Abstract
Asthma is a highly heterogeneous disease, often manifesting with wheeze, dyspnea, chest tightness, and cough as prominent symptoms. The eliciting factors, natural history, underlying molecular biology, and clinical management of asthma highly vary among affected individuals. Because of this variation, many efforts have gone into subtyping asthma. Endotypes are subtypes of disease based on distinct pathophysiological mechanisms. Endotypes can be clinically useful because they organize mechanistic understanding of heterogeneous diseases and can direct treatment toward modalities that are likely to be the most effective. Asthma endotyping can be shaped by clinical features, laboratory parameters, and/or -omic approaches. We discuss the application of -omic approaches—including transcriptomics, epigenomics, microbiomics, metabolomics, and proteomics—to asthma endotyping. -Omic approaches have provided supporting evidence for many existing endotyping paradigms and also suggested novel ways to conceptualize asthma endotypes. While endotypes based on single -omic approaches are relatively common, their integrated multi-omic application to asthma endotyping has been more limited thus far. We discuss paths forward to integrate multi-omics with clinical features and laboratory parameters to achieve the goal of precise asthma endotypes.
Keywords: asthma, endotype, -ome, -omic, transcriptome, epigenome, microbiome, metabolome, proteome, multi-ome, phenotype, cluster, integrate
Introduction
Asthma is a heterogeneous disease with varied contributing mechanisms and diverse clinical presentations. Despite variations in underlying molecular biology and phenotypic manifestation, there is often a convergent suite of symptoms including wheeze, dyspnea, chest tightness, and/or cough.
Many efforts have gone into subtyping asthma. Phenotypes are classifications based on common sets of observable characteristics that are often a downstream result of genetics and the environment. Disease severity, response to treatment, and exacerbating factors have long been used to delineate asthma phenotypes. Phenotypes may not reflect unified molecular and cellular mechanisms underlying disease. Endotypes, on the other hand, are subtypes of disease based on distinct pathophysiological mechanisms. Endotypes can be clinically useful because they help organize mechanistic understanding of heterogeneous diseases like asthma, directing treatment toward modalities that target specific pathways that may be disrupted within a given endotype. This is especially important in asthma, where patients respond to medications with varying efficacy. The growing availability of biologics that modulate particular pathways makes it increasingly important to accurately characterize asthma endotypes so that therapy can be appropriately prescribed to improve asthma control.1
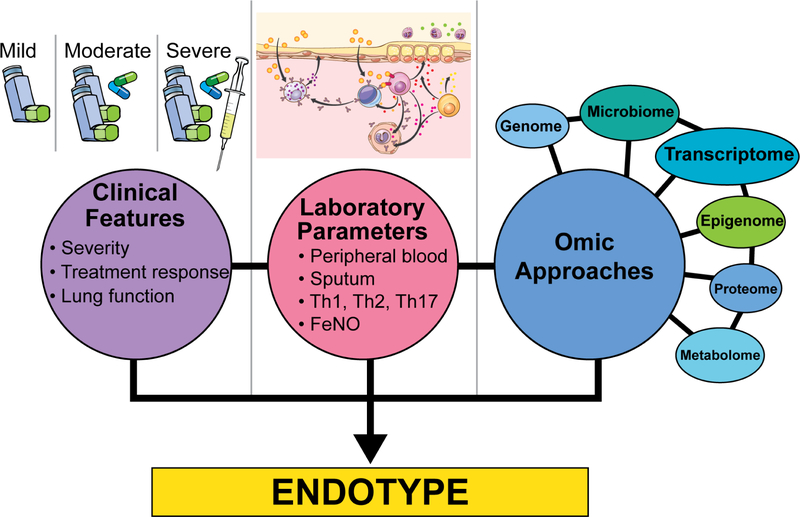
Approaches to asthma classification and endotyping can be organized into three broad categories including strategies based on clinical features, laboratory parameters, and molecular profiling by -omic approaches (Figure 1). While clinical features and common laboratory parameters have long been used to classify asthma, more recent -omic approaches offer the potential to refine current paradigms and identify novel endotypes. Integration of these approaches is emerging as a way to coalesce strategies toward more meaningful asthma endotypes. Here we first briefly describe classification of asthma based on clinical features and laboratory parameters and then focus on how -omic approaches have contributed to endotyping thus far. We then discuss emerging efforts toward integrating -omics and incorporating them with clinical features and laboratory parameters to better identify asthma endotypes.
Figure 1: Approaches to endotyping asthma.
Endotyping, or the subtyping of asthma based on pathophysiological mechanisms, can be informed by clinical features, laboratory parameters, and -omic approaches. Clinical features often serve as the basis for phenotyping asthma. Asthma endotyping based on laboratory parameters is common, spanning the well-recognized T2-high and T2-low endotypes based on induced sputum cells and/or fractional exhaled nitric oxide (FeNO), as well as eosinophilic, neutrophilic, mixed-granulocytic, and pauci-granulocytic endotypes based on sputum, BAL, and/or blood inflammatory cells. The application of -omic approaches to asthma endotyping has provided supporting evidence for many existing endotyping paradigms but also suggested novel ways to conceptualize asthma endotypes. While -omic approaches to asthma endotyping have been largely based on transcriptomic data thus far, there is emerging evidence that other -omic modalities can inform endotyping. The successful integration of clinical features, laboratory parameters, and -omic approaches will advance asthma endotyping.
Classification of asthma based on clinical features
Asthma is often classified based on clinical characteristics such as severity of disease, response to treatment, exacerbating factors, age of onset, and comorbid conditions (Figure 2). This classification into clinical phenotypes provides a first step to addressing heterogeneity in asthma. For example, the Severe Asthma Research Program (SARP), a multicenter network studying asthma in adults and children, used unsupervised cluster analysis to identify five strata of asthma based primarily on lung function and age of onset.2 However, the overlap between these phenotypically-defined strata and underlying pathophysiologic mechanisms (endotypes) was not clear. For example, although adults in SARP with combined elevation of sputum eosinophils and neutrophils had the most severe asthma, these subjects were spread among SARP phenotype clusters.2 Because the underlying molecular etiology of a disease is the mechanism by which phenotypes emerge, one might expect greater overlap between endotypes and phenotypes. This is often not the case, however, likely due to the multidimensional heterogeneity in clinical presentations as well as their underlying mechanisms.
Figure 2: Clinical features can be used to classify asthma.
Phenotypes of asthma are often based on clinical features, including severity of disease (e.g. mild intermittent, moderate persistent, severe persistent), response to treatment, and exacerbating factors.
Endotyping of asthma based on laboratory parameters
While clinical features can sometimes offer clues toward underlying mechanisms of asthma, the depth of mechanistic insight from them alone is limited. Laboratory parameters can provide deeper evidence for the mechanistic underpinnings of an endotype. Laboratory parameters used for asthma endotyping include Type 2 (T2)-related attributes, airway and peripheral blood inflammatory cell patterns, and T-helper cell subtype involvement.
T2-high vs. T2-low asthma represents one of the most recognized endotyping strategies for asthma.3 In T2-high asthma, genetic susceptibility and allergen exposure lead to skewing toward T-helper cell 2 (Th2)-cytokine related inflammation, which upregulates airway inflammation and airway hyperresponsiveness causing asthma (Figure 3). Several laboratory parameters can provide evidence for the T2-high vs. T2-low endotype, including peripheral blood eosinophilia, BAL eosinophilia, sputum eosinophilia, serum total and allergen-specific IgE levels, and fractional exhaled nitric oxide (FeNO).4, 5 While these measures are often associated, a subject with T2 asthma may not show high levels in all of these dimensions. Similarly, there is a growing appreciation for the role that innate lymphoid cells (ILCs) play in T2 asthma. In particular, ILC2 cells secrete a Th2-related milieu of cytokines including IL4, IL5, and IL13.6 The relative abundance of ILC2 cells is highly concordant with the relative abundance of sputum eosinophils – a hallmark of T2-high asthma.7
Figure 3: Asthma endotypes based on laboratory parameters.
Laboratory parameters, such as inflammatory cell fractions in sputum, BAL, and peripheral blood, and fractional exhaled nitric oxide (FeNO), have been used to define asthma endotypes. As an example, the T2-high endotype is shown here, reflecting a pathophysiologic mechanism driven by Th2 and ILC2 cells. In this scenario, Th2 and ILC2 cells produce IL4, IL5, and IL13, leading to goblet cell hyperplasia, increased mucin production, local and systemic eosinophilia, and B-cell production of IgE. IgE coats mast cells that are then primed to release histamine upon IgE cross-linking induced by environmental allergens, leading to the bronchospasm and airway inflammation that characterizes T2-high asthma. Signs of this pathophysiologic mechanism can be detected by measuring inflammatory cell patterns in sputum, bronchoalveolar lavage fluid, and peripheral blood, FeNO, serum total and specific IgE levels, and cytokine levels in sputum and peripheral blood. Such laboratory parameters are often used to define asthma endotypes.
The cellular pattern of inflammation can provide an alternate asthma endotyping strategy beyond the binary T2-high vs. T2-low endotypes.8 Bronchoalveolar lavage (BAL) fluid, induced sputum, and peripheral blood can show varied cellular compositions in patients with asthma. The cellular pattern of these samples can be eosinophilic, neutrophilic, mixed eosinophilic/neutrophilic, or demonstrating few cells of either class (i.e. paucigranulocytic9). Although there is no consensus on specific definitions for asthma endotypes based on inflammatory cell patterns, one schema classifies the eosinophilic endotype based on elevated sputum (>3%) and/or blood (≥400 cells/ul) eosinophils on at least two occasions and positive response to treatment strategies that suppress eosinophils.8 This eosinophilic endotype highly overlaps with T2-high and allergic asthma. The neutrophilic endotype is based on elevated sputum neutrophils (≥64%) but not eosinophils (<3%) with an increased total cell count (≥9.7 million cells/g) on at least two occasions with poor response to treatments that suppress eosinophils.8 Neutrophilic asthma tends to be more severe and refractory to corticosteroids than non-neutrophilic asthma.10 The mixed eosinophilic/neutrophilic endotype is defined by evidence for both neutrophils and eosinophils, independently or concurrently, on at least two occasions.8Paucigranulocytic asthma is an endotype of exclusion when neither elevated sputum eosinophils (<3%) or neutrophils (<64%) were observed and treatment targeting eosinophils and neutrophils were ineffective.8
While the T2-high vs. T2-low distinction represents a classical approach for asthma endotyping, classification based on other T helper cell classes (e.g. Th17 and Th1) has more recently come to the fore.11, 12 Th17 cells are a subset of T-cells that are often involved in humoral response to extracellular pathogens such as bacteria and fungi; these cells can also recruit neutrophils to the site of inflammation.13 There is evidence that IL17F, a key cytokine produced by Th17 cells, has a direct involvement in the development of asthma. Mutations in the 1L17F gene increase the probability of developing asthma, while overexpression of IL17F results in neutrophil recruitment to the lung, amplifying allergic pulmonary inflammation in an animal model.14 The Th17-related, neutrophilic asthma endotype may arise from treatment of T2 asthma with corticosteroids or anti-IL13 that in turn induces a Th17 profile marked by corticosteroid resistance.15 The most notable clinical differences observed between T2 and Th17 asthma were higher FeNO and sputum eosinophils in T2 compared to Th17 asthma. Diagnostically, it has been suggested that a select set of marker genes can define the T2 vs. Th17 asthma endotypes, including POSTN, CLCA1, SERPINB2 for T2 asthma,4 and CXCL1, CXCL2, CXCL3, IL8, CSF3 for Th17 asthma.15 In addition, high relative protein expression of CECAM6 in bronchial epithelium and neutrophils is associated with Th17 vs. T2 asthma.16
Another T-helper cell type, Th1 cells, may also play a role in asthma pathobiology. Typically, Th1 immunity is associated with intracellular pathogen response. The most notable cytokine produced by Th1 cells is interferon gamma, and CCR5 is a surface marker of Th1 cells.17 A Th1 asthma endotype has recently been proposed based on levels of CD4+/CCR5+ T-cells in the lower airways of children with severe asthma.17 Th1 cells as well as high neutrophils were found in the BAL of both allergic and non-allergic children with asthma, suggesting that this feature is associated with severity independent of allergen sensitivity.17 Additionally, CD8+ cytotoxic T-cells were elevated in the BAL of subjects with the highest levels of interferon gamma.17 Because this cohort was comprised exclusively of severe asthmatics, follow up studies quantifying the absolute number of Th1 cells across a wider range of asthma severity are needed. In a separate study of SARP subjects not stratified by interferon gamma levels, CD8+ T-cell levels were not significantly different across healthy controls, non-severe, and severe asthma.18 However, differences associated with Th1 cells were still observed across severity.18 This study found that BAL from subjects with severe asthma demonstrated a higher ratio of Th1-enriched T-cells (CD4+/CCR6−/CCR4−) to NK cells (CD3−NKp46+ with lymphoid morphology), compared to non-severe asthma subjects or healthy controls (>14% vs. 4–14%).18 This ratiometric measure also suggested the potential involvement of Th1 cells in severe asthma with concordant NK depletion. Given Th1 cells mediate immune response to viruses, this may fit with clinical observations that many asthma exacerbations are triggered by viral infection.19
Whether asthma endotypes are defined based on laboratory-based quantification of airway and peripheral blood inflammatory cell patterns, or T helper cell subtype involvement, it is clear that the characteristics of such endotypes overlap to some degree (Figure 4). Whether endotypes are stable over time is also not known. In theory, for example, a T2-high subject could develop characteristics of a Th1-high endotype during a viral infection, then return to a T2-high-only endotype following resolution of the infection. Longitudinal studies of deeply phenotyped subjects can help determine if these or other related scenarios elicit endotype changes over time.
Figure 4: Asthma endotypes based on T-helper cell subtype involvement.
Endotypes of asthma based on the relative signal strengths of Th2, Th17, Th1, and T cells overall have been described and associated with eosinophil, neutrophil, serum total and specific IgE levels as well as with variable treatment response. These T-helper cell-based endotypes likely have some overlap between one another.
Endotyping of asthma based on -omics
Studies employing -omic approaches can provide unbiased molecular evidence for refining existing endotypes and defining new ones. In this section we will review how -omic approaches have contributed to asthma endotyping (Figure 1). We begin by discussing transcriptomics, the -omic approach that has provided the largest contribution to asthma endotyping thus far.
Transcriptomics
Transcriptomics is the systematic, quantitative, and qualitative characterization of RNA transcripts across the genome.20 Because RNA reflects dynamic processes at play that underlie pathophysiology in a given tissue, profiling its expression via transcriptomics in disease-relevant tissue such as airway brushings, endobronchial biopsies, BAL, and peripheral blood can help with accurately distinguishing asthma status,21–23 classifying asthma phenotypes,24 and defining its endotypes.25–29
Transcriptional profiles for the T2-high vs. T2-low endotypes (Figure 3) were first identified in 2009 by Woodruff et al. through microarray analysis of airway brushings from 42 patients with mild-to-moderate asthma and 28 healthy controls.4 T2-high and T2-low profiles among the subjects with asthma were determined by relative expression of IL13-inducible genes.4 Compared to their T2-low counterparts, subjects demonstrating the T2-high gene expression signature had higher expression of IL5 and IL13 in bronchial biopsies, and significantly higher serum IgE levels, blood and airway eosinophilia, subepithelial fibrosis, and airway mucin gene expression.4 The lung function of subjects belonging to this T2-high endotype improved with inhaled corticosteroid treatment, while those belonging to the T2-low endotype did not. The fact that the efficacy of inhaled corticosteroids differed among endotypes suggests clinical utility in considering T2-high vs. -low endotype when selecting asthma therapy.4
More recent transcriptomic studies of asthma have supported T2-based endotypes,30 and also suggested other endotyping paradigms.25–29 Among 299 young adult participants of the Childhood Asthma Management Cohort, Howrylak et al. identified five longitudinally stable phenotypic clusters of asthma.25 Their examination of gene expression profiles from peripheral blood CD4+ cells revealed that each of these five phenotypic clusters was associated with a corresponding CD4+ gene expression pattern.25 Reorganization of these five clusters into two subgroups—more atopic vs. less atopic, akin to T2-high vs. T2-low—helped to identify a CD4+ gene signature based on five genes (SRM, HDAC2, SLC33A1, P2RY10, ADD3) that could classify atopic patients in an independent cohort with high accuracy.25 The expression levels of HDAC2, SLC33A1, P2RY10 and ADD3 were relatively upregulated in the more atopic subgroup compared to the less atopic subgroup, whereas SRM was relatively downregulated. Serum total IgE, serum eosinophil count, and methacholine PC20 were significantly different between the more atopic vs. less atopic subgroups defined by CD4+ gene signature.25 Further, serum IgE level was positively correlated with all 5 genes in the CD4+ gene signature, and serum eosinophil counts were positively correlated with SRM expression levels.25
Studies from the Unbiased BIO markers in PREDiction of respiratory disease outcomes (U-BIOPRED) cohort similarly suggest the possibility of new endotype clusters based on transcriptomics in addition to support for T2-based endotyping. Among 120 U-BIOPRED subjects, semi-supervised clustering of 508 genes differentially expressed in sputum between healthy volunteer, eosinophil- and non-eosinophil-predominant inflammation identified the expected three transcriptome clusters.26 While these three clusters did not cleanly separate paucigranulocytic, eosinophilic, mixed, and neutrophilic asthma phenotypes, there were differences in the granulocytic content across these clusters. The first cluster overlapped most with the T2-high endotype and was characterized by sputum gene expression of IL33R, CCR3, and TSLPR, and association with sputum eosinophilia, high FeNO, and severe asthma with oral corticosteroid dependency, frequent exacerbations, and severe airflow obstruction. The second and third transcriptome clusters exhibited T2-low traits. Cluster 2 was identified by expression of interferon, TNF, and inflammasome-related genes, and subjects in this cluster demonstrated high sputum neutrophilia, elevated serum CRP level, and atopic dermatitis. Cluster 3 was identified based on genes related to metabolic and mitochondrial pathways and included subjects with better lung function; subjects with paucigranulocytic inflammation mainly clustered to Cluster 3, but Cluster 3 also included subjects with eosinophilic, neutrophilic, and mixed inflammatory sputum composition.26 More recent U-BIOPRED cohort studies have explored sputum, nasal brushing, endobronchial brushing, and biopsy transcriptome data together.27, 28 Based on these data, a gene signature for adult-onset severe asthma was identified and found to be associated with eosinophilic airway inflammation, mast cells, and group 3 innate lymphoid cells, while a gene signature for childhood-onset severe asthma was found to be associated with induced lung injury.28 The gene set variation analysis in this study was based on a mix of data from in vivo and in vitro human sample studies as well as murine models; these inferences should therefore be validated directly in human subjects with asthma.
The concept of a “T2-ultrahigh” endotype was suggested by a study of sputum transcriptomics from 84 subjects with asthma and 27 healthy controls.29 Consistent with other studies, T2-high and T2-low clusters were identified, but the investigators also highlighted a subgroup of T2-high subjects with especially high T2 signal based on their analysis. Clinically, these subjects were older with more severe airflow obstruction.29
Transcriptomic studies have also enhanced our understanding of asthma endotypes defined by airway inflammatory cell patterns. Using microarray-based gene expression profiles of sputum from 47 asthmatic patients, Baines et al. identified a 6-gene (CLC, DNASE1L3, IL1B, alkaline phosphatase, ALPL, CXCR2) signature that could discriminate eosinophilic asthma from non-eosinophilic, paucigranulocytic, and neutrophilic asthma as well as controls.31 Targeted qRT-PCR measures of this 6-gene signature in sputum to predict future asthma exacerbation demonstrated that this signature performed better (i.e. had greater areas under the ROC curves) than predictions based on peripheral eosinophil counts, sputum neutrophil counts, or combined sputum eosinophil and neutrophil counts.32
Epigenomics
While transcriptomic approaches have contributed most to asthma endotyping thus far, epigenomic approaches may also have a role to play because they can characterize interactions between the environment and transcriptome.33 As asthma is a disease where environmental factors influence pathophysiology and natural history, leveraging epigenomics toward endotyping is likely to become increasingly important.
Epigenome-wide studies have identified peripheral blood methylation loci associated with serum IgE concentration in populations with asthma34, 35 Liang et al. found that 13% of variance in circulating IgE levels could be explained by the peripheral blood methylation status at three loci (SLC25A33, LPCAT2, and L2HGDH).34 Several of the identified IgE-associated methylation loci annotated to genes that encode proteins characteristic of eosinophils, including IL5RA, CCR3, IL1RL1, PGR2, PGR3, and GATA1.34 Given the central roles that IgE and eosinophils play in T2-high asthma (Figure 3), these epigenomic data could be used to support Th-related endotypes of asthma. In line with this, Yang et al. identified 81 loci with differential methylation in peripheral blood mononuclear cells (PBMCs) of subjects with atopic asthma (n=97) vs. healthy controls (n=97).36 Loci relevant to Th2 immunity tracked with IgE content and the expression of Th2-related genes including IL13, RUNX3, CAT, PRF1, and TIGIT.36
Epigenomic studies of tissue beyond blood also suggest that epigenomics can contribute to asthma endotyping. A DNA methylation study of airway epithelial cells used correlation-based network analysis to classify CpG sites differentially methylated between 74 asthmatics and 41 non-asthmatics into four modules.37 These four modules were associated with clinical features and laboratory parameters commonly used for asthma endotyping. Specifically, one module was associated with eosinophilia in BAL fluid, and another module was associated with FeNO; the remaining two modules were associated with the clinical feature of inhaled corticosteroid use.37
Microbiomics
Given recognized roles for viruses19 and bacteria38 in the development and course of asthma, it is logical to consider microbiome data when conceptualizing asthma endotypes. Upper and lower airway microbiota are known to vary by asthma status and disease activity,38–40 and there is some evidence from microbiome studies that there could be distinct microbiota associated with particular endotypes of asthma. Results to date have been heterogeneous in nature, however.
In a study of bronchial brushings from 40 subjects with severe asthma, no taxa were found to be associated with a T2-related epithelial gene signature.40 Proteobacteria abundance was found to be negatively correlated with biopsy eosinophil abundance while Proteobacteria abundance was positively associated with the expression of Th17 genes.40 This also further supports the negative correlation between T2 and Th17 asthma previously described by Choy et al.15
Separately, Sverrild et al. examined the relationship between bronchoalveolar lavage fluid inflammatory patterns and airway microbial diversity in a study of 23 steroid-free nonsmoking asthmatics and 10 healthy control subjects.41 They examined airway microbial composition differences in relation to eosinophil and neutrophil content.41 Specifically, airway microbiome composition among asthmatics with low BAL eosinophils significantly differed from both healthy controls and eosinophil high subjects. Unexpectedly however, there was no significant difference in composition when comparing asthmatic subjects with high BAL eosinophils to healthy subjects.41 Asthmatic subjects with low BAL eosinophils had high abundances of Neisseria, Bacteroides, and Rothia compared to their counterparts with high BAL eosinophils and healthy subjects.41 They also observed lower beta-diversity in subjects with higher levels of BAL neutrophils, suggesting lower bacterial diversity within subjects that have neutrophilic asthma.
More recently, the microbial composition and inflammatory patterns of induced sputum samples collected from 167 adults with severe asthma were studied.42 The investigators found that airway microbiota were less diverse and more dissimilar in subjects with neutrophilic vs. eosinophilic sputum inflammatory patterns. Diversity measures were significantly correlated with sputum neutrophil proportions but not eosinophil proportions; furthermore, sputum neutrophil proportion was the strongest predictor of microbiota composition.42 In subjects with neutrophilic asthma, there was reduced relative abundance of Streptococcus, Gemella, and Porphyromonas, while Haemophilus was negatively correlated with eosinophil percentage.42
The bronchial microbiota of adults with steroid-naive atopic asthma (n=42), atopy without asthma (n=21), and healthy controls (n=21) has also been studied within the AsthmaNet clinical research network.43 Microbial composition differed significantly between these groups. Asthmatics showed higher enrichment of several taxa including Haemophilus, Neisseria, Fusobacterium, and Porphyromonas, and the bronchial microbiome of asthmatics at baseline differed by responsiveness to treatment with inhaled corticosteroids.43 Additionally, subjects identified as T2-high based on the gene expression levels of 3 bronchial epithelial genes (CLCA1, SERPINB2, and POSTN) were found to have lower bronchial bacterial load.43
The findings from these and other studies suggest that airway microbiome composition can be associated with inflammatory cell phenotypes, with measures of microbial diversity and composition more frequently associated with neutrophilic asthma thus far. The mechanistic links between airway microbiome and asthma endotypes warrant further investigation.
Metabolomics
Metabolomics provides a snapshot of active physiology that has been used to classify individuals with asthma and asthma phenotypes.20, 44, 45 Metabolomic profiling involves the systematic assessment, identification, and quantification of low molecular weight compounds in biological samples, typically by liquid or gas chromatography mass spectrometry (LC-MS or GC-MS) or by nuclear magnetic resonance (NMR).20 Biospecimens for which metabolites are frequently measured in studies of asthma include plasma, serum, urine, and exhaled breath.20, 44
A study of exhaled breath samples from 35 patients with asthma and 23 controls showed that a model based on 15 volatile organic compounds in the exhaled breath samples could classify asthma as the eosinophilic (>2% sputum eosinophilia) endotype with AUC 0.98 or the neutrophilic ( ≥40% sputum neutrophilia) endotype with AUC 0.90.46 Separately, a study of exhaled breath and steroid responsiveness in 25 patients with mild/moderate asthma found that exhaled breath volatile organic compound levels could predict steroid responsiveness with greater accuracy than FeNO or sputum eosinophils.47 Using NMR spectroscopy and machine learning, another group of investigators found that NMR spectra of exhaled breath condensate could cluster 89 asthmatic subjects into three phenotypic groups, one of which was consistent with the neutrophilic endotype with low peripheral blood eosinophil and high neutrophils.48 More recently, cluster analysis based on metabolomic data from exhaled breath from 78 subjects identified three clusters of individuals with significant differences in peripheral blood eosinophil and neutrophil percentages, as well as differences in oral corticosteroid needs.49
Because of heterogeneity in findings and the relatively small sample sizes studied thus far,44 it is not yet clear the extent to which metabolomics will impact asthma endotyping. However, the fact that metabolomic profiling can be done on exhaled breath is an attractive characteristic, especially for endotyping asthma in children and individuals who may be averse to more invasive sampling.50
Proteomics
Proteomics can offer a descriptive window into the current inflammatory state of the airway. Within 240 U-BIOPRED cohort subjects, proteomic profiles were recently used to identify 10 clusters with distinct proteomic signatures that were dubbed “proteotypes.”51 Overlay of these proteotypes with sputum granulocyte counts showed strong associations. Specifically, proteotypes 1, 2, and 3 were associated with high sputum eosinophilia (mean counts 19%, 23%, and 17%) and were thought to represent a common, highly eosinophilic phenotype; transcriptomic study of sputum cells showed cytokine gene expression patterns consistent with eosinophilia.51 Proteotypes 5 and 6 had atopy as a common feature, and proteotypes 8, 9, and 10 were associated with high neutrophil counts (mean counts 64%, 82%, and 73%).51 Proteotypes 4 and 7 where less defined by granulocyte count and had the highest levels of smoking (current or former) with the least aberrant inflammatory cell proportions relative to healthy controls.51 There was low concordance between the eosinophil high proteotypes and atopy. The proteotypes with the highest sputum eosinophils showed high IL13 and periostin, yet low total serum IgE. The most atopic proteotypes however showed moderate levels of eosinophilia, IL13, and periostin, but high levels of total IgE.51 This suggests that differing underlying mechanisms may exist for some forms of T2 asthma. Further study of the identified proteotypes will be needed to assess relationship to treatment response, and validation of proteotypes such as these will be needed to assess reproducibility and temporality.
Multi-omic endotyping
-Omic approaches systematically capture a dimension of molecular processes underlying disease, whether at the level of genome, epigenome, transcriptome, proteome, metabolome, or microbiome. Although comprehensive at each level, each single -ome can be thought of as capturing just one of many dimensions underlying disease pathology. In some ways, this is akin to the parable of the blind men and the elephant, where none has encountered an elephant before and each must learn and conceptualize what the elephant is by touching a different part of the elephant’s body. Although each claims a truth based on their individual touch, none can accurately describe the actual elephant. Such an analogy may be relevant to asthma endotyping, where we may learn a part of “truth” by each -omic approach (and deeply so), but accurately defining robust asthma endotypes will require integration of multiple -omics.
To this end, there have been limited multi-omic studies relevant to asthma endotyping. In a small study of 20 subjects with asthma, McGeachie et al. integrated metabolomic data from plasma samples with genome-wide genotype, peripheral blood CD4+ T-cell methylation profiles, and peripheral blood CD4+ transcriptomes to generate a conditional Gaussian Bayesian network of asthma control.52 Analysis of this network implicated two pathways (arachidonic acid metabolism and linoleic acid metabolism) in asthma control and altered sphingolipid metabolism as an underlying feature of uncontrolled asthma and cellular response to albuterol.52
A U-BIOPRED study sought to assess the relationship between asthma clusters identified by clinical features and sputum transcriptomic and proteomic profiles.53 Among 266 subjects, cluster analysis based on eight clinical features (age of onset, pack years of smoking, BMI, FEV1 percent predicted, FEV1/FVC, ACQ-related score, number of exacerbations in the past year, and daily dose of oral prednisolone) identified four clusters of asthmatic patients.53 The investigators found significant differences in sputum proteomics and transcriptomics between these clusters, with higher sputum eosinophilia in cluster 1 compared to the other three clusters of subjects with severe asthma.53 Interesting, there were no differences in sputum neutrophil counts, FeNO, or serum IgE levels between the clusters.53 In a subsequent study of sputum transcriptomics and proteomics among U-BIOPRED subjects, investigators found that neutrophilic asthma was associated NLRP3 inflammasome expression, and IL1R1 gene expression was associated with eosinophilic severe asthma, leading them to suggest that Th2-driven eosinophilic inflammation and neutrophil-associated inflammasome activation are interacting pathways in severe asthma.54
Paving a path to multi-omics for asthma endotyping
Although there is a limited track record to date of integrating multiple -omics to define asthma endotypes, the raw materials and tools for doing so are growing. These methods help provide a path forward to maximize the utility of the high dimensional data sets that are being generated in asthma studies. Numerous asthma cohorts and asthma research networks have gathered and continue to build rich phenotypic and molecular data including -omic profiles of their participants. Strategies for integrating multi-omic data from cohorts are also becoming increasingly available, many of which have been developed in the cancer field where -omic data have been more widely available. In envisioning next steps for how multi-omics can play a more prominent role in asthma endotyping, we describe some important considerations and approaches for multi-omic analysis below.
The successful generation and analysis of meaningful multi-omic data hold the promise of revealing systems-level insights, but carry several challenges.20 As is true for all studies-- multi-omic, single-omic and non-omic alike-- careful study design is important to ensure the validity and generalizability of results generated. Temporality, potential confounders, missing data, and comparability across cohorts and populations must all be considered. For –omic and multi-omic studies in particular, we previously reviewed several current challenges of high-throughput approaches, including access to systems-wide profiling technologies, cost of system-wide profiling, data transfer and management, data quality and control, accurate modeling of biological systems through data integration, and acceptance by communities accustomed to reductionist approaches.20 Increasing accessibility to core facilities, high performance computing systems, software innovations, standardization of data and data sharing, and iterative development of tools and software platforms are some solutions to these challenges.20 Working toward these solutions will enable more multi-omic studies of asthma that can inform on endotypes and are reproducible across studies and populations.20
A common first step to explore data is cluster analysis, in which subjects are clustered into groups based on similarities of features of interest. Within a multi-omic study, cluster analysis could be used to identify subjects who are similar to one other within and across - omes. For example, within a cohort in which both bronchial transcriptome and bronchial microbiome were profiled, cluster analysis could reveal 4 clusters: a cluster of subjects with a T2-high bronchial epithelial transcriptomic signature and a bronchial microbiome that differs from healthy subjects, another cluster also with a T2-high transcriptomic signature but a “healthy” bronchial microbiome, and two additional clusters that each have a T2-low bronchial epithelial transcriptomic signature, yet show distinct microbiome signatures. While clustering single -omics can reveal patterns in the data, as several studies of asthma have already shown, clustering of multiple -omic profiles generated from the same subject can reveal findings not evident from a single data type.55
Multi-omic clustering methods can themselves be classified into three broad categories based on the stage at which each -ome is integrated for clustering.55 Late integration methods treat each -ome separately, and then integrate the clustering results of each individual -ome.55 Intermediate integration performs some form of -ome specific calculation, integrating these for clustering.55 Early integration methods often perform clustering only once on a combined multi- omic dataset.55
Late integration methods for multi-omic clustering involve first clustering on each -ome independently, allowing the user to specify various clustering approaches tailored to each -ome. The results of the per -ome clustering are then combined into a net cluster result.55 Methods for integrating these per -ome clustering results can vary however. The Cluster-Of-Clusters Assignments (COCA) algorithm concatenates binary vectors representing cluster membership for each -ome, then performs final clustering on this matrix.56 In contrast, the Perturbation clustering for data INtegration and disease Subytping (PINS) algorithm creates a connectivity matrix for each -ome in which each sample is compared to all other samples, creating a binary matrix with 1’s indicating that the samples were clustered together, and 0’s representing that samples were not clustered together in the given -ome.57 The connectivity matrices for all -omes are then averaged for the final connectivity matrix that is then used for clustering.57
Intermediate integration of -omic data involves processing each -ome independently, then integrating these processed representations for clustering.55 In some methods, within -ome similarity is calculated, then each is dimensionally reduced by matrix decomposition, taking the first k eigenvectors for concatenation across -omes and clustering; this can be seen as a form of combined spectral clustering.58, 59 Similarity network fusion (SNF) is a method that builds similarity networks within each -ome, then uses message passing for clustering; this ensures that any low level similarity between subjects that is conserved across -omes is taken into account as well as high similarity within a single -ome.60 This approach creates a weighted network graph from a k-nearest neighbor embedding of subject-subject similarities for each - ome. SNF then merges all of the network graphs from each -ome for a ‘fused’ final graph, which is then used for clustering.60 SNF has been used successfully to identify sub-strata of glioblastoma multiforme through integrating mRNA, methylation, and micro-RNA measures.60 The algorithm could meaningfully stratify subjects of several cancer types as measured by differences in survival across identified groups.60
Many early integration methods are performed on a matrix of data from each -ome concatenated together; this combined matrix is then deconvolved through some form of dimension reduction to orthogonal components in which each component captures a correlated suite of features, including those across -omes.61 While the high level representation can vary (i.e. correlation,62 Bayesian networks,63 decomposed matrix components64),55 the overarching theme between these methods is similar: find the correlated variables, and represent them as a lower dimensional ‘hub’, eigenvector, or correlated component upon which to perform clustering. Alternatively, the correlations between variables from all combined -omes can be used to build a network graph model of combined -omes, as in xMWAS.65 Many of these approaches will be effective when –omic data are all continuous (rather than categorical or ordinal) in nature. These approaches may not be as appropriate when trying to integrate categorical information such as allele status or ordinal variables such as copy number variation with continuous variables such as gene expression, microbial abundances, and metabolite levels. There therefore currently exists a need for methodologies that will, in an automated fashion, appropriately integrate these different data types for unsupervised clustering.
Conclusions
With asthma embodying such a heterogeneous disease, the quest to subtype it into endotypes based on cohesive, underlying pathophysiologic mechanisms will not only organize our thinking on this complex disease, but also help target treatment toward precision medicine.66 Clinical features, laboratory parameters, and –omic approaches all have a role to play in defining and refining asthma endotypes toward robust subtypes that can meaningfully contribute to our understanding and treatment of asthma (Figure 1). The transcriptomic identification of T2-high asthma4 and identification of other endotypes of asthma through –omic approaches as reviewed here reveal the potential for multi-omically defined endotypes to inform treatment approach and candidacy for targeted biologics67, 68 Further refining existing endotypes and discovering new endotypes through multi-omic methods can lead to improved asthma outcomes if they can inform treatment beyond the T2-centric paradigm.
As multi-omic studies continue to develop to inform asthma endotypes, it will be important to keep several considerations at the forefront. First, careful study design, well-characterized cohorts, appropriate address of confounding variables, and reproducibility across cohorts will be needed to ensure the robustness of the data generated and quality of results produced. Rigorous methodology and attention to data quality and control via standardization and sharing can advance this. Tissue-specificity and relevance of the profiles generated is also important to consider; thus far, -omic studies of peripheral blood in asthma have predominantly yielded findings related to atopy25, 34, 35 while studies of local samples (e.g. airway brushings and biopsies, BAL, and sputum) in asthma have enabled more refined endotyping.4, 17, 18, 26–28, 40–43, 51 Additionally, longitudinal studies can help to provide evidence for the relative stability of endotypes; this will help clinicians decide on the temporal appropriateness of endotyping efforts and endotype-tailored treatments.
While –omic approaches including transcriptomics, epigenomics, microbiomics, metabolomics, and proteomics have each been applied as modalities to support existing endotypes and to conceptualize new endotypes, their integrated multi-omic application has been more limited thus far. Although we have discussed paths forward for expanding multi-omic approaches for asthma endotyping, this will have to be combined with clinical features and laboratory parameters to achieve the goal of precision medicine for asthma .
Key Concepts and Therapeutic Implications.
While reversible airway obstruction is a defining feature of asthma, the eliciting factors, natural history, underlying molecular biology, and clinical management of asthma highly vary among affected individuals.
Endotypes are subtypes of disease based on pathophysiological mechanisms.
Endotypes help organize mechanistic understanding of heterogeneous diseases like asthma and can direct treatment toward modalities that are likely to be the most effective due to their targeting of specific pathophysiologic pathways.
Approaches to asthma classification and endotyping can be organized into three broad categories including strategies based on clinical features, laboratory parameters, and molecular profiling by -omic approaches.
-Omic approaches—including transcriptomics, epigenomics, microbiomics, metabolomics, and proteomics—have supported and refined current endotyping paradigms while also suggesting novel ways to conceptualize asthma endotypes.
The application of multi-omics, including their integration with clinical features and laboratory parameters, can help coalesce strategies toward robust asthma endotypes that will enable precision medicine.
Funding:
SB is funded by the National Institutes of Health (NIH) R01 AI118833 and NIH U19 AI136053
Abbreviations:
- ACQ
Asthma control questionnaire
- BAL
Bronchoalveolar lavage
- BMI
Body mass index
- COCA
Cluster-of-clusters assignments
- FeNO
Fractional exhaled nitric oxide
- FEV1
Forced expiratory volume in 1 second
- FVC
Forced vital capacity
- GC-MS
Gas chromatography mass spectrometry
- LC-MS
Liquid chromatography mass spectrometry
- NMR
Nuclear magnetic resonance
- PINS
Perturbation clustering for data integration and disease subtyping
- SARP
Severe Asthma Research Program
- SNF
Similarity network fusion
- T2
Type 2
- Th
T helper
- U-BIOPRED
Unbiased BIOmarkers in PREDiction of respiratory disease outcomes
MS 19–00215 – Clinical glossary
- Clustering
An approach to segregating samples of a dataset into discrete groups based on how similar samples are to one another. In the context of -omics and disease, clustering algorithms can be used to group subjects into different subgroups that may demonstrate distinct characteristics within the dataset.
- CpG sites
Regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide in the linear sequence of bases along its 5’ → 3’ direction. CpG sites occur frequently in genomic regions called CpG islands where methylation of the cytosine within a gene can change its expression.
- DNA methylation
A process by which methyl groups are added to DNA, often modifying gene expression.
- Endotype
Subtypes of disease based on shared pathophysiological mechanisms. While a disease label may be given to subjects with a given suite of symptoms and signs, the underlying mechanisms that lead to these symptoms and signs may not be uniform across subjects. Endotypes can help organize mechanistic understanding of diseases that have heterogeneous pathophysiology.
- Epigenome
Modifications made to the genome including chemical modifications to DNA or modifications to the chromatin packaging of DNA. These can include but are not limited to DNA methylation or hydroxy-methylation and post-translational modifications to histones. These modifications frequently alter the transcription levels of nearby genes.
- FeNO
An acronym for fractional exhaled nitric oxide. Nitric oxide is an exhaled gas that is produced by cells involved in inflammation associated with allergic or eosinophilic asthma. FeNO is often checked to assess inflammation and response to steroid medications.
- Gas chromatography mass spectrometry (GC-MS)
An instrumental technique that incorporates a gas chromatograph (GC) coupled to a mass spectrometer (MS), by which complex mixtures of chemicals may be separated, identified and quantified. This is a reliable means to analyze hundreds of relatively low molecular weight compounds found in environmental materials.
- Genome
The complete set of genetic information for an organism, including genes and non-coding sequences. The genome contains the information needed to build and maintain the organism. The human genome is over 3 billion DNA base pairs.
- Innate lymphoid cells
Innate immune cells from the lymphoid lineage that produce cytokines associated with T-helper cell subtypes; these cells however do not express antigen specific T-cell or B-cell receptors. Innate lymphoid cells include those in Group 1 (ILC1s) that produce Th1 cytokines, Group 2 (ILC2s) that produce Th2 related cytokines, and Group 3 (ILC3s) that produce similar cytokines to Th17 cells
- Inflammasome
A multiprotein oligomer of the innate immune system that is responsible for the activation of inflammatory responses primarily through promoting the maturation and secretion of pro-inflammatory cytokines Interleukin 1β (IL-1β) and Interleukin 18 (IL-18) leading to pyroptosis, a form of programmed cell death.
- Liquid chromatography–mass spectrometry (LC-MS)
An analytical chemistry technique that combines the physical separation of liquid chromatography (or HPLC) with the mass analysis of mass spectrometry. Liquid chromatography separates mixtures with multiple components, mass spectrometry provides structural identity of the individual components with high molecular specificity and detection sensitivity.
- Metabolome
The set of all metabolites found in a sample. Nuclear magnetic resonance (NMR), gas chromatography mass spectrometry (GC-MS), or liquid chromatography mass spectrometry (LC-MS) are typically used. Metabolites can be specific to certain body fluids, such as urine, serum, plasma, and exhaled breath condensate.
- Microbiome
The sum of microbes and their genomic elements in a particular environment.
- Multi-omics
Combined measures and/or analyses of multiple system-wide molecular profiles.
- NK cells
Cytotoxic immune cells; small granules in their cytoplasm contain special proteins such as perforin and proteases known as granzymes which induce apoptosis in the target cell. An innate lymphoid cell, NK cells rapidly contain viruses while the adaptive immune system is producing antigen-specific cytotoxic T cells.
- Omics
The study of the role, relationships, and actions of a system-wide measure of a given molecular type; for example, transcriptomics measure all transcripts in a sample, proteomics measure all proteins in a sample, etc.
- Phenotype
Classification based on a common set of observable characteristics that is often a downstream result of genetics and the environment
- Proteome
The set of all proteins expressed in a sample.
- Transcriptome
The set of all RNA molecules in a sample.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Conflicts of interest: The authors declare no conflicts of interest.
References
- 1.Casale TB. Biologics and biomarkers for asthma, urticaria, and nasal polyposis. J Allergy Clin Immunol 2017; 139:1411–21. [DOI] [PubMed] [Google Scholar]
- 2.Fitzpatrick AM, Moore WC. Severe Asthma Phenotypes — How Should They Guide Evaluation and Treatment? The Journal of Allergy and Clinical Immunology: In Practice 2017; 5:901–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Skloot GS. Asthma phenotypes and endotypes: a personalized approach to treatment. Curr Opin Pulm Med 2016; 22:3–9. [DOI] [PubMed] [Google Scholar]
- 4.Woodruff PG, Modrek B, Choy DF, Jia G, Abbas AR, Ellwanger A, et al. T-helper type 2-driven inflammation defines major subphenotypes of asthma. Am J Respir Crit Care Med 2009; 180:388–95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Nagasaki T, Matsumoto H, Izuhara K, Kanemitsu Y, Tohda Y, Horiguchi T, et al. Utility of serum periostin in combination with exhaled nitric oxide in the management of asthma. Allergology International 2017; 66:404–10. [DOI] [PubMed] [Google Scholar]
- 6.Tini G, Marchetti L, Priami C, Scott-Boyer MP. Multi-omics integration—a comparison of unsupervised clustering methodologies. Brief Bioinform 2017; doi: 10.1093/bib/bbx167. [DOI] [PubMed] [Google Scholar]
- 7.Liu T, Wu J, Zhao J, Wang J, Zhang Y, Liu L, et al. Type 2 innate lymphoid cells: A novel biomarker of eosinophilic airway inflammation in patients with mild to moderate asthma. Respiratory Medicine 2015; 109:1391–6. [DOI] [PubMed] [Google Scholar]
- 8.Svenningsen S, Nair P. Asthma Endotypes and an Overview of Targeted Therapy for Asthma. Front Med (Lausanne) 2017; 4:158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tliba O, Panettieri RA Jr. Paucigranulocytic asthma: Uncoupling of airway obstruction from inflammation. J Allergy Clin Immunol 2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ray A, Kolls JK. Neutrophilic Inflammation in Asthma and Association with Disease Severity. Trends in Immunology 2017; 38:942–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Liu W, Liu S, Verma M, Zafar I, Good JT, Rollins D, et al. Mechanism of TH2/TH17-predominant and neutrophilic TH2/TH17-low subtypes of asthma. J Allergy Clin Immunol 2017; 139:1548–58 e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ricciardolo FLM, Sorbello V, Folino A, Gallo F, Massaglia GM, Favata G, et al. Identification of IL-17F/frequent exacerbator endotype in asthma. J Allergy Clin Immunol 2017; 140:395–406. [DOI] [PubMed] [Google Scholar]
- 13.Rathore JS, Wang Y. Protective role of Th17 cells in pulmonary infection. Vaccine 2016; 34:1504–14. [DOI] [PubMed] [Google Scholar]
- 14.Hizawa N, Kawaguchi M, Huang SK, Nishimura M. Role of interleukin 17F in chronic inflammatory and allergic lung disease. Clinical & Experimental Allergy 2006; 36:1109–14. [DOI] [PubMed] [Google Scholar]
- 15.Choy DF, Hart KM, Borthwick LA, Shikotra A, Nagarkar DR, Siddiqui S, et al. TH2 and TH17 inflammatory pathways are reciprocally regulated in asthma. Sci Transl Med 2015; 7:301ra129. [DOI] [PubMed] [Google Scholar]
- 16.Shikotra A, Choy DF, Siddiqui S, Arthur G, Nagarkar DR, Jia G, et al. A CEACAM6-High Airway Neutrophil Phenotype and CEACAM6-High Epithelial Cells Are Features of Severe Asthma. The Journal of Immunology 2017:1600606. [DOI] [PubMed] [Google Scholar]
- 17.Wisniewski JA, Muehling LM, Eccles JD, Capaldo BJ, Agrawal R, Shirley DA, et al. TH1 signatures are present in the lower airways of children with severe asthma, regardless of allergic status. J Allergy Clin Immunol 2018; 141:2048–60.e13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Duvall MG, Barnig C, Cernadas M, Ricklefs I, Krishnamoorthy N, Grossman NL, et al. Natural killer cell-mediated inflammation resolution is disabled in severe asthma. Science immunology 2017; 2:eaam5446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Jartti T, Gern JE. Role of viral infections in the development and exacerbation of asthma in children. J Allergy Clin Immunol 2017; 140:895–906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bunyavanich S, Schadt EE. Systems biology of asthma and allergic diseases: A multiscale approach. Journal of Allergy and Clinical Immunology 2015; 135:31–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Pandey G, Pandey OP, Rogers AJ, Ahsen ME, Hoffman GE, Raby BA, et al. A Nasal Brush-based Classifier of Asthma Identified by Machine Learning Analysis of Nasal RNA Sequence Data. Sci Rep 2018; 8:8826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yick CY, Zwinderman AH, Kunst PW, Grünberg K, Mauad T, Dijkhuis A, et al. Transcriptome sequencing (RNA-Seq) of human endobronchial biopsies: asthma versus controls. European Respiratory Journal 2013; 42:662–70. [DOI] [PubMed] [Google Scholar]
- 23.Bigler J, Boedigheimer M, Schofield JPR, Skipp PJ, Corfield J, Rowe A, et al. A Severe Asthma Disease Signature from Gene Expression Profiling of Peripheral Blood from U-BIOPRED Cohorts. Am J Respir Crit Care Med 2017; 195:1311–20. [DOI] [PubMed] [Google Scholar]
- 24.Yan X, Chu JH, Gomez J, Koenigs M, Holm C, He X, et al. Noninvasive Analysis of the Sputum Transcriptome Discriminates Clinical Phenotypes of Asthma. Ann Am Thorac Soc 2016; 13 Suppl 1:S104–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Howrylak JA, Moll M, Weiss ST, Raby BA, Wu W, Xing EP. Gene expression profiling of asthma phenotypes demonstrates molecular signatures of atopy and asthma control. J Allergy Clin Immunol 2016; 137:1390–7.e6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kuo CS, Pavlidis S, Loza M, Baribaud F, Rowe A, Pandis I, et al. T-helper cell type 2 (Th2) and non-Th2 molecular phenotypes of asthma using sputum transcriptomics in U-BIOPRED. Eur Respir J 2017; 49. [DOI] [PubMed] [Google Scholar]
- 27.Hekking PP, Loza MJ, Pavlidis S, De Meulder B, Lefaudeux D, Baribaud F, et al. Transcriptomic gene signatures associated with persistent airflow limitation in patients with severe asthma. Eur Respir J 2017; 50. [DOI] [PubMed] [Google Scholar]
- 28.Hekking PP, Loza MJ, Pavlidis S, de Meulder B, Lefaudeux D, Baribaud F, et al. Pathway discovery using transcriptomic profiles in adult-onset severe asthma. J Allergy Clin Immunol 2018; 141:1280–90. [DOI] [PubMed] [Google Scholar]
- 29.Peters MC, Ringel L, Dyjack N, Herrin R, Woodruff PG, Rios C, et al. A Transcriptomic Method to Determine Airway Immune Dysfunction in T2-High and T2-Low Asthma. Am J Respir Crit Care Med 2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Peters MC, Mekonnen ZK, Yuan S, Bhakta NR, Woodruff PG, Fahy JV. Measures of gene expression in sputum cells can identify TH2-high and TH2-low subtypes of asthma. J Allergy Clin Immunol 2014; 133:388–94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Baines KJ, Simpson JL, Wood LG, Scott RJ, Fibbens NL, Powell H, et al. Sputum gene expression signature of 6 biomarkers discriminates asthma inflammatory phenotypes. J Allergy Clin Immunol 2014; 133:997–1007. [DOI] [PubMed] [Google Scholar]
- 32.Fricker M, Gibson PG, Powell H, Simpson JL, Yang IA, Upham JW, et al. A sputum 6 gene signature predicts future exacerbations of poorly controlled asthma. J Allergy Clin Immunol 2019. [DOI] [PubMed] [Google Scholar]
- 33.Yang IV, Lozupone CA, Schwartz DA. The environment, epigenome, and asthma. J Allergy Clin Immunol 2017; 140:14–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liang L, Willis-Owen SAG, Laprise C, Wong KCC, Davies GA, Hudson TJ, et al. An epigenome-wide association study of total serum immunoglobulin E concentration. Nature 2015; 520:670–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Chen W, Wang T, Pino-Yanes M, Forno E, Liang L, Yan Q, et al. An epigenome-wide association study of total serum IgE in Hispanic children. J Allergy Clin Immunol 2017; 140:571–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yang IV, Pedersen BS, Liu A, O’Connor GT, Teach SJ, Kattan M, et al. DNA methylation and childhood asthma in the inner city. J Allergy Clin Immunol 2015; 136:69–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Nicodemus-Johnson J, Myers RA, Sakabe NJ, Sobreira DR, Hogarth DK, Naureckas ET, et al. DNA methylation in lung cells is associated with asthma endotypes and genetic risk. JCI Insight 2016; 1:e90151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Chung KF. Airway microbial dysbiosis in asthmatic patients: A target for prevention and treatment? J Allergy Clin Immunol 2017; 139:1071–81. [DOI] [PubMed] [Google Scholar]
- 39.Fazlollahi M, Lee TD, Andrade J, Oguntuyo K, Chun Y, Grishina G, et al. The nasal microbiome in asthma. J Allergy Clin Immunol 2018; 142:834–43.e2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Huang YJ, Nariya S, Harris JM, Lynch SV, Choy DF, Arron JR, et al. The airway microbiome in patients with severe asthma: Associations with disease features and severity. J Allergy Clin Immunol 2015; 136:874–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sverrild A, Kiilerich P, Brejnrod A, Pedersen R, Porsbjerg C, Bergqvist A, et al. Eosinophilic airway inflammation in asthmatic patients is associated with an altered airway microbiome. J Allergy Clin Immunol 2017; 140:407–17.e11. [DOI] [PubMed] [Google Scholar]
- 42.Taylor SL, Leong LEX, Choo JM, Wesselingh S, Yang IA, Upham JW, et al. Inflammatory phenotypes in patients with severe asthma are associated with distinct airway microbiology. J Allergy Clin Immunol 2018; 141:94–103 e15. [DOI] [PubMed] [Google Scholar]
- 43.Durack J, Lynch SV, Nariya S, Bhakta NR, Beigelman A, Castro M, et al. Features of the bronchial bacterial microbiome associated with atopy, asthma, and responsiveness to inhaled corticosteroid treatment. Journal of Allergy and Clinical Immunology 2017; 140:63–75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Kelly RS, Dahlin A, McGeachie MJ, Qiu W, Sordillo J, Wan ES, et al. Asthma Metabolomics and the Potential for Integrative Omics in Research and the Clinic. Chest 2017; 151:262–77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Turi KN, Romick-Rosendale L, Ryckman KK, Hartert TV. A review of metabolomics approaches and their application in identifying causal pathways of childhood asthma. Journal of Allergy and Clinical Immunology 2018; 141:1191–201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Ibrahim B, Basanta M, Cadden P, Singh D, Douce D, Woodcock A, et al. Non-invasive phenotyping using exhaled volatile organic compounds in asthma. Thorax 2011; 66:804–9. [DOI] [PubMed] [Google Scholar]
- 47.van der Schee MP, Palmay R, Cowan JO, Taylor DR. Predicting steroid responsiveness in patients with asthma using exhaled breath profiling. Clin Exp Allergy 2013; 43:1217–25. [DOI] [PubMed] [Google Scholar]
- 48.Sinha A, Desiraju K, Aggarwal K, Kutum R, Roy S, Lodha R, et al. Exhaled breath condensate metabolome clusters for endotype discovery in asthma. J Transl Med 2017; 15:262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Brinkman P, Wagener AH, Hekking PP, Bansal AT, Maitland-van der Zee AH, Wang Y, et al. Identification and prospective stability of electronic nose (eNose)-derived inflammatory phenotypes in patients with severe asthma. J Allergy Clin Immunol 2018. [DOI] [PubMed] [Google Scholar]
- 50.Neerincx AH, Vijverberg SJH, Bos LDJ, Brinkman P, van der Schee MP, de Vries R, et al. Breathomics from exhaled volatile organic compounds in pediatric asthma. Pediatr Pulmonol 2017; 52:1616–27. [DOI] [PubMed] [Google Scholar]
- 51.Schofield JPR, Burg D, Nicholas B, Strazzeri F, Brandsma J, Staykova D, et al. on behalf of the U-BIOPRED Study Group. Stratification of asthma phenotypes by airway proteomic signatures. Journal of Allergy and Clinical Immunology 2019. [DOI] [PubMed] [Google Scholar]
- 52.McGeachie MJ, Dahlin A, Qiu W, Croteau-Chonka DC, Savage J, Wu AC, et al. The metabolomics of asthma control: a promising link between genetics and disease. Immun Inflamm Dis 2015; 3:224–38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Lefaudeux D, De Meulder B, Loza MJ, Peffer N, Rowe A, Baribaud F, et al. U-BIOPRED clinical adult asthma clusters linked to a subset of sputum omics. J Allergy Clin Immunol 2017; 139:1797–807. [DOI] [PubMed] [Google Scholar]
- 54.Rossios C, Pavlidis S, Hoda U, Kuo CH, Wiegman C, Russell K, et al. Sputum transcriptomics reveal upregulation of IL-1 receptor family members in patients with severe asthma. J Allergy Clin Immunol 2018; 141:560–70. [DOI] [PubMed] [Google Scholar]
- 55.Rappoport N, Shamir R. Multi-omic and multi-view clustering algorithms: review and cancer benchmark. Nucleic Acids Research 2018; 46:10546–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Hoadley KA, Yau C, Wolf DM, Cherniack AD, Tamborero D, Ng S, et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 2014; 158:929–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Nguyen T, Tagett R, Diaz D, Draghici S. A novel approach for data integration and disease subtyping. Genome research 2017:gr. 215129.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Zhou D, Burges CJ. Spectral clustering and transductive learning with multiple views. Proceedings of the 24th international conference on Machine learning: ACM, 2007:1159–66. [Google Scholar]
- 59.Kumar A, Rai P, Daume H. Co-regularized multi-view spectral clustering. Advances in neural information processing systems, 2011:1413–21. [Google Scholar]
- 60.Wang B, Mezlini AM, Demir F, Fiume M, Tu Z, Brudno M, et al. Similarity network fusion for aggregating data types on a genomic scale. Nature methods 2014; 11:333. [DOI] [PubMed] [Google Scholar]
- 61.Priami C, Tini G, Marchetti L, Scott-Boyer M-P. Multi-omics integration—a comparison of unsupervised clustering methodologies. 2017. [DOI] [PubMed] [Google Scholar]
- 62.Rohart F, Gautier B, Singh A, Le Cao K-A. mixOmics: An R package for ‘omics feature selection and multiple data integration. PLoS computational biology 2017; 13:e1005752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Mo Q, Shen R, Guo C, Vannucci M, Chan KS, Hilsenbeck SG. A fully Bayesian latent variable model for integrative clustering analysis of multi-type omics data. Biostatistics 2017; 19:71–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hidru D, Goldenberg A. EquiNMF: Graph regularized multiview nonnegative matrix factorization. arXiv preprint arXiv:1409.4018 2014. [Google Scholar]
- 65.Ma C, Jones DP, Go Y-M, Uppal K. xMWAS: a data-driven integration and differential network analysis tool. Bioinformatics 2017; 34:701–2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Muraro A, Lemanske RF Jr., Hellings PW, Akdis CA, Bieber T, Casale TB, et al. Precision medicine in patients with allergic diseases: Airway diseases and atopic dermatitis-PRACTALL document of the European Academy of Allergy and Clinical Immunology and the American Academy of Allergy, Asthma & Immunology. J Allergy Clin Immunol 2016; 137:1347–58. [DOI] [PubMed] [Google Scholar]
- 67.Chipps BE, Lanier B, Milgrom H, Deschildre A, Hedlin G, Szefler SJ, et al. Omalizumab in children with uncontrolled allergic asthma: Review of clinical trial and real-world experience. Journal of Allergy and Clinical Immunology 2017; 139:1431–44. [DOI] [PubMed] [Google Scholar]
- 68.Barnes PJ. Targeting cytokines to treat asthma and chronic obstructive pulmonary disease. Nature Reviews Immunology 2018; 18:454–66. [DOI] [PubMed] [Google Scholar]