Supplemental Digital Content is available in the text
Keywords: gene expression, immune infiltration, nasopharyngeal carcinoma
Abstract
Immune infiltration of nasopharyngeal carcinoma (NPC) is closely associated with the patients’ prognosis. However, previous studies have not interpreted the difference of infiltrating immune cells in NPC.
We comprehensively analyzed the tumor-infiltrating immune cells present in NPC for the first time, which was based on a scientific deconvolution algorithm (CIBERSORT) and the gene expression data of GSE64634. The fractions of 22 immune cells were assessed to reveal the associations between normal samples and NPC samples.
Profiles of immune infiltration vary significantly between normal samples and NPC samples, and the variation could characterize the individual differences. NPC samples contained a higher proportion for M1 macrophages, whereas memory B cells and CD4 memory resting T cells were relatively lower.
Our data suggest that the differences in the infiltrating immune cells in NPC and these differences would probably facilitate patient consultation and individualized treatment.
1. Introduction
Nasopharyngeal carcinoma (NPC) is a common malignancy of head and neck in Southern China and Southeast Asia, which is highly invasive and has a relatively high metastasis rate.[1,2] With the development of radiotherapeutic techniques, chemotherapy regimens, and surgical techniques, the prognosis of NPC was improved significantly.[2] The tumor–node–metastasis (TNM) cancer staging system, made by the American Joint Committee on Cancer and the International Union for Cancer Control (AJCC/UICC), provided a useful standard for suggesting the treatment strategy or estimating patients’ prognosis. But it remained large variations in the patients’ prognosis who were also undergoing the similar treatment with the same stage. Therefore, the TNM system may be not enough to evaluate the entire NPC status or guide treatments. The malignant phenotypes of cancers are also defined by the immune cells activated in the tumor microenvironment (TME).[3] TME consists of immune cells, endothelial cells, mesenchymal cells, inflammatory mediators and extracellular matrix molecules.[4] In TME, immune cells are one major type of nontumor components and have been validated for diagnostic and prognostic assessment of tumors.[5–8] However, the roles of immune cells in TME of NPC remain still poorly understood.
Tumor-infiltrating immune cells (TIICs) have been reported to be effective targets of drugs and were related to the clinical outcomes.[9–11] Mechanism studies confirmed that TIICs have a dual role by conducting both host defense and tumor progression.[9,11] TIICs infiltrated in NPC, including T cells, macrophages, dendritic cells, and mast cells, and it had been reported that TIICs in NPC had significant prognostic values.[12–14]
CIBERSORT, a gene expression-based deconvolution algorithm, was developed by Newman et al to assess the abundance of member cell types in a mixed-cell population.[15] The landscape of TIICs can properly be presented by this method. Due to the excellent performance of CIBERSORT, its application in researching cell heterogeneity has attached more importance.[11,16] Therefore, we are the first to apply CIBERSORT to quantify 22 TIICs in NPC.
Importantly, we have validated our results in The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov) database. We hoped that our study would facilitate patient consultation and individualized treatment.
2. Methods
2.1. Data acquisition
This study was performed by using the data from the GEO database (https://www.ncbi.nlm.nih.gov/geo/). The available dataset (GSE64634) contained gene expression profiles of NPC patients’ tumor and normal tissues was detected and downloaded, which was updated on 25 March 2019. GSE64634, based on GPL570 platform ([HG-U133_Plus_2] Affymetrix Human Genome U133 Plus 2.0 Array), was submitted by Xiong, Zeng, and Li. The GSE64634 dataset contained 16 samples, including 12 NPC samples and 4 normal nasopharyngeal samples.
2.2. Evaluation of TIICs
ID conversion and the grouping of samples were conducted by Perl (The Perl Programming Language, version 5.28.1, http://www.perl.org). The extracted gene expression data were normalized by using the limma package of R (The R Project for Statistical Computing, version 3.5.3, https://www.r-project.org). Then, the normalized gene expression data were included to infer the relative proportions of 22 types of infiltrating immune cells by utilizing the CIBERSORT algorithm. CIBERSORT is a deconvolution algorithm based on the gene expression data, which uses a “signature matrix” of 547 genes’ expression values to characterize immune cell composition. So, we uploaded the normalized gene expression data to the CIBERSORT website (http://cibersort.stanford.edu/) by using the default algorithm of the signature matrix at 1000 permutations. CIBERSORT was applied to calculate the P value for the deconvolution for each sample by using Monte Carlo sampling, which provided a measure of confidence in the result.[15] The samples with P < .05 were included.
2.3. Visualization and evaluation of the filtered data
The filtered data were utilized to draw the bar plot, the heat map, the correlation heat map, and the violin plot by R. Principal component analysis (PCA) was also performed by using the ggplot2 package of R. The bar plot and the heat map were drawn to visualize the proportions of 22 types of infiltrating immune cells. The correlation heat map was drawn to visualize the correlations of 22 types of infiltrating immune cells by using the corrplot package of R. Variables had been decided by average linkage clustering. The violin plot was drawn to visualize the expression differences in 22 types of infiltrating immune cells by using the vioplot package of R. PCA is one of the most extensive data dimensionality reduction algorithms.[17–19]
2.4. Validation in the TGGA database
TGGA database was applied to validate our results. Clinical data of survival and outcome were also downloaded from the TCGA database.
3. Results
3.1. Data normalization, CIBERSORT analysis, and data filtering
Before performing CIBERSORT analysis, data normalization was conducted (Supplementary Table 1). Owing to the current technical limitations, the landscape of immune infiltration in NPC has not yet been entirely revealed. By using CIBERSORT algorithm, it would be the first time to observe the difference of immune infiltration between cancer and normal tissue of 22 subpopulations of immune cells in NPC relative studies. Based on the criteria of P < .05, 16 samples were all included (Table 1).
Table 1.
The filtered data after CIBERSORT analysis between normal samples and NPC samples.

3.2. The landscape of immune infiltration in NPC
Figure 1 summarizes the results between 4 normal nasopharyngeal samples and 12 NPC samples. The proportions of some immune cells in NPC vary significantly between both intragroup and intergroup. Therefore, we inferred that variation in TIICs proportions may be an intrinsic feature, which could be utilized to characterize the individual differences. Then, the proportions of immune cells from 16 samples showed the obvious group-bias clusterings and individual differences by the PCA plot (Fig. 2), whereas the proportions of some TIICs subpopulations were strongly correlated (Fig. 3). Compared with normal samples, NPC samples contained a higher proportion for M1 macrophages, whereas memory B cells and CD4 memory resting T cells were relatively lower (Fig. 4). Furthermore, the NPC samples and normal samples were separated into 2 discrete groups by using the hierarchical clustering based on 22 immune cells subpopulation (Fig. 5). Therefore, these results indicated that aberrant immune infiltration may have important clinical values in NPC.
Figure 1.
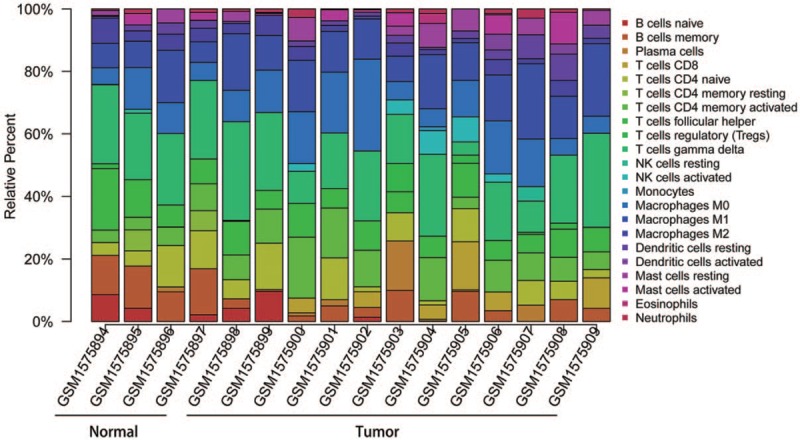
The bar plot of immune infiltration between normal samples and NPC samples. NK = Natural killer.
Figure 2.
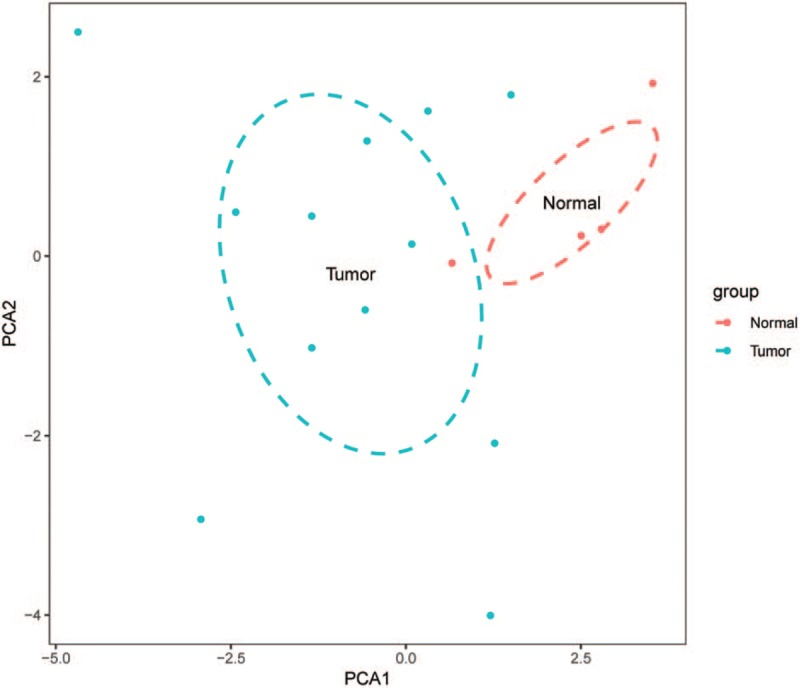
Principal components analysis performed on normal samples and NPC samples.
Figure 3.
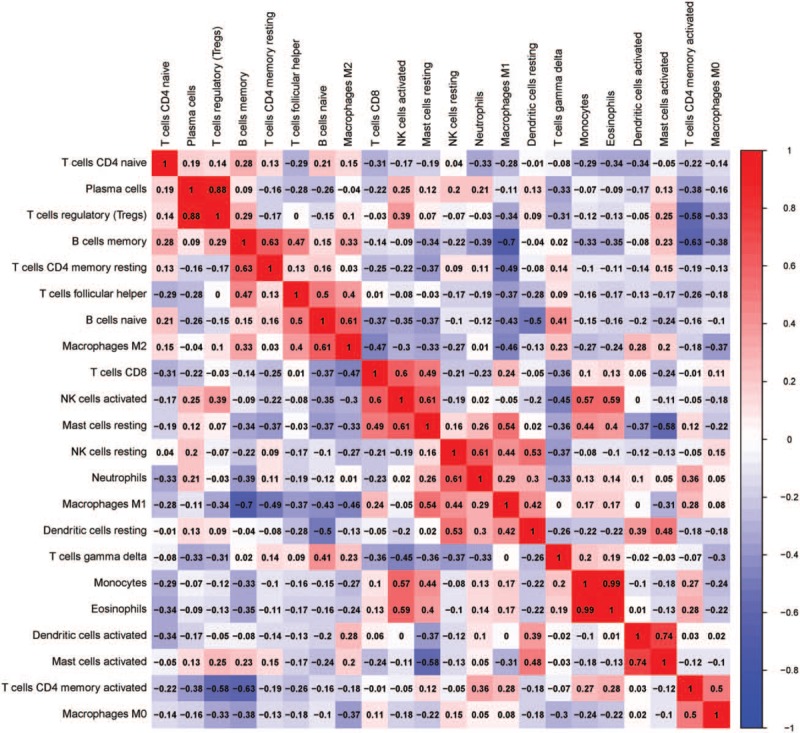
Correlation matrix of all 22 immune cell proportions and immune cytolytic activity between normal samples and NPC samples.
Figure 4.
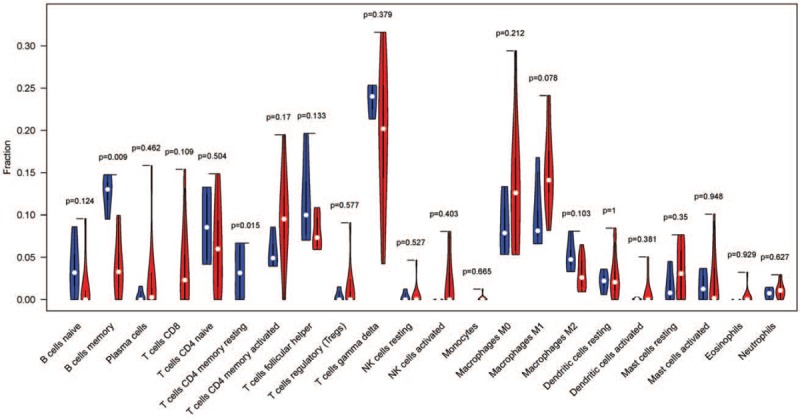
The violin plot of the 22 immune cell proportions between normal samples and NPC samples.
Figure 5.
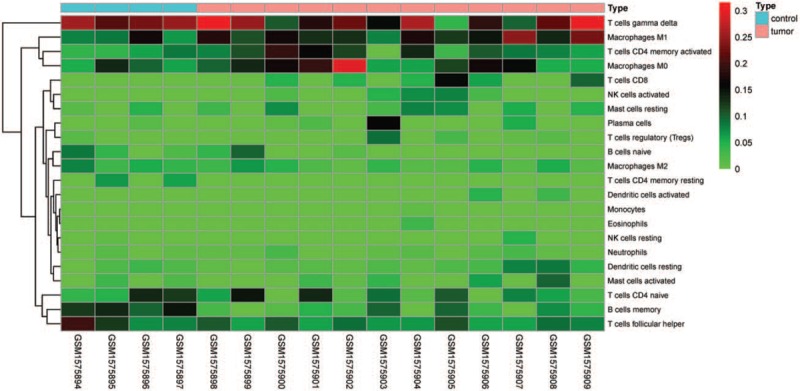
Heat map of the 22 immune cell proportions between normal samples and NPC samples.
3.3. Survival analyses in the TGGA database
Based on the TGGA database, we conducted the survival analyses (Fig. 6) of the significant tumor-infiltrating immune cells (M1 macrophages, memory B cells, and CD4 memory resting T cells).
Figure 6.
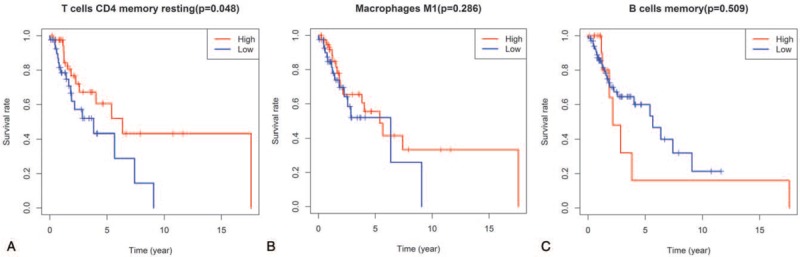
Survival analyses the significant tumor-infiltrating immune cells (A, CD4 memory resting T cells; B, M1 macrophages; C, memory B cells, and) in the TGGA database.
4. Discussion
Except for malignant neoplastic cells, immune cells, endothelial cells, fibroblasts, cytokines, growth factors, and chemokines are also included in tumor tissues.[11,12,14,16] Those components and their interactions can exert inhibitory effects on the malignant cells, whereas, during the tumor progression, tumor cells may circumvent these inhibitory signals and exploit immune cells or others for tumor growth, metastasis, or invasion [20]. A growing number of studies reported that TIICs may predict the clinical outcome and treatment response of various tumors.[11,12,16,21] Actually, many studies showed that increased mature T cells, memory T-cells, and dendritic cells infiltration are closely related to a favorable prognosis, but immune suppressive regulatory T cells are opposite.[9,10,22–25] However, because of the technical limitations, current studies have been limited to show a narrow view of the immune response. To identify TIICs, researchers always evaluated them with the single surface marker by immunohistochemistry-based analysis. But this method was less effective for discriminating related cell types and may be misled by other marker proteins, whereas inconsistent results would often be reported by different researchers.
Nowadays, integrating genomic profiles and the deconvolution algorithm can possibly resolve relative proportions of TIICs and overcome the defect of immunohistochemistry-based analysis.[26] However, as an emerging technology, CIBERSORT analysis in NPC was not validated. Thus, CIBERSORT analysis was first applied to infer the proportions of 22 immune cells from NPC transcriptome. Gentles et al reported that the CIBERSORT results were in accordance with the results of flow cytometry and immunohistochemistry experiment.[21]
By using CIBERSORT, we can directly compare the alteration of 22 immune cells between normal samples and NPC samples for the first time and found the significant changes occurred in both intragroup and intergroup. Furthermore, PCA displayed that there were the distinct group-bias clusterings in proportions of 22 immune cells from 16 samples, which indicated that the variation of TIICs may characterize the individual differences and have clinical values. The comprehensive assessment of immune infiltration in NPC was conducted based on the deconvolution of the normalized gene expression data from GSE64634. This would be of great interest in immunomodulatory therapies of NPC. We found that the increase of M1 macrophages and the decrease of memory B cells and CD4 memory resting T cells may be associated with tumorigenesis of NPC. These differences might be important determinants for the patients’ prognosis. Our unbiased method confirmed that the immune infiltration is associated with NPC tumorigenesis. The survival analyses based on the TCGA database showed that CD4 momory resting T cells would be a predicitive outcome signature in NPC.
Olkhanud et al reported that the primary role of designated tumor-evoked Bregs in lung metastases was to induce TGF-β-dependent conversion of FoxP3(+) T-regulatory cells from resting CD4(+) T cells.[27] Increase of regulatory T cells (Tregs) in the TME predicted worse survival of patients with various types of cancer.[28–31] Yu et al performed a study to explore the predictive role of tumor-associated macrophages (TAMs) in NPC.[32] The results showed that the patient's risk group based on the level of CD163+ TAMs was an independent predictor of survival, which may facilitate patients’ individualized treatments. Ooft et al reported M2 spectrum macrophages were probably more prominent in EBV-negative NPC.[33] Huang et al proved that tumor infiltration of TAMs was closely correlated with serum EBV infection titers and NPC progression.[34] Lin et al reported that CD4 T cell memory and antibody responses were directed against the pneumococcal histidine triad proteins PhtD and PhtE following nasopharyngeal colonization and immunization.[23] Voigt et al and Vahidi Y both proved that CD4+ T cells were related to tumor process.[25,35] Therefore, we could conclude that M1 macrophages, memory B cells, and CD4 memory resting T cells may play pivotal roles in the development of NPC and the differentiation may be possible therapeutic targets.
Moreover, our data first revealed the detail of infiltration of 22 TIICs in NPC. Expression of plasma cells was strongly correlated with expression of regulatory T cells (Tregs), expression of monocytes was strongly correlated with expression of eosinophils, and expression of activated dendritic cells was strongly correlated with expression of activated mast cells. Jang et al revealed that regulatory T cells control humoral autoimmunity by inhibiting the development of plasma cells.[36] Koyabu et al conducted an analysis of regulatory T cells and IgG4-positive plasma cells in patients of autoimmune liver diseases and IgG4-related sclerosing cholangitis, which indicated that abundant infiltration of regulatory T cells (Tregs) might impact the switching of B cells to IgG4-producing plasmacytes.[22] Podgorny et al indicated that low counts of monocytes and eosinophils were related to posten-graftment infections following allogeneic hematopoietic cell transplantation.[37] Choi et al showed that perivascular dendritic cells elicited anaphylaxis by relaying allergens to mast cells.[38] Leon et al proved that the characteristic location of dendritic cells play a role as antigen-receptor cells, whereas mast cells may play dual roles in the tumor.[39] Wachter et al revealed that coronary atherosclerosis in chronic kidney disease patients shows a significantly higher number of mast cells, whereas dendritic cells were less frequent.[24]
However, there were several limitations in our study. First, the most significant limitation is that the CIBERSORT analysis was based on the limited gene data, which may deviate in cells’ heterotypic interactions, disease-induced dysregulation, or phenotypic plasticity. Second, the included samples are unpaired. So more larger paired gene data are required to validate our conclusion. Third, CIBERSORT showed a lower estimation bias than other methods, but it may overestimate or underestimate some cell types. So, necessary experiments are needed to warrant our conclusion.
As a consequence, our analysis of 22 immune cells in NPC was associated with tumorigenesis, which had the potential to help NPC patients to benefit from immunomodulatory therapies, as well as highlighting possible targets for new drugs. To uncover the clinical and biological significance of TIICs in NPC, more larger paired gene data and necessary experiments are required to validate our conclusion.
Author contributions
Conceptualization: Guan-Jiang Huang, Bao-Xinzi Liu.
Data curation: Meng-Si Luo, Guan-Jiang Huang.
Formal analysis: Meng-Si Luo.
Funding acquisition: Bao-Xinzi Liu.
Investigation: Meng-Si Luo, Guan-Jiang Huang.
Methodology: Meng-Si Luo, Guan-Jiang Huang, Bao-Xinzi Liu.
Project administration: Bao-Xinzi Liu.
Resources: Guan-Jiang Huang, Bao-Xinzi Liu.
Software: Guan-Jiang Huang.
Supervision: Bao-Xinzi Liu.
Validation: Meng-Si Luo.
Visualization: Meng-Si Luo, Guan-Jiang Huang.
Writing – original draft: Meng-Si Luo, Guan-Jiang Huang.
Writing – review & editing: Guan-Jiang Huang, Bao-Xinzi Liu.
Supplementary Material
Footnotes
Abbreviations: AJCC/UICC = American Joint Committee on Cancer and the International Union for Cancer Control, NPC = nasopharyngeal carcinoma, PCA = principal component analysis, TCGA = The Cancer Genome Atlas, TIIC = tumor-infiltrating immune cell, TME = tumor microenvironment.
How to cite this article: Luo MS, Huang GJ, Liu BX. Immune infiltration in nasopharyngeal carcinoma based on gene expression. Medicine. 2019;98:39(e17311).
M-SL and G-JH have contributed equally to this work as joint first authors.
Ethics: Ethics approval and patient written informed consent were not required because all analyses in our study were performed based on data from the GEO database.
Disclosure: The authors have no finding and conflicts of interest to disclose. The authors are responsible for the content and writing of the paper.
Supplemental Digital Content is available for this article.
References
- [1].Tang XR, Li YQ, Liang SB, et al. Development and validation of a gene expression-based signature to predict distant metastasis in locoregionally advanced nasopharyngeal carcinoma: a retrospective, multicentre, cohort study. Lancet Oncol 2018;19:382–93. [DOI] [PubMed] [Google Scholar]
- [2].Lee AW, Ma BB, Ng WT, et al. Management of nasopharyngeal carcinoma: current practice and future perspective. J Clin Oncol 2015;33:3356–64. [DOI] [PubMed] [Google Scholar]
- [3].Yan L, Xu F, Dai CL. Relationship between epithelial-to-mesenchymal transition and the inflammatory microenvironment of hepatocellular carcinoma. J Exp Clin Cancer Res 2018;37:203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [4].Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell 2011;144:646–74. [DOI] [PubMed] [Google Scholar]
- [5].Guo G, Wang Y, Zhou Y, et al. Immune cell concentrations among the primary tumor microenvironment in colorectal cancer patients predicted by clinicopathologic characteristics and blood indexes. J Immunother Cancer 2019;7:179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Zhou C, Qian W, Li J, et al. High glucose microenvironment accelerates tumor growth via SREBP1-autophagy axis in pancreatic cancer. J Exp Clin Cancer Res 2019;38:302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [7].Zhang Q, Zhang F, Li S, et al. A multifunctional nanotherapy for targeted treatment of colon cancer by simultaneously regulating tumor microenvironment. Theranostics 2019;9:3732–53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [8].Jiang T, Shi J, Dong Z, et al. Genomic landscape and its correlations with tumor mutational burden, PD-L1 expression, and immune cells infiltration in Chinese lung squamous cell carcinoma. J Hematol Oncol 2019;12:75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Kim HS, Lee JH, Nam SJ, et al. Association of PD-L1 expression with tumor-infiltrating immune cells and mutation burden in high-grade neuroendocrine carcinoma of the lung. J Thorac Oncol 2018;13:636–48. [DOI] [PubMed] [Google Scholar]
- [10].Dadabayev AR, Sandel MH, Menon AG, et al. Dendritic cells in colorectal cancer correlate with other tumor-infiltrating immune cells. Cancer Immunol Immunother 2004;53:978–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [11].Zhang S, Zhang E, Long J, et al. Immune infiltration in renal cell carcinoma. Cancer Sci 2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [12].Ono T, Azuma K, Kawahara A, et al. Prognostic stratification of patients with nasopharyngeal carcinoma based on tumor immune microenvironment. Head Neck 2018;40:2007–19. [DOI] [PubMed] [Google Scholar]
- [13].Tan L, Wong B, Gangodu NR, et al. Enhancing the immune stimulatory effects of cetuximab therapy through TLR3 signalling in Epstein-Barr virus (EBV) positive nasopharyngeal carcinoma. Oncoimmunology 2018;7:e1500109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [14].Chan OS, Kowanetz M, Ng WT, et al. Characterization of PD-L1 expression and immune cell infiltration in nasopharyngeal cancer. Oral Oncol 2017;67:52–60. [DOI] [PubMed] [Google Scholar]
- [15].Newman AM, Liu CL, Green MR, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 2015;12:453–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [16].Zhou R, Zhang J, Zeng D, et al. Immune cell infiltration as a biomarker for the diagnosis and prognosis of stage I-III colon cancer. Cancer Immunol Immunother 2019;68:433–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [17].Azadeh S, Hobbs BP, Ma L, et al. Integrative Bayesian analysis of neuroimaging-genetic data through hierarchical dimension reduction. Proc IEEE Int Symp Biomed Imaging 2016;2016:824–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Reich D, Price AL, Patterson N. Principal component analysis of genetic data. Nat Genet 2008;40:491–2. [DOI] [PubMed] [Google Scholar]
- [19].Stewart S, Ivy MA, Anslyn EV. The use of principal component analysis and discriminant analysis in differential sensing routines. Chem Soc Rev 2014;43:70–84. [DOI] [PubMed] [Google Scholar]
- [20].Klemm F, Joyce JA. Microenvironmental regulation of therapeutic response in cancer. Trends Cell Biol 2015;25:198–213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Gentles AJ, Newman AM, Liu CL, et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat Med 2015;21:938–45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Koyabu M, Uchida K, Miyoshi H, et al. Analysis of regulatory T cells and IgG4-positive plasma cells among patients of IgG4-related sclerosing cholangitis and autoimmune liver diseases. J Gastroenterol 2010;45:732–41. [DOI] [PubMed] [Google Scholar]
- [23].Khan MN, Pichichero ME. CD4 T cell memory and antibody responses directed against the pneumococcal histidine triad proteins PhtD and PhtE following nasopharyngeal colonization and immunization and their role in protection against pneumococcal colonization in mice. Infect Immun 2013;81:3781–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Wachter DL, Neureiter D, Campean V, et al. In-situ analysis of mast cells and dendritic cells in coronary atherosclerosis in chronic kidney disease (CKD). Histol Histopathol 2018;33:871–86. [DOI] [PubMed] [Google Scholar]
- [25].Vahidi Y, Faghih Z, Talei AR, et al. Memory CD4(+) T cell subsets in tumor draining lymph nodes of breast cancer patients: a focus on T stem cell memory cells. Cell Oncol (Dordr) 2018;41:1–1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26].Chen B, Khodadoust MS, Liu CL, et al. Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol Biol 2018;1711:243–59. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Olkhanud PB, Damdinsuren B, Bodogai M, et al. Tumor-evoked regulatory B cells promote breast cancer metastasis by converting resting CD4(+) T cells to T-regulatory cells. Cancer Res 2011;71:3505–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [28].Shah W, Yan X, Jing L, et al. A reversed CD4/CD8 ratio of tumor-infiltrating lymphocytes and a high percentage of CD4(+)FOXP3(+) regulatory T cells are significantly associated with clinical outcome in squamous cell carcinoma of the cervix. Cell Mol Immunol 2011;8:59–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].de Ruiter EJ, Ooft ML, Devriese LA, et al. The prognostic role of tumor infiltrating T-lymphocytes in squamous cell carcinoma of the head and neck: a systematic review and meta-analysis. Oncoimmunology 2017;6:e1356148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [30].Boucek J, Mrkvan T, Chovanec M, et al. Regulatory T cells and their prognostic value for patients with squamous cell carcinoma of the head and neck. J Cell Mol Med 2010;14:426–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [31].Yu Y, Ke L, Lv X, et al. The prognostic significance of carcinoma-associated fibroblasts and tumor-associated macrophages in nasopharyngeal carcinoma. Cancer Manag Res 2018;10:1935–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [32].Ooft ML, van Ipenburg JA, Sanders ME, et al. Prognostic role of tumour-associated macrophages and regulatory T cells in EBV-positive and EBV-negative nasopharyngeal carcinoma. J Clin Pathol 2018;71:267–74. [DOI] [PubMed] [Google Scholar]
- [33].Huang D, Song SJ, Wu ZZ, et al. Epstein-Barr virus-induced VEGF and GM-CSF drive nasopharyngeal carcinoma metastasis via recruitment and activation of macrophages. Cancer Res 2017;77:3591–604. [DOI] [PubMed] [Google Scholar]
- [34].Voigt C, May P, Gottschlich A, et al. Cancer cells induce interleukin-22 production from memory CD4(+) T cells via interleukin-1 to promote tumor growth. Proc Natl Acad Sci USA 2017;114:12994–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [35].Jang E, Cho WS, Cho ML, et al. Foxp3+ regulatory T cells control humoral autoimmunity by suppressing the development of long-lived plasma cells. J Immunol 2011;186:1546–53. [DOI] [PubMed] [Google Scholar]
- [36].Podgorny PJ, Pratt LM, Liu Y, et al. Low counts of B cells, natural killer cells, monocytes, dendritic cells, basophils, and eosinophils are associated with postengraftment infections after allogeneic hematopoietic cell transplantation. Biol Blood Marrow Transplant 2016;22:37–46. [DOI] [PubMed] [Google Scholar]
- [37].Choi HW, Suwanpradid J, Kim IH, et al. Perivascular dendritic cells elicit anaphylaxis by relaying allergens to mast cells via microvesicles. Science 2018;362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [38].Leon A, Ceausu Z, Ceausu M, et al. Mast cells and dendritic cells in basal cell carcinoma. Rom J Morphol Embryol 2009;50:85–90. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


