In this study, Gable et al. follow a family with early onset pulmonary fibrosis and report the discovery of a new genetic cause of pulmonary fibrosis. They use multidimensional analysis methods, involving molecular studies, mouse model, and transcriptome-wide studies to show that heterozygous loss-of-function of the exosomal targeting protein ZCCHC8 to identify a novel cause of telomerase insufficiency in human disease.
Keywords: lung disease, ciliopathy, nuclear RNA exosome, RNA processing, telomerase RNA
Abstract
Short telomere syndromes manifest as familial idiopathic pulmonary fibrosis; they are the most common premature aging disorders. We used genome-wide linkage to identify heterozygous loss of function of ZCCHC8, a zinc-knuckle containing protein, as a cause of autosomal dominant pulmonary fibrosis. ZCCHC8 associated with TR and was required for telomerase function. In ZCCHC8 knockout cells and in mutation carriers, genomically extended telomerase RNA (TR) accumulated at the expense of mature TR, consistent with a role for ZCCHC8 in mediating TR 3′ end targeting to the nuclear RNA exosome. We generated Zcchc8-null mice and found that heterozygotes, similar to human mutation carriers, had TR insufficiency but an otherwise preserved transcriptome. In contrast, Zcchc8−/− mice developed progressive and fatal neurodevelopmental pathology with features of a ciliopathy. The Zcchc8−/− brain transcriptome was highly dysregulated, showing accumulation and 3′ end misprocessing of other low-abundance RNAs, including those encoding cilia components as well as the intronless replication-dependent histones. Our data identify a novel cause of human short telomere syndromes-familial pulmonary fibrosis and uncover nuclear exosome targeting as an essential 3′ end maturation mechanism that vertebrate TR shares with replication-dependent histones.
Telomerase, the specialized ribonucleoprotein that synthesizes telomere DNA, has two essential components: the telomerase reverse transcriptase, TERT, and, a specialized non-coding RNA, TR (also known as TERC), which provides the template for telomere repeat addition (Greider and Blackburn 1985, 1987, 1989; Feng et al. 1995; Lingner et al. 1997). Mutant telomerase genes have their most common manifestation in the age-related disease idiopathic pulmonary fibrosis (IPF) (Armanios et al. 2007; Armanios 2012; McNally et al. 2019). Among IPF families with known mutations, half carry mutations in TERT or TR (Armanios et al. 2007; Tsakiri et al. 2007). The rest carry mutations in one of five other telomere maintenance genes, RTEL1, PARN, NAF1, TINF2, and DKC1 (Alder et al. 2013, 2015; Cogan et al. 2015; Stuart et al. 2015; Stanley et al. 2016). Four of these seven known mutant telomere genes in familial IPF affect TR itself, its deadenylation, trafficking, or stability as in the case for PARN, NAF1, and DKC1, respectively (Mitchell et al. 1999b; Moon et al. 2015; Stanley et al. 2016). However, for 50%–60% of IPF families, the genetic etiology is unknown, and within this subset there are individuals with unexplained TR insufficiency and short telomere syndrome features (Stanley et al. 2016).
Here, we used an unbiased genome-wide linkage approach to discover a novel familial IPF disease gene that is required for TR maturation and telomerase function. Human TR has its own RNA polymerase II transcriptional unit (Feng et al. 1995); it shares a 3′ end box H/ACA motif with a subset of non-coding RNAs, but, relative to them, it has low abundance (Mitchell et al. 1999a; Chen et al. 2000; Stanley et al. 2016). Telomere maintenance is vulnerable to small reductions in TR levels (Greider 2006; McNally et al. 2019), and TR is haploinsufficient in both humans and mice (Hathcock et al. 2002; Hao et al. 2005; Armanios et al. 2009). TR+/− mice show genetic anticipation with later generations eventually developing short telomere phenotypes including bone marrow failure and immunodeficiency (Hathcock et al. 2002; Hao et al. 2005; Armanios et al. 2009; Wagner et al. 2018). The determinants of vertebrate TR integrity are incompletely understood in part because the TR sequence is divergent, and different organisms have adapted distinct posttranscriptional processing strategies (Chen et al. 2000; Podlevsky and Chen 2016). Tetrahymena thermophila TR, for example, is transcribed by RNA polymerase III and relies on runs of T’s for transcriptional termination (Greider and Blackburn 1989). S. pombe, on the other hand, uses a modified spliceosome-dependent cleavage mechanism as an initial 3′ end processing step (Box et al. 2008; Coy et al. 2013). Vertebrate TR is transcribed as a longer RNA (Feng et al. 1995; Goldfarb and Cech 2013), and a subset of these extended forms, the short extended oligoadenylated ones, were recently found to be precursors for the mature functional TR (Roake et al. 2019). While TR’s degradation has been linked to the RNA exosome in some cell-based systems (Nguyen et al. 2015; Tseng et al. 2015), the mechanisms by which a longer extended TR matures into a shorter, functional RNA are not known. We report here mutant ZCCHC8, the zinc finger CCHC-type domain containing 8 protein, and a component of the nuclear exosome targeting complex (NEXT) (Lubas et al. 2011), is a cause of familial IPF. We show that ZCCHC8 is required for TR maturation and uncover two distinct RNA dysregulation disease phenotypes caused by partial and complete loss of nuclear RNA exosome targeting.
Results
Genome-wide linkage analysis identifies mutant ZCCHC8 in a family with pulmonary fibrosis
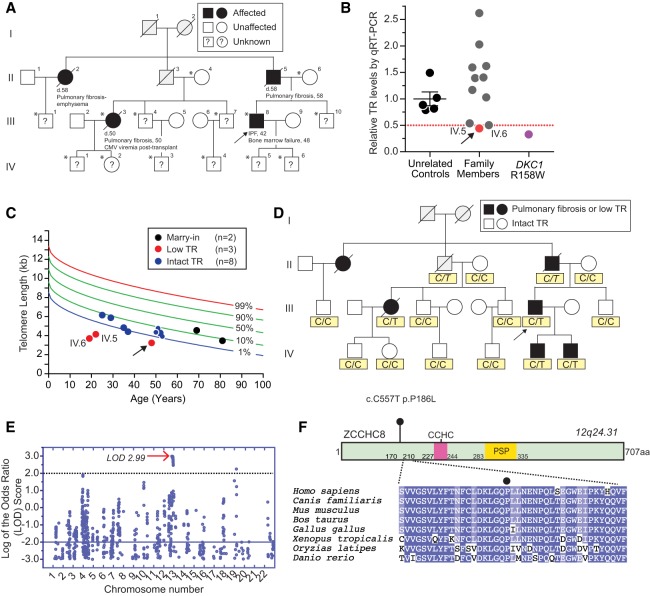
We studied an adult with IPF who showed classic short telomere syndrome features including bone marrow failure (Fig. 1A). His family history was consistent with autosomal dominant pulmonary fibrosis (Fig. 1A), and he had genetically unexplained low TR levels [lymphoblastoid cell lines (LCLs), Fig. 1B]. To identify the genetic basis, we recruited and assessed 13 of his family members, but found there were no living affected individuals (Fig. 1A). This led us to determine preclinical status by measuring TR levels in LCLs and we found the proband's two children also had 50% of control TR levels, while the remaining relatives had TR levels similar to healthy controls (Fig. 1B). Telomere length measurement confirmed both of these “low TR” asymptomatic individuals were at risk for disease since they had abnormally short telomere length, below the first age-adjusted percentile, and shorter than the proband (Fig. 1C). These data supported assigning affected status to the two presymptomatic individuals, and the other family members as unaffected (Fig. 1D). We used these designations to perform a SNP array-based genome-wide linkage analysis and identified a single 17.3-Mb linkage peak on chromosome 12 (1,073,260–1,246,120) with a maximum log of the odds ratio (LOD) score of 2.99 (Fig. 1E). Whole-genome sequencing identified three rare coding variants within this interval in the proband (Supplemental Table S1), but only one of them fell in an RNA-related gene, ZCCHC8. The variant, c.C557T predicted a p.P186L missense, had not been reported in >140,000 individuals in the databases (Supplemental Table S1) and segregated with both the IPF and low TR phenotypes (Fig. 1D). In a multi-species alignment, p.P186 fell in an unstructured, highly conserved domain of ZCCHC8 (Fig. 1F; Supplemental Fig. S1). We screened 42 other genetically uncharacterized families with pulmonary fibrosis and other short telomere syndrome phenotypes but did not identify additional ZCCHC8 variants, suggesting these mutations are rare (1 of 43, 2%), and consistent with a locus heterogeneity for short telomere syndromes.
Figure 1.
Linkage analysis and whole-genome sequencing identify novel disease gene ZCCHC8 in familial pulmonary fibrosis with low telomerase RNA (TR). (A) Pedigree with pulmonary fibrosis proband (arrow) with affected relatives are indicated by the shaded symbols (key). The clinical history below each of the four shaded pedigree symbols refers to the age of onset of lung disease including idiopathic pulmonary fibrosis (IPF). (?) Asymptomatic individuals who had unknown affected status at the time of clinical assessment; (gray shading) unknown cause of death; (*) individuals with DNA who were included in the linkage analysis. (B) TR levels measured by quantitative real time PCR (qRT-PCR) in lymphoblastoid cell lines (LCLs). Arrow refers to proband (red) and pedigree identifiers refer to A. TR level from a DKC1 mutation carrier is a positive control. The data represent a mean of three experiments, each from independent RNA isolations. (C) Telogram shows age-adjusted lymphocyte telomere length by flow cytometry and fluorescence in situ hybridization (flowFISH) in the proband (arrow) and family (pedigree designations as in A). The validated telogram is based on 192 controls. (D) Phenotype assignments used in linkage (key) and genotype below each individual refers to ZCCHC8 SNP. Italicized genotypes refer to obligate carriers. (E) Log of the odds (LOD) ratio across autosomal chromosomes calculated from SNP data from 14 individuals, with arrow on chromosome 12 pointing to maximum LOD. (F) p.P186L conservation across eight vertebrate ZCCHC8 species with darker shading denoting more conserved residues. CCHC refers to Zinc-knuckle domain; PSP refers to proline-rich domain.
ZCCHC8P186L is a loss-of-function mutation
We first tested the functional effects of ZCCHC8P186L and found that all three mutation carriers had 50% lower ZCCHC8 protein levels compared with controls, including noncarrier relatives (LCLs; P = 0.038, Mann-Whitney U-test) (Fig. 2A; antibody validation in Supplemental Fig. S2A). Low protein levels were also seen in the proband's primary skin fibroblasts and were specific to ZCCHC8 as the levels of the other two NEXT complex proteins, RBM7 and SKIV2L2, were normal (Fig. 2B; antibody validation in Supplemental Fig. S2B,C). We transfected Myc-tagged wild-type and mutant ZCCHC8 and found ZCCHC8P186L levels were decreased (P < 0.01, three independent transfections, Student's t-test) (Fig. 2C). Total ZCCHC8 mRNA levels were also intact in mutation carriers and the mutation was detected in the mRNA (Fig. 2D,E), additionally confirming p.P186L compromised protein stability. Short hairpin (shRNA) knockdown of endogenous ZCCHC8 decreased total TR levels in HeLa cells as quantified by both qRT-PCR and northern (Fig. 2F–H; northern specificity in Supplemental Fig. S2D). In contrast, shRNA knockdown of RBM7 and SKIV2L2 increased TR levels (Supplemental Fig. S2E,F), as had been seen previously (Tseng et al. 2015). In this experiment, the depletion of one component of NEXT by shRNA affected the stability of others, in contrast to patient-derived primary cells where low ZCCHC8 levels did not affect RBM7 or SKIV2L2 levels (Fig. 2B). These collective data suggested that ZCCHC8 loss-of-function was sufficient to cause TR depletion.
Figure 2.
ZCCHC8 loss of function is sufficient to cause low TR levels. (A) Immunoblot of ZCCHC8 in lymphoblastoid cell lines (LCLs) from healthy controls (C1 and C2) and unaffected relatives and mutation carriers labeled with pedigree identifiers from Figure 1A. Quantification from one blot and result replicated twice from independently harvested protein lysates. (B) Immunoblot of ZCCHC8, SKIV2L2, and RBM7 levels in proband's primary skin fibroblasts. (C) Immunoblot of transfected Myc-tagged (293FT cells) and endogenous ZCCHC8. (D) Mean ZCCHC8 mRNA levels ± SEM from LCLs in unaffected family members (n = 4) and ZCCHC8 p.P186L mutation carriers (n = 3). (E) Chromatogram showing that ZCCHC8 p.P186L mutation is expressed in LCL mRNA from proband (also verified in two other mutation carriers). (F) Immunoblot showing efficiency of shRNA knockdown of Luciferase (Luc), ZCCHC8, and NAF1 in HeLa cells. (G) Total TR levels measured by qRT-PCR (mean ± SEM from three independent knockdowns and RNA isolations). (H) Northern blot of TR after stable knockdown of ZCCHC8 and NAF1 (replicated twice with independent RNA isolations). (**) P < 0.01; (***) P < 0.001 (Student's t-test, two-sided).
ZCCHC8 is required for TR maturation and telomerase function
To test the consequences of ZCCHC8 loss on telomerase function, we disrupted its locus in pseudodiploid HCT116 lines using CRISPR/Cas9 editing (Fig. 3A,B). Since the NEXT complex is thought to target a subset of RNAs for degradation and/or 3′ end processing by the nuclear RNA exosome (Lubas et al. 2011), we characterized TR 3′ ends using rapid amplification of cDNA ends combined with next-generation sequencing (3′RACE-seq) (Fig. 3C). In control cells, nearly all of the nonmature TR species were oligoadenylated and had genomically encoded short extensions (<15 nt) (Fig. 3D), as had been seen previously (Goldfarb and Cech 2013). Compared with isogenic wild-types, ZCCHC8−/− HCT116 cells had a relative decrease in the mature TR fraction (451 nt; single clone, three isolations, and sequencing replicates) (Fig. 3D; speciation of sequencing products shown in Supplemental Fig. S3). Concurrently, ZCCHC8−/− cells had an increase in short, extended TR forms which comprised >90% of all the aberrant forms detected (Fig. 3D). These short extended TR molecules have been recently shown to be nascent precursors to mature TR (Roake et al. 2019). The remaining genomically encoded extensions were longer, reaching 20, 51, and 784 nt beyond the mature TR 3′ end, and these also showed a fourfold to 10-fold increase as quantified by qRT-PCR (Fig. 3E). We tested whether the accumulation of extended TR caused by ZCCHC8 deletion may affect TR levels. We measured total TR (as a surrogate for mature TR) by Northern blot and found ZCCHC8−/− HCT116 cells had TR insufficiency with mean of 21% decrease (P = 0.038) (Fig. 3F,G); this effect underestimated the mature TR amounts since Northern blot lacks the resolution for distinguishing mature from the short extended TR detected by 3′RACE-seq. Since extended TR has recently been shown to be nonfunctional (Deng et al. 2019), the Northern quantification may also underestimate the consequences of ZCCHC8 loss on TR function. To assess this directly, we measured telomerase activity using the telomere repeat amplification protocol (TRAP) and found that, similar to NAF1 mutant cells (Stanley et al. 2016), ZCCHC8−/− cells showed decreased activity (mean 52% of isogenic ZCCHC8+/+ cells, P = 0.001, Student's t-test) (Fig. 3H,I). These collective data indicated that ZCCHC8 is required for TR 3′ end maturation and for telomerase function.
Figure 3.
ZCCHC8 is required for its 3′ end maturation and telomerase function. (A) Compound heterozygous frameshift (fs) mutations introduced using CRISPR/Cas9 in HCT116 pseudodiploid cells. (B) Immunoblot for ZCCHC8 in HCT116-edited cells. (C) Scheme summarizing TR 3′ rapid amplification of cDNA ends sequencing (3′RACE-seq). TR 3′ ends were generally divided into mature (451 bp) and extended (>451 bp) where extensions are denoted by gray N's. (D) Summary of TR 3′RACE-seq fractions in isogenic ZCCHC8+/+ and ZCCHC8−/− cells. Color-coded key shows four categories of TR forms: mature (451 nt), adenylated (A)n, short genomically extended (g)n (<465 nt), and long genomically extended (>465 nt). Data are mean of three independent 3′RACE-seq analyses from three RNA isolations each from a different aliquot of a single clone. (E) qRT-PCR of extended TR forms beyond the 451 mature end (>20, >51, >784 nt). Data are mean of three independent RNA isolations similar to D. (F) TR Northern blot of edited ZCCHC8+/+ and ZCCHC8−/− cells. (G) Total TR levels by Northern bot (six blots from three RNA isolations). (H) Telomerase activity measured by telomere repeat amplification protocol (TRAP) assay in ZCCHC8+/+, ZCCHC8−/−, and NAF1S329/S329 HCT116 cell extracts. Activity was quantified on serially diluted extracts (1, 1/5, 1/25, and 1/125) against a PCR-amplified internal control (IC). RNase-treated wild-type extract and no template PCR reaction are included as negative controls. (I) Mean TRAP activity of 1/5× diluted extracts (three independent TRAP assays, each from a different lysate). (J) Summary of 3′RACE-seq of TR forms from control and proband's primary skin fibroblasts with speciation as in D. (K) qRT-PCR values of extended TR forms in primary skin fibroblasts as in E, mean of three technical replicates). (L) Amplified TR from input and Myc-ZCCHC8 immunprecipitated fractions (293FT cells) using primers falling within the mature TR sequence. (M) qRT-PCR of extended TR (>51 nt extended beyond the 3′ mature TR end) after transfection of tagged ZCCHC8, DIS3, EXOSC10/RRP6, and PARN into HCT116 ZCCHC8−/− cells (three to four independent transfections/experiment). Data are expressed as mean ± SEM (*) P < 0.05; (**) P < 0.01 (Student's t-test, two-sided).
We next examined the clinical relevance of these findings by performing 3′RACE-seq on the proband's primary fibroblasts and found a similar TR 3′ end distribution as ZCCHC8−/− lines with an accumulation of short 3′ extended TR fractions (Fig. 3J,K; Supplemental Fig. S4). However, there was heterogeneity in the stability of these TR precursors across cell types as we could not amplify them readily from Epstein-Barr virus-transformed LCLs, consistent with published data pointing to herpes viruses interfering with RNA turnover in hematopoietic-derived cells (Glaunsinger and Ganem 2004).
ZCCHC8 associates with TR precursors and their accumulation is rescued by a nuclear RNA exosome catalytic component
If ZCCHC8 is involved in TR targeting, we would expect an association, we therefore performed RNA immunoprecipitation of Myc-tagged ZCCHC8 and found TR was enriched in the precipitate (Fig. 3L; Supplemental Fig. S5A). This physical association was independent of an interaction with dyskerin or NAF1 (Supplemental Fig. S5B). We characterized these TR species by performing TR-specific 3′RACE-seq of ZCCHC8-immunoprecipitated RNA and found 3′ genomically extended TR was enriched (Supplemental Figs. S5C, S6). Nearly all of these extensions were short, consistent with a role of ZCCHC8 in nascent TR processing/targeting. We directly tested whether the accumulation of extended TR forms may be rescued by overexpression of DIS3 or EXOSC10 (also known as RRP6), both 3′ > 5′ exoribonucleases of the nuclear RNA exosome and found a modest reduction of TR extending >51 nt (Fig. 3M). ZCCHC8 overexpression in these cells also rescued extended TR levels, consistent with an on-target effect. In contrast, there was no significant change with forced expression of the PARN deadenylase (Fig. 3M). These data implicated ZCCHC8 in the targeting of genomically encoded extended TR to the nuclear RNA exosome.
ZCCHC8 shows a dose-dependent requirement for TR maturation
To study the role of ZCCHC8 in vivo, we generated knockout mice. These studies were particularly relevant since we noted in the literature mention of one consanguineous family with autosomal recessive intellectual disability that carried a homozygous ZCCHC8 mutation p.L90X (Najmabadi et al. 2011). We designed the CRISPR/Cas9 genome editing strategy to insert a 33-bp insertion in exon 2 of Zcchc8 introducing a premature stop codon at amino acid 93 (Supplemental Fig. S7A). We interbred heterozygous mutant mice and detected viable heterozygous and homozygous null progeny and they had half and no detectable ZCCHC8 protein, respectively (Fig. 4A; antibody cross-validation in Supplemental Fig. S2A). We first examined the stability of RBM7 and SKIV2L2 in adult Zcchc8+/− fibroblasts and found their levels were preserved similar to the patient cells (Figs. 4B,C and 2B, respectively), while Zcchc8−/− cells had minimally decreased levels (Fig. 4B,C; Supplemental Fig. S2B,C). The stability of RBM7 and SKIV2L2 thus does not depend on the presence of ZCCHC8 also in this physiologic model, in contrast to the results seen with shRNA knockdown in cancer cells (Fig. 4B,C; Supplemental Fig. S2E,F). We next quantified total TR levels by Northern blot and found that, similar to mTR+/−, Zcchc8+/− and Zcchc8−/− cells had a dose-dependent decrease (P = 0.02 for both comparisons, n = 3–4 mice/genotype, Student's t-test) (Fig. 4D,E). To test whether ZCCHC8 also plays a role in TR end-processing in mice, we amplified genomically encoded TR extensions that are longer than 20 nt beyond the mature 3′ end, and found that Zcchc8+/− and Zcchc8−/− mice showed a dose-dependent increase with knockouts having the highest levels (Fig. 4F).
Figure 4.
Zcchc8-null mice have TR insufficiency. (A–C) Immunoblot for ZCCHC8, SKIV2L2, and RBM7, respectively, on lysates from mouse ear fibroblasts. (D,E) Northern blot for mouse TR and quantification. For E, mean reflects mice Zcchc8+/+ (n = 4, 2M/2F), Zcchc8+/− (n = 4, 2M/2F), Zcchc8−/− (n = 3M), mTR+/− (n = 2, sex unknown) and mTR−/− (n = 2, sex unknown). (F) TR 3′ extended levels (>20 bp) relative to Hprt as measured by qRT-PCR. Mouse numbers and M/F designations as in E. Data are expressed as mean ± SEM. (*) P < 0.05; (**) P < 0.01 (Student's t-test, two-sided).
When we interbred Zcchc8+/− mice, we noted a distortion in the Mendelian ratios with Zcchc8−/− progeny representing 5% of adult mice (P < 0.001, X2 test, total n = 170 weaned pups by 30 d) (Supplemental Fig. S7B). None of these knockout mice survived beyond 70 d. The Zcchc8−/− lethality occurred postnatally as the Mendelian ratios were preserved at E12.5 and at birth (Supplemental Fig. S7B). Thus, while ZCCHC8 is not required for viable embryogenesis, its complete loss causes a postnatal fatal phenotype.
Zcchc8−/− mice develop a progressive neurodevelopmental defect
We characterized the phenotype by first focusing on Zcchc8+/− adults and found that these mice were grossly and histologically indistinguishable from wild-type littermates (Fig. 5A). Specifically, there were no abnormalities in the brain, viscera, and hematopoietic system including complete blood counts. These observations were consistent with Zcchc8+/− mice primarily having TR insufficiency that would be predicted to cause telomere shortening after many successive generations, as has been documented for mTR+/− mice (Armanios et al. 2009). In contrast, Zcchc8−/− mice had obvious cranial deformities with domed-shaped heads, a sign of hydrocephalus (Fig. 5A, top panel). Computed tomography (CT) imaging confirmed the presence of dilated cranial sutures, another indicator of hydrocephalus (Fig. 5A; Supplemental Fig. S7C), and nearly all adult Zcchc8−/− mice had severe hydrocephalus with ventriculomegaly postmortem (91%, nine of 11) (Fig. 5B; Supplemental Fig. S7D,E). Hydrocephalus is a common feature of ciliopathies; it arises because of defective motile cilia on ependymal cells that are required for normal cerebrospinal fluid circulation (Kousi and Katsanis 2016). We looked for other ciliopathy features and found a high prevalence of severe otitis media in Zcchc8−/− mice (80%, eight of 10) (Supplemental Fig. S7D,E), but there were no laterality defects, such as situs inversus or heterotaxy (Supplemental Fig. S8A), and there was no evidence of upper airway disease. To assess the cause of the cranial defects, we traced brain development in Zcchc8−/− embryos and found they had small brain volumes during embryogenesis and at birth (Fig. 5C–F), indicating that defective neurogenesis preceded the onset of hydrocephalus. We saw no gross difference in proliferation or cell death during embryonic brain development by immunohistochemical staining (Supplemental Fig. S9A). These collective data indicated that central nervous development was particularly sensitive to complete loss of ZCCHC8, and that a progressive neurodevelopmental defect with features of ciliopathy was the primary cause of death in Zcchc8−/− mice.
Figure 5.
ZCCHC8 complete loss causes progressive and fatal neurodevelopmental phenotype. (A, top row) Images showing head profile of Zcchc8 wild-type, heterozygous, and homozygous null mice (41–46 d-old). The labels show the genotype with a male (left) and a female (right) for each genotype. Zcchc8−/− mice have abnormal head profiles with domed crania, as outlined by the dashed line. (Middle row) CT head mid-sagittal images show Zcchc8−/− mice have dome-shaped crania. (Bottom row). Volume-rendered (VR) CT images of mouse calvaria show widened cranial sutures in Zcchc8−/− mice (seen in three of six imaged). None of Zcchc8+/+ or Zcchc8+/− mice (four mice imaged/genotype) had this feature. Each vertical image group is from the same mouse except the last column (two different females). (B) Representative H&E coronal sections from 8-wk-old heads (all male) show no differences in Zcchc8+/− mice (11 examined) compared with Zcchc8+/+ mice (10 examined). In contrast, Zcchc8−/− mice had severe ventricular dilation (nine of 11 examined). (C) E12.5 brain sections show Zcchc8−/− have microcephaly but intact brain structures with no ventriculomegaly in utero. (D) Image of E12.5 embryos from a single dam showing expected Mendelian ratios but Zcchc8−/− embryos have small crania. (E) Cranial area of newborn (P0) pups measured on VR CT images and corrected to left femur length on the same images (Zcchc8+/+ n = 7, 3M/4F; Zcchc8+/− n = 10, 3M/7F; Zcchc8−/− n = 4, 3M/1F). (F) Mean cranial area ± SEM relative to age for all three genotypes showing that Zcchc8−/− mice develop macrocephy after birth. Newborn mice include those listed in E and older mice (Zcchc8+/+, n = 4, 3M/1F; Zcchc8+/−, n = 4, 3M/1F; Zcchc8−/−, n = 5, 3M/2F). Dashed lines denote 95% confidence intervals. (**) P < 0.01 (Student's t-test, two-sided).
Global analysis shows that Zcchc8−/− developing brains have defective turnover of low abundance RNA polymerase II transcripts
To better understand the basis for the neurodevelopment defect in Zcchc8−/− mice, we performed RNA-seq on E12.5 brains. We used an adenylated RNA selection library to focus on protein-coding genes. Comparison of the RNA profiles in a hierarchical analysis, using no supervision, showed each of the three Zcchc8 genotypes clustered, but there was a striking deviation in the Zcchc8−/− pattern from wild-type and heterozygous transcriptomes (Fig. 6A). The volcano plot analyses also showed little difference between the Zcchc8+/− versus Zcchc8+/+ transcriptomes, but the Zcchc8−/− brain showed a skewed pattern towards higher expression of multiple RNAs (Fig. 6B,C). We examined the identity of these differentially expressed RNAs (defined as >2SD-fold change, n = 197) and found they were all transcribed by RNA polymerase II and fell within the lowest FPKM values of the wild-type adenylated transcriptome distribution (P < 10−8, Mann-Whitney test and Bootstrap analysis) (Fig. 6D). In contrast, the distribution of the most down-regulated genes was broad (Fig. 6D; Supplemental Table S2B). Among the up-regulated RNAs, TR had a mean 6.8-fold increase (Zcchc8−/− vs. Zcchc8+/+, P = 1.72 × 10−8, one-way two-tailed ANOVA), and the increase was due to 3′ genomically encoded extended forms (Supplemental Fig. S9B). This pattern was similar to what we documented in patient-derived and HCT116-edited cancer cells. We also noted that this pattern was specific, as no 3′ extensions were seen for RNA polymerase III transcribed genes such as Rmrp (Supplemental Fig. S9B). These data suggested that ZCCHC8 is involved in the turnover and/or processing of low-abundance RNA polymerase II transcribed RNAs other than TR.
Figure 6.
The Zcchc8−/− transcriptome shows up-regulation and misprocessing of low-abundance intronless RNAs other than TR. (A) Heat map with dendrogram of gene expression showing unsupervised analysis of 9788 high-quality genes from brain RNA-seq analysis. Colors denote mean-subtracted FPKM expression values on a log2 scale (Zcchc8+/+, n = 5; Zcchc8+/−, n = 3; Zcchc8−/−, n = 6 embryonic brains sequenced). Each column is labeled below by WT, HET, and KO followed by the embryo number (1, 2, 3, etc.), referring to respective Zcchc8 genotypes. The log2 expression value was subtracted from the mean log2 expression value of the entire cohort. The dendrogram showing relatedness of the samples is above, and relatedness of the gene transcripts is at the left. The differential change in RNA expression is shown as positive and negative change on color scale in the key above the top right corner. (B,C) Volcano plots depicting the log2-fold changes (X-axis) versus −log10 P-values calculated by two-tailed one-way ANOVA (Y-axis) for the Zcchc8+/− and Zcchc8−/− versus Zcchc8+/+ comparisons, respectively. Each dot represents a single transcript. (D) Histogram of number of genes at each expression value denoted on the x-axis by the mean log2FPKM values obtained from Zcchc8 wild-type embryos (n = 5). RNAs that have more than two SD higher levels in the Zcchc8+/+ versus Zcchc8−/− comparison are shown in red (n = 197) and fall on the low end of the histogram with TR and its mean FPKM in wild-type embryos shown. Down-regulated RNAs, defined as less than two SD (n = 43), are shown in blue appear uniformly distributed on the distribution. (E) Histogram of the most up-regulated (>2SD) transcripts in the Zcchc8−/− versus Zcchc8+/+ by exon number shows the largest subset is intronless RNAs (42 of 188 with known gene structure, 22%). The pie chart divides the intronless RNAs by functional category. (F) Annotation of 28 up-regulated intronless RNAs (TR, histones and cilia) shows a majority of the histones represented are replication-dependent histones (RDH) (23 of 24, 96%). The majority have an annotated transcript size in the range of TR between 400 and 560 (22 of 28). Columns referring to 5′ end and 3′ end refer to visualized additional reads beyond annotated gene boundaries with 5′ end reads referring to upstream reads that are not necessarily contiguous (manually identified in the Integrative Genome Viewer [IGV]). (G,H) Genome browser read coverage plots from IGV viewer showing extended 3′ ends as labeled above from two histone genes in each of Zcchc8+/+, Zcchc8+/−, Zcchc8−/− transcriptomes. (I,J) Coverage plots for two coiled-coil domain containing cilia genes Ccdc89 and Ccdc182, respectively, by genotype. For Ccdc89, there is also an increase in discontinuous upstream of gene 5′end reads that resemble so-called promoter upstream transcripts (PROMPTs).
ZCCHC8 is required for posttranscriptional processing of a subset of replication-dependent histones and some motile cilia components in addition to TR
We probed the identity of up-regulated RNAs in the Zcchc8−/− developing brain and found the largest subset was, like TR, intronless (22%, 42 of 188 with known gene structure) (Fig. 6E; Supplemental Table S2A). These intronless pre-mRNAs also generally had similar length to TR, between 400 and 500 nt (Fig. 6F). Among this class of RNAs derived from single-exon genes, there was enrichment of two subsets. The first were histones which comprised more than half (23 of 42, 55%), and almost all of these were replication-dependent histones (RDH) (22 of 23, 96%, Fig. 6F). The mouse genome has 65 RDH genes and the majority, 51 of 65 (78%), fall in the major histone cluster on chromosome 13 (Marzluff et al. 2002). Our RNA-seq analysis enriched for nearly half of these RDH (P < 0.0001, 22 of 51 vs. 55 of 9788 high-quality transcripts analyzed in the RNA-seq data set, Fisher's exact test). The second subset involved RNAs that encode cilia protein components (Fig. 6E,F; Supplemental Table S2A). Dysfunction of motile cilia on epithelial cells could explain the hydrocephalus-otitis phenotype of Zcchc8−/− mice. Indeed, while the overall expression of ZCCHC8 in the brain was low, we found that it was highly enriched in ependymal cells (Supplemental Fig. S9C–E). Among the 197 most up-regulated genes, 13 (7%) had cilia-related functions (Supplemental Table S2A). In addition to coiled-coil domain proteins involved in ciliagenesis (n = 5), there were transcripts encoding transmembrane proteins (Tmems) that are involved in the formation of ciliary transition zones (n = 4), Dnah7b, an essential component of the dynein axonemal heavy chain of motile cilia (Zhang et al. 2002), and Ttc26, the intraflagellar transporter which when mutated causes hydrocephalus in mice (Supplemental Table S2A; Swiderski et al. 2014). These data, along with the enriched expression of ZCCHC8 in ependymal cells, suggest that RNA dysregulation may provoke a secondary ciliopathy that manifests as hydrocephalus in Zcchc8−/− mice.
We asked whether the up-regulation of RNAs in the Zcchc8−/− transcriptome reflected 3′ end misprocessing as we had seen for TR. We manually examined the RDH RNAs and found that, in addition to their increased abundance, most had 3′ genomically encoded extensions (22 of 28) (Fig. 6F–H). These extensions were most prominent in the Zcchc8−/− transcriptome and generally subtle or absent in heterozygotes (Fig. 6G,H). A similar pattern of 3′ extensions was seen for the cilia-related transcripts (Fig. 6F,I,J). The 3′ extended transcripts were specific to the up-regulated genes as there was no misprocessing of RNA polymerase II transcripts that had preserved levels in the Zcchc8−/− transcriptome (e.g., Arf1, Arf3). For half of the RDH and cilia genes we examined (14 of 28), there were additionally upstream RNA reads that were discontiguous with the 5′ gene end (Fig. 6G). These resembled promoter upstream transcripts (PROMPTS), ncRNAs that are known to be degraded by the nuclear RNA exosome and that have been reported with depletion of RBM7 as well as some nuclear RNA exosome components (Andersen et al. 2013).
Discussion
We report here heterozygous mutations in ZCCHC8 as a novel cause of familial pulmonary fibrosis and the human short telomere phenotype. This report further establishes the intimate connection between the susceptibility to IPF and the short telomere defect. We used molecular phenotyping to assign preclinical affected status for an age-dependent, rapidly progressive disease phenotype in order to strengthen the power of linkage analysis. ZCCHC8 is a vertebrate-specific Zinc-knuckle containing protein; it was identified as part of the NEXT trimer in mammalian cells as a cofactor and adaptor of the nuclear RNA exosome. It, along with RBM7, are nuclear specific and excluded from nucleoli presumably to provide substrate specificity for the RNA exosome's catalytic activity (Lubas et al. 2011). An interaction between SKIV2L2 and the RNA exosome has been proposed to activate exonucleolytic activity involved in the maturation as well as degradation of a subset of RNA polymerase II transcripts (Lubas et al. 2011). The human and mouse genetic studies we report here establish a role for ZCCHC8 in the posttranscriptional maturation of functional TR. TR is transcribed as a longer RNA with a majority of detected species extending up to 15 beyond its eventual mature end. Our data, across cell and model organisms, show ZCCHC8 loss causes an accumulation of these short extended TR forms at the expense of mature TR and support a role for ZCCHC8-directed targeting for exosome-dependent exonucleolytic activity. Our model is consistent with the known functions of NEXT in 3′ end maturation of other RNAs (Lubas et al. 2015) and the data showing short extended TR represents a nascent precursor (Roake et al. 2019). Notably, in contrast to the consequences of ZCCHC8 loss, the knockdown of RBM7 and SKIV2L2 increased TR levels. The interpretation of these data requires further follow up given the fact that mutant ZCCHC8 human and mouse cells, including those derived from affected patients, had intact levels of other NEXT components, in contrast to shRNA knockdown. Overall, our findings highlight the complexities regulating TR biogenesis and the exquisite susceptibility for genetic disturbances in this pathway to manifest in the human IPF phenotype. The PARN deadenylase, which is also mutated in human short telomere disease, is required to prevent exosome-dependent degradation of TR (Moon et al. 2015; Shukla et al. 2016). Our data showing accumulation of genomically encoded extended TR in ZCCHC8 mutant cells and animals suggest that ZCCHC8-dependent exosome targeting is an early process likely preceding PARN deadenylation. They point to the RNA exosome differentially playing a role in TR maturation as well as degradation during different stages of TR biogenesis.
We uncovered two distinct RNA dysregulation disease phenotypes with partial and complete loss of ZCCHC8. Heterozygous null humans and mice developed TR insufficiency and this manifested as a classic short telomere syndrome that is indistinguishable from TR or TERT mutation carriers. On the other hand, complete ZCCHC8 loss caused a severe and ultimately fatal neurodevelopmental defect in mice. This neurodevelopmental phenotype showed features of impaired neurogenesis as well as a postnatal ciliopathy. The murine phenotype we report is notable given a prior report of a single family with autosomal recessive intellectual disability that carried a homozygous null ZCCHC8 mutation (Najmabadi et al. 2011). Although we found ZCCHC8 was ubiquitously expressed in visceral organs, its levels on the brain were overall low but showed a particular enrichment in ependymal cells. We also detected a large number of misprocessed cilia-encoding RNAs in the Zcchc8−/− developing brain, suggesting a link to cilia dysfunction and hydrocephalus. The central nervous system predilection of disease in ZCCHC8-null mice, as well as human mutation carriers with intellectual disability, is consistent with the exquisite sensitivity of neurodevelopment to RNA dysregulation as has been seen for exosomopathies (Müller et al. 2015; Giunta et al. 2016). Importantly, our data identify RNA misprocessing as a potential cause for secondary ciliopathies.
We show that there is an in vivo requirement for ZCCHC8 in the 3′ end processing of low abundance polymerase II RNAs beyond TR, including RDH and some cilia components. This RNA subset is enriched for short, intronless RNAs that are of similar length as TR. The mechanisms by which ZCCHC8 selectively targets this subset of RNAs to the nuclear RNA exosome remains unclear, but a similar signal for low abundance RNAs was also seen recently when RRP40, a noncatalytic nuclear RNA exosome component, was depleted in mouse embryonic stem cells (Lloret-Llinares et al. 2018). RDH mRNAs have been shown to have a unique posttranscriptional processing pathway that did not involve adenylation (Marzluff et al. 2008). Recently, however, RDHs were found to be adenylated in terminally differentiated tissues (Lyons et al. 2016). Our RNA-seq data, in the relevant setting of embryonic brain development, indicate that RDH posttranscriptional processing may include 3′ adenylation and an end-processing step that, as for TR, may rely on nuclear exosome targeting. Our data therefore identify a role for ZCCHC8-dependent targeting to the nuclear RNA exosome in the maturation of TR as well as the processing of other low abundance RNA polymerase II genes.
Materials and methods
Subjects and study approval
Patients were recruited from 2007 to 2017 as part of the Johns Hopkins Telomere Syndrome Registry as described previously (Jonassaint et al. 2013; Alder et al. 2018). The study was approved by the Johns Hopkins Medicine Institutional Review Board and all the subjects gave written informed consent. The proband was identified in a screen for individuals who had low TR levels in lymphoblastoid cell lines (LCLs) by qRT-PCR as described previously (Stanley et al. 2016). LCLs were generated by Epstein-Barr virus transduction (Penno et al. 1993), and primary skin fibroblasts from the index case were derived from a skin punch biopsy using standard methods. Control skin fibroblasts were purchased from American Type Culture Collection (ATCC; BJ, CRL-2522). The screen of 42 patients for mutant ZCCHC8 included genetically uncharacterized familial pulmonary fibrosis-emphysema probands (n = 31) and patients with a classic short telomere syndrome phenotype and short telomere length as previously defined (n = 11) (Stanley et al. 2016).
Telomere length measurement and DNA extraction
Telomere length was measured on peripheral blood-derived cells by flow cytometry and fluorescence in situ hybridization (flowFISH) at the Johns Hopkins Pathology Laboratories (Baerlocher et al. 2006; Alder et al. 2018). Genomic DNA was extracted from fresh blood or PBMCs using the Gentra Puregene Kit B (Qiagen) and from formalin-fixed paraffin-embedded (FFPE) using the FFPE tissue DNA isolation kit (MO BIO Laboratories).
Quantitative real-time-PCR (qRT-PCR) for TR
RNA levels were measured using the SYBR Green qRT-PCR method as described previously (Stanley et al. 2016). Briefly, RNA was isolated (RNeasy, Qiagen) and reverse transcribed using random hexamer primers and Superscript III (Invitrogen). Within any given experiment, RNA isolation and cDNA reverse transcription were performed side-by-side for all the samples. TR levels were normalized to ARF3 (human) and Hprt (mouse). Supplemental Table S3 includes primer sequences for hTR and mTR qRT-PCR both total and extended forms. Primers for quantifying hTR extended forms were from obtained from Nguyen et al. (2015) and Tseng et al. (2015).
Whole-genome sequencing
Whole-genome sequencing was performed on blood-derived DNA (Macrogen) using a HiSeq X platform (Illumina). Variants were called and aligned to GRCh37 reference genome using the Isaac variant caller (v.2.0.13) and aligner (v.01.15.02.08). The mean depth of coverage was 48X with 98% of the genome covered at 20X and 93% at 30X. Annotation and filtering was performed using PhenoDB (Hamosh et al. 2013). Nonsynonymous exonic and splice junctions were defined as rare if they were absent in dbSNP builds 129, 131, and 135, and also had minor allele frequency (MAF) <0.0001 in 1000 genomes and the NHLBI GO Exome Sequencing Project (accessed June 19, 2017 and updated January 6, 2019).
Genome-wide linkage analysis
To identify the disease locus, we genotyped a total of 14 individuals using a SNP array (548K, Infinium HumanCoreExome-24v1-0, Johns Hopkins Genetic Center Resource Facility). SNPs were ranked in 400-kb bins by quality metrics based on call rates, position, and tightness of the genotype cluster. The 6710 high-quality autosomal SNPs were then used in a singlepoint linkage analysis using MERLIN software (Abecasis et al. 2002). Subjects were assigned “affected” status if they had a short telomere syndrome phenotype such as IPF, were obligate carriers, or had low TR levels and abnormally short telomere length. We used an autosomal dominant model with prevalence of 0.0001 and penetrance of 1.0 and 0.0001 in carriers and noncarriers, respectively.
ZCCHC8 multiple species alignment
Human ZCCHC8 (NP_060082) was aligned to the following vertebrate ZCCHC8 protein sequences (Canus familiaris: XP_005636220; Mus musculus: NP_081770; Bos taurus: NP_001192619; Gallus gallus: NP_001192619, Xenopus tropicalis: NP_001135573; Oryzias latipes: XP_004072626; and Danio rerio: NP_001077287) within Jalview (version 2.10.2b2) using Clustal Omega (Waterhouse et al. 2009).
Targeted sequencing of ZCCHC8
Genotyping for ZCCHC8 P186L was done using Sanger sequencing of the 102 bp magnetic bead purified PCR product (AMPure XP, Beckman Coulter Life Sciences) using primers listed (Supplemental Table S3). To screen for ZCCHC8 mutations in short telomere syndrome patients, we sequenced gDNA for the coding sequence and exon–intron junctions on a customized targeted nextgen sequencing panel (Truseq, Illumina) as shown previously (n = 21 cases) (Stanley et al. 2015). The mean depth of coverage was 50X. For remaining cases (n = 21), we Sanger sequenced the ZCCHC8 cDNA derived from LCLs bidirectionally (primers available upon request).
Western blots
Total protein was isolated from cells in RIPA buffer (Cell Signaling Technology) with cOmplete Mini protease inhibitor (Sigma-Aldrich) and lysates were quantified using Pierce BCA protein assay kit (Thermo Fisher Scientific). Using the NuPAGE SDS-polyacrylamide gel system, protein (10–20 µg) was run on 10% Bis-Tris gels with MOPS-SDS running buffer at 150 V and transferred to a PVDF membrane by wet transfer using XCell SureLock mini-cell elecrophoresis system or by the iBlot 2 dry blotting system at 30 V (Thermo Fisher Scientific). Immunoblotting was performed using the LI-COR system as previously (Stanley et al. 2016). Membranes were blocked for 1 h to overnight prior to antibody staining. The following antibodies were used with clone identifiers listed for the two monoclonal antibodies (Actin and Myc): Actin (mouse, 1:2000; Abcam, ab8226, mAbcam8226), DIS3 (rabbit, GR:195524-7, 1:500; Abcam, ab179933), Myc (mouse, clone 4A6, 1:1000; Millipore, 4A6), NAF1 (rabbit, 1:1000; Abcam, ab157106), RBM7 (rabbit, HPA013993, 1:250; Sigma), SKIV2L2 (rabbit, 1:500; Abcam, ab187884), Tubulin (rabbit, 1:5000; Abcam, ab6046), ZCCHC8 (mouse, 1:500; Abcam, ab68739). Membranes were stained with anti-mouse or anti-rabbit IRDye secondary antibodies (IR680 or IR800, donkey, 1:10,000 for both) before visualization on an Odyssey scanner (LI-COR). Quantification of protein levels was performed using an internal loading control (e.g., actin or tubulin) and then normalized relative to control sample. Band intensities were measured using ImageJ (Schneider et al. 2012).
Northern quantification of TR
We optimized a protocol for quantifying TR levels using a modified Northern method to maximize visualization of TR, a low-abundance RNA. Total RNA (0.75–6 µg) was mixed with RNA loading buffer (Sigma R4268-1VI) and heated to 95°C for 3 min, then cooled on ice for 3 min. Samples were then immediately loaded onto a prerun Novex 6% TBE-UREA gel and run at 180 V in heated TBE buffer (65°C) for 55 min on an Isotemp hot plate set to 70°C (Thermo Fisher Scientific). Gels were transferred at 20 V onto an Amerhsam Hybond-XL membrane for 90 min (GE Healthcare Life Sciences). The cross-linking, hybridization, and probe preparation were performed as previously described for hTR, mTR, and 5.8S (Stanley et al. 2016).
Cloning and site-directed mutagenesis
hZCCHC8 (NM_017612), hRBM7 (NM_001286045), hSKIV2L2 (NM_015360), hDIS3 (NM_014953.4), and mRBM7 (NM_144948), mSkiv2l2 (NM_028151), hPARN (NM_002582), hEXOSC10/RRP6 (NM_001001998) and were cloned from total cDNA with addition of a tag into a CMV promoter-driven pcDNA5/FRT/TO expression vector using restriction digestion and Gibson cloning (New England Biolabs). hZCCHC8, mZCCHC8, mRBM7, and mSKIV2L2 were all N terminus-tagged (Myc-Flag-Gly-) except that mZCCHC8 included Myc-Gly4-Ser- sequence. Myc-tagged mouse ZCCHC8 cDNA (NM_028151) was purchased in a pCMV3 expression vector (MG51487-NM, Sino Biological). RBM7, SKIV2L2, RRP6/EXOSC10, and PARN were C-terminal tagged (−Gly1-2-Myc-Flag). PrimerX was used to design primers for site-directed mutagenesis (http://www.bioinformatics.org/primerx). Plasmids were transfected using Lipofectamine 2000 or 3000 (Invitrogen) and protein lysates were harvested after 24–48 h.
Transduction, lentivirus production, and shRNA knockdown
Virus production in HEK-293FT cells, as well as the stable lentiviral shRNA knockdown in HeLa cells, were performed as previously described (Stanley et al. 2016). shRNAs were in PLKO.1 cloning vectors: Luciferase (SHC007, Sigma), NAF1 (TRCN000135721, Open Biosystems), ZCCHC8 (TRCN0000075158, Open Biosystems), RBM7 (TRCN0000074528, Open Biosystems), SKIV2L2 (TRCN0000051976, Open Biosystems).
Generation of ZCCHC8-null human cells
We used CRISPR/Cas9 editing to generate ZCCHC8 knockout HCT116 cells since they are pseudodiploid. The sgRNA (5′-CCCATCCTTCGGAAAGCTGA-3′, 20 bp) was cloned into the pX458 vector (gift from Feng Zhang, Addgene #62988) (Ran et al. 2013) and transfected into HCT116 cells. GFP-positive cells were flow-sorted in bulk and replated using single-cell dilutions. Mutants were screened by PCR and Sanger sequencing (F: 5′-ACACATCAAAGTCTGGCTCCTT-3′, R: 5′-CGGAAAGCTGAGGGTTTTC-3′).
Cell line authentication
HCT116 cells (gift from Dr. Ben Ho Park, Johns Hopkins University) were authenticated by short tandem-repeat profiling according to ATCC guidelines using the PowerPlex 16HS short tandem-repeat profiling kit (Promega) and matched ATCC CCL-247 HCT116 human colon carcinoma lines according to the ANSI/ATCC ASN-0002-2011 standard (ANSI eStandard) (Johns Hopkins Genetic Resources Core Facility).
3′ rapid amplification of cDNA ends sequencing (3′RACE-seq)
3′RACE for TR was performed as previously described (Goldfarb and Cech 2013; Moon et al. 2015) with modifications. Briefly, DNase I-treated (Qiagen) total RNA was ligated to 5 µM pre-adenylated linker (Universal miRNA Cloning Linker, New England Bio Labs) with 280 U of T4 RNA ligase 2, truncated KO (New England BioLabs) in a 20-µL reaction at 25°C for 16 h with RNaseOUT and 25% PEG8000 (New England BioLabs). The ligation reaction was then cleaned using RNA Clean and Concentrator (Zymo Research) and cDNA was synthesized with a universal RT primer (5′-CTACGTAACGATTGATGGTGCCTACAG) using the SuperScript III reverse transcriptase (Thermo Fisher Scientific). TR PCR was carried out using hTR_RACE_F and hTR_RACELinker_R (final 0.4 µM) (primer sequences in Supplemental Table S3), dNTPs (final 200 µM) and Q5 high-fidelity DNA Polymerase (at 0.02 U/µL) (New England Biolabs) with these parameters: 1 min at 98°C, 22 cycles of 10 sec at 98°C and 30 sec at 67°C, and a final extension for 2 min at 72°C. The product was then purified (QIAquick, Qiagen) and visualized on agarose gels for quality control prior to library preparation. Libraries were prepared using the TruSeq Nano DNA LT library preparation kit (Illumina) following the manufacturer's instructions but starting with adenylation of 3′ ends. Samples were quantified by Bioanalyzer diluted to 2 nM each, then pooled. The combined libraries, containing 15%–20% phiX control (Illumina), were then run on an Illumina MiSeq with 250 paired-end reads (Johns Hopkins Genetic Resources Core Facilities).
3′RACE-seq analysis
Reads were demultiplexed and the read 1 and read 2 fastq files for each sample were generated. Reads were trimmed using a custom script. To quantify the length and sequence of short extensions (defined as up to 15 bases beyond the 451st nucleotide of the mature hTR end with or without an oligoA tail), reads were mapped to the reference genomic TR locus (NR_001566) appended with the linker sequence using Bowtie2 (Langmead and Salzberg 2012) and extensions were determined as insertion blocks in the alignments. The mean number of reads mapped per sample was 200,745 (range, 79,000–301,000) and the median alignment rate was 80% (range, 42–89). Because of the limited amount of RNA, the 3′RACE-seq from the ZCCHC8 RNA IP had a lower alignment (17%), corresponding to 132,550 reads. Using a customized script, reads were then filtered to contain both at least 11 bases of the 3′ end of mature TR and at least nine of the 17 bases of the linker sequence. The relative abundance of different 3′ termini for TR was determined by normalizing to the percentage of total trimmed and filtered reads.
Long genomic extensions (i.e., >15 bases beyond the mature TR end) were quantified separately using Bowtie and Sim4db (Walenz and Florea 2011) to ensure mapping to the entire TR locus including ±5-kb sequences flanking the mature TR. SAMtools were subsequently used to extract reads corresponding to the extended region >15 bases beyond the end of mature TR. The mean number of reads mapped was 181,460 (range, 70,000–282,000) and the median alignment rate was 71% (range, 31–85). Percent alignment for RNA immunoprecipitate (IP) was 16% with 124,020 reads aligning to the target TR sequence. Customized scripts for TR 3′RACEp-seq analysis are available upon request.
Telomerase repeat amplification protocol (TRAP)
TRAP was performed as described previously (Kim et al. 1994). HCT116 cells were treated with CHAPS lysis buffer (104 cells/µL) and isolated protein was quantified using a BCA assay.
Protein and RNA IP
293FT cells were grown to 75%–90% confluence and transiently transfected with pcDNA5/FRT/TO/ZCCHC8-Myc-DDK plasmid using lipofectamine 2000 reagent (Thermo Fisher). After 48 h, cells were washed with PBS supplemented with protease inhibitors (Roche) and lysed cells in buffer (20 mM Tris-HCl at pH 7.5, 150 mM NaCl, 0.1% NP40, 2 mM MgCl2, 1× EDTA-free Halt protease, phosphatase inhibitor cocktail [ThermoFisher], RNAsin [Promega]). The lysate was rotated end to end for 15 min at 4°C and clarified by spinning at 12.3 rpm for 10 min at 4°C. The supernatant lysate was incubated with anti-c-Myc magnetic beads (Pierce) for 2 h at room temperature, rotating end to end. After washing in 1× TBS-T (25 mM Tris, 0.15 M NaCl, 0.05% Tween-20) and lysis buffer, one-third of the sample was processed for protein IP and the rest for RNA IP. For protein IP, the bound protein was eluted with NuPAGE LDS sample buffer (4×) and then diluted to 2× with lysis buffer. For RNA IP, the beads were resuspended in TriZol, incubated overnight at −80°C, and RNA was extracted as a coprecipitate based on the manufacturer's protocol (Life Technologies). The isolated RNA was treated with DNase (Ambion, AM1906) and used as a template for cDNA synthesis using SuperScript III reverse transcriptase (Invitrogen) and random hexamer primer mix (Thermo Fisher Scientific). TR was then amplified by RT-PCR using the same primer set used for total TR quantification described elsewhere here. Alternatively total RNA was used for TR 3′RACE-seq.
Mouse study approval, maintenance, and genotyping
The mouse studies were reviewed and approved by the Johns Hopkins Institutional Animal Care and Use Committee and the procedures conformed with the Guide for the Care and Use of Laboratory Animals (National Research Council 2011). Mice were housed in the Johns Hopkins University School of Medicine East Baltimore campus. All the mice studied were on a pure C57BL/6J background. mTR-null mice were derived and maintained as previously described (Blasco et al. 1997). The derivation of Zcchc8-null mice is described below, and these mice were genotyped using primers listed in Supplemental Table S3 (mZcchc8_ Genotyping_F and mZcchc8_ Genotyping_R) which amplified 363- and 396-bp products for the wild-type and mutant allele, respectively. Sperm from mice carrying the null Zcchc8 allele was deposited in the Jackson Laboratories Biorepository (stock ID 404814).
Generation of Zcchc8-null mice
We designed a CRISPR/Cas9 strategy to disrupt the Zcchc8 allele by zygote injection of a sgRNA targeting exon 2 (5′-TGAACATTCTGACAAGACCC-3′, 20 bp), and a DNA oligo for homology directed repair (HDR) and Cas9 protein. The HDR oligo (5′-AAGAACTTAAAAGAAAGTTGAACATTCTGACAAGAGGATCCGATTACAAGGACGACGATGACAAGTAGCCCAGGTATGACATCTTGAAATTACACACGAGCTG-3′, 103 bp) targeted a 33-bp insertion using 35-bp flanking homology arms which introduced a premature stop codon at amino acid 93 (P83delinsGSDYKDDDDK*) as well as a BamHI site and Flag tag sequence (Supplemental Fig. S8A). To facilitate a genetic screen for the edited alleles, we used DDK sequence-specific primers that fell within the Flag tag sequence (Supplemental Fig. S8A). Injections into C57BL/6J zygotes were performed at the Johns Hopkins Transgenic Mouse Core Facility. Founder mice were backcrossed to wild-type C57BL/6J twice prior to interbreeding of Zcchc8+/− mice.
Mouse phenotyping and pathology
Mouse ear fibroblasts (Merfs) were derived and maintained as described previously (Stanley et al. 2016). For histopathology studies, mice were individually euthanized by exposure to gradually increasing concentrations of CO2, then perfusion with heparinized saline followed by 10% NBF via left cardiac ventricle. Perfused tissues were fixed in 10% NBF for at least 24 h prior to trimming. The head and hindlimb were separately decalcified with Formical 4 (StatLab Medical Products). Gross examination and tissue collection was performed as described in Brayton et al. (2014). Sections (∼5 µm) were stained with hematoxylin and eosin and reviewed by two veterinary pathologists (B. Kang and C. Brayton). Image capture was performed on a Nikon 55i microscope and slides were scanned at 20x on an Aperio AT2 instrument (Leica Biosystems). Immunohistochemistry was performed using standard methods on formalin-fixed paraffin-embedded tissues using these antibodies: Ki67 (SP6, Abcam, ab16667), Cleaved caspase 3 (Asp175, 5A1E, rabbit mAb #9664, Cell Signaling Technology), and ZCCHC8 (R34469, polyclonal rabbit, Sigma-Aldrich).
Computed tomography (CT) imaging and image analysis
Mice were anesthetized using isoflurane prior to and during imaging. Gross images of the mice's heads were taken using a phone camera. CT imaging was performed using the nano PET/CT Small Animal Imager (Mediso) (Johns Hopkins Center for Infection and Inflammation Imaging Research). CT images were visualized on RadiAnt DICOM Viewer (v.4.6.5.18450, 64-bit, Medixant). Using the CT Bone setting, mid-sagittal CT images were analyzed using the 3D Multiplanar Reconstruction viewer to align the axes to the middle of the nasal cavity and incisors in the coronal and axial settings, respectively. Virtual reconstruction CT images were generated using the 3D Volume Rendering function and saved in the Bones and Skin 3 setting. Cranial area was measured as a surrogate for brain volume using the polygon selection tool of ImageJ. Using the 3D maximum intensity projection setting, the left femur length of each pup was measured. To ensure there are no sex-specific phenotypes, for newborn pup imaging studies we assessed male/female status by tail DNA PCR amplification as described in McFarlane et al. (2013).
RNA isolation for RNA-seq, library preparation, and sequencing
E12.5 brains were isolated and the tissue was placed into RNAlater (Qiagen) then homogenized in a Bullet Blender using zirconium oxide-coated beads (Next Advance). RNA was prepared using RNAeasy kit (Qiagen). Libraries were prepared for RNA-seq using the standard protocol for Truseq Stranded mRNA Library preparation (Illumina). The sequencing was completed using NovaSeq S1 flowcell paired-end 150 bp. This generated 253 million 150-bp-long paired-end reads per sample (range 195–321 million), of which 95.8% on average (range 90.9%–96.6%) mapped to the mouse genome following the procedure described below.
RNA-seq bioinformatics
Transcriptomic data collected by RNA-seq were analyzed to determine the genes that are present in each sample, their expression levels, and the differences between expression levels among different genotypes. For alignment and annotation, following quality checking with the software FastQC (v. 0.10.0; https://wwwbioinformaticsbabrahamacuk/projects/fastqc), reads were end-trimmed to 100 bp and mapped to the mouse genome version mm10 with the alignment tool Tophat2 v.2.1.0 (Kim et al. 2013). Between 91% and 97% of reads per sample mapped to the mouse genome, of which a very small fraction (1.5%–1.9%) had multiple matches, and the concordant read fraction was 87.8%–94.4%. The aligned reads were assembled with CLASS2 v.2.1.7 (Song et al. 2016) to create partial gene and transcript models (transfrags). Transfrags from all samples were further merged with Cuffmerge (v. 2.2.1) (Trapnell et al. 2010) and mapped to the GENCODE v.M17 (https://www.genecodegenes.org) gene models, to create a unified set of gene annotations for differential analyses. Gene and transcript expression levels (in FPKM) were then computed with the tool Cuffdiff2 v.2.2.1 (Trapnell et al. 2013) and differentially expressed genes (transcripts) were determined by statistical analysis. The alignment and annotation were performed in August 2018.
Gene expression analysis
Differential gene expression analyses were performed using the FPKM files comprising 67,811 transcripts, each generated as described above for each embryonic sample. The transcript identifiers were derived from these FPKM files’ “gene” column and updated to current nomenclature. The raw FPKM values of 0.0 were treated as nulls and actual values were transformed into log2 annotation and quantile normalized across the 14 samples. The three biological classes’ (KO, HET, and WT) normalized signals underwent differential expression analysis with a two-tailed one-way t-test ANOVA using the Partek GS 6.16.0812 platform. A standard deviation analysis was performed for each class–class comparison using those transcripts that had high-quality data: an NCBI Entrez gene ID, a mean FPKM linear value >2.0 in at least one cell class, and a nonzero FPKM signal value from all the 14 samples. On the order of 9500] transcripts met these three criteria for each cell–class comparison and were used for subsequent downstream functional analyses. The raw data were deposited in the National Center for Biotechnology Information Gene Expression Omnibus database GEO (GSE126108).
RNA-seq 3′-end analysis
To elucidate the presence of 3′ extensions for mTR and mRmrp, and for relative abundance comparisons among Zcchc8+/+, Zcchc8+/−, and Zcchc8−/− mice (14 samples), we extracted RNA-seq read pairs partially aligning to the gene from the mouse RNA-seq Fastq files generated as part of the RNA-seq bioinformatic analysis. Reads were then processed using the protocol described for 3′RACE-seq data with some modifications, described herein. Briefly, reads (and fragments) mapping to the target gene were determined with Bowtie2 (Langmead and Salzberg 2012) with the option “-local”. Reads were then trimmed of Illumina adapters using the tool “cutadapt” (Martin 2011), and an artificial “tag” consisting of the 28-bp linker sequence ACTGTAGGCACCATCAATCGTTACGTAG was added to the 3′ ends of forward mapping reads, whereas the reverse complement linker sequence was prepended to the 5′ end of the reverse mapping reads. Similarly, the linker sequence was appended to the 3′ end of the gene(s) to generate reference sequences. Unlike with 3′RACE-seq reads, where the start of the linker marked the precise position of the gene's end, here the linker merely indicates that the end of the gene is at or downstream from the mapping location of the read. Hence, reads spanning the gene-linker junction collectively encompass 3′ RNA extensions, both adenylated and genomic, as well as potential genomic and other types of contamination. The relative abundance of these reads was then compared across the genotypes.
Statistical analyses
Single-point parametric linkage was performed using logarithm of the odds scores. qRT-PCR analyses were reported as standard error of the mean (SEM), and P-values were calculated using GraphPad Prism software. For each PCR run, samples were done in triplicate. All P-values shown are two-sided. The number of replicates and statistical tests used are indicated in the text and legends with the respective data.
Supplementary Material
Acknowledgments
We are indebted to the proband and family for their generosity, time and participation. We are grateful to Carolyn Applegate for help with the clinical characterization, Emily McNally with the targeted nextgen sequencing and Corina Antonescu for help with the RNA-seq bioinformatics. We appreciate comments on the manuscript from Carol Greider, Jeffry Corden, and Andrew Holland. We appreciate support from the Mouse Transgenic Core, the Genetic Center Resource Facility, especially the Cell Center team, Phenotyping Core, Computational Biology Core, the Oncology Tissue Services Core, and the Center for Infection and Inflammation Imaging Research, all at Johns Hopkins University School of Medicine. This work was supported by National Institutes of Health grants RO1CA225027 and RO1HL119476, an award from the Maryland Cigarette Restitution Fund, the Commonwealth Foundation, the Gary Williams Foundation, and an award from the S&R Foundation Kuno Award (to M.A.). D.L.G. and S.E.S. received support from T32GM007309, and D.L.G. was supported by the Turock Scholars Fund to the Telomere Center at Johns Hopkins. V.G. received support from T32HL007534 and F32HL142207 and L.F. from National Science Foundation ABI-1356078. M.A. acknowledges a gift to the Telomere Center in the name of Mrs. P. Godrej. The Oncology Tissue Services Core was supported by P30CA006973.
Author contributions: D.L.G. identified the disease gene, generated the null cell lines, and the knockout mouse and designed, performed, and analyzed other experiments. V.G. designed, performed and analyzed the 3′RACE-seq, immunoprecipitation experiments and analyzed the RNA-seq data. C.C.A. performed the mouse imaging studies and characterized mouse phenotypes with D.L.G. C.C.T. analyzed RNA-seq data with V.G. and D.L.G. B.K. and C.B. performed the mouse histopathology analyses. S.E.S. analyzed clinical data, identified the proband in a screen for low TR families and analyzed whole-genome sequencing data. E.W.P. analyzed the SNP array data and performed the linkage analysis. N.A.C. performed transfection, qPCR, and immunoblot experiments. K.M.S. performed the P186L transfection experiment. M.O.A. analyzed clinical data. L.F. implemented the 3′RACE-seq protocol with V.G. and performed the annotation and differential expression analysis of the RNA-seq. M.A. oversaw all aspects of the project, reviewed the raw data, and wrote the manuscript with D.L.G. and V.G.
Footnotes
Supplemental material is available for this article.
Article published online ahead of print. Article and publication date are online at http://www.genesdev.org/cgi/doi/10.1101/gad.326785.119.
Freely available online through the Genes & Development Open Access option.
References
- Abecasis GR, Cherny SS, Cookson WO, Cardon LR. 2002. Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 30: 97–101. 10.1038/ng786 [DOI] [PubMed] [Google Scholar]
- Alder JK, Parry EM, Yegnasubramanian S, Wagner CL, Lieblich LM, Auerbach R, Auerbach AD, Wheelan SJ, Armanios M. 2013. Telomere phenotypes in females with heterozygous mutations in the dyskeratosis congenita 1 (DKC1) gene. Hum Mutat 34: 1481–1485. 10.1002/humu.22397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alder JK, Stanley SE, Wagner CL, Hamilton M, Hanumanthu VS, Armanios M. 2015. Exome sequencing identifies mutant TINF2 in a family with pulmonary fibrosis. Chest 147: 1361–1368. 10.1378/chest.14-1947 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alder JK, Hanumanthu VS, Strong MA, DeZern AE, Stanley SE, Takemoto CM, Danilova L, Applegate CD, Bolton SG, Mohr DW, et al. 2018. Diagnostic utility of telomere length testing in a hospital-based setting. Proc Natl Acad Sci 115: E2358–E2365. 10.1073/pnas.1720427115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersen PR, Domanski M, Kristiansen MS, Storvall H, Ntini E, Verheggen C, Schein A, Bunkenborg J, Poser I, Hallais M, et al. 2013. The human cap-binding complex is functionally connected to the nuclear RNA exosome. Nat Struct Mol Biol 20: 1367–1376. 10.1038/nsmb.2703 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Armanios M. 2012. Telomerase and idiopathic pulmonary fibrosis. Mutat Res 730: 52–58. 10.1016/j.mrfmmm.2011.10.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Armanios MY, Chen JJ, Cogan JD, Alder JK, Ingersoll RG, Markin C, Lawson WE, Xie M, Vulto I, Phillips JA III, et al. 2007. Telomerase mutations in families with idiopathic pulmonary fibrosis. N Engl J Med 356: 1317–1326. 10.1056/NEJMoa066157 [DOI] [PubMed] [Google Scholar]
- Armanios M, Alder JK, Parry EM, Karim B, Strong MA, Greider CW. 2009. Short telomeres are sufficient to cause the degenerative defects associated with aging. Am J Hum Genet 85: 823–832. 10.1016/j.ajhg.2009.10.028 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baerlocher GM, Vulto I, de Jong G, Lansdorp PM. 2006. Flow cytometry and FISH to measure the average length of telomeres (flow FISH). Nat Protoc 1: 2365–2376. 10.1038/nprot.2006.263 [DOI] [PubMed] [Google Scholar]
- Blasco MA, Lee HW, Hande MP, Samper E, Lansdorp PM, DePinho RA, Greider CW. 1997. Telomere shortening and tumor formation by mouse cells lacking telomerase RNA. Cell 91: 25–34. 10.1016/S0092-8674(01)80006-4 [DOI] [PubMed] [Google Scholar]
- Box JA, Bunch JT, Tang W, Baumann P. 2008. Spliceosomal cleavage generates the 3′ end of telomerase RNA. Nature 456: 910–914. 10.1038/nature07584 [DOI] [PubMed] [Google Scholar]
- Brayton C, McKerlie C, Brown S. 2014. Analysis of the phenotype. In Transgenic animal technology, 3rd ed. (ed. Pinkert CA), pp. 431–487. Elsevier, London, UK: 10.1016/B978-0-12-410490-7.00016-5 [DOI] [Google Scholar]
- Chen JL, Blasco MA, Greider CW. 2000. Secondary structure of vertebrate telomerase RNA. Cell 100: 503–514. 10.1016/S0092-8674(00)80687-X [DOI] [PubMed] [Google Scholar]
- Cogan JD, Kropski JA, Zhao M, Mitchell DB, Rives L, Markin C, Garnett ET, Montgomery KH, Mason WR, McKean DF, et al. 2015. Rare variants in RTEL1 are associated with familial interstitial pneumonia. Am J Respir Crit Care Med 191: 646–655. 10.1164/rccm.201408-1510OC [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coy S, Volanakis A, Shah S, Vasiljeva L. 2013. The Sm complex is required for the processing of non-coding RNAs by the exosome. PLoS One 8: e65606 10.1371/journal.pone.0065606 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deng T, Huang Y, Weng K, Lin S, Li Y, Shi G, Chen Y, Huang J, Liu D, Ma W, et al. 2019. TOE1 acts as a 3′ exonuclease for telomerase RNA and regulates telomere maintenance. Nucleic Acids Res 47: 391–405. 10.1093/nar/gky1019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng J, Funk WD, Wang SS, Weinrich SL, Avilion AA, Chiu CP, Adams RR, Chang E, Allsopp RC, Yu J, et al. 1995. The RNA component of human telomerase. Science 269: 1236–1241. 10.1126/science.7544491 [DOI] [PubMed] [Google Scholar]
- Giunta M, Edvardson S, Xu Y, Schuelke M, Gomez-Duran A, Boczonadi V, Elpeleg O, Muller JS, Horvath R. 2016. Altered RNA metabolism due to a homozygous RBM7 mutation in a patient with spinal motor neuropathy. Hum Mol Genet 25: 2985–2996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Glaunsinger B, Ganem D. 2004. Lytic KSHV infection inhibits host gene expression by accelerating global mRNA turnover. Mol Cell 13: 713–723. 10.1016/S1097-2765(04)00091-7 [DOI] [PubMed] [Google Scholar]
- Goldfarb KC, Cech TR. 2013. 3′ terminal diversity of MRP RNA and other human noncoding RNAs revealed by deep sequencing. BMC Mol Biol 14: 23 10.1186/1471-2199-14-23 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greider CW. 2006. Telomerase RNA levels limit the telomere length equilibrium. Cold Spring Harb Symp Quant Biol 71: 225–229. 10.1101/sqb.2006.71.063 [DOI] [PubMed] [Google Scholar]
- Greider CW, Blackburn EH. 1985. Identification of a specific telomere terminal transferase activity in Tetrahymena extracts. Cell 43: 405–413. 10.1016/0092-8674(85)90170-9 [DOI] [PubMed] [Google Scholar]
- Greider CW, Blackburn EH. 1987. The telomere terminal transferase of Tetrahymena is a ribonucleoprotein enzyme with two kinds of primer specificity. Cell 51: 887–898. 10.1016/0092-8674(87)90576-9 [DOI] [PubMed] [Google Scholar]
- Greider CW, Blackburn EH. 1989. A telomeric sequence in the RNA of Tetrahymena telomerase required for telomere repeat synthesis. Nature 337: 331–337. 10.1038/337331a0 [DOI] [PubMed] [Google Scholar]
- Hamosh A, Sobreira N, Hoover-Fong J, Sutton VR, Boehm C, Schiettecatte F, Valle D. 2013. PhenoDB: a new web-based tool for the collection, storage, and analysis of phenotypic features. Hum Mutat 34: 566–571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hao LY, Armanios M, Strong MA, Karim B, Feldser DM, Huso D, Greider CW. 2005. Short telomeres, even in the presence of telomerase, limit tissue renewal capacity. Cell 123: 1121–1131. 10.1016/j.cell.2005.11.020 [DOI] [PubMed] [Google Scholar]
- Hathcock KS, Hemann MT, Opperman KK, Strong MA, Greider CW, Hodes RJ. 2002. Haploinsufficiency of mTR results in defects in telomere elongation. Proc Natl Acad Sci 99: 3591–3596. 10.1073/pnas.012549799 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jonassaint NL, Guo N, Califano JA, Montgomery EA, Armanios M. 2013. The gastrointestinal manifestations of telomere-mediated disease. Aging Cell 12: 319–323. 10.1111/acel.12041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim NW, Piatyszek MA, Prowse KR, Harley CB, West MD, Ho PL, Coviello GM, Wright WE, Weinrich SL, Shay JW. 1994. Specific association of human telomerase activity with immortal cells and cancer. Science 266: 2011–2015. 10.1126/science.7605428 [DOI] [PubMed] [Google Scholar]
- Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. 2013. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14: R36 10.1186/gb-2013-14-4-r36 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kousi M, Katsanis N. 2016. The genetic basis of hydrocephalus. Annu Rev Neurosci 39: 409–435. 10.1146/annurev-neuro-070815-014023 [DOI] [PubMed] [Google Scholar]
- Langmead B, Salzberg SL. 2012. Fast gapped-read alignment with Bowtie 2. Nat Methods 9: 357–359. 10.1038/nmeth.1923 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lingner J, Hughes TR, Shevchenko A, Mann M, Lundblad V, Cech TR. 1997. Reverse transcriptase motifs in the catalytic subunit of telomerase. Science 276: 561–567. 10.1126/science.276.5312.561 [DOI] [PubMed] [Google Scholar]
- Lloret-Llinares M, Karadoulama E, Chen Y, Wojenski LA, Villafano GJ, Bornholdt J, Andersson R, Core L, Sandelin A, Jensen TH. 2018. The RNA exosome contributes to gene expression regulation during stem cell differentiation. Nucleic Acids Res 46: 11502–11513. 10.1093/nar/gky817 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lubas M, Christensen MS, Kristiansen MS, Domanski M, Falkenby LG, Lykke-Andersen S, Andersen JS, Dziembowski A, Jensen TH. 2011. Interaction profiling identifies the human nuclear exosome targeting complex. Mol Cell 43: 624–637. 10.1016/j.molcel.2011.06.028 [DOI] [PubMed] [Google Scholar]
- Lubas M, Andersen PR, Schein A, Dziembowski A, Kudla G, Jensen TH. 2015. The human nuclear exosome targeting complex is loaded onto newly synthesized RNA to direct early ribonucleolysis. Cell Rep 10: 178–192. 10.1016/j.celrep.2014.12.026 [DOI] [PubMed] [Google Scholar]
- Lyons SM, Cunningham CH, Welch JD, Groh B, Guo AY, Wei B, Whitfield ML, Xiong Y, Marzluff WF. 2016. A subset of replication-dependent histone mRNAs are expressed as polyadenylated RNAs in terminally differentiated tissues. Nucleic Acids Res 44: 9190–9205. 10.1093/nar/gkw418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin M. 2011. Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnetjournal 17: 10–12. [Google Scholar]
- Marzluff WF, Gongidi P, Woods KR, Jin J, Maltais LJ. 2002. The human and mouse replication-dependent histone genes. Genomics 80: 487–498. 10.1006/geno.2002.6850 [DOI] [PubMed] [Google Scholar]
- Marzluff WF, Wagner EJ, Duronio RJ. 2008. Metabolism and regulation of canonical histone mRNAs: life without a poly(A) tail. Nat Rev Genet 9: 843–854. 10.1038/nrg2438 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McFarlane L, Truong V, Palmer JS, Wilhelm D. 2013. Novel PCR assay for determining the genetic sex of mice. Sex Dev 7: 207–211. 10.1159/000348677 [DOI] [PubMed] [Google Scholar]
- McNally EJ, Luncsford PJ, Armanios M. 2019. Long telomeres and cancer risk: the price of cellular immortality. J Clin Invest 130: 120851 10.1172/JCI120851 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchell JR, Cheng J, Collins K. 1999a. A box H/ACA small nucleolar RNA-like domain at the human telomerase RNA 3′ end. Mol Cell Biol 19: 567–576. 10.1128/MCB.19.1.567 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mitchell JR, Wood E, Collins K. 1999b. A telomerase component is defective in the human disease dyskeratosis congenita. Nature 402: 551–555. 10.1038/990141 [DOI] [PubMed] [Google Scholar]
- Moon DH, Segal M, Boyraz B, Guinan E, Hofmann I, Cahan P, Tai AK, Agarwal S. 2015. Poly(A)-specific ribonuclease (PARN) mediates 3′-end maturation of the telomerase RNA component. Nat Genet 47: 1482–1488. 10.1038/ng.3423 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller JS, Giunta M, Horvath R. 2015. Exosomal protein deficiencies: How abnormal RNA metabolism results in childhood-onset neurological diseases. J Neuromuscul Dis 2: S31–S37. 10.3233/JND-150086 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Najmabadi H, Hu H, Garshasbi M, Zemojtel T, Abedini SS, Chen W, Hosseini M, Behjati F, Haas S, Jamali P, et al. 2011. Deep sequencing reveals 50 novel genes for recessive cognitive disorders. Nature 478: 57–63. 10.1038/nature10423 [DOI] [PubMed] [Google Scholar]
- National Research Council. 2011. Guide for the care and use of laboratory animals (8th ed). The National Academies Press, Washington, DC. [Google Scholar]
- Nguyen D, Grenier St-Sauveur V, Bergeron D, Dupuis-Sandoval F, Scott MS, Bachand F. 2015. A polyadenylation-dependent 3′ end maturation pathway is required for the synthesis of the human telomerase RNA. Cell Rep 13: 2244–2257. 10.1016/j.celrep.2015.11.003 [DOI] [PubMed] [Google Scholar]
- Penno MB, Pedrotti-Krueger M, Ray T. 1993. Cryopreservation of whole blood and isolated lymphocytes for B-cell immortalization. J Tissue Cult Methods 15: 43–47. 10.1007/BF02387289 [DOI] [Google Scholar]
- Podlevsky JD, Chen JJ. 2016. Evolutionary perspectives of telomerase RNA structure and function. RNA Biol 13: 720–732. 10.1080/15476286.2016.1205768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. 2013. Genome engineering using the CRISPR-Cas9 system. Nat Protoc 8: 2281–2308. 10.1038/nprot.2013.143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roake CM, Chen L, Chakravarthy AL, Ferrell JJ Jr, Raffa GD, Artandi SE. 2019. Disruption of telomerase RNA maturation kinetics precipitates disease. Mol Cell 74: 688–700.e3. 10.1016/j.molcel.2019.02.033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneider CA, Rasband WS, Eliceiri KW. 2012. NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9: 671–675. 10.1038/nmeth.2089 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shukla S, Schmidt JC, Goldfarb KC, Cech TR, Parker R. 2016. Inhibition of telomerase RNA decay rescues telomerase deficiency caused by dyskerin or PARN defects. Nat Struct Mol Biol 23: 286–292. 10.1038/nsmb.3184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song L, Sabunciyan S, Florea L. 2016. CLASS2: accurate and efficient splice variant annotation from RNA-seq reads. Nucleic Acids Res 44: e98 10.1093/nar/gkw158 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stanley SE, Chen JJ, Podlevsky JD, Alder JK, Hansel NN, Mathias RA, Qi X, Rafaels NM, Wise RA, Silverman EK, et al. 2015. Telomerase mutations in smokers with severe emphysema. J Clin Invest 125: 563–570. 10.1172/JCI78554 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stanley SE, Gable DL, Wagner CL, Carlile TM, Hanumanthu VS, Podlevsky JD, Khalil SE, DeZern AE, Rojas-Duran MF, Applegate CD, et al. 2016. Loss-of-function mutations in the RNA biogenesis factor NAF1 predispose to pulmonary fibrosis-emphysema. Sci Transl Med 8: 351ra107 10.1126/scitranslmed.aaf7837 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stuart BD, Choi J, Zaidi S, Xing C, Holohan B, Chen R, Choi M, Dharwadkar P, Torres F, Girod CE, et al. 2015. Exome sequencing links mutations in PARN and RTEL1 with familial pulmonary fibrosis and telomere shortening. Nat Genet 47: 512–517. 10.1038/ng.3278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swiderski RE, Nakano Y, Mullins RF, Seo S, Bánfi B. 2014. A mutation in the mouse ttc26 gene leads to impaired hedgehog signaling. PLoS Genet 10: e1004689 10.1371/journal.pgen.1004689 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, Salzberg SL, Wold BJ, Pachter L. 2010. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28: 511–515. 10.1038/nbt.1621 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trapnell C, Hendrickson DG, Sauvageau M, Goff L, Rinn JL, Pachter L. 2013. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat Biotechnol 31: 46–53. 10.1038/nbt.2450 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsakiri KD, Cronkhite JT, Kuan PJ, Xing C, Raghu G, Weissler JC, Rosenblatt RL, Shay JW, Garcia CK. 2007. Adult-onset pulmonary fibrosis caused by mutations in telomerase. Proc Natl Acad Sci 104: 7552–7557. 10.1073/pnas.0701009104 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tseng CK, Wang HF, Burns AM, Schroeder MR, Gaspari M, Baumann P. 2015. Human telomerase RNA processing and quality control. Cell Rep 13: 2232–2243. 10.1016/j.celrep.2015.10.075 [DOI] [PubMed] [Google Scholar]
- Wagner CL, Hanumanthu VS, Talbot CJ Jr, Abraham RS, Hamm D, Gable DL, Kanakry CG, Applegate CD, Siliciano J, Jackson JB, et al. 2018. Short telomere syndromes cause a primary T cell immunodeficiency. J Clin Invest 128: 5222–5234. 10.1172/JCI120216 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walenz B, Florea L. 2011. Sim4db and Leaff: utilities for fast batch spliced alignment and sequence indexing. Bioinformatics 27: 1869–1870. 10.1093/bioinformatics/btr285 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waterhouse AM, Procter JB, Martin DM, Clamp M, Barton GJ. 2009. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25: 1189–1191. 10.1093/bioinformatics/btp033 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang YJ, O'Neal WK, Randell SH, Blackburn K, Moyer MB, Boucher RC, Ostrowski LE. 2002. Identification of dynein heavy chain 7 as an inner arm component of human cilia that is synthesized but not assembled in a case of primary ciliary dyskinesia. J Biol Chem 277: 17906–17915. 10.1074/jbc.M200348200 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.