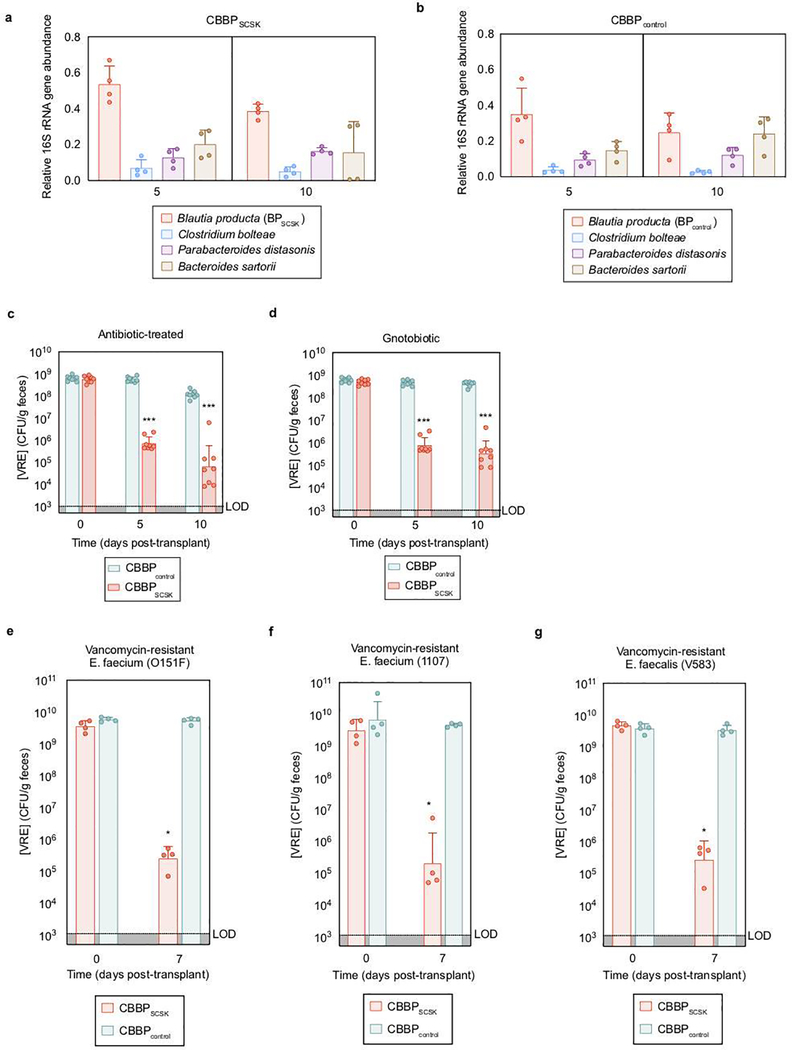
Extended Data Figure 2 |. BPSCSK, but not BPcontrol, reduces in vivo VRE colonization.
a,b, fecal samples collected from antibiotic- treated, VRE-dominated mice (n = 4 mice/1 independent experiment) orally gavaged with CBBP (a) or CBBPcontrol (b) were shotgun sequenced and the relative abundance of each species was determined by 16S rRNA. c,d, Antibiotic-treated (c) or germ- free (d) mice (n = 8 mice/2 independent experiments) were orally gavaged with VRE. Three days later, VRE-dominated mice received an oral gavage of CBBP or CBBPcontrol and VRE colonization was monitored by quantifying VRE in fecal samples. e-g, antibiotic-treated mice (n = 4 mice/1 independent experiment) were orally gavaged with different strains of clinical VRE isolates. Three days later, VRE-dominated mice received an oral gavage of CBBP or CBBPcontrol and VRE colonization was monitored by quantifying VRE in fecal samples. VRE strain 0151F is an E. faecium MLST type ST80 (e), VRE strain 1107 is an E. faceium MLST type ST412 (f), VRE strain V583 is an E. faecalis strain (g). VRE strains used were VRE (ATCC 700221) (a-d), VRE (0151F) (e), VRE (1107) (f), and VRE (V583) (g). *** p-value < 0.001. Center values (geometric mean), error bars (geometric s.d.). All statistical analyses were performed using the Mann-Whitney rank sum test (two-tailed) comparing two experimental conditions. *p- value < 0.05 (= 0.0286), ***p-value < 0.001.