Abstract
Doublecortin (DCX) is a microtubule-associated protein required for neuronal migration to the cerebral cortex. DCAMKL1 consists of an N terminus that is 65% similar to DCX throughout the entire length of DCX, but also contains an additional 360 amino acid C-terminal domain encoding a putative Ca2+/calmodulin-dependent protein kinase. The homology to DCX suggested that DCAMKL1 may regulate microtubules, as well as mediate a phosphorylation-dependent signal transduction pathway. Here we show that DCAMKL1 is expressed throughout the CNS and PNS in migrating neuronal populations and overlaps in its expression with DCX and microtubules. Purified DCAMKL1 associates with microtubules and stimulates polymerization of purified tubulin and the formation of aster-like microtubule structures. Overexpressed DCAMKL1 leads to striking microtubule bundling in cell lines and cultured primary neural cells. Time-lapse imaging of cells transfected with a DCAMKL1-green fluorescent protein fusion protein shows that the microtubules associated with the protein remain dynamic. DCAMKL1 also encodes a functional kinase capable of phosphorylating myelin basic protein and itself. However, elimination of the kinase activity of DCAMKL1 has no detectable effect on its microtubule polymerization activity. Because DCAMKL1 is coexpressed with DCX, the two proteins form a potentially mutually regulatory network linking calcium signaling and microtubule dynamics.
Keywords: neuronal migration, microtubule, kinase, microtubule-associated protein, doublecortin, lissencephaly
Dynamic rearrangements of microtubules are required for many aspects of neuronal development and function, including migration, process outgrowth, and synaptic plasticity (Matus, 1994; Brandt, 1996; Baas, 1999). The role of microtubules in neuronal process outgrowth has been best studied in terms of specialized neuronal microtubule-associated proteins (MAPs) such as tau and MAP2 (Caceres and Kosik, 1990; Caceres et al., 1992;Brandt, 1996). These proteins bind microtubules through specialized microtubule-binding domains (Goode and Feinstein, 1994; Felgner et al., 1997), regulate microtubule polymerization (Kosik, 1993; Avila et al., 1994; Brandt, 1996), and are essential for neurite outgrowth (Caceres and Kosik, 1990; Kosik and Caceres, 1991; Kosik, 1993). Microtubules are normally dynamically unstable, undergoing rounds of polymerization and depolymerization (Mitchison and Kirschner, 1984; Desai and Mitchison, 1997). The MAPs do not abolish this dynamic instability, but induce a relative increase in the polymerization phase (Kaech et al., 1996).
The requirement for distinct MAPs in neuronal migration has been more recently recognized. For example, LIS1 mutations cause human lissencephaly (“smooth brain”) in which the cerebral cortex shows massive arrest of neuronal migration (Dobyns et al., 1993). LIS1 encodes a protein without obvious microtubule-binding domains (Reiner et al., 1993). Nonetheless, LIS1 appears to bind microtubules in some settings, and to regulate microtubule catastrophe events (Sapir et al., 1997, 1999). Mutations in a second gene, doublecortin (des Portes et al., 1998; Gleeson et al., 1998; Sossey-Alaoui et al., 1998), also cause lissencephaly that is similar to that seen with LIS1 mutations (Berg et al., 1998; Pilz et al., 1998). Doublecortin also binds microtubules and regulates their polymerization, and also fulfills the criteria of a MAP (Francis et al., 1999; Gleeson et al., 1999; Horesh et al., 1999). Doublecortin (DCX) shows no sequence homology to LIS1 and also shows no obvious microtubule-binding motif. The N terminus of DCX is required for microtubule binding (Horesh et al., 1999; Sapir et al., 2000; Taylor et al., 2000), suggesting that it defines a new microtubule-binding structure.
The closet single homolog to DCX in GenBank (des Portes et al., 1998;Gleeson et al., 1998), called KIAA0369 (now referred to as DCAMKL1), also is a protein of unknown function (Nagase et al., 1998). DCX and DCAMKL1 share homology throughout the entire DCX sequence, but DCAMKL1 is twice as large. The unique C terminus of DCAMKL1 encodes a domain similar to Ca2+/calmodulin-dependent (CAM) kinases. Both DCX and DCAMKL1 are highly expressed in developing brain (Omori et al., 1998; Sossey-Alaoui et al., 1998; Burgess et al., 1999;Sossey-Alaoui and Srivastava, 1999), suggesting that they may function together in migrating cerebral cortical neurons.
Here we show that DCAMKL1 is expressed in a pattern that overlaps with the expression of DCX in migrating neurons. We also show that DCAMKL1 has microtubule binding and polymerization activities. Although full-length DCAMKL1 retains kinase activity for itself and for myelin basic protein (MBP), its effects on microtubules are not detectably dependent on its kinase activity. These data suggest that DCAMKL1 and DCX may together form a signaling pathway that regulates microtubules in migrating neurons.
MATERIALS AND METHODS
Overexpression of DCAMKL1. Human DCAMKL1was cloned by RT-PCR using primers directed toward the human sequence and was subsequently sequenced. The isoform isolated corresponds to the K369-A variant of Omori et al. (1998). Full-length humanDCAMKL1 was subsequently cloned into the KpnI site of pcDNA3.1C(−) (Invitrogen, Carlsbad, CA) and overexpressed in COS7 cells by transient transfection with Superfectamine (Qiagen, Chatsworth, CA) according to the manufacturer's recommendations. Cells were harvested 2 d later by adding boiling protein sample buffer (20 mm Tris, 5% glycerol, 0.625% SDS, and 5% B-mercaptoethanol).
Generation of DCAMKL1 antisera. To generate antisera reactive to both human and mouse DCAMKL1 protein, a 15 mer DCAMKL1 N-terminal polypeptide (MSFGRDMELEHFDER-CYS-amide) corresponding to both the mouse and human sequences was used as an immunogen. This N terminus shows only 2 of 15 shared amino acids (1 of 14 amino acids other than the initial methionine) with the N-terminal region of DCX, which was used for the creation of an anti-DCX antibody previously described (Gleeson et al., 1999). α−DCAMKL1 was used at 1:250 for Western analysis and 1:50 for immunofluorescence.
Western analysis. Total protein was run on a 10% SDS-PAGE gel, transferred to nitrocellulose membranes, then probed with either α−DCAMKL1 or α-pan MAP2 (1:500) and developed using an HRP anti-rabbit or anti-mouse secondary (Bio-Rad, Hercules, CA) at 1:3000 and ECL (Amersham, Arlington Heights, IL). Fifty micrograms of total protein were loaded for expression studies, and 50% of the pellet was loaded for DCAMKL1 microtubule experiments. Blots were stripped and probed with β-tubulin to control for protein loading.
Immunohistochemistry. Mouse brains were perfusion-fixed in 4% paraformaldehyde in PBS, pH 7.4, followed by immersion in 30% sucrose in PBS until the brains had sunk. Brains were frozen in OCT compound and sectioned on a Leica freezing microtome at 15 μm. Sections were incubated overnight with primary rabbit antiserum. Sections were then rinsed with PBS and incubated with Cy3-labeled goat anti-rabbit secondary antibody (The Jackson Laboratory, Bar Harbor, ME) (diluted 1:300), rinsed several times in PBS, mounted, and examined under an Olympus fluorescent microscope. Negative controls were run in parallel using preimmune sera as the primary antisera. Colabeling was done with TuJ-1 (1:200) (Research Diagnostics, Flanders, NJ), calbindin (1:200) (Sigma, St. Louis, MO), GFAP (1:200) (Sigma), and Tag1 (1:200) (Developmental Hybridoma Studies Bank at the University of Iowa, Iowa City, IA). Colabeled sections were incubated with FITC-conjugated anti-mouse secondary antibody (The Jackson Laboratory), diluted 1:75.
Imprint assay. Imprint assays were performed as previously described (Anton et al., 1999). E17 mouse cortex was removed and sectioned at 75 μm, incubated in CMF-PBS with 10 U/ml papain for 10 min, then in MEM with 2 mg/ml ovomucoid inhibitor and 100 mg/ml DNase for 10 min, washed in media, and placed overnight on glass coverslips that had been treated with Cell-Tak (Becton Dickinson). The cortex was removed with excess media, and cortical imprints containing radial migration units were observed and fixed for subsequent immunofluorescence. Imprints were stained with DCAMKL1 antisera and Rat-401 antibody (Developmental Studies Hybridoma Bank) and counterstained with DAPI.
Immunofluorescence of cultured neurons and transfection of primary cortical cultures. Primary cortical neurons were harvested from embryonic day 17 (E17) rats, dissociated in papain, plated at a density of 5 × 105/ml of culture media in MEM with 10% fetal calf serum, and cultured for 2 d on microwell poly-d-lysine-coated coverslips. Colchicine (Sigma) (10 μg/ml) or vehicle was added for 0–3 hr. Cold-treated cells were placed at 0°C in an ice-water bath for 30 min. Cells were rinsed briefly in K-PIPES (in mm: 80 K-PIPES, pH 6.8, 5 EGTA, and 2 MgCl2); fixed in 0.5% glutaraldehyde with 0.1% Triton X-100; quenched in 1 mg/ml NaBH4; blocked with 1% BSA, 0.25% saponin, and 5 mm lysine in PBS for 10 min; incubated with primary antibodies as above for 1 hr; rinsed extensively with K-PIPES; and incubated with Cy3- or FITC-conjugated secondary antibody (diluted 1:50) for 1 hr; rinsed extensively in K-PIPES; and post-fixed in 4% paraformaldehyde for 30 min. Cells were then mounted in Aquamount and examined using a Bio-Rad 1100 confocal microscope.
Isolation of microtubules from brain. Four grams of newborn rat brain was homogenized in MES buffer (in mm: 100 MES, pH 6.6, 1 EGTA, 1 MgSO4, 25 NaF, and benzamidine, leupeptin, pepstatin A, aprotinin, and AEBSF protease inhibitors) at 4°C. The homogenate was centrifuged at 25,000 rpm for 15 min in a Beckman Optima centrifuge in the TLA 100.3 rotor, and the supernatant was subsequently centrifuged similarly at 75,000 rpm for 90 min, giving a tubulin-rich supernatant. Half of the supernatant was treated with 10 μm taxol (Calbiochem, La Jolla, CA) and 1 mm GTP, and half was treated with only 1 mm GTP. Both fractions were incubated for 25 min at 37°C to allow microtubules to polymerize and added over one volume of a 10% sucrose–MES buffer in a centrifuge tube, and centrifuged at 25,000 rpm at 35°C for 30 min. The supernatant and microtubule pellet were boiled in sample buffer and analyzed by Western blot for the presence of DCAMKL1, MAP2, and tubulin.
Purification of recombinant DCAMKL1 protein. To generate recombinant protein, DCAMKL1 was cloned into theBamHI–HindIII sites of the pET21a+ (Novagen, Madison, WI) vector, encoding for DCAMKL1 with a 6XHIS tag on both the N and C termini. This construct was also mutagenized to remove the 6XHIS tag from the amino terminus, so that the first encoded amino acid is the methionine of DCAMKL1. Recombinant protein was produced in BL21 DE3 Escherichia coli (Stratagene, La Jolla, CA), according to the manufacturer's recommendations. Briefly, a single clone was grown until OD 600 = 0.4–0.6, and induced with 1 mmisopropyl-β-d-thiogalactoside. After 2 hr, the pellet was resuspended in 1× binding buffer, sonicated, cleared by centrifugation, and the supernatant was run on an equilibrated Ni2+ column (Novagen, Madison, WI). The column was washed with 10 volumes of 1× binding buffer, six volumes of 1× wash buffer, then eluted with six volumes of 1× elution buffer, dialyzed overnight against 100 mm HEPES, pH 7.5, 200 mm NaCl, and 10 mm MgCl2 at 4°C, and concentrated to 1 mg/ml on a Centricon column (Amicon, Beverly, MA).
Microtubule polymerization assays. Assessment of aster formation was performed as previously described (Gleeson et al., 1999). Briefly, rhodamine-labeled tubulin (Cytoskeleton, Denver, CO) was mixed 1:4 with phosphocellulose-purified (PCP) tubulin (Cytoskeleton) to a final concentration of 0.5 mg/ml. DCAMKL1 (0.01–0.7 mg/ml) was added in BRB80 buffer (in mm: 80 PIPES, pH 6.8, 1 MgCl2, and 1 EGTA) in 1 mm GTP for 15 min at 37°C. Samples were fixed with 0.1% glutaraldehyde and 20% glycerol in BRB80. Aliquots were then visualized with fluorescent microscopy.
For the quantitative analysis of microtubule polymerization, PCP tubulin at 1 mg/ml was added to specific concentrations of DCAMKL1, in 1 mm GTP, in PEM buffer. Each experimental sample was mixed briefly and assayed for polymerization as measured by the change in diffraction (Gaskin et al., 1974). The kinetics of the diffraction for each sample was measured by right angle scattering in a quartz cuvette. The transmitted wavelength was 340 nm, and the detection wavelength was 345 nm, at a 90° angle to the transmitted beam, and the recording was performed for 16 min at 37°C in a Perkin-Elmer (Emeryville, CA) LS50B fluorimeter. The initial diffraction was set to zero at time 0 for each sample.
Construction of DCAMKL1-green fluorescent protein and kinase dead construct. To create a green fluorescent protein (GFP)-labeled version of DCAMKL1, a cDNA fragment containing EGFP was cut out of the pEGFP-1 vector (Clontech, Palo Alto, CA) with EcoRI and NotI. The fragment was blunt-ended and then cloned into the DCAMKL1-Myc construct at the HindIII site. The construct was checked for proper orientation of the insert through restriction digest. A kinase-inactive DCAMKL1-GFP construct was created by mutating Lys419 → Arg419 (K419R) (Hanson and Schulman, 1992) using site-directed mutagenesis of the DCAMKL1-Myc construct using the QuikChange site-directed mutagenesis kit (Stratagene). Subsequently, the altered construct was checked for errors by sequencing through the open reading frame. Insertion of the EGFP cDNA in theHindIII site was then performed as described above.
Transfection of COS7 and NIH 3T3 cells. Both NIH 3T3 and COS7 cells were grown in 10 cm tissue culture plates containing DMEM and 10% fetal calf serum (FCS) to ∼60% confluency. Cells were then transfected with various plasmid constructs with Superfectamine (Qiagen). After transfection, cells were cultured in normal growth media and maintained in a humidified incubator (5% CO2) at 37°C for 2 d. COS cells were transfected with DCAMKL1-Myc for kinase assays and GFP analysis, whereas 3T3 cells were transfected for GFP analysis and time-lapse alone. Latrunculin B (0.10 μg/ml) was used to induce actin depolymerization and process outgrowth.
Time-lapse video imaging. Imaging was performed essentially as described (M. Lu, L. Orecchio, and K. Kosik, unpublished observations). Cells transfected with DCAMKL1 constructs were imaged using a Nikon Diaphot 300 microscope with 40× [1.0 numerical aperture (NA)] and 100× (1.4 NA) oil immersion lenses, with the GFP filter set (Chroma Tech). Fluorescent images were captured with a cooled CCD camera (Princeton Instruments, Princeton, NJ), and processed using Metamorph software and Adobe Photoshop.
Immunoprecipitation. Control, DCAMKL1-Myc, and DCAMKL1–419R-transfected COS7 cells were immunoprecipitated as follows. Cells were washed once with ice-cold PBS and lysed (10 min on ice) in buffer H (in mm: 50 B-glycerophosphate, pH 7.3, 1.5 EGTA, 1 EDTA, 1 DTT, 0.1 sodium vanadate, 1 benzamidine, 10 μg/ml aprotinin, 10 μg/ml leupeptin, and 2 μg/ml pepstatin-A) (Seger et al., 1994). Cells were then scraped from the plate, pelleted, and homogenized using three strokes of a homogenizer. Cells were then centrifuged (14,000 × g; 20 min, 4°C). The supernatant was collected and incubated with 2 μg of anti-DCAMKL1 antibody (4°C) for 1 hr, then precipitated with protein A-agarose (Sigma) for 1 hr. Complexes were washed (4°C) three times with PBS + 0.1% NP-40, once with 0.5 m LiCl in PBS, and twice with buffer A (in mm: 50B-glycerophosphate, pH 7.3, 1.5 EGTA, 1 EDTA, 1 DTT, and 0.1 Na3VO4) (Seger et al., 1994). Agarose beads were split into fractions and directly added to the kinase assay mixture or run on a Western blot.
Autophosphorylation and in vitro kinase assays. Kinase assays were performed as previously described (Silverman et al., 1999). DCAMKL1-Myc phosphorylation was performed at 30°C in a buffer containing 50 mm HEPES, pH 8.5, 10 mm magnesium acetate, 10 μg/ml leupeptin, 10 μg/ml aprotinin, 50 mmB-glycerophosphate, 100 μm orthovanadate, 50 μm γ-32P[ATP] (20 cpm/fmol) in a final volume of 30 μl. Reactions were initiated by adding immunoprecipitation complexes to kinase buffer, with immediate incubation at 30°C. All reactions were run for 5 min. Reactions were terminated by adding 2× SDS-PAGE sample buffer and boiled for 5 min. All in vitro kinase assays were performed using the same reaction mixture with varying substrates. MBP was purchased from Sigma, and 0.066 μg was used per reaction.
RESULTS
Characterization of polyclonal antisera specific to DCAMKL1
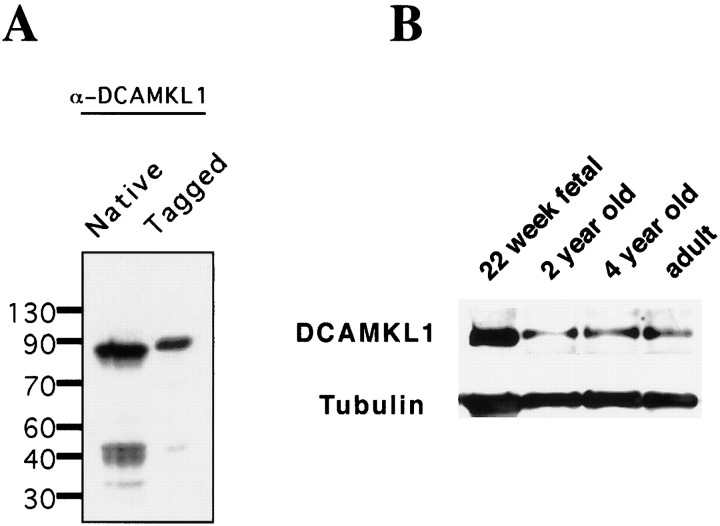
To determine the spatial and temporal expression of DCAMKL1, a polyclonal antibody was generated to a peptide immunogen corresponding to the N terminus of DCAMKL1. On a Western blot, the antibody recognized a major band at 80 kDa in whole-cell lysate from brain and a major band of slightly larger size in COS7 cells transfected with a Myc-tagged construct containing full-length DCAMKL1 (Fig.1A). A lower band at 35 kDa, which has been observed with other N-terminal DCAMKL1 antisera as well (Mizuguchi et al., 1999), likely represents an N terminal breakdown product of DCAMKL1. The α-DCAMKL1 sera did not recognize recombinant DCX, and the peptide immunogen contained only 2 of 15 amino acids in common DCX, indicating that the antisera could be used to study the expression of DCAMKL1 independently.
Fig. 1.
Developmental Western and antibody specificity of DCAMKL1 antibody. A, Immunoblot analysis demonstrating specificity of α-DCAMKL1 antisera. Whole-cell extract from cultured neurons or COS7 cells transfected with an epitope-tagged expression vector encoding full-length DCAMKL1 was probed with α-DCAMKL1. α-DCAMKL1 produces a specific band at 80 kDa both in cultured neurons and as a Myc-tagged fusion protein in overexpressing COS7 cells (90 kDa). B, Developmental regulation of DCAMKL1 in various aged human occipital cortex. Blots were probed with α-DCAMKL1 and subsequently probed with α-tubulin to control for protein loading. DCAMKL1 is expressed highly during embryonic life, then rapidly downregulated, with continued expression in the adult.
To define the developmental time course of DCAMKL1 protein expression, occipital cortex from human postmortem specimens at several ages was subjected to Western blot analysis with α-DCAMKL1 (Fig.1B). DCAMKL1 has a regulated expression during development, with highest levels of expression in 22 week fetal brain and lower levels throughout adulthood. The results of the Western blot analysis confirm previous Northern analysis (Omori et al., 1998;Burgess et al., 1999) showing that various isoforms of DCAMKL1 continue to be expressed throughout adulthood as well.
Spatial and temporal expression of DCAMKL1
To analyze the temporal and spatial pattern of DCAMKL1 expression further, a series of immunofluorescent stainings of mouse embryonic sections was performed (Fig.2A–F). DCAMKL1 immunoreactivity at E14 was widespread throughout the nervous system. DCAMKL1 immunoreactivity was high in the developing retina (Fig.2A), spinal cord and dorsal root ganglion (Fig.2B), cerebellum (Fig. 2C), and cerebral cortex (Fig. 2D). Throughout all areas of the peripheral nervous system studied, such as the trigeminal ganglion, dorsal root ganglia (Fig. 2B), enteric plexus, and sympathetic ganglion, DCAMKL1 was expressed as well (data not shown). Furthermore, at P8, during migration of granule cells in the cerebellum (Fig. 2C), DCAMKL1 was expressed highly in areas that were rich in migrating granule cells, although the high levels of expression made it difficult to ascertain whether the staining was specific to migrating granule cells. There were lower levels of immunoreactivity in Purkinje cells and in the external granule layer. Comparison to the expression of DCX showed that DCAMKL1 and DCX are coexpressed in regions containing postmitotic, MAP2-positive migrating neurons (Fig.2D–F), but low levels of DCAMKL1 immunoreactivity are also present in regions containing dividing cells such as the ventricular zone of the cortex (Fig.3A–C). Thus, DCAMKL1 expression may not be completely neuron-specific, although studies of mRNA (Omori et al., 1998; Burgess et al., 1999) and protein localization using some other antisera and immunoperoxidase staining (Mizuguchi et al., 1999) showed lower levels of DCAMKL1 immunoreactivity in the cortical ventricular zone than observed with our antiserum. Within the developing cortex, DCAMKL1 expression was present at all ages examined. There was intense staining in the intermediate zone and cortical plate compared with the less intense ventricular zone staining at earlier stages (Fig. 3A,B) and more diffuse staining at later postnatal ages (Fig. 3C).
Fig. 2.
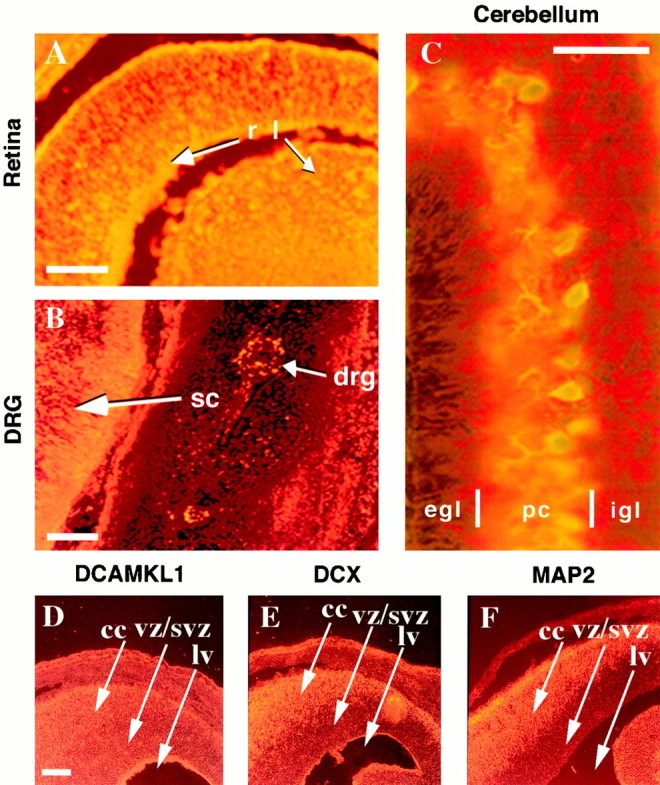
DCAMKL1 is expressed by populations of migrating neurons in the CNS and PNS. All immunostained tissues are from mouse at E14 in the sagittal plane except for the cerebellum (P8). DCAMKL1 is expressed by retinal cells predominantly in the ganglion cell layer (A), by spinal cord neurons as well as peripheral neurons in the dorsal root ganglia (B), and by cells in the internal granule layer (igl) and deeper parts of the external granule layer (egl) of the cerebellum (C). In C, sections were colabeled with α-DCAMKL1 and Calbindin. Scale bars:A, 100 μm; B, 200 μm;C, 70 μm. D–F, DCAMKL1 is expressed in similar populations of neurons as DCX, but also more widely in ventricular zones. Whereas DCX is excluded from the ventricular zone (E), DCAMKL1 immunoreactivity is present in the cortical plate (cc) as well as in the vz/svz (D). MAP2 is expressed specifically in postmitotic neurons and shows a distribution similar to DCX (F). Scale bar: D, 200 μm.cc, Cortical plate; vz, ventricular zone;lv, lateral ventricle; egl, external granule layer; pc, Purkinje cell layer;igl, internal granule layer; r, retina;l, lens; sc, spinal cord;drg, dorsal root ganglion.
Fig. 3.
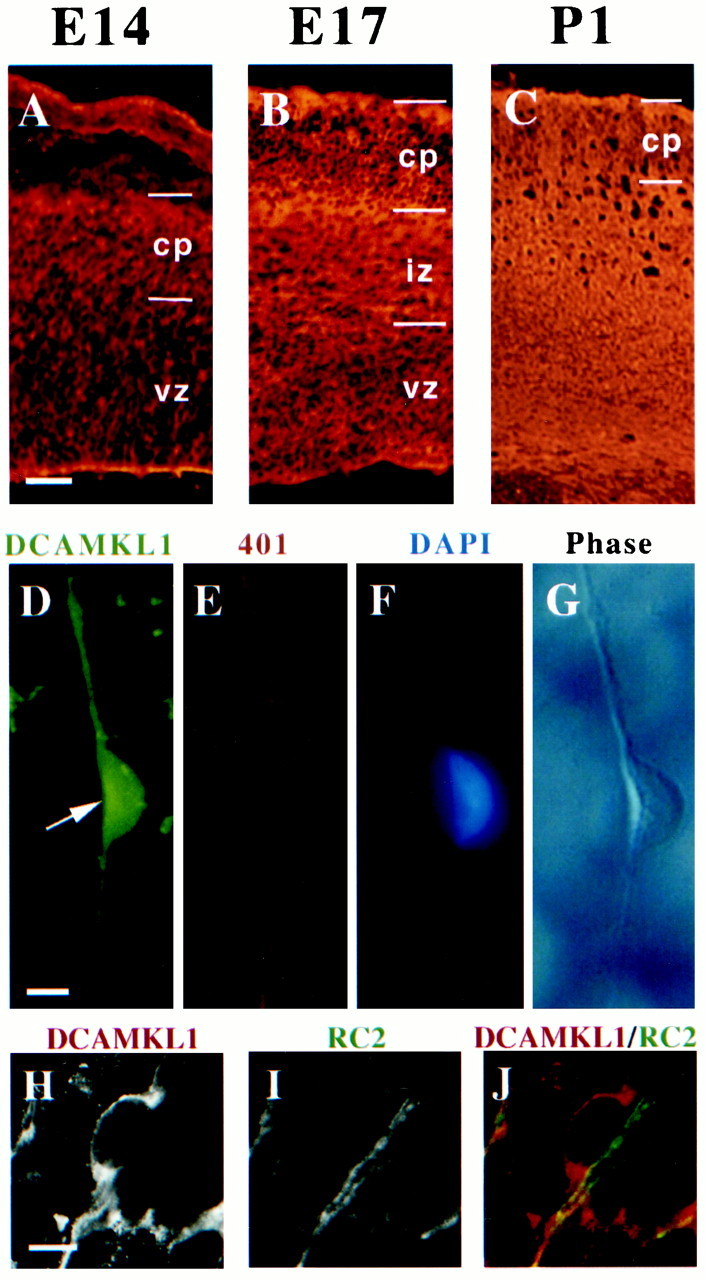
DCAMKL1 is present in migrating neurons in the cortex. A–C, Expression of DCAMKL1 in the cortex at different embryonic stages (E14, E17, and P1). DCAMKL1 is expressed widely throughout the entire cortical thickness, with less staining in the vz and intense staining in the iz, cp, and mz. Scale bars:A–C, 90 μm. vz, Ventricular zone; iz, intermediate zone; cp, cortical plate; mz, marginal zone. D–G, DCAMKL1 is expressed in bipolar cells (probably neurons) that are apposed to radial glia isolated by the cortical imprint method (Anton et al., 1999). Cortical imprint of an individual radial unit demonstrates a cell expressing DCAMKL1 (D) closely apposed to a radial glial fiber stained with Rat-401 antibody (E); F shows a DAPI nuclear counterstain, whereas G shows a matching phase-contrast image. Scale bar, 10 μm. H–J, DCAMKL1 is expressed in bipolar cells apposed to radial glia, highly suggestive of radially migrating neurons, seen in E14 cortical sections costained with anti-DCAMKL1 and RC2 antibody (Misson et al., 1988) to label radial glial cells. A radially oriented cell (H, andred in J) is seen apposed to a radial glial fiber stained with RC2 (I, andgreen in J). Scale bar, 10 μm.
DCAMKL1 is present in radially migrating neurons in the cerebral cortex
Because DCAMKL1 is expressed highly by embryonic neurons that seem to be migrating radially in the intermediate zone and cortical plate, we performed confocal microscopy and stained individual radial migratory units to confirm that DCAMKL1 was indeed present in at least some radially migrating neurons (Fig. 3D–G). By using the cortical imprint method, individual radial migration units were isolated (Anton et al., 1999) and immunostained with a variety of cell type-specific antibodies. DCAMKL1 expression was present in the migrating neuron, whereas the glial fibers were immunoreactive for the monoclonal rat-401 antibody (Hockfield and McKay, 1985). Moreover, double-label confocal microscopy using the RC2 monoclonal antibody to stain radial glial fibers (Misson et al., 1988) confirmed that DCAMKL1 expression is present in bipolar profiles that strongly resemble migrating neurons (Fig. 3H–J). These data and previous data (Mizuguchi et al., 1999) suggest that DCAMKL1 is coexpressed in migrating neurons along with DCX (Gleeson et al., 1999), although DCAMKL1 may have some lower level of expression more broadly.
DCAMKL1 coassembles with microtubules from brain and leads to increased polymerization of microtubules
To define the subcellular localization of DCAMKL1, primary cortical neural cultures were immunostained with α-DCAMKL1. DCAMKL1 immunoreactivity was located predominantly along the length of cells with the morphological appearance of neurons (Fig.4A). DCAMKL1 immunoreactivity was fibrillar under high power, suggesting an association with cytoskeletal elements (Fig. 4A; data not shown). Double-labeling of cells with antibodies directed against cytoskeletal proteins and α-DCAMKL1 revealed some overlapping localization with anti-α-tubulin antibodies (Fig.4A–C). The colocalization was not absolute, but suggested the possibility that DCAMKL1, like DCX, may have areas of overlap with tubulin. In contrast, no consistent colocalization with actin or neurofilaments was observed (data not shown).
Fig. 4.
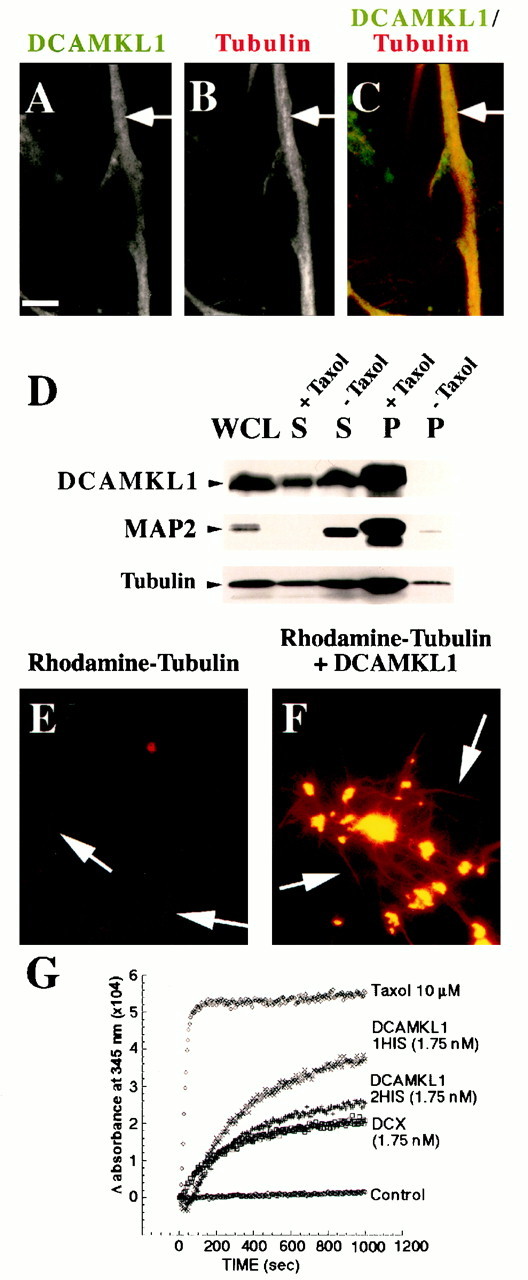
DCAMKL1 associates with microtubules and leads to microtubule polymerization. A–C, Subcellular localization of DCAMKL1 shows some localization that overlaps with microtubules. DCAMKL1 immunostaining of untreated E17 rat primary cortical neural cells (likely a neuron) demonstrates overlapping patterns of expression of DCAMKL1 with microtubules using α-tubulin antibody. DCAMKL1 and tubulin both were localized mainly to the processes of neurons. Scale bar, 10 μm. D, DCAMKL1 coprecipitates with taxol-stabilized microtubules from rat brain. Whole brain lysates from P5 newborn rat pups were cleared by centrifugation to isolate a tubulin-rich fraction and divided into two equal aliquots. To one aliquot, taxol and GTP were added and to the other aliquot, GTP alone was added, and microtubules were isolated by centrifugation. DCAMKL1 is enriched in the taxol-stabilized microtubule pellet (P, +Taxol), with smaller amounts remaining in the supernatant (S, +Taxol), whereas DCAMKL1 is not present in the pellet in the absence of taxol (P, −Taxol) and is retained in the supernatant (S, −Taxol). Reprobing a similar blot with α-pan-MAP2 indicates that DCAMKL1 distribution is similar to that of MAP2C. WCL,Whole-cell lysate; p, microtubule pellet;S, supernatant. E, F, Fluorescent images of rhodamine-tubulin plus phosphocellulose-purified tubulin in the absence of DCAMKL1 (E) and in the presence of 0.01 mg/ml recombinant DCAMKL1 (F) show that addition of DCAMKL1 leads to an increase in the number of microtubules (arrows) and the formation of aster-like stars.G, Turbidimetric analysis of microtubule polymerization induced by DCAMKL1. Recombinant DCAMKL1 with a C-terminal (DCAMKL1 1HIS) histidine tag, or with both C-terminal and N-terminal HIS tags (DCAMKL1 2 HIS) were incubated with PCP tubulin along with a comparable concentration of HIS-tagged DCX, or taxol as a positive control. Refracted light at 345 nm indicates light scattering because of microtubule polymerization, and shows that DCX and DCAMKL1 have comparable molar effects.
To examine whether the overlapping localization of DCAMKL1 with microtubules in cultured neurons reflected a functional interaction, microtubules from a cytosolic fraction of bovine brain were precipitated in the presence of taxol and analyzed for the presence of DCAMKL1. DCAMKL1 was substantially enriched in the taxol-stabilized pellet, whereas DCAMKL1 was not present in the pellet in the absence of taxol (Fig. 4D). A portion of DCAMKL1 (20%) remained in the supernatant in the presence of taxol as well, indicating that there may be distinct pools of DCAMKL1 not associated with microtubules. A MAP2C antibody was used as a positive control for the procedure, showing that MAP2C was exclusively present in the Taxol-stabilized microtubule pellet fraction as expected.
Because of the homology of DCX with DCAMKL1, we next tested whether there are direct effects of DCAMKL1 on microtubule polymerization, similar to that observed for DCX (Gleeson et al., 1999), by incubating purified DCAMKL1 with rhodamine-labeled tubulin. Coincubation led to a dramatic stimulation of microtubule polymerization and to the formation of aster-like stars of polymerized rhodamine-labeled tubulin (Fig.4E,F), suggesting that DCAMKL1, like DCX, can directly stimulate polymerization or stabilization of microtubules. To provide an estimate of the quantitative effects of DCAMKL1 in inducing microtubule polymerization, we used a turbidimetric assay (Gaskin et al., 1974) to compare the effects of DCAMKL1 and DCX. DCAMKL1 was HIS-tagged at both its N and C terminus (DCAMKL1 2HIS) or only at its C terminus end (DCAMKL1 1HIS). Equimolar quantities of either DCAMKL1 construct had very similar quantitative effects in this assay to DCX (Fig. 4G). We have previously shown (Gleeson et al., 1999) that the stimulation of microtubule polymerization by DCX occurs in the same order of magnitude of concentration as MAP2C, suggesting that DCAMKL1, like DCX, can stimulate microtubule polymerization at physiological concentrations.
Overexpression of DCAMKL1 leads to microtubule bundling and cold and drug stability
To determine whether DCAMKL1 can alter microtubule structure, Myc-tagged DCAMKL1 was over-expressed in primary cortical cultures and analyzed for effects on microtubules. Whereas cells overexpressing a negative control lacZ gene showed diffuse immunoreactivity and no effect on microtubules (Fig.5A–C), overexpression of the epitope-tagged construct led to striking changes in microtubule morphology, with the development of rod-like bundles of microtubules throughout the transfected cells (Fig. 5D–F). Bundled microtubule patterns were quite common and observed in 70–90% of overexpressing cells; bundled patterns also tended to be more apparent and more complex in cells with very bright DCAMKL1 immunofluorescence, suggesting higher levels of overexpression. Differences were most noticeable in non-neuronal cells, probably oligodendrocytes and astrocytes, which were larger and flatter than neurons and had a more visible cytoskeleton. Neurons, which normally express DCAMKL1 and DCX, showed no apparent change in the microtubule cytoskeleton (data not shown) after DCAMKL1 overexpression as compared with untransfected neurons, although the elongated morphology of neurons may make microtubule bundling more difficult to detect.
Fig. 5.

Overexpression of DCAMKL1 in cultured neural cells leads to a microtubule bundling phenotype that is resistant to depolymerization with either cold or colchicine treatment.A–C, Transfection of neural cells (in this case a likely glial cell based on morphology) with lacZ-encoding vector leads to no change in microtubule bundling and a diffuse distribution of lacZ within the cytoplasm. D–F, Transfection of cells with DCAMKL1-Myc leads to microtubule bundling (F, arrows). Scale bar, 10 μm. G–I,Overexpression of DCAMKL1 stabilizes microtubules to colchicine treatment (H, I, arrows), whereas surrounding untransfected cells (data not shown) or cells expressing low levels of DCAMKL1 have destabilized microtubules. J–L,Microtubule bundles induced by overexpression of DCAMKL1 are partially resistant to cold (0°C) treatment (K, L, arrows), whereas microtubules in neighboring cells are disrupted (data not shown). Scale bars: A, D, G, J, 10 μm.
Overexpression of other MAPs has been shown to lead to a relative stabilization of microtubules to drug and cold depolymerization (Takemura et al., 1992). We subjected DCAMKL1-transfected primary cortical cultures to cold treatment and colchicine treatment to examine whether DCAMKL1 overexpression also would stabilize microtubules to these treatments. After 2 hr of colchicine treatment or 30 min of cold treatment at 0°C, cells were then fixed with glutaraldehyde and processed to visualize DCAMKL1-Myc as well as microtubules. Treatment of transfected cells with colchicine led to a disruption of most microtubules and loss of some DCAMKL1-microtubule colocalization in cells expressing low levels of DCAMKL1 (Fig. 5G). However, transfected cells expressing higher levels of DCAMKL1 still maintained the presence of some stabilized microtubules (Fig.5G–I). A significant effect of DCAMKL1 on cold stability of microtubule bundles was also seen in transfected cells expressing high levels of DCAMKL1 (Fig. 5J–L) as compared with nontransfected cells in the same culture (data not shown). Cells transfected with DCAMKL1 maintained some bundles of microtubules after cold treatment when compared with neighboring nontransfected cells, which contained deteriorated microtubule structures.
DCAMKL1 is a functional kinase
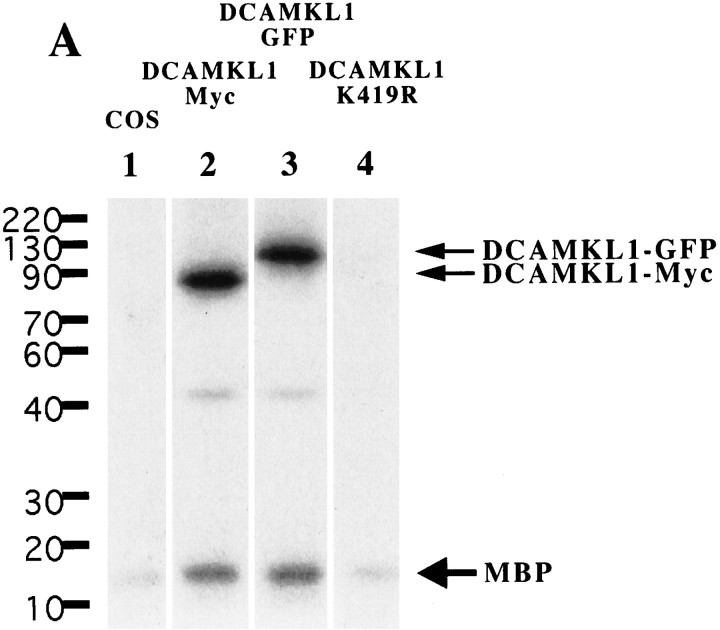
Based on sequence homology, the primary structure of the C terminus of DCAMKL1 encodes for a protein serine–threonine kinase similar to CaM kinase II. Previous reports have shown that CPG16, encoded by an alternative transcript that includes only the C-terminal domain of DCAMKL1, is a functional cAMP-dependent kinase that does not respond to calcium or calmodulin stimulation (Hevroni et al., 1998;Silverman et al., 1999). To determine whether full-length DCAMKL1 is a functional kinase, an in vitro kinase assay was performed. A Myc- or GFP-tagged construct containing the cDNA of DCAMKL1 was overexpressed in COS7 cells, and the DCAMKL1 fusion proteins were immunoprecipitated with α-DCAMKL1 antiserum. Precipitated DCAMKL1 was then used in an in vitro kinase assay with MBP protein. The results of the assay showed that both MBP and DCAMKL1 were phosphorylated by DCAMKL1 (Fig. 6,lane 2). Untransfected COS7 cell extracts subjected to immunoprecipitation showed some low background level of MBP phosphorylation (Fig. 6, lane 1).
Fig. 6.
In vitro kinase activity of DCAMKL1 fusion proteins is blocked by a K419R point mutation. The various constructs of DCAMKL1 used in this study are labeled above the blot. COS7 cell extracts were transfected with either no construct (COS, 1), DCAMKL1-Myc (2), DCAMKL1-GFP (3), or the kinase-inactive K419R DCAMKL1-GFP (4) constructs. Transfection and immunoprecipitation with anti-Myc or anti-DCAMKL1 antibody was performed as described in Materials and Methods. Samples from the following assay mixtures were loaded: lane 1, nontransfected cells with MBP substrate; lane 2, DCAMKL1-Myc transfected cells with MBP; lane 3, DCAMKL1-GFP transfected cells with MBP; lane 4, K419R transfected cells with MBP. Ten micrograms of MBP were used for each reaction. Some phosphorylation of MBP is seen in untransfected COS cells (1) that is comparable with the level of phosphorylation seen in the kinase-dead DCAMKL1 construct (4). Expression of DCAMKL1 as a Myc or GFP fusion shows phosphorylation of DCAMKL1 and MBP, as well as an unidentified ≈40 kDa band that is probably a degradation product of DCAMKL1. This degradation product was not seen in other experiments with a shorter interval between reaction and loading and was more prominent with a longer postreaction interval (data not shown). All reactions were performed simultaneously and separated and transferred on the same gel and developed on the same blot and same piece of film at the same time; however, some lanes of the final gel are not illustrated for simplicity.
An alternate trial involving the DCAMKL1-GFP construct showed a similar kinase activity to the Myc-tagged DCAMKL1 (Fig. 6, lane 3). A smaller band was present in some assays (Fig. 6, lanes 2,3). This smaller product likely represents a phosphorylated cleavage product of DCAMKL1, because it was less apparent when the kinase assay was performed immediately after the immunoprecipitation and was more prominent when the kinase assay was performed several hours after the immunoprecipitation (data not shown). Contrary to previous results showing that CPG16 kinase activity was very low in the absence of 8-bromo-cAMP or forskolin (Silverman et al., 1999), full-length DCAMKL1 required no activation for functional kinase activity. Moreover, in our hands, full-length DCAMKL1 showed equivalent amounts of phosphorylation in the absence and presence of 8-bromo-cAMP (data not shown).
We created a point mutation in the DCAMKL1-GFP construct by site-directed mutagenesis of a previously identified site critical for kinase activity (Hanson and Schulman, 1992; Hanson et al., 1994). This mutated (K419R) DCAMKL1-GFP construct showed no ability to autophosphorylate or to phosphorylate MBP (Fig. 6, lane 4) above baseline.
Kinase-independent effects of DCAMKL1-GFP on microtubule bundling
Fusion of microtubule-associated proteins to the GFP allows for study of the real-time dynamics of cytoskeletal arrangements in cells in culture (Kaech et al., 1996). A DCAMKL1-GFP fusion construct was created to analyze the effect of DCAMKL1 on microtubule dynamics in real time. Expression of the GFP construct alone in transfected COS7 cells showed no effect on microtubule bundling (data not shown). Overexpression of the DCAMKL1-GFP construct showed strong colocalization and striking bundling of microtubules in a variety of patterns (Fig. 7A). In the majority of cells, microtubules appeared in bright bundles at the periphery of cells or in whorl-like spirals within the cell. All transfected cells showed microtubule bundling to some degree. Whereas cells with lower levels of expression appeared to contain a noticeable microtubule-organizing center (MTOC) in the form of an aster-like center, higher levels of DCAMKL1 expression produced a lack of visible asters or MTOCs within the cells. These phenomena, as described with other MAPs (Kaech et al., 1996), suggest that DCAMKL1 may act to nucleate microtubule polymerization.
Fig. 7.
Overexpression of GFP-tagged DCAMKL1 in COS7 cells and NIH 3T3 cells induces microtubule bundling with maintained microtubule dynamics (A). Transfection of COS7 cells with DCAMKL1-GFP leads to a striking amount of microtubule bundling (arrows). B, Transfection of COS7 cells with kinase-dead DCAMKL1-GFP K419R leads to no apparent difference in degree of microtubule bundling from that of wild-type construct. C, Addition of 0.2 μmLatrunculin B to COS7 cells transfected with DCAMKL1-GFP leads to the development of multiple processes containing bundled microtubules (arrows). D–H, Video frames taken from a time-lapse recording of a single NIH 3T3 cell (which show similar microtubule bundling effects to COS7 cells) showing DCAMKL1-GFP labeled microtubules and maintenance of microtubule dynamics. During this time, a single microtubule (arrow) is seen to collapse and reextend. Scale bars: A–C, 10 μm.
The ability of DCAMKL1 to nucleate and bundle microtubules is similar to that of DCX (Gleeson et al., 1999), suggesting that it is the shared DCX-like domain that is responsible for these effects. Moreover, transfection of the DCAMKL1-GFP kinase-dead (K419R) construct showed no apparent difference versus the wild-type construct in the ability to bundle microtubules (Fig. 7B). Addition of latrunculin B, a potent actin depolymerization agent, led to the formation of processes containing bundled microtubules (Fig. 7C), confirming that the DCAMKL1 construct is localizing with microtubules (Edson et al., 1993; Matus, 1994; Kaech et al., 1996).
Video time-lapse imaging of the DCAMKL1-transfected cells showed that, despite the potent microtubule-polymerizing effects of DCAMKL1, microtubules continued to maintain dynamic cycles of polymerization and depolymerization. To analyze whether DCAMKL1-associated microtubules retained the capacity for dynamic change, the DCAMKL1-GFP construct was transfected into NIH 3T3 cells for video analysis. NIH 3T3 cells showed similar patterns of microtubule bundling after DCAMKL1 overexpression as did COS7 cells (data not shown). In this experiment, microtubules were visualized only by their association with DCAMKL1-GFP, because tubulin itself was unlabeled. Real-time analysis of microtubule dynamics revealed that microtubules associated with DCAMKL1 remained dynamic, as shown by the presence of active depolymerization and polymerization during observation (Fig. 7D–H). During the period of observation, single microtubules (arrows) were seen to grow and to shrink.
DISCUSSION
Here we show that full-length DCAMKL1 fulfills the criteria to be a member of a new family of microtubule-associated proteins, because it colocalizes with microtubules, coprecipitates with microtubules, and dramatically stimulates microtubule elongation when provided in purified form or transfected into cell lines or primary neural cells. The effects on microtubules do not require kinase activity, because they are retained in a kinase-dead protein. We also show that DCAMKL1 has kinase activity for itself, confirm that it has active kinase activity for MBP, and show that it can phosphorylate MBP in vitro without requiring stimulation. Finally, we show that DCAMKL1 is expressed in migrating neurons with DCX, suggesting that these two MAPs may function in concert to regulate microtubule dynamics in migrating neurons.
DCAMKL1 is expressed in the developing and adult brain as a number of alternative transcripts that include either its full-length, or else consist of only the amino terminal DCX-like region or the kinase-encoding C terminus alone (Omori et al., 1998; Burgess et al., 1999). The full-length transcript is most highly expressed during development (Omori et al., 1998), whereas the adult brain shows expression predominantly of a truncated form that lacks the entire region of similarity to DCX (Omori et al., 1998; Burgess et al., 1999). Intriguingly, the C-terminal splice form of DCAMKL1 had been previously cloned in rat as a candidate plasticity gene (CPG 16), defined as a transcript whose expression in the hippocampus is induced by the glutamate agonist kainic acid, suggesting a potential role in synaptic remodeling (Hevroni et al., 1998; Silverman et al., 1999). Two other recently cloned cDNAs induced by kainate (Vreugdenhil et al., 1999) or dopamine (Berke et al., 1998) stimulation also show regions of sequence identity to rat CPG16, and homology to human DCAMKL1, and may also be alternative splice forms of the same gene.
In their analysis of the kinase activity of CPG16, Silverman et al. (1999) found no stimulatory effect of calcium-calmodulin on kinase activity, and instead found stimulation of kinase activity by forskolin and 8-bromo-cAMP. They suggested that CPG16 may function as a cAMP-dependent kinase and not as a CAM kinase, despite its lack of structural homology to cAMP-dependent kinases and strong structural homology to CAM kinases. In contrast, we found readily evident levels of spontaneous kinase activity of DCAMKL1, and no stimulatory effect of 8-bromo-cAMP. Although we did not test effects of calcium-calmodulin on DCAMKL1, our results suggest that there are either technical differences in the assay, or that the full-length DCAMKL1 does not require 8-bromo-cAMP for its activity. Thus, lacking definitive evidence that full-length DCAMKL1 resembles a cAMP-dependent kinase, its structural homology to CAM kinases still warrants consideration that the full-length protein may show stimulation of its activity by Ca-calmodulin.
Our analysis is limited to full-length DCAMKL1. The full-length form is the predominant form expressed during the development of the cortex, whereas truncated isoforms such as CPG16 are persistently expressed in the adult (Omori et al., 1998). Our data are consistent with the DCX homology domain as being required for the MAP activity, and recent dissection of DCX using truncated constructs suggests that the microtubule binding domain is in the N-terminal 2/3 of DCX (Horesh et al., 1999). For example, missense mutations that disrupt neuronal migration (Gleeson et al., 1999), presumably because of loss of DCX function, all cluster in the N-terminal domain of DCX. These missense mutations occur in two evolutionarily conserved structural domains that have now been recognized in a number of molecules besides DCX and DCAMKL1, and these mutations disrupt the microtubule binding and polymerization activities of DCX (Sapir et al., 2000; Taylor et al., 2000). The effects of DCX missense mutations strongly suggest that interactions between DCX and tubulin are essential for normal neuronal migration. Therefore, the microtubule-binding domain of DCAMKL1 is likely to be encoded by the N terminus of DCAMKL1, which is highly homologous to the microtubule binding domain of DCX.
Microtubule function and neuronal migration
What is the role of microtubule reorganization in migrating neurons, and what role might DCAMKL1 play in this process? LIS1 shows a well conserved ortholog in Aspergillus nidulans called nudF (Xiang et al., 1995), whose role may represent a potential model for the role of LIS1, DCX, and potentially DCAMKL1 in neuronal migration. NudF is required for nuclear translocation along the fungal mycelium, and interacts genetically with genes that encode microtubule-related proteins such as α-tubulin and dynein (Willins et al., 1995, 1997;Morris et al., 1998). Nuclear translocation must also occur during the migration of neurons, and the nucleus of migrating neurons shows a unique microtubule structure surrounding it in the form of a “cage” (Rivas and Hatten, 1995). This perinuclear microtubule structure may be specialized for translocating the nucleus into the leading process of the migrating neuron.
Because DCAMKL1 is a bifunctional molecule, with both MAP and kinase activities, its in vivo role may prominently reflect one of the other of these functions, or may represent an interaction of the two. For example, DCAMKL1 (and DCX) may function primarily as MAPs whose activity is regulated by phosphorylation. Phosphorylation is often a means for negatively regulating the interactions of MAPs and microtubules (Raffaelli et al., 1992), raising the possibility that phosphorylation of DCAMKL1 by itself or other kinases may regulate its roles on microtubules. LIS1 preferentially associates with microtubules in its dephosphorylated form (Sapir et al., 1999), so that phosphorylation of DCAMKL1 may similarly block its effects on microtubule stabilization. Neuronal migration to the cortex is known to be saltatory, and is associated with calcium transients, so that calcium transients could potentially give rise to pulsatile cycles of microtubule stability and instability. The initial phase of neuronal migration consists of neurite elongation, which is associated with microtubule lengthening. The second phase of migration involves neurite shortening. This process of neurite shortening is complex and poorly understood, but presumably requires rapid microtubule reorganization and perhaps shortening or disassembly. This second phase is when nuclear translocation takes place. Interestingly, the second phase of migration is associated with increases in intracellular calcium (Komuro and Rakic, 1992, 1996, 1998), suggesting that intracellular calcium may regulate specific stages of microtubule reorganization. Because CAM kinases are activated downstream of intracellular calcium, phosphorylation of MAPs such as DCX and DCAMKL1 by DCAMKL1 or other kinases could represent a rapid mechanism for linking calcium transients to microtubule reorganization.
An alternative model suggests that DCAMKL1 plays a primary signaling function through its kinase domain and that the kinase activity of DCAMKL1 may be modulated by interactions with microtubules, DCX, or both. Thus, DCAMKL1 shows baseline autophosphorylation activity and strong kinase activity on MBP. This kinase activity may be negatively regulated by truncation of the N-terminal DCX domain of DCAMKL1, because CPG16, a DCAMKL1 isoform that lacks the DCX domain, shows low levels of unstimulated kinase activity (Silverman et al., 1999). Further analysis of the primary amino acid structure of DCAMKL1 and DCX revealed several highly conserved CAM kinase II consensus phosphorylation sequences, when searched with the PhosphoBase program (http://www.cbs.dtu.dk/htbin/pbase predict.pl). T-42 of DCX (RTRTLQA) and T-46 of DCAMKL1 (RTRTLQT), S-79 of DCX (RFRSFDA) and S-83 of DCAMKL1 (RFRSFEA), S-92 of DCX (RSLSDNI) and S-96 of DCAMKL1 (RTLSDNV), and S-306 of DCX (RSKSPAD) and S-307 of DCAMKL1 (RSKSPAS) are all highly conserved between the two proteins and closely match the consensus substrate sites for CAM kinase II. Although we have not determined the site of DCAMKL1 autophosphorylation, or determined whether DCX can be phosphorylated by DCAMKL1, several of these sites are intriguing. For example, two of the phosphorylation consensus sequences in DCX (S-79, S-92) correspond to regions of the protein that have been shown to be essential for microtubule binding (Horesh et al., 1999). Therefore, phosphorylation of these sites could affect the interaction of DCX with tubulin.
If the primary function of DCAMKL1 is through its kinase domain, interaction of the N terminal domain of DCAMKL1 with DCX or microtubules may regulate its kinase activity. Because kinase proteins frequently dimerize, DCAMKL1 may normally dimerize as well. If so, DCX and DCAMKL1 might form nonfunctional heterodimers in a way analogous to the competitive regulation of POU homeodomain proteins by I-POU proteins (Treacy et al., 1991; Verrijzer and Van der Vliet, 1993) or regulation of helix-loop-helix proteins by truncated Id proteins (Benezra et al., 1990; Sun et al., 1991). Alternatively, there may be competition between DCX and DCAMKL1 for binding to additional proteins (such as microtubules) that activates the kinase function of DCAMKL1. In any event, the strong homology of structure of the two proteins and their correlated activity and expression strongly suggest that theirin vivo functions are linked as well.
Footnotes
This work was supported by the Howard Hughes Medical Institute Medical Student Research Training Program (P.T.L.), Neurological Sciences Academic Development Award Fellowship 5K12NS01701 (J.G.G.), National Institute of Neurological Diseases and Stroke Grants RO1 NS38097 and PO1 39404, and the National Alliance for Autism Research and the National Association for Research in Schizophrenia and Depression (C.A.W.). We thank R. Ratan for providing primary cortical neurons, E. Anton for preparing and staining radial migratory units, M. Lu and K. Kosik for use of their time-lapse imaging equipment, Y. Feng, E. Olson, A. Chenn, and R. Segal for helpful comments, and G. Cummings for help preparing this manuscript.
Correspondence should be addressed to Christopher A. Walsh, Division of Neurogenetics, Department of Neurology, Beth Israel Deaconess Medical Center, HIM 846, 77 Avenue Louis Pasteur, Boston, MA 02115. E-mail: cwalsh@caregroup.harvard.edu.
Dr. Gleeson's present address: Division of Pediatric Neurology, University of California San Diego, San Diego, CA.
REFERENCES
- 1.Anton ES, Kreidberg JA, Rakic P. Distinct functions of alpha3 and alpha(v) integrin receptors in neuronal migration and laminar organization of the cerebral cortex. Neuron. 1999;22:277–289. doi: 10.1016/s0896-6273(00)81089-2. [DOI] [PubMed] [Google Scholar]
- 2.Avila J, Dominguez J, Diaz-Nido J. Regulation of microtubule dynamics by microtubule-associated protein expression and phosphorylation during neuronal development. Int J Dev Biol. 1994;38:13–25. [PubMed] [Google Scholar]
- 3.Baas PW. Microtubules and neuronal polarity: lessons from mitosis. Neuron. 1999;22:23–31. doi: 10.1016/s0896-6273(00)80675-3. [DOI] [PubMed] [Google Scholar]
- 4.Benezra R, Davis RL, Lockshon D, Turner DL, Weintraub H. The protein Id: a negative regulator of helix-loop-helix DNA binding proteins. Cell. 1990;61:49–59. doi: 10.1016/0092-8674(90)90214-y. [DOI] [PubMed] [Google Scholar]
- 5.Berg MJ, Schifitto G, Powers JM, Martinez-Capolino C, Fong CT, Myers GJ, Epstein LG, Walsh CA. X-linked female band heterotopia-male lissencephaly syndrome. Neurology. 1998;50:1143–1146. doi: 10.1212/wnl.50.4.1143. [DOI] [PubMed] [Google Scholar]
- 6.Berke JD, Paletzki RF, Aronson GJ, Hyman SE, Gerfen CR. A complex program of striatal gene expression induced by dopaminergic stimulation. J Neurosci. 1998;18:5301–5310. doi: 10.1523/JNEUROSCI.18-14-05301.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Brandt R. The tau proteins in neuronal growth and development. Front Biosci. 1996;1:118–130. doi: 10.2741/a120. [DOI] [PubMed] [Google Scholar]
- 8.Burgess HA, Martinez S, Reiner O. KIAA0369, doublecortin-like kinase, is expressed during brain development [In Process Citation]. J Neurosci Res. 1999;58:567–575. [PubMed] [Google Scholar]
- 9.Caceres A, Kosik KS. Inhibition of neurite polarity by tau antisense oligonucleotides in primary cerebellar neurons. Nature. 1990;343:461–463. doi: 10.1038/343461a0. [DOI] [PubMed] [Google Scholar]
- 10.Caceres A, Mautino J, Kosik KS. Suppression of MAP2 in cultured cerebellar macroneurons inhibits minor neurite formation. Neuron. 1992;9:607–618. doi: 10.1016/0896-6273(92)90025-9. [DOI] [PubMed] [Google Scholar]
- 11.des Portes V, Pinard JM, Billuart P, Vinet MC, Koulakoff A, Carrie A, Gelot A, Dupuis E, Motte J, Berwald-Netter Y, Catala M, Kahn A, Beldjord C, Chelly J. A novel CNS gene required for neuronal migration and involved in X-linked subcortical laminar heterotopia and lissencephaly syndrome. Cell. 1998;92:51–61. doi: 10.1016/s0092-8674(00)80898-3. [DOI] [PubMed] [Google Scholar]
- 12.Desai A, Mitchison TJ. Microtubule polymerization dynamics. Annu Rev Cell Dev Biol. 1997;13:83–117. doi: 10.1146/annurev.cellbio.13.1.83. [DOI] [PubMed] [Google Scholar]
- 13.Dobyns WB, Reiner O, Carrozzo R, Ledbetter DH. Lissencephaly. A human brain malformation associated with deletion of the LIS1 gene located at chromosome 17p13. JAMA. 1993;270:2838–2842. doi: 10.1001/jama.270.23.2838. [DOI] [PubMed] [Google Scholar]
- 14.Edson K, Weisshaar B, Matus A. Actin depolymerisation induces process formation on MAP2-transfected non-neuronal cells. Development. 1993;117:689–700. doi: 10.1242/dev.117.2.689. [DOI] [PubMed] [Google Scholar]
- 15.Felgner H, Frank R, Biernat J, Mandelkow EM, Mandelkow E, Ludin B, Matus A, Schliwa M. Domains of neuronal microtubule-associated proteins and flexural rigidity of microtubules. J Cell Biol. 1997;138:1067–1075. doi: 10.1083/jcb.138.5.1067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Francis F, Koulakoff A, Boucher D, Chafey P, Schaar B, Vinet MC, Friocourt G, McDonnell N, Reiner O, Kahn A, McConnell SK, Berwald-Netter Y, Denoulet P, Chelly J. Doublecortin is a developmentally regulated, microtubule-associated protein expressed in migrating and differentiating neurons. Neuron. 1999;23:247–256. doi: 10.1016/s0896-6273(00)80777-1. [DOI] [PubMed] [Google Scholar]
- 17.Gaskin F, Cantor CR, Shelanski ML. Turbidimetric studies of the in vitro assembly and disassembly of porcine neurotubules. J Mol Biol. 1974;89:737–755. doi: 10.1016/0022-2836(74)90048-5. [DOI] [PubMed] [Google Scholar]
- 18.Gleeson JG, Allen KM, Fox JW, Lamperti ED, Berkovic S, Scheffer I, Cooper EC, Dobyns WB, Minnerath SR, Ross ME, Walsh CA. Doublecortin, a brain-specific gene mutated in human X-linked lissencephaly and double cortex syndrome, encodes a putative signaling protein. Cell. 1998;92:63–72. doi: 10.1016/s0092-8674(00)80899-5. [DOI] [PubMed] [Google Scholar]
- 19.Gleeson JG, Lin PT, Flanagan LA, Walsh CA. Doublecortin is a microtubule-associated protein and is expressed widely by migrating neurons. Neuron. 1999;23:257–271. doi: 10.1016/s0896-6273(00)80778-3. [DOI] [PubMed] [Google Scholar]
- 20.Gleeson JG, Minnerath SR, Fox JW, Allen KM, Luo RF, Hong SE, Berg MJ, Kuzniecky R, Reitnauer PJ, Borgatti R, Mira AP, Guerrini R, Holmes GL, Rooney CM, Berkovic S, Scheffer I, Cooper EC, Ricci S, Cusmai R, Crawford TO, Leroy R, Andermann E, Wheless JW, Dobyns WB, Ross ME, Walsh CA. Characterization of mutations in the gene doublecortin in patients with double cortex syndrome. Ann Neurol. 1999;45:146–153. doi: 10.1002/1531-8249(199902)45:2<146::aid-ana3>3.0.co;2-n. [DOI] [PubMed] [Google Scholar]
- 21.Goode BL, Feinstein SC. Identification of a novel microtubule binding and assembly domain in the developmentally regulated inter-repeat region of tau. J Cell Biol. 1994;124:769–782. doi: 10.1083/jcb.124.5.769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hanson PI, Schulman H. Inhibitory autophosphorylation of multifunctional Ca2+/calmodulin-dependent protein kinase analyzed by site-directed mutagenesis. J Biol Chem. 1992;267:17216–17224. [PubMed] [Google Scholar]
- 23.Hanson PI, Meyer T, Stryer L, Schulman H. Dual role of calmodulin in autophosphorylation of multifunctional CaM kinase may underlie decoding of calcium signals. Neuron. 1994;12:943–956. doi: 10.1016/0896-6273(94)90306-9. [DOI] [PubMed] [Google Scholar]
- 24.Hevroni D, Rattner A, Bundman M, Lederfein D, Gabarah A, Mangelus M, Silverman MA, Kedar H, Naor C, Kornuc M, Hanoch T, Seger R, Theill LE, Nedivi E, Richter-Levin G, Citri Y. Hippocampal plasticity involves extensive gene induction and multiple cellular mechanisms. J Mol Neurosci. 1998;10:75–98. doi: 10.1007/BF02737120. [DOI] [PubMed] [Google Scholar]
- 25.Hockfield S, McKay RD. Identification of major cell classes in the developing mammalian nervous system. J Neurosci. 1985;5:3310–3328. doi: 10.1523/JNEUROSCI.05-12-03310.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Horesh D, Sapir T, Francis F, Wolf SG, Caspi M, Elbaum M, Chelly J, Reiner O. Doublecortin, a stabilizer of microtubules. Hum Mol Genet. 1999;8:1599–1610. doi: 10.1093/hmg/8.9.1599. [DOI] [PubMed] [Google Scholar]
- 27.Kaech S, Ludin B, Matus A. Cytoskeletal plasticity in cells expressing neuronal microtubule-associated proteins. Neuron. 1996;17:1189–1199. doi: 10.1016/s0896-6273(00)80249-4. [DOI] [PubMed] [Google Scholar]
- 28.Komuro H, Rakic P. Selective role of N-type calcium channels in neuronal migration. Science. 1992;257:806–809. doi: 10.1126/science.1323145. [DOI] [PubMed] [Google Scholar]
- 29.Komuro H, Rakic P. Intracellular Ca2+ fluctuations modulate the rate of neuronal migration. Neuron. 1996;17:275–285. doi: 10.1016/s0896-6273(00)80159-2. [DOI] [PubMed] [Google Scholar]
- 30.Komuro H, Rakic P. Orchestration of neuronal migration by activity of ion channels, neurotransmitter receptors, and intracellular Ca2+ fluctuations. J Neurobiol. 1998;37:110–130. [PubMed] [Google Scholar]
- 31.Kosik KS. The molecular and cellular biology of tau. Brain Pathol. 1993;3:39–43. doi: 10.1111/j.1750-3639.1993.tb00724.x. [DOI] [PubMed] [Google Scholar]
- 32.Kosik KS, Caceres A. Tau protein and the establishment of an axonal morphology. J Cell Sci [Suppl] 1991;15:69–74. doi: 10.1242/jcs.1991.supplement_15.10. [DOI] [PubMed] [Google Scholar]
- 33.Matus A. Stiff microtubules and neuronal morphology. Trends Neurosci. 1994;17:19–22. doi: 10.1016/0166-2236(94)90030-2. [DOI] [PubMed] [Google Scholar]
- 34.Misson J-P, Edwards MA, Yamamoto M, Caviness VS. Identification of radial glial cells within the developing murine central nervous system: Studies based upon a new immunohistochemical marker. Dev Brain Res. 1988;44:95–108. doi: 10.1016/0165-3806(88)90121-6. [DOI] [PubMed] [Google Scholar]
- 35.Mitchison T, Kirschner M. Dynamic instability of microtubule growth. Nature. 1984;312:237–242. doi: 10.1038/312237a0. [DOI] [PubMed] [Google Scholar]
- 36.Mizuguchi M, Qin J, Yamada M, Ikeda K, Takashima S. High expression of doublecortin and KIAA0369 protein in fetal brain suggests their specific role in neuronal migration. Am J Pathol. 1999;155:1713–1721. doi: 10.1016/S0002-9440(10)65486-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Morris NR, Efimov VP, Xiang X. Nuclear migration, nucleokinesis and lissencephaly. Trends Cell Biol. 1998;8:467–470. doi: 10.1016/s0962-8924(98)01389-0. [DOI] [PubMed] [Google Scholar]
- 38.Nagase T, Ishikawa K, Miyajima N, Tanaka A, Kotani H, Nomura N, Ohara O. Prediction of the coding sequences of unidentified human genes. IX. The complete sequences of 100 new cDNA clones from brain which can code for large proteins in vitro. DNA Res. 1998;5:31–39. doi: 10.1093/dnares/5.1.31. [DOI] [PubMed] [Google Scholar]
- 39.Omori Y, Suzuki M, Ozaki K, Harada Y, Nakamura Y, Takahashi E, Fujiwara T. Expression and chromosomal localization of KIAA0369, a putative kinase structurally related to Doublecortin. J Hum Genet. 1998;43:169–177. doi: 10.1007/s100380050063. [DOI] [PubMed] [Google Scholar]
- 40.Pilz DT, Matsumoto N, Minnerath S, Mills P, Gleeson JG, Allen KM, Walsh CA, Barkovich AJ, Dobyns WB, Ledbetter DH, Ross ME. LIS1 and XLIS (DCX) mutations cause most classical lissencephaly, but different patterns of malformation. Hum Mol Genet. 1998;7:2029–2037. doi: 10.1093/hmg/7.13.2029. [DOI] [PubMed] [Google Scholar]
- 41.Raffaelli N, Yamauchi PS, Purich DL. Microtubule-associated protein autophosphorylation alters in vitro microtubule dynamic instability. FEBS Lett. 1992;296:21–24. doi: 10.1016/0014-5793(92)80394-v. [DOI] [PubMed] [Google Scholar]
- 42.Reiner O, Carrozzo R, Shen Y, Wehnert M, Faustinella F, Dobyns WB, Caskey CT, Ledbetter DH. Isolation of a Miller-Dieker lissencephaly gene containing G protein beta-subunit-like repeats. Nature. 1993;364:717–721. doi: 10.1038/364717a0. [DOI] [PubMed] [Google Scholar]
- 43.Rivas RJ, Hatten ME. Motility and cytoskeletal organization of migrating cerebellar granule neurons. J Neurosci. 1995;15:981–989. doi: 10.1523/JNEUROSCI.15-02-00981.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sapir T, Elbaum M, Reiner O. Reduction of microtubule catastrophe events by LIS1, platelet-activating factor acetylhydrolase subunit. EMBO J. 1997;16:6977–6984. doi: 10.1093/emboj/16.23.6977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sapir T, Cahana A, Seger R, Nekhai S, Reiner O. LIS1 is a microtubule-associated phosphoprotein. Eur J Biochem. 1999;265:181–188. doi: 10.1046/j.1432-1327.1999.00711.x. [DOI] [PubMed] [Google Scholar]
- 46.Sapir T, Horesh D, Caspi M, Atlas R, Burgess HA, Wolf SG, Francis F, Chelly J, Elbaum M, Pietrokovski S, Reiner O. Doublecortin mutations cluster in evolutionarily conserved functional domains. Hum Mol Genet. 2000;9:703–712. doi: 10.1093/hmg/9.5.703. [DOI] [PubMed] [Google Scholar]
- 47.Seger R, Seger D, Reszka AA, Munar ES, Eldar-Finkelman H, Dobrowolska G, Jensen AM, Campbell JS, Fischer EH, Krebs EG. Overexpression of mitogen-activated protein kinase kinase (MAPKK) and its mutants in NIH 3T3 cells. Evidence that MAPKK involvement in cellular proliferation is regulated by phosphorylation of serine residues in its kinase subdomains VII and VIII. J Biol Chem. 1994;269:25699–25709. [PubMed] [Google Scholar]
- 48.Silverman MA, Benard O, Jaaro H, Rattner A, Citri Y, Seger R. CPG16, a novel protein serine/threonine kinase downstream of cAMP-dependent protein kinase. J Biol Chem. 1999;274:2631–2636. doi: 10.1074/jbc.274.5.2631. [DOI] [PubMed] [Google Scholar]
- 49.Sossey-Alaoui K, Hartung AJ, Guerrini R, Manchester DK, Posar A, Puche-Mira A, Andermann E, Dobyns WB, Srivastava AK. Human doublecortin (DCX) and the homologous gene in mouse encode a putative Ca2+-dependent signaling protein which is mutated in human X-linked neuronal migration defects. Hum Mol Genet. 1998;7:1327–1332. doi: 10.1093/hmg/7.8.1327. [DOI] [PubMed] [Google Scholar]
- 50.Sossey-Alaoui K, Srivastava AK. DCAMKL1, a brain-specific transmembrane protein on 13q12.3 that is similar to doublecortin (DCX). Genomics. 1999;56:121–126. doi: 10.1006/geno.1998.5718. [DOI] [PubMed] [Google Scholar]
- 51.Sun XH, Copeland NG, Jenkins NA, Baltimore D. Id proteins Id1 and Id2 selectively inhibit DNA binding by one class of helix-loop-helix proteins. Mol Cell Biol. 1991;11:5603–5611. doi: 10.1128/mcb.11.11.5603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Takemura R, Okabe S, Umeyama T, Kanai Y, Cowan NJ, Hirokawa N. Increased microtubule stability and alpha tubulin acetylation in cells transfected with microtubule-associated proteins MAP1B, MAP2 or tau. J Cell Sci. 1992;103:953–964. doi: 10.1242/jcs.103.4.953. [DOI] [PubMed] [Google Scholar]
- 53.Taylor KR, Holzer A, Walsh CA, Bazan JF, Gleeson JG (2000) Patient mutations in Doublecortin define a repeated microtubule-binding domain. J Biol Chem, in press. [DOI] [PubMed]
- 54.Treacy MN, He X, Rosenfeld MG. I-POU: a POU-domain protein that inhibits neuron-specific gene activation. Nature. 1991;350:577–584. doi: 10.1038/350577a0. [DOI] [PubMed] [Google Scholar]
- 55.Verrijzer CP, Van der Vliet PC . POU domain transcription factors. Biochim Biophys Acta. 1993;1173:1–21. doi: 10.1016/0167-4781(93)90237-8. [DOI] [PubMed] [Google Scholar]
- 56.Vreugdenhil E, Datson N, Engels B, de Jong J, van Koningsbruggen S, Schaaf M, de Kloet ER. Kainate-elicited seizures induce mRNA encoding a CaMK-related peptide: a putative modulator of kinase activity in rat hippocampus. J Neurobiol. 1999;39:41–50. [PubMed] [Google Scholar]
- 57.Willins DA, Xiang X, Morris NR. An alpha tubulin mutation suppresses nuclear migration mutations in Aspergillus nidulans. Genetics. 1995;141:1287–1298. doi: 10.1093/genetics/141.4.1287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Willins DA, Liu B, Xiang X, Morris NR. Mutations in the heavy chain of cytoplasmic dynein suppress the nudF nuclear migration mutation of Aspergillus nidulans. Mol Gen Genet. 1997;255:194–200. doi: 10.1007/s004380050489. [DOI] [PubMed] [Google Scholar]
- 59.Xiang X, Osmani AH, Osmani SA, Xin M, Morris NR. NudF, a nuclear migration gene in Aspergillus nidulans, is similar to the human LIS-1 gene required for neuronal migration. Mol Biol Cell. 1995;6:297–310. doi: 10.1091/mbc.6.3.297. [DOI] [PMC free article] [PubMed] [Google Scholar]