Abstract
Terminally differentiated B cell, the plasma cell, is the sole cell type capable of producing antibodies in our body. Over the past 30 years, the identification of many key molecules controlling B cell activation and differentiation has elucidated the molecular pathways for generating antibody-producing plasma cells. Several types of regulation modulating the functions of the important key molecules in B cell activation and differentiation add other layers of complexity in shaping B cell responses following antigen exposure in the absence or presence of T cell help. Further understanding of the mechanisms contributing to the proper activation and differentiation of B cells into antibody-secreting plasma cells may enable us to develop new strategies for managing antibody humoral responses during health and disease. Herein, we reviewed the effect of different types of regulation, including transcriptional regulation, post-transcriptional regulation and epigenetic regulation, on B cell activation, and on mounting memory B cell and antibody responses. We also discussed the link between the dysregulation of the abovementioned regulatory mechanisms and B cell-related disorders.
Keywords: B cell, Antibody, Plasma cell, Transcription factor, microRNA, Epigenetic regulation, B cell malignancy, Autoimmune disease
Background
The life journey of B cells – from development to activation and differentiation
B cell development begins in the fetal liver and continues in hematopoietic stem cells (HSCs) in the bone marrow where the stromal cells provide cytokines and chemokines, such as C-X-C motif chemokine 12 (CXCL12) and interleukin (IL)-7, for early stage B cell development [1]. The signals from the stromal cells allow HSCs to differentiate into common lymphoid progenitor cells (CLPs), which express c-kit and IL-7 receptors to provide the survival and proliferation signals for CLPs once they encounter the ligands. Upon expression of the transcription factors, E2A and early B-cell factor (EBF), CLPs further develop into pro-B cells [2]. Starting from pro-B cells, B cells in the bone marrow experience a sequential genetic rearrangement of heavy-chain and light-chain immunoglobulin genes, the V(D)J recombination, resulting in the generation of the IgM-expressing immature B cells [3]. The immature B cells migrate from the bone marrow to the spleen, where they further differentiate into T1 and T2 stages. B cells finally become mature B cells that co-express IgD and IgM, after which they wait to be activated by foreign antigens [4].
For activation and differentiation into antibody-secreting plasma cells, mature B cells in the periphery lymphoid organs require two signals. The first signal is derived from antigen-coupled B cell receptors (BCRs), and the second signal can be delivered in a T cell-dependent (TD) or T cell-independent (TI) manner. TI antigens, such as lipopolysaccharides (LPS) and glycolipids, mostly give rise to short-lived plasma cells that produce low-affinity antibodies. TD responses, initiated by antigen encounter and interaction with follicular helper T (Tfh) cells [5], allow B cells to either quickly become short-lived plasma cells or enter the germinal center (GC) to differentiate into plasma cells or memory B cells with higher affinity toward the antigens. The GC can be polarized into the dark zone, where B cells undergo somatic hypermutation (SHM) at the variable regions of the BCR genes and clonal expansion, or the light zone, where B cells go through affinity maturation via interaction with Tfh cells and follicular dendritic cells (FDCs) to select B cell clones with high affinity BCRs [6]. Tfh cells produce the CD40 ligand for maintaining B cell survival, and IL-21 for promoting cell proliferation and differentiation [7]. In GC B cells, class switch recombination (CSR) that changes the constant region of the immunoglobulin from one isotype to another also occurs. GC B cells that are not positively selected by FDCs are eliminated by apoptosis, while the selected B cells may re-enter the dark zone to re-evolve BCRs with better affinity. The GC reaction allows B cells with high affinity receptors to further differentiate into plasma cells or memory B cells [8]. The GC-derived plasma cells circulate to the bone marrow and secrete antigen-specific antibodies to become long-lived plasma cells that provide long-term protection against specific antigens [9].
Main text
Transcriptional network in mature B cells and plasma cells
B cell differentiation is tightly controlled by a transcription regulation network. It involves the coordination of several transcription factors to promote the expression of antibody-secretion and plasma cell-related genes, and downregulate the B cell identity genes. B lymphocyte-induced maturation protein-1 (Blimp-1) is a critical transcription regulator of plasma cell formation, which mainly functions as a transcription repressor [10]. Deficiency in Blimp-1, encoded by the PR domain zinc finger protein 1 (Prdm1) gene, in mice impaired plasma cell differentiation, but did not affect B cell development [11]. Blimp-1 is expressed at low levels during the plasmablast stage, and at high levels in mature plasma cells [12]. Mechanistically, Blimp-1 represses the genes important for B cell identity, such as paired box protein 5 (PAX5), B-cell lymphoma-6 (BCL6) and BTB domain and CNC Homolog 2 (BACH2) [13, 14], and induces the activation of interferon regulatory factor 4 (IRF4) and X-Box Binding Protein 1 (XBP-1) [15, 16], suggesting that it has a multifunctional role in transcription regulation. PAX5 is expressed throughout the early B cell developmental stages and in mature B cells, and is crucial for the maintenance of the identity of the B cell lineage [17]. Furthermore, it regulates the expression of BCR component genes, such as CD19, CD21 and IgH, and other transcription factors important for B cells, like IRF4, IRF8, BACH2, Ikaros family zinc finger protein 3 (IKZF3) and PRDM1 [18]. After the B cells are activated and enter the GC B-cell stage in the secondary lymphoid organs, BCL6 and BACH2 expression begins. Upregulation of BCL6 is critical for the formation of GC and the prevention of plasma cell differentiation [19, 20]. Signaling through IL-21 receptor in proliferating GC B cells sustains the expression of BCL6 [21]. BACH2 is expressed in the pro-B to mature B cell stages, and is absent in plasma cells. Loss of BACH2 causes the lack of GC and Aicda, encoding activation-induced cytidine deaminase (AID), which is critical for SHM and CSR [22]. Both BCL6 and BACH2 suppress the expression of PRDM1 [23, 24]. In addition to Blimp-1, plasma cell formation requires IRF4, which represses Bcl6, therefore inducing Blimp-1 expression [25, 26]. Loss of IRF4 leads to impaired antibody production [27]. XBP-1 functions as a transcription regulator that is essential for Ig secretion and remodeling of the endoplasmic reticulum in plasma cells [28]. Absence of Blimp-1 causes impaired expression of XBP-1 and its downstream genes, suggesting that Blimp-1 is necessary for XBP-1 induction [29].
miRNA in B cell activation and differentiation
MicroRNAs (miRNAs) are small non-coding RNAs containing approximately 22–23 nucleotides (nts) in length that play important roles in post-transcriptional regulation in several biological processes, including apoptosis, cell proliferation, cell cycle, cell differentiation, hematopoiesis and cancer [30]. Studies on miRNA functions have revealed that one miRNA can specifically target hundreds of different mRNAs, and every single mRNA can be regulated by several different miRNAs [31, 32].
More than 1000 miRNAs have been identified in the human genome, which target about 60% of the human protein-encoding genes [33]. More than 100 different miRNAs are expressed by the immune system cells [34–36]. MiRNAs have the potential to broadly influence the molecular pathways that control the development and functions of innate and adaptive immune responses. Global miRNA expression profiling in various B cell stages has been reported [37]. An atlas of human mature B cell miRNAs (“miRNome”) was constructed with mature B cell line-specific short-RNA libraries coupled with low throughput sequencing [36]. Furthermore, miRNA array has been extensively used to identify miRNA expression profiles. For example, miRNA array profiling of CD5+-activated and CD5−-resting B cells from human peripheral blood and tonsils revealed that 34 miRNAs were enriched in CD5+-activated B cells, and eight of them, including miR-323, miR-138, miR-9*, miR-211, miR-129, miR-373, miR-135a and miR-184, were highly expressed miRNAs capable of co-targeting ZEB1 and TP53 [38]. The importance of miRNAs in B cell lineage was emphasized by a study on a mouse gene knockout model in which Dicer, encoding a key enzyme responsible for the generation of miRNAs from their precursors [39], is deleted in a B cell-specific manner. B cell-specific deletion of Dicer exhibited a developmental block at the pro-B to pre-B stages and revealed that miRNAs may have a role in controlling V(D)J recombination for generating antibody diversity in the early stage of B cell development [40].
We have investigated the changes in the miRNA expression inherent to the transcription network in plasma cell differentiation (Fig. 1) [41]. Two large scale analyses, deep-sequencing and miRNA microarray, were used to elucidate the changes in the expression of miRNAs during human plasma cell differentiation. In this study, human peripheral blood B cells were treated with the stimuli provided by Tfh-mimicking signals. Our computational analysis revealed that 34 and 60 miRNAs with significant reads were upregulated and downregulated, respectively, during human plasma cell differentiation. We characterized the relationship between differentially expressed miRNAs and transcription factors during plasma cell differentiation. We found that several differentially expressed miRNAs commonly target a single key transcription factor. We thus called these miRNAs a “miRNA hub”. It is noteworthy that these miRNA hubs collaboratively regulate the expression of key transcription factors, thereby enabling the formation of human plasma cells in culture. Specifically, we found that upregulated miRNA hubs, including miR-34a-5p, miR-148a-3p, miR-183-5p and miR-365a-3p, directly repressed endogenous BCL6, BACH2 and FOXP1 expression during plasma cell differentiation. However, downregulated miRNA hubs, including miR-101-3p, miR-125b-5p and miR-223-3p, target the PRDM1 3′ untranslated region (UTR). We further showed that NF-κB and PRDM1 contribute to the induction and repression of upregulated and downregulated miRNA hubs, respectively, during plasma cell differentiation. Moreover, our computational analysis unveiled that the transcription factor, FOXP1, is regulated by an induced miRNA hub and plays a role in prohibiting plasma cell differentiation.
Fig. 1.
The action of miRNAs and key transcription factors in coordinately directing plasma cell differentiation. Several factors are involved in the negative regulation of PRDM1 in mature B cells, including BCL6/BACH2/FOXP1 and the miR-101-3p, miR-125b-5p, miR-223-3p miRNA hub. During B cell activation, NF-κB induces not only PRDM1 for the initiation of plasma cell differentiation, but also the miR-34a-5p, miR-148a-3p, miR-183-5p and miR-365a-5p hub. The induced miRNA hub including miR-34a-5p, miR-148a-3p, miR-183-5p and miR-365a-5p downregulates BCL6/BACH2/FOXP1, thereby establishing elevated levels of PRDM1 for driving plasma cell differentiation. Induced PRDM1 in turn suppresses the expression of the miR-101-3p, miR-222-3p and miR-223-3p hub, and BCL6/BACH2/FOXP1, resulting in commitment to the plasma cell fate. Lines with arrow and bars indicate upregulation and downregulation, respectively. miRNAs in a red or green box represent upregulated or downregulated expression, respectively
With regard to the function of miRNAs in B cells, accumulating reports have demonstrated their roles in regulating B cell development and shaping the properties of the effector functions of B cells. One of the first miRNAs identified with functional significance to B cell development is miR-181a (now called miR-181a-5p). Overexpression of miR-181a-5p in HSCs increased the number of B cells in vitro and in vivo [42]. miR-181a-5p targets the differentiation inhibitor, ID2, which suppresses the early differentiation of B cells [43]. MiR-150 regulates the differentiation of normal B cells into antibody-secreting plasma cells. Several studies have indicated that miR-150 is highly expressed in mature B cells, relatively lowly expressed in immature B cells, and has the lowest expression during the pro-B to pre-B cell transition [44, 45]. One key target of miR-150 is c-Myb, which is required for the development of pro-B cells [46, 47]. Another study demonstrated that the p53-induced miRNA, miR-34a (now called miR-34a-5p), impaired B cell development. MiR-34a targets the 3’UTR of Forkhead box protein P1 (Foxp1) mRNA, which regulates the expression of recombination-activating genes (Rag1 and Rag2) in the pro-B to pre-B transition stages [48, 49]. MiR-155 is a highly expressed miRNA in GC B cells, however it is expressed at relatively low levels in HSCs and mature B cells [34, 50]. Knockout of the miR-155 gene in mice caused defective CSR and impaired differentiation of antibody-secreting plasma cells, by targeting Spi1 (encoding PU.1) and Aicda [51–53]. Besides miR-155, miR-181b has been shown to negatively regulate CSR by targeting Aicda [54]. Additionally, several other studies have indicated that miR-9, miR-125b, the miR-17–92 cluster and the miR-30 family are expressed in GC B cells and enhance plasma cell differentiation [37, 55]. Deletion of the miR-17–92 cluster in B cells in mice caused enhanced homing of plasma cells to the bone marrow upon TD immunization, likely owing to the effect of miR-17–92 on S1pr1, a gene important for the egress of lymphocytes from the lymphoid organs [56].
miRNAs in B cell malignancy and autoimmune diseases
Lymphoma, including B and T-cell lymphoma, is malignancy of lymphatic cells, which affects more than a million people worldwide. Many miRNAs that contribute to B cell lymphomagenesis are also key regulators in normal hematopoiesis and lymphopoiesis. MiRNAs that affect tumorigenesis are called onco-miRs or tumor suppressor miRs [57, 58]. The first reported onco-miR is miR-155, which is upregulated in normal plasma cell differentiation and overexpressed in several types of B cell lymphomas [59]. It is noteworthy that mice with miR-155 overexpression in a B cell-specific manner develop high-grade B-cell lymphoma resembling diffuse large B-cell lymphoma (DLBCL) [59], likely owing to the effect of miR-155 on SHIP1, which promotes TNFα-dependent cell proliferation [60]. MiR-155 is also a key regulator of the PI3K/AKT pathway in DLBCL. It promotes cell proliferation and inhibits apoptosis of DLBCL cells [61]. The most studied tumor-suppressor miRNA is miR-34a, which forms parts of the p53 network [62]. p53 directly induces miR-34a expression, but at the same time miR-34a enhances p53 expression via inhibiting SIRT1, a regulator of p53 deacetylation, resulting in a positive feedback loop [63]. An additional study has shown that miR-34a reduces tumor growth in mice by targeting Foxp1 [64]. Another well-studied tumor-suppressor miRNA in B cell malignancy is miR-101 (now called miR-101-3p). The decreased expression of miR-101 correlated with the pathogenesis and prognosis of DLBCL, while upregulation of miR-101 in DLBCL inhibited cell proliferation and facilitated apoptosis by targeting MEK1 [65]. Furthermore, miR-183 is differentially expressed in the three Hodgkin’s lymphoma (HL) subtypes and in EBV+ and EBV− HLs. However, elucidation of the exact mode of action of miR-183 in HL requires further investigation [66]. Another study has shown that the expression of miR-223 and miR-125b in DLBCL is higher than in follicular lymphoma (FL), suggesting that the high expression of miR-223 and miR-125b may contribute to the transformation of DLBCL [67]. The aberrant expression of miR-125b in mantle cell lymphoma (MCL) has also been reported. A miRNA expression profile study was able to segregate MCLs into three different groups with distinct biological and clinical features [68].
Unlike DLBCL, which is usually formed from mutated or dysregulated normal GC B cells [69], multiple myeloma (MM) arises from malignant plasma cells in the bone marow [70, 71]. The molecular mechanisms underlying the dysregulation of p53 in MM have been intensively investigated for many years [72, 73]. Recently, it has been suggested that many miRNAs reported to negatively regulate p53 expression may also have implications in MM cells. For example, miR-125b is an onco-miR in hematologic malignancies as it targets p53 [74] and other components of the p53 pro-apoptotic network, including BAK1, PUMA, BMF, TRP53INP1 and Krupple-like factor 13 (KLF13) [75, 76]. Furthermore, dexamethasone induces the expression of miR-34a in MM cells [75, 77]. MiR-34a suppresses the expression of SIRT1 deacetylase, and thus allows the maintenance of acetylation and inactivation of p53. Results from a gene microarray study showed that the onco-miRs, miR-19b and miR-20a, were differently expressed in patients with MM and normal controls. MiR19b/20a promotes cell proliferation and migration, and inhibits cell apoptosis by targeting PTEN in patients with MM [78]. Moreover, miR-9 is another onco-miR that represses apoptosis in MM patients and MM cell lines by regulating the TRIM56/NF-κB pathway [79]. Taken together, these studies indicate the roles of miRNAs in the progression of B cell malignancy and their therapeutic potential for the treatment of B cell malignancy.
Recently, miRNAs have also been reported to play a role in immune dysfunction and autoimmune diseases. Systemic lupus erythematosus (SLE) is a multisystem autoimmune disorder that mostly affects women at childbearing age [80]. B cells play a cardinal role in the pathogenesis of SLE; more than 90% of SLE patients have high levels of antinuclear antibodies in the sera, including anti-dsDNA antibodies and anti-snRNP antibodies [81]. Several miRNAs have been identified as biomarkers during the development of SLE. miRNA microarray analysis first revealed that miR-21 and miR-17-5p are differently expressed in peripheral blood mononuclear cells (PBMCs) of SLE patients [82]. The increased expression of miR-7 downregulates the expression of PTEN in B cells of SLE patients and contributes to hyper-activation of B cells [83]. Moreover, miR-17-5p is downregulated in PBMCs of SLE patients [82]. MiR-17-5p downregulates c-MYC expression during SLE formation. Transfection of miR-17-5p mimics into PBMCs from SLE patients caused a dramatic reduction in E2F1 and c-MYC expression, which resulted in reduced mRNA levels of the IFN-inducible gene, MxA [84]. Screening of SLE-specific miRNAs from 42 B cell-related miRNAs by a miRNA PCR Array identified that 14 miRNAs, including miR-103, miR-150, miR-20a, miR-223, miR-27, miR-15b, miR-16, miR-181a, miR-19b, miR-22, miR-23a, miR-25, miR-92a and miR-93, were significantly downregulated in the plasma of SLE patients, compared with the plasma of healthy donors [85]. Moreover, six plasma miRNAs, including miR-92a, miR-27a, miR-19b, miR-23a, miR-223 and miR-16, were expressed at significantly decreased levels in SLE patients than in rheumatoid arthritis (RA) patients. These results demonstrated that these differentially expressed miRNAs in the plasma of SLE and RA patients can potentially be used as a diagnostic signature to distinguish between these two diseases. Another study using miRNA expression array revealed the serum miRNA expression profiles of SLE patients and healthy donors [86], showing that miR-371b-5p, miR-5100 and miR-146a-5p were increased in active SLE. These studies suggest the potential roles of these identified miRNAs in the development of SLE. The function of miRNAs in B cell malignancy and autoimmune diseases was summarized in Table 1.
Table 1.
miRNAs in B cell malignancy and autoimmune diseases
| miRNA | Biological function | Target genes | Disease type | References |
|---|---|---|---|---|
| miR-155 | Promote TNFα-dependent proliferation | SHIP1 | DCBCL | [60] |
| miR-155 | Promote proliferation and inhibits apoptosis | PI3K/AKT pathway | DCBCL | [61] |
| miR-101 | Inhibit proliferation and facilitate apoptosis | MEK1 | DCBCL | [65] |
| miR-125b | Inhibit apoptosis | p53 | MM | [74, 75] |
| miR-19b/20a | Promote proliferation, migration, and inhibit apoptosis | PTEN | MM | [78] |
| miR-34a | Regulate cell cycle progression, cellular senescence and apoptosis | SIRT1 | MM | [77] |
| miR-34a | Reduce tumor growth | Foxp1 | DLBCL | [49] |
| miR-9 | Inhibit apoptosis | TRIM56/NF-κB pathway | MM | [79] |
| miR-7 | Promote B cell hyperresponsiveness | PTEN | SLE | [83] |
| miR-17-5p | Inhibit IFN-inducible gene | c-MYC, and E2F1 | SLE | [84] |
miRNA therapeutics
MiRNA-targeted therapeutics can be divided into miRNA mimics [87] and inhibitors (also called anti-miRs) [88]. The effect of modulation of miRNAs’ levels on B cell malignancy has been demonstrated. Studies on a mouse model of miR-155-induced lymphoma, in which mir-155 is expressed under the control of doxycycline, demonstrated that doxycycline withdrawal resulted in suppression of mir-155 expression and subsequent tumor shrinkage [89]. In this mouse model, anti-miR-155 treatment resulted in decreased tumor burden, indicating that miR-155 inhibition has therapeutic potential [89]. In contrast, miR-34a has been identified as a tumor suppressor miRNA by repressing several target genes, such as cyclin-dependent kinase 4 (CDK4), CDK6, BCL2, MET, Notch, c-MYC, AXL and FOXP1 [48, 49, 90]. Several preclinical studies using miR-34 mimics have demonstrated their potential as anticancer therapeutics. For instance, miR-34a mimics showed promising anti-tumor activity in mouse models of lung [91], liver [92] and prostate [93] cancer. In these cases, significant inhibition of tumor growth was observed, which correlated with reduced expression of target proteins, such as c-MYC and BCL-2, in tumors. As a result of the above-mentioned studies, several miRNA-targeted therapeutics have reached clinical development. Currently, there are more than 20 clinical trials applying miRNA and siRNA-based therapeutics [94]. For instance, anti-miRs are single-stranded first-generation antisense oligonucleotides, which have been modified and designed to block the function of miRNAs. Anti-miRs with a 2′-O-methoxyethyl modification (2′-OM) are called antagomiRs [95]. These synthetic small RNA molecules have a complementary sequence to the target miRNA, and are able to strongly bind to the target miRNA and thereby block its function. MiRNA mimics are synthetic double-stranded small RNA molecules matching the corresponding miRNA sequence, and therefore functionally able to restore the loss of miRNA expression in diseases. MiR-34a mimics reached phase I clinical trials for treating cancer, but this trial was halted at phase I owing to immune-related adverse events [94]. Effective delivery of RNA-based therapeutics to the target tissues has been a challenge in their therapeutic application [96]. Development of better in vivo delivery systems to reach the target specifically and efficiently to overcome the bottleneck of RNA-based therapy (including miRNA) in the clinic is the next important task.
Epigenetic regulation in B cell activation and differentiation
Epigenetic regulation is critical for coordination with the abovementioned transcription regulation networks in molecular programming during B cell activation and differentiation. The synergistic effects of both genetically and environmentally induced epigenetic modifications have been demonstrated to contribute to plasma cell differentiation and the etiopathogenetic mechanisms of the generation of B cell- or plasma cell-related diseases, such as autoimmune disorders and lymphomagenesis [97]. In general, the quiescent naïve B cells in peripheral lymphoid organs display inactive chromatin structures that show genome-wide DNA hypermethylation [98] and methylation of histone 3 K9 (H3K9) and H3K27 [99]. At this stage, the expression level of genes important for regulating B cell identify and antigen recognition is regulated by histone deacetylase 7 (HDAC7) [100]. During early B cell development, HDAC7 represses myeloid and T cell genes in early B cell progenitors [100]. Enhancer of zeste homolog 2 (Ezh2) is able to catalyze H3K27me3, which is associated with long-term repression [101]. In GC B cells, Ezh2 is highly expressed [102]. Deletion of Ezh2 in mice in a GC-specific manner caused impaired GC response, memory B cell formation and antibody responses compared with the control mice, suggesting that Ezh2 is essential for B cell functions [103]. In GC reactions, a number of histone modifications, including acetylated H3 and H4, and DNA double-strand breaks (DSBs)-induced phosphorylated H2AX (γH2AX), are associated with CS [104, 105]. However, acetylated H3 and H4 may not be linked with SHM activation. Instead, the histone modification pattern of SHM consists of phosphorylation of histone H2B on serine 14 (H2BSer14P), which is also responsive to DSBs [106].
In addition, in GC, the expression and action of AID is regulated by a series of epigenetic mechanisms. The suppression of Aicda in naïve B cells is due to DNA hypermethylation at the promoter region [107]. The H3 acetylation level of the Aicda gene locus in naïve B cells is low compared with the global H3 acetylation levels of other nearby genes. After B cells are stimulated, the Aicda gene locus is demethylated and becomes enriched with H3K4me3, H3K9ac and H3K14ac, which are associated with active histone marks [108]. Downregulation of Aicda in memory B cells and plasma cells may result from re-methylation of the Aicda gene locus. The histone chaperone, Spt6, regulates CSR and AID expression through two distinct types of histone modifications to generate the euchromatin status, namely, H3K4me3 and H3K36me3, respectively. Spt6 is also required for the establishment of H3K4me3 marks in the IgH variable region during SHM [109]. In terms of the functional mode of action of AID, it interacts with ubiquitinated chromatin. Specifically, ubiquitination of H2BK120 and H2AK119 is colocalized with mismatched DNA polymerase η in the AID-containing region [110].
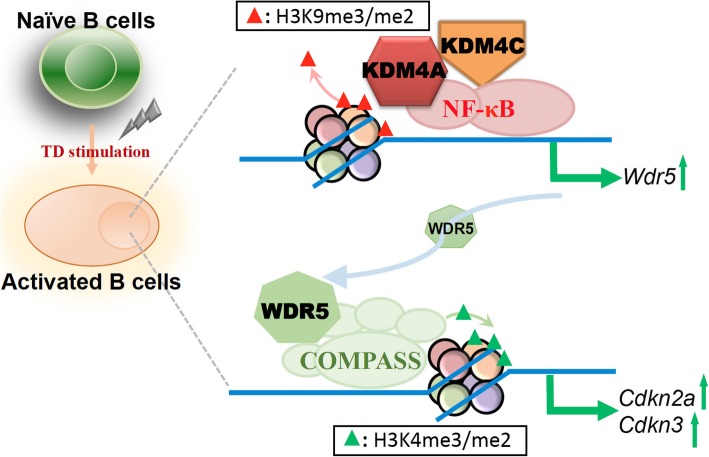
The global levels of H3K9me2/me3 and H3K4me2 are all upregulated after LPS and IL-4 stimulation in a B cell culture [111]. We have also reported changes in histone modifications in B cells treated with Tfh cell-mimicking signals (Fig. 2) [99]. We found that the global levels of H3K9me3/me2 were reduced after stimulating mouse spleen B cells with Tfh cell-mediated signals. Furthermore, a systemic search of the epigenetic modifiers that contribute to the downregulation of H3K9me3/me2 revealed that the histone demethylases, KDM4A and KDM4C, were upregulated in mouse spleen B cells treated with Tfh cell-mimicking signals, whereas stimulation with LPS did not induce a similar pattern of KDM4A/KDM4C-mediated epigenetic changes. Functionally, depletion of KDM4A and KDM4C in response to Tfh cell-mimicking signals accelerated B cell activation and proliferation. Our genome-wide analysis using chromatin immunoprecipitation sequencing (ChIP-seq) combined with cDNA microarray analyses further revealed KDM4A and KDM4C targets during B cell activation. Among these, WDR5, an MLL complex member that facilitates H3K4 methylation [112], was further demonstrated to regulate the cell cycle; in particular, the cell cycle inhibitors, Cdkn2 and Cdkn3. Mechanistically, de novo motif analysis of the ChIP-seq data of KDM4A and KDM4C revealed that NF-κB p65 interacts with KDM4A and KDM4C to regulate gene expression, including WDR5.
Fig. 2.
Proposed model of the role of KDM4A and KDM4C in B cell activation. During activation by Tfh cell-mimicking signals, the induced demethylases, KDM4A and KDM4C, cooperate with NF-κB to upregulate the expression of Wdr5 by removing H3K9me3/me2. WDR5, a core subunit of the COMPASS histone H3K4 methyltransferase complex, in turn facilitates the transcription of Cdkn2a and Cdkn3 by elevating H3K4me3/me2. Both CDKN2A and CDKN3 are involved in the regulation of stimulated B cell proliferation
Less is known about epigenetic modification during plasma cell differentiation. Genes expressed during plasma cell differentiation correlated with the acquisition of H3K4me1 and H3K4me3 histone marks, which are the markers of active promoters and distal enhancers [113]. Blimp-1 is a key transcription factor in directing plasma cell differentiation [15], thus, epigenetic regulation of PRDM1 expression and function would affect the humoral responses. BCL6 and HDAC4, − 5 or − 7 form stable complexes to mediate the decreased levels of histone acetylation on the PRDM1 promoter in GC B cells [114, 115]. Furthermore, treatment of B cells with HDAC inhibitors, such as trichostatin A (TSA) and butyrate, induced the expression of Blimp-1 and J chain, but decreased the expression of c-Myc and Pax-5. TSA treatment also promotes the expression of CD138 and downregulates surface IgM [116]. Several Blimp-1 target genes, including Pax5 and Spib, showed decreased histone acetylation in plasma cells [117, 118]. Therefore, histone acetylation regulates the transcription of genes controlling B cell differentiation. In general, Blimp-1 is a transcription repressor able to recruit co-repressor proteins and histone modifiers to induce transcription repression. We have previously shown that a proline-rich domain of Blimp-1 directly interacts with LSD1, an H3K4me2/1 and H3K9me2/1-specific demethylase [117]. These proteins collaboratively remodel the chromatin accessibility of Blimp-1 targets, and thereby modulate the expression of Blimp-1 targets [117]. Disruption of the Blimp-1 interaction with LSD1, depletion of LSD1 expression and inhibition of LSD1 function all reduced the formation of antibody-secreting plasma cells. The five C2H2 zinc fingers of Blimp-1 contain a nuclear localization signal and show DNA binding activity. They interact with HDAC2 [118] and H3K9-specific methyltransferase G9a [12], both of which are histone modifiers that facilitate the inactive chromatin and reduce transcription. These findings suggest that Blimp-1 regulates the differentiation of plasma cells by interacting with multiple chromatin modifiers.
Dysregulated epigenetic mechanisms in B cell malignancy and autoimmune diseases
The dysregulated epigenetic mechanism resulting in inadequate cell cycle is one of the major causes leading to malignant B cells. For instance, the H3K27-specific methyltransferase EZH2 is highly expressed in the GC, where it cooperates with Polycomb Repressive Complex-2 (PRC2) [119]. Mutation of Ezh2 in mice resulted in GC hyperplasia [120], which in part may stem from the enhanced repression of CDKN1A [120]. Furthermore, EZH2 can restrain plasma cell differentiation by establishing the H3K27me3 marks at the Prdm1 and Irf4 loci [103]. In human DLBCL cells, a mutant form of EZH2 silences PRDM1 through the recruitment of PRC2, resulting in the promotion of growth of the GC-type DLBCL [103]. Hence, 22% of GC-type DLBCL cases carry EZH2 mutations [121]. In addition, KMT2D (also known as MLL2 or MLL4) is a member of the SET1 family of histone methyltransferases (HMTs) that facilitates the establishment of H3K4me2/me1 at enhancer regions and was frequently found to be mutated in FL (accounting for 70–80% of cases) and DLBCL [122–124]. KMT2D regulates genes involved in the CD40, JAK-STAT, TLR and BCR signaling pathways [123]. Mice harboring a Kmt2d deletion in B cells showed B cell proliferation advantages and B cell lymphoproliferative diseases through collaboration with BCL-2 [124]. The CREBBP gene encodes a H3K18 and H3K27-specific lysine acetyltransferase that tags the transcription activation [125, 126]. Further inactivating mutations and/or copy number losses of CREBBP occur in more than half of the FL cases and in nearly 20% of the DLBCL cases [127]. Accordingly, Crebbp deletion in the GC stage in mice in the presence of BCL-2 overexpression promotes the development of FL [128].
MM is plasma-cell malignancy with a slow progress feature [129]. One of the well studied HMTs in MM is the H3K36me2 and H4K20me2/me3-specific lysine methyltransferase, MMSET (also known as WHSC1 or NSD2) [130, 131]. MMEST is a DNA damage-responsive protein that catalyzes H4K20 methylation and induces the accumulation of 53BP1 at DNA damage sites [132]. Fifteen to 20 % of MM cells carry the t(4;14) translocation, which generates the fusion of MMSET to the IgH locus, resulting in the upregulation MMSET, which correlates with poor prognosis [133]. MMSET forms a complex with epigenetic repressors by recruiting Sin3a, HDAC1, HDAC2 and the H3K4-specific demethylase, LSD1/KDM1A. The MMSET complex then indirectly induces c-MYC levels by repressing miR-126, thereby sustaining the proliferation of MM cells [134, 135]. Nevertheless, the t [4, 14] translocation in MM cells remodels the chromatin structure that carries the H3K36me2 histone activation mark and induces global reduction in H3K27me3 by upregulating EZH2 at the oncogenic loci [131, 133]. The extent of H3K27 methylation is associated with the malignancy of plasma cells. EZH2 is upregulated during MM progression [136], resulting in enhanced IL-6R expression, c-MYC activation, miR-26a downregulation and long non-coding RNA expression, thereby affecting the proliferation and apoptosis of MM cells [137, 138]. In contrast, mutations of the H3K27-specific demethylase, KDM6A, are found in 10% of primary MM samples [138, 139]. Inhibition of EZH2 decreases the growth of KDM6A-depleted MM cells. The KDM6A-mutated MM cells are more sensitive to EZH2 inhibitor-induced apoptosis through reactivation of BCL6 and subsequently repression of IRF4 and c-MYC [140]. These results showed the synergetic effect of EZH2 and KDM6A, which collaboratively control the expression of a set of oncogenic genes. The levels of the H3K9-specific demethylase, KDM3A, have been demonstrated to be increased in MM cells. It has been shown that the KDM3A-KLF2-IRF4 axis promotes homing of MM cells to the bone marrow and their adherence to bone marrow stromal cells. KDM3A maintains the transcriptional activity of KLF2 and IRF4 [141, 142]. KDM3A is upregulated by hypoxia-induced HIF1α that induces the expression of the long non-coding RNA, MALAT1, which in turn facilitates the upregulation of glycolytic and anti-apoptotic genes in MM cells [143, 144].
Recent reports have shown the significance of abnormal epigenetic regulation in the pathogenesis of SLE. A significantly reduced level of DNMT1 and DNMT3A transcripts was found in SLE patients compared with healthy controls [145]. The high IL-6 levels produced by SLE patients resulted in impaired induction of DNMT1, which in turn caused the demethylation of DNA in CpG islands in the cytoplasmic isoform of CD5, CD5-E1B. CD5-E1B is a negative regulator of BCR signaling, thereby establishing the immune tolerance in SLE B cells [146, 147]. Beyond the coding genes, the DNA methylation state of non-coding regions in the genome of SLE patients was also found to be altered. The hypomethylated long interspersed nuclear elements, but not the short interspersed nuclear elements, in SLE B cells correlate with the disease prognosis [148, 149]. Furthermore, our previously identified KDM4A/KDM4C/WDR5/CDKNs epigenetic pathway induced by Tfh cell-mimicking signals is dysregulated in B cells isolated from SLE patients [99]. We found that activated normal human peripheral blood B cells exhibited a significant reduction in H3K9me2 and H3K9me3, while the levels of H3K9me2 and H3K9me3 in stimulated SLE B cells did not change significantly. Accordingly, KDM4A and KDM4C mRNA levels were significantly reduced in the steady state and the stimulated SLE B cells, compared with normal B cells. Together, the effects of dysregulated histone modifiers on B cell malignancy and autoimmune diseases were summarized in Table 2.
Table 2.
Epigenetic controls in B cell malignancy and autoimmune diseases
| Histone Modifier | Biological function | Target genes (direct or indirect) |
Disease type | References |
|---|---|---|---|---|
| EZH2 | Promote growth | CDKN1A | DCBCL | [120] |
| EZH2 | Retrain plasma cell differentiation | PRDM1 and IRF4 | Non-Hodgkin lymphomas | [103] |
| KDM4A | Promote B cell activation and differentiation | WDR5 | SLE | [99] |
| KDM4C | Promote B cell activation and differentiation | WDR5 | SLE | [99] |
| LSD1 | Promote differentiation | Blimp-1 targets, such as CIITA and c-MYC | MM | [117] |
| MMSET | Promote growth | c-MYC | MM | [135] |
| KDM6A | Promote oncogenesis | BCL-6, IRF4 and c-MYC | MM | [140] |
| KDM3A | Promote homing of MM cells | KLF2, and IRF4 | MM | [141] |
| WDR5 | Inhibit proliferation | CDKN2, and CDKN3 | SLE | [99] |
| CREBBP | Regulate GC reaction and proliferation | BCL-6 targets, such as BCL-2 and PRDM1 | FL and DCBCL | [128] |
| KMT2D | Repress lymphoma development | Genes in CD40, JAK-STAT, TLR and BCR pathways | FL and DCBCL | [123] |
| DNMT1 | Promote BCR signaling | CD5, and CD5-E1B | SLE | [146] |
Epigenetic therapeutics
The aberrant epigenetic profiles of malignant cells, such as in MM, have been established in past decade. To target these aberrant epigenetic regulation mechanisms in MM, there are three categories of epigenetic modulating therapeutic agents under development: DNA methyltransferase inhibitors (DNMTi), histone deacetylase inhibitors (HDACi) and histone lysine methyltransferase inhibitor (HKMTi). Several DNMTis, such as 5-azacytidine (AZA, Vidaza) and 2-deoxy-5-aza-cytidine (DAC, Decitabine, Dacogen), have been ideal therapeutics for myelodysplastic syndrome [150, 151]. In MM, AZA and DAC have been demonstrated to have anti-MM effects, which cause cell cycle arrest and generation of oxidative stress to induce necrosis and apoptosis [152, 153]. In recent years, great progress has been achieved with HDACis in drug development for cancer therapy. The anti-MM effects of HDACis rely on their chromatin remodeling activity to induce apoptosis, cell cycle arrest and autophagy, as well as to suppress angiogenesis [154]. In particular, HDAC6 regulates deacetylation of α-tubulin and heat shock protein 90α (HSP90α), thereby affecting cell motility and cell adhesion, as well as the aggresome degradation pathway in response to misfolded proteins in MM cells [155]. Furthermore, inhibition of HDAC6 shows minimal side effects on healthy cells [156]. Hence, several clinical trials have applied HDAC6-specific inhibitors, such as Rocilinostat [157], ITF2357 [158] and Panobinostat [159, 160], in MM therapy.
The bromodomain (BRD) and extra-terminal (BET) family of BRD-containing proteins is a group of proteins that recognize acetylated lysine residues of histones and regulate gene expression. Hence, suppression of the activity of BRD-containing proteins is an effective way to control the histone-acetylation-dependent gene activation. The pan-BET inhibitor, GSK525762, inhibits growth of a broad spectrum of human hematological cancer cells, including MM [161]. Other ongoing clinical studies showed that BET inhibitors, such as OTX015 and CPI-0610, which selectively block BRD2, BRD3 and BRD4, were also used in the pre-clinical or clinical trials for MM or lymphoma [162, 163].
In addition, EZH2 inhibitors have been developed to block MM. Currently, the EZH2 inhibitor, Tazemetostat (EPZ-6438), is under clinical trials in combination with immunomodulatory imide drugs (IMiDs) for treating a subgroup of MM patients [164]. GSK2816126, another EZH2 inhibitor, is also in clinical trials for MM. It induces apoptosis in MM cells by downregulating mitochondrial activity [165].
Conclusions
Accumulating research efforts have been made to elucidate the molecular pathways regulating the B cell responses and antibody production. Studying the regulatory mechanisms of B cell responses has become an emerging research topic with the need to further understand the pathways that control the new coming pathogens through vaccination or to combat cancers. In addition to the above-described regulatory mechanisms in B cell activation and differentiation, there are other types of regulation involved, such as glycosylation and SUMOylation. Studies on these regulatory mechanisms open opportunities for identifying new druggable targets to control B cell-related diseases such as autoimmune diseases and B cell malignancies.
Acknowledgements
Not applicable.
Abbreviations
- 2′-OM
2′-O-methoxyethyl
- AID
Activation induced cystidine deaminase
- ASOs
Antisense oligonucleotides
- BACH2
BTB domain and CNC homologue 2
- BCL6
B-cell lymphoma 6
- BCRs
B cell receptors
- BET
Bromodomain extra-terminal
- Blimp-1
B lymphocyte-induced maturation protein-1
- BRD
Bromodomain
- CDK4
Cyclin-dependent kinase 4
- CDK6
Cyclin-dependent kinase 6
- CLPs
Common lymphoid progenitors
- CSR
Class switch recombination
- DLBCL
Diffuse large B-cell lymphoma
- DNMTi
DNA methyltransferase inhibitors
- EBF
Early B-cell factor
- ER
Endoplasmic reticulum
- Ezh2
Enhancer of zeste homolog 2
- FDCs
Follicular dendritic cells
- FL
Follicular lymphoma
- FOXP1
Forkhead box protein P1
- GC
Germinal center
- H3K27
methylation of histone 3 K27
- H3K9
methylation of histone 3 K9
- HDAC7
Histone deacetylase 7
- HDACi
Histone deacetylase inhibitor
- HKMTi
Histone lysine methyltransferase inhibitor
- HL
Hodgkin’s lymphoma
- HMTs
Histone methyltransferases
- HSCs
Hematopoietic stem cells
- IKZF3
Ikaros family zinc finger protein 3
- IMiDs
Immunomodulatory imide drugs
- IRF4
Interferon-regulatory factor 4
- IRF8
Interferon-regulatory factor 8
- KLF13
Krupple-like factor 13
- LPS
Lipopolysaccharides
- MCL
Mantle cell lymphoma
- miRNA
microRNA
- MM
Multiple myeloma
- MMSET
H4K20me2/me3-specific lysine methyltransferase
- PAX5
Paired box gene 5
- PBMCs
Peripheral blood mononuclear cells
- PRC2
Polycomb Repressive Complex-2
- PRDM1
PR domain zinc finger protein 1
- RA
Rheumatoid arthritis
- RAG-1
Recombination-activating gene-1
- RAG-2
Recombination-activating gene-2
- SHM
Somatic hypermutation
- SLE
Systemic lupus erythematosus
- TD
T-cell dependent
- Tfh
follicular helper T
- TI
T-cell independent
- TSA
Trichostatin A
- XBP-1
X-box-binding protein 1
Authors’ contributions
DYT, KHH, CWC and KIL wrote the manuscript. All authors read and approved the final manuscript.
Funding
This work was supported by grants from the Taiwan Bio-Development Foundation, Ministry of Science and Technology (MOST 106–2320-B-001-011-MY3 and MOST 107–0210-01-19-01), National Health Research Institute (NHRI-EX108-10835SI) and Academia Sinica (AS-IA-107-L05).
Availability of data and materials
Not applicable.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Dong-Yan Tsai and Kuo-Hsuan Hung contributed equally to this work.
References
- 1.Nagasawa T. The chemokine CXCL12 and regulation of HSC and B lymphocyte development in the bone marrow niche. Adv Exp Med Biol. 2007;602:69–75. doi: 10.1007/978-0-387-72009-8_9. [DOI] [PubMed] [Google Scholar]
- 2.O'Riordan M, Grosschedl R. Coordinate regulation of B cell differentiation by the transcription factors EBF and E2A. Immunity. 1999;11(1):21–31. doi: 10.1016/S1074-7613(00)80078-3. [DOI] [PubMed] [Google Scholar]
- 3.Krangel MS. Gene segment selection in V(D)J recombination: accessibility and beyond. Nat Immunol. 2003;4(7):624–630. doi: 10.1038/ni0703-624. [DOI] [PubMed] [Google Scholar]
- 4.Chung JB, Silverman M, Monroe JG. Transitional B cells: step by step towards immune competence. Trends Immunol. 2003;24(6):343–349. doi: 10.1016/S1471-4906(03)00119-4. [DOI] [PubMed] [Google Scholar]
- 5.Nutt SL, Hodgkin PD, Tarlinton DM, Corcoran LM. The generation of antibody-secreting plasma cells. Nat Rev Immunol. 2015;15(3):160–171. doi: 10.1038/nri3795. [DOI] [PubMed] [Google Scholar]
- 6.Manser T. Textbook Germinal Centers? J Immunol. 2004;172(6):3369–3375. doi: 10.4049/jimmunol.172.6.3369. [DOI] [PubMed] [Google Scholar]
- 7.Bishop GA, Hostager BS. Signaling by CD40 and its mimics in B cell activation. Immunol Res. 2001;24(2):97–109. doi: 10.1385/IR:24:2:097. [DOI] [PubMed] [Google Scholar]
- 8.Phan Tri Giang, Paus Didrik, Chan Tyani D., Turner Marian L., Nutt Stephen L., Basten Antony, Brink Robert. High affinity germinal center B cells are actively selected into the plasma cell compartment. The Journal of Experimental Medicine. 2006;203(11):2419–2424. doi: 10.1084/jem.20061254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kurosaki Tomohiro, Kometani Kohei, Ise Wataru. Memory B cells. Nature Reviews Immunology. 2015;15(3):149–159. doi: 10.1038/nri3802. [DOI] [PubMed] [Google Scholar]
- 10.Shaffer A.L., Lin Kuo-I, Kuo Tracy C., Yu Xin, Hurt Elaine M., Rosenwald Andreas, Giltnane Jena M., Yang Liming, Zhao Hong, Calame Kathryn, Staudt Louis M. Blimp-1 Orchestrates Plasma Cell Differentiation by Extinguishing the Mature B Cell Gene Expression Program. Immunity. 2002;17(1):51–62. doi: 10.1016/S1074-7613(02)00335-7. [DOI] [PubMed] [Google Scholar]
- 11.Shapiro-Shelef Miriam, Lin Kuo-I, McHeyzer-Williams Louise J, Liao Jerry, McHeyzer-Williams Michael G, Calame Kathryn. Blimp-1 Is Required for the Formation of Immunoglobulin Secreting Plasma Cells and Pre-Plasma Memory B Cells. Immunity. 2003;19(4):607–620. doi: 10.1016/S1074-7613(03)00267-X. [DOI] [PubMed] [Google Scholar]
- 12.Győry Ildikó, Wu Jian, Fejér György, Seto Edward, Wright Kenneth L. PRDI-BF1 recruits the histone H3 methyltransferase G9a in transcriptional silencing. Nature Immunology. 2004;5(3):299–308. doi: 10.1038/ni1046. [DOI] [PubMed] [Google Scholar]
- 13.Lin K.-I, Angelin-Duclos C., Kuo T. C., Calame K. Blimp-1-Dependent Repression of Pax-5 Is Required for Differentiation of B Cells to Immunoglobulin M-Secreting Plasma Cells. Molecular and Cellular Biology. 2002;22(13):4771–4780. doi: 10.1128/MCB.22.13.4771-4780.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Vasanwala Farha H., Kusam Saritha, Toney Lisa M., Dent Alexander L. Repression of AP-1 Function: A Mechanism for the Regulation of Blimp-1 Expression and B Lymphocyte Differentiation by the B Cell Lymphoma-6 Protooncogene. The Journal of Immunology. 2002;169(4):1922–1929. doi: 10.4049/jimmunol.169.4.1922. [DOI] [PubMed] [Google Scholar]
- 15.Minnich Martina, Tagoh Hiromi, Bönelt Peter, Axelsson Elin, Fischer Maria, Cebolla Beatriz, Tarakhovsky Alexander, Nutt Stephen L, Jaritz Markus, Busslinger Meinrad. Multifunctional role of the transcription factor Blimp-1 in coordinating plasma cell differentiation. Nature Immunology. 2016;17(3):331–343. doi: 10.1038/ni.3349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tellier Julie, Shi Wei, Minnich Martina, Liao Yang, Crawford Simon, Smyth Gordon K, Kallies Axel, Busslinger Meinrad, Nutt Stephen L. Blimp-1 controls plasma cell function through the regulation of immunoglobulin secretion and the unfolded protein response. Nature Immunology. 2016;17(3):323–330. doi: 10.1038/ni.3348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Cobaleda César, Schebesta Alexandra, Delogu Alessio, Busslinger Meinrad. Pax5: the guardian of B cell identity and function. Nature Immunology. 2007;8(5):463–470. doi: 10.1038/ni1454. [DOI] [PubMed] [Google Scholar]
- 18.Horcher Markus, Souabni Abdallah, Busslinger Meinrad. Pax5/BSAP Maintains the Identity of B Cells in Late B Lymphopoiesis. Immunity. 2001;14(6):779–790. doi: 10.1016/S1074-7613(01)00153-4. [DOI] [PubMed] [Google Scholar]
- 19.Dent A. L. Control of Inflammation, Cytokine Expression, and Germinal Center Formation by BCL-6. Science. 1997;276(5312):589–592. doi: 10.1126/science.276.5312.589. [DOI] [PubMed] [Google Scholar]
- 20.Cattoretti G, Chang CC, Cechova K, Zhang J, Ye BH, Falini B, et al. BCL-6 protein is expressed in germinal-center B cells. Blood. 1995;86(1):45–53. [PubMed]
- 21.Linterman Michelle A., Beaton Laura, Yu Di, Ramiscal Roybel R., Srivastava Monika, Hogan Jennifer J., Verma Naresh K., Smyth Mark J., Rigby Robert J., Vinuesa Carola G. IL-21 acts directly on B cells to regulate Bcl-6 expression and germinal center responses. The Journal of Experimental Medicine. 2010;207(2):353–363. doi: 10.1084/jem.20091738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Muto Akihiko, Tashiro Satoshi, Nakajima Osamu, Hoshino Hideto, Takahashi Satoru, Sakoda Eiichirou, Ikebe Dai, Yamamoto Masayuki, Igarashi Kazuhiko. The transcriptional programme of antibody class switching involves the repressor Bach2. Nature. 2004;429(6991):566–571. doi: 10.1038/nature02596. [DOI] [PubMed] [Google Scholar]
- 23.Tunyaplin Chainarong, Shaffer A. L., Angelin-Duclos Cristina D., Yu Xin, Staudt Louis M., Calame Kathryn L. Direct Repression of prdm1 by Bcl-6 Inhibits Plasmacytic Differentiation. The Journal of Immunology. 2004;173(2):1158–1165. doi: 10.4049/jimmunol.173.2.1158. [DOI] [PubMed] [Google Scholar]
- 24.Ochiai Kyoko, Katoh Yasutake, Ikura Tsuyoshi, Hoshikawa Yutaka, Noda Tetsuo, Karasuyama Hajime, Tashiro Satoshi, Muto Akihiko, Igarashi Kazuhiko. Plasmacytic Transcription Factor Blimp-1 Is Repressed by Bach2 in B Cells. Journal of Biological Chemistry. 2006;281(50):38226–38234. doi: 10.1074/jbc.M607592200. [DOI] [PubMed] [Google Scholar]
- 25.Angelin-Duclos Cristina, Cattoretti Giorgio, Lin Kuo-I, Calame Kathryn. Commitment of B Lymphocytes to a Plasma Cell Fate Is Associated with Blimp-1 Expression In Vivo. The Journal of Immunology. 2000;165(10):5462–5471. doi: 10.4049/jimmunol.165.10.5462. [DOI] [PubMed] [Google Scholar]
- 26.Falini B, Fizzotti M, Pucciarini A, Bigerna B, Marafioti T, Gambacorta M, et al. A monoclonal antibody (MUM1p) detects expression of the MUM1/IRF4 protein in a subset of germinal center B cells, plasma cells, and activated T cells. Blood. 2000;95(6):2084–92. [PubMed]
- 27.Mittrucker H.-W. Requirement for the Transcription Factor LSIRF/IRF4 for Mature B and T Lymphocyte Function. Science. 1997;275(5299):540–543. doi: 10.1126/science.275.5299.540. [DOI] [PubMed] [Google Scholar]
- 28.Reimold Andreas M., Iwakoshi Neal N., Manis John, Vallabhajosyula Prashanth, Szomolanyi-Tsuda Eva, Gravallese Ellen M., Friend Daniel, Grusby Michael J., Alt Frederick, Glimcher Laurie H. Plasma cell differentiation requires the transcription factor XBP-1. Nature. 2001;412(6844):300–307. doi: 10.1038/35085509. [DOI] [PubMed] [Google Scholar]
- 29.Shaffer A.L, Shapiro-Shelef Miriam, Iwakoshi Neal N, Lee Ann-Hwee, Qian Shu-Bing, Zhao Hong, Yu Xin, Yang Liming, Tan Bruce K, Rosenwald Andreas, Hurt Elaine M, Petroulakis Emmanuel, Sonenberg Nahum, Yewdell Jonathan W, Calame Kathryn, Glimcher Laurie H, Staudt Louis M. XBP1, Downstream of Blimp-1, Expands the Secretory Apparatus and Other Organelles, and Increases Protein Synthesis in Plasma Cell Differentiation. Immunity. 2004;21(1):81–93. doi: 10.1016/j.immuni.2004.06.010. [DOI] [PubMed] [Google Scholar]
- 30.Huang Yong, Shen Xing Jia, Zou Quan, Zhao Qiao Ling. Biological functions of MicroRNAs. Russian Journal of Bioorganic Chemistry. 2010;36(6):684–689. doi: 10.1134/S1068162010060026. [DOI] [PubMed] [Google Scholar]
- 31.Selbach Matthias, Schwanhäusser Björn, Thierfelder Nadine, Fang Zhuo, Khanin Raya, Rajewsky Nikolaus. Widespread changes in protein synthesis induced by microRNAs. Nature. 2008;455(7209):58–63. doi: 10.1038/nature07228. [DOI] [PubMed] [Google Scholar]
- 32.Zhao G, Yu D, Weiss MJ. MicroRNAs in erythropoiesis. Curr Opin Hematol. 2010;17(3):155–62. [DOI] [PubMed]
- 33.Hogg Daniel R., Harries Lorna W. Human genetic variation and its effect on miRNA biogenesis, activity and function. Biochemical Society Transactions. 2014;42(4):1184–1189. doi: 10.1042/BST20140055. [DOI] [PubMed] [Google Scholar]
- 34.Georgantas R. W., Hildreth R., Morisot S., Alder J., Liu C.-g., Heimfeld S., Calin G. A., Croce C. M., Civin C. I. CD34+ hematopoietic stem-progenitor cell microRNA expression and function: A circuit diagram of differentiation control. Proceedings of the National Academy of Sciences. 2007;104(8):2750–2755. doi: 10.1073/pnas.0610983104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Guo S., Lu J., Schlanger R., Zhang H., Wang J. Y., Fox M. C., Purton L. E., Fleming H. H., Cobb B., Merkenschlager M., Golub T. R., Scadden D. T. MicroRNA miR-125a controls hematopoietic stem cell number. Proceedings of the National Academy of Sciences. 2010;107(32):14229–14234. doi: 10.1073/pnas.0913574107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Basso Katia, Sumazin Pavel, Morozov Pavel, Schneider Christof, Maute Roy L., Kitagawa Yukiko, Mandelbaum Jonathan, Haddad Joseph, Chen Chang-Zheng, Califano Andrea, Dalla-Favera Riccardo. Identification of the Human Mature B Cell miRNome. Immunity. 2009;30(5):744–752. doi: 10.1016/j.immuni.2009.03.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhang Jenny, Jima Dereje D., Jacobs Cassandra, Fischer Randy, Gottwein Eva, Huang Grace, Lugar Patricia L., Lagoo Anand S., Rizzieri David A., Friedman Daphne R., Weinberg J. Brice, Lipsky Peter E., Dave Sandeep S. Patterns of microRNA expression characterize stages of human B-cell differentiation. Blood. 2009;113(19):4586–4594. doi: 10.1182/blood-2008-09-178186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Malpeli G, Barbi S, Zupo S, Tosadori G, Scardoni G, Bertolaso A, et al. Identification of microRNAs implicated in the late differentiation stages of normal B cells suggests a central role for miRNA targets ZEB1 and TP53. Oncotarget. 2017;8(7):11809–26. [DOI] [PMC free article] [PubMed]
- 39.Tijsterman Marcel, Ketting René F., Plasterk Ronald H. A. The Genetics of RNA Silencing. Annual Review of Genetics. 2002;36(1):489–519. doi: 10.1146/annurev.genet.36.043002.091619. [DOI] [PubMed] [Google Scholar]
- 40.Koralov Sergei B., Muljo Stefan A., Galler Gunther R., Krek Azra, Chakraborty Tirtha, Kanellopoulou Chryssa, Jensen Kari, Cobb Bradley S., Merkenschlager Matthias, Rajewsky Nikolaus, Rajewsky Klaus. Dicer Ablation Affects Antibody Diversity and Cell Survival in the B Lymphocyte Lineage. Cell. 2008;132(5):860–874. doi: 10.1016/j.cell.2008.02.020. [DOI] [PubMed] [Google Scholar]
- 41.Tsai DY, Hung KH, Lin IY, Su ST, Wu SY, Chung CH, et al. Uncovering MicroRNA regulatory hubs that modulate plasma cell differentiation. Sci Rep. 2015;5:17957. [DOI] [PMC free article] [PubMed]
- 42.Chen C.-Z. MicroRNAs Modulate Hematopoietic Lineage Differentiation. Science. 2004;303(5654):83–86. doi: 10.1126/science.1091903. [DOI] [PubMed] [Google Scholar]
- 43.Jensen Kristin, Brusletto Berit Sletbakk, Aass Hans Christian Dalsbotten, Olstad Ole K., Kierulf Peter, Gautvik Kaare M. Transcriptional Profiling of mRNAs and microRNAs in Human Bone Marrow Precursor B Cells Identifies Subset- and Age-Specific Variations. PLoS ONE. 2013;8(7):e70721. doi: 10.1371/journal.pone.0070721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Vasilatou D, Papageorgiou S, Pappa V, Papageorgiou E, Dervenoulas J. The role of microRNAs in normal and malignant hematopoiesis. Eur J Haematol. 2010;84(1):1–16. doi: 10.1111/j.1600-0609.2009.01348.x. [DOI] [PubMed] [Google Scholar]
- 45.Zhou B, Wang S, Mayr C, Bartel DP, Lodish HF. miR-150, a microRNA expressed in mature B and T cells, blocks early B cell development when expressed prematurely. Proc Natl Acad Sci U S A. 2007;104(17):7080–7085. doi: 10.1073/pnas.0702409104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Xiao Changchun, Calado Dinis Pedro, Galler Gunther, Thai To-Ha, Patterson Heide Christine, Wang Jing, Rajewsky Nikolaus, Bender Timothy P., Rajewsky Klaus. MiR-150 Controls B Cell Differentiation by Targeting the Transcription Factor c-Myb. Cell. 2007;131(1):146–159. doi: 10.1016/j.cell.2007.07.021. [DOI] [PubMed] [Google Scholar]
- 47.Fahl SP, Crittenden RB, Allman D, Bender TP. C-Myb is required for pro-B cell differentiation. J Immunol. 2009;183(9):5582–5592. doi: 10.4049/jimmunol.0901187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Rao DS, O'Connell RM, Chaudhuri AA, Garcia-Flores Y, Geiger TL, Baltimore D. MicroRNA-34a perturbs B lymphocyte development by repressing the forkhead box transcription factor Foxp1. Immunity. 2010;33(1):48–59. doi: 10.1016/j.immuni.2010.06.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Craig V. J., Cogliatti S. B., Imig J., Renner C., Neuenschwander S., Rehrauer H., Schlapbach R., Dirnhofer S., Tzankov A., Muller A. Myc-mediated repression of microRNA-34a promotes high-grade transformation of B-cell lymphoma by dysregulation of FoxP1. Blood. 2011;117(23):6227–6236. doi: 10.1182/blood-2010-10-312231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.O'Connell RM, Chaudhuri AA, Rao DS, Gibson WS, Balazs AB, Baltimore D. MicroRNAs enriched in hematopoietic stem cells differentially regulate long-term hematopoietic output. Proc Natl Acad Sci U S A. 2010;107(32):14235–14240. doi: 10.1073/pnas.1009798107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Rodriguez A., Vigorito E., Clare S., Warren M. V., Couttet P., Soond D. R., van Dongen S., Grocock R. J., Das P. P., Miska E. A., Vetrie D., Okkenhaug K., Enright A. J., Dougan G., Turner M., Bradley A. Requirement of bic/microRNA-155 for Normal Immune Function. Science. 2007;316(5824):608–611. doi: 10.1126/science.1139253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Thai T.-H., Calado D. P., Casola S., Ansel K. M., Xiao C., Xue Y., Murphy A., Frendewey D., Valenzuela D., Kutok J. L., Schmidt-Supprian M., Rajewsky N., Yancopoulos G., Rao A., Rajewsky K. Regulation of the Germinal Center Response by MicroRNA-155. Science. 2007;316(5824):604–608. doi: 10.1126/science.1141229. [DOI] [PubMed] [Google Scholar]
- 53.Vigorito Elena, Perks Kerry L., Abreu-Goodger Cei, Bunting Sam, Xiang Zou, Kohlhaas Susan, Das Partha P., Miska Eric A., Rodriguez Antony, Bradley Allan, Smith Kenneth G.C., Rada Cristina, Enright Anton J., Toellner Kai-Michael, MacLennan Ian C.M., Turner Martin. microRNA-155 Regulates the Generation of Immunoglobulin Class-Switched Plasma Cells. Immunity. 2007;27(6):847–859. doi: 10.1016/j.immuni.2007.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Timmer AM, Nizet V. IKKbeta/NF-kappaB and the miscreant macrophage. J Exp Med. 2008;205(6):1255–1259. doi: 10.1084/jem.20081056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Malumbres R., Sarosiek K. A., Cubedo E., Ruiz J. W., Jiang X., Gascoyne R. D., Tibshirani R., Lossos I. S. Differentiation stage-specific expression of microRNAs in B lymphocytes and diffuse large B-cell lymphomas. Blood. 2009;113(16):3754–3764. doi: 10.1182/blood-2008-10-184077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Xu S, Ou X, Huo J, Lim K, Huang Y, Chee S, et al. Mir-17-92 regulates bone marrow homing of plasma cells and production of immunoglobulin G2c. Nat Commun. 2015;6:6764. [DOI] [PubMed]
- 57.Lu Jun, Getz Gad, Miska Eric A., Alvarez-Saavedra Ezequiel, Lamb Justin, Peck David, Sweet-Cordero Alejandro, Ebert Benjamin L., Mak Raymond H., Ferrando Adolfo A., Downing James R., Jacks Tyler, Horvitz H. Robert, Golub Todd R. MicroRNA expression profiles classify human cancers. Nature. 2005;435(7043):834–838. doi: 10.1038/nature03702. [DOI] [PubMed] [Google Scholar]
- 58.Thomson JM, Newman M, Parker JS, Morin-Kensicki EM, Wright T, Hammond SM. Extensive post-transcriptional regulation of microRNAs and its implications for cancer. Genes Dev. 2006;20(16):2202–2207. doi: 10.1101/gad.1444406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Costinean S., Zanesi N., Pekarsky Y., Tili E., Volinia S., Heerema N., Croce C. M. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in E -miR155 transgenic mice. Proceedings of the National Academy of Sciences. 2006;103(18):7024–7029. doi: 10.1073/pnas.0602266103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Pedersen Irene M., Otero Dennis, Kao Elaine, Miletic Ana V., Hother Christoffer, Ralfkiaer Elisabeth, Rickert Robert C., Gronbaek Kirsten, David Michael. Onco‐miR‐155 targets SHIP1 to promote TNFα‐dependent growth of B cell lymphomas. EMBO Molecular Medicine. 2009;1(5):288–295. doi: 10.1002/emmm.200900028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Huang Xin, Shen Yulei, Liu Miao, Bi Chengfeng, Jiang Chunsun, Iqbal Javeed, McKeithan Timothy W., Chan Wing C., Ding Shi-Jian, Fu Kai. Quantitative Proteomics Reveals that miR-155 Regulates the PI3K-AKT Pathway in Diffuse Large B-Cell Lymphoma. The American Journal of Pathology. 2012;181(1):26–33. doi: 10.1016/j.ajpath.2012.03.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.He Lin, He Xingyue, Lim Lee P., de Stanchina Elisa, Xuan Zhenyu, Liang Yu, Xue Wen, Zender Lars, Magnus Jill, Ridzon Dana, Jackson Aimee L., Linsley Peter S., Chen Caifu, Lowe Scott W., Cleary Michele A., Hannon Gregory J. A microRNA component of the p53 tumour suppressor network. Nature. 2007;447(7148):1130–1134. doi: 10.1038/nature05939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Yamakuchi M, Lowenstein CJ. MiR-34, SIRT1 and p53: the feedback loop. Cell Cycle. 2009;8(5):712–715. doi: 10.4161/cc.8.5.7753. [DOI] [PubMed] [Google Scholar]
- 64.Craig V J, Tzankov A, Flori M, Schmid C A, Bader A G, Müller A. Systemic microRNA-34a delivery induces apoptosis and abrogates growth of diffuse large B-cell lymphoma in vivo. Leukemia. 2012;26(11):2421–2424. doi: 10.1038/leu.2012.110. [DOI] [PubMed] [Google Scholar]
- 65.Huang Y, Zou Y, Lin L, Ma X, Zheng R. miR101 regulates the cell proliferation and apoptosis in diffuse large Bcell lymphoma by targeting MEK1 via regulation of the ERK/MAPK signaling pathway. Oncol Rep. 2019;41(1):377–386. doi: 10.3892/or.2018.6821. [DOI] [PubMed] [Google Scholar]
- 66.Navarro A., Gaya A., Martinez A., Urbano-Ispizua A., Pons A., Balague O., Gel B., Abrisqueta P., Lopez-Guillermo A., Artells R., Montserrat E., Monzo M. MicroRNA expression profiling in classic Hodgkin lymphoma. Blood. 2007;111(5):2825–2832. doi: 10.1182/blood-2007-06-096784. [DOI] [PubMed] [Google Scholar]
- 67.Lawrie Charles H., Chi Jianxiang, Taylor Stephen, Tramonti Daniela, Ballabio Erica, Palazzo Stefano, Saunders Nigel J., Pezzella Francesco, Boultwood Jacqueline, Wainscoat James S., Hatton Christian S. R. Expression of microRNAs in diffuse large B cell lymphoma is associated with immunophenotype, survival and transformation from follicular lymphoma. Journal of Cellular and Molecular Medicine. 2009;13(7):1248–1260. doi: 10.1111/j.1582-4934.2008.00628.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Iqbal J., Shen Y., Liu Y., Fu K., Jaffe E. S., Liu C., Liu Z., Lachel C. M., Deffenbacher K., Greiner T. C., Vose J. M., Bhagavathi S., Staudt L. M., Rimsza L., Rosenwald A., Ott G., Delabie J., Campo E., Braziel R. M., Cook J. R., Tubbs R. R., Gascoyne R. D., Armitage J. O., Weisenburger D. D., McKeithan T. W., Chan W. C. Genome-wide miRNA profiling of mantle cell lymphoma reveals a distinct subgroup with poor prognosis. Blood. 2012;119(21):4939–4948. doi: 10.1182/blood-2011-07-370122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Pasqualucci L, Dalla-Favera R. The genetic landscape of diffuse large B-cell lymphoma. Semin Hematol. 2015;52(2):67–76. doi: 10.1053/j.seminhematol.2015.01.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Boyle EM, Davies FE, Leleu X, Morgan GJ. Understanding the multiple biological aspects leading to myeloma. Haematologica. 2014;99(4):605–612. doi: 10.3324/haematol.2013.097907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Vicente-Dueñas Carolina, Romero-Camarero Isabel, González-Herrero Inés, Alonso-Escudero Esther, Abollo-Jiménez Fernando, Jiang Xiaoyu, Gutierrez Norma C, Orfao Alberto, Marín Nieves, Villar Luisa María, Criado Ma Carmen Fernández, Pintado Belén, Flores Teresa, Alonso-López Diego, De Las Rivas Javier, Jiménez Rafael, Criado Francisco Javier García, Cenador María Begoña García, Lossos Izidore S, Cobaleda César, Sánchez-García Isidro. A novel molecular mechanism involved in multiple myeloma development revealed by targeting MafB to haematopoietic progenitors. The EMBO Journal. 2012;31(18):3704–3717. doi: 10.1038/emboj.2012.227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Hideshima Teru, Cottini Francesca, Nozawa Yoshihisa, Seo Hyuk-Soo, Ohguchi Hiroto, Samur Mehmet K., Cirstea Diana, Mimura Naoya, Iwasawa Yoshikazu, Richardson Paul G., Munshi Nikhil C., Chauhan Dharminder, Massefski Walter, Utsugi Teruhiro, Dhe-Paganon Sirano, Anderson Kenneth C. p53-related protein kinase confers poor prognosis and represents a novel therapeutic target in multiple myeloma. Blood. 2017;129(10):1308–1319. doi: 10.1182/blood-2016-09-738500. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Herrero Ana, Rojas Elizabeta, Misiewicz-Krzeminska Irena, Krzeminski Patryk, Gutiérrez Norma. Molecular Mechanisms of p53 Deregulation in Cancer: An Overview in Multiple Myeloma. International Journal of Molecular Sciences. 2016;17(12):2003. doi: 10.3390/ijms17122003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Shaham L, Binder V, Gefen N, Borkhardt A, Izraeli S. MiR-125 in normal and malignant hematopoiesis. Leukemia. 2012;26(9):2011–2018. doi: 10.1038/leu.2012.90. [DOI] [PubMed] [Google Scholar]
- 75.Murray MY, Rushworth SA, Zaitseva L, Bowles KM, Macewan DJ. Attenuation of dexamethasone-induced cell death in multiple myeloma is mediated by miR-125b expression. Cell Cycle. 2013;12(13):2144–2153. doi: 10.4161/cc.25251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Zhou Ming, Liu Zixing, Zhao Yuhua, Ding Yan, Liu Hao, Xi Yaguang, Xiong Wei, Li Guiyuan, Lu Jianrong, Fodstad Oystein, Riker Adam I., Tan Ming. MicroRNA-125b Confers the Resistance of Breast Cancer Cells to Paclitaxel through Suppression of Pro-apoptotic Bcl-2 Antagonist Killer 1 (Bak1) Expression. Journal of Biological Chemistry. 2010;285(28):21496–21507. doi: 10.1074/jbc.M109.083337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Abdi J, Rastgoo N, Li L, Chen W, Chang H. Role of tumor suppressor p53 and micro-RNA interplay in multiple myeloma pathogenesis. J Hematol Oncol. 2017;10(1):169. doi: 10.1186/s13045-017-0538-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Yuan Jie, Su Zhangyao, Gu Wenchao, Shen Xianjuan, Zhao Qiumin, Shi Linying, Jin Chunjing, Wang Xudong, Cong Hui, Ju Shaoqing. MiR-19b and miR-20a suppress apoptosis, promote proliferation and induce tumorigenicity of multiple myeloma cells by targeting PTEN. Cancer Biomarkers. 2019;24(3):279–289. doi: 10.3233/CBM-182182. [DOI] [PubMed] [Google Scholar]
- 79.Huang G, Liu X, Zhao X, Zhao J, Hao J, Ren J, et al. MiR-9 promotes multiple myeloma progression by regulating TRIM56/NF-kappaB pathway. Cell Biol Int. 2019. 10.1002/cbin.11104. [DOI] [PubMed]
- 80.Rothfield N. Clinical aspects and treatment of systemic lupus erythematosus. Curr Opin Rheumatol. 1989;1(3):327–331. doi: 10.1097/00002281-198901030-00014. [DOI] [PubMed] [Google Scholar]
- 81.Kil LP, Hendriks RW. Aberrant B cell selection and activation in systemic lupus erythematosus. Int Rev Immunol. 2013;32(4):445–470. doi: 10.3109/08830185.2013.786712. [DOI] [PubMed] [Google Scholar]
- 82.Dai Y., Huang Y.-S., Tang M., Lv T.-Y., Hu C.-X., Tan Y.-H., Xu Z.-M., Yin Y.-B. Microarray analysis of microRNA expression in peripheral blood cells of systemic lupus erythematosus patients. Lupus. 2007;16(12):939–946. doi: 10.1177/0961203307084158. [DOI] [PubMed] [Google Scholar]
- 83.Wu X.-n., Ye Y.-x., Niu J.-w., Li Y., Li X., You X., Chen H., Zhao L.-d., Zeng X.-f., Zhang F.-c., Tang F.-l., He W., Cao X.-t., Zhang X., Lipsky P. E. Defective PTEN regulation contributes to B cell hyperresponsiveness in systemic lupus erythematosus. Science Translational Medicine. 2014;6(246):246ra99–246ra99. doi: 10.1126/scitranslmed.3009131. [DOI] [PubMed] [Google Scholar]
- 84.Sarhan RA, Aboelenein HR, Sourour SK, Fawzy IO, Salah S, Abdelaziz AI. Targeting E2F1 and c-Myc expression by microRNA-17-5p represses interferon-stimulated gene MxA in peripheral blood mononuclear cells of pediatric systemic lupus erythematosus patients. Discov Med. 2015;19(107):419–425. [PubMed] [Google Scholar]
- 85.Zhang HD, Huang XX, Ye LL, Guo GQ, Li X, Chen CS, et al. B cell-related circulating MicroRNAs with the potential value of biomarkers in the differential diagnosis, and Distinguishment between the disease activity and lupus nephritis for systemic lupus erythematosus. Front Immunol. 2018;9:1473. [DOI] [PMC free article] [PubMed]
- 86.Zeng Li, Wu Jia-li, Liu Li-min, Jiang Ju-qing, Wu Hai-jing, Zhao Ming, Lu Qian-jin. Serum miRNA-371b-5p and miRNA-5100 act as biomarkers for systemic lupus erythematosus. Clinical Immunology. 2018;196:103–109. doi: 10.1016/j.clim.2018.10.004. [DOI] [PubMed] [Google Scholar]
- 87.Iorio MV, Croce CM. MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol Med. 2012;4(3):143–159. doi: 10.1002/emmm.201100209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Li Z, Rana TM. Therapeutic targeting of microRNAs: current status and future challenges. Nat Rev Drug Discov. 2014;13(8):622–638. doi: 10.1038/nrd4359. [DOI] [PubMed] [Google Scholar]
- 89.Babar I. A., Cheng C. J., Booth C. J., Liang X., Weidhaas J. B., Saltzman W. M., Slack F. J. Nanoparticle-based therapy in an in vivo microRNA-155 (miR-155)-dependent mouse model of lymphoma. Proceedings of the National Academy of Sciences. 2012;109(26):E1695–E1704. doi: 10.1073/pnas.1201516109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Misso Gabriella, Di Martino Maria Teresa, De Rosa Giuseppe, Farooqi Ammad Ahmad, Lombardi Angela, Campani Virginia, Zarone Mayra Rachele, Gullà Annamaria, Tagliaferri Pierosandro, Tassone Pierfrancesco, Caraglia Michele. Mir-34: A New Weapon Against Cancer? Molecular Therapy - Nucleic Acids. 2014;3:e195. doi: 10.1038/mtna.2014.47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Trang Phong, Wiggins Jason F, Daige Christopher L, Cho Chris, Omotola Michael, Brown David, Weidhaas Joanne B, Bader Andreas G, Slack Frank J. Systemic Delivery of Tumor Suppressor microRNA Mimics Using a Neutral Lipid Emulsion Inhibits Lung Tumors in Mice. Molecular Therapy. 2011;19(6):1116–1122. doi: 10.1038/mt.2011.48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Wiggins J. F., Ruffino L., Kelnar K., Omotola M., Patrawala L., Brown D., Bader A. G. Development of a Lung Cancer Therapeutic Based on the Tumor Suppressor MicroRNA-34. Cancer Research. 2010;70(14):5923–5930. doi: 10.1158/0008-5472.CAN-10-0655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Liu Can, Kelnar Kevin, Liu Bigang, Chen Xin, Calhoun-Davis Tammy, Li Hangwen, Patrawala Lubna, Yan Hong, Jeter Collene, Honorio Sofia, Wiggins Jason F, Bader Andreas G, Fagin Randy, Brown David, Tang Dean G. The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nature Medicine. 2011;17(2):211–215. doi: 10.1038/nm.2284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Chakraborty C, Sharma AR, Sharma G, Doss CGP, Lee SS. Therapeutic miRNA and siRNA: moving from bench to clinic as next generation medicine. Mol Ther-Nucl Acids. 2017;8:132–143. doi: 10.1016/j.omtn.2017.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Geary RS, Watanabe TA, Truong L, Freier S, Lesnik EA, Sioufi NB, et al. Pharmacokinetic properties of 2′-O-(2-methoxyethyl)-modified oligonucleotide analogs in rats. J Pharmacol Exp Ther. 2001;296(3):890–7. [PubMed]
- 96.Broderick JA, Zamore PD. MicroRNA therapeutics. Gene Ther. 2011;18(12):1104–1110. doi: 10.1038/gt.2011.50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Jiang Y, Dominguez PM, Melnick AM. The many layers of epigenetic dysfunction in B-cell lymphomas. Curr Opin Hematol. 2016;23(4):377–384. doi: 10.1097/MOH.0000000000000249. [DOI] [PubMed] [Google Scholar]
- 98.Shaknovich R., Cerchietti L., Tsikitas L., Kormaksson M., De S., Figueroa M. E., Ballon G., Yang S. N., Weinhold N., Reimers M., Clozel T., Luttrop K., Ekstrom T. J., Frank J., Vasanthakumar A., Godley L. A., Michor F., Elemento O., Melnick A. DNA methyltransferase 1 and DNA methylation patterning contribute to germinal center B-cell differentiation. Blood. 2011;118(13):3559–3569. doi: 10.1182/blood-2011-06-357996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Hung KH, Woo YH, Lin IY, Liu CH, Wang LC, Chen HY, et al. The KDM4A/KDM4C/NF-kappaB and WDR5 epigenetic cascade regulates the activation of B cells. Nucleic Acids Res. 2018;46(11):5547–60. [DOI] [PMC free article] [PubMed]
- 100.Azagra Alba, Román-González Lidia, Collazo Olga, Rodríguez-Ubreva Javier, de Yébenes Virginia G., Barneda-Zahonero Bruna, Rodríguez Jairo, Castro de Moura Manuel, Grego-Bessa Joaquim, Fernández-Duran Irene, Islam Abul B.M.M.K., Esteller Manel, Ramiro Almudena R., Ballestar Esteban, Parra Maribel. In vivo conditional deletion of HDAC7 reveals its requirement to establish proper B lymphocyte identity and development. The Journal of Experimental Medicine. 2016;213(12):2591–2601. doi: 10.1084/jem.20150821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Kalushkova Antonia, Fryknäs Mårten, Lemaire Miguel, Fristedt Charlotte, Agarwal Prasoon, Eriksson Maria, Deleu Sarah, Atadja Peter, Österborg Anders, Nilsson Kenneth, Vanderkerken Karin, Öberg Fredrik, Jernberg-Wiklund Helena. Polycomb Target Genes Are Silenced in Multiple Myeloma. PLoS ONE. 2010;5(7):e11483. doi: 10.1371/journal.pone.0011483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Raaphorst Frank M., van Kemenade Folkert J., Fieret Elly, Hamer Karien M., Satijn David P. E., Otte Arie P., Meijer Chris J. L. M. Cutting Edge: Polycomb Gene Expression Patterns Reflect Distinct B Cell Differentiation Stages in Human Germinal Centers. The Journal of Immunology. 2000;164(1):1–4. doi: 10.4049/jimmunol.164.1.1. [DOI] [PubMed] [Google Scholar]
- 103.Caganova Marieta, Carrisi Chiara, Varano Gabriele, Mainoldi Federica, Zanardi Federica, Germain Pierre-Luc, George Laura, Alberghini Federica, Ferrarini Luca, Talukder Asoke K., Ponzoni Maurilio, Testa Giuseppe, Nojima Takuya, Doglioni Claudio, Kitamura Daisuke, Toellner Kai-M., Su I-hsin, Casola Stefano. Germinal center dysregulation by histone methyltransferase EZH2 promotes lymphomagenesis. Journal of Clinical Investigation. 2013;123(12):5009–5022. doi: 10.1172/JCI70626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Chen H. T. Response to RAG-Mediated V(D)J Cleavage by NBS1 and gamma-H2AX. Science. 2000;290(5498):1962–1964. doi: 10.1126/science.290.5498.1962. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Petersen Simone, Casellas Rafael, Reina-San-Martin Bernardo, Chen Hua Tang, Difilippantonio Michael J., Wilson Patrick C., Hanitsch Leif, Celeste Arkady, Muramatsu Masamichi, Pilch Duane R., Redon Christophe, Ried Thomas, Bonner William M., Honjo Tasuku, Nussenzweig Michel C., Nussenzweig André. AID is required to initiate Nbs1/γ-H2AX focus formation and mutations at sites of class switching. Nature. 2001;414(6864):660–665. doi: 10.1038/414660a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Odegard VH, Kim ST, Anderson SM, Shlomchik MJ, Schatz DG. Histone modifications associated with somatic hypermutation. Immunity. 2005;23(1):101–110. doi: 10.1016/j.immuni.2005.05.007. [DOI] [PubMed] [Google Scholar]
- 107.Fujimura S, Matsui T, Kuwahara K, Maeda K, Sakaguchi N. Germinal center B-cell-associated DNA hypomethylation at transcriptional regions of the AID gene. Mol Immunol. 2008;45(6):1712–1719. doi: 10.1016/j.molimm.2007.09.023. [DOI] [PubMed] [Google Scholar]
- 108.Crouch Elizabeth E., Li Zhiyu, Takizawa Makiko, Fichtner-Feigl Stefan, Gourzi Polyxeni, Montaño Carolina, Feigenbaum Lionel, Wilson Patrick, Janz Siegfried, Papavasiliou F. Nina, Casellas Rafael. Regulation of AID expression in the immune response. The Journal of Experimental Medicine. 2007;204(5):1145–1156. doi: 10.1084/jem.20061952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Begum NA, Stanlie A, Nakata M, Akiyama H, Honjo T. The histone chaperone Spt6 is required for activation-induced cytidine deaminase target determination through H3K4me3 regulation. J Biol Chem. 2012;287(39):32415–32429. doi: 10.1074/jbc.M112.351569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Borchert GM, Holton NW, Edwards KA, Vogel LA, Larson ED. Histone H2A and H2B are monoubiquitinated at AID-targeted loci. PLoS One. 2010;5(7):e11641. doi: 10.1371/journal.pone.0011641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Baxter Jonathan, Sauer Stephan, Peters Antoine, John Rosalind, Williams Ruth, Caparros Marie-Laure, Arney Katharine, Otte Arie, Jenuwein Thomas, Merkenschlager Matthias, Fisher Amanda G. Histone hypomethylation is an indicator of epigenetic plasticity in quiescent lymphocytes. The EMBO Journal. 2004;23(22):4462–4472. doi: 10.1038/sj.emboj.7600414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Ruthenburg Alexander J, Wang Wooikoon, Graybosch Daina M, Li Haitao, Allis C David, Patel Dinshaw J, Verdine Gregory L. Histone H3 recognition and presentation by the WDR5 module of the MLL1 complex. Nature Structural & Molecular Biology. 2006;13(8):704–712. doi: 10.1038/nsmb1119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Shi Wei, Liao Yang, Willis Simon N, Taubenheim Nadine, Inouye Michael, Tarlinton David M, Smyth Gordon K, Hodgkin Philip D, Nutt Stephen L, Corcoran Lynn M. Transcriptional profiling of mouse B cell terminal differentiation defines a signature for antibody-secreting plasma cells. Nature Immunology. 2015;16(6):663–673. doi: 10.1038/ni.3154. [DOI] [PubMed] [Google Scholar]
- 114.Ochiai K, Muto A, Tanaka H, Takahashi S, Igarashi K. Regulation of the plasma cell transcription factor Blimp-1 gene by Bach2 and Bcl6. Int Immunol. 2008;20(3):453–460. doi: 10.1093/intimm/dxn005. [DOI] [PubMed] [Google Scholar]
- 115.Lemercier C, Brocard MP, Puvion-Dutilleul F, Kao HY, Albagli O, Khochbin S. Class II histone deacetylases are directly recruited by BCL6 transcriptional repressor. J Biol Chem. 2002;277(24):22045–22052. doi: 10.1074/jbc.M201736200. [DOI] [PubMed] [Google Scholar]
- 116.Lee SC, Bottaro A, Insel RA. Activation of terminal B cell differentiation by inhibition of histone deacetylation. Mol Immunol. 2003;39(15):923–932. doi: 10.1016/S0161-5890(03)00029-4. [DOI] [PubMed] [Google Scholar]
- 117.Su ST, Ying HY, Chiu YK, Lin FR, Chen MY, Lin KI. Involvement of histone demethylase LSD1 in Blimp-1-mediated gene repression during plasma cell differentiation. Mol Cell Biol. 2009;29(6):1421–1431. doi: 10.1128/MCB.01158-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Yu J, Angelin-Duclos C, Greenwood J, Liao J, Calame K. Transcriptional repression by blimp-1 (PRDI-BF1) involves recruitment of histone deacetylase. Mol Cell Biol. 2000;20(7):2592–2603. doi: 10.1128/MCB.20.7.2592-2603.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Czermin B, Melfi R, McCabe D, Seitz V, Imhof A, Pirrotta V. Drosophila enhancer of Zeste/ESC complexes have a histone H3 methyltransferase activity that marks chromosomal Polycomb sites. Cell. 2002;111(2):185–196. doi: 10.1016/S0092-8674(02)00975-3. [DOI] [PubMed] [Google Scholar]
- 120.Béguelin Wendy, Popovic Relja, Teater Matt, Jiang Yanwen, Bunting Karen L., Rosen Monica, Shen Hao, Yang Shao Ning, Wang Ling, Ezponda Teresa, Martinez-Garcia Eva, Zhang Haikuo, Zheng Yupeng, Verma Sharad K., McCabe Michael T., Ott Heidi M., Van Aller Glenn S., Kruger Ryan G., Liu Yan, McHugh Charles F., Scott David W., Chung Young Rock, Kelleher Neil, Shaknovich Rita, Creasy Caretha L., Gascoyne Randy D., Wong Kwok-Kin, Cerchietti Leandro, Levine Ross L., Abdel-Wahab Omar, Licht Jonathan D., Elemento Olivier, Melnick Ari M. EZH2 Is Required for Germinal Center Formation and Somatic EZH2 Mutations Promote Lymphoid Transformation. Cancer Cell. 2013;23(5):677–692. doi: 10.1016/j.ccr.2013.04.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Morin RD, Johnson NA, Severson TM, Mungall AJ, An J, Goya R, et al. Somatic mutations altering EZH2 (Tyr641) in follicular and diffuse large B-cell lymphomas of germinal-center origin. Nat Genet. 2010;42(2):181–5. [DOI] [PMC free article] [PubMed]
- 122.Shilatifard A. The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu Rev Biochem. 2012;81:65–95. doi: 10.1146/annurev-biochem-051710-134100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Ortega-Molina Ana, Boss Isaac W, Canela Andres, Pan Heng, Jiang Yanwen, Zhao Chunying, Jiang Man, Hu Deqing, Agirre Xabier, Niesvizky Itamar, Lee Ji-Eun, Chen Hua-Tang, Ennishi Daisuke, Scott David W, Mottok Anja, Hother Christoffer, Liu Shichong, Cao Xing-Jun, Tam Wayne, Shaknovich Rita, Garcia Benjamin A, Gascoyne Randy D, Ge Kai, Shilatifard Ali, Elemento Olivier, Nussenzweig Andre, Melnick Ari M, Wendel Hans-Guido. The histone lysine methyltransferase KMT2D sustains a gene expression program that represses B cell lymphoma development. Nature Medicine. 2015;21(10):1199–1208. doi: 10.1038/nm.3943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Zhang Jiyuan, Dominguez-Sola David, Hussein Shafinaz, Lee Ji-Eun, Holmes Antony B, Bansal Mukesh, Vlasevska Sofija, Mo Tongwei, Tang Hongyan, Basso Katia, Ge Kai, Dalla-Favera Riccardo, Pasqualucci Laura. Disruption of KMT2D perturbs germinal center B cell development and promotes lymphomagenesis. Nature Medicine. 2015;21(10):1190–1198. doi: 10.1038/nm.3940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Wang Zhibin, Zang Chongzhi, Rosenfeld Jeffrey A, Schones Dustin E, Barski Artem, Cuddapah Suresh, Cui Kairong, Roh Tae-Young, Peng Weiqun, Zhang Michael Q, Zhao Keji. Combinatorial patterns of histone acetylations and methylations in the human genome. Nature Genetics. 2008;40(7):897–903. doi: 10.1038/ng.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Lunning MA, Green MR. Mutation of chromatin modifiers; an emerging hallmark of germinal center B-cell lymphomas. Blood Cancer J. 2015;5:e361. doi: 10.1038/bcj.2015.89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Pasqualucci Laura, Dominguez-Sola David, Chiarenza Annalisa, Fabbri Giulia, Grunn Adina, Trifonov Vladimir, Kasper Lawryn H., Lerach Stephanie, Tang Hongyan, Ma Jing, Rossi Davide, Chadburn Amy, Murty Vundavalli V., Mullighan Charles G., Gaidano Gianluca, Rabadan Raul, Brindle Paul K., Dalla-Favera Riccardo. Inactivating mutations of acetyltransferase genes in B-cell lymphoma. Nature. 2011;471(7337):189–195. doi: 10.1038/nature09730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Zhang Jiyuan, Vlasevska Sofija, Wells Victoria A., Nataraj Sarah, Holmes Antony B., Duval Romain, Meyer Stefanie N., Mo Tongwei, Basso Katia, Brindle Paul K., Hussein Shafinaz, Dalla-Favera Riccardo, Pasqualucci Laura. The CREBBP Acetyltransferase Is a Haploinsufficient Tumor Suppressor in B-cell Lymphoma. Cancer Discovery. 2017;7(3):322–337. doi: 10.1158/2159-8290.CD-16-1417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Rollig C, Knop S, Bornhauser M. Multiple myeloma. Lancet. 2015;385(9983):2197–2208. doi: 10.1016/S0140-6736(14)60493-1. [DOI] [PubMed] [Google Scholar]
- 130.Avet-Loiseau Hervé, Leleu Xavier, Roussel Murielle, Moreau Philippe, Guerin-Charbonnel Catherine, Caillot Denis, Marit Gérald, Benboubker Lotfi, Voillat Laurent, Mathiot Claire, Kolb Brigitte, Macro Margaret, Campion Loïc, Wetterwald Marc, Stoppa Anne-Marie, Hulin Cyrille, Facon Thierry, Attal Michel, Minvielle Stéphane, Harousseau Jean-Luc. Bortezomib Plus Dexamethasone Induction Improves Outcome of Patients With t(4;14) Myeloma but Not Outcome of Patients With del(17p) Journal of Clinical Oncology. 2010;28(30):4630–4634. doi: 10.1200/JCO.2010.28.3945. [DOI] [PubMed] [Google Scholar]
- 131.Kuo Alex J., Cheung Peggie, Chen Kaifu, Zee Barry M., Kioi Mitomu, Lauring Josh, Xi Yuanxin, Park Ben Ho, Shi Xiaobing, Garcia Benjamin A., Li Wei, Gozani Or. NSD2 Links Dimethylation of Histone H3 at Lysine 36 to Oncogenic Programming. Molecular Cell. 2011;44(4):609–620. doi: 10.1016/j.molcel.2011.08.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Pei Huadong, Zhang Lindsey, Luo Kuntian, Qin Yuxin, Chesi Marta, Fei Frances, Bergsagel P. Leif, Wang Liewei, You Zhongsheng, Lou Zhenkun. MMSET regulates histone H4K20 methylation and 53BP1 accumulation at DNA damage sites. Nature. 2011;470(7332):124–128. doi: 10.1038/nature09658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Martinez-Garcia E., Popovic R., Min D.-J., Sweet S. M. M., Thomas P. M., Zamdborg L., Heffner A., Will C., Lamy L., Staudt L. M., Levens D. L., Kelleher N. L., Licht J. D. The MMSET histone methyl transferase switches global histone methylation and alters gene expression in t(4;14) multiple myeloma cells. Blood. 2010;117(1):211–220. doi: 10.1182/blood-2010-07-298349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Marango J., Shimoyama M., Nishio H., Meyer J. A., Min D.-J., Sirulnik A., Martinez-Martinez Y., Chesi M., Bergsagel P. L., Zhou M.-M., Waxman S., Leibovitch B. A., Walsh M. J., Licht J. D. The MMSET protein is a histone methyltransferase with characteristics of a transcriptional corepressor. Blood. 2008;111(6):3145–3154. doi: 10.1182/blood-2007-06-092122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Min D-J, Ezponda T, Kim M K, Will C M, Martinez-Garcia E, Popovic R, Basrur V, Elenitoba-Johnson K S, Licht J D. MMSET stimulates myeloma cell growth through microRNA-mediated modulation of c-MYC. Leukemia. 2012;27(3):686–694. doi: 10.1038/leu.2012.269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Pawlyn C, Bright M D, Buros A F, Stein C K, Walters Z, Aronson L I, Mirabella F, Jones J R, Kaiser M F, Walker B A, Jackson G H, Clarke P A, Bergsagel P L, Workman P, Chesi M, Morgan G J, Davies F E. Overexpression of EZH2 in multiple myeloma is associated with poor prognosis and dysregulation of cell cycle control. Blood Cancer Journal. 2017;7(3):e549–e549. doi: 10.1038/bcj.2017.27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Agarwal P, Alzrigat M, Parraga AA, Enroth S, Singh U, Ungerstedt J, et al. Genome-wide profiling of histone H3 lysine 27 and lysine 4 trimethylation in multiple myeloma reveals the importance of Polycomb gene targeting and highlights EZH2 as a potential therapeutic target. Oncotarget. 2016;7(6):6809–23. [DOI] [PMC free article] [PubMed]
- 138.Pawlyn C, Kaiser MF, Heuck C, Melchor L, Wardell CP, Murison A, et al. The Spectrum and clinical impact of epigenetic modifier mutations in myeloma. Clin Cancer Res. 2016;22(23):5783–94. [DOI] [PMC free article] [PubMed]
- 139.van Haaften G, Dalgliesh GL, Davies H, Chen L, Bignell G, Greenman C, et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nat Genet. 2009;41(5):521–3. [DOI] [PMC free article] [PubMed]
- 140.Ezponda Teresa, Dupéré-Richer Daphné, Will Christine M., Small Eliza C., Varghese Nobish, Patel Tej, Nabet Behnam, Popovic Relja, Oyer Jon, Bulic Marinka, Zheng Yupeng, Huang Xiaoxiao, Shah Mrinal Y., Maji Sayantan, Riva Alberto, Occhionorelli Manuela, Tonon Giovanni, Kelleher Neil, Keats Jonathan, Licht Jonathan D. UTX/KDM6A Loss Enhances the Malignant Phenotype of Multiple Myeloma and Sensitizes Cells to EZH2 inhibition. Cell Reports. 2017;21(3):628–640. doi: 10.1016/j.celrep.2017.09.078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Ohguchi H, Hideshima T, Bhasin MK, Gorgun GT, Santo L, Cea M, et al. The KDM3A-KLF2-IRF4 axis maintains myeloma cell survival. Nat Commun. 2016;7:10258. [DOI] [PMC free article] [PubMed]
- 142.Uemura M., Yamamoto H., Takemasa I., Mimori K., Hemmi H., Mizushima T., Ikeda M., Sekimoto M., Matsuura N., Doki Y., Mori M. Jumonji Domain Containing 1A Is a Novel Prognostic Marker for Colorectal Cancer: In vivo Identification from Hypoxic Tumor Cells. Clinical Cancer Research. 2010;16(18):4636–4646. doi: 10.1158/1078-0432.CCR-10-0407. [DOI] [PubMed] [Google Scholar]
- 143.Cho SF, Chang YC, Chang CS, Lin SF, Liu YC, Hsiao HH, et al. MALAT1 long non-coding RNA is overexpressed in multiple myeloma and may serve as a marker to predict disease progression. BMC Cancer. 2014;14:809. [DOI] [PMC free article] [PubMed]
- 144.Handa Hiroshi, Kuroda Yuko, Kimura Kei, Masuda Yuta, Hattori Hikaru, Alkebsi Lobna, Matsumoto Morio, Kasamatsu Tetsuhiro, Kobayashi Nobuhiko, Tahara Ken-ichi, Takizawa Makiko, Koiso Hiromi, Ishizaki Takuma, Shimizu Hiroaki, Yokohama Akihiko, Tsukamoto Norifumi, Saito Takayuki, Murakami Hirokazu. Long non-coding RNA MALAT1 is an inducible stress response gene associated with extramedullary spread and poor prognosis of multiple myeloma. British Journal of Haematology. 2017;179(3):449–460. doi: 10.1111/bjh.14882. [DOI] [PubMed] [Google Scholar]
- 145.Nawrocki MJ, Majewski D, Puszczewicz M, Jagodzinski PP. Decreased mRNA expression levels of DNA methyltransferases type 1 and 3A in systemic lupus erythematosus. Rheumatol Int. 2017;37(5):775–783. doi: 10.1007/s00296-017-3711-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Garaud Soizic, Le Dantec Christelle, Jousse-Joulin Sandrine, Hanrotel-Saliou Catherine, Saraux Alain, Mageed Rizgar A., Youinou Pierre, Renaudineau Yves. IL-6 Modulates CD5 Expression in B Cells from Patients with Lupus by Regulating DNA Methylation. The Journal of Immunology. 2009;182(9):5623–5632. doi: 10.4049/jimmunol.0802412. [DOI] [PubMed] [Google Scholar]
- 147.Hippen KL, Tze LE, Behrens TW. CD5 maintains tolerance in anergic B cells. J Exp Med. 2000;191(5):883–890. doi: 10.1084/jem.191.5.883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Nakkuntod J, Avihingsanon Y, Mutirangura A, Hirankarn N. Hypomethylation of LINE-1 but not Alu in lymphocyte subsets of systemic lupus erythematosus patients. Clin Chim Acta. 2011;412(15–16):1457–1461. doi: 10.1016/j.cca.2011.04.002. [DOI] [PubMed] [Google Scholar]
- 149.Fali Tinhinane, Le Dantec Christelle, Thabet Yosra, Jousse Sandrine, Hanrotel Catherine, Youinou Pierre, Brooks Wesley H., Perl Andras, Renaudineau Yves. DNA methylation modulates HRES1/p28 expression in B cells from patients with Lupus. Autoimmunity. 2013;47(4):265–271. doi: 10.3109/08916934.2013.826207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Silverman Lewis R., Demakos Erin P., Peterson Bercedis L., Kornblith Alice B., Holland Jimmie C., Odchimar-Reissig Rosalie, Stone Richard M., Nelson Douglas, Powell Bayard L., DeCastro Carlos M., Ellerton John, Larson Richard A., Schiffer Charles A., Holland James F. Randomized Controlled Trial of Azacitidine in Patients With the Myelodysplastic Syndrome: A Study of the Cancer and Leukemia Group B. Journal of Clinical Oncology. 2002;20(10):2429–2440. doi: 10.1200/JCO.2002.04.117. [DOI] [PubMed] [Google Scholar]
- 151.Kantarjian Hagop, Issa Jean-Pierre J., Rosenfeld Craig S., Bennett John M., Albitar Maher, DiPersio John, Klimek Virginia, Slack James, de Castro Carlos, Ravandi Farhad, Helmer Richard, Shen Lanlan, Nimer Stephen D., Leavitt Richard, Raza Azra, Saba Hussain. Decitabine improves patient outcomes in myelodysplastic syndromes. Cancer. 2006;106(8):1794–1803. doi: 10.1002/cncr.21792. [DOI] [PubMed] [Google Scholar]
- 152.Kiziltepe T., Hideshima T., Catley L., Raje N., Yasui H., Shiraishi N., Okawa Y., Ikeda H., Vallet S., Pozzi S., Ishitsuka K., Ocio E. M., Chauhan D., Anderson K. C. 5-Azacytidine, a DNA methyltransferase inhibitor, induces ATR-mediated DNA double-strand break responses, apoptosis, and synergistic cytotoxicity with doxorubicin and bortezomib against multiple myeloma cells. Molecular Cancer Therapeutics. 2007;6(6):1718–1727. doi: 10.1158/1535-7163.MCT-07-0010. [DOI] [PubMed] [Google Scholar]
- 153.Cao Yang, Qiu Guo-Qiang, Wu Hao-Qing, Wang Zhi-Lin, Lin Yan, Wu Wei, Xie Xiao-Bao, Gu Wei-Ying. Decitabine enhances bortezomib treatment in RPMI 8226 multiple myeloma cells. Molecular Medicine Reports. 2016;14(4):3469–3475. doi: 10.3892/mmr.2016.5658. [DOI] [PubMed] [Google Scholar]
- 154.New M, Olzscha H, La Thangue NB. HDAC inhibitor-based therapies: can we interpret the code? Mol Oncol. 2012;6(6):637–656. doi: 10.1016/j.molonc.2012.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155.Seidel C, Schnekenburger M, Dicato M, Diederich M. Histone deacetylase 6 in health and disease. Epigenomics. 2015;7(1):103–118. doi: 10.2217/epi.14.69. [DOI] [PubMed] [Google Scholar]
- 156.Hideshima Teru, Qi Jun, Paranal Ronald M., Tang Weiping, Greenberg Edward, West Nathan, Colling Meaghan E., Estiu Guillermina, Mazitschek Ralph, Perry Jennifer A., Ohguchi Hiroto, Cottini Francesca, Mimura Naoya, Görgün Güllü, Tai Yu-Tzu, Richardson Paul G., Carrasco Ruben D., Wiest Olaf, Schreiber Stuart L., Anderson Kenneth C., Bradner James E. Discovery of selective small-molecule HDAC6 inhibitor for overcoming proteasome inhibitor resistance in multiple myeloma. Proceedings of the National Academy of Sciences. 2016;113(46):13162–13167. doi: 10.1073/pnas.1608067113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157.Santo L., Hideshima T., Kung A. L., Tseng J.-C., Tamang D., Yang M., Jarpe M., van Duzer J. H., Mazitschek R., Ogier W. C., Cirstea D., Rodig S., Eda H., Scullen T., Canavese M., Bradner J., Anderson K. C., Jones S. S., Raje N. Preclinical activity, pharmacodynamic, and pharmacokinetic properties of a selective HDAC6 inhibitor, ACY-1215, in combination with bortezomib in multiple myeloma. Blood. 2012;119(11):2579–2589. doi: 10.1182/blood-2011-10-387365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158.Galli Monica, Salmoiraghi Silvia, Golay Joseé, Gozzini Antonella, Crippa Claudia, Pescosta Norbert, Rambaldi Alessandro. A phase II multiple dose clinical trial of histone deacetylase inhibitor ITF2357 in patients with relapsed or progressive multiple myeloma. Annals of Hematology. 2009;89(2):185–190. doi: 10.1007/s00277-009-0793-8. [DOI] [PubMed] [Google Scholar]
- 159.Richardson PG, Hungria VT, Yoon SS, Beksac M, Dimopoulos MA, Elghandour A, et al. Panobinostat plus bortezomib and dexamethasone in previously treated multiple myeloma: outcomes by prior treatment. Blood. 2016;127(6):713–21. [DOI] [PMC free article] [PubMed]
- 160.Dupere-Richer D, Licht JD. Epigenetic regulatory mutations and epigenetic therapy for multiple myeloma. Curr Opin Hematol. 2017;24(4):336–344. doi: 10.1097/MOH.0000000000000358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161.Chaidos A., Caputo V., Gouvedenou K., Liu B., Marigo I., Chaudhry M. S., Rotolo A., Tough D. F., Smithers N. N., Bassil A. K., Chapman T. D., Harker N. R., Barbash O., Tummino P., Al-Mahdi N., Haynes A. C., Cutler L., Le B., Rahemtulla A., Roberts I., Kleijnen M., Witherington J. J., Parr N. J., Prinjha R. K., Karadimitris A. Potent antimyeloma activity of the novel bromodomain inhibitors I-BET151 and I-BET762. Blood. 2013;123(5):697–705. doi: 10.1182/blood-2013-01-478420. [DOI] [PubMed] [Google Scholar]
- 162.Siu K T, Ramachandran J, Yee A J, Eda H, Santo L, Panaroni C, Mertz J A, Sims III R J, Cooper M R, Raje N. Preclinical activity of CPI-0610, a novel small-molecule bromodomain and extra-terminal protein inhibitor in the therapy of multiple myeloma. Leukemia. 2016;31(8):1760–1769. doi: 10.1038/leu.2016.355. [DOI] [PubMed] [Google Scholar]
- 163.Amorim Sandy, Stathis Anastasios, Gleeson Mary, Iyengar Sunil, Magarotto Valeria, Leleu Xavier, Morschhauser Franck, Karlin Lionel, Broussais Florence, Rezai Keyvan, Herait Patrice, Kahatt Carmen, Lokiec François, Salles Gilles, Facon Thierry, Palumbo Antonio, Cunningham David, Zucca Emanuele, Thieblemont Catherine. Bromodomain inhibitor OTX015 in patients with lymphoma or multiple myeloma: a dose-escalation, open-label, pharmacokinetic, phase 1 study. The Lancet Haematology. 2016;3(4):e196–e204. doi: 10.1016/S2352-3026(16)00021-1. [DOI] [PubMed] [Google Scholar]
- 164.Alzrigat Mohammad, Jernberg-Wiklund Helena, Licht Jonathan. Targeting EZH2 in Multiple Myeloma—Multifaceted Anti-Tumor Activity. Epigenomes. 2018;2(3):16. doi: 10.3390/epigenomes2030016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165.Zeng D, Liu M, Pan J. Blocking EZH2 methylation transferase activity by GSK126 decreases stem cell-like myeloma cells. Oncotarget. 2017;8(2):3396–3411. doi: 10.18632/oncotarget.13773. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.