Abstract
Objective
Clinical data of patients’ measurements and treatment history stored in electronic health record (EHR) systems are starting to be mined for better treatment options and disease associations. A primary challenge associated with utilizing EHR data is the considerable amount of missing data. Failure to address this issue can introduce significant bias in EHR-based research. Currently, imputation methods rely on correlations among the structured phenotype variables in the EHR. However, genetic studies have shown that many EHR-based phenotypes have a heritable component, suggesting that measured genetic variants might be useful for imputing missing data. In this article, we developed a computational model that incorporates patients’ genetic information to perform EHR data imputation.
Materials and Methods
We used the individual single nucleotide polymorphism’s association with phenotype variables in the EHR as input to construct a genetic risk score that quantifies the genetic contribution to the phenotype. Multiple approaches to constructing the genetic risk score were evaluated for optimal performance. The genetic score, along with phenotype correlation, is then used as a predictor to impute the missing values.
Results
To demonstrate the method performance, we applied our model to impute missing cardiovascular related measurements including low-density lipoprotein, heart failure, and aortic aneurysm disease in the electronic Medical Records and Genomics data. The integration method improved imputation's area-under-the-curve for binary phenotypes and decreased root-mean-square error for continuous phenotypes.
Conclusion
Compared with standard imputation approaches, incorporating genetic information offers a novel approach that can utilize more of the EHR data for better performance in missing data imputation.
Keywords: electronic health record, imputation, genetic risk score, missing data, single nucleotide polymorphisms
INTRODUCTION
Electronic health record (EHR) data present a wealth of information for biomedical knowledge discovery on an unprecedented scale.1 However, EHR data present significant challenges for research use as they have been collected for clinical and billing purposes.2 As a result, a significant amount of EHR data are missing due to, among other factors, the financial burden of testing and diagnostics,3 underdiagnoses,4 and differences in methods in classifying disease phenotypes.5 Failure to account for the missing data can reduce power to detect true signals from the data and can have a significant effect on the research conclusions.6
Imputing structured EHR data (eg, quantifiable measurements) has been explored previously.2,7 The most common approach to handle missing EHR data is complete case analysis, where the missing samples are omitted during analysis. However, deleting samples will greatly reduce power and may not be a viable option if multiple variables are involved. Imputations based on the variable distributions (eg, mean, median) of the existing data are also widely used, nevertheless, they generally result in the same estimate as the complete case analysis.2 Statistical approaches that consider the correlation among the clinical variables have also been utilized. Most notably, multiple imputation using chained equations (MICE) can impute different variable types by estimating the posterior distribution of each variable by regressing it on all other variables.8 Machine learning approaches that use dimension reduction,9 similarity,9 and network modeling10 have also been applied for missing data imputation. However, to the best of our knowledge, all current imputation methods applied to EHR data use the distribution and correlations among the clinical variables for imputing the missing values.
In the past decade, genome-wide association studies (GWAS) have identified numerous genetic variants that are associated with human traits.11,12 Whereas GWAS focuses on identifying genotype–phenotype associations through statistical hypothesis testing, some of the discovered genetic variants are predictive of the phenotype as well. As a result, much of the research has examined and validated the predictive ability of genetic data for various phenotypes. Single nucleotide polymorphism (SNP) based prediction has been shown to reach 80% area-under-the-curve prediction for lifetime Alzheimer’s disease.13 Thousands of common alleles of small effects have been combined to capture the genetic risk of bipolar disorder.14 Utilizing genotyping information on 10–30 SNPs have been demonstrated to improve the prediction of breast cancer in women.15,16 In addition, as the cost of genotyping and sequencing continues to decrease, an increasing number of EHRs have linked genetic data available for hundreds of thousands of people.17–19 To this end, we propose an integrative approach to incorporate both clinical and genetic variables to impute missing values in structured EHR data.
In this study, we evaluated our integrative imputation approach on several cardiovascular-related phenotypes in the electronic Medical Records and Genomics (eMERGE) EHR data.20 Imputation on binary disease diagnosis, heart failure (HF), and aortic aneurysm disease (AAA), and continuous measurement, low-density lipoprotein (LDL) showed that incorporating genetic information improved imputation accuracy compared to methods that omit this information. We also applied the method to impute missing labels for a HF data set and found improved power to detect previously identified HF-associated SNPs. Compared to existing imputation methods, our method is the first EHR-specific data imputation method that integrates patients’ genetic information.
MATERIALS AND METHODS
eMERGE EHR data
All patients’ clinical and genetic data were obtained from the electronic medical records and genomics network (dbGaP accession: phs000888.v1.p1). Within eMERGE, patients’ high-density lipoprotein (HDL, mg/dL), low-density lipoprotein (LDL, mg/dL), and recorded ages were obtained from the Geisinger_AAA_Labs data set. For binary phenotype, patients’ HF and AAA disease statuses along with their gender were obtained from 3 different consent groups: Health/Medical/Biomedical (HMB), Health/Medical/Biomedical - Genetic Studies Only-No Insurance Companies (HM_B_GSO_NIC), and Health/Medical/Biomedical (GSO) (HMB_GSO). The 3 consent groups contain nonoverlapping patients from 9 different EHRs: Children's Hospital of Pennsylvania, Cincinnati Children's Hospital Medical Center/Boston's Children's Hospital, Geisinger Health System, Group Health/University of Washington, Essentia Institute of Rural Health, Marshfield Clinic, Pennsylvania State University (Marshfield), Mayo Clinic, Icahn School of Medicine at Mount Sinai School, Northwestern University, and Vanderbilt University. There were multiple categories of disease status for the phenotypes including case, control, and neither case nor control (Supplementary File 1). We only retained patients that have either case or control status. SNP genotyping was performed using the Illumina 660W-Quad BeadChip at the Center for Genotyping and Analysis at the Broad Institute, Cambridge, MA. Whole genome imputation based on the genotyped SNPs were performed by eMERGE according to the standard pipeline.21
Quality control
A patient could have multiple HDL or LDL measurements at different ages in their clinical records. Due to the unequal number of recorded measurements per patient, the median value of HDL and LDL were used. The ages associated with the median value of HDL, Age(HDL), and the median value of LDL, Age(LDL), showed high concordance by the linear regression analysis:
Thus, the mean value of the 2 median ages was calculated as the patient’s age. To avoid extreme values, HDL and LDL values that lie 4 standard deviations from the respected means were removed. SNP genotyping data were filtered to satisfy the following criteria: missing rate < 5%, minor allele frequency > 1%, and Hardy–Weinberg equilibrium < 0.00001. 38 040 165 SNP genotypes passed QC and were used for the subsequent analysis.
Cross-validation
To test the validity of the method, we only retained patients with complete clinical records. If more than 1 variable was used in the model, we kept only pairwise complete records. We randomly selected 50%, 70%, or 90% of patients for training the model and the remaining 50%, 30%, or 10% of patients for testing the model. The model performance was determined by comparing the predicted value on the testing data to the actual patients’ record. For the continuous phenotype, the performance metric used was the root-mean-square error (RMSE):
Where n is the sample size of the testing data, and are the predicted and actual phenotype value for patient i, respectively. For the binary variable, we used the area under the curve (AUC) to evaluate performance on the testing data. Cross-validation was performed 100 times to assess the consistency of the results (Figure 1).
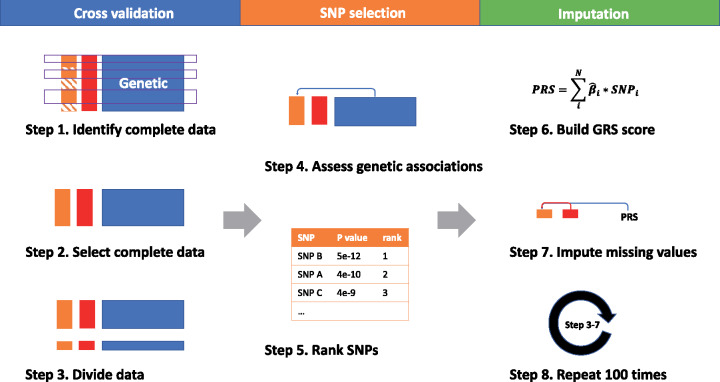
Figure 1.
Overview of the imputation model. Complete data were used to assess each SNP’s association to the phenotype (Steps 1–5). A GRS is then used to summarize multiple SNPs based on their associations (Step 6). The GRS as well as other clinical variables are then used to impute the missing values (Step 7). The variability of the imputation is assessed using 100 different cross-validations (Step 8).
Candidate SNPs
For a phenotype, each SNP genotype’s marginal association with the phenotype was determined using logistic regression or linear regression implemented in Plink.22 Previous research has shown that certain phenotypes may be associated with up to thousands of SNPs.23–26 Thus, we selected the SNPs whose P value are less than .005 to capture all of the candidate SNPs (Figure 1).
Genetic risk score
We reduced the high dimensional SNPs genotyping data using the genetic risk score (GRS) developed and applied in population genetics.27,28 Within each cross-validation, the candidate SNPs were first LD pruned using Plink22 with the parameter (indep -50 5 2) and then re-evaluated for their associations to the phenotype in the training data. Each SNP’s , P value, (continuous phenotype) and AUC (binary phenotype) were obtained using the regression models. A SNP’s value was calculated as:
Where n is the number of patients, is the predicted value of the phenotype for individual i and is the mean value of the phenotype. A SNP’s AUC was calculated by ROCR package.29 To calculate an AUC, the fitted probability (p) of a sample being a case was obtained from the logistic regression. The fitted probability was then compared to a threshold cutoff, c, to classify a patient as a case (P > c) or a control (P < c). The sensitivity and specificity of the classification can be calculated for a particular cutoff c. By varying c from 0 to 1, we were able to plot sensitivity against (1-specificity), which is commonly known as the receiver operating characteristic (ROC) curve. The area under the ROC curve was calculated as AUC.
Using P value, , or AUC, the SNPs were ranked by their association with respect to the phenotype. For all ranking criterion, the GRS for the top N SNPs in the training data was calculated as:
where is the effect size of individual SNPi and is the ’s minor allele count in the training data. The GRS for the testing data was similarly calculated as:
Here we used obtained from the training data and from the testing data (Figure 1).
Imputation model
Imputation model was trained on the training data, and the missing values were imputed on the testing data where the true values were known.
For HF and AAA phenotypes, a logistic regression model was first trained on the training data to obtain regression coefficients:
Then, imputation on the testing data was carried out by using these coefficients:
Similarly, imputation for LDL was performed using the following linear regression models:
Comparison with other methods
We compared several imputation models that do not incorporate genetic information. First, we compared to a baseline model using the diseases, probabilities, or phenotype distributions in the training data.
Then, we compared the imputation model with only clinical variables
Finally, we imputed the missing values using the MICE package with default settings.30 For the binary variable, we included gender as a covariate; and for LDL measurement, we included gender, HDL, and age.
Validation on previously reported HF GWAS associations
Previously reported SNPs associated with HF were downloaded from NHGRI-EBI catalog (https://www.ebi.ac.uk/gwas/).31,32 The 17 SNPs were re-analyzed for their associations in the HMB consent group data set, which has the most balanced diagnosis labels. Associations were performed on half of the data where the labels are known and again on the full data after imputing the missing label. We recorded the P values of the association from the incomplete data and complete data using GRSpval or GRSauc imputation. We retained the SNPs that showed genome-wide Bonferroni corrected significance (P < 10−8) in any of the settings.
RESULTS
We selected 2 cardiovascular related binary phenotypes (AAA and HF) from the eMERGE data. Within eMERGE, disease status was obtained from 3 different consent groups: HMB, HMB_GSO, and HM_B_GSO_NIC. There were about 22 000 AAA and 13 000 HF patients included in the study (Supplementary File 1).
Continuous phenotypes including patients’ HDL, LDL, and age were obtained from eMERGE’s Geisinger lab data set. After quality controls, 12 752 patients were kept for the subsequent analysis. HDL and LDL measurements appear to be normally distributed. Patients’ age is slightly left-skewed and resembles an older population (Supplementary File 2).
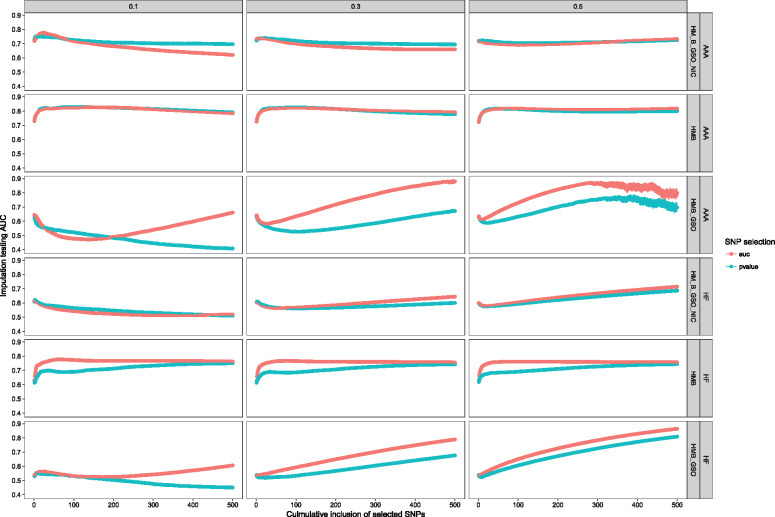
To assess the contribution of genetic information to imputing binary and continuous EHR clinical variables, we required known values to be compared with imputed values. Thus, 10%, 30% or 50% of clinical outcomes were made unavailable during the training stage and later compared with the imputed values (Figure 2, vertical panels). We performed a separate analysis for each disease (AAA and HF) and consent groups (HM_B_GSO_NIC, HMB, HMB_GSO) to ensure homogeneity within each patient group (Figure 2, horizontal panels). Here 2 different criteria for selecting SNPs (Figure 2, colored bands) as well as the number of included SNPs on the imputation performance were also evaluated. Generally, imputation accuracy improved as more SNPs were included in the model. However, the accuracy reached plateaus at different rates. For example, imputing AAA in HMB consent group achieved the highest accuracy with around 50 SNPs included in the GRS. On the contrary, phenotypes in HMB_GSO required more than 500 SNPs to achieve a stable accuracy. SNPs selection criteria also affected the imputation performance. Using AUC to select SNPs to construct GRS outperformed P value in almost all evaluations. The degree of improvement varied across different data sets.
Figure 2.
Impact of incorporating genetic information on imputation accuracy of AAA and HF. The 3 vertical panels indicate different percentages of missing data (10%, 30%, and 50%). Horizontal panels show the 6 different disease and consent group combinations. The red color band represents accuracies using SNPs selected by AUC from 100 repetitions. The green color band represents P value selection. From left to right, the x-axis represents GRSs calculated from increasing number of SNPs, eg, SNP(1), SNP(1, 2), and SNP(1, 2, 3… 500). The y-axis shows the imputation AUC on the testing data.
We also compared several imputation methods that do not utilize genetic information. Overall, GRS-based imputation achieved the best performance in most data sets. The improved accuracies compared with other models reflect the added prediction due to the genetic data (Table 1). For the analysis that used 10% of patients as testing data, GRS showed slightly less efficient performance that could be caused by the low prevalence of the cases (Supplementary File 1).
Table 1.
Comparison of imputation models on AAA and HF. MICE, disease probability, and regression model use only clinical variables for imputation. GRS P value and GRS auc represent model contains P value or AUC selected GRS constructed from the top 500 SNPs. Cell values are AUCs on the testing data and their standard deviations (in parenthesis) in 100 cross-validations. The best accuracy in each column is in bold
| Missing percentage = 10% |
||||||
|---|---|---|---|---|---|---|
| AAA |
HF |
|||||
| HM_B_GSO_NIC | HMB | HMB_GSO | HM_B_GSO_NIC | HMB | HMB_GSO | |
| MICE(gender) | 0.5 (0.02) | 0.54 (0.03) | 0.51 (0.03) | 0.51 (0.02) | 0.51 (0.02) | 0.51 (0.03) |
| Disease probability | 0.5 (0.01) | 0.5 (0.03) | 0.5 (0.02) | 0.5 (0.02) | 0.5 (0.02) | 0.5 (0.03) |
| Regression | 0.72 (0.06) | 0.67 (0.03) | 0.64 (0.04) | 0.6 (0.02) | 0.55 (0.02) | 0.55 (0.03) |
| GRS P value | 0.7 (0.05) | 0.79 (0.02) | 0.41 (0.06) | 0.51 (0.03) | 0.75 (0.02) | 0.45 (0.03) |
| GRS auc | 0.62 (0.05) | 0.79 (0.02) | 0.66 (0.05) | 0.52 (0.03) | 0.76 (0.02) | 0.61 (0.04) |
|
| ||||||
|
Missing percentage = 30% |
||||||
|
AAA |
HF |
|||||
| HM_B_GSO_NIC | HMB | HMB_GSO | HM_B_GSO_NIC | HMB | HMB_GSO | |
| MICE(gender) | 0.5 (0.01) | 0.55 (0.02) | 0.5 (0.01) | 0.51 (0.01) | 0.5 (0.01) | 0.5 (0.02) |
| Disease probability | 0.5 (0.01) | 0.5 (0.02) | 0.5 (0.01) | 0.5 (0.01) | 0.5 (0.01) | 0.5 (0.02) |
| Regression | 0.71 (0.03) | 0.67 (0.01) | 0.64 (0.02) | 0.6 (0.01) | 0.55 (0.01) | 0.54 (0.02) |
| GRS P value | 0.69 (0.03) | 0.78 (0.01) | 0.67 (0.05) | 0.6 (0.01) | 0.74 (0.01) | 0.68 (0.02) |
| GRS auc | 0.66 (0.03) | 0.79 (0.01) | 0.88 (0.06) | 0.64 (0.01) | 0.76 (0.01) | 0.79 (0.02) |
|
| ||||||
|
Missing percentage = 50% |
||||||
|
AAA |
HF |
|||||
| HM_B_GSO_NIC | HMB | HMB_GSO | HM_B_GSO_NIC | HMB | HMB_GSO | |
| Mice(gender) | 0.5 (0.01) | 0.55 (0.01) | 0.51 (0.01) | 0.51 (0.01) | 0.51 (0.01) | 0.5 (0.01) |
| Disease probability | 0.5 (0.01) | 0.5 (0.01) | 0.5 (0.01) | 0.5 (0.01) | 0.5 (0.01) | 0.5 (0.01) |
| Regression | 0.71 (0.02) | 0.68 (0.01) | 0.64 (0.01) | 0.6 (0.01) | 0.55 (0.01) | 0.55 (0.01) |
| GRS P value | 0.72 (0.03) | 0.8 (0.01) | 0.7 (0.16) | 0.69 (0.01) | 0.74 (0.01) | 0.81 (0.01) |
| GRS auc | 0.73 (0.03) | 0.82 (0.01) | 0.8 (0.15) | 0.71 (0.01) | 0.76 (0.01) | 0.86 (0.01) |
data set
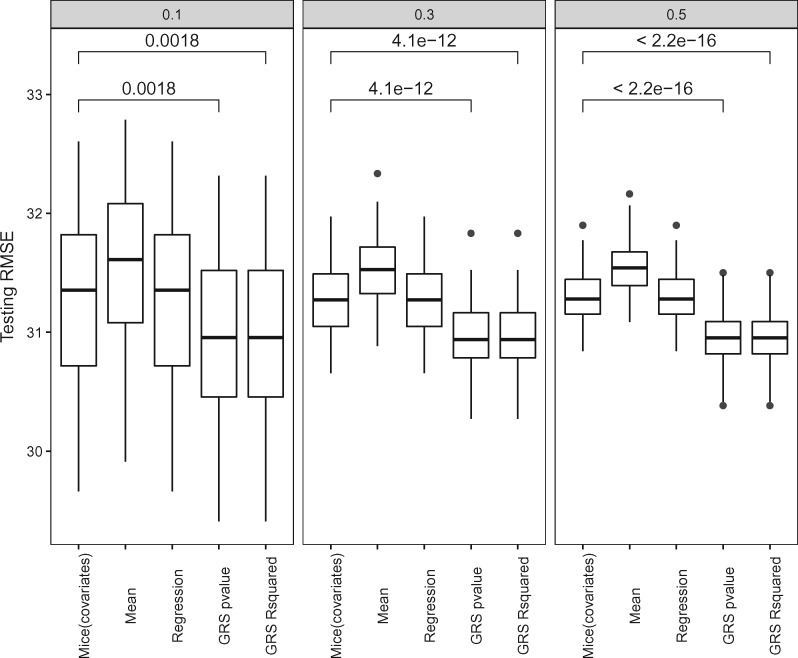
Similarly, we evaluated imputation of LDL using genetic information on the eMERGE Geisinger data set. Across different percentage of missing LDL, GRS composed of selected SNPs decreased similar RMSE compared with that of P value (Figure 3). The mean value of LDL was used as the baseline model for comparison. HDL, LDL, and age were used as predictors in MICE using the norm.predict function and linear regression model as comparisons for the imputation model that account for correlations between clinical variables. Models that use GRS achieved the lowest RMSE in imputing the missing values (Figure 3). The amount of variance explained by GRS is shown in the Supplementary File 3.
Figure 3.
Comparison of imputation models on LDL. Vertical panels show different percentages of missing data. GRS P value and GRS R2 consist of top 500 SNPs selected by P value or r-squared, respectively. Statistical significances were obtained using t-test.
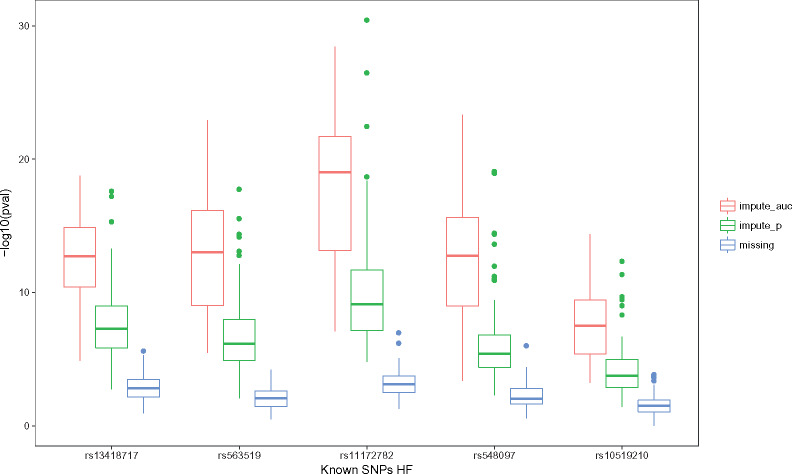
To demonstrate the power of imputation using genetic information, we performed association analyses of known HF SNPs with and without performing imputation. We obtained the 17 known SNPs from the NHGRI-EBI catalog (https://www.ebi.ac.uk/gwas/) and validated in the HMB consent group, which was the most balanced data set. In this data set, 5 SNPs showed significant association (P < 10−8) in any analysis. Imputing the missing HF label resulted in more significant SNP associations compared with omitting the samples with missing labels (Figure 4).
Figure 4.
Improved power on known HF associated SNPs. Five HF associated SNPs showed significance (P < 10−8, or equivalently -log10(p) > 8) in at least 1 setting. The box-plots show the p values associated with each of the 5 SNPs without imputation (missing), imputing using AUC selected SNPs (impute_auc), and imputing using P value selected SNPs (impute_P) over 100 repetitions.
DISCUSSION
Dealing with missing data is often the first challenge that many researchers face when conducting EHR-based research. For structured EHR clinical variables such as lab measurements and disease diagnosis, imputing the missing value often relies on the distributions and correlations among the clinical variables. In this study, we evaluated the added benefit of incorporating genetic information when performing imputation. We showed that for various common clinical variables, integrating genetic information greatly increased the imputation accuracy.
One challenge in utilizing genetic information in the imputation model is the high dimensionality of the data set. Because of this, typical machine learning and statistical methods cannot be directly used to incorporate the genetic data. Thus, we utilized the GRS to select and combine important genetic risk factors, in this case SNPs, to summarize the high dimensional genetic data. We evaluated multiple SNP selection criteria focusing on either the traditional significance test (P value) or the predictive property ( and AUC). The results showed that selecting SNPs based on their predictive abilities performed similar or better than the P value selection (Table 1 and Figure 3). This is likely due to selecting SNPs based on their predictive property, which can lead to a better predictive power of the GRS. Thus, for prediction purpose, GRS may be better constructed using AUC or criteria, rather than P value. The number of SNPs included in the GRS also affected the imputation accuracy (Figure 2). Generally, including more SNPs in the GRS would improve the prediction accuracy on the training data. To avoid this bias, we performed all model evaluations on the testing data, which does not guarantee improved accuracy when more SNPs are included. Increasing the number of SNPs improved the testing accuracy until a plateau was reached; however, the rates of reaching the plateaus were different for each data set, which likely reflects the heterogeneity of the underlying genetic associations within each data set. One concern with including a large number of variables in a model is the potential for overfitting. However, the notable observation here is that including noninformative SNPs in GRS generally did not degrade the imputation accuracy (Figure 2). This advantageous property could be due to the near-zero weight associated with noninformative SNPs in GRS calculation thus reducing their impact on the final score. There are several exceptions to this observation that occurred when the missing percentage is 10% and the SNPs were selected based on P values. As we have demonstrated, P values do not directly translate to predictive power, thus the GRS can perform poorly (AUC < 50%) on the testing data when adding noninformative SNPs. In addition, in some cases, the methods performed better when the percentage of missing data is 50% compared with that of 10%. Intuitively, the imputation efficiency should be similar between these 2 settings because the missing data were generated randomly. However, we believe that this can be attributed to the low case prevalence in some of the data sets. The combination of low case prevalence and a small percentage of samples (10%) for the testing data could lead to very few cases in the testing data. For data sets with a moderate case prevalence (HMB, Supplementary File 1), the method performed similarly across 10%, 30%, and 50% of missing data. Furthermore, all data sets achieved similar accuracies between 30% and 50% of missing data.
To demonstrate the added benefits of the genetic information, we compared the prediction performance of models with GRS and other clinical variables to those without using GRS. GRS-based imputation methods generally performed the best followed by the regression model (Table 1). Whereas the contribution of genetic data to imputation was consistent, its efficiency varied across consent groups. In the eMERGE EHR data, overall, around 80% of the patients are White and 20% are non-White. The differences in efficiency could be due to different genetic compositions in each consent group, as genetic associations are sensitive to population background. The sample size variations across consent groups could also affect the power to detect genetic associations (eg, HM_B_GSO_NIC is about 4 times larger than HMB in AAA (Supplementary File 1). As alluded to previously, different consent groups also have different case prevalences. For consent groups with a low number of diagnosed patients, their power to detect genetic association could be reduced.33
Identifying phenotype–genotype associations is one of the major knowledge discoveries being carried out in EHR data. Among factors such as underdiagnosis and insufficient phenotyping algorithms, many patients do not have the complete set of disease status. The missing disease labels limit the sample size available for identifying genetic association, which reduces the power to detect the true signals. For the 5 known HF associations that are replicated in our data set, imputing the missing labels using genetic information as well as demographic variables improved the power to detect SNP associations (Figure 4).
Integrating genetic information in EHR data imputation offers promising results, but there remain several limitations worthy of follow up studies. Genetic data provide valuable information on the average prediction of the disease status but do not account for the temporal changes of the phenotype. This is shown by the significantly improved power for imputing “stationary” binary HF and AAA phenotypes compared to the mild improvement in the “temporal” LDL levels. Using the improved EHR phenotypes to perform additional genetic association analysis could potentially introduce some bias because the genetic data have been used for imputation. However, the bias should be small for individual SNP associations because a GRS is calculated from hundreds or thousands of SNPs and each SNP’s contribution to a GRS is relatively small. EHR data can often have complex missing patterns including differential missingness and missing-not-at-random patterns. The impact of genetic data on missing patterns should be thoroughly explored in future studies. Furthermore, eMERGE EHR data have only released a limited number of clinical and demographic records of the patients. We used imputation accuracies from these variables as the baseline to show the added effect from GRS. As a result, our study only evaluated the additive effects between clinical variables and GRS on imputation accuracies. Nonlinear effects between clinical variables such as comorbidity, temporal medication history, and unstructured doctors’ notes have been used to predict EHR phenotypes.34,35 These effects could be integrated with genetic information to maximize imputation accuracy in other EHR data sets. Finally, the effectiveness of the proposed method depends on the availability of the EHR linked genetic data and the prevalence of the phenotypes. Whereas a growing number of EHRs are starting to have linked patients’ genetic data, the majority of EHRs do not yet have this type of information available.
CONCLUSION
Existing EHR imputation methods only take advantage of the patterns and structures found in the clinical variables. In this study, we demonstrated the utility of incorporating genetic data in EHR phenotype imputation. Using several continuous and binary EHR phenotype variables, we showed that incorporating genetic information through GRS significantly improved imputation accuracies. In addition, GRS calculated using prediction thresholds generally outperformed the P value threshold. Future research should consider investigating nonrandom missing data patterns and additional approaches to integrate clinical and genetic data. Nevertheless, our results showed the value to integrate informative genetic data in EHR data imputation.
FUNDING
This work was supported by National Institutes of Health grants AI116794, DK112217, ES013508, HL134015, LM010098, LM011360, LM012601, and TR001263.
AUTHOR CONTRIBUTIONS
RL, YC, and JHM conceived of the study. RL performed data processing and analyses. RL, YC, and JHM wrote the manuscript, and all authors revised and approved the final manuscript.
SUPPLEMENTARY MATERIAL
Supplementary material is available at Journal of the American Medical Informatics Association online.
CONFLICT OF INTEREST STATEMENT
None declared.
Supplementary Material
REFERENCES
- 1. Prokosch HU, Ganslandt T.. Perspectives for medical informatics. Reusing the electronic medical record for clinical research. Methods Inf Med 2009; 481: 38–44. [PubMed] [Google Scholar]
- 2. Wells BJ, Chagin KM, Nowacki AS, Kattan MW.. Strategies for handling missing data in electronic health record derived data. EGEMS (Washington, DC )2013; 13: 1035.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. McClatchey KD. Clinical Laboratory Medicine. Philadelphia, PA: Lippincott Wiliams & Wilkins; 2002: 1693. [Google Scholar]
- 4. Banerjee D, Chung S, Wong EC, Wang EJ, Stafford RS, Palaniappan LP.. Underdiagnosis of hypertension using electronic health records. Am J Hypertens 2012; 251: 97–102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Shivade C, Raghavan P, Fosler-Lussier E, et al. A review of approaches to identifying patient phenotype cohorts using electronic health records. J Am Med Inform Assoc 2014; 212: 221–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Graham JW. Missing data analysis: making it work in the real world. Annu Rev Psychol 2009; 601: 549–76. [DOI] [PubMed] [Google Scholar]
- 7. Beaulieu-Jones BK, Lavage DR, Snyder JW, Moore JH, Pendergrass SA, Bauer CR.. Characterizing and managing missing structured data in electronic health records: data analysis. JMIR Med Inform 2018; 61: e11.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. White IR, Royston P, Wood AM.. Multiple imputation using chained equations: issues and guidance for practice. Statist Med 2011; 304: 377–99. [DOI] [PubMed] [Google Scholar]
- 9. Troyanskaya O, Cantor M, Sherlock G, et al. Missing value estimation methods for DNA microarrays. Bioinformatics [Internet] 2001; 176: 520–5. [DOI] [PubMed] [Google Scholar]
- 10. Beaulieu-Jones BK, Moore JH, Consortium TPRO-AACT. Missing data imputation in the electronic health record using deeply learned autoencoders. Pac Symp Biocomput 2016; 22: 207–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Visscher PM, Wray NR, Zhang Q, et al. 10 Years of GWAS discovery: biology, function, and translation; 2017. https://www.cell.com/ajhg/pdf/S0002-9297(17)30240-9.pdf. Accessed May 24, 2018. [DOI] [PMC free article] [PubMed]
- 12. Donnelly P, Price AL, Spencer CCA. Progress and promise in understanding the genetic basis of common diseases. 10.1098/rspb.2015.1684. Accessed May 24, 2018. [DOI] [PMC free article] [PubMed]
- 13. Escott-Price V, Shoai M, Pither R, Williams J, Hardy J.. Polygenic score prediction captures nearly all common genetic risk for Alzheimer’s disease. Neurobiol Aging 2017; 49: 214.e7–.e11. [DOI] [PubMed] [Google Scholar]
- 14. Purcell SM, Wray NR, Stone JL, et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009; 4607256: 748.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Darabi H, Czene K, Zhao W, Liu J, Hall P, Humphreys K.. Breast cancer risk prediction and individualised screening based on common genetic variation and breast density measurement. Breast Cancer Res 2012; 141: R25.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Li H, Feng B, Miron A, et al. Breast cancer risk prediction using a polygenic risk score in the familial setting: a prospective study from the Breast Cancer Family Registry and kConFab. Genet Med 2017; 191: 30–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Gottesman O, Kuivaniemi H, Tromp G, et al. The electronic Medical Records and Genomics (eMERGE) Network: past, present, and future. Genet Med 2013; 1510: 761–71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Wolford BN, Willer CJ, Surakka I.. Electronic health records: the next wave of complex disease genetics. Hum Mol Genet [Internet] 2018; 27 (R1): R14–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Kohane IS. Using electronic health records to drive discovery in disease genomics. Nat Rev Genet 2011; 126: 417–28. [DOI] [PubMed] [Google Scholar]
- 20. McCarty CA, Chisholm RL, Chute CG, et al. The eMERGE Network: a consortium of biorepositories linked to electronic medical records data for conducting genomic studies. BMC Med Genomics 2011; 4: 13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Verma SS, de Andrade M, Tromp G, et al. Imputation and quality control steps for combining multiple genome-wide datasets. Front Genet 2014; 5: 370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Purcell S, Neale B, Todd-Brown K, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 813: 559–75. http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1950838&tool=pmcentrez&rendertype=abstract [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Chakravarti A, Turner TN.. Revealing rate-limiting steps in complex disease biology: the crucial importance of studying rare, extreme-phenotype families. BioEssays [Internet] 2016; 386: 578–86. [DOI] [PubMed] [Google Scholar]
- 24. Weiner DJ, Wigdor EM, Ripke S, et al. Polygenic transmission disequilibrium confirms that common and rare variation act additively to create risk for autism spectrum disorders. Nat Genet 2017; 497: 978–85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Shi H, Kichaev G, Pasaniuc B.. Contrasting the genetic architecture of 30 complex traits from summary association data. Am J Hum Genet 2016; 991: 139–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Wood AR, Esko T, Yang J, et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat Genet 2014; 4611: 1173–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Spiliopoulou A, Nagy R, Bermingham ML, et al. Genomic prediction of complex human traits: relatedness, trait architecture and predictive meta-models. Hum Mol Genet 2015; 2414: 4167–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Lewis CM, Vassos E.. Prospects for using risk scores in polygenic medicine. Genome Med 2017; 91: 96.. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Sing T, Sander O, Beerenwinkel N, Lengauer T.. ROCR: visualizing classifier performance in R. Bioinformatics 2005; 2120: 3940–1. [DOI] [PubMed] [Google Scholar]
- 30. Buuren S. V, Groothuis-Oudshoorn K.. MICE: multivariate imputation by chained equations in R. J Stat Softw 2011; 453: 1–67. [Google Scholar]
- 31. Larson MG, Atwood LD, Benjamin EJ, et al. Framingham Heart Study 100K project: genome-wide associations for cardiovascular disease outcomes. BMC Med Genet 2007; 8 (Suppl 1): S5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Smith NL, Felix JF, Morrison AC, et al. Association of genome-wide variation with the risk of incident heart failure in adults of European and African ancestry: a prospective meta-analysis from the cohorts for heart and aging research in genomic epidemiology (CHARGE) consortium. Circ Cardiovasc Genet 2010; 33: 256–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. King G, Zeng L.. Logistic regression in rare events data. Polit Anal 2001; 902: 137–63. [Google Scholar]
- 34. Yu S, Liao KP, Shaw SY, et al. Toward high-throughput phenotyping: unbiased automated feature extraction and selection from knowledge sources. J Am Med Inform Assoc 2015; 225: 993–1000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Hripcsak G, Albers DJ.. Next-generation phenotyping of electronic health records. Am Med Inform Assoc 2013; 201: 117–21. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.