The absence of eIFiso4G decreases the levels of some proteins, whereas overexpression of eIFiso4G1 restores synthesis of the decreased proteins, normal chloroplast function/appearance, and plant fitness.
Abstract
The plant-specific translation initiation complex eIFiso4F is encoded by three genes in Arabidopsis (Arabidopsis thaliana)—genes encoding the cap binding protein eIFiso4E (eifiso4e) and two isoforms of the large subunit scaffolding protein eIFiso4G (i4g1 and i4g2). To quantitate phenotypic changes, a phenomics platform was used to grow wild-type and mutant plants (i4g1, i4g2, i4e, i4g1 x i4g2, and i4g1 x i4g2 x i4e [i4f]) under various light conditions. Mutants lacking both eIFiso4G isoforms showed the most obvious phenotypic differences from the wild type. Two-dimensional differential gel electrophoresis and mass spectrometry were used to identify changes in protein levels in plants lacking eIFiso4G. Four of the proteins identified as measurably decreased and validated by immunoblot analysis were two light harvesting complex binding proteins 1 and 3, Rubisco activase, and carbonic anhydrase. The observed decreased levels for these proteins were not the direct result of decreased transcription or protein instability. Chlorophyll fluorescence induction experiments indicated altered quinone reduction kinetics for the double and triple mutant plants with significant differences observed for absorbance, trapping, and electron transport. Transmission electron microscopy analysis of the chloroplasts in mutant plants showed impaired grana stacking and increased accumulation of starch granules consistent with some chloroplast proteins being decreased. Rescue of the i4g1 x i4g2 plant growth phenotype and increased expression of the validated proteins to wild-type levels was obtained by overexpression of eIFiso4G1. These data suggest a direct and specialized role for eIFiso4G in the synthesis of a subset of plant proteins.
The synthesis of proteins is a fundamental process and is mechanistically similar across all organisms. Higher eukaryotes (plants, animals, and fungi) have evolved a much more complex scheme for the initiation of translation compared to prokaryotes, requiring more than 10 known initiation factors compared to three in prokaryotes (Hinnebusch, 2014). The first steps in initiation involve a group of eukaryotic initiation factors (eIF) that recognize and bind to the m7GpppX cap group of the mRNA (Hinnebusch, 2014). These factors, eIF4F, eIF4A, and eIF4B, are known collectively as the “eIF4 group.” The factor eIF4F is composed of two subunits, eIF4E and eIF4G (Merrick, 2015). eIF4E is ∼24 kD in eukaryotes and binds the m7GpppX cap group. eIF4G is a large scaffolding protein, ∼180 kD, and has consensus binding sites for eIF4E and contains 1–3 HEAT (Huntington, eIF3, protein phosphatase A, and TOR) domains (one in yeast, two in plants, and three in mammals) that function to bind other initiation factors (eIF4A, eIF4B, eIF3, and eIF5). These factors all form interactions with the mRNA and the 40S ribosome to assemble the preinitiation complex (Valásek, 2012; Hinnebusch, 2014, 2017; Dever et al., 2016).
Plant translation has a similar number of initiation factors to other eukaryotes, but has a second form of eIF4F, namely eIFiso4F, that is specific to land plants (Browning and Bailey-Serres, 2015). eIFiso4F contains two subunits—eIFiso4E and eIFiso4G—with functions similar to the subunits of eIF4F (van Heerden and Browning, 1994). eIFiso4G is considerably smaller than eIF4G (∼80 kD versus ∼180 kD); however, eIFiso4G retains the conserved eIF4E binding site and the two C-terminal 1–3 HEAT domains, but lacks the N-terminal region present in eIF4G (Patrick and Browning, 2012). Evolutionary conservation of the distinct eIFiso4G and eIF4G isoforms is apparent throughout the plant lineage with eIFiso4E appearing more recently and only noted in flowering plant species. These observations imply each isoform has some important nonoverlapping function(s) that have, to date, been difficult to characterize but are fundamentally necessary throughout land plants and green algae (Patrick and Browning, 2012). eIFiso4G binds eIF4A in a similar manner to eIF4G but has overlapping binding regions for eIF4B and poly(A) binding protein, suggesting they may bind competitively (Gallie, 2014, 2018). Arabidopsis (Arabidopsis thaliana) has two genes for eIFiso4G—eIFiso4g1 (At5g57870) and eIFiso4g2 (At2g24050; Lellis et al., 2010). Both mRNAs are expressed, although eIFiso4g1 is expressed at a much higher level than eIFiso4g2 (Lellis et al., 2010). Recombinant expression and in vitro activity of eIFiso4F1 or eIFiso4F2 suggest that both are able to support initiation of translation of mRNAs (Mayberry et al., 2009). Point mutations or deletions in either eIFiso4E or eIF4E, as well as eIF4G and eIFiso4G to a lesser extent, confer resistance to a variety of plant viruses (Yoshii et al., 2004; Charron et al., 2008; German-Retana et al., 2008; Jenner et al., 2010; Wang and Krishnaswamy, 2012; Contreras-Paredes et al., 2013; Estevan et al., 2014; Konečná et al., 2014; Moury et al., 2014; Nicaise, 2015; Sanfaçon, 2015), suggesting a role for cap-binding protein complexes and/or translation in the virus/host interaction. Deletion of the Arabidopsis genes for eIFiso4E or either of the two genes for eIFiso4G had minimal impact on plant growth or viability (Duprat et al., 2002; Lellis et al., 2002, 2010); however, deletion of both of the genes for eIFiso4G displays a substantial phenotype that includes poor growth, lower chlorophyll levels, and lower fertility (Lellis et al., 2010; Chen et al., 2014). Deletion of both eIFiso4G genes was further observed to decrease levels of light harvesting complex II (LHCII) increase photosensitivity, and increase levels of nonphotochemical quenching; however, there was no detected change in photorespiration or dark respiration (Chen et al., 2014). It was observed that the loss of both eIFiso4G genes resulted in an increase in the level of violaxanthin de-epoxidase expression and increased activity of the xanthophyll cycle (Chen et al., 2014). A point mutation of eIFiso4G1 was also found to affect seed lipid synthesis due to a decrease in the transcript and expression levels of the beta subunit of the plastid heteromeric acetyl CoA carboxylase, but it also elevated the levels of cytoplasmic homomeric ACCase, suggesting an interplay between cytosol and plastid components involved in lipid biosynthesis (Li et al., 2017). Distinct interactions of isoforms of poly A binding protein with eIF4G or eIFiso4G1/2 have also been described in Gallie (2018). Phosphorylation of eIFiso4G1 by SnRK1 during hypoxia and subsequent changes in the translatome suggest a key role in the response to environmental changes (Cho et al., 2019). Recently, eIFiso4G1 and eIFiso4G2 have been linked to abscisic acid signaling and regulation (Bi et al., 2019), suggesting an important role for these proteins in plants.
In this report we show that plants lacking both subunits of eIFiso4G show decreased levels of specific plant proteins. The decreased protein levels do not correlate with corresponding decreased mRNA levels or protein stability, indicating posttranscriptional regulation of specific mRNAs in the absence of eIFiso4G subunits. Interestingly, several of the decreased proteins are protein components associated with photosynthesis and/or chloroplast function. The observation that removal of eIFiso4G leads to decreased protein levels for proteins with plant-specific functionality that can be restored with overexpression of eIFiso4G1 suggests that eIFiso4G has an evolutionarily conserved role necessary for normal synthesis of a subset of plant proteins.
RESULTS
Phenotypic Response to Varying Light Intensity
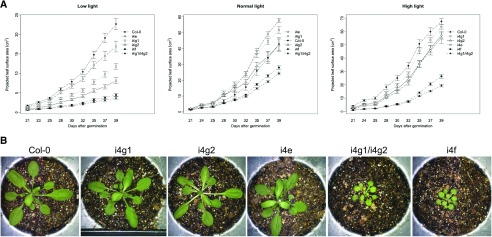
We had previously characterized large subunit (At5g57870 [i4g1]; At2g24050 [i4g2]) mutants of the plant specific initiation complex eIFiso4F (Lellis et al., 2010). The i4g1 x i4g2 double mutant plants have a severe phenotype, while individual mutants have subtle phenotypes in comparison. The deletion of the entire eIFiso4F complex (i4f, i4e x i4g1 x i4g2) was generated by crossing the large subunit double mutant (i4g1 x i4g2) to the corresponding cap binding mutant (i4e; Duprat et al., 2002). Initial observations of the triple mutant i4f plant showed a phenotype similar to the i4g1 x i4g2 double mutant, indicating that the loss of eIFiso4E did not appear to contribute any further detriment to plant fitness. To obtain a quantitative analysis of the various eIFiso4F mutant phenotypes, a high-throughput phenotyping platform (Scanalyzer HTS; LemnaTec), was used to monitor the growth of plants grown under low (100–120 μE/m2/s), normal (200–220 μE/m2/s), and high light (320–350 μE/m2/s) by measuring several parameters. As shown in Figure 1A, the i4f triple mutant plants were no more impaired in projected leaf area than the i4g1 x i4g2 double mutant, suggesting that the severe phenotype of these mutants is a result of the loss of the eIFiso4G1 and eIFiso4G2 subunits. The single mutant parents grew similarly to the wild-type controls under normal and high light; however, under low light more growth differentiation of the parent lines was observed. These results and the comparison of plant physical phenotype (Fig. 1B) confirm that the loss of eIFiso4E in the i4f mutant does not appear to affect overall fitness beyond that observed for the loss of eIFiso4G1 and eIFiso4G2 subunits. In planta, chlorophyll fluorescence was measured as an estimation of overall photosynthetic activity and was lowest in mutants lacking eIFiso4G1 and eIFiso4G2 (Supplemental Fig. S1). In other parameters measured, caliper length (rosette diameter) and convex hull area (minimal polygon that encloses the entire rosette area of the plant), the i4g1 x i4g2 double and i4f triple mutant plants behaved similarly to each other, and the single mutant parents resembled wild-type plants (Supplemental Fig. S2, A and B). Water content for all plants was lowest under low light conditions, but was largely consistent with no significant differences observed between the mutant or wild-type plants (Supplemental Fig. S2C).
Figure 1.
Plant high throughput phenotyping of single, double, and triple mutants. A, Projected leaf area of Arabidopsis lines grown under low, normal, or high white light conditions. Note that the scale differs for each light level. Statistical analysis P-value adjustment used the Tukey test for n = 15 and had a 0.05 significance level. Error bars = (se). B, Representative images of the size difference of Arabidopsis lines grown in soil (normal light, 16 h light) and photographed at 30-d postgermination.
2D-Differential Gel Electrophoresis Analysis and Confirmation of Protein Changes by Western Blotting
Previously, it was shown that there were no obvious global changes in translation in the i4g1 x i4g2 double mutant by incorporation of [35s]-Met using 1D SDS-PAGE (Lellis et al., 2010). Additional [35s]-Met labeling and separation by 2D gels confirmed that there were no gross changes in overall translation (see Supplemental Fig. S3). A more sensitive method using 2D differential gel electrophoresis (2D-DIGE; Applied Biomics) was used to identify more subtle changes in protein levels. Protein extracts were made from 14-d–old seedlings (i4g1 x i4g2 double mutant plants, i4f triple mutant plants, and the single mutant parent lines) grown under normal conditions and compared to wild-type Col-0 extracts (Fig. 2). There were 94 unique proteins that showed measurable changes in expression and were identified by mass spectrometry (see Supplemental Table S1). Among the top 10 proteins with decreased protein expression in the i4g1 x i4g2 double and i4f triple mutant plants, but not in the single mutant plants, were LHC binding protein 3 (Lhcb3; At5g54270), Lhcb1 (also known as chlorophyll A/B binding protein 1; At1g29930), carbonic anhydrase (CA1; At3g01500), and Rubisco activase (RCA; At2g39730). Three of these proteins (Lhcb3, CA1, and RCA) had multiple isoelectric forms on the 2D-DIGE that were all decreased in both the double and triple mutants. These multiple forms indicate that these proteins have various forms of posttranslational modifications, but protein levels were similarly decreased in the isoforms by the absence of eIFiso4G. Other proteins were observed to change in the single mutants, but these were not investigated further in this study as they were not changed in the double or triple mutant to the same extent.
Figure 2.
2D-DIGE of proteins extracted from double or triple mutants. Protein extracts were cy2 (Col-0) or cy3 (i4g1 x i4g2 or i4f) dye-labeled and run on 2D-PAGE (Applied Biomics). A, Wild type (Col-0) and mutant (i4g1 x i4g2; i4f) 2D gels are shown. B, Superimposed images comparing wild-type and mutant protein extracts as indicated. Green indicates the protein is decreased relative to Col-0, red indicates the protein is increased relative to Col-0, and yellow indicates that the protein level remained the same. Proteins that were measurably increased or decreased were identified by mass spectrometry. See Supplemental Table S1 for all mutants.
Antibodies to Lhcb3, Lhcb1, CA1, and RCA were used to validate the observed protein level changes in the 2D-DIGE in single, double, and triple mutant plant extracts by western blotting (Fig. 3A; Supplemental Fig. S4). Decreased protein levels were observed for all four targets in i4g1 x i4g2 double mutant and i4f triple mutant plants, but not in single mutant plants, suggesting that the decreased levels of these specific proteins are due to the lack of a functional eIFiso4G subunit. Lhcb3 and Lhcb1 showed the greatest decrease in the i4g1 x i4g2 double mutant and i4f triple mutant plants. CA1 and RCA showed more modest decreases in the double and triple mutant plants in agreement with 2D-DIGE observations. The actin control did not show any decrease in the double or triple mutant plants. Western blots with antibodies for eIFiso4G1 and eIFiso4E verified that the appropriate subunits were missing in the respective parents and double and triple mutants. Given the much lower level of eIFiso4G2 mRNA relative to eIFiso4G1 mRNA (Lellis et al., 2010) and a 5-fold–less cross-reaction to the eIFiso4G1 antibody (see Supplemental Fig. S5), it is likely that eIFiso4G2 is below the detection limit in the eIFiso4g1 mutant line. Interestingly, the i4g1 x i4g2 double mutant also shows a decrease in eIFiso4E. Analysis by quantitative western blot indicated that the amounts of eIFiso4E and eIFiso4G1 in wild-type plant extracts were roughly stoichiometric (see Supplemental Fig. S6). These results suggested that there may be coordinated regulation to maintain the stoichiometry for the eIFiso4F complex.
Figure 3.
Confirmation by western blotting of protein targets identified as decreased by 2D-DIGE in double or triple mutants. Total plant extracts were probed with antibodies to protein targets identified by 2D-DIGE in wild-type and mutant plants. A, Proteins that were the most decreased evidenced by 2D-DIGE: Lhcb3, Lhcb1, RCA, and CA1; i4G and i4E and actin are included as controls. B, Additional proteins identified as decreased in the 2D-DIGE: PsbP, VIPP1, PsbQ, and PsbO. See Supplemental Figure S4A for an example of the Stain-Free gel for protein loading comparison. MW, molecular weight.
A number of additional proteins from the 2D-DIGE data with more modest decreases (Vesicle-Inducing Protein In Plastid1 [VIPP1], PSII Subunit O [PsbO], PSII Subunit P [PsbP], and PSII Subunit Q [PsbQ]) were also analyzed by western blotting. As shown in Figure 3B, VIPP1, PsbP, and PsbQ appeared slightly decreased in the double and triple mutants, while PsbO displayed little or no change. In total, results from western blotting confirmed 2D-DIGE observations for decreased protein levels of specific proteins. Among the proteins that were increased in protein levels, there were no obvious candidates to validate that displayed a consistent increase in both the i4g1 x i4g2 double and i4f triple mutant plants, but not in the single mutants. It should be noted that only the most abundant proteins are likely to be observed by this method and proteins with low expression would likely be below the threshold for quantification.
Photosynthetic Efficiency
We had previously shown (Lellis et al., 2010) decreased chlorophyll in the double eIFiso4G mutant and apparent decreases in photosynthetic proteins, a phenotype that suggested photosynthetic efficiency might be impaired. Fast fluorescence kinetics were used to examine the initial rise to maximum fluorescence levels. Analysis of chlorophyll a fluorescence indicated that the overall quantum yield (QY_max) was not affected despite decreased levels of PSII components Lhcb1 and Lhcb3 (Fig. 4A). Statistically significant (P < 0.01) increases in the energy absorption/reaction center (ABS/RC), electron trapping efficiency (ETo/RC), transport (TRo/RC), and electron transfer per trapped photon (ETo/TRo) were observed, as well as an increase in the dissipation of excitation energy (DIo/RC), which is associated with stress or photodamage. The OJIP transients for the single mutants were similar to wild type; however, the double and triple mutants exhibited altered induction kinetics (Fig. 4B). The O-J rise was similar in wild type and the mutants, indicating that the rate of reduction of quinone A and quinone B was similar in all strains. The double- and triple mutants, however, exhibited slower thermal phase transitions (J→I and I→P), which are associated with reduction of the plastoquinone pool and electron acceptors beyond the cytochrome b6/f complex (Guo and Tan, 2011; Stirbet and Govindjee, 2011). Overall, the data suggested that PSII was functional, as there was little change in the QY_max and OJ transient. However, the mutant plants do appear to be in a constitutive “high light” state (State 2), likely due to the decreased levels of Lhcb1 and/or Lhcb3 antenna proteins (Damkjaer et al., 2009; Pietrzykowska et al., 2014). This conclusion is consistent with the slower transients (JI and IP) that indicate slower reduction of the electron carriers plastocyanin, ferredoxin, and NADP+, which lie beyond the cytochrome b6/f complex (Strasser et al., 2004; Baker, 2008).
Figure 4.
Photosynthetic efficiency of single, double, and triple mutants. A, Measurements of PSII photosynthetic efficiency in 21-d–old wild-type and mutant plant lines as determined by chlorophyll a fluorescence induction: QY_max, ABS/RC, TRo/RC, ETo/RC, ETo/TRo, and DIo/RC. Error bars = true mean ± se (n = 15–16) and asterisks indicate measurements that passed a two-tailed t test with P value < 0.01. B, OJIP fluorescence transient observed for the same lines (average curves normalized to FO and FM). C, Western blot of CYP38. See Supplemental Figure S4B for Stain-Free gel for loading control. MW, molecular weight.
The OJIP transient measurements bear similarity to those reported for cyp38 mutant plants (Lepedus et al., 2009). Cyclophilin 38 (CYP38) is a protein involved in PSII turnover and repair, and cyp38 mutants show sensitivity to light stress (Fu et al., 2007). The level of CYP38 protein was analyzed by western blotting (Fig. 4C) and found to be decreased in the double or triple mutants, but not in the parents. The decreased levels of antenna proteins such as Lhcb1 and Lhcb3 and the repair protein CYP38 may all contribute to poor repair and recovery of PSII during the dark resulting in a similar OJIP curve and phenotypic similarities to cyp38 mutant plants (Lepedus et al., 2009) and contribute to the overall impaired growth phenotype of plants lacking eIFiso4G.
Chloroplast Defects in i4g1 x i4g2 Plants
The i4g1 x i4g2 double mutant plants displayed decreased chlorophyll, altered chlorophyll a fluorescence kinetics, and 2D-DIGE/western blot observations indicated decreased levels of important proteins involved in photosynthesis/chloroplast function. These results strongly suggested that chloroplast assembly/function may be impaired. Transmission electron microscopy (TEM) analysis was used to examine plants for phenotypic changes in chloroplasts. Sections were prepared from 14-d–old seedlings and examined for morphological differences. In the i4g1 x i4g2 double mutant plants, the grana were thinner with fewer stacks compared to wild-type plants (Fig. 5). The decreased levels of the antenna proteins Lhcb1 and Lhcb3 protein, as well as the decreased amount of CYP38, would be consistent with i4g1 x i4g2 double mutant plants having an increased level of time in State 2 (high light), which may result in decreased grana stacking similar to that observed for Lhcb1 or Lhcb3 mutant plants (Damkjaer et al., 2009; Pietrzykowska et al., 2014). The decreased level of VIPP1 may also contribute to the altered grana stacking as VIPP1 has essential roles in chloroplast membrane function/assembly and thylakoid formation (Zhang et al., 2016a, 2016b; Ohnishi et al., 2018). The TEM results indicated a higher prevalence of starch granules in i4g1 x i4g2 double mutant plants (see Supplemental Table S2), suggesting compromised sugar utilization/signaling in i4g1 x i4g2 double mutant chloroplasts.
Figure 5.
TEM of wild-type and i4g1 x i4g2 double mutant chloroplasts. A to F, Cross sections of Col-0 (A–C) and i4g1 x i4g2 double mutant plants (D–F). Scale bars = 1 μm (A and D), 0.5 μm (B and E), or 0.2 μm (C and F). Mutant chloroplasts show more diffuse grana with fewer stacks and a higher incidence of starch granules (see Supplemental Table S2).
mRNA Levels and Stability/Turnover of Proteins Affected in eIFiso4G Mutants
To examine whether decreased levels of specific proteins in plants lacking eIFiso4G can be explained by altered levels of transcription, mRNA transcript levels from total RNA were obtained using RNA sequencing (RNA-Seq; see Table 1). Seven of the eight proteins validated by western blotting showed minimal changes in the total RNA levels (−0.37 to −0.03); however, Lhcb1 showed a modest increase (0.54) in mRNA. These results suggest that for the proteins validated by western blotting, the decrease in protein level was not directly due to substantial changes in transcription for these mRNAs.
Table 1. Summary of mRNA and protein levels for validated genes.
See Supplemental Table S1 for values of single mutants. ND, not detected in 2D-DIGE set.
| AT Number | Gene Name | mRNAa | Proteinb (Double Mutant) | Proteinc (Triple Mutant) |
|---|---|---|---|---|
| AT5G54270 | Lhcb3 | −0.16 (0.54) | −5.0 | −4.8 |
| AT1G29930 | Lhcb1 | 0.54 (0.048) | −3.1 | −2.4 |
| AT3G01500 | CA1 | −0.19 (0.41) | −2.8 | −2.2 |
| AT2G39730 | RCA | −0.20 (0.49) | −2.0 | −1.9 |
| AT1G06680 | PsbP1 | −0.29 (0.21) | −1.9 | −1.9 |
| AT1G65260 | VIPP1 | −0.29 (0.064) | −1.7 | −1.7 |
| AT4G05180 | PsbQ2 | −0.37 (0.059) | −1.7 | −1.8 |
| AT5G66570 | PsbO1 | −0.03 (0.93) | −1.6 | −1.7 |
| AT5G57870 | eIFiso4G1 | −3.0 (**)c | ND | ND |
| AT2G24050 | eIFiso4G2 | −8.3 (**)d | ND | ND |
| AT5G35620 | eIFiso4E | 0.38 (0.073) | ND | ND |
| AT3G01480 | CYP38 | −0.43 (0.011) | ND | ND |
Log2-fold change in RNA-SEQ of i4g1 x i4g2 mutant compared to Col-0 (wild type).
Log2-fold change from 2D-DIGE compared to Col-0 (wild type).
False discovery rate for eIFiso4G1 (**) was 7.4e-33.
False discovery rate for eIFiso4G2 (**) was 4.7e-82.
To determine whether the decreased protein levels observed in i4g1 x i4g2 double and i4f triple mutant plants were due to an issue in protein stability, 10-d–old seedlings were treated with cycloheximide (CHX) to inhibit new translation and the protein stability of several proteins was measured by western blotting (Gilkerson et al., 2015, 2016). The proteins all appeared similarly stable in the presence or absence of CHX, suggesting that the observed decrease in levels of the antibody validated proteins in the double and triple mutants was not due to significant changes in protein turnover (Fig. 6). HA-tagged indole-3-acetic acid1 (IAA1) serves as a positive control that is subject to degradation under these conditions (Gilkerson et al., 2016).
Figure 6.
Protein stability analysis of proteins in the i4g1 x i4g2 double mutant. Protein extracts were prepared from tissue treated with CHX to inhibit new translation. Extracts of Col-0 and the i4g1 x i4g2 double mutant were compared with and without CHX treatment. Antibodies were used to detect Lhcb3, Lhcb1, RCA, CA1, i4G and i4E, and actin. HA-tagged IAA1 is used as a control for stability. See Supplemental Figure S4C for Stain-Free gel for loading control. MW, molecular weight.
Rescue of the i4g1 x i4g2 Double Mutant Phenotype
Plants with the double mutation (i4g1 x i4g2) were complemented with transgenic cDNA for eIFiso4G1 under the control of the CaMV 35S promoter. Three homozygous plant lines with a single insertion for eIFiso4G1 were obtained and shown to rescue the phenotype of the double mutant (see Fig. 7A). Western blot analysis showed eIFiso4G1 overexpression in the three lines obtained. In addition to rescuing the plant growth phenotype, overexpression also restored, to wild-type levels, the antibody-validated proteins from the 2-DIGE analysis. In addition, overexpression of eIFiso4G1 also restored normal levels of eIFiso4E, but had no effect on the levels of PsbO or actin expression compared to wild type (see Fig. 7B). eIFiso4G1 was selected for making the overexpression line as the rabbit antibodies are more specific for eIFiso4G1 (see Supplemental Fig. S5); however, similar results would be expected for eIFiso4G2 because the severe phenotype is only observed in the double mutant. These results suggest there is a subset of proteins whose normal level of synthesis occurs in the presence of eIFiso4G.
Figure 7.
Effect of CaMV 35S overexpression of eIFiso4G1 in i4g1 xi 4g2 plants. A, 30-d–old plants grown in soil: Col-0 wild type, i4g1 xi 4g2 mutant, A1, E1, and F2 independent lines overexpressing eIFiso4G1 (16-h light). B, Western blot showing effect of overexpression of eIFiso4G1 with antibodies to Lhcb3, Lhcb1, RCA, CA1, i4E, i4G1, cyclophilin 38 (CYP38), and actin. See Supplemental Figure S4D for Stain-Free gel for loading control. MW, molecular weight.
DISCUSSION
The absence of the eIFiso4G1 and eIFiso4G2 subunits of the plant-specific translation initiation complex eIFiso4F has a profound effect on plant growth and fitness (Lellis et al., 2010). Here we have further characterized the effects of the removal of these subunits and examined the phenotypic and molecular changes in higher detail. Plants lacking eIFiso4G1 and eIFiso4G2, whether in the double or triple mutant, consistently showed similarly decreased growth under all light conditions. Removal of eIFiso4E does not appear to provide any additional impairment over the absence of eIFiso4G1 and eIFiso4G2, with both the double and triple mutants showing similar phenotypes under all growth conditions examined. This suggests some special functionality for the plant-specific scaffolding isoform eIFiso4G, as well as functional redundancy for the cap-binding subunits eIFiso4E and eIF4E (Callot and Gallois, 2014; Patrick et al., 2014). This apparent redundancy of the cap-binding protein is consistent with the observation that mixed complexes of eIF4G and eIFiso4E or eIFiso4G and eIF4E were able to form translationally active complexes in vitro and the activity of those complexes tracked with the large subunit and not the cap-binding subunit (Mayberry et al., 2011). It should be noted that the loss of eIFiso4G appeared to reduce the amount of eIFiso4E protein made, but conversely, loss of eIFiso4E did not result in any reduction of eIFiso4G protein, suggesting eIFiso4G has a role in the expression and/or regulation of eIFiso4E.
To examine changes in protein expression levels, we used 2D-DIGE to compare wild-type and mutant plant lines lacking one or more of the eIFiso4F subunits. A subset of proteins with decreased expression levels in the double and triple mutant plants was observed in the 2D-DIGE dataset. Eight of these proteins (Lhcb1, Lhcb3, RCA, CA1, VIPP1, PsbO, PsbP, and PsbQ) were selected for validation by western blot analysis, and protein levels were confirmed to be decreased to varying degrees in the double and triple mutant plants. Further analysis of the mRNA levels and protein stability suggests the decreased level of proteins in the double and triple mutants was not due to significant changes in transcription or protein stability, implying some specificity of function provided by eIFiso4G.
The apparent defects in synthesis for some of the components of the photosynthetic apparatus and the mutant phenotype suggested that photosynthesis efficiency may be impaired. Analysis by chlorophyll a fluorescence indicated that the overall QY_max was not impaired despite decreased levels of PSII components Lhcb1 and Lhcb3, but changes in the ETo and transport were noted, as well as an increase in the dissipation of excitation energy, which is associated with stress or photodamage. This may explain the similar OJIP pattern of the eIFiso4G mutants, with the decreased levels of antenna proteins Lhcb1 and Lhcb3, to cyp38 mutants, which show a sensitivity to light stress due to the inability to turnover PSII antenna components (Fu et al., 2007). The decrease in the amount of Lhcb1 and Lhcb3, as well as CYP38, is also consistent with the report that LHC II was reduced in plants lacking eIFiso4G (Chen et al., 2014). Collectively, the data point to defects in photosynthesis and/or chloroplasts as the primary cause for the phenotype in the double and triple mutant plants. In addition, all the proteins that were validated by western blotting had one or more gene ontology terms associated with photosynthesis or chloroplast function. TEM analysis was carried out to determine if there were physical defects in the chloroplasts of the eIFiso4G mutant plants. Indeed, the chloroplasts had substantial defects in the grana that were longer and thinner with fewer thylakoid stacks and a higher prevalence of starch granules compared to wild-type plants. It should be noted that glyceraldehyde-3-P dehydrogenase (AT3G26650) is slightly decreased in i4g1 x i4g2 double mutant plants (although not in the triple mutant) at both the protein (−1.1) and RNA levels (−0.37). Reduction in glyceraldehyde-3-P dehydrogenase activity results in an increase in starch, which may account, at least in part, for the observed increase in starch granules in the i4g1 x i4g2 double mutant plants (Yang et al., 2015).
Transgenic complementation of eIFiso4G1 in i4g1 x i4g2 mutant plants reverses the physical phenotype and restores wild-type levels of the protein levels decreased by the loss of eIFiso4G, including eIFiso4E. These data specifically show that eIFiso4G is necessary for normal translation of a subset of mRNAs including eIFiso4E mRNA. In vitro translation of mRNAs for two of the validated proteins, Lhcb3 or Lhcb1, surprisingly did not show a particular preference for eIFiso4F over eIF4F in the wheat germ system (see Supplemental Fig. S7). However, this result is consistent with other observations of mRNA translation in vitro that show most capped mRNAs prefer eIF4F over eIFiso4F in vitro (Mayberry et al., 2009). It appears that in vivo translation is subject to control mechanisms that the wheat germ in vitro system cannot recapitulate. The wheat germ system is derived from dormant tissue that is not actively photosynthesizing and there could be additional RNA binding proteins or other factors that may promote translation of specific mRNAs by eIFiso4F in actively photosynthesizing tissue in vivo. Further work will be needed to elucidate the molecular mechanism of this specificity, whether it is through the properties of the RNA itself, mediated through RNA-binding proteins, or some other unknown mechanism(s).
From analysis of the phenotype and changes in specific protein levels, photosynthetic efficiency, TEM analysis, and rescue of physical and molecular phenotype by overexpression, it appears that eIFiso4G is necessary for the normal translation of a subset of proteins associated with PSII (Lhcb1, Lhcb3, Cyp38, and PsbQ), chloroplast development (VIPP1), and/or carbon metabolism (RCA and CA1). This does not exclude that other proteins whose translation may be affected by the absence of eIFiso4F may fall below the threshold of detection or were otherwise not identified in the 2D-DIGE analysis, but could also contribute to the phenotype, e.g. CYP38. This study supports the role for eIFiso4G in the translation of a specific set of proteins contributing to plant-specific functions such as photosynthesis, chloroplast development, and carbon metabolism that are consistent with the plant-specific evolutionary origin for the eIFiso4F complex (Patrick and Browning, 2012).
MATERIALS AND METHODS
Plant Material and Growth Conditions
All experiments were performed with mutants derived from Arabidopsis (Arabidopsis thaliana) ecotype Columbia-0 (Col-0). Homozygous knockout lines were recovered for At5g57870 (i4g1, SALK line 009905) and At2g24050 (i4g2, SALK line 076633) through the Signal Salk T-DNA express program (http://signal.salk.edu/cgi-bin/tdnaexpress) as described in Lellis et al. (2010). A homozygous knockout line for At5g35620 (i4e) was obtained from the Sainsbury Laboratory’s SLAT library as described in Duprat et al. (2002). The i4g1 x i4g2 double mutant was described in Lellis et al. (2010). The i4f triple mutant was generated through crossing, self-fertilization, and PCR screening for a homozygous line lacking all three genes (see Supplemental Table S3). All plants were grown at 22°C in a 16-h photoperiod unless otherwise noted. Plants grown in soil were vernalized for 3–4 d at 4°C before germination. In experiments using 10- or 14-d–old seedlings, plants were grown aseptically. Seeds were sterilized with a 20% (v/v) bleach 1% (w/v) SDS solution for 15–20 min and rinsed 5× with sterile deionized water and plated on 0.5× Murashige and Skoog (MS) nutrient agar (0.8% w/v agar at pH 5.7) supplemented with 1% (w/v) Suc under sterile conditions and vernalized for 3–4 d at 4°C before germination. All tissue was harvested 6-h post-light, frozen in liquid nitrogen, powdered using a mortar and pestle, and stored at −80°C.
Plant High-Throughput Phenotyping
Arabidopsis seeds were surface-sterilized sequentially with 70% (v/v) ethanol, 50% (v/v) bleach, 0.05% (v/v) TWEEN 20 (Millipore-Sigma), and finally with sterile water before being planted on MS media supplemented with 3% (w/v) Suc. Seeds were vernalized for 3 d at 4°C before being transferred to a Plant Growth Chamber (Conviron) at 22 ± 1°C, 65 ± 5% relative humidity, and 16-h day:8-h night at 160 µE/m2/s. After true leaves had formed, uniformly sized, vigorous seedlings were transferred into Quick Pot 15 RW trays (HerkuPlast) containing Arabidopsis growing media (Lehle Seeds). Plants were grown under low (100–120 μE/m2/s), normal (200–220 μE/m2/s), or high (320–350 μE/m2/s) white light conditions. Images were captured every 2–3 d from seedling establishment to full vegetative growth using a Scanalyzer HTS system (LemnaTec) controlled by the LemnaControl software (LemnaTec). Specification of the visible, fluorescence, and near-infrared cameras used in this study as well as the detailed description of the image analysis pipeline were as described in Acosta-Gamboa et al. (2017). Statistical analysis P-value adjustment used the Tukey test for n = 15 and was 0.05 for the data shown in Figure 1A and Supplemental Figures S1 and S2.
2D-DIGE Sample Preparation
Samples for 2D-DIGE analysis were prepared from 14-d–old aseptically grown seedlings (wild type, i4g1, i4g2, i4e, i4g1 x i4g2, and i4f) under normal conditions described above in "Plant Material and Growth Conditions". Protein extracts were prepared as described in Deng et al. (2007). Briefly, ∼300 mg of powdered seedling tissue was ground in a microfuge tube with plastic pestle containing 700 µL of extraction buffer (100-mM Tris-Cl at pH 8.0, 2% [w/v] SDS, 1% [v/v] β-mercaptoethanol, 10-mm EDTA, 5-mm EGTA, 1× Pierce protease inhibitor mini tablet [Pierce Protein Biology/Thermo Fisher Scientific], and 1-mm phenylmethylsulfonyl fluoride)and extracted with an equal volume of phenol buffered with 50-mm Tris-Cl at pH 8. The organic layer was extracted with an equal volume of 50-mm Tris-Cl at pH 8.0 two times, precipitated with five volumes of 0.1-m ammonium acetate prepared in methanol and stored at −20°C for 3 h. The precipitates were collected by centrifugation and the pellets sent to Applied Biomics where proteins were dye-labeled, run with 2D gel electrophoresis, and differentially expressed proteins were identified through mass spectroscopy (see Supplemental Table S1).
Western Blotting
Samples to validate identified 2D-DIGE targets were prepared from 14-d–old seedlings grown as described in "Plant Material and Growth Conditions". Extracts were prepared as described in "2D-DIGE Sample Analysis" (Figs. 3 and 4) and the precipitate was resuspended in twice the volume (in microliters) of 2× Laemmli sample buffer (Bio-Rad) containing 1× protease inhibitor (Thermo Fisher Scientific) as the initial tissue weight (in milligrams). The protein concentration was determined using a BCA kit (Pierce Protein Biology/Thermo Fisher Scientific). Alternatively, extracts were prepared from frozen-powdered tissue using 2× Laemmli sample buffer (Figs. 6 and 7). Briefly, frozen weighed tissue (n miligrams) was crushed in twice the volume (2n μL) of 2× Laemmli sample buffer (Bio-Rad) containing protease inhibitor in a 1.5-mL microfuge tube with a plastic pestle, centrifuged briefly in a microfuge, and re-extracted with the pestle. The supernatant was carefully removed after a final centrifugation at maximum speed for 10 min at 4°C.
Equal amounts of protein (phenol extraction) as determined by the BCA assay (Pierce Protein Biology/Thermo Fisher Scientific) or equal volumes of extract from the 2× Laemmli sample buffer tissue extracts were resolved on 12% Stain Free polyacrylamide gels (Bio-Rad), visualized in a Chem-Doc (Bio-Rad) before transfer to a polyvinylidene difluoride (Millipore) membrane via semidry transfer (Bio-Rad) using transfer buffers described in Browning et al. (1990). Examples of Stain-Free gel images used for western blots are shown in Supplemental Figure S4. Membranes were blocked in 1× phosphate buffered saline (8-mm Na2HPO4, 150-mm NaCl, 2-mm KH2PO4, 3-mm KCl) containing 0.05% (v/v) TWEEN 20 (PBST) and 5% (w/v) nonfat dry milk. Membranes were probed with primary antibody (1/2,000 in PBST containing 5% [w/v] dry milk) for 1 h at room temperature followed by 3–4 washes with PBST and followed by secondary anti-rabbit HRP, anti-mouse HRP, or anti-goat HRP (Kendrick Labs) at 1:20,000 (in PBST containing 5% [w/v] dry milk) for 1 h and washed 3–4 times with PBST before visualization with chemiluminescent substrate (Genscript or Agrisera) using a Chem-Doc (Bio-Rad). Antibodies to Lhcb1 (goat) and RCA (goat) were obtained from Santa Cruz Biotechnology; actin (mouse) was from Millipore Sigma; Lhcb3 (rabbit) and VIPP1 (rabbit) antibodies were from Agrisera; and CA1 antibody (rabbit) and CYP38 antibody (rabbit) were gifts from James Moroney (LA State University, Baton Rouge, LA) and Björn Ingelsson (Linköping University, Linköping, Sweden), respectively. Antibodies to Arabidopsis eIFiso4G1 (Lellis et al., 2010), eIFiso4E (Duprat et al., 2002), and PsbO, PsbP, and PsbQ (Yi et al., 2006, 2008) were made as described in the indicated references. All extractions and western blots were performed at least in triplicate with similar results.
Total RNA Preparation and RNA-Seq
RNA samples were prepared from aseptically grown 14-d–old seedlings. The RNA was extracted using a RNeasy Plant Mini Kit (Qiagen) and treated with DNAse I (Roche) according to the manufacturer’s instructions. Poly(A) RNA libraries were constructed and sequenced by the Genome Sequencing and Analysis Facility at The University of Texas at Austin. All samples were prepared and sequenced in triplicate. Log-fold changes were estimated using negative binomial generalized linear models with log-link function fit to RNA-seq read counts as implemented in the “R” package “edgeR” (McCarthy et al., 2012). The fold changes shown in Table 1 compare the double mutant i4g1 x i4g2 to wild type (Col-0). The PCA analysis of the RNA-seq data points for the proteins identified in the 2D-DIGE is shown in Supplemental Figure S8A. A scatterplot of the mRNA levels and protein levels from 2D-DIGE is shown in Supplemental Figure S8B. Full analysis of the RNA-seq data set will be published elsewhere (R.M. Patrick and K.S. Browning, unpublished data).
Protein Stability Assay
Protein stability experiments were carried out as described in Gilkerson et al. (2015, 2016) and the HA-tagged IAA1 plant line was generously provided by Judy Callis (University of California-Davis). Approximately 250 mg of 10-d–old aseptically grown seedlings were placed into 3 mL of liquid 0.5× MS, 1% (w/v) Suc, and 200-μm CHX solution in a 35 × 10 mm petri dish. Controls were incubated in 0.5× MS and 1% (w/v) Suc without CHX. Plants were rotated on an orbital platform shaker at 50 rpm under light, at 22°C for 3 h. The tissue was rinsed with water, blotted dry with paper towels, frozen in liquid N2, and powdered with a mortar and pestle. The powdered sample was then brought up in two volumes (relative to initial weight in milligrams) of 2× Laemmli buffer (Bio-Rad) as described in "Western Blotting". Samples were run on 12% polyacrylamide Stain Free gels (Bio-Rad), imaged on a Chem-Doc (Bio-Rad), and western blotting was performed as described in "Western Blotting". Rabbit anti-HA was from ICL and used at 1:2,000 dilution as described in "Western Blotting". Assays were performed three times with similar results.
Chlorophyll a Fluorescence
Measurements and analysis of chlorophyll a fluorescence induction were made on 21-d–old, dark-adapted wild-type or mutant plants (n = 15–16) grown on soil as previously described in Roose et al. (2011). Data were collected and analyzed using a FluorCam 7 (Photon Systems Instruments) and its associated proprietary software.
TEM
Initial TEM shown in Figure 5 was carried out through Science Exchange (https://www.scienceexchange.com/) with Dr. Anza Darehshouri (University of Texas Southwestern Medical School). Samples of wild-type and i4g1 x i4g2 aseptically grown plants were cut into ∼2-mm pieces, fixed in 2.5% (v/v) glutaraldehyde, 4% (v/v) paraformaldehyde, and 2.5-mm CaCl2 in 100-mm sodium cacodylate buffer overnight at 4°C. After three rinses in 100-mm sodium cacodylate buffer, they were postfixed in 1% (w/v) osmium tetroxide and 1.5% (w/v) K3(Fe[CN6]) in 100-mm sodium cacodylate buffer for 1.5 h at room temperature. They were rinsed in double-distilled water and dehydrated in an ethanol series and propylene oxide, infiltrated with a graded series of Spurr resin, and embedded in the same resin. Blocks were polymerized overnight at 70°C. Ultrathin sections (60–70 nm) were obtained with a diamond knife (Diatome) on an Ultracut UCT Ultramicrotome (Leica Microsystems), and stained with uranyl acetate and lead citrate and viewed with a Tecnai G2 Spirit BioTwin transmission electron microscope (FEI) using a voltage of 120 kV. Additional analysis was carried out at the University of Texas at Austin Center for Biomedical Research Support Microscopy facility to further analyze the starch granule phenotype (Supplemental Table S2). The tissue was similarly fixed and ultrathin sections obtained using a model no. UC7 diamond knife (Leica) and visualized on a Tecnai Spirit BioTwin transmission electron microscope (FEI) using a voltage of at 80 kV.
CaMV 35S Overexpression of eIFiso4G1
The Arabidopsis eIFiso4G1 coding sequence was amplified by PCR from pET22b-Ati4g1 (Mayberry et al., 2009) and cloned into pG125 (a modified Gateway entry vector) via EcoRI-SpeI restriction sites (see Supplemental Table S3). The eIFiso4G1 coding sequence was then recombined into the pEarleyGate100 plant transformation vector (Earley et al., 2006) by LR recombination (Invitrogen). The resulting pEarleyGate100-Ati4g1 construct was transformed into Agrobacterium tumefaciens (strain GV3101) to be used for plant transformation. i4g1 x i4g2 double mutant plants were transformed by floral dip as described in Davis et al. (2009). Herbicide-resistant (BASTA; Bayer) plants were screened for overexpression of eIFiso4G1 by western blot analysis. The i4g1 x i4g2 background was confirmed by PCR using primers described in Lellis et al. (2010). Three independent overexpression, single-insertion, homozygous lines were obtained.
Accession Numbers
At5g57870 (i4g1), At2g24050 (i4g2), and At5g35620 (i4e) were used in this work. Other accession numbers are available in Supplemental Table S1.
Supplemental Data
The following supplemental materials are available.
Supplemental Figure S1. Chlorophyll fluorescence in leaves.
Supplemental Figure S2. Comparison of caliper, convex hull, and water content under low, normal, and high light.
Supplemental Figure S3. 2D gel images.
Supplemental Figure S4. Stain-free gel images for western blots.
Supplemental Figure S5. Rabbit anti-AteIFiso4G1 antibody cross-reaction with AteIFiso4G2.
Supplemental Figure S6. eIFiso4G and eIFiso4E exist in equimolar concentrations in wild-type Arabidopsis seedlings.
Supplemental Figure S7. In vitro translation assay for Lhcb3 and Lhcb1(chlorophyll A/B binding protein 1) mRNAs.
Supplemental Figure S8. PCA and scatter plot analysis of mRNA and protein data sets.
Supplemental Table S1. 2D-DIGE protein analysis and identification.
Supplemental Table S2. Average percent area of starch granules in chloroplasts.
Supplemental Table S3. Oligonucleotides used.
Acknowledgments
We thank Dr. Dwight Romanovicz (University of Texas at Austin Center for Biomedical Research Support Microscopy facility) for training and assistance with TEM. We also thank the Freshman Research Initiative BioBricks Stream led by Grace S. Choy and Karen S. Browning. for assistance with cloning the eIFiso4G1 cDNA used in the overexpression and Lhcb1 and Lhcb3 cDNAs used for mRNA transcription.
Footnotes
This work was supported by the National Science Foundation (grant no. MCB1052530 to K.S.B.), the Plant Imaging Consortium (NSF Award nos. IIA-1430427 and IIA-1430428 to A.L.), and the United States Department of Energy, Office of Basic Energy Sciences (grant DE-FG02-09ER20310 to T.M.B. and L.K.F.).
Articles can be viewed without a subscription.
References
- Acosta-Gamboa LM, Liu S, Langley E, Campbell Z, Castro-Guerrero N, Mendoza-Cozatl D, Lorence A (2017) Moderate to severe water limitation differentially affects the phenome and ionome of Arabidopsis. Funct Plant Biol 44: 94–106 [DOI] [PubMed] [Google Scholar]
- Baker NR. (2008) Chlorophyll fluorescence: A probe of photosynthesis in vivo. Annu Rev Plant Biol 59: 89–113 [DOI] [PubMed] [Google Scholar]
- Bi C, Ma Y, Jiang SC, Mei C, Wang XF, Zhang DP (2019) Arabidopsis translation initiation factors eIFiso4G1/2 link repression of mRNA cap-binding complex eIFiso4F assembly with RNA-binding protein SOAR1-mediated ABA signaling. New Phytol 223: 1388–1406 [DOI] [PubMed] [Google Scholar]
- Browning KS, Bailey-Serres J (2015) Mechanism of cytoplasmic mRNA translation. Arabidopsis Book 13: e0176 doi:10.1199/tab.0176 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Browning KS, Humphreys J, Hobbs W, Smith GB, Ravel JM (1990) Determination of the amounts of the protein synthesis initiation and elongation factors in wheat germ. J Biol Chem 265: 17967–17973 [PubMed] [Google Scholar]
- Callot C, Gallois JL (2014) Pyramiding resistances based on translation initiation factors in Arabidopsis is impaired by male gametophyte lethality. Plant Signal Behav 9: e27940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Charron C, Nicolaï M, Gallois JL, Robaglia C, Moury B, Palloix A, Caranta C (2008) Natural variation and functional analyses provide evidence for co-evolution between plant eIF4E and potyviral VPg. Plant J 54: 56–68 [DOI] [PubMed] [Google Scholar]
- Chen Z, Jolley B, Caldwell C, Gallie DR (2014) Eukaryotic translation initiation factor eIFiso4G is required to regulate violaxanthin de-epoxidase expression in Arabidopsis. J Biol Chem 289: 13926–13936 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cho HY, Lu MJ, Shih MC (2019) The SnRK1-eIFiso4G1 signaling relay regulates the translation of specific mRNAs in Arabidopsis under submergence. New Phytol 222: 366–381 [DOI] [PubMed] [Google Scholar]
- Contreras-Paredes CA, Silva-Rosales L, Daròs JA, Alejandri-Ramírez ND, Dinkova TD (2013) The absence of eukaryotic initiation factor eIF(iso)4E affects the systemic spread of a Tobacco etch virus isolate in Arabidopsis thaliana. Mol Plant Microbe Interact 26: 461–470 [DOI] [PubMed] [Google Scholar]
- Damkjaer JT, Kereïche S, Johnson MP, Kovacs L, Kiss AZ, Boekema EJ, Ruban AV, Horton P, Jansson S (2009) The photosystem II light-harvesting protein Lhcb3 affects the macrostructure of photosystem II and the rate of state transitions in Arabidopsis. Plant Cell 21: 3245–3256 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis AM, Hall A, Millar AJ, Darrah C, Davis SJ (2009) Protocol: Streamlined sub-protocols for floral-dip transformation and selection of transformants in Arabidopsis thaliana. Plant Methods 5: 3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng Z, Zhang X, Tang W, Oses-Prieto JA, Suzuki N, Gendron JM, Chen H, Guan S, Chalkley RJ, Peterman TK, et al. (2007) A proteomics study of brassinosteroid response in Arabidopsis. Mol Cell Proteomics 6: 2058–2071 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dever TE, Kinzy TG, Pavitt GD (2016) Mechanism and regulation of protein synthesis in Saccharomyces cerevisiae. Genetics 203: 65–107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duprat A, Caranta C, Revers F, Menand B, Browning KS, Robaglia C (2002) The Arabidopsis eukaryotic initiation factor (iso)4E is dispensable for plant growth but required for susceptibility to potyviruses. Plant J 32: 927–934 [DOI] [PubMed] [Google Scholar]
- Earley KW, Haag JR, Pontes O, Opper K, Juehne T, Song K, Pikaard CS (2006) Gateway-compatible vectors for plant functional genomics and proteomics. Plant J 45: 616–629 [DOI] [PubMed] [Google Scholar]
- Estevan J, Maréna A, Callot C, Lacombe S, Moretti A, Caranta C, Gallois JL (2014) Specific requirement for translation initiation factor 4E or its isoform drives plant host susceptibility to Tobacco etch virus. BMC Plant Biol 14: 67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fu A, He Z, Cho HS, Lima A, Buchanan BB, Luan S (2007) A chloroplast cyclophilin functions in the assembly and maintenance of photosystem II in Arabidopsis thaliana. Proc Natl Acad Sci USA 104: 15947–15952 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallie DR. (2014) The role of the poly(A) binding protein in the assembly of the Cap-binding complex during translation initiation in plants. Translation (Austin) 2: e959378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallie DR. (2018) Plant growth and fertility requires functional interactions between specific PABP and eIF4G gene family members. PLoS One 13: e0191474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- German-Retana S, Walter J, Doublet B, Roudet-Tavert G, Nicaise V, Lecampion C, Houvenaghel MC, Robaglia C, Michon T, Le Gall O (2008) Mutational analysis of plant cap-binding protein eIF4E reveals key amino acids involved in biochemical functions and potyvirus infection. J Virol 82: 7601–7612 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilkerson J, Kelley DR, Tam R, Estelle M, Callis J (2015) Lysine residues are not required for proteasome-mediated proteolysis of the auxin/indole acidic acid protein IAA1. Plant Physiol 168: 708–720 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilkerson J, Tam R, Zhang AH, Dreher K, Callis J (2016) Cycloheximide assays to measure protein degradation in vivo in plants. Bio Protoc 6: e1919 [Google Scholar]
- Guo Y, Tan J (2011) Modeling and simulation of the initial phases of chlorophyll fluorescence from Photosystem II. Biosystems 103: 152–157 [DOI] [PubMed] [Google Scholar]
- Hinnebusch AG. (2014) The scanning mechanism of eukaryotic translation initiation. Annu Rev Biochem 83: 779–812 [DOI] [PubMed] [Google Scholar]
- Hinnebusch AG. (2017) Structural insights into the mechanism of scanning and start codon recognition in eukaryotic translation initiation. Trends Biochem Sci 42: 589–611 [DOI] [PubMed] [Google Scholar]
- Jenner CE, Nellist CF, Barker GC, Walsh JA (2010) Turnip mosaic virus (TuMV) is able to use alleles of both eIF4E and eIF(iso)4E from multiple loci of the diploid Brassica rapa. Mol Plant Microbe Interact 23: 1498–1505 [DOI] [PubMed] [Google Scholar]
- Konečná E, Šafářová D, Navrátil M, Hanáček P, Coyne C, Flavell A, Vishnyakova M, Ambrose M, Redden R, Smýkal P (2014) Geographical gradient of the eIF4E alleles conferring resistance to potyviruses in pea (Pisum) germplasm. PLoS One 9: e90394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lellis AD, Kasschau KD, Whitham SA, Carrington JC (2002) Loss-of-susceptibility mutants of Arabidopsis thaliana reveal an essential role for eIF(iso)4E during potyvirus infection. Curr Biol 12: 1046–1051 [DOI] [PubMed] [Google Scholar]
- Lellis AD, Allen ML, Aertker AW, Tran JK, Hillis DM, Harbin CR, Caldwell C, Gallie DR, Browning KS (2010) Deletion of the eIFiso4G subunit of the Arabidopsis eIFiso4F translation initiation complex impairs health and viability. Plant Mol Biol 74: 249–263 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lepedus H, Tomasic A, Juric S, Katanic Z, Cesar V, Fulgosi H (2009) Photochemistry of PSII in CYP38 Arabidopsis thaliana deletion mutant. Food Technol Biotechnol 47: 275–280 [Google Scholar]
- Li Q, Shen W, Zheng Q, Tan Y, Gao J, Shen J, Wei Y, Kunst L, Zou J (2017) Effects of eIFiso4G1 mutation on seed oil biosynthesis. Plant J 90: 966–978 [DOI] [PubMed] [Google Scholar]
- Mayberry LK, Allen ML, Dennis MD, Browning KS (2009) Evidence for variation in the optimal translation initiation complex: Plant eIF4B, eIF4F, and eIF(iso)4F differentially promote translation of mRNAs. Plant Physiol 150: 1844–1854 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayberry LK, Allen ML, Nitka KR, Campbell L, Murphy PA, Browning KS (2011) Plant cap-binding complexes eukaryotic initiation factors eIF4F and eIFISO4F: Molecular specificity of subunit binding. J Biol Chem 286: 42566–42574 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCarthy DJ, Chen Y, Smyth GK (2012) Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res 40: 4288–4297 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merrick WC. (2015) eIF4F: A retrospective. J Biol Chem 290: 24091–24099 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moury B, Charron C, Janzac B, Simon V, Gallois JL, Palloix A, Caranta C (2014) Evolution of plant eukaryotic initiation factor 4E (eIF4E) and potyvirus genome-linked protein (VPg): A game of mirrors impacting resistance spectrum and durability. Infect Genet Evol 27: 472–480 [DOI] [PubMed] [Google Scholar]
- Nicaise V. (2015) Lost in translation: An antiviral plant defense mechanism revealed. Cell Host Microbe 17: 417–419 [DOI] [PubMed] [Google Scholar]
- Ohnishi N, Zhang L, Sakamoto W (2018) VIPP1 involved in chloroplast membrane integrity has GTPase activity in vitro. Plant Physiol 177: 328–338 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patrick RM, Browning KS (2012) The eIF4F and eIFiso4F complexes of plants: An evolutionary perspective. Comp Funct Genomics 2012: 287814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patrick RM, Mayberry LK, Choy G, Woodard LE, Liu JS, White A, Mullen RA, Tanavin TM, Latz CA, Browning KS (2014) Two Arabidopsis loci encode novel eukaryotic initiation factor 4E isoforms that are functionally distinct from the conserved plant eukaryotic initiation factor 4E. Plant Physiol 164: 1820–1830 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pietrzykowska M, Suorsa M, Semchonok DA, Tikkanen M, Boekema EJ, Aro EM, Jansson S (2014) The light-harvesting chlorophyll a/b binding proteins Lhcb1 and Lhcb2 play complementary roles during state transitions in Arabidopsis. Plant Cell 26: 3646–3660 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roose JL, Frankel LK, Bricker TM (2011) Developmental defects in mutants of the PsbP domain protein 5 in Arabidopsis thaliana. PLoS One 6: e28624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanfaçon H. (2015) Plant translation factors and virus resistance. Viruses 7: 3392–3419 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stirbet A, Govindjee (2011) On the relation between the Kautsky effect (chlorophyll a fluorescence induction) and Photosystem II: Basics and applications of the OJIP fluorescence transient. J Photochem Photobiol B 104: 236–257 [DOI] [PubMed] [Google Scholar]
- Strasser RJ, Tsimilli-Michael M, Srivastava A (2004) Analysis of the chlorophyll a fluorescence transient. In Papageorgiou GC, Govindjee, eds, Chlorophyll a Fluorescence: A Signature of Photosynthesis. Springer, Dordrecht, The Netherlands, pp 321–362 [Google Scholar]
- Valásek LS. (2012) ‘Ribozoomin’—translation initiation from the perspective of the ribosome-bound eukaryotic initiation factors (eIFs). Curr Protein Pept Sci 13: 305–330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Heerden A, Browning KS (1994) Expression in Escherichia coli of the two subunits of the isozyme form of wheat germ protein synthesis initiation factor 4F. Purification of the subunits and formation of an enzymatically active complex. J Biol Chem 269: 17454–17457 [PubMed] [Google Scholar]
- Wang A, Krishnaswamy S (2012) Eukaryotic translation initiation factor 4E-mediated recessive resistance to plant viruses and its utility in crop improvement. Mol Plant Pathol 13: 795–803 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang T, Wang L, Li C, Liu Y, Zhu S, Qi Y, Liu X, Lin Q, Luan S, Yu F (2015) Receptor protein kinase FERONIA controls leaf starch accumulation by interacting with glyceraldehyde-3-phosphate dehydrogenase. Biochem Biophys Res Commun 465: 77–82 [DOI] [PubMed] [Google Scholar]
- Yi X, Hargett SR, Frankel LK, Bricker TM (2006) The PsbQ protein is required in Arabidopsis for photosystem II assembly/stability and photoautotrophy under low light conditions. J Biol Chem 281: 26260–26267 [DOI] [PubMed] [Google Scholar]
- Yi X, Hargett SR, Frankel LK, Bricker TM (2008) The effects of simultaneous RNAi suppression of PsbO and PsbP protein expression in photosystem II of Arabidopsis. Photosynth Res 98: 439–448 [DOI] [PubMed] [Google Scholar]
- Yoshii M, Nishikiori M, Tomita K, Yoshioka N, Kozuka R, Naito S, Ishikawa M (2004) The Arabidopsis cucumovirus multiplication 1 and 2 loci encode translation initiation factors 4E and 4G. J Virol 78: 6102–6111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L, Kondo H, Kamikubo H, Kataoka M, Sakamoto W (2016a) VIPP1 has a disordered C-terminal tail necessary for protecting photosynthetic membranes against stress. Plant Physiol 171: 1983–1995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang L, Kusaba M, Tanaka A, Sakamoto W (2016b) Protection of chloroplast membranes by VIPP1 rescues aberrant seedling development in Arabidopsis nyc1 mutant. Front Plant Sci 7: 533. [DOI] [PMC free article] [PubMed] [Google Scholar]