, Chk2 and p53. Treatment of v-cyclin expressing cells with an inhibitor of the ATM kinase (KU-55933) led to a reduction in DDR markers and a release of the S-phase arrest which resulted in aberrant mitosis [101]. This could be due to v-cyclin induced centrosome amplification which can lead to aberrant mitosis in the absence of a functional DDR [102]. The DDR pathway was activated in KSHV infected endothelial cells after they overcame an early period of arrest. The effect of the DDR could be bypassed by ablation of the p53 or Rb signaling [101].
Figure 3. Kaposi’s sarcoma-associated herpesvirus and the DNA damage response.
a) DDR activation during KSHV latency. The induction of cell cycle progression by the viral cyclin (v-cyclin) in complex with CDK6 leads to the activation of an ATM-mediated DDR.The consequence of this activation is p53 induction, oncogene-induced senescence (OIS), and autophagy. However, co-expression of the KSHV v-FLIP protein overrides v-cyclin-mediated (OIS) by suppressing autophagy. b) KSHV and the DDR during lytic reactivation. Little is understood about KSHV lytic viral DNA replication and the DDR. However, the expression of the lytic cycle vIRF1 gene is capable of suppressing p53 activity and, in particular, the ability of ATM to activate p53. This suggests that if ATM were to be activated by KSHV lytic DNA replication (as occurs in EBV, HPV, and MHV68 infection), then a plausible mechanism to overcome checkpoint activation or apoptosis would be in place.
This work was followed up by Leidal, et al. who demonstrated that KSHV-infected primary human fibroblasts activated the DDR but did not undergo senescence [103]. This is in contrast to v-cylin expressing cells which had a greater induction of the DDR as well as senescence markers. The authors hypothesized that one of the KSHV latency genes must be capable of abrogating the v-cyclin induced senescence. To address this they performed an ectopic screen of a number of latency proteins and identified v-FLIP as a suppressor of v-cyclin mediated senescence.
vFLIP was previously shown to inhibit autophagy in KSHV-infected endothelial cells and lymphocytes [100] and autophagy is necessary for the efficient establishment of senescence [104]. Therefore, the authors explored the link between the loss of oncogene induced senescence (OIS) and the suppression of autophagy. KSHV v-cyclin induced autophagy similar to the potent oncogene HrasV12 [103]. Temporally regulated expression of v-cyclin revealed an initial period of hyper-proliferation followed by a sharp decrease in the rate of proliferation coinciding with the onset of OIS. p53 was strongly activated in cells expressing v-cyclin and its activity was required for the induction of OIS through the transcriptional up-regulation of the key autophagy regulators Sestrin1 and damage-regulated autophagy modulator (DRAM). Notably, the mTOR pathway was inactivated concomitant with OIS induction partially dependent on AMP-responsive protein kinase (AMPK), a target of Sestrin1. While co-expression of vFLIP with v-cyclin had no effect on DDR markers, the induction of autophagy and OIS was dramatically reduced. Surprisingly, in contrast to its role in cell survival through NFκB activity [105, 106], the specific antagonism of autophagy by vFLIP was responsible for its ability to override v-cyclin mediated OIS. This collaboration likely contributes significantly to KSHV-mediated tumorigenesis (Figure 3A).
3.3. The role of the DDR in KSHV lytic replication
While the default pathway for KSHV infection is latency it can be induced to reactivate by a variety of stimuli resulting in the generation of a full set of viral proteins and the production of new virions. The switch between latency and lytic replication is primarily mediated by the protein RTA, encoded by ORF5 [107]. The expression of RTA and other immediate early genes are followed by the expression of the delayed-early genes. These genes include a viral DNA polymerase, primase, helicase and single stranded binding proteins. These proteins can form a stable replication complex on the OriLyt promoter in the host cell nucleus. The final stage of lytic reactivation is the production of late genes which encodes structural proteins necessary for viral particle formation [107].
As with all of the DNA viruses discussed, KSHV needs to modulate the host cell to ensure that the there is an appropriate S phase-like environment for replication as well as the evasion of host cell defenses. KSHV evade host innate immunity by encoding several homologue of host genes such as viral interleukin-6 (vIL-6), viral Bcl2 and four homologues of interferon regulatory factors (vIRF1–4) (reviewed in [108]). vIRF1 was initially reported to interact with and inactivate p53 by suppressing its acetylation and its transcriptional activation potential [109]. This was further explored by Shin, et al., who used ectopic expression of a tetracycline regulated vIRF1 in a KSHV-null B lymphoma line, BJAB [110]. Expression of vIRF1 resulted in an increase in p53 ubiquitination and the inhibition of the activating phosphorylation of S15. Further analysis revealed that vIRF1 interacted with ATM through its C-terminal domain and that this lead to a decrease in its activation as revealed by phosphorylation of S1981. The activation of downstream targets of ATM such as Chk2, H2AX and p53 were also decreased by vIRF1, while the expression of the cell cycle regulatory protein, Cdc25, was increased consistent with a role for vIRF1 in suppressing downstream checkpoint activation by ATM. Therefore, these studies suggest a possible mechanism by which KSHV oncoproteins subvert downstream DDR signaling during lytic reactivation (Figure 3B). It will be of great interest in the future to discern whether, like in the case of EBV, KSHV also utilizes aspects of ATM signaling to promote lytic viral gene expression and DNA replication. Indeed, evidence from the KSHV-related gamma-herpesvirus, MHV68, suggests a role for ATM and H2AX in virus replication [111].
4. Human papillomaviruses and the DNA damage response
4.1. Human papillomaviruses
Human papillomaviruses (HPVs) are small double stranded DNA viruses that generally cause benign skin lesions. Over 120 serotypes of HPV have been identified that can be further classified as low or high risk according to their association with benign or malignant lesions, respectively. Infection with a high risk variant, such as HPV 16 or 18, HPV has been associated with 99% of cervical cancer [112] and ~25% of head and neck cancer [113].
The HPV lifecycle is closely linked with the differentiation state of epithelial cells (reviewed in [114, 115]). Initially, the virus infects the basal layer of squamous epithelia and establishes itself as a low copy number extra-chromosomal episome. These cells represent the only actively replicating cells within the epithelia and are exposed to infection through abrasions or wounding. Infection leads to an initial burst of viral DNA replication that is followed by a period where the virus DNA replicates in concert with the host chromosome. As uninfected epithelia divide they leave the basement membrane layer and undergo terminal differentiation until they are eventually shed as dead and dying cells. HPV undergoes its final replication cycle, called vegetative replication, in the differentiated cell layer. Newly generated virions can then be shed and spread to new targets.
Restricting its replicative life-cycle to differentiated cells allows the virus to remain undetected by the immune system for extended periods of time. However, terminal differentiation entails an exit from the cell cycle which would deprive the virus of the host replicative machinery. To circumvent this, HPV encodes several proteins (E5, E6 and E7) that promote proliferation by delaying exit from the cell cycle. E6 and E7 are the primary transforming genes of HPV through their antagonism of p53 and Rb, respectively [116–118]. The E1 and E2 proteins are essential for viral DNA replication and episome maintenance and may play a role in promoting genomic instability and cancer progression as will be discussed below [119, 120].
4.2. The role of the DDR in suppressing HPV-induced cell proliferation
Despite the high frequency of HPV infection, the rate of cancer progression is low and only occurs if the virus has not been cleared over an extended period of time. One striking observation is that a majority of the cervical cancer biopsies show integration of the high risk HPVs [114]. While this appears to occur randomly there does seem to be a preference for common fragile sites within the genome. One consequence of HPV genome integration is the loss of the E2 protein which is responsible for repressing viral gene expression in addition to its role as a replication factor. This could lead to uncontrolled expression of the E6 and E7 [121] which can independently transform cells and lead to genomic instability through mitotic defects [122].
The Ustav group demonstrated that integrated HPV genomes in HeLa and SiHa cells undergo an “onion skip” type of replication in the presence of exogenous E1 [123]. This leads to DNA replication structures that are recognized by the ATM arm of the DDR (Figure 4A). This ultimately leads to repair by either HR or NHEJ, the latter of which can produce mutations or chromosome translocations. In fact, genomic instability was observed in SiHa cells transfected with E1. In other studies, E1 over-expression from diverse HPV strains induced cell cycle arrest and a DDR in undifferentiated keratinocytes in the absence of viral DNA [124–126]. Expression of E2 alone or coupled with the viral origin was able to partially suppress this effect in one study [124] but not in another [125]. The ability of E1 to induce a DDR was seen even with basal level expression of the protein indicating that it is not simply an artifact of overexpression. The DDR foci partially co-localized with E1 and E2 as well as sites of DNA synthesis [125]. The level of nucleotide incorporation detected was below what would normally be detected for S phase progression and could indicate regions of DNA repair. Consistent with this, the DDR foci were also TUNEL positive indicating regions of fragmented DNA [125] and had double stranded breaks detected by alkaline comet assay [126].
Figure 4. Human papillomavirus and the DNA damage response.
a) HPV infection of basal layer epithelial cells and DDR activation. The infection of basal epithelial cells by HPV leads to a constitutive ATM-dependent DDR. This may be due to the role of HPV E1 in promoting replication of integrated HPV genomes or perhaps promoting DNA damage at other sites through the host genome (left). Normally, in this layer, E2 protein suppresses the expression of E6 and E7. If E2 is abrogated due to integration, then E7 up-regulation could lead to increased S-phase entry promoting the DDR or through direct engagement and activation of ATM (bottom, right). b) During vegetative DNA replication upon differentiation of the basal keratinocyte layer the ATM pathway is activated and DDR factors co-localize with viral genomes (right). ATM and Chk2 activity are important for viral DNA replication in this phase of the HPV life cycle. The uppermost layer of keratinocytes in the figure indicates virion release (stars).
High risk HPV E6 and E7 expression in undifferentiated keratinocytes is sufficient to induce replicative stress, which could result in collapsed replication forks and the activation of ATR and ATM pathways [127, 128]. While HPV oncoprotein expression drives aberrant S phase promoting a DDR, the potential for senescence or apoptosis is strongly mitigated by the ability of E6 to leverage E6AP to degrade p53 [129]. Therefore, HPV infection in undifferentiated keratinocytes can drive an oncogenic stress, but cell proliferation in the context of viral genome integration is maintained due to potent anti-apoptotic activities. The role of the DDR in suppressing HPV tumorigenesis may then be restricted to its functions as a regulator of viral DNA replication as discussed below.
4.3. The role of the DDR in facilitating HPV vegetative DNA replication
Emerging evidence suggests that the DDR plays a critical role in HPV amplification in differentiated keratinocytes. Keratinocytes containing the HPV-31 episome differentiate upon calcium treatment. Differentiation increases markers of the canonical ATM pathway which localize to nuclear foci [127]. These foci also contain the HPV episome as well as enzymes responsible for DNA repair and homologous recombination such as PCNA, pRPAS32-S33, Rad51 and Brca1 [130] (Figure 4B). Inhibition of ATM led to diminished replication foci and suppressed episome amplification [127]. ATM driven activation of Chk2 is required for caspase 7 activation [127] which was previously shown to be necessary for the cleavage of E1 and genome amplification [131] further solidifying the link between the DDR and viral replication. The up-regulation of DDR markers was also observed in primary human keratinocytes that underwent differentiation in organotypic raft cultures [132]. Strong pATM-S1981 was seen in differentiated cells that expressed either the HPV-31 [127] or HPV-18 genome [132].
DDR activation appears to be primarily induced by E7 in differentiated cells. Banerjee, et al. compared the expression of the HPV-18 genome, E7 alone or vector controls in organotypic raft cultures using primary human keratinocytes. The full viral genome or E7 alone were able to induce up-regulation of pATM-S1981as well as the activation of Chk2 and the ATR effector Chk1 [132]. Additionally, E7 was previously shown to directly interact with ATM and induce the activation of Chk2 [127]. Cultures expressing E7 alone or the HPV-18 genome exhibited a prolonged G2 phase [132, 133] as evidenced by the accumulation of inactive Cdk1 and Cdc25 in the cytoplasm and the lack of mitotic cells in culture. Active Chk1, Chk2 and JNK are known to inactive Cdc25 and therefore the activation of the DDR could lead to prolonging G2 phase for viral amplification [132].
A model is starting to emerge in which HPV is able to exploit the intrinsic DDR to amplify its genome. E7 can interact with and activate ATM in differentiated cells leading to the activation of its downstream effectors such as Chk2 and γH2AX (Figure 4). This results in the co-localization of many repair proteins at the viral genome. Chk2 can then activate caspase 7 resulting in the cleavage of E1 an essential replication factor. The activated DDR would then induce a G2 checkpoint by inactivating Cdk1 and Cdc25 to prevent entry into mitosis. This allows the virus to amplify its genome out of sync with normal cellular S phase thus preventing competition for replication factors. The inhibition of p53 by E6 prevents apoptosis normally induced by the activation of the DDR and thereby facilitates viral DNA replication.
5. Conclusions
DNA tumor viruses activate the host DNA damage response in multiple ways. Viral oncoproteins drive cells from quiescence into cell cycle and often lead to aberrant cellular DNA replication that can provoke the host DDR. This process may occur during viral latency as exemplified by the oncogenic herpesviruses or during acute infection. Furthermore, viral DNA replication, either latent or lytic, can produce structures that directly activate the DDR. Once activated, the DDR can promote DNA repair and recombination as well as activate cell cycle checkpoints, senescence, or apoptosis. In the case of latent viral oncoprotein-driven cellular DNA replicative stress leading to DDR activation, the consequence is a decrease in cellular proliferation which is ultimately detrimental for the virus. In contrast, during lytic replication the activation of the DDR through viral DNA replication structures is beneficial for the virus. Therefore, the host DDR plays divergent roles in the replication of human DNA tumor viruses depending on the mode of virus replication.
DNA tumor viruses drive cellular transformation under circumstances where virus replication and the ability to produce progeny virions is inhibited. For example, in the case of high risk HPVs, unintentional integration of viral DNA into the host genome leads to the aberrant high expression of oncoproteins E6 and E7, which can lead to cell transformation and contribute to tumorigenesis. Early expression of these oncoproteins in basal keratinocytes may mimic aspects of this process. As shown by Bester, et al., HPV-16 E6/E7 expression can activate a growth suppressive DDR due to increased cellular DNA replicative stress [128]. Similarly, in the case of EBV and KSHV, viral latent oncoproteins drive aberrant cellular DNA proliferation and a growth-suppressive DDR [43, 101]. Interestingly, recent work by the McCormick lab has shown that KSHV vFLIP is capable of overcoming the DDR-dependent senescence mediated by the v-cyclin protein in KSHV-infected endothelial cells [103]. This activity depends on induction of autophagy by vFLIP and mitigates the far downstream consequences of the DDR. Similarly, in the context of EBV infection of primary B cells, the cell proliferation rate is transiently high concomitant with DDR activation, but then wanes through immortalization dependent on expression of the viral EBNA3C protein [43]. Therefore, DDR activation in latently infected cells (in the case of the gamma-herpesivruses) leads to growth suppression that ultimately compromises viral infection. However, viral oncoproteins are able to mitigate these pathways either downstream through modulating autophagy or upstream through attenuating the strength of the oncogenic signal.
The nature of the molecular signal downstream of oncogenic stress that triggers the DDR in these settings remains controversial. Evidence for increased DNA replicative stress [134, 135], increased fragile sites [135], increased irreparable DNA damage at telomeres [136] and telomere dysfunction [137], as well as changes in chromatin structure associated with hyper-activation of transcription at certain loci have all been implicated in DDR activation downstream of activated oncogenes. The careful genetic dissection of the critical sensors and mediators of each of these aberrant DNA structures in the context of viral oncogene expression will be needed to discern precisely how the DDR is activated and whether these signals engender specificity in the downstream signaling pathways following oncogenic stress.
During lytic viral DNA replication, two mechanisms can give rise to an activated DDR. First, the set of proteins expressed early during infection or reactivation can generate a pseudo-S phase environment important for viral DNA replication. Often the consequence of this aberrant cell cycle promotion is the activation of the DDR. However, the downstream consequences of the DDR are mitigated enabling virus replication. In the case of EBV, for example, the promotion of S phase entry through the BPLF1 protein leads to an activated DDR including ATM phosphorylation. However, the downstream signaling at sites of cellular DNA damage are mitigated in part through the activity of BPLF1 in preventing PCNA mono-ubuiqitination. Furthermore, the viral Z protein is capable of degrading p53 once activated at late stages in lytic reactivation. Thus, as often observed in the context of virus infection, a multi-faceted approach to ensuring cell survival is used to facilitate DNA replication in the context of an activated DDR.
A second signal for DDR activation during lytic replication is the replication of viral DNA. In both the EBV and HPV systems, strong co-localization of DDR factors with sites of viral DNA replication corroborate their induction by aberrant viral DNA structures sensed as damaged DNA. Interestingly, DDR signaling, and ATM in particular, is required for efficient DNA replication and progeny virion formation. It is likely that the recruitment of factors necessary for viral DNA repair and/or recombination is critical in the process of faithful DNA replication and packaging necessary to produce competent infectious progeny. The precise set of ATM targets important for DNA tumor virus replication is an active and exciting area of current research. The mechanistic delineation of the beneficial DDR in virus replication has the potential to lead to new and interesting therapeutic targets in these and other viral diseases. Moreover, it is likely that using these viruses to probe the specificities of DDR activation will lead to new insights into the recognition, repair, and signaling from cellular sites of DNA damage as well.
Abbreviations
- DDR
DNA damage response
- EBV
Epstein-Barr virus
- KSHV
Kaposi’s sarcoma-associated herpesvirus
- HPV
human papillomavirus
- ATM
ataxia-telangectasia mutated
- ATR
ataxia-telangectasia and RAD3-related
- MRN
Mre11-Rad50-Nbs1
- BL
Burkitt’s lymphoma
- LCL
lymphoblastoid cell line
- DSB
double-stranded break
- ssDNA
single-stranded DNA
- EBNA
Epstein-Barr virus nuclear antigen
- LMP
(EBV) latent membrane protein
- vFLIP
(KSHV) viral FLICE (FADD-like IL-1β-converting enzyme) inhibitory protein
References
- 1.Bartek J, Bartkova J, and Lukas J, DNA damage signalling guards against activated oncogenes and tumour progression. Oncogene, 2007. 26(56): p. 7773–9. [DOI] [PubMed] [Google Scholar]
- 2.Smith J, et al. , The ATM-Chk2 and ATR-Chk1 pathways in DNA damage signaling and cancer. Adv Cancer Res, 2010. 108: p. 73–112. [DOI] [PubMed] [Google Scholar]
- 3.Derheimer FA and Kastan MB, Multiple roles of ATM in monitoring and maintaining DNA integrity. FEBS Lett, 2010. 584(17): p. 3675–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Falck J, Coates J, and Jackson SP, Conserved modes of recruitment of ATM, ATR and DNA-PKcs to sites of DNA damage. Nature, 2005. 434(7033): p. 605–11. [DOI] [PubMed] [Google Scholar]
- 5.So S, Davis AJ, and Chen DJ, Autophosphorylation at serine 1981 stabilizes ATM at DNA damage sites. J Cell Biol, 2009. 187(7): p. 977–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bakkenist CJ and Kastan MB, DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature, 2003. 421(6922): p. 499–506. [DOI] [PubMed] [Google Scholar]
- 7.Sun Y, et al. , A role for the Tip60 histone acetyltransferase in the acetylation and activation of ATM. Proc Natl Acad Sci U S A, 2005. 102(37): p. 13182–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Goodarzi AA, et al. , Autophosphorylation of ataxia-telangiectasia mutated is regulated by protein phosphatase 2A. EMBO J, 2004. 23(22): p. 4451–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Shreeram S, et al. , Wip1 phosphatase modulates ATM-dependent signaling pathways. Mol Cell, 2006. 23(5): p. 757–64. [DOI] [PubMed] [Google Scholar]
- 10.Burma S, et al. , ATM phosphorylates histone H2AX in response to DNA double-strand breaks. The Journal of biological chemistry, 2001. 276(45): p. 42462–7. [DOI] [PubMed] [Google Scholar]
- 11.Stewart GS, et al. , MDC1 is a mediator of the mammalian DNA damage checkpoint. Nature, 2003. 421(6926): p. 961–6. [DOI] [PubMed] [Google Scholar]
- 12.Huen MS, et al. , RNF8 transduces the DNA-damage signal via histone ubiquitylation and checkpoint protein assembly. Cell, 2007. 131(5): p. 901–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mailand N, et al. , RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell, 2007. 131(5): p. 887–900. [DOI] [PubMed] [Google Scholar]
- 14.Kolas NK, et al. , Orchestration of the DNA-damage response by the RNF8 ubiquitin ligase. Science, 2007. 318(5856): p. 1637–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Doil C, et al. , RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell, 2009. 136(3): p. 435–46. [DOI] [PubMed] [Google Scholar]
- 16.Matsuoka S, et al. , ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science, 2007. 316(5828): p. 1160–6. [DOI] [PubMed] [Google Scholar]
- 17.Nam EA and Cortez D, ATR signalling: more than meeting at the fork. Biochem J, 2011. 436(3): p. 527–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zou L and Elledge SJ, Sensing DNA damage through ATRIP recognition of RPA-ssDNA complexes. Science, 2003. 300(5625): p. 1542–8. [DOI] [PubMed] [Google Scholar]
- 19.Kumagai A, et al. , TopBP1 activates the ATR-ATRIP complex. Cell, 2006. 124(5): p. 943–55. [DOI] [PubMed] [Google Scholar]
- 20.Jeong SY, et al. , Phosphorylated claspin interacts with a phosphate-binding site in the kinase domain of Chk1 during ATR-mediated activation. The Journal of biological chemistry, 2003. 278(47): p. 46782–8. [DOI] [PubMed] [Google Scholar]
- 21.Kumagai A and Dunphy WG, Repeated phosphopeptide motifs in Claspin mediate the regulated binding of Chk1. Nat Cell Biol, 2003. 5(2): p. 161–5. [DOI] [PubMed] [Google Scholar]
- 22.Bartek J and Lukas J, Chk1 and Chk2 kinases in checkpoint control and cancer. Cancer Cell, 2003. 3(5): p. 421–9. [DOI] [PubMed] [Google Scholar]
- 23.Stracker TH, Usui T, and Petrini JH, Taking the time to make important decisions: the checkpoint effector kinases Chk1 and Chk2 and the DNA damage response. DNA Repair (Amst), 2009. 8(9): p. 1047–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Stiff T, et al. , ATR-dependent phosphorylation and activation of ATM in response to UV treatment or replication fork stalling. Embo J, 2006. 25(24): p. 5775–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Stracker TH, Carson CT, and Weitzman MD, Adenovirus oncoproteins inactivate the Mre11-Rad50-NBS1 DNA repair complex. Nature, 2002. 418(6895): p. 348–52. [DOI] [PubMed] [Google Scholar]
- 26.Weitzman MD, Lilley CE, and Chaurushiya MS, Genomes in conflict: maintaining genome integrity during virus infection. Annu Rev Microbiol, 2010. 64: p. 61–81. [DOI] [PubMed] [Google Scholar]
- 27.Nikitin PA and Luftig MA, The DNA damage response in viral-induced cellular transformation. Br J Cancer, 2012. 106(3): p. 429–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nikitin PA and Luftig MA, At a crossroads: human DNA tumor viruses and the host DNA damage response. Future Virol, 2011. 6(7): p. 813–830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Rickinson A and Kieff E, Epstein-Barr virus in Fields Virology, Knipe DM and Howley PM, Editors. 2007, Lippincott, Williams, and Wilkins: Philadelphia: p. 2603–2654. [Google Scholar]
- 30.Yates JL, Warren N, and Sugden B, Stable replication of plasmids derived from Epstein-Barr virus in various mammalian cells. Nature, 1985. 313(6005): p. 812–5. [DOI] [PubMed] [Google Scholar]
- 31.Kaiser C, et al. , The proto-oncogene c-myc is a direct target gene of Epstein-Barr virus nuclear antigen 2. J Virol, 1999. 73(5): p. 4481–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sinclair AJ, et al. , EBNA-2 and EBNA-LP cooperate to cause G0 to G1 transition during immortalization of resting human B lymphocytes by Epstein-Barr virus. EMBO J, 1994. 13(14): p. 3321–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Johannsen E, et al. , EBNA-2 and EBNA-3C extensively and mutually exclusively associate with RBPJkappa in Epstein-Barr virus-transformed B lymphocytes. J Virol, 1996. 70(6): p. 4179–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zhao B, Marshall DR, and Sample CE, A conserved domain of the Epstein-Barr virus nuclear antigens 3A and 3C binds to a discrete domain of Jkappa. J Virol, 1996. 70(7): p. 4228–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lin J, et al. , Epstein-Barr virus nuclear antigen 3C putative repression domain mediates coactivation of the LMP1 promoter with EBNA-2. J Virol, 2002. 76(1): p. 232–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Cahir McFarland ED, Izumi KM, and Mosialos G, Epstein-barr virus transformation: involvement of latent membrane protein 1-mediated activation of NF-kappaB. Oncogene, 1999. 18(49): p. 6959–64. [DOI] [PubMed] [Google Scholar]
- 37.Miller CL, et al. , Integral membrane protein 2 of Epstein-Barr virus regulates reactivation from latency through dominant negative effects on protein- tyrosine kinases. Immunity, 1995. 2(2): p. 155–66. [DOI] [PubMed] [Google Scholar]
- 38.Thorley-Lawson DA, Epstein-Barr virus: exploiting the immune system. Nat Rev Immunol, 2001. 1(1): p. 75–82. [DOI] [PubMed] [Google Scholar]
- 39.Kelly G, Bell A, and Rickinson A, Epstein-Barr virus-associated Burkitt lymphomagenesis selects for downregulation of the nuclear antigen EBNA2. Nat Med, 2002. 8(10): p. 1098–104. [DOI] [PubMed] [Google Scholar]
- 40.Babcock GJ, Hochberg D, and Thorley-Lawson AD, The expression pattern of Epstein-Barr virus latent genes in vivo is dependent upon the differentiation stage of the infected B cell. Immunity, 2000. 13(4): p. 497–506. [DOI] [PubMed] [Google Scholar]
- 41.Henderson E, et al. , Efficiency of transformation of lymphocytes by Epstein-Barr virus. Virology, 1977. 76(1): p. 152–63. [DOI] [PubMed] [Google Scholar]
- 42.Sugden B and Mark W, Clonal transformation of adult human leukocytes by Epstein-Barr virus. J Virol, 1977. 23(3): p. 503–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Nikitin PA, et al. , An ATM/Chk2-mediated DNA damage-responsive signaling pathway suppresses Epstein-Barr virus transformation of primary human B cells. Cell Host Microbe, 2010. 8(6): p. 510–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Price AM, et al. , Analysis of Epstein-Barr Virus-Regulated Host Gene Expression Changes through Primary B-Cell Outgrowth Reveals Delayed Kinetics of Latent Membrane Protein 1-Mediated NF-kappaB Activation. J Virol, 2012. 86(20): p. 11096–106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Forte E, et al. , The Epstein-Barr virus induced tumor suppressor miR-34a is growth promoting in EBV-infected B cells. J Virol, 2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kieff E and Rickinson A, Epstein-Barr virus and its replication, in Fields Virology, Knipe DM and Howley PM, Editors. 2007. p. 2603–2654. [Google Scholar]
- 47.Skalska L, et al. , Epigenetic repression of p16(INK4A) by latent Epstein-Barr virus requires the interaction of EBNA3A and EBNA3C with CtBP. PLoS Pathog, 2010. 6(6): p. e1000951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Maruo S, et al. , Epstein-Barr virus nuclear antigens 3C and 3A maintain lymphoblastoid cell growth by repressing p16INK4A and p14ARF expression. Proc Natl Acad Sci U S A, 2011. 108(5): p. 1919–24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Rodier F, et al. , DNA-SCARS: distinct nuclear structures that sustain damage-induced senescence growth arrest and inflammatory cytokine secretion. J Cell Sci, 2011. 124(Pt 1): p. 68–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Bajaj BG, et al. , Epstein-Barr virus nuclear antigen 3C interacts with and enhances the stability of the c-Myc oncoprotein. J Virol, 2008. 82(8): p. 4082–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Rosendorff A, et al. , EBNA3C coactivation with EBNA2 requires a SUMO homology domain. J Virol, 2004. 78(1): p. 367–77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Subramanian C, et al. , Epstein-Barr virus nuclear antigen 3C and prothymosin alpha interact with the p300 transcriptional coactivator at the CH1 and CH3/HAT domains and cooperate in regulation of transcription and histone acetylation. J Virol, 2002. 76(10): p. 4699–708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Touitou R, et al. , Epstein-Barr virus EBNA3 proteins bind to the C8/alpha7 subunit of the 20S proteasome and are degraded by 20S proteasomes in vitro, but are very stable in latently infected B cells. J Gen Virol, 2005. 86(Pt 5): p. 1269–77. [DOI] [PubMed] [Google Scholar]
- 54.Choudhuri T, et al. , The ATM/ATR signaling effector Chk2 is targeted by Epstein-Barr virus nuclear antigen 3C to release the G2/M cell cycle block. J Virol, 2007. 81(12): p. 6718–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Saha A, et al. , EBNA3C attenuates the function of p53 through interaction with inhibitor of growth family proteins 4 and 5. J Virol, 2011. 85(5): p. 2079–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Knight JS and Robertson ES, Epstein-Barr virus nuclear antigen 3C regulates cyclin A/p27 complexes and enhances cyclin A-dependent kinase activity. J Virol, 2004. 78(4): p. 1981–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Lacoste S, et al. , Chromosomal rearrangements after ex vivo Epstein-Barr virus (EBV) infection of human B cells. Oncogene, 2010. 29(4): p. 503–15. [DOI] [PubMed] [Google Scholar]
- 58.Gruhne B, et al. , EBV and genomic instability--a new look at the role of the virus in the pathogenesis of Burkitt’s lymphoma. Semin Cancer Biol, 2009. 19(6): p. 394–400. [DOI] [PubMed] [Google Scholar]
- 59.Kamranvar SA, et al. , Epstein-Barr virus promotes genomic instability in Burkitt’s lymphoma. Oncogene, 2007. 26(35): p. 5115–23. [DOI] [PubMed] [Google Scholar]
- 60.Gruhne B, et al. , The Epstein-Barr virus nuclear antigen-1 promotes genomic instability via induction of reactive oxygen species. Proc Natl Acad Sci U S A, 2009. 106(7): p. 2313–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Kamranvar SA and Masucci MG, The Epstein-Barr virus nuclear antigen-1 promotes telomere dysfunction via induction of oxidative stress. Leukemia, 2011. 25(6): p. 1017–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Gruhne B, Sompallae R, and Masucci MG, Three Epstein-Barr virus latency proteins independently promote genomic instability by inducing DNA damage, inhibiting DNA repair and inactivating cell cycle checkpoints. Oncogene, 2009. 28(45): p. 3997–4008. [DOI] [PubMed] [Google Scholar]
- 63.Chen YR, et al. , Epstein-Barr virus latent membrane protein 1 represses DNA repair through the PI3K/Akt/FOXO3a pathway in human epithelial cells. J Virol, 2008. 82(16): p. 8124–37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Knight JS, Sharma N, and Robertson ES, Epstein-Barr virus latent antigen 3C can mediate the degradation of the retinoblastoma protein through an SCF cellular ubiquitin ligase. Proc Natl Acad Sci U S A, 2005. 102(51): p. 18562–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Saha A, et al. , Epstein-Barr virus nuclear antigen 3C augments Mdm2-mediated p53 ubiquitination and degradation by deubiquitinating Mdm2. J Virol, 2009. 83(9): p. 4652–69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Hammerschmidt W and Sugden B, Replication of epstein-barr viral DNA. Cold Spring Harb Perspect Biol, 2013. 5(1). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Rawlins DR, et al. , Sequence-specific DNA binding of the Epstein-Barr virus nuclear antigen (EBNA-1) to clustered sites in the plasmid maintenance region. Cell, 1985. 42(3): p. 859–68. [DOI] [PubMed] [Google Scholar]
- 68.Ritzi M, et al. , Complex protein-DNA dynamics at the latent origin of DNA replication of Epstein-Barr virus. J Cell Sci, 2003. 116(Pt 19): p. 3971–84. [DOI] [PubMed] [Google Scholar]
- 69.Deng Z, et al. , Telomeric proteins regulate episomal maintenance of Epstein-Barr virus origin of plasmid replication. Mol Cell, 2002. 9(3): p. 493–503. [DOI] [PubMed] [Google Scholar]
- 70.Deng Z, et al. , Telomere repeat binding factors TRF1, TRF2, and hRAP1 modulate replication of Epstein-Barr virus OriP. J Virol, 2003. 77(22): p. 11992–2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Palm W and de Lange T, How shelterin protects mammalian telomeres. Annu Rev Genet, 2008. 42: p. 301–34. [DOI] [PubMed] [Google Scholar]
- 72.Norseen J, et al. , RNA-dependent recruitment of the origin recognition complex. Embo J, 2008. 27(22): p. 3024–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Tatsumi Y, et al. , Involvement of human ORC and TRF2 in pre-replication complex assembly at telomeres. Genes to cells : devoted to molecular & cellular mechanisms, 2008. 13(10): p. 1045–59. [DOI] [PubMed] [Google Scholar]
- 74.Zhou J, Snyder AR, and Lieberman PM, Epstein-Barr virus episome stability is coupled to a delay in replication timing. J Virol, 2009. 83(5): p. 2154–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Buscemi G, et al. , The shelterin protein TRF2 inhibits Chk2 activity at telomeres in the absence of DNA damage. Curr Biol, 2009. 19(10): p. 874–9. [DOI] [PubMed] [Google Scholar]
- 76.Zhou J, et al. , Regulation of Epstein-Barr virus origin of plasmid replication (OriP) by the S-phase checkpoint kinase Chk2. J Virol, 2010. 84(10): p. 4979–87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Dheekollu J, et al. , A role for MRE11, NBS1, and recombination junctions in replication and stable maintenance of EBV episomes. PLoS One, 2007. 2(12): p. e1257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Kudoh A, et al. , Epstein-Barr virus lytic replication elicits ATM checkpoint signal transduction while providing an S-phase-like cellular environment. J Biol Chem, 2005. 280(9): p. 8156–63. [DOI] [PubMed] [Google Scholar]
- 79.Zhang Q, Gutsch D, and Kenney S, Functional and physical interaction between p53 and BZLF1: implications for Epstein-Barr virus latency. Mol Cell Biol, 1994. 14(3): p. 1929–38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Mauser A, et al. , The Epstein-Barr virus immediate-early protein BZLF1 regulates p53 function through multiple mechanisms. J Virol, 2002. 76(24): p. 12503–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Kato M, et al. , Frequent inactivation of A20 in B-cell lymphomas. Nature, 2009. 459(7247): p. 712–6. [DOI] [PubMed] [Google Scholar]
- 82.Li R, et al. , Conserved herpesvirus kinases target the DNA damage response pathway and TIP60 histone acetyltransferase to promote virus replication. Cell Host Microbe, 2011. 10(4): p. 390–400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Hagemeier SR, et al. , The cellular ataxia telangiectasia-mutated kinase promotes epstein-barr virus lytic reactivation in response to multiple different types of lytic reactivation-inducing stimuli. J Virol, 2012. 86(24): p. 13360–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Guo Z, et al. , ATM activation by oxidative stress. Science, 2010. 330(6003): p. 517–21. [DOI] [PubMed] [Google Scholar]
- 85.Saridakis V, et al. , Structure of the p53 binding domain of HAUSP/USP7 bound to Epstein-Barr nuclear antigen 1 implications for EBV-mediated immortalization. Mol Cell, 2005. 18(1): p. 25–36. [DOI] [PubMed] [Google Scholar]
- 86.Gastaldello S, et al. , A deneddylase encoded by Epstein-Barr virus promotes viral DNA replication by regulating the activity of cullin-RING ligases. Nat Cell Biol, 2010. 12(4): p. 351–61. [DOI] [PubMed] [Google Scholar]
- 87.Whitehurst CB, et al. , Epstein-Barr virus BPLF1 deubiquitinates PCNA and attenuates polymerase eta recruitment to DNA damage sites. J Virol, 2012. 86(15): p. 8097–106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Yang K, Moldovan GL, and D’Andrea AD, RAD18-dependent recruitment of SNM1A to DNA repair complexes by a ubiquitin-binding zinc finger. The Journal of biological chemistry, 2010. 285(25): p. 19085–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Sato Y and Tsurumi T, Noise cancellation: viral fine tuning of the cellular environment for its own genome replication. PLoS Pathog, 2010. 6(12): p. e1001158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Chang Y, et al. , Identification of herpesvirus-like DNA sequences in AIDS-associated Kaposi’s sarcoma. Science, 1994. 266(5192): p. 1865–9. [DOI] [PubMed] [Google Scholar]
- 91.Mesri EA, Cesarman E, and Boshoff C, Kaposi’s sarcoma and its associated herpesvirus. Nat Rev Cancer, 2010. 10(10): p. 707–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Ballestas ME, Chatis PA, and Kaye KM, Efficient persistence of extrachromosomal KSHV DNA mediated by latency-associated nuclear antigen. Science, 1999. 284(5414): p. 641–4. [DOI] [PubMed] [Google Scholar]
- 93.Friborg J Jr., et al. , p53 inhibition by the LANA protein of KSHV protects against cell death. Nature, 1999. 402(6764): p. 889–94. [DOI] [PubMed] [Google Scholar]
- 94.Si H and Robertson ES, Kaposi’s sarcoma-associated herpesvirus-encoded latency-associated nuclear antigen induces chromosomal instability through inhibition of p53 function. J Virol, 2006. 80(2): p. 697–709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Radkov SA, Kellam P, and Boshoff C, The latent nuclear antigen of Kaposi sarcoma-associated herpesvirus targets the retinoblastoma-E2F pathway and with the oncogene Hras transforms primary rat cells. Nat Med, 2000. 6(10): p. 1121–7. [DOI] [PubMed] [Google Scholar]
- 96.Verma SC, Borah S, and Robertson ES, Latency-associated nuclear antigen of Kaposi’s sarcoma-associated herpesvirus up-regulates transcription of human telomerase reverse transcriptase promoter through interaction with transcription factor Sp1. J Virol, 2004. 78(19): p. 10348–59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Mittnacht S and Boshoff C, Viral cyclins. Rev Med Virol, 2000. 10(3): p. 175–84. [DOI] [PubMed] [Google Scholar]
- 98.Verschuren EW, et al. , The role of p53 in suppression of KSHV cyclin-induced lymphomagenesis. Cancer Res, 2004. 64(2): p. 581–9. [DOI] [PubMed] [Google Scholar]
- 99.Guasparri I, Keller SA, and Cesarman E, KSHV vFLIP is essential for the survival of infected lymphoma cells. J Exp Med, 2004. 199(7): p. 993–1003. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 100.Lee JS, et al. , FLIP-mediated autophagy regulation in cell death control. Nat Cell Biol, 2009. 11(11): p. 1355–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Koopal S, et al. , Viral oncogene-induced DNA damage response is activated in Kaposi sarcoma tumorigenesis. PLoS Pathog, 2007. 3(9): p. 1348–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Verschuren EW, et al. , The oncogenic potential of Kaposi’s sarcoma-associated herpesvirus cyclin is exposed by p53 loss in vitro and in vivo. Cancer Cell, 2002. 2(3): p. 229–41. [DOI] [PubMed] [Google Scholar]
- 103.Leidal AM, et al. , Subversion of autophagy by Kaposi’s sarcoma-associated herpesvirus impairs oncogene-induced senescence. Cell Host Microbe, 2012. 11(2): p. 167–80. [DOI] [PubMed] [Google Scholar]
- 104.Young AR, et al. , Autophagy mediates the mitotic senescence transition. Genes Dev, 2009. 23(7): p. 798–803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Chaudhary PM, et al. , Modulation of the NF-kappa B pathway by virally encoded death effector domains-containing proteins. Oncogene, 1999. 18(42): p. 5738–46. [DOI] [PubMed] [Google Scholar]
- 106.Liu L, et al. , The human herpes virus 8-encoded viral FLICE inhibitory protein physically associates with and persistently activates the Ikappa B kinase complex. The Journal of biological chemistry, 2002. 277(16): p. 13745–51. [DOI] [PubMed] [Google Scholar]
- 107.Ganem D, KSHV-induced oncogenesis. 2007. [PubMed]
- 108.Jacobs SR and Damania B, The viral interferon regulatory factors of KSHV: immunosuppressors or oncogenes? Front Immunol, 2011. 2: p. 19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Seo T, et al. , Viral interferon regulatory factor 1 of Kaposi’s sarcoma-associated herpesvirus binds to p53 and represses p53-dependent transcription and apoptosis. J Virol, 2001. 75(13): p. 6193–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Shin YC, et al. , Inhibition of the ATM/p53 signal transduction pathway by Kaposi’s sarcoma-associated herpesvirus interferon regulatory factor 1. J Virol, 2006. 80(5): p. 2257–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Tarakanova VL, et al. , Gamma-herpesvirus kinase actively initiates a DNA damage response by inducing phosphorylation of H2AX to foster viral replication. Cell Host Microbe, 2007. 1(4): p. 275–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Walboomers JM, et al. , Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J Pathol, 1999. 189(1): p. 12–9. [DOI] [PubMed] [Google Scholar]
- 113.Gillison ML, Human papillomavirus-associated head and neck cancer is a distinct epidemiologic, clinical, and molecular entity. Semin Oncol, 2004. 31(6): p. 744–54. [DOI] [PubMed] [Google Scholar]
- 114.Moody CA and Laimins LA, Human papillomavirus oncoproteins: pathways to transformation. Nat Rev Cancer, 2010. 10(8): p. 550–60. [DOI] [PubMed] [Google Scholar]
- 115.McBride AA, et al. , Hitchhiking on host chromatin: how papillomaviruses persist. Biochim Biophys Acta, 2012. 1819(7): p. 820–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Dyson N, et al. , The human papilloma virus-16 E7 oncoprotein is able to bind to the retinoblastoma gene product. Science, 1989. 243(4893): p. 934–7. [DOI] [PubMed] [Google Scholar]
- 117.Munger K, et al. , Complex formation of human papillomavirus E7 proteins with the retinoblastoma tumor suppressor gene product. The EMBO journal, 1989. 8(13): p. 4099–105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Munger K, et al. , The E6 and E7 genes of the human papillomavirus type 16 together are necessary and sufficient for transformation of primary human keratinocytes. J Virol, 1989. 63(10): p. 4417–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Amin AA, et al. , Identification of domains of the HPV11 E1 protein required for DNA replication in vitro. Virology, 2000. 272(1): p. 137–50. [DOI] [PubMed] [Google Scholar]
- 120.Androphy EJ, Lowy DR, and Schiller JT, Bovine papillomavirus E2 trans-activating gene product binds to specific sites in papillomavirus DNA. Nature, 1987. 325(6099): p. 70–3. [DOI] [PubMed] [Google Scholar]
- 121.Schwarz E, et al. , Structure and transcription of human papillomavirus sequences in cervical carcinoma cells. Nature, 1985. 314(6006): p. 111–4. [DOI] [PubMed] [Google Scholar]
- 122.Duensing S, et al. , The human papillomavirus type 16 E6 and E7 oncoproteins cooperate to induce mitotic defects and genomic instability by uncoupling centrosome duplication from the cell division cycle. Proc Natl Acad Sci U S A, 2000. 97(18): p. 10002–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Kadaja M, et al. , Mechanism of genomic instability in cells infected with the high-risk human papillomaviruses. PLoS Pathog, 2009. 5(4): p. e1000397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Fradet-Turcotte A, et al. , Nuclear accumulation of the papillomavirus E1 helicase blocks S-phase progression and triggers an ATM-dependent DNA damage response. J Virol, 2011. 85(17): p. 8996–9012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Sakakibara N, Mitra R, and McBride AA, The papillomavirus E1 helicase activates a cellular DNA damage response in viral replication foci. J Virol, 2011. 85(17): p. 8981–95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Reinson T, et al. , Engagement of the ATR-dependent DNA damage response at the human papillomavirus 18 replication centers during the initial amplification. J Virol, 2013. 87(2): p. 951–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Moody CA and Laimins LA, Human papillomaviruses activate the ATM DNA damage pathway for viral genome amplification upon differentiation. PLoS Pathog, 2009. 5(10): p. e1000605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Bester AC, et al. , Nucleotide deficiency promotes genomic instability in early stages of cancer development. Cell, 2011. 145(3): p. 435–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Scheffner M, et al. , The HPV-16 E6 and E6-AP complex functions as a ubiquitin-protein ligase in the ubiquitination of p53. Cell, 1993. 75(3): p. 495–505. [DOI] [PubMed] [Google Scholar]
- 130.Gillespie KA, et al. , Human papillomaviruses recruit cellular DNA repair and homologous recombination factors to viral replication centers. J Virol, 2012. 86(17): p. 9520–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Moody CA, et al. , Human papillomaviruses activate caspases upon epithelial differentiation to induce viral genome amplification. Proc Natl Acad Sci U S A, 2007. 104(49): p. 19541–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Banerjee NS, et al. , Human papillomavirus (HPV) E7 induces prolonged G2 following S phase reentry in differentiated human keratinocytes. The Journal of biological chemistry, 2011. 286(17): p. 15473–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Wang X, et al. , Oncogenic HPV infection interrupts the expression of tumor-suppressive miR-34a through viral oncoprotein E6. RNA, 2009. 15(4): p. 637–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Bartkova J, et al. , Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature, 2006. 444(7119): p. 633–7. [DOI] [PubMed] [Google Scholar]
- 135.Di Micco R, et al. , Oncogene-induced senescence is a DNA damage response triggered by DNA hyper-replication. Nature, 2006. 444(7119): p. 638–42. [DOI] [PubMed] [Google Scholar]
- 136.Fumagalli M, et al. , Telomeric DNA damage is irreparable and causes persistent DNA-damage-response activation. Nat Cell Biol, 2012. 14(4): p. 355–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Suram A, et al. , Oncogene-induced telomere dysfunction enforces cellular senescence in human cancer precursor lesions. Embo J, 2012. 31(13): p. 2839–51. [DOI] [PMC free article] [PubMed] [Google Scholar]