Abstract
Background
Lactobacilli are gatekeepers of vaginal ecosystem impeding growth of pathogenic microbes and their diversity varies across populations worldwide. The present study investigated diversity of human vaginal microbiota among females of Northeast India, who are distinct in dietary habits, lifestyle, and genomic composition from rest of India.
Results
Altogether, 154 bacterial isolates were obtained from vaginal swab samples of 40 pregnant and 29 non-pregnant females. The samples were sequenced for 16 s rRNA gene and analysed for identification using a dual approach of homology search and maximum likelihood based clustering. Molecular identification based on 16S rRNA gene sequence confirmed the isolates belonging to 31 species. Lactobacilli constituted 37.7% of the bacterial isolates with 10 species and other Lactic Acid Bacteria (39.61%) represented another 10 species, some of which are opportunistic pathogens. The remaining of the communities are mostly dominated by species of Staphylococcus (14.28%) and rarely by Propionibacterium avidum (3.90%), Bacillus subtilis, Escherchia coli, Janthinobacterium lividum, and Kocuria kristinae (each 0.64%). Interestingly Lactobacillus mucosae and Enterococcus faecalis, which are globally uncommon vaginal microbes is found dominant among women of Northeast India. This tentatively reflects adaptability of particular Lactobacillus species, in distinct population, to better compete for receptors and nutrients in vaginal epithelium than other species. Further, intrageneric 16S rRNA gene exchange was observed among Enterococcus, Staphylococcus, and two species of Lactobacillus, and deep intraspecies divergence among L. mucosae, which pinpointed possibility of emergence of new strains with evolved functionality. Lactobacilli percentage decreased from young pregnant to aged non-pregnant women with maximum colonization in trimester II.
Conclusion
The study highlighted importance of assessment of vaginal microbiota, Lactobacillus in particular, across different population to gain more insight on female health.
Electronic supplementary material
The online version of this article (10.1186/s12866-019-1568-6) contains supplementary material, which is available to authorized users.
Keywords: Vaginal microbiota, Lactobacillus, Northeast India, Non-pregnant and pregnant women
Background
Lactobacilli are the predominant microbes in the vaginal microenvironment of healthy females [1]. A key feature that favours colonization of Lactobacillus is low vaginal pH due to metabolism of glycogen to lactic acid; glycogen deposits in vaginal epithelial cells in response to rise in circulating estrogen [2, 3]. The predominance of Lactobacillus promotes vaginal health by inhibiting colonization of pathogenic microbes at acidic pH and also by producing antimicrobial bacteriocins [4]. However, the vaginal pH and Lactobacillus levels vary over the life course of women [5]. During bacterial vaginosis (BV), Lactobacillus predominance is lost and replaced by pathogenic microbes, which adversely affects women health making susceptible to HIV, HSV-2, pathogenic microorganisms such as Neisseria gonorrhoeae, Gardnerella vaginalis, Escherchia coli, Staphylococcus aureus, Peptostreptococcus anaerobius, P. bivia, and Chlamydia trachomatis, pelvic inflammatory disease and pre-term delivery [6–13]. Therefore, Lactobacilli are considered gatekeepers of vaginal ecosystem and understanding their qualitative and quantitative diversity is an emphasized area of research [1].
The human microbiome project set out vaginal microenvironment as one of the priority site related to human health and disease and reported around 60 bacterial species that colonizes lower urinogenital tract of female dominated by Lactobacillus crispatus, L. iners, L. gasseri, L. jensenii [1, 14, 15]. Apparently, the diversity and dominance of Lactobacillus varies among different populations, ethnic groups, regions etc. Ravel et al [16] studied Lactobacillus composition in four ethnic groups and found that Asian and White women have higher vaginal Lactobacillus composition, dominated by L. iners and L. crispatus respectively, than Hispanic and Black women, who have higher proportion of strictly anaerobic bacteria and dominant Lactobacillus species is L. iners. In some other studies Lactobacillus composition has been further reported to vary among different populations and as such L. iners, L. crispatus, and L. gasseri were reported predominant among Chinese women [17], L. acidophilus among Mexicans and Swedish [18, 19], L. crispatus, L. iners, L. gasseri, L. jenseni, and L. vaginalis among South Africans [20]. A comparative study on Korean and Uganda women reported that L. fermentum was common in Korean, and L. gasseri, L. reuteri, and L. vaginalis in Ugandan women [21]. In India the biogeographical condition and population varies across different regions and as such the vaginal Lactobacillus species L. iners, L. crispatus, L. reuteri, L. gasseri, and L. jensenii have been found predominant in Mumbai (southwest region of India) [22], L. reuteri, L. fermentum, and L. salivarius in Delhi (central region of India) [23], L. crispatus, L. gasseri, and L. jenseni in Mysore (southern region of India) [24] and many of the regions still remain unexplored. A consensus of all studies explains four Lactobacillus species viz., L. crispatus, L. jenseni, L. gasseri, and L. iners [2] are predominant in vagina and among them L. crispatus and L. jenseni are considered most beneficial [25, 26] and their colonization corresponds to higher vaginal glycogen content [2].
Lactobacillus also restores vaginal health during pregnancy and its imbalance may cause BV and subsequent post-abortal infection, early and late miscarriage, histological chorioamnionitis, postpartum endometritis, preterm premature rupture of membranes and preterm birth [27–29]. In this context, a number of studies [30–35] had characterized vaginal microbiota during different stages of pregnancy in different biogeographical condition. However, such important studies are lacking from India.
Therefore, it is quite clear that the composition of Lactobacillus in vaginal ecosystem varies across population and dependent on many factors such as hormonal de-regulation, poor genital hygiene, improper vaginal douching, random sexual behaviour and in labouring women. Also different species of Lactobacillus offer different level of protection under different vaginal condition. Hence, structuring of vaginal microbiota in general and Lactobacillus in particular in different populations and their link with female health in normal and pregnant condition is highly appealing.
The present study evaluated the prevalence and the quantities of vaginal microbiota in the vagina of healthy fertile and pregnant women of Northeast India using culture-dependent techniques and subsequent 16S rRNA gene sequencing and analysis. Northeast India is genetically distinct from other parts of India and comprises of many tribes mainly of Austro-Asiatic and Tibeto-Burman background [36]. Further, they have unique dietary habits and lifestyle and these features in combination provide unique opportunity and challenges to decipher microbial composition in different habitats of human body [37] and we focussed on vaginal microbiota.
Results
Screening and characterization of samples, culture of bacterial isolates and sequencing
Vaginal swab samples were initially collected from 83 number of reproductive age females having age range 18–35. Among them 14 cases either had history of urinary tract infection, under antibiotic treatment, or has ongoing vaginal infection and excluded from the study (Additional file 1: Table S1). The remaining 69 samples according to age may be categorized as; 23 cases in age group 18–23, 25 cases in age group 24–29, and 21 cases in age group 30–35. The 69 samples further may be categorized as 29 non-pregnant and 40 pregnant cases and the pregnant cases may be categorized as; 12 first (I) trimester, 16 s (II) trimester, and 12 third ((III) trimester. The swab samples were streak plated in Nutrient Agar (NA), de Man, Rogosa and Sharpe (MRS), and Blood Agar (BA) media for microbial growth under ambient condition. While bacterial colonies appeared in NA and MRS media and a total of 154 bacterial isolates were obtained, no growth was observed in BA media. The number of isolates obtained from each sample, their gram staining and colony morphology, sample and colony code and corresponding sequencing code and accession numbers were detailed in Additional file 2: Table S2.
Vaginal microbial diversity of females of Northeast India
The 16S rRNA gene based identification of the samples were based on a dual approach of homology search with database (Additional file 3: Table S3) and characteristic ML clustering (Additional file 6: Figure S1) of derived and their corresponding closely match database sequence. ML clustering was done with a total 270 sequences, that includes 154 derived and 116 database sequences (Additional file 4: Table S4), modelled by Kimura’s 2 parameter with gamma distributed rate across sites that describes best substitution pattern for the dataset according to Bayesian information criterion [38] (Additional file 5: Table S5). The homology search and ML clustering results are presented in a manageable form in Table 1.
Table 1.
Identification of the bacterial isolates based on 16 s rRNA gene sequence
| Sl. No. | Accession No. | No. of sequences derived | Species identified by homology search with database | Characteristic ML clustering as seen in Additional file 6: Figure S1 |
|---|---|---|---|---|
| 1 | KU184494 | 1 | Bacillus subtilis | Clustered distinct |
| 2 | KT906577 KT906579 KT906580 | 3 | Bifidobacterium breve | Clustered distinct |
| 3 | KR264992 | 1 | Enterococcus avium | The four species of Enterococcus genus clustered cohesively |
| 4 | KR265008 KT906587 KU184461 | 39 | Enterococcus faecalis | |
| KU184462 KU184464 KU184465 | ||||
| KU184460 KT906572 KU184469 | ||||
| KU184496 KU184470 KU184471 | ||||
| KU184472 KT906573 KU184498 | ||||
| KU184474 KT589112 KT589114 | ||||
| KT589115 KT589124 KR265004 | ||||
| KT589118 KT005519 KT005522 | ||||
| KT589119 KT589120 KT589122 | ||||
| KT589125 KT589126 KT589137 | ||||
| KT589136 KT589128 KT589105 | ||||
| KT589107 KT589108 KT589109 | ||||
| KT589110 KU184479 KT589111 | ||||
| 5 | KP775931 KP775933 | 2 | Enterococcus faecium | |
| 6 | KU184480 KR264993 KR264996 | 6 | Enterococcus hirae | |
| KU184476 KU184477 KU184478 | ||||
| 7 | KT005520 KT589121 | 2 | Escherichia coli | Clustered distinct |
| 8 | KP747671 KT597700 KP775922 | 3 | Janthinobacterium lividum | Clustered distinct |
| 9 | KT906585 | 1 | Kocuria kristinae | Clustered distinct |
| 10 | KT361204 KT589116 KT589117 | 5 | Lactobacillus brevis | Clustered distinct |
| KT589132 KT589133 | ||||
| 11 | KU184473 KU184499 KP775929 | 10 | Lactobacillus fermentum | Clustered distinct |
| KT589130 KP775934 KP775923 | ||||
| KT589106 KR264983 KR264984 | ||||
| KR264986 | ||||
| 12 | KP775927 | 1 | Lactobacillus gallinarum | Clustered together with Lactobacillus helviticus |
| 13 | KT991842 KT991843 KU184463 | 6 | Lactobacillus gasseri | Clustered distinct |
| KU184466 KP775925 KR264987 | ||||
| 14 | KT361211 KT361209 KP775926 | 3 | Lactobacillus helveticus | Clustered together with Lactobacillus gallinarum |
| 15 | KU184500 KU184475 | 2 | Lactobacillus ingluviei | Clustered distinct |
| 16 | KT906576 KT906578 KT906588 | 3 | Lactobacillus jensenii | Clustered distinct |
| 17 | KT835013 KT835015 KT835018 | 25 | Lactobacillus mucosae* | Clustered distinct and showed deep intraspecies divergence |
| KT835007 KT835008 KT835009 | ||||
| KT835010 KT835011 KT835012 | ||||
| KT991844 KU184467 KU184468 | ||||
| KU184495 KU184497 KT597696 | ||||
| KU184483 KU184484 KU184485 | ||||
| KU184486 KU184487 KU184488 | ||||
| KU184489 KU184490 KU184491 | ||||
| KR264981 | ||||
| 18 | KP775924 | 1 | Lactobacillus parafarraginis | Clustered distinct |
| 19 | KR264991 KR264988 | 2 | Lactobacillus reuteri | Clustered distinct |
| 20 | KT906581 KT906582 KT906583 | 6 | Propionibacterium avidum | Clustered distinct |
| KT906584 KT906571 KU184493 | ||||
| 21 | KT906569 | 1 | Staphylococcus capitis | The six species of Staphylococcus genus clustered cohesively |
| 22 | KR264994 KR264997 KR265000 | 3 | Staphylococcus caprae | |
| 23 | KR265006 KT361206 KT361207 | 8 | Staphylococcus epidermidis | |
| KP747672 KT589123 KT589127 | ||||
| KT589131 KT589135 | ||||
| 24 | KT005521 | 1 | Staphylococcus haemolyticus | |
| 25 | KT589113 KP775928 KR264995 | 8 | Staphylococcus hominis | |
| KR264998 KR265001 KP775932 | ||||
| KR264999 KR264989 | ||||
| 26 | KT906586 | 1 | Staphylococcus kloosii | |
| 27 | KT906589 KT906574 KT906575 | 3 | Streptococcus agalactiae | Clustered distinct |
| 28 | KT835014 KT906568 KT906570 | 3 | Streptococcus anginosus | Clustered distinct |
| 29 | KT835016 KT835017 | 2 | Streptococcus gallolyticus | Clustered distinct |
| 30 | KR264982 | 1 | Streptococcus infantarius | Clustered distinct |
| 31 | KT361205 | 1 | Weissella confusa | Clustered distinct |
The Genbank accession number (column 2) of 16sRNA gene sequence of each of the 154 bacterial isolates and their corresponding similarity search result (column 4) (detailed in Additional file 3: Table S3) and characteristic ML clustering (column 5) (detailed in Additional file 6: Figure S1) is shown. Overall the 154 bacterial isolates were identified belonging to 31 species. Enterococcus and Staphylococcus, and two species of Lactobacillus (L. gallinarum and L. helveticus) showed cohesive intrageneric clustering in contrast to, cohesive cluster of all sequences of a particular species and distinct cluster with all other species sequences, and are shown here as bold. Lactobacillus mucosae showed deep intraspecies split and marked with asterisk (*)
Consensus match of derived sequences with a particular microbe in both the databases (NCBI and EZ-Taxon) in the range of similarity 97–100% and cohesive clustering of conspecific derived and database sequences confirmed straight forward identification of 80 sequences into 19 species. This includes Bacillus subtilis, Bifidobacterium breve, Escherichia coli, Janthinobacterium lividum, Kocuria kristinae, Lactobacillus brevis, L. fermentum, L. gasseri, L. ingluviei, L. jensenii, L. mucosae, L. parafarraginis, L. reuteri, Propionibacterium avidum, Streptococcus agalactiae, S. anginosus, S. gallolyticus, S. infantarius, and Weissella confusa. Among them Lactobacillus mucosae showed deep intraspecies split. One derived sequence (KP775927) that showed high similarity (98–99%) with Lactobacillus gallinarum and 3 sequences (KT361211, KT361209, and KP775926) that showed high similarity (97–99%) with L. helveticus based on homology search, clustered together with database sequences of L. gallinarum and L. helveticus. Similarly, 48 sequences identified as four species of Enterococcus (E. avium, E. faecalis, E. faecium, and E. hirae) based on homology search, and database sequences of the four Enterococcus species clustered together. Also, 22 sequences identified as 6 species of Staphylococcus (S. capitis, S. caprae, S. epidermidis, S. haemolyticus, S. hominis, and S. kloosii) and database sequences of the six Staphylococcus species clustered together.
In summary Enterococcus faecalis (25.3%) and Lactbacillus mucosae (16.2%) were found dominant species among the study population. 37.7% of the bacterial isolates were Lactobacilli, that includes 10 species, and their relative dominance is; L. mucosae (16.2%) L. fermentum (6.4%), L. gasseri (3.9%), L. brevis (3.2%), L helveticus and L. jensenii each 2%, L. ingluviei and L. reuteri each 1.3%, and L. gallinarum and L. parafarraginis each 0.6%. Clearly Lactobacilli percentage significantly varied among age group (p < 0.001) and we found in age group 18–23, 87% of females harbour Lactobacillus, in age group 24–29, 76% females harbour Lactobacillus, and in age group 30–35, 71.4% females harbour Lactobacillus.
Lactobacillus composition in non-pregnant and pregnant women
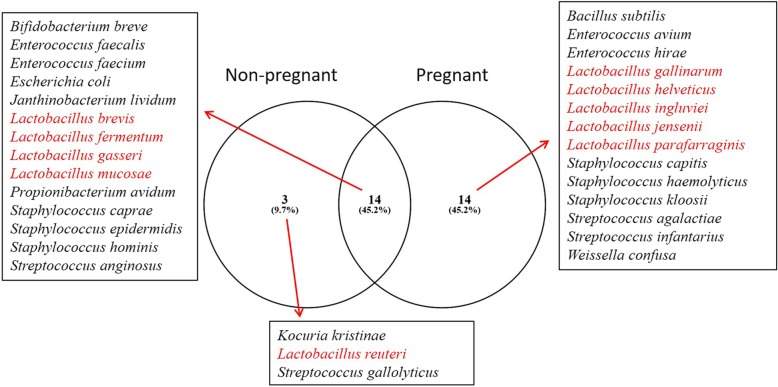
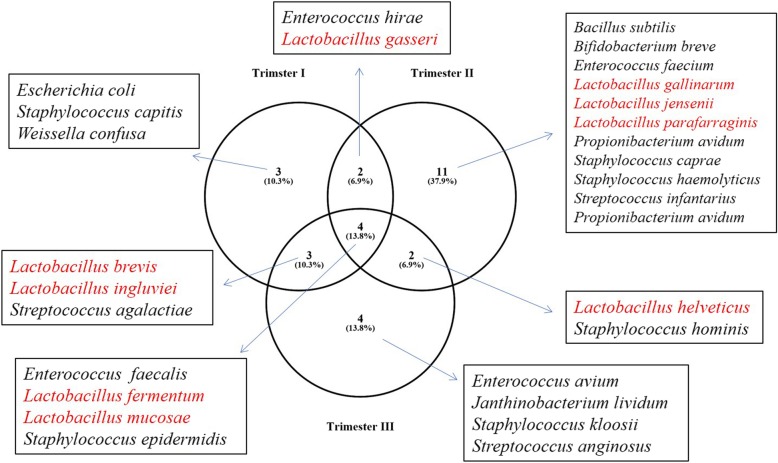
One major objective of the present study is to categorize Lactobacillus diversity in non-pregnant and pregnant women and in three trimester of pregnant women. It is clearly evident in this study that the diversity of Lactobacillus increases in labouring women. As such we found only L. reuteri colonized non-pregnant women; 4 species L. brevis, L. fermentum, L. gasseri, and L. mucosae colonized both non-pregnant and pregnant women; and 5 species L. gallinarum, L. helveticus, L. ingluviei, L. jensenii, and L. parafarraginis colonized only pregnant women (Fig. 1). Trimester-wise the diversity of Lactobacillus is as follows: L. fermentum and L. mucosae overlapped in all three trimester, L. brevis and L. ingluviei overlapped in I and III trimester, L. gasseri overlapped in I and II trimester, L. helveticus overlapped in II and III trimester, while 3 species L. gallinarum, L. jensenii, and L. parafarraginis were found in only II trimester (Fig. 2). Again Lactobacilli percentage significantly varied among non-pregnant and pregnant women (p < 0.0002) and we found 58.6% of studied non-pregnant females harbour Lactobacillus while 90% of studied pregnant females harbour Lactobacillus, which may be further categorized as 83.3% trimester I, 93.8% trimester II, and 91.7% trimester III.
Fig. 1.
Bacterial colonization in non-pregnant and pregnant women. The numbers under each circle corresponds to number of species found under each condition and in parenthesis relative percentage shown. Clearly colonization is more in pregnant women than non-pregnant women
Fig. 2.
Bacterial colonization in three trimester of pregnant women. The numbers under each circle corresponds to number of species found under each trimester of pregnant women and in parenthesis relative percentage shown. Clearly bacterial colonization is higher in trimester II
Discussion
In women the vaginal ecosystem is dominated by Lactobacilli and this system has evolved to protect females health by inhibiting growth of pathogenic microbes. However, the characteristic microbial diversity varies across population [4, 16, 39–42]. Women in different settings face different geographical condition, dietary habits, and lifestyle. In particular, peoples of Northeast India are from Austro-Asiatic and Tibeto-Burman background and are genetically different from other parts of India [36, 43, 44]. Therefore, it is tempting to screen and compare how the vaginal microbiota is structured in females in Northeast India and how it differs from other population.
In this study altogether we isolated 154 bacterial isolates in NA and MRS media from 69 females; which includes 40 pregnant and 29 non-pregnant cases. BA was used as a selective media for Lactobacilli, especially Lactobacillus iners, however, no colonies appeared in BA. Previous studies [45–47] advocates that L. iners grows in transitional phase of BV and healthy state, which is not the case in this study, and this tentatively is the reason for no growth in BV. However, we believe a detail sampling including healthy and BV cases in future studies may provide a comprehensive insight on Lactobacillus diversity among the females of Northeast India.
A dual approach of homology search and ML clustering confirmed the 154 bacterial isolates belonging to 31 species and 37.7% of the bacterial isolates are Lactobacillus. Interestingly, we found L. mucosae is the dominant species and L. fermentum, L. gasseri, L. brevis, L. helveticus, L. jensenii, L. ingluviei, L. reuteri, L. gallinarum, and L. parafarraginis together forms the Lactobacilli complex that colonizes vaginal microenvironment among the females of Northeast India. This is the first report of its type from Northeast India and has implications for global comparison of Lactobacilli composition. Clearly, vaginal Lactobacillus composition among Northeast India population is different from other parts of India [22–24] and various regions/countries of the World [16–21]. Lactobacillus mucosae has been reported as less common vaginal microflora in previous studies [41, 48], however dominance of L. mucosae than L. crispatus and L. jensenii (as observed in most of the previous studies [2, 16]) observed in this study in Northeast Indian females is an intriguing finding and tentatively reflects different vaginal set-up. In particular, different Lactobacillus species possess unique repertoire of protein families which helps in specific community adaptations [49, 50]. Lactobacillus mucosae has been reported to express Lam29 protein in intestinal epithelial cells, that significantly improves its adhesion for human blood group A and B antigens and out competes pathogens to protect intestinal mucosa [51]. In human vagina, the epithelial layer similarly acts as a barrier for mucosal layer, which the pathogen wants to penetrate and Lactobacillus wants to protect by adhesion and outcompeting pathogens [52–55]. However, this adhesion is dependent on genetic repertoire of both the host and the bacteria [49, 50]. Further, the adhesion and colonization of Lactobacillus is also dependent on hormonal level (especially estrogen), glycogen content of epithelial cell, and availability of nutrients [3]. We hypothesize Lactobacillus mucosae outcompetes other organisms for receptors and nutrients in the vaginal epithelial cells of females of Northeast India and this claim needs more insight from future studies. Nevertheless, this finding is also tempting to explore probiotic potential of L. mucosae than L. crispatus and L. jensenii among women of Northeast India and correlation with estrogen level and deposition of glycogen in vaginal epithelium.
Besides Lactobacillus that constituted 37.7% of bacterial isolates, we also found 10 other lactic acid bacteria species (39.61%) viz., Bifidobacterium breve, Enterococcus avium, E. faecalis, E. faecium, E. hirae, Streptococcus agalactiae, S. anginosus, S. gallolyticus, S. infantarius, and Weissella confusa with predominance of Enterococcus faecalis (25.3%). Among them S. infantarius, S. gallolyticus, S. agalactiae, S. anginosus, E. avium, E. hirae are opportunistic pathogens and causes severe invasive infections [56–60].
The study also reports 4 species of Enterococcus viz., E. avium, E. faecalis, E. faecium, and E. hirae, 6 species of Staphylococcus (14.28%) viz., S. capitis, S. caprae, S. epidermidis, S. haemolyticus, S. hominis, and S. kloosii and 2 species of Lactobacillus viz., L. gallinarum, and L. helveticus with cohesive congeneric clustering. This clearly reveals congeneric exchange of 16S rRNA gene segment, may be due to horizontal gene transfer or homologous recombination, which is common among microbes [61]. These key findings together with the observation of deep intraspecies split in Lactobacillus mucosae warrants genome-wide study of Enterococcus, Staphylococcus and Lactobacillus in future studies to understand intraspecies genomic variability, possible emergence of new strains/variants and its impact on female health of Northeast India. Nevertheless, Propionibacterium avidum (3.90%), Bacillus subtilis, Escherchia coli, Janthinobacterium lividum, and Kocuria kristinae (each 0.64%) represented renaming of the microbiota found in this study. Lactobacillus ingluviei and L. parafarraginis found in this study have never been previously reported from human vagina [62, 63] and so this is the first case of their report from human vagina.
The vaginal microbiota have a major impact during different stages of pregnancy [30] and therefore we assessed how the microbes, and Lactobacillus in particular, are structured among non-pregnant and different stages of pregnant women. We found Lactobacillus percentage is highest in the age group 18–23 (87%) and successively decreases (71–76%). This finding is consistent with previous studies [64] and therefore as female attains age and Lactobacillus composition decreases, the pathogenic microbes have higher chance of colonization and thus extra precaution must be followed. In particular, aged women may be supplemented with Lactobacillus probiotic, which is predominant in their population, since it is observed herein and some previous studies [4, 16, 39–42] that predominance of Lactobacillus species varies across population. The Lactobacillus composition in pregnant women showed a sharp increase (90%) compared to non-pregnant women (58.6%) similar to earlier study [65] and this perhaps is the best way to protect vaginal health in labour periods. We further evidenced higher colonization of Lactobacilli in trimester II followed by trimester III and I. These features must be explored in more details for specific functional characterization of different species of Lactobacillus in different stages of pregnancy.
Conclusion
The study clearly highlighted importance of assessment of vaginal microbiota in different populations, groups, ethnics, religions etc. The vaginal microbial composition studied herein revealed some interesting new information’s such as: i) Lactobacillus mucosae rather than L. crispatus or L. iners is dominant among females of Northeast India and this may be related to Lacotbacillus adaptation in particular population mainly due to genetic variability, ii) deep intraspecies split among L. mucosae revealed population level divergence and may be due to evolution of various strains active under specific conditions, iii) Lactobacilli percentage among Northeast India females is less and tentatively compensated by other LABs, iv) parallel to L. mucosae, Enterococcus faecalis is also dominant among females of Northeast India and its specific role must be explored. The Lactobacilli composition in non-pregnant and pregnant cases is consistent to previous studies however specific trimester-wise colonization of particular Lactobacillus species needs to be explored in more details. This study is the first report of its type from Northeast India and although we used culture-dependent technique, we identified around 31 vaginal microbial species and importantly revealed unique Lactobacilli diversity and dominance. We believe our study provided an interesting clue on vaginal Lactobacillus composition. This is tempting for future studies to include broad spectrum culture independent technique together with culture dependent technique for better understanding deficiency and dominance of particular vaginal Lactobacillus species and its relation to female health of Northeast India; Northeast India females have high rate of cervical cancer and Lactobacilli have major role in acquisition and persistence of Human Papilloma Virus [66] and regulation of cervical cancer [67].
Methods
Sample collection and culture of microbes
Vaginal swab samples were collected from 83 numbers of healthy pregnant and non-pregnant women during the period from 23 July 2012 to 29 November 2015 (Additional file 1: Table S1). A conscious effort was made to collect samples from volunteers who did not have any apparent vaginal infection. The present experiment excluded the volunteers who had antibiotic treatment in last one and half months, and who has history of urinary tract infection in last 6 months, any homogenous white discharge from vagina, and whose pH higher than 4.5, and who have the problem of itching, bleeding from vagina prior to sampling. All women were at reproductive age i.e. 18–35. Necessary clearance from the Institutional Ethical Committee, Karimganj College (KC/IEC/2012/M-1/10, dated: 23 Jun 2012) before the commencement of the study was obtained. The samples were collected aseptically by a practicing gynaecologist of the Red Cross Society of India, Karimganj, Assam (India). Initially, the experimental procedure was explained to the volunteers and when they agreed to participate in the study, they were asked to sign a written consent which was prepared as per the recommendation of the Institutional Ethical Committee. The collection of the samples was made after obtaining formal consent. A sterile swab (Hi Media Laboratory Pvt. Ltd., India) was inserted into the vaginal lumen and the samples were collected from the posterior fornix of vagina. The swabs were collected separately in sterile tubes. The swabs were used within 3 h of collection. The microbes from the swabs were transferred to sterile NA, MRS Agar, and BA media by streak-plate method. It is to be mentioned that while MRS and BA are selective media for Lactobacilli, NA was used as a general starter media to get a general view on microbes present apart from Lactobacillus in vaginal microenvironment of females of NE India. The plates were incubated for 48 h at 37 °C both under aerobic and anaerobic condition to allow the growth of microbial colonies. During collection of samples a statistical approach called convenience method/sampling was applied [68] (Additional file 1: Table S1).
DNA isolation and PCR
The microbial genomic DNA was isolated following the protocol as mentioned in our previous paper [69]. The amplification of the 16S rRNA gene was done using published universal primers 27F Forward primer (AGAGTTTGATCMTGGCTCAG) and 1492R Reverse primer (5ˈ-GGTTACCTTGTTACGACTT-3ˈ) [70]. The 10 μl PCR reaction mixture contained template DNA 1 μl, 20 picomole of each primer, Master Mix 2X (HiMedia, India) 5 μl, and nuclease-free water 2 μl. The PCR thermal condition includes initial denaturation at 94 °C for 2 min, 35 cycles of 94 °C for 30 s, 49.3 °C for 30 s, 72 °C for 1 min and final extension at 72 °C for 3 min in S1100 thermal cycler (BioRad, USA). The PCR products were visualized in 1% agarose gel and images were captured by a Gel DOC machine (BioRad, USA).
16S rRNA gene sequencing
The 1.5 Kb amplicons were sequenced using the facilities provided by the MERK, Bangalore, India and Xcelris, Hyderabad, India. After obtaining the sequencing data, these were analysed and edited using Sequence scanner software v1.0 [57] (Applied Biosystem Inc., CA, USA). Alignments are prepared by ClustalX [71] and DNA contigs. Thereafter, the sequence was matched using BLASTn (NCBI database) [72] to identify its closest relatives. The sequences were submitted in the NCBI database using the submission tool BankIt and the accession numbers were obtained.
Identification of the microbes
For Identification of the microbes we followed a dual approach of sequence based homology search and clustering (Additional file 7: Figure S2). For homology search we compared a particular sequence for similarity with two separate databases NCBI using BLASTn approach [72] and EZ-Taxon (www.eztaxon.org/). For clustering we used Maximum Likelihood method [73] structured by Kimura’s 2 parameter [57] model and the computation platform used was MEGAX [74]. The high similarity (97–100%) of a sequence with database sequences of a particular microbe coupled with cohesive clustering of developed and database sequence of the particular microbe and distinct cluster with respect of other developed and database congeners sequences confirmed identification of the microbe with high precision.
The Lactobacilli percentage across different age group and pregnant and non-pregnant women were compared using Student’s t-test.
Additional files
Table S1. Sample collection information. (DOCX 28 kb)
Table S2. Cell morphology, Gram staining of the bacterial isolates and their corresponding GenBank accession numbers of 16S rRNA gene sequence. (DOCX 36 kb)
Table S3. Homology search of the sequences in database. (DOCX 38 kb)
Table S4. Developed and database sequences. (XLSX 13 kb)
Table S5. Model test result. (XLS 49 kb)
Figure S1. Maximum Likelihood based clustering of developed and database sequences. The sequences of same species clustered together and distinct with respect to sequences of other species. (PDF 50 kb)
Figure S2. 16S rRNA gene based identification of the microbes is based on dual approach of homology search and ML clustering. To confirm identification we considered consensus match of our query sequence with a particular microbe on both the database and cohesive clustering of developed and database sequence of the particular microbe (here shown as Species A and Species B) and distinct with respect to other microbes (all species A members clustered distinct with all species B members). (JPG 250 kb)
Acknowledgements
The authors are very grateful to all the volunteer women involved in this study, who dedicated their time, effort and samples for this research. Authors acknowledge with thanks financial assistance received from Department of Biotechnology (DBT), Govt. of India (GOI) under NER TWINNING PROJECT vide DBT Sanction Order No. BT/220/NE/TBP/2011 (dated 30th May, 2012) and DBT, GOI sponsored Advanced Institutional Biotech Hub, Karimganj College (BCIL/NER-BPMC/2010; dated 30th November, 2010) for providing infrastructural facility.
Authors’ contributions
SDP, MKB, HKP, KP and HU, conceived this study. SDL, provided samples. SDP, MKB, HKP, designed wet-lab experiments. SDP, MD, carried out wet-lab experiments. MJB, designed and performed computational analysis, prepared tables and figures and interpreted results. GDS, KP, HKP, MJB, SDL and HU, offered technical consultation and professional suggestions. MJB and SDP, wrote the manuscript. All authors have read and approved the final manuscript.
Funding
Department of Biotechnology (DBT), Govt. of India (GOI) under NER TWINNING PROJECT vide DBT Sanction Order No. BT/220/NE/TBP/2011 (dated 30th May, 2012) funded the project for design, collection, analysis, and interpretation of the data and fellowship to S.D.P. DBT sponsored Advanced Institutional Biotech hub (BCIL/NER-BPMC/2010; dated 30 Nov, 2010) provided infrastructural support.
Availability of data and materials
All the sequences used in this study have been submitted to NCBI and the details can be checked using their respective accession numbers (please see Additional file 2: Table S2) in NCBI (https://www.ncbi.nlm.nih.gov/).
Ethics approval and consent to participate
Ethical approval was obtained for this study from the Institutional Ethical Committee, Karimganj College (Ref No. KC/IEC/2012/M-1/10, dated: 23 Jun 2012). Written informed consent was obtained from all the participants.
Consent for publication
Not applicable
Competing interests
The authors declare that they have no competing interest.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Mrinal Kanti Bhattacharya, Email: mrinalkxj@yahoo.co.in.
Himanshu Kishore Prasad, Email: himanshu17@gmail.com.
Maloyjo Joyraj Bhattacharjee, Email: mjoyraj@gmail.com.
References
- 1.Amabebe E, Anumba DOC. The vaginal microenvironment: the physiologic role of lactobacilli. Front Med (Lausanne) 2018;5:181. doi: 10.3389/fmed.2018.00181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Mirmonsef P, Hotton AL, Gilbert D, Burgad D, Landay A, Weber KM, Cohen M, Ravel J, Spear GT. Free glycogen in vaginal fluids is associated with Lactobacillus colonization and low vaginal pH. PLoS One. 2014;9(7):e102467. doi: 10.1371/journal.pone.0102467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Mirmonsef P, Hotton AL, Gilbert D, Gioia CJ, Maric D, Hope TJ, Landay AL, Spear GT. Glycogen levels in undiluted genital fluid and their relationship to vaginal pH, estrogen, and progesterone. PLoS One. 2016;11(4):e0153553. doi: 10.1371/journal.pone.0153553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Borges S, Silva J, Teixeira P. The role of lactobacilli and probiotics in maintaining vaginal health. Arch Gynecol Obstet. 2014;289(3):479–489. doi: 10.1007/s00404-013-3064-9. [DOI] [PubMed] [Google Scholar]
- 5.Huang B, Fettweis JM, Brooks JP, Jefferson KK, Buck GA. The changing landscape of the vaginal microbiome. Clin Lab Med. 2014;34(4):747–761. doi: 10.1016/j.cll.2014.08.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Castro J, Martins AP, Rodrigues ME, Cerca N. Lactobacillus crispatus represses vaginolysin expression by BV associated Gardnerella vaginalis and reduces cell cytotoxicity. Anaerobe. 2018;50:60–63. doi: 10.1016/j.anaerobe.2018.01.014. [DOI] [PubMed] [Google Scholar]
- 7.Marrazzo JM. Biomedical prevention of Hiv in women: challenges and approaches, with particular reference to the vaginal microbiome. Trans Am Clin Climatol Assoc. 2018;129:63–73. [PMC free article] [PubMed] [Google Scholar]
- 8.Graver MA, Wade JJ. The role of acidification in the inhibition of Neisseria gonorrhoeae by vaginal lactobacilli during anaerobic growth. Ann Clin Microbiol Antimicrob. 2011;10:8. doi: 10.1186/1476-0711-10-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Matu MN, Orinda GO, Njagi EN, Cohen CR, Bukusi EA. In vitro inhibitory activity of human vaginal lactobacilli against pathogenic bacteria associated with bacterial vaginosis in Kenyan women. Anaerobe. 2010;16(3):210–215. doi: 10.1016/j.anaerobe.2009.11.002. [DOI] [PubMed] [Google Scholar]
- 10.Skarin A, Sylwan J. Vaginal lactobacilli inhibiting growth of Gardnerella vaginalis, Mobiluncus and other bacterial species cultured from vaginal content of women with bacterial vaginosis. Acta Pathol Microbiol Immunol Scand B. 1986;94(6):399–403. doi: 10.1111/j.1699-0463.1986.tb03074.x. [DOI] [PubMed] [Google Scholar]
- 11.Strus M, Malinowska M, Heczko PB. In vitro antagonistic effect of Lactobacillus on organisms associated with bacterial vaginosis. J Reprod Med. 2002;47(1):41–46. [PubMed] [Google Scholar]
- 12.Tyssen D, Wang YY, Hayward JA, Agius PA, DeLong K, Aldunate M, Ravel J, Moench TR, Cone RA, Tachedjian G. Anti-HIV-1 Activity of Lactic Acid in Human Cervicovaginal Fluid. mSphere. 2018;3(4). [DOI] [PMC free article] [PubMed]
- 13.Srinivasan S, Hoffman NG, Morgan MT, Matsen FA, Fiedler TL, Hall RW, Ross FJ, McCoy CO, Bumgarner R, Marrazzo JM, et al. Bacterial communities in women with bacterial vaginosis: high resolution phylogenetic analyses reveal relationships of microbiota to clinical criteria. PLoS One. 2012;7(6):e37818. doi: 10.1371/journal.pone.0037818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Aagaard K, Petrosino J, Keitel W, Watson M, Katancik J, Garcia N, Patel S, Cutting M, Madden T, Hamilton H, et al. The human microbiome project strategy for comprehensive sampling of the human microbiome and why it matters. FASEB J. 2013;27(3):1012–1022. doi: 10.1096/fj.12-220806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kroon SJ, Ravel J, Huston WM. Cervicovaginal microbiota, women’s health, and reproductive outcomes. Fertil Steril. 2018;110(3):327–336. doi: 10.1016/j.fertnstert.2018.06.036. [DOI] [PubMed] [Google Scholar]
- 16.Ravel J, Gajer P, Abdo Z, Schneider GM, Koenig SS, McCulle SL, Karlebach S, Gorle R, Russell J, Tacket CO, et al. Vaginal microbiome of reproductive-age women. Proc Natl Acad Sci U S A. 2011;108(Suppl 1):4680–4687. doi: 10.1073/pnas.1002611107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhang R, Daroczy K, Xiao B, Yu L, Chen R, Liao Q. Qualitative and semiquantitative analysis of Lactobacillus species in the vaginas of healthy fertile and postmenopausal Chinese women. J Med Microbiol. 2012;61(Pt 5):729–739. doi: 10.1099/jmm.0.038687-0. [DOI] [PubMed] [Google Scholar]
- 18.Hernandez-Rodriguez C, Romero-Gonzalez R, Albani-Campanario M, Figueroa-Damian R, Meraz-Cruz N, Hernandez-Guerrero C. Vaginal microbiota of healthy pregnant Mexican women is constituted by four Lactobacillus species and several vaginosis-associated bacteria. Infect Dis Obstet Gynecol. 2011;2011:851485. doi: 10.1155/2011/851485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Vasquez A, Jakobsson T, Ahrne S, Forsum U, Molin G. Vaginal lactobacillus flora of healthy Swedish women. J Clin Microbiol. 2002;40(8):2746–2749. doi: 10.1128/JCM.40.8.2746-2749.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pendharkar S, Magopane T, Larsson PG, de Bruyn G, Gray GE, Hammarstrom L, Marcotte H. Identification and characterisation of vaginal lactobacilli from south African women. BMC Infect Dis. 2013;13:43. doi: 10.1186/1471-2334-13-43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jin L, Tao L, Pavlova SI, So JS, Kiwanuka N, Namukwaya Z, Saberbein BA, Wawer M. Species diversity and relative abundance of vaginal lactic acid bacteria from women in Uganda and Korea. J Appl Microbiol. 2007;102(4):1107–1115. doi: 10.1111/j.1365-2672.2006.03147.x. [DOI] [PubMed] [Google Scholar]
- 22.Pramanick R, Parab S, Mayadeo N, Warke H, Aranha C. Diversity of cultivable vaginal microbiota in asymptomatic women of reproductive age in Mumbai, India. Can J Biotechnol. 2017;1:129. doi: 10.24870/cjb.2017-a115. [DOI] [Google Scholar]
- 23.Garg KB, Ganguli I, Das R, Talwar GP. Spectrum of Lactobacillus species present in healthy vagina of Indian women. Indian J Med Res. 2009;129(6):652–657. [PubMed] [Google Scholar]
- 24.Madhivanan P, Alleyn HN, Raphael E, Krupp K, Ravi K, Nebhrajani R, Arun A, Reingold AL, Riley LW, Klausner JD. Identification of culturable vaginal Lactobacillus species among reproductive age women in Mysore, India. J Med Microbiol. 2015;64(6):636–641. doi: 10.1099/jmm.0.000070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Martinez RC, Franceschini SA, Patta MC, Quintana SM, Nunes AC, Moreira JL, Anukam KC, Reid G, De Martinis EC. Analysis of vaginal lactobacilli from healthy and infected Brazilian women. Appl Environ Microbiol. 2008;74(14):4539–4542. doi: 10.1128/AEM.00284-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Verstraelen H, Verhelst R, Claeys G, De Backer E, Temmerman M, Vaneechoutte M. Longitudinal analysis of the vaginal microflora in pregnancy suggests that L crispatus promotes the stability of the normal vaginal microflora and that L gasseri and/or L iners are more conducive to the occurrence of abnormal vaginal microflora. BMC Microbiol. 2009;9:116. doi: 10.1186/1471-2180-9-116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Chu DM, Seferovic M, Pace RM, Aagaard KM. The microbiome in preterm birth. Best Pract Res Clin Obstet Gynaecol. 2018. [DOI] [PubMed]
- 28.Jacobsson B, Pernevi P, Chidekel L, Jorgen Platz-Christensen J. Bacterial vaginosis in early pregnancy may predispose for preterm birth and postpartum endometritis. Acta Obstet Gynecol Scand. 2002;81(11):1006–1010. doi: 10.1034/j.1600-0412.2002.811103.x. [DOI] [PubMed] [Google Scholar]
- 29.Petricevic L, Domig KJ, Nierscher FJ, Sandhofer MJ, Fidesser M, Krondorfer I, Husslein P, Kneifel W, Kiss H. Characterisation of the vaginal Lactobacillus microbiota associated with preterm delivery. Sci Rep. 2014;4:5136. doi: 10.1038/srep05136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Avershina E, Slangsvold S, Simpson MR, Storro O, Johnsen R, Oien T, Rudi K. Diversity of vaginal microbiota increases by the time of labor onset. Sci Rep. 2017;7(1):17558. doi: 10.1038/s41598-017-17972-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Freitas AC, Chaban B, Bocking A, Rocco M, Yang S, Hill JE, Money DM, Group VR The vaginal microbiome of pregnant women is less rich and diverse, with lower prevalence of Mollicutes, compared to non-pregnant women. Sci Rep. 2017;7(1):9212. doi: 10.1038/s41598-017-07790-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Haque MM, Merchant M, Kumar PN, Dutta A, Mande SS. First-trimester vaginal microbiome diversity: a potential indicator of preterm delivery risk. Sci Rep. 2017;7(1):16145. doi: 10.1038/s41598-017-16352-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.MacIntyre DA, Chandiramani M, Lee YS, Kindinger L, Smith A, Angelopoulos N, Lehne B, Arulkumaran S, Brown R, Teoh TG, et al. The vaginal microbiome during pregnancy and the postpartum period in a European population. Sci Rep. 2015;5:8988. doi: 10.1038/srep08988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nuriel-Ohayon M, Neuman H, Koren O. Microbial changes during pregnancy, birth, and infancy. Front Microbiol. 2016;7:1031. doi: 10.3389/fmicb.2016.01031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Vuillermin PJ, Macia L, Nanan R, Tang ML, Collier F, Brix S. The maternal microbiome during pregnancy and allergic disease in the offspring. Semin Immunopathol. 2017;39(6):669–675. doi: 10.1007/s00281-017-0652-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Basu A, Mukherjee N, Roy S, Sengupta S, Banerjee S, Chakraborty M, Dey B, Roy M, Roy B, Bhattacharyya NP, et al. Ethnic India: a genomic view, with special reference to peopling and structure. Genome Res. 2003;13(10):2277–2290. doi: 10.1101/gr.1413403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Shetty SA, Marathe NP, Shouche YS. Opportunities and challenges for gut microbiome studies in the Indian population. Microbiome. 2013;1(1):24. doi: 10.1186/2049-2618-1-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Schwarz G. Estimating the dimension of a model. Ann Stat. 1978;6(2):461–464. doi: 10.1214/aos/1176344136. [DOI] [Google Scholar]
- 39.Gajer P, Brotman RM, Bai G, Sakamoto J, Schutte UM, Zhong X, Koenig SS, Fu L, Ma ZS, Zhou X, et al. Temporal dynamics of the human vaginal microbiota. Sci Transl Med. 2012;4(132):132ra152. doi: 10.1126/scitranslmed.3003605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yamamoto T, Zhou X, Williams CJ, Hochwalt A, Forney LJ. Bacterial populations in the vaginas of healthy adolescent women. J Pediatr Adolesc Gynecol. 2009;22(1):11–18. doi: 10.1016/j.jpag.2008.01.073. [DOI] [PubMed] [Google Scholar]
- 41.Pavlova SI, Kilic AO, Kilic SS, So JS, Nader-Macias ME, Simoes JA, Tao L. Genetic diversity of vaginal lactobacilli from women in different countries based on 16S rRNA gene sequences. J Appl Microbiol. 2002;92(3):451–459. doi: 10.1046/j.1365-2672.2002.01547.x. [DOI] [PubMed] [Google Scholar]
- 42.Fettweis JM, Brooks JP, Serrano MG, Sheth NU, Girerd PH, Edwards DJ, Strauss JF, The vaginal microbiome C. Jefferson KK, Buck GA. Differences in vaginal microbiome in African American women versus women of European ancestry. Microbiology. 2014;160(Pt 10):2272–2282. doi: 10.1099/mic.0.081034-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lourembam SD, Sawian CE, Baruah S. Differential association of KIR gene loci to risk of malaria in ethnic groups of Assam, Northeast India. Infect Genet Evol. 2011;11(8):1921–1928. doi: 10.1016/j.meegid.2011.08.017. [DOI] [PubMed] [Google Scholar]
- 44.Reddy BM, Langstieh BT, Kumar V, Nagaraja T, Reddy AN, Meka A, Reddy AG, Thangaraj K, Singh L. Austro-Asiatic tribes of Northeast India provide hitherto missing genetic link between south and Southeast Asia. PLoS One. 2007;2(11):e1141. doi: 10.1371/journal.pone.0001141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Jakobsson T, Forsum U. Lactobacillus iners: a marker of changes in the vaginal flora? J Clin Microbiol. 2007;45(9):3145. doi: 10.1128/JCM.00558-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Macklaim JM, Gloor GB, Anukam KC, Cribby S, Reid G. At the crossroads of vaginal health and disease, the genome sequence of Lactobacillus iners AB-1. Proc Natl Acad Sci U S A. 2011;108(Suppl 1):4688–4695. doi: 10.1073/pnas.1000086107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Vaneechoutte M. Lactobacillus iners, the unusual suspect. Res Microbiol. 2017;168(9–10):826–836. doi: 10.1016/j.resmic.2017.09.003. [DOI] [PubMed] [Google Scholar]
- 48.Madhivanan P, Raphael E, Rumphs A, Krupp K, Ravi K, Srinivas V, Arun A, Reingold AL, Klausner JD, Riley LW. Characterization of culturable vaginal Lactobacillus species among women with and without bacterial vaginosis from the United States and India: a cross-sectional study. J Med Microbiol. 2014;63(Pt 7):931–935. doi: 10.1099/jmm.0.073080-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Mendes-Soares H, Suzuki H, Hickey RJ, Forney LJ. Comparative functional genomics of Lactobacillus spp. reveals possible mechanisms for specialization of vaginal lactobacilli to their environment. J Bacteriol. 2014;196(7):1458–1470. doi: 10.1128/JB.01439-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.O'Sullivan O, O'Callaghan J, Sangrador-Vegas A, McAuliffe O, Slattery L, Kaleta P, Callanan M, Fitzgerald GF, Ross RP, Beresford T. Comparative genomics of lactic acid bacteria reveals a niche-specific gene set. BMC Microbiol. 2009;9:50. doi: 10.1186/1471-2180-9-50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Watanabe M, Kinoshita H, Nitta M, Yukishita R, Kawai Y, Kimura K, Taketomo N, Yamazaki Y, Tateno Y, Miura K, et al. Identification of a new adhesin-like protein from Lactobacillus mucosae ME-340 with specific affinity to the human blood group a and B antigens. J Appl Microbiol. 2010;109(3):927–935. doi: 10.1111/j.1365-2672.2010.04719.x. [DOI] [PubMed] [Google Scholar]
- 52.Witkin SS, Linhares IM, Giraldo P. Bacterial flora of the female genital tract: function and immune regulation. Best Pract Res Clin Obstet Gynaecol. 2007;21(3):347–354. doi: 10.1016/j.bpobgyn.2006.12.004. [DOI] [PubMed] [Google Scholar]
- 53.Zarate G, Nader-Macias ME. Influence of probiotic vaginal lactobacilli on in vitro adhesion of urogenital pathogens to vaginal epithelial cells. Lett Appl Microbiol. 2006;43(2):174–180. doi: 10.1111/j.1472-765X.2006.01934.x. [DOI] [PubMed] [Google Scholar]
- 54.Zarate G, Santos V, Nader-Macias ME. Protective effect of vaginal Lactobacillus paracasei CRL 1289 against urogenital infection produced by Staphylococcus aureus in a mouse animal model. Infect Dis Obstet Gynecol. 2009;2009:48358. doi: 10.1155/2007/48358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Boris S, Suarez JE, Vazquez F, Barbes C. Adherence of human vaginal lactobacilli to vaginal epithelial cells and interaction with uropathogens. Infect Immun. 1998;66(5):1985–1989. doi: 10.1128/iai.66.5.1985-1989.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Corredoira J, Alonso MP, Coira A, Varela J. Association between Streptococcus infantarius (formerly S. bovis II/1) bacteremia and noncolonic cancer. J Clin Microbiol. 2008;46(4):1570. doi: 10.1128/JCM.00129-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980;16(2):111–120. doi: 10.1007/BF01731581. [DOI] [PubMed] [Google Scholar]
- 58.Rajagopal L. Understanding the regulation of group B streptococcal virulence factors. Future Microbiol. 2009;4(2):201–221. doi: 10.2217/17460913.4.2.201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Patel R, Keating MR, Cockerill FR, 3rd, Steckelberg JM. Bacteremia due to Enterococcus avium. Clin Infect Dis. 1993;17(6):1006–1011. doi: 10.1093/clinids/17.6.1006. [DOI] [PubMed] [Google Scholar]
- 60.Bourafa N, Loucif L, Boutefnouchet N, Rolain JM. Enterococcus hirae, an unusual pathogen in humans causing urinary tract infection in a patient with benign prostatic hyperplasia: first case report in Algeria. New Microbes New Infect. 2015;8:7–9. doi: 10.1016/j.nmni.2015.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Tian RM, Cai L, Zhang WP, Cao HL, Qian PY. Rare events of Intragenus and Intraspecies horizontal transfer of the 16S rRNA gene. Genome Biol Evol. 2015;7(8):2310–2320. doi: 10.1093/gbe/evv143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Baele M, Vancanneyt M, Devriese LA, Lefebvre K, Swings J, Haesebrouck F. Lactobacillus ingluviei sp. nov., isolated from the intestinal tract of pigeons. Int J Syst Evol Microbiol. 2003;53(Pt 1):133–136. doi: 10.1099/ijs.0.02206-0. [DOI] [PubMed] [Google Scholar]
- 63.Liu QH, Yang FY, Zhang JG, Shao T. Characteristics of Lactobacillus parafarraginis ZH1 and its role in improving the aerobic stability of silages. J Appl Microbiol. 2014;117(2):405–416. doi: 10.1111/jam.12530. [DOI] [PubMed] [Google Scholar]
- 64.Aagaard K, Riehle K, Ma J, Segata N, Mistretta TA, Coarfa C, Raza S, Rosenbaum S, Van den Veyver I, Milosavljevic A, et al. A metagenomic approach to characterization of the vaginal microbiome signature in pregnancy. PLoS One. 2012;7(6):e36466. doi: 10.1371/journal.pone.0036466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Romero R, Hassan SS, Gajer P, Tarca AL, Fadrosh DW, Nikita L, Galuppi M, Lamont RF, Chaemsaithong P, Miranda J, et al. The composition and stability of the vaginal microbiota of normal pregnant women is different from that of non-pregnant women. Microbiome. 2014;2(1):4. doi: 10.1186/2049-2618-2-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Mitra A, MacIntyre DA, Marchesi JR, Lee YS, Bennett PR, Kyrgiou M. The vaginal microbiota, human papillomavirus infection and cervical intraepithelial neoplasia: what do we know and where are we going next? Microbiome. 2016;4(1):58. doi: 10.1186/s40168-016-0203-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Yang X, Da M, Zhang W, Qi Q, Zhang C, Han S. Role of Lactobacillus in cervical cancer. Cancer Manag Res. 2018;10:1219–1229. doi: 10.2147/CMAR.S165228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Suen LJ, Huang HM, Lee HH. A comparison of convenience sampling and purposive sampling. Hu Li Za Zhi. 2014;61(3):105–111. doi: 10.6224/JN.61.3.105. [DOI] [PubMed] [Google Scholar]
- 69.Purkayastha SD, Bhattacharya MK, Prasad HK, Upadhyaya H, Lala SD, Pal K, Sharma GD. Antibacterial activity of Weissella confusa isolated from vaginal swab of Indian women. Int J Adv Chem Eng Biol Sci. 2017;4(1):98–102. [Google Scholar]
- 70.Turner S, Pryer KM, Miao VP, Palmer JD. Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J Eukaryot Microbiol. 1999;46(4):327–338. doi: 10.1111/j.1550-7408.1999.tb04612.x. [DOI] [PubMed] [Google Scholar]
- 71.Thompson JD, Gibson TJ, Higgins DG. Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinformatics. 2002;Chapter 2:Unit 2 3. doi: 10.1002/0471250953.bi0203s00. [DOI] [PubMed] [Google Scholar]
- 72.Dias R, Xavier MG, Rossi FD, Neves MV, Lange TA, Giongo A, De Rose CA, Triplett EW. MPI-blastn and NCBI-TaxCollector: improving metagenomic analysis with high performance classification and wide taxonomic attachment. J Bioinforma Comput Biol. 2014;12(3):1450013. doi: 10.1142/S0219720014500139. [DOI] [PubMed] [Google Scholar]
- 73.Tamura K, Nei M, Kumar S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci U S A. 2004;101(30):11030–11035. doi: 10.1073/pnas.0404206101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35(6):1547–1549. doi: 10.1093/molbev/msy096. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Table S1. Sample collection information. (DOCX 28 kb)
Table S2. Cell morphology, Gram staining of the bacterial isolates and their corresponding GenBank accession numbers of 16S rRNA gene sequence. (DOCX 36 kb)
Table S3. Homology search of the sequences in database. (DOCX 38 kb)
Table S4. Developed and database sequences. (XLSX 13 kb)
Table S5. Model test result. (XLS 49 kb)
Figure S1. Maximum Likelihood based clustering of developed and database sequences. The sequences of same species clustered together and distinct with respect to sequences of other species. (PDF 50 kb)
Figure S2. 16S rRNA gene based identification of the microbes is based on dual approach of homology search and ML clustering. To confirm identification we considered consensus match of our query sequence with a particular microbe on both the database and cohesive clustering of developed and database sequence of the particular microbe (here shown as Species A and Species B) and distinct with respect to other microbes (all species A members clustered distinct with all species B members). (JPG 250 kb)
Data Availability Statement
All the sequences used in this study have been submitted to NCBI and the details can be checked using their respective accession numbers (please see Additional file 2: Table S2) in NCBI (https://www.ncbi.nlm.nih.gov/).