Abstract
The Parkinson’s disease (PD)-related ubiquitin ligase Parkin and mitochondrial kinase PINK1 function together in the clearance of damaged mitochondria. Upon mitochondrial depolarization, Parkin translocates to mitochondria in a PINK1-dependent manner to ubiquitinate outer mitochondrial membrane proteins. According to the current model, the ubiquitin- and LC3-binding adaptor protein SQSTM1 is recruited to mitochondria, followed by their selective degradation through autophagy (mitophagy). However, the role of the ubiquitin proteasome system (UPS), although essential for this process, still remains largely elusive. Here, we investigated the role of the UPS and autophagy by applying the potassium ionophore Valinomycin in PINK1-deficient human fibroblasts and isogenic neuroblastoma cell lines generated by CRISPR/Cas9. Although identical to the commonly used CCCP/FCCP in terms of dissipating the mitochondrial membrane potential and triggering complete removal of mitochondria, Valinomycin did not induce conversion of LC3 to its autophagy-related form. Moreover, FCCP-induced conversion of LC3 occurred even in mitophagy-incompetent, PINK1-deficient cell lines. While both stressors required a functional UPS, the removal of depolarized mitochondria persisted in cells depleted of LC3A and LC3B. Our study highlights the importance of the UPS in PINK1-/Parkin-mediated mitochondrial quality control. In contrast, activation of autophagy, monitored through conversion of LC3, is likely induced by depolarizing-agent-induced toxicity in a PINK1-/Parkin-independent manner.
Subject terms: Neuroscience, Diseases
Introduction
Mutations in the E3 ubiquitin ligase Parkin and the mitochondrial kinase PINK1 cause autosomal recessive forms of Parkinson’s disease (PD) [1]. PINK1 acts upstream of Parkin in a pathway for the maintenance of mitochondrial function and morphology [2–4]. It has been shown that reduction of the mitochondrial membrane potential (ΔΨm) leads to aggregation of PINK1 on mitochondria, which recruits Parkin to mitochondria. Parkin polyubiquitinates outer mitochondrial membrane (OMM) proteins [5] and recruits the ubiquitin- and LC3-binding autophagic adaptor protein SQSTM1 to aggregates of damaged mitochondria [6, 7], supposedly promoting their degradation by autophagy (mitophagy) [8, 9]. This model, however, requires further clarification because depletion of SQSTM1 fails to inhibit Parkin-mediated mitophagy [10, 11], suggesting that another mechanism, independent of SQSTM1, may contribute to this process. Indeed, a growing body of evidence supports the notion that the ubiquitin proteasome system (UPS) is necessary for PINK1-/Parkin-mediated removal of depolarized mitochondria [12, 13]. In this respect, it has been demonstrated that Parkin polyubiquitinates numerous OMM proteins [5], followed by their degradation via the UPS [12–15].
In this context, is important to bear in mind that all of the >100 published studies on PINK1-/Parkin-mediated mitophagy are based on an artificial experimental set-up using overexpression of Parkin and agents inducing a complete loss of the ΔΨm. Of particular note, the vast majority of studies on PINK1-/Parkin-mediated mitophagy have employed almost exclusively one type of respiratory chain uncouplers, i.e., the hydrogen ionophores carbonyl cyanide m-chlorophenyl hydrazone (CCCP) or carbonyl cyanide-4-(trifluoromethoxy) phenylhydrazone (FCCP).
Another classical respiratory chain uncoupler is the potassium ionophore Valinomycin that has been extensively used outside of the Parkin/PINK1 field in studies requiring dissipation of the ΔΨm [16–18]. CCCP/FCCP and Valinomycin have identical effects on mitochondria in that they both (i) induce immediate dissipation of the ΔΨm [17, 19], (ii) increase the mitochondrial volume by ~65% without effect on the surface area [16], (iii) initiate stabilization of PINK1 on mitochondria and mitochondrial translocation of Parkin [8, 20], and (iv) promote complete removal of depolarized mitochondria [5, 6, 8]. Furthermore, both CCCP/FCCP and Valinomycin do not specifically act on mitochondria, but rather non-selectively bind to lysosomal [21, 22] and plasma membranes [23, 24]. However, in contrast to Valinomycin, CCCP/FCCP alkalinize lysosomes and thus affect their function and autophagic flux [25]. Furthermore, CCCP inhibits autophagy at both the initiation and lysosomal degradation stages [25].
Taken together, the uncertainty about the exact mechanism of PINK1-/Parkin-mediated mitochondrial clearance and the fact that the current experimental systems affect the very pathway under study, prompted us to revisit the role of the UPS and lysosomal systems in removing depolarized mitochondria using not only FCCP but also Valinomycin as a type of uncoupler not directly impacting on lysosomal function.
Results
Generation of PINK1 knockout neuroblastoma cell lines
To knock out PINK1 in neuroblastoma (SH-SY5Y) cells we used CRISPR/Cas9 technology (Fig. 1a). We identified one clonal cell line carrying compound-heterozygous mutations in PINK1 ([c.84_142del58bp]+[c.135_136ins95bp]) (Fig. 1b) resulting in frameshift and a premature stop codon (Fig. 1c). To confirm the absence of PINK1, control and PINK1 knockout SH-SY5Y (PINK1KO) cells were incubated with Valinomycin for 6 h. As expected, we detected the accumulation of endogenous PINK1 in Valinomycin-treated control but not PINK1KO cells (Fig. 1d). Furthermore, PINK1 mRNA levels in PINK1KO cells were only 10 ± 3% of PINK1 mRNA levels in control cells, suggesting that the vast majority of PINK1 mRNA in PINK1KO cells is removed by nonsense-mediated decay (Fig. 1e).
Fig. 1.
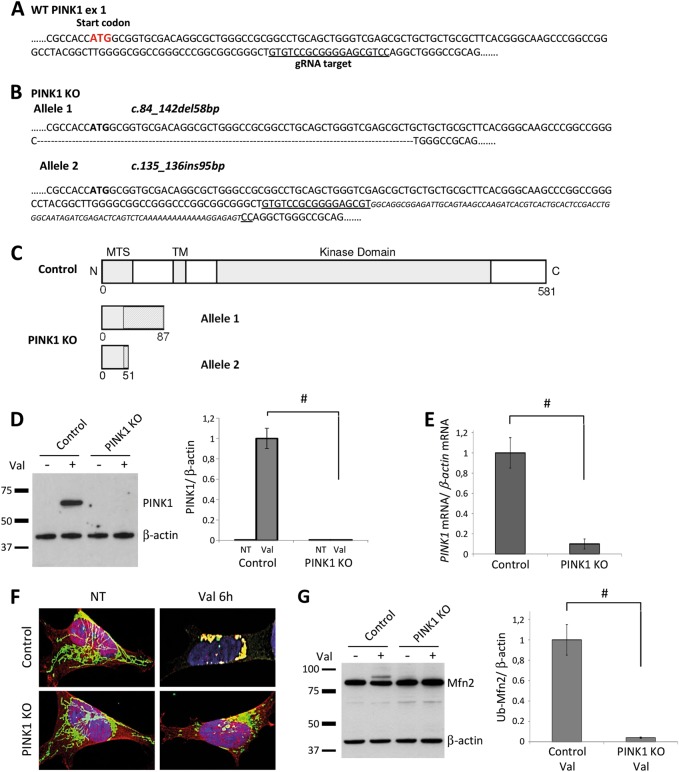
CRISPR/Cas9-mediated knockout of PINK1 in neuroblastoma cells. a Neuroblastoma (SH-SY5Y) cells were transfected with episomal vectors expressing Cas9 and gRNA targeting the underlined sequence located in exon 1 of the PINK1 gene. b The PINK1 knockout (PINK1KO) clonal cell line carries compound-heterozygous mutations in PINK1 ([c.84_142del58bp]+[c.135_136ins95bp]). c Schematic representation of the wildtype PINK1 protein and putative truncated forms of the PINK1 protein in PINK1KO cells. Areas shaded in gray represent a frame-shifted protein. d To detect endogenous PINK1 protein, control and PINK1KO cells were treated with Valinomycin for 6 h and analyzed by western blotting using antibodies against PINK1. Protein levels of PINK1 were quantified and normalized to levels of β-actin. e PINK1 mRNA expression in control and PINK1KO SH-SY5Y cells. The values represent means ± SD from three independent measurements. f Control and PINK1KO cells treated with Valinomycin for 6 h were fixed and immunostained using antibodies against Parkin (red) and the mitochondrial matrix protein GRP75 (green). g Immunoblot of untreated and Valinomycin-treated control and PINK1KO cells probed with an antibody against MFN2. Levels of ubiquitinated MFN2 (Ub-MFN2) in Valinomycin-treated cells were normalized to levels of β-actin. #p < 0.01
To further characterize this PINK1KO cellular model, we analyzed its ability to perform two well established PINK1-dependent activities: translocation of Parkin to depolarized mitochondria [6, 8] and Parkin-mediated ubiquitination of the OMM protein MITOFUSIN 2 (MFN2) [15, 26]. As expected and consistent with previous studies, mitochondrial translocation of Parkin (Fig. 1f) and ubiquitination of MFN2 (Fig. 1g) occurred only in Valinomycin-treated controls but not in PINK1KO cells. These data suggest that PINK1KO cells represent a useful new model to study PINK1 deficiency-related cellular processes.
The UPS is essential for removal of depolarized mitochondria
While previous studies demonstrated that mitochondrial depolarization leads to autophagy-mediated removal of mitochondria (mitophagy) [6, 8, 9], we and others showed that mitochondrial depolarization leads to PINK1-dependent, Parkin-mediated polyubiquitination [5] and degradation of OMM proteins that can be prevented by inhibiting the UPS [12].
Accordingly, we sought to dissect the precise role of the UPS in removing depolarized mitochondria. For this, we used control and PINK1KO cells stably overexpressing wildtype Parkin since levels of endogenous Parkin are not sufficient to initiate detectable mitophagy [5, 12]. To analyze mitophagy, we performed immunoblotting to monitor steady-state levels of mitochondrial proteins localized in the OMM, the inner mitochondrial membrane (IMM), and the mitochondrial matrix (Fig. 2). This method to monitor the reduction in mitochondrial mass is comparable to the commonly used MitoTracker dye labeling [5] but has the added advantage that we can simultaneously follow changes in all mitochondrial compartments during mitophagy. Upon Valinomycin treatment, we detected a reduction in the levels of almost all analyzed mitochondrial proteins in control cells but not in PINK1KO cells, indicative of PINK1-dependent mitochondrial removal (Fig. 2, second lanes). The only exceptions were two matrix proteins, HSP60 and SOD2, which remained unaffected by Valinomycin treatment. Inhibition of the UPS by MG132 prevented Valinomycin-induced removal of mitochondria (Fig. 2, third lanes). Specifically, inhibition of the UPS prevented the loss of IMM proteins and the matrix protein GRP75 and preserved the ubiquitinated forms of larger OMM proteins (e.g., MFN2, TOM70). This result is consistent with previous reports [12, 13] showing that degradation of OMM proteins occurs via the UPS independently of autophagy. To further support this finding, we treated cells with both Valinomycin and the lysosomal inhibitor Bafilomycin A1 (Fig. 2, fourth lanes). Similar to proteasomal inhibition, lysosomal inhibition preserved IMM and matrix proteins, as well as smaller OMM proteins, such as VDAC1 and TOM20. However, Bafilomycin A1 failed to prevent Valinomycin-induced loss of larger OMM proteins.
Fig. 2.
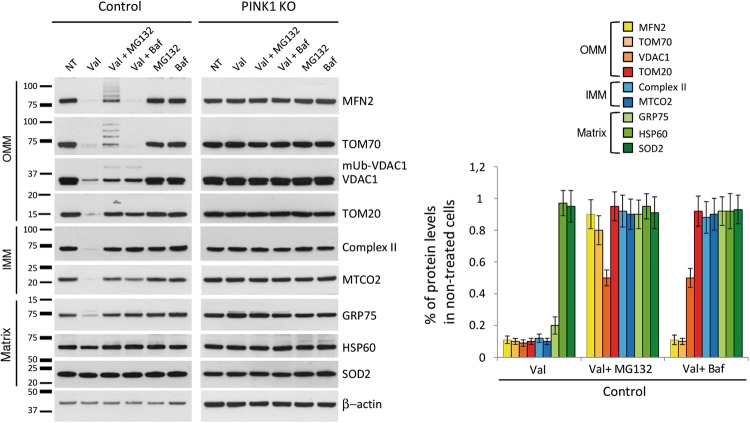
Inhibition of the UPS or lysosomal system prevents removal of depolarized mitochondria in neuroblastoma cells. Control or PINK1KO neuroblastoma cells engineered to stably overexpress Parkin were treated with Valinomycin alone or in combination with MG132 or Bafilomycin A1 for 16 h. In addition, cells were treated with either MG132 or Bafilomycin A1. Cells were harvested and analyzed by western blot using antibodies against mitochondrial proteins localized in different mitochondrial compartments (outer mitochondrial membrane (OMM), inner mitochondrial membrane (IMM), and matrix). Protein levels in control neuroblastoma cells were quantified and normalized to levels of β-actin. Values represent means ± SD from three independent measurements.
We further examined the role of the UPS and lysosomal systems in human dermal fibroblasts. For this, we used cells obtained from a healthy control and from a PD patient carrying a homozygous mutation (c.509T>G; p.V170G) in PINK1 (PINK1mut), again both engineered to stably overexpress wildtype Parkin. Valinomycin-treated control (Fig. 3a) and PINK1mut (Fig. 3b) fibroblasts were analyzed upon inhibition of the UPS or lysosomes with epoxomicin or Bafilomycin A1, respectively. The OMM proteins MFN2, and TOM70 were exclusively degraded via the UPS (Fig. 3a), whereas the smaller OMM proteins, TOM40 and TOM20, were only partially ubiquitinated, but mostly degraded through lysosomal-mediated proteolysis. Valinomycin-induced degradation of the IMM proteins, Complex II Fp subunit (Complex II), F1F0ATPase (α subunit), and MT-CO2, and of the mitochondrial matrix proteins, GRP75 and TFAM, could be protected using either one of the inhibitors (Fig. 3a). Again, protein levels of two other matrix proteins, HSP60 and SOD2 were unaffected after 16 h of Valinomycin treatment (Fig. 3a). While HSP60 was degraded after 24 h of Valinomycin treatment, levels of SOD2 remained unaffected also at this time point. In PINK1mut cells, as expected, neither treatment affected the levels of the mitochondrial proteins studied (Fig. 3b).
Fig. 3.
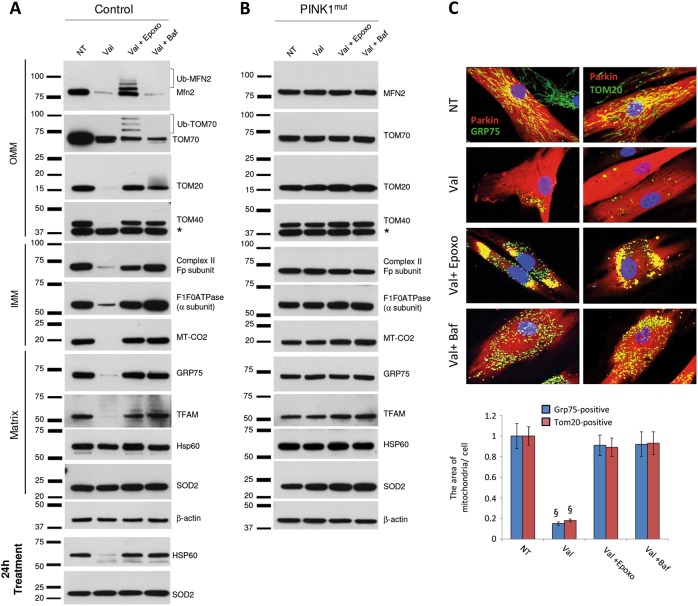
Inhibition of the UPS or lysosomal system prevents removal of depolarized mitochondria in human fibroblasts. a Control and b PINK1mut fibroblasts stably overexpressing Parkin were treated with Valinomycin only or in combination either with Epoxomicin or with Bafilomycin A1 for 16 h. Cells were immunoblotted using antibodies against mitochondrial proteins localized in the three different mitochondrial compartments. β-actin served as loading control. Results with longer (24 h) Valinomycin treatment are also shown for HSP60 and SOD2. *: non-specific band. c Control fibroblasts stably expressing Parkin were treated with Valinomycin only or in combination either with Epoxomicin or with Bafilomycin A1 for 16 h. Cells were immunostained using antibodies against Parkin (red) together with either GRP75 (green) or TOM20 (green). Average area of either GRP75-positive or TOM20-positive mitochondria per cell. Values represent means ± SD from three independent measurements. §p < 0.01 vs. non-treated cells
Immunocytochemistry was used to confirm and extend the above immunoblot results. Control fibroblasts overexpressing Parkin, were treated with Valinomycin alone or in combination with epoxomicin or Bafilomycin A1 (Fig. 3c). In non-treated cells, Parkin (red) was diffusely distributed in the cytosol and mitochondria were organized in an intact network (green). Valinomycin treatment resulted in the nearly complete removal of mitochondria (detected with antibodies to GRP75 and TOM20), while Parkin remained diffusely distributed in the cytosol. However, when the UPS is inhibited, Valinomycin treatment resulted in the mitochondria becoming arrested within perinuclear aggregates that colocalized with Parkin, and no loss of mitochondrial was detected. Inhibition of the lysosomal system resulted in a punctate, homogenous distribution of GRP75 and TOM20, with Parkin diffusely localized throughout the cytoplasm. The area of GRP75- or TOM20-positive mitochondria per cell was comparable to non-treated cells, suggesting that no loss of mitochondria was observed.
Our data indicate that mitochondrial proteins can be degraded through two different mechanisms: the UPS or lysosomal-mediated proteolysis. Of note, they are differentially degraded, with some of them being rapidly removed and others remaining unaffected even upon prolonged treatment.
Valinomycin induces removal of depolarized mitochondria without LC3 conversion
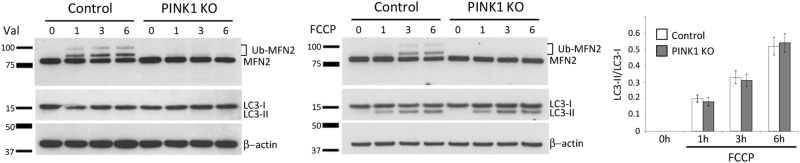
Previous studies using FCCP/CCCP demonstrated that PINK1-/Parkin-dependent removal of depolarized mitochondria is mediated by autophagy [6, 9]. During this process, Microtubule-associated protein 1A/1B-light chain 3 (LC3) is converted from its cytosolic form LC3-I into the autophagy-related form, LC3-II. Detecting LC3 conversion by immunoblotting or immunofluorescence is a reliable method of monitoring autophagy and autophagy-related processes [27]. While prior studies used FCCP/CCCP to induce mitochondrial depolarization, we sought to analyze the process of autophagy upon Valinomycin treatment. For this, control SH-SY5Y cells stably overexpressing Parkin were treated with FCCP or Valinomycin for 16 h and assessed by immunoblot analysis of mitochondrial proteins at different time points (Fig. 4). We observed no difference between these two stressors in the rate of mitochondrial protein degradation (Fig. 4a). In contrast, the conversion of LC3 differed markedly: While FCCP induced rapid and pronounced conversion of LC3 (Fig. 4b), Valinomycin did not. To exclude the possibility that the observed differences in LC3 conversion result from differential toxicity of Valinomycin and FCCP, we performed a trypan blue exclusion assay (Fig. 4c). Here, we did not detect any difference in cell toxicity between the two compounds. Furthermore, no difference in cell toxicity between controls and PINK1KO cells using either Valinomycin or FCCP (Fig. 4c).
Fig. 4.
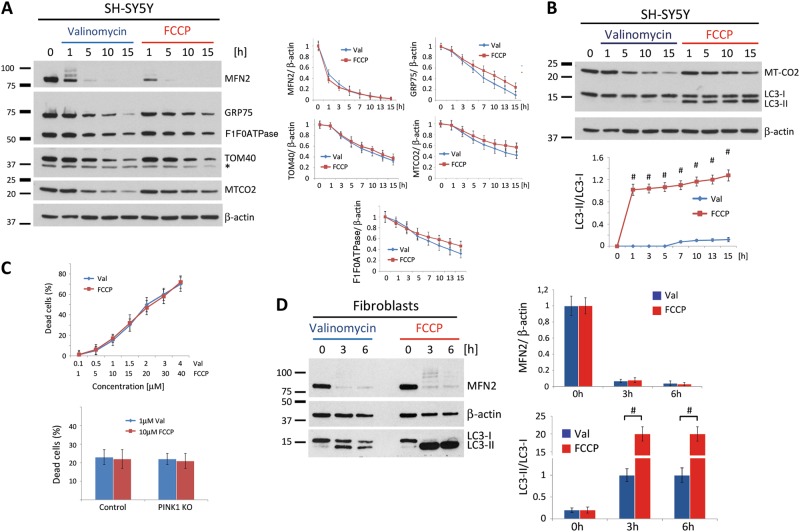
Valinomycin-induced clearance of mitochondria does not require activation of autophagy. Neuroblastoma cells stably overexpressing Parkin were harvested at different time points and analyzed by western blotting using antibodies against a proteins localized in different mitochondrial compartments or against b the autophagy marker LC3. Protein levels were quantified and normalized to levels of β-actin. c Trypan blue exclusion assay test in control neuroblastoma cells overexpressing Parkin treated with various concentration of Valinomycin or FCCP for 16 h (upper panel). Trypan blue exclusion assay in control and PINK1KO neuroblastoma cells overexpressing Parkin treated with either Valinomycin or FCCP for 16 h (lower panel). d Human dermal fibroblasts from a healthy control overexpressing Parkin were treated with either Valinomycin or FCCP. Cells were harvested 7.5 and 15 h upon Valinomycin treatment and immunoblotted using antibodies against MFN2 and LC3. Values represent means ± SD from three independent measurements. #p < 0.01 vs. Valinomycin. Asterisk: nonspecific band
Taken together, we conclude that LC3 conversion, which is indicative of autophagy, is not necessary for Valinomycin-induced removal of mitochondria. To support this hypothesis, we analyzed LC3 conversion in human fibroblasts overexpressing Parkin upon either Valinomycin or FCCP treatment (Fig. 4d). As in SH-SY5Y cells, both stressors induced rapid reduction in MFN2 protein levels (Fig. 4d), but differed significantly with respect to LC3 conversion. While Valinomycin induced a partial conversion of LC3, FCCP resulted in essentially complete conversion (Fig. 4d).
In conclusion, our data indicate that FCCP induces LC3 conversion, which is independent of the removal of depolarized mitochondria. To test this hypothesis, we treated control and PINK1KO cells with either Valinomycin or FCCP (Fig. 5). As shown previously, these cells fail to remove depolarized mitochondria most likely due to either low levels of endogenous Parkin [5, 6, 12] or lack of functional PINK1. While both treatments had identical effects on MFN2, i.e., rapid monoubiquitination in a PINK1-dependent manner, a difference in LC3 conversion was observed. Valinomycin did not induce detectable LC3 conversion in either control or PINK1KO cells, whereas FCCP induced a strong time-dependent LC3 conversion in both control and PINK1KO cells, confirming our hypothesis.
Fig. 5.
FCCP initiates LC3 conversion in mitophagy-incompetent cells. Control and PINK1KO neuroblastoma cells expressing endogenous levels of Parkin were treated with either Valinomycin or FCCP for 6 h and harvested at different time points. Whole cell lysates were analyzed by western blotting using antibodies against MFN2 or LC3. Protein levels of LC3 were quantified and normalized to levels of β-actin. Values represent means ± SD from three independent measurements
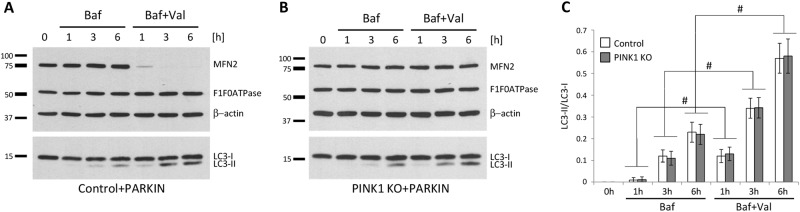
Finally, we compared Valinomycin-induced conversion of LC3 between control and PINK1KO cells overexpressing Parkin in the presence of lysosomal inhibitor (Fig. 6), as this is a more accurate way to asses autophagy [27]. To determine basic autophagy flux, control (Fig. 6a) and PINK1KO (Fig. 6b) cells were treated with Bafilomycin alone. As expected, Bafilomycin induced an increase in levels of LC3-II by preventing its degradation in the lysosomes. When cells were treated with Bafilomycin and Valinomycin together, we detected an increase in the LC3 conversion in comparison to cells treated with Bafilomycin alone (Fig. 6c) most likely due to their combined toxicity. More importantly, we did not observe a difference in autophagic flux between controls and PINK1KO cells.
Fig. 6.
Autophagy flux is comparable between Valinomycin-treated control and mitophagy-incompetent PINK1 knockout cells. a Control and b PINK1KO neuroblastoma cells overexpressing Parkin were treated with Bafilomycin A1 alone or in combination with Valinomycin for 6 h and harvested at different time points. Whole cell lysates were analyzed by western blotting using antibodies against MFN2, the α subunit of F1F0ATPase or LC3. c Protein levels of LC3 were quantified and normalized to levels of β-actin. Values represent means ± SD from three independent measurements.
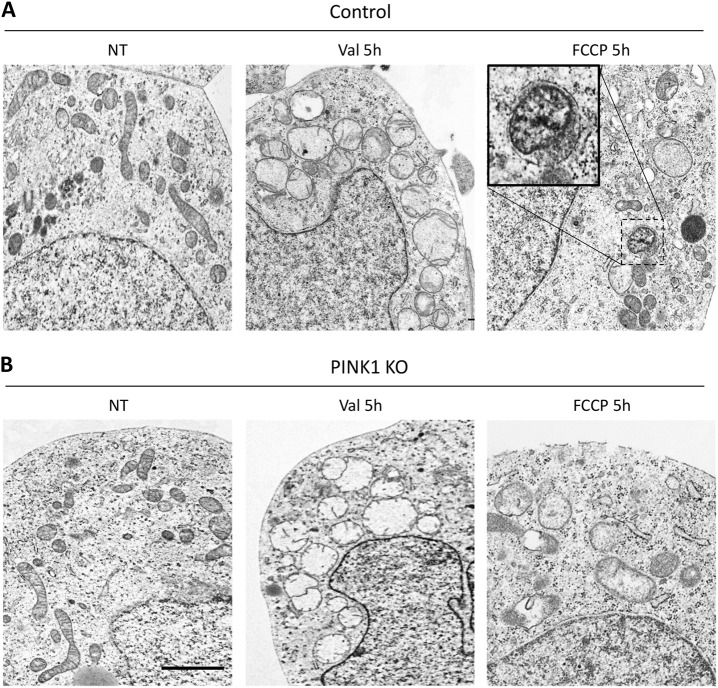
The transmission electron microscopy (TEM) is still considered as the “gold standard” for verifying macroautophagy [28]. Since the formation of autophagic vacuoles in CCCP-treated controls overexpressing Parkin has already been demonstrated by TEM [9, 13], we sought to compare Valinomycin- and FCCP-treated controls and PINK1KO cells using TEM (Fig. 7). For this, cells were treated with either Valinomycin or FCCP for 5 h, as we already observed a loss of mitochondria at this time point (Fig. 4a). In controls, Valinomycin induced swelling and accumulation of mitochondria in the perinuclear region, however, we did not detect any morphological evidence of autophagic structures around mitochondria. In contrast, in FCCP-treated controls, mitochondrial remnants were detected in membrane-surrounded vacuoles consistent with mitophagy. In PINK1KO cells, Valinomycin and FCCP both induced mitochondrial swelling but no morphological signs of macroautophagy were detectable.
Fig. 7.
Valinomycin does not induce the formation of autophagic structures. a Control and b PINK1KO neuroblastoma cells stably overexpressing Parkin were treated with either Valinomycin or FCCP for 5 h and compared to non-treated cells. Cells were fixed and prepared for electron microscopy. Treatment with Valinomycin induced mitochondrial swelling and accumulation in the perinuclear region in both controls and PINK1KO cells. Importantly, no isolation membranes or any other autophagic structures were observed. In contrast, upon FCCP treatment, some mitochondria were surrounded by autophagosome-like structures (inset) only in control but not in PINK1KO cells. Val Valinomycin, NT non-treated; scale bar: 2 μm
Our LC3 conversion and TEM data suggest that FCCP-induced removal of mitochondria is paralleled by strong activation of autophagy, whereas Valinomycin-initiated mitochondrial degradation induces no or very mild autophagy. Furthermore, our data indicate that conversion of LC3 might not be an ideal indicator of PINK1-/Parkin-mediated removal of depolarized mitochondria, as no difference in LC3 lipidation was observed between control and PINK1KO cells. Rather, the observed increase in LC3-II levels occurs most likely as a result of cell injury.
Knockdown of LC3A or LC3B fails to prevent PINK1-/Parkin-mediated removal of depolarized mitochondria
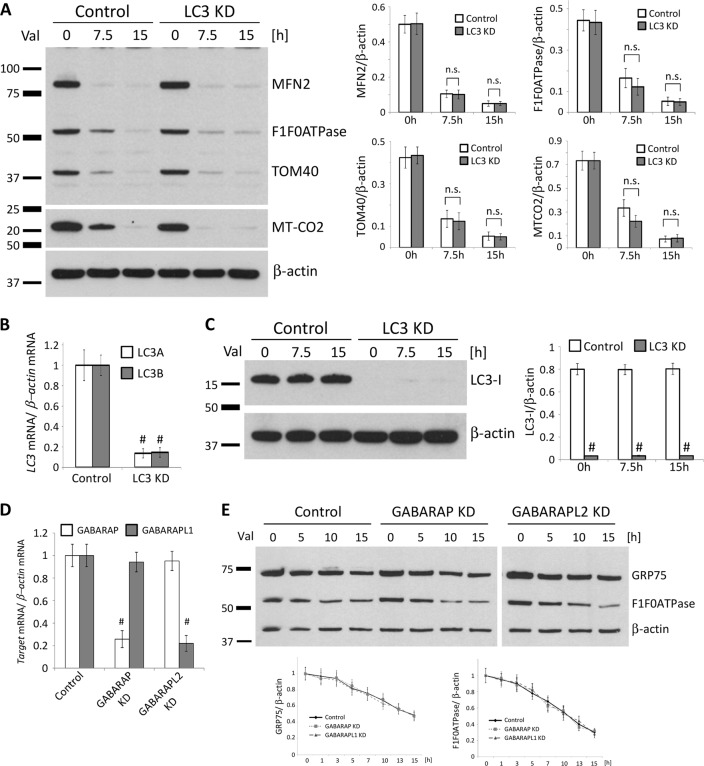
It has been reported previously that degradation of OMM proteins occurs via the proteasome system and is independent of autophagy [12]. Since the removal of depolarized mitochondria does not seem to require LC3 conversion, we sought to test whether LC3 is dispensable for PINK1-/Parkin-dependent mitochondrial clearance. Previous findings in cancer cells showed that the isoforms LC3A and LC3B are mainly located in the perinuclear region and equally distributed throughout the cytoplasm, whereas LC3C is mainly present in the nuclei [29] and therefore most likely less relevant for autophagy. Here, we knocked down both the A and B isoforms of LC3 in control SH-SY5Y cells overexpressing Parkin (LC3KD), using lentiviral particles expressing a small hairpin RNAs (shRNA) against both LC3A and LC3B mRNAs. To test the ability of LC3KD cells to remove depolarized mitochondria, we treated them with Valinomycin for 15 h and examined various mitochondrial proteins by immunoblotting in order to simultaneously detect changes in the OMM and IMM (Fig. 8a). LC3KD cells proved to be as efficient as control cells in the removal of all tested mitochondrial proteins. Knockdown efficiency was more than 90% (Fig. 8b) and no LC3 protein was detectable by western blot (Fig. 8c).
Fig. 8.
LC3, GABARAP, or GABARAPL1 are not required for Valinomycin-induced removal of mitochondria. Neuroblastoma cells (SH-SY5Y) stably overexpressing Parkin were transduced with lentiviral particles expressing scrambled shRNA (Control) or shRNAs against both LC3A and LC3B (LC3KD). a Control and LC3KD cells were treated with Valinomycin for 15 h and harvested at different time points. Whole cell lysates were analyzed by western blotting using antibodies against different mitochondrial proteins. Protein levels of mitochondrial proteins were quantified and normalized to levels of β-actin. The knockdown efficiency of LC3 was confirmed b by real-time PCR and c by western blotting. d GABARAP or GABARAPL2 knockdown efficiency in SH-SY5Y overexpressing Parkin was determined by real-time PCR. e Control, GABARAPKD, or GABARAPL1KD cells were treated with Valinomycin for 15 h and harvested at different time points. Whole cell lysates were analyzed by western blotting using antibodies against mitochondrial proteins GRP75 and α subunit of F1F0ATPase. Protein levels of mitochondrial proteins were quantified and normalized to levels of β-actin. Values represent means ± SD from three independent measurements. #p < 0.01 vs. control. Val Valinomycin, n.s. not significant
A previous study has shown that the autophagy adaptor proteins GABARAP and GABARAPL2/GATE-16 are recruited to damaged mitochondria and promote their elimination [30, 31]. Therefore, we generated GABARAP (GABARAPKD) and GABARAPL2/GATE-16 (GABARAPL2KD) knockdown SH-SY5Y cells overexpressing Parkin using lentiviral particles expressing shRNAs against either GABARAP mRNA or GABARAPL2/GATE-16 mRNA (Fig. 8d). These cells were treated with Valinomycin for 15 h and levels of mitochondrial proteins were measured at different time points. Here, we observed a comparable reduction in levels of mitochondrial proteins between control and either GABARAPKD or GABARAPL2KD cells, suggesting that these two autophagy adaptors are not involved in Valinomycin-induced mitochondrial clearance. Our data are consistent with recently published data showing that GABARAP proteins do not translocate to damaged mitochondria during mitophagy [32] and do not play an important role in PINK1-mediated mitochondrial removal.
Knockout of ATG7 fails to prevent PINK1-/Parkin-mediated removal of outer mitochondrial proteins
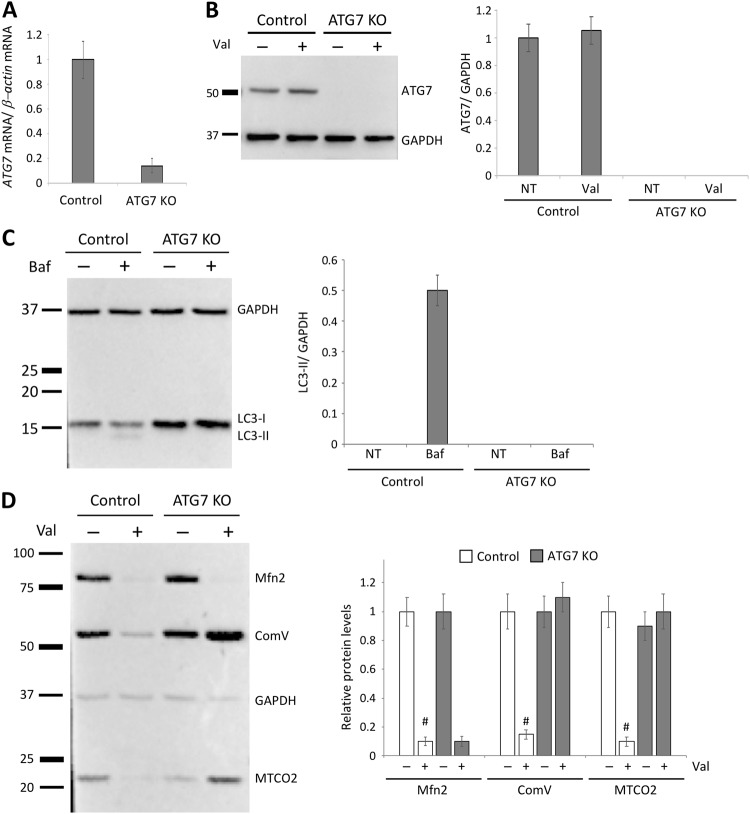
To investigate whether inactivation of autophagy upstream of LC3 affects Valinomycin-induced mitochondrial removal, we generated ATG7 knockout neuroblastoma cells (ATG7KO) using CRISPR/Cas9 technology (Fig. 9). We identified one clone carrying a homozygous deletion in ATG7 (c.del157_158, p.N53fsX1) resulting in frameshift and a premature stop codon. In ATG7KO cells, the ATG7 mRNA levels were only 15 ± 2% of ATG7 mRNA levels in control cells (Fig. 9a) and no ATG7 protein could be detected (Fig. 9b). Since ATG7 acts as an E1-like activating enzyme facilitating both lipidation of LC3-phosphatidylethanolamine and conjugation of ATG12 [33], we investigated autophagy flux in control and ATG7KO cells. For this, cells were treated with Bafilomycin A1 for 12 h and conversion of LC3 was analyzed using western blotting. As expected, we detected LC3 conversion only in control cells (Fig. 9c), which is indicative of impaired autophagy in ATG7KO cells.
Fig. 9.
Autophagy is not involved in the removal of outer mitochondrial membrane proteins. a ATG7 mRNA expression in control and ATG7 knockout neuroblastoma cells (ATG7KO). b Protein levels of ATG7 in control and ATG7KO levels. c Lack of the Bafilomycin A-induced conversion of non-autophagic LC3-I to autophagic LC3-II confirmed impaired autophagic flux in ATG7KO cells. d Control and ATG7KO 7 cells stably expressing Parkin were treated with Valinomycin for 15 h. Whole cell lysates of non-treated and Valinomycin-treated cells were analyzed by western blotting using antibodies against the outer mitochondrial membrane protein MFN2 and the inner mitochondrial membrane proteins α subunit of F1F0ATPase (ComV) and MTCO2. Protein levels of mitochondrial proteins were quantified and normalized to levels of GAPDH. Values represent means ± SD from three independent measurements. #p < 0.01 vs. control. Val Valinomycin
Finally, we assessed Valinomycin-induced removal of proteins localized in different mitochondrial compartments (Fig. 9d). In control cells, we detected loss of all mitochondrial proteins. However, in ATG7KO cells, loss of autophagy could not prevent removal of the OMM protein MFN2, suggesting that a process other than autophagy is responsible for the initiation of removal of damaged mitochondria.
Discussion
In this study, we demonstrated that PINK1-/Parkin-mediated removal of depolarized mitochondria can occur independently of macroautophagy and that both the UPS and lysosomal system play an essential role in this process. In contrast to previous studies using only one kind of respiratory chain uncoupler, i.e., CCCP or FCCP to induce loss of the mitochondrial membrane potential (ΔΨm), we here compared the effects of the commonly used, hydrogen-specific uncoupler FCCP with those of the potassium ionophore Valinomycin.
Conversion of LC3 from its cytosolic form LC3-I into its autophagy-related form LC3-II [26] or colocalization of exogenously expressed GFP-/RFP-tagged LC3 with mitochondria and Parkin [9, 34] have been proposed as reliable indicators of mitophagy. Consistent with previous studies [26], we showed that FCCP-induced removal of mitochondria was accompanied by strong LC3 conversion. However, we observed comparable conversion of LC3 between control cells and PINK1KO cells, which were not able to remove damaged mitochondria, suggesting that FCCP itself strongly activates the autophagy independently of removal of damaged mitochondria. On the other hand, Valinomycin was equally potent to dissipate the ΔΨm, to initiate mitochondrial translocation of Parkin [20], and to induce complete removal of mitochondria in a PINK1-dependent manner [5]. However, in comparison to FCCP, Valinomycin did not induce LC3-related autophagy since no (or only minimal) conversion of LC3 was observed. Moreover, our experiments demonstrated that lack of the isoforms LC3A and LC3B does not impair removal of depolarized mitochondria, however, although unlikely, we cannot exclude the possibility that the isoform LC3C cannot compensate for this loss. We further corroborated our findings by electron microscopy showing that Valinomycin-induced removal of mitochondria is not paralleled by the formation of autophagic structures. Our data are thus consistent with previous findings showing that knocking down of all isoforms of LC3 severely inhibited macroautophagy, however only marginally suppressed starvation-based mitochondrial clearance [35].
Our data imply that both the UPS and the lysosomal system jointly play an essential role in mitochondrial clearance. Indeed, this notion is in keeping with data showing that Parkin ubiquitinates OMM proteins, followed by their degradation by the UPS [12–14]. However, in contrast to larger OMM proteins, which are degraded exclusively by the UPS, smaller OMM proteins are degraded by both the UPS and the lysosomal system. This observation can be explained by a previously proposed model in which degradation of some (larger) OMM proteins destabilizes the OMM, eventually resulting in its fragmentation [13]. This mechanism is further supported by the fact that (i) uncouplers-induced mitochondrial depolarization is associated with a considerable increase in mitochondrial volume (by 65–140%) without change in surface area of the membranes [16] and (ii) the surface of the IMM is approximately 5-fold larger than that of the OMM, which additionally increases pressure and tension on the already destabilized OMM. In the next step, those OMM fragments that carry partially ubiquitinated proteins, are further degraded by the lysosomes. In contrast, IMM proteins and those of the mitochondrial matrix are almost exclusively degraded through the lysosomes. However, their degradation still depends on a functional UPS system.
Interestingly, the kinetics of Valinomycin-induced degradation of HSP60 and SOD2 differed from those of all other mitochondrial proteins analyzed in that their levels remained unchanged after even up to 24 h of treatment. This is consistent with a previous study showing that not only HSP60 and SOD2 but also another mitochondrial matrix protein, i.e., ATP synthase F1-beta-subunit (F1β), is not affected by mitochondrial depolarization [12]. In case of autophagy (mitophagy), the degradation rate of all IMM and mitochondrial matrix proteins would be expected to be more or less comparable since the content of the autophagosomes is non-selectively degraded via acidic lysosomal hydrolases [28]. On the other hand, this does not necessarily negate a role of autophagy in this process, as it has been shown before that some proteins, including SOD2, are partially resistant to lysosomal proteases [36].
Finally, the experiments using autophagy-incompetent cells, i.e., ATG7KO cells showed that removal of damaged mitochondria starts independently of autophagy. Most likely, autophagy serves only to remove debris of mitochondria that are previously identified by PINK1 and partially lysed by the UPS. Therefore, future studies should be focused on identifying and targeting a mechanism that can complement the impaired PINK1/Parkin/UPS machinery rather than boosting autophagy.
Our data collectively suggest that PINK1-/Parkin-dependent removal of damaged mitochondria can be partially accomplished via a mechanism other than macroautophagy. This mechanism, however, requires both a functional proteasome and lysosomal system. It is tempting to speculate that the uncoupler-initiated, UPS-mediated destabilization of mitochondrial membranes leads to their disruption, followed by the lysosomal-mediated removal of mitochondrial debris (Fig. 10). Taking the above-mentioned into account, “UPS-initiated, lysosome-executed mitolysis” would be a mechanism that takes place in parallel with autophagy. A recent study suggested that an alternative, MAPK1- and MAPK14-mediated form of autophagy induced by starvation or hypoxia is involved in mitophagy [35]. However, as the authors pointed out in their study, starvation or hypoxia did not decrease the levels of several mitochondrial proteins studied, indicating that only a small fraction of the mitochondrial pool was degraded. In contrast, treatment with CCCP/FCCP or Valinomycin results in efficient, complete loss of mitochondria. Furthermore, their study relied mainly on one mitochondrial marker, i.e., mt-Keima. Here, we demonstrate that a comprehensive investigation of mitochondrial clearance requires the simultaneous analysis of several mitochondrial proteins localized in all three mitochondrial compartments, as they are differentially degraded through UPS- and lysosome-mediated proteolysis. Finally, although initially suggested as a suitable alternative to CCCP, Antimycin A and Oligomycin were able to depolymerize mitochondria without activation of macroautophagy [25]. However, unlike Valinomycin, they failed to induce mitochondrial clearance [35].
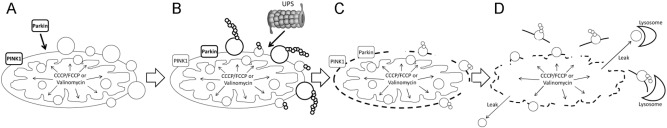
Fig. 10.
Putative model of PINK1-/Parkin-dependent removal of depolarized mitochondria. a Valinomycin (or CCCP/FCCP)-mediated mitochondrial depolarization initiates mitochondrial accumulation of PINK1 and mitochondrial translocation of Parkin. In parallel, both Valinomycin and CCCP/FCCP increase intramitochondrial pressure. b Parkin-mediated ubiquitination of outer mitochondrial membrane (OMM) proteins and UPS-mediated degradation of (large) OMM proteins. c UPS-mediated proteolysis of large OMM, increased intramitochondrial pressure and intrinsic pressure of IMM on the OMM leads to its destabilization and rupture. d Prolonged Valinomycin- or CCCP/FCCP-induced increase of the intramitochondrial pressure and lack of OMM destabilize the IMM and lead to its rupture. Mitochondrial debris is removed by lysosome-mediated proteolysis and lipolysis
Our findings thus warrant further studies using Valinomycin as a mitochondrial stressor to (i) investigate the exact role of the UPS in mitophagy, (ii) further dissect and elucidate the important mechanism of (impaired) removal of dysfunctional mitochondria in the pathophysiology of PD, and (iii) to conclusively clarify the present controversy regarding mitochondrial clearance in the context of PINK1- and Parkin-based models.
Materials and methods
Cell culture
SH-SY5Y cells and human dermal fibroblasts from a healthy control and a PD patient harboring a homozygous PINK1 missense mutation (c.509T>G; p.V170G) were grown at 37 °C under a 5% CO2 humidified atmosphere in Dulbecco’s modified Eagle medium (DMEM, PAA) supplemented with 10% FBS (PAA) and 1% penicillin/streptomycin (PAA). Cells used in the present studies were between passages 4 and 12.
CRISPR/Cas9-mediated generation of PINK1 and ATG7 knockout cells
PINK1 and ATG7 knockout SH-SY5Y cells were generated using an RNA-guided CRISPR/Cas9 endonuclease. For this, SH-SY5Y cells were transiently transfected with episomal vectors expressing both a human codon-optimized Cas9 and a guide RNA (gRNA) containing a 19-base long sequence that matches the human PINK1 target sequence 5′-TGTCCGCGGGGAGCGTCC-3′. To knockout ATG7, a gRNAs containing a 19-base long sequence that matches the human ATG7 target sequence 5′-GGGTTATTACTACAATGGT-3′ was used. Upon transfection, cells were resuspended in growth medium, counted and plated onto Petri dishes at a density of 1 cell/cm2. Cells were grown until they formed distinct, monoclonal colonies. The colonies were scraped off, transferred into different wells of a 6-well plate, and propagated to obtain enough material for the DNA extraction.
Antibodies
Proteins were separated by SDS polyacrylamide gel electrophoresis (SDS PAGE) and detected by western blot analysis using the following antibodies: anti-β-actin (Sigma), anti-ATG5 (Cell Signaling), anti-ATG7 (Cell Signaling), anti-Complex II Fp subunit (Mitosciences), F1F0ATPase (α subunit) (Mitosciences), anti-GAPDH (Cell Signaling), anti-GRP75 (Abcam), anti-HSP60 (Abcam), anti-MT-CO2 (Mitosciences), anti-LC3 (Cell Signaling), anti-MITOFUSIN 2 (Abcam), anti-Parkin (Cell Signaling), anti-PINK1 (Cell Signaling), anti-SOD2 (Santa Cruz), anti-Transcription factor A, mitochondrial (TFAM) (Abcam), anti-TOM20 (Santa Cruz), anti-TOM40 (Santa Cruz), anti-TOM70 (Abcam), and anti-VDAC1 (Abcam).
To quantify band intensities of immunoblots, the TotalLab TL100 v2006 1D-gel analysis software (Nonlinear Dynamics) was used.
Immunofluorescence
Fibroblasts and differentiated neurons stably expressing Parkin or/and MitoDsRed were grown on glass coverslips, fixed in 4% formaldehyde for 15 min, permeabilized with 0.1% Triton X-100 and blocked in 4% normal goat serum in PBS for 1 h. Immunofluorescence staining was performed using primary antibodies against Parkin (1:200; Cell Signalling), TOM20 (1:400; Santa Cruz Biotechnology), and GRP75 (1:400; Abcam). Appropriate secondary antibodies were obtained from Santa Cruz.
Cell viability analysis
Cell viability was evaluated with the trypan blue exclusion assay. For this, cells were treated with various concentrations of either Valinomycin or FCCP for 16 h. Both detached and adherent cells were collected by accutase (Thermo Fisher Scientific) and centrifuged at 300g for 5 min. Resulting cell pellets were resuspended in 1× PBS and mixed with 0.4% trypan. The ratio between dead and live cells was measured using an automated cell counter (Countess, Thermo Fisher Scientific).
Lentiviral particles expressing wildtype Parkin or small hairpin RNA against LC3s, GABARAP, or GABARAPL2 mRNA
A lentiviral plasmid overexpressing Parkin (pER4-Parkin) has been described previously [37]. Lentiviral particles expressing a small hairpin RNA against LC3 mRNAs were generated using pLKO.1 puro vector (a gift from Bob Weinberg (Addgene plasmid # 8453)) [38]. The following sequences in LC3A and LC3B mRNAs were targeted: in LC3A 5′-GCGAGUUGGUCAAGAUCAUCC-3′, in LC3B 5′-GCUUACAGCUCAAUGCUAAUC-3′. To knock down GABARAP and GABARAPL2, the following sequences were targeted using lentiviral particles expressing shRNAs: in GABARAP 5′-GCCUACAGUGACGAAAGUGUC-3′, in GABARAPL2 5′-GCGAAGAUUCGAGCGAAAUAU-3′.
Transmission electron microscopy
After treatment, cells were scraped off, washed in 1× PBS, centrifuged at 300g, and the supernatant was removed. Cells were resuspended in Monti’s fixative (2% glutardialdehyde, 0.6% paraformaldehyde, and 0.03% CaCl2 in 0.06 M cacodylate buffer, pH 7.35) for 1 h, centrifuged and kept in fixative for at least 24 h. The pellet was washed in 0.1 M cacodylate buffer pH 7.35 and subsequently treated for 2 h with 2% osmium tetroxide in water. After washing in cacodylate buffer, the pellet was dehydrated with increasing concentrations of ethanol, transferred to propylene oxide followed by mixtures of propylene oxide and araldite (2+1 and 1+2), embedded in araldite and polymerized at 60 °C for 48 h. Thin sections were cut (approximately 70 nm thick) using an ultramicrotome (Ultracut E, Leica, Bensheim, Germany) and transferred to 200 mesh nickel grids (Plano, Wetzlar, Germany). Sections were stained with 0.5% uranyl acetate followed by 3% lead citrate using an automated grid stainer (Ultrastainer, Leica). Sections were evaluated using a Jeol 1011 transmission electron microscope (Jeol, Freising, Germany) and documented with a Morada TEM Camera (Olympus, Hamburg, Germany).
Acknowledgements
This work was supported by the DFG (KL-1134/11-1, FOR2488 (P2 to C.K. and P3 to A.R.)), the BMBF (MitoPD), and EU (SysMedPD). C.K. is a recipient of a career development award from the Hermann and Lilly Schilling Foundation. We would like to thank Dr. Richard Youle for critical reading and his valuable comments and suggestions. We would like to thank Mr. Harry Manfeldt (Institute of Anatomy, University of Lübeck, Germany) for preparing samples for electron microscopy.
Compliance with ethical standards
Conflict of interest
The authors declare that they have no conflict of interest.
Footnotes
Edited by H. Zhang
References
- 1.Kasten M, et al. Genotype–phenotype relations for the Parkinson’s disease genes Parkin, PINK1, DJ1: MDSGene systematic review. Mov Disord. 2018;33:730–41. doi: 10.1002/mds.27352. [DOI] [PubMed] [Google Scholar]
- 2.Clark IE, et al. Drosophila PINK1 is required for mitochondrial function and interacts genetically with Parkin. Nature. 2006;441:1162–6. doi: 10.1038/nature04779. [DOI] [PubMed] [Google Scholar]
- 3.Exner N, et al. Loss-of-function of human PINK1 results in mitochondrial pathology and can be rescued by Parkin. J Neurosci. 2007;27:12413–8. doi: 10.1523/JNEUROSCI.0719-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Park J, et al. Mitochondrial dysfunction in Drosophila PINK1 mutants is complemented by Parkin. Nature. 2006;441:1157–61. doi: 10.1038/nature04788. [DOI] [PubMed] [Google Scholar]
- 5.Rakovic A, et al. Phosphatase and tensin homolog (PTEN)-induced putative kinase 1 (PINK1)-dependent ubiquitination of endogenous Parkin attenuates mitophagy: study in human primary fibroblasts and induced pluripotent stem cell-derived neurons. J Biol Chem. 2013;288:2223–37. doi: 10.1074/jbc.M112.391680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Geisler S, et al. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat Cell Biol. 2010;12:119–31. doi: 10.1038/ncb2012. [DOI] [PubMed] [Google Scholar]
- 7.Ding WX, et al. Nix is critical to two distinct phases of mitophagy, reactive oxygen species-mediated autophagy induction and Parkin-ubiquitin-p62-mediated mitochondrial priming. J Biol Chem. 2010;285:27879–90. doi: 10.1074/jbc.M110.119537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Narendra DP, et al. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol. 2010;8:e1000298. doi: 10.1371/journal.pbio.1000298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Vives-Bauza C, et al. PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc Natl Acad Sci USA. 2010;107:378–83. doi: 10.1073/pnas.0911187107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Narendra D, Kane LA, Hauser DN, Fearnley IM, Youle RJ. p62/SQSTM1 is required for Parkin-induced mitochondrial clustering but not mitophagy; VDAC1 is dispensable for both. Autophagy. 2010;6:1090–106. doi: 10.4161/auto.6.8.13426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Okatsu K, et al. p62/SQSTM1 cooperates with Parkin for perinuclear clustering of depolarized mitochondria. Genes Cells. 2010;15:887–900. doi: 10.1111/j.1365-2443.2010.01426.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chan NC, et al. Broad activation of the ubiquitin-proteasome system by Parkin is critical for mitophagy. Hum Mol Genet. 2011;20:1726–37. doi: 10.1093/hmg/ddr048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yoshii SR, Kishi C, Ishihara N, Mizushima N. Parkin mediates proteasome-dependent protein degradation and rupture of the outer mitochondrial membrane. J Biol Chem. 2011;286:19630–40. doi: 10.1074/jbc.M110.209338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tanaka A, et al. Proteasome and p97 mediate mitophagy and degradation of mitofusins induced by Parkin. J Cell Biol. 2010;191:1367–80. doi: 10.1083/jcb.201007013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Rakovic A, et al. Mutations in PINK1 and Parkin impair ubiquitination of Mitofusins in human fibroblasts. PLoS One. 2011;6:e16746. doi: 10.1371/journal.pone.0016746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Safiulina D, Veksler V, Zharkovsky A, Kaasik A. Loss of mitochondrial membrane potential is associated with increase in mitochondrial volume: physiological role in neurones. J Cell Physiol. 2006;206:347–53. doi: 10.1002/jcp.20476. [DOI] [PubMed] [Google Scholar]
- 17.Smith TA, Blaylock MG. Treatment of breast tumor cells in vitro with the mitochondrial membrane potential dissipater valinomycin increases 18F-FDG incorporation. J Nucl Med. 2007;48:1308–12. doi: 10.2967/jnumed.107.041665. [DOI] [PubMed] [Google Scholar]
- 18.Felber SM, Brand MD. Valinomycin can depolarize mitochondria in intact lymphocytes without increasing plasma membrane potassium fluxes. FEBS Lett. 1982;150:122–4. doi: 10.1016/0014-5793(82)81317-3. [DOI] [PubMed] [Google Scholar]
- 19.Joshi DC, Bakowska JC. Determination of mitochondrial membrane potential and reactive oxygen species in live rat cortical neurons. J Vis Exp. 2011 doi: 10.3791/27042704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Rakovic A, et al. Effect of endogenous mutant and wild-type PINK1 on Parkin in fibroblasts from Parkinson disease patients. Hum Mol Genet. 2010;19:3124–37. doi: 10.1093/hmg/ddq215. [DOI] [PubMed] [Google Scholar]
- 21.Steinberg BE, et al. A cation counterflux supports lysosomal acidification. J Cell Biol. 2010;189:1171–86. doi: 10.1083/jcb.200911083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ishida Y, Nayak S, Mindell JA, Grabe M. A model of lysosomal pH regulation. J Gen Physiol. 2013;141:705–20. doi: 10.1085/jgp.201210930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Park KS, et al. FCCP depolarizes plasma membrane potential by activating proton and Na+ currents in bovine aortic endothelial cells. Pflug Arch. 2002;443:344–52. doi: 10.1007/s004240100703. [DOI] [PubMed] [Google Scholar]
- 24.Nicholls DG. Simultaneous monitoring of ionophore- and inhibitor-mediated plasma and mitochondrial membrane potential changes in cultured neurons. J Biol Chem. 2006;281:14864–74. doi: 10.1074/jbc.M510916200. [DOI] [PubMed] [Google Scholar]
- 25.Padman BS, Bach M, Lucarelli G, Prescott M, Ramm G. The protonophore CCCP interferes with lysosomal degradation of autophagic cargo in yeast and mammalian cells. Autophagy. 2013;9:1862–75. doi: 10.4161/auto.26557. [DOI] [PubMed] [Google Scholar]
- 26.Gegg ME, et al. Mitofusin 1 and Mitofusin 2 are ubiquitinated in a PINK1/Parkin-dependent manner upon induction of mitophagy. Hum Mol Genet. 2010;19:4861–70. doi: 10.1093/hmg/ddq419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Mizushima N, Yoshimori T. How to interpret LC3 immunoblotting. Autophagy. 2007;3:542–5. doi: 10.4161/auto.4600. [DOI] [PubMed] [Google Scholar]
- 28.Feng Y, He D, Yao Z, Klionsky DJ. The machinery of macroautophagy. Cell Res. 2014;24:24–41. doi: 10.1038/cr.2013.168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Koukourakis MI, et al. Autophagosome proteins LC3A, LC3B and LC3C have distinct subcellular distribution kinetics and expression in cancer cell lines. PLoS One. 2015;10:e0137675. doi: 10.1371/journal.pone.0137675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Novak I, et al. Nix is a selective autophagy receptor for mitochondrial clearance. EMBO Rep. 2010;11:45–51. doi: 10.1038/embor.2009.256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Vaites LP, Paulo JA, Huttlin EL, Harper JW. Systematic analysis of human cells lacking ATG8 proteins uncovers roles for GABARAPs and the CCZ1/MON1 regulator C18orf8/RMC1 in macro and selective autophagic flux. Mol Cell Biol. 2017 doi: 10.1128/MCB.00392-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lazarou M, et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature. 2015;524:309–14. doi: 10.1038/nature14893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tanida I, Yamasaki M, Komatsu M, Ueno T. The FAP motif within human ATG7, an autophagy-related E1-like enzyme, is essential for the E2-substrate reaction of LC3 lipidation. Autophagy. 2012;8:88–97. doi: 10.4161/auto.8.1.18339. [DOI] [PubMed] [Google Scholar]
- 34.Narendra D, Tanaka A, Suen DF, Youle RJ. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol. 2008;183:795–803. doi: 10.1083/jcb.200809125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hirota Y, et al. Mitophagy is primarily due to alternative autophagy and requires the MAPK1 and MAPK14 signaling pathways. Autophagy. 2015;11:332–43. doi: 10.1080/15548627.2015.1023047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rabouille C, Strous GJ, Crapo JD, Geuze HJ, Slot JW. The differential degradation of two cytosolic proteins as a tool to monitor autophagy in hepatocytes by immunocytochemistry. J Cell Biol. 1993;120:897–908. doi: 10.1083/jcb.120.4.897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Seibler P, et al. Mitochondrial Parkin recruitment is impaired in neurons derived from mutant PINK1 induced pluripotent stem cells. J Neurosci. 2011;31:5970–6. doi: 10.1523/JNEUROSCI.4441-10.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Stewart SA, et al. Lentivirus-delivered stable gene silencing by RNAi in primary cells. RNA. 2003;9:493–501. doi: 10.1261/rna.2192803. [DOI] [PMC free article] [PubMed] [Google Scholar]