The model Solanaceous plant Nicotiana benthamiana is established as a system for analysis of the role of EDS1 complexes in signaling downstream of TIR-domain containing immune receptors
Abstract
Heterodimeric complexes containing the lipase-like protein ENHANCED DISEASE SUSCEPTIBILITY1 (EDS1) are regarded as central regulators of plant innate immunity. In this context, a complex of EDS1 with PHYTOALEXIN DEFICIENT4 (PAD4) is required for basal resistance and signaling downstream of immune receptors containing an N-terminal Toll-interleukin-1 receptor-like domain (TNLs) in Arabidopsis (Arabidopsis thaliana). Here we analyze EDS1 functions in the model Solanaceous plant Nicotiana benthamiana (Nb). Stable Nb mutants deficient in EDS1 complexes are not impaired in basal resistance, a finding which contradicts a general role for EDS1 in immunity. In Nb, PAD4 demonstrated no detectable immune functions, but TNL-mediated resistance responses required EDS1 complexes incorporating a SENESCENCE ASSOCIATED GENE101 (SAG101) isoform. Intriguingly, SAG101 is restricted to those genomes also encoding TNL receptors, and we propose it may be required for TNL-mediated immune signaling in most plants, except the Brassicaceae. Transient complementation in Nb was used for accelerated mutational analyses while avoiding complex biotic interactions. We identify a large surface essential for EDS1-SAG101 immune functions that extends from the N-terminal lipase domains to the C-terminal EDS1-PAD4 domains and might mediate interaction partner recruitment. Furthermore, this work demonstrates the value of genetic resources in Nb, which will facilitate elucidation of EDS1 functions.
INTRODUCTION
Plants lack mobile immune cells but have evolved an elaborate innate immune system to defend against invading pathogens (Spoel and Dong, 2012; Jones et al., 2016). Cell surface-resident pattern recognition receptors (PRRs) can detect pathogen-/microbe-associated molecular patterns (PAMPs/MAMPs). MAMP perception and PRR activation induces PRR-triggered immunity (PTI; also referred to as MAMP/PAMP-triggered immunity), a multifaceted, low-level immune response efficient against a broad spectrum of nonadapted pathogens (Macho and Zipfel, 2014; Yu et al., 2017). However, many host-adapted pathogens use effector proteins, which are secreted directly into the host cell cytoplasm, to suppress PTI (Macho and Zipfel, 2015; Büttner, 2016). As a second layer of the plant immune system, effectors can become recognized by plant resistance proteins (R proteins) in resistant plant lines. Effector recognition induces a rapid and efficient immune response termed effector-triggered immunity (ETI; Jones et al., 2016; Khan et al., 2016). The ETI response commonly culminates in programmed cell death at infection sites, the hypersensitive response (HR), which in most cases correlates with inhibition of plant colonization by the pathogen (Cui et al., 2015; Büttner, 2016).
Most plant R proteins are nucleotide-binding domain–leucine-rich repeat (NLR)–type immune receptors. The canonical NLR architecture consists of a C-terminal leucine-rich repeat domain, a central nucleotide-binding domain, and a variable N-terminal domain (Monteiro and Nishimura, 2018). Structurally similar NLR receptors operate in animal innate immunity and often function by ligand-dependent oligomerization and recruitment of signaling adapters via oligomeric N-terminal domain assemblies (Bentham et al., 2017; Shen et al., 2019). Until recently, plant NLRs were less well understood than animal NLRs. However, according to a general working model, the leucine-rich repeat domain often defines specificity, the nucleotide-binding domain acts as an ATP-driven switch controlling the transition of the receptor from a resting to an active signaling state, and the N-terminal domain conveys downstream signaling (Bernoux et al., 2011, 2016; Maekawa et al., 2011a, 2011b; Takken and Goverse, 2012). The N-terminal domains of most plant NLRs are Toll-interleukin-1 receptor (TIR) or coiled-coil (CC) domains. Beyond analogy to animal NLRs, a function in downstream signaling is supported by the induction of HR-like cell death upon expression of TIR or CC domains alone (Swiderski et al., 2009; Bernoux et al., 2011; Collier et al., 2011; Maekawa et al., 2011b).
Recent structural elucidation of the Arabidopsis (Arabidopsis thaliana) immune receptor HOPZ-ACTIVATED RESISTANCE1 (ZAR1) provided groundbreaking new insights to our knowledge of CC-type plant NLRs (CNLs; Wang et al., 2019a, 2019b): ZAR1 persists as an ADP-bound monomer in the resting state and forms a pentameric “resistosome” upon effector-triggered nucleotide exchange. In the resistosome, the ZAR1 CC domain assembly forms a funnel-shaped structure that may directly insert into the plasma membrane to interfere with membrane integrity or to function as an ion channel. Thus, CNLs most likely initiate the HR directly, and indirectly regulate downstream immune signaling (Wang et al., 2019a). Although several lines of evidence suggest that additional CNLs form ZAR1-like resistosomes (Wang et al., 2019a), such a direct function in HR initiation is unlikely to apply to TIR-type NLRs (TNLs). First, TNLs were recently shown to require so-called helper NLRs of the CC-type to mediate immunity (Qi et al., 2018; Castel et al., 2019; Wu et al., 2019). Furthermore, all known responses mediated by TNLs are dependent on ENHANCED DISEASE SUSCEPTIBILITY1 (EDS1; Aarts et al., 1998; Wirthmueller et al., 2007).
EDS1 was identified in Arabidopsis in a screen for mutants impaired in resistance to the obligate biotrophic oomycete Hyaloperonospora arabidopsidis (Hpa; Parker et al., 1996; Falk et al., 1999). EDS1 interacts with two sequence-related proteins, PHYTOALEXIN DEFICIENT4 (PAD4) and SENESCENCE-ASSOCIATED GENE101 (SAG101; Feys et al., 2001, 2005). All three proteins share homology with eukaryotic lipases (α/β-hydrolases) in their N termini and contain a C-terminal EP (EDS1-PAD4) domain. Occurrence of the unique EP domain together with an N-terminal lipase-like domain defines the EDS1 family (Wagner et al., 2013). Although a catalytic triad (S-D-H including a GXSXG motif) is conserved in EDS1 and PAD4 orthologs, it is not required for immune functions of EDS1-PAD4 complexes in Arabidopsis, suggesting a noncatalytic mode of action (Wagner et al., 2013). Elucidation of the EDS1-SAG101 heterodimer structure and modeling of the EDS1-PAD4 complex showed that EDS1 engages in mutually exclusive heterodimers with PAD4 or SAG101 (Wagner et al., 2013), which differentially contribute to immunity in Arabidopsis: Loss of EDS1-PAD4 complexes (in pad4 mutant plants) severely impairs immune signaling, whereas loss of EDS1-SAG101 (in sag101 mutant plants) is largely compensated by presence of EDS1-PAD4 (Feys et al., 2005; Wagner et al., 2013). Complete loss of EDS1-based complexes (in eds1 single or pad4 sag101 double mutant plants) abolishes TNL-mediated resistance signaling. In agreement with genetic data, structure-guided mutations untethering EDS1 from PAD4 and SAG101 provided strong evidence for heterodimeric assemblies executing immune functions (Wagner et al., 2013).
In addition to its strict requirement for TNL-mediated immune responses, EDS1 also contributes to resistance mediated by some CNLs and to basal resistance (Wiermer et al., 2005; Venugopal et al., 2009; Cui et al., 2017). Basal resistance is a somewhat promiscuously employed term, generally used to describe the residual resistance observed upon challenge of a wild-type plant with a virulent pathogen isolate. To that end, for example, Pseudomonas syringae (Pst) strain DC3000 bacteria grow significantly better in eds1 mutants than in wild-type lines. Similarly, Arabidopsis accession Columbia-0 (Col-0) is resistant to Pst bacteria translocating AvrRpt2, recognized by the CNL RPS2 (Bent et al., 1994; Mindrinos et al., 1994), but resistance is impaired in plants lacking EDS1 and the defense-associated hormone salicylic acid (SA; Venugopal et al., 2009; Vlot et al., 2009; Cui et al., 2017). It remains unclear whether EDS1 has distinct functions in basal, TNL-, and CNL-mediated resistance. An alternative explanation is that abolishment of weak TNL signaling, provoked by minor effector recognition in interactions termed “compatible,” is at the basis of enhanced pathogen growth in eds1 plants (Jones and Dangl, 2006; Poland et al., 2009; Krasileva et al., 2011; Kushalappa et al., 2016; Wei et al., 2018). Similarly, combined loss of weak recognition mediated by TNLs (eds1) and SA-mediated bolstering of immune responses (sid2) may lower immune capacities in eds1 sid2 plants below a critical threshold, allowing plant colonization by otherwise incompatible Pst AvrRpt2 bacteria without any direct contribution of EDS1 to CNL-mediated responses. This explanation is supported by the notion that EDS1 and SA function additively, and not redundantly, in RPS2-mediated resistance, and by induction of residual cell death by AvrRpt2 in the absence of EDS1 and SA (Cui et al., 2017). A general function for EDS1 complexes in plant immunity is suggested by the conservation of EDS1 and PAD4, but not SAG101, in plant lineages that lost TNLs in the course of evolution, such as monocotyledonous plants and the eudicots Aequilegia courulea and Mimulus guttatus (Collier et al., 2011; Jacob et al., 2013; Wagner et al., 2013). Accordingly, it was proposed that EDS1-PAD4 form an ancient module regulating basal resistance, which has been co-opted for TNL-mediated immunity in eudicots (Feys et al., 2005; Rietz et al., 2011), but EDS1 immune functions in the absence of TNLs remain largely unexplored (Chen et al., 2018). Indeed, EDS1 functions have mainly been analyzed in Arabidopsis.
We and others generated eds1 mutant lines in the model Solanaceous plant Nicotiana benthamiana (Ordon et al., 2017; Schultink et al., 2017). In N. benthamiana, genetic analyses are hampered by allotetraploidy and incomplete genome sequences, but a key advantage is the efficient transient protein expression after agroinfiltration (Goodin et al., 2008; Bombarely et al., 2012; Naim et al., 2012). Stable Nbeds1 mutant lines confirmed the conserved role of EDS1 in TNL signaling and revealed EDS1-dependent recognition of the effector protein XopQ from Xanthomonas campestris pv vesicatoria (Xcv; also Xanthomonas euvesicatoria) in this species (Peart et al., 2002; Hu et al., 2005; Adlung et al., 2016; Adlung and Bonas, 2017; Qi et al., 2018). Xcv is the causal agent of bacterial spot disease on pepper (Capsicum annuum) and tomato (Solanum lycopersicum) and is virulent in N. benthamiana after deletion of xopQ or in Nbeds1 mutant plants. The corresponding TNL receptor, Recognition of XopQ 1 (Roq1), was subsequently identified (Schultink et al., 2017), completing the signal transduction pathway from XopQ toward EDS1-conditioned immunity in the Xcv – N. benthamiana interaction. Also, N requirement gene 1 (NRG1) was identified as a novel component in TNL signaling downstream of NbEDS1 (Qi et al., 2018). However, the molecular role of EDS1 complexes in TNL signaling remains elusive.
In this study, we genetically dissected the EDS1 family in N. benthamiana. Although major immune functions in Arabidopsis reside in EDS1-PAD4, we show that an EDS1-SAG101 complex is essential for TNL-mediated resistance responses in N. benthamiana. TNL activation and signaling can be uncoupled from complex biotic interactions in transient complementation assays in N. benthamiana, which facilitates straightforward and simplified gene functional analyses. This system will prove seminal for future functional analyses of EDS1 complexes toward a mechanistic understanding.
RESULTS
Duplication of SAG101 in Solanaceous Plants
We previously mined genomes of tomato and N. benthamiana for homologs of EDS1 and PAD4 (Ordon et al., 2017). Single genes were identified for PAD4 in both species. For EDS1, a single copy was detected in tomato, whereas the N. benthamiana genome contains two loci with similarity to EDS1 but one is a pseudogene (Adlung et al., 2016). N. benthamiana EDS1 and PAD4 loci were targeted by genome editing to generate Nbeds1 and Nbpad4 mutant plants (Ordon et al., 2017). Growth restriction of Xcv (strain 85-10) bacteria due to Roq1-mediated XopQ recognition was abolished in Nbeds1 (Adlung et al., 2016; Schultink et al., 2017) but not in Nbpad4 plants (see later sections).
We suspected that EDS1-PAD4 and EDS1-SAG101 complexes might function redundantly in Roq1 signaling and extended genome mining to SAG101 orthologs from tomato and N. benthamiana. Two different SAG101 isoforms, termed SAG101a and SAG101b, were identified in tomato and were further duplicated in N. benthamiana in agreement with its allotetraploid genome (Supplemental Figure 1; Supplemental Data Set 1). However, NbSAG101a2 and NbSAG101b2 are likely pseudogenes, as experimentally supported gene models encode for truncated proteins (Supplemental Figure 1), and were not further inspected.
To analyze the possible biological significance of the apparent duplication of SAG101, additional genomes from Solanaceae and other species were analyzed for SAG101 orthologs. The genomes of the core eudicots Coffea canephora (Cc) and Mimulus guttatus (Mg), belonging like Solanaceae to the asterids, and of the rosid Arabidopsis were analyzed. Musa accuminata (Ma) was included as a monocotyledonous plant. A phylogeny was constructed using predicted EDS1 family proteins of these species (Figure 1). SAG101 homologs were not detected in species lacking TNL receptors (here Ma and Mg), as previously described by Wagner et al. (2013). In agreement with the distant relationship, the rosid AtSAG101 was clearly separated from SAG101 orthologs of asterid plants. Genes encoding SAG101a and SAG101b isoforms were detected in all analyzed Solanaceae genomes except pepper (Capsicum annuum, Ca), and respective proteins formed two distinct groups within the SAG101 branch (Figure 1A). The single copy SAG101 from pepper grouped within the SAG101b branch. The asterid CcSAG101 grouped more closely with Solanaceae SAG101a but with low branch-point support. This suggests that diversification of SAG101a and SAG101b most likely occurred at a time point similar to separation of the lineages leading to Cc and Solanaceae, and that SAG101a was lost in Ca relatively recently.
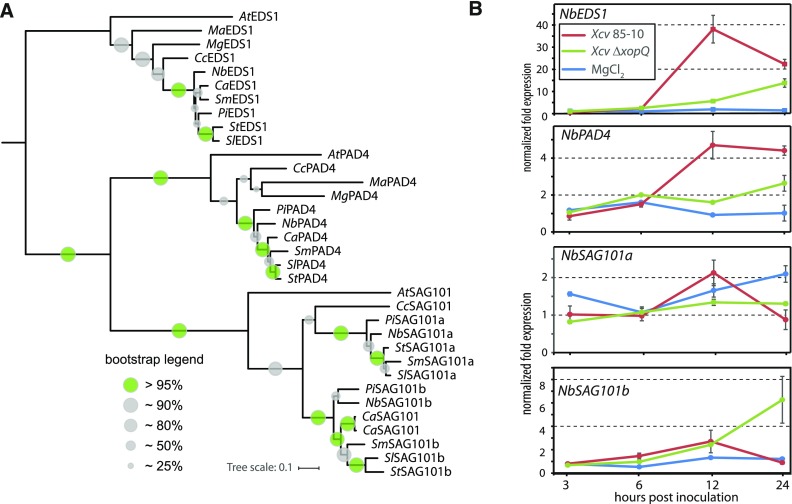
Figure 1.
Occurrence and Expression of EDS1 Family Genes in Solanaceae.
(A) Phylogenetic clustering of putative EDS1 family proteins from Solanaceae and control species. The tree was midpoint rooted. At, Arabidopsis thaliana; Ca, Capsicum annuum (SAG101 orthologs from cultivars Zunla and CM334 shown); Cc, Coffea canephora; Ma, Musa accuminata; Mg, Mimulus guttatus; Pi, Petunia inflata; Nb, Nicotiana benthamiana; Sl, Solanum lycopersicum; Sm, Solanum melongena.
(B) Expression of EDS1 family genes in N. benthamiana. Plants were challenged with virulent (Xcv ΔxopQ) or avirulent (Xcv 85-10) Xanthomonas campestris pv vesicatoria bacteria or mock treated (MgCl2). RNA was extracted at indicated time points, and expression of EDS1 family genes measured by quantitative RT-PCR. Displayed data originates from normalization to PP2A expression, and similar results were obtained when using Elongation Factor 1-α (EF1α) for normalization. Data points represent means of four biological replicates with se shown.
Retention of two SAG101 isoforms in most Solanaceae genomes is indicative of functional diversification. To get an insight into expression of SAG101 isoforms and EDS1 family genes in general, we assessed a public RNA sequencing (RNA-seq) data set from tomato plants treated with different bacterial isolates or the MAMP flgII-28 (Rosli et al., 2013) using the TomExpress platform (Supplemental Figure 2; Zouine et al., 2017). All four genes were expressed in control (mock-treated) plants. Treatment with Pseudomonas fluorescens and Pseudomonas putida bacteria, inducing robust PTI responses in Solanaceae (Chakravarthy et al., 2010; Rosli et al., 2013), moderately induced expression of EDS1 family genes (6 h after treatment).
We also examined the expression of EDS1 family genes in N. benthamiana. Plants were infected with avirulent (Xcv 85-10) or virulent (Xcv ΔxopQ) Xcv bacteria. Gene expression was analyzed by RT-qPCR in a time course experiment (Figure 1B). Expression of NbEDS1 was strongly upregulated in response to avirulent Xcv 85-10 and, to a lesser extent, by the ΔxopQ mutant strain (Figure 1B). Expression peaked 12 h after infection and then declined. A similar expression profile was observed for NbPAD4, although its overall induction (∼35-fold for NbEDS1, fivefold for NbPAD4; Figure 1B) was less pronounced. NbSAG101a and NbSAG101b were expressed but not regulated under infection conditions. These data indicate that two SAG101 isoforms are expressed in tomato and N. benthamiana, and they may form heterocomplexes with EDS1 and thus contribute to immune signaling.
Localization and Complex Formation of tomato EDS1 Family Proteins
EDS1 family proteins from tomato (Solanum lycopersicum, Sl) are very similar to those of N. benthamiana (percentage of identity and similarity, 79% and 86% , respectively [EDS1], 77% and 85% [PAD4], 81% and 87% [SAG101a], and 72% and 79% [SAG101b]; comparison of tomato proteins to Arabidopsis: 40% and 57% [EDS1], 38% and 53% [PAD4], 33% and 45% [SAG101a], and 34% and 45% [SAG101b]), suggesting that they can likely functionally replace each other. For simplicity, we decided to first clone cDNAs of EDS1 family genes from tomato for functional characterization, although we aimed at genetic dissection of functions of EDS1 complexes in the N. benthamiana model system.
For subcellular localization studies, SlEDS1 was fused to mCherry, and SlPAD4, SlSAG101a, and SlSAG101b were fused to mEGFP, respectively, in 35S promoter-controlled expression constructs. Fusions proteins were transiently expressed alone or in combination in N. benthamiana, and tissues were analyzed by live-cell imaging (Figure 2; Supplemental Figure 3). SlEDS1 and SlPAD4 were detected in both the cytoplasm and the nucleus, whereas SlSAG101a was located exclusively in the nucleus. Similar subcellular localization patterns were described for Arabidopsis orthologs (Feys et al., 2005; García et al., 2010). In coexpression with SlSAG101a, SlEDS1 was detected exclusively in the nucleus (compared with Figure 2A and Supplemental Figure 3A). In contrast, SlSAG101b was nucleocytoplasmically distributed both alone and in combination with SlEDS1. In summary, SlEDS1 family proteins from tomato localize to cytoplasm and/or nucleus, although we cannot fully exclude cell wall or membrane association. All proteins were detected on immunoblots, and the GFP-tagged SlPAD4, SlSAG101a, and SlSAG101b appeared to be stabilized by coexpression of SlEDS1 (Supplemental Figure 3B).
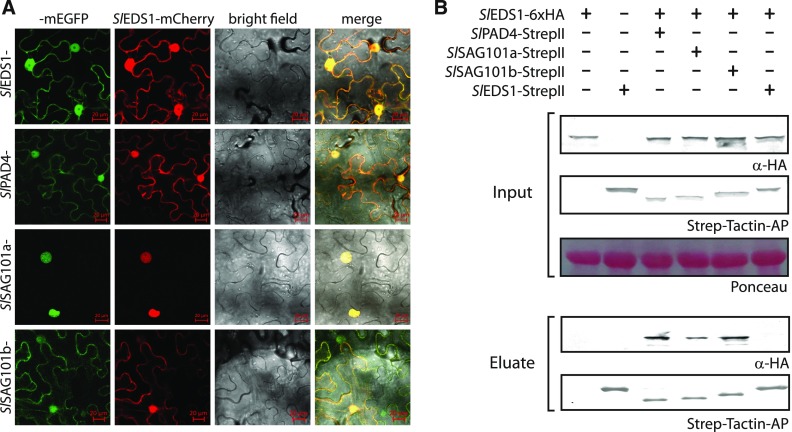
Figure 2.
Complex Formation and Localization of Tomato EDS1 Family Proteins.
(A) Protein localization in living cells detected by confocal laser scanning microscopy. Indicated proteins (from tomato) were transiently coexpressed as GFP fusions together with SlEDS1 in N. benthamiana leaf tissues by agroinfiltration, and protein localization was analyzed 3 dpi. Localization of single proteins and integrity of fluorophore fusions is shown in Supplemental Figure 3. Scale bar = 20 µm.
(B) Formation of complexes by tomato EDS1 proteins. Indicated proteins were transiently (co)expressed in N. benthamiana by agroinfiltration. At 3 dpi, extracts were used for StrepII purification, and total extracts and eluates analyzed by immunoblotting. Ponceau staining is shown as loading control.
Förster resonance energy transfer (FRET) and acceptor photobleaching (FRET-APB) was used to probe formation of SlEDS1-based complexes in living cells. mCherry and mEGFP-tagged proteins were expressed from a single T-DNA for reduced variation in coexpression rates for FRET analyses (Hecker et al., 2015). Robust FRET was detected upon coexpression of SlEDS1-mCherry and SlPAD4, SlSAG101a or SlSAG101b, but not SlEDS1, fused with mEGFP (Supplemental Fig. 3C). Complex formation was further analyzed by protein copurification (Figure 2B). In line with results obtained by FRET-APB, SlEDS1 (fused to a 6xHA tag) copurified with Strep-tagged SlPAD4, SlSAG101a and SlSAG101b, but not SlEDS1. We conclude that SlEDS1 engages in heterocomplexes containing SlPAD4, SlSAG101a or SlSAG101b, but does not form homodimers. Hence, considering the high similarity of NbEDS1 and SlEDS1 family proteins, three different heterocomplexes with NbEDS1 are most likely also established by the orthologous NbPAD4, NbSAG101a and NbSAG101b proteins.
Identification of EDS1 Complexes Functioning in XopQ Recognition
Gene expression analyses and interaction studies suggested that two different EDS1-SAG101 complexes exist in N. benthamiana. Furthermore, we could not exclude functionality of the SAG101a2 and SAG101b2 isoforms we considered pseudogenes. To reveal a potential role of SAG101 isoforms in TNL-mediated immunity, all four SAG101 genes were targeted for mutagenesis by SpCas9 (Supplemental Figure 4). A respective genome editing construct was transformed into the previously generated Nbpad4-1 mutant line (Ordon et al., 2017), and primary transformants (T0) were tested for recognition of XopQ. Plants were inoculated with a P. fluorescens derivative engineered for Type III system-dependent translocation (Thomas et al., 2009) and expressing the effector XopQ fused to a secretion signal of the P. syringae effector AvrRpt2. This Pfo xopQ strain induces a strong EDS1-dependent HR in wild-type N. benthamiana (Gantner et al., 2018) and the Nbpad4-1 mutant line (Figure 3).
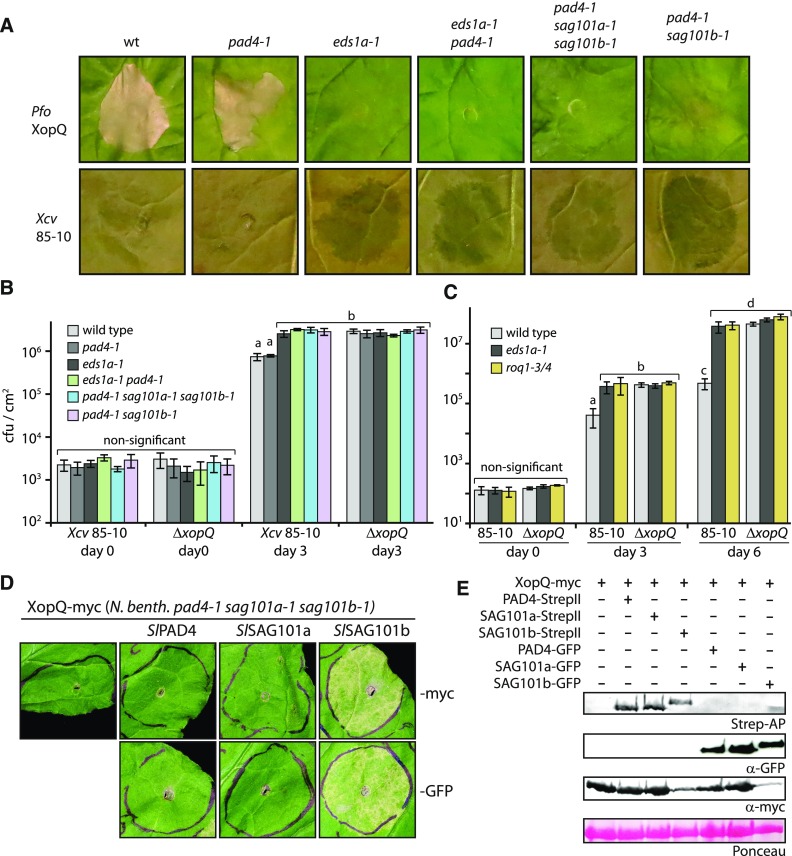
Figure 3.
Immune Responses of N. benthamiana Mutant Lines Deficient in EDS1 Family Genes or Roq1
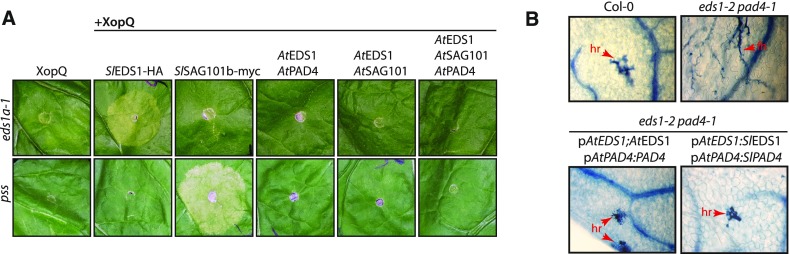
(A) Recognition of XopQ in different mutant lines. Indicated N. benthamiana lines were challenged with XopQ-translocating P. fluorescens bacteria (top; infiltrated at OD600 = 0.2) or Xcv strain 85-10 bacteria (bottom; infiltrated at OD600 = 0.4). Phenotypes were documented at 4 dpi. Similar results were obtained in three independent experiments, and multiple plants of the indicated genotypes were infiltrated in each experiment.
(B) Bacterial growth of Xcv bacteria on mutant lines. Indicated lines were infected with Xcv strain 85-10 or a corresponding mutant strain lacking XopQ (ΔxopQ). Means and sd of four biological replicates are shown. Letters indicate statistically significant differences as determined by one-way ANOVA and Fisher LSD post hoc test (P < 0.01).
(C) Bacterial growth in eds1 and roq1 mutant lines. As in (B), but means and SD of eight biological replicates are shown for days 3 and 6. The roq1 mutant line was a T1 line segregating for two disruptive alleles at the Roq1 locus (Supplemental Fig. 4F).
(D) Reconstitution of XopQ detection in the pss triple mutant line. By agroinfiltration, XopQ was expressed alone or in combination with PAD4, SAG101a, or SAG101b (from tomato and fused to a StrepII and 4 × c-myc tag or GFP). Phenotypes were documented 5 dpi.
(E) Immunodetection of fusion proteins expressed in (D). Samples were taken 3 dpi from a second infiltration on the same leaf shown in (D). Ponceau staining of the membrane is shown as loading control.
One T0 plant failed to initiate the HR in response to XopQ and thus phenocopied the Nbeds1 line. Sequencing revealed that the assumed pseudogenes SAG101a2 and SAG101b2 did not contain any mutations in this line and thus can be dismissed. Hereafter, only SAG101a1 and SAG101b1 will be considered, and they will be referred to as SAG101a and SAG101b for simplicity. The primary line nonresponsive to XopQ was homozygous for a sag101a-1 mutation, and heterozygous for two different sag101b alleles (Supplemental Figure 4). From the resulting T1 plants, a line lacking the genome editing transgene and homozygous for the sag101b-1 allele (i.e., a pad4-1 sag101a-1 sag101b-1 triple mutant; pss) was selected. An additional double mutant line containing the pad4-1 and sag101b-1 mutant alleles (pSs) was isolated from a cross (pss x wild type). We failed to isolate a sag101b-1 single mutant line from the same cross. Allele identifiers of N. benthamiana lines will be omitted hereafter.
N. benthamiana mutant lines were tested alongside control plants for recognition of XopQ by inoculation with Pfo xopQ and Xcv bacteria (Figure 3A). Both bacterial strains induced the HR in leaves of wild-type and pad4 plants. The remaining mutant lines (eds1 pad4 (ep), pss, pSs) failed to initiate the HR and were indistinguishable from eds1 mutant plants (Figure 3A). In addition, in planta bacterial titers of Xcv and Xcv ΔxopQ bacteria were determined for quantitative analysis of immune responses in different N. benthamiana mutant lines (Figure 3B). The growth of Xcv, but not ΔxopQ bacteria, was restricted in wild-type and pad4 plants. Xcv replication in leaves of eds1 and any of the pad4 sag101 mutant lines was similar, and not different from that of ΔxopQ bacteria on wild-type plants (Figure 3B). Therefore, the SAG101a isoform present in the pad4 sag101b (pSs) double mutant does not contribute to immunity, and loss of PAD4 and SAG101b phenocopies eds1 mutants.
Notably, none of the mutant lines showed an enhanced susceptibility or basal resistance phenotype: bacterial titers of ΔxopQ (in any plant genotype) and Xcv in eds1 or pad4 sag101 mutant plants were identical. We also compared bacterial growth in eds1 and roq1 mutant plants (Supplemental Figure 4) up to 6 d (Figure 3C). Identical titers for Xcv were observed in eds1 and roq1 plants, and were similar to bacterial growth of ΔxopQ bacteria. This result further corroborates that N. benthamiana plants deficient in EDS1 complexes lack a basal resistance phenotype, and that XopQ is indeed the only Xcv effector inducing EDS1-dependent defenses (Adlung et al., 2016).
Full susceptibility of Nbpad4 sag101b mutant lines to the Xcv wild-type strain suggests that EDS1-PAD4 and EDS1-SAG101b complexes might function redundantly in immune signaling. Alternatively, only EDS1-SAG101b might have immune functions in N. benthamiana. To discriminate between these scenarios, XopQ and tomato SlPAD4 or SlSAG101 proteins were transiently expressed in the pss background. Coexpression of XopQ with SlSAG101b, but not SlPAD4 or SlSAG101a, restored HR induction (Figure 3D). All proteins were detected on immunoblots using two different epitope tags (Figure 3E). The same complementation assay was performed expressing untagged proteins with identical results. We also compared NbEDS1 and NbSAG101b with their respective tomato orthologs for restoration of XopQ recognition (Supplemental Figure 5). EDS1 or SAG101b orthologs from the two species were equally efficient for restoration of the XopQ-induced HR in respective mutant backgrounds (Supplemental Figures 5A and 5B), and SlPAD4, SlSAG101a and SlSAG101b engaged into complexes with NbEDS1 (Supplemental Figure 5C). We concluded that tomato EDS1 family proteins can functionally replace N. benthamiana orthologs, as anticipated, and decided to use tomato proteins for further functional characterization.
Taking together the observations (1) that pad4 mutant N. benthamiana lines are not impaired in XopQ-induced resistance (Figures 3A and 3B) and (2) that only expression of SAG101b can restore XopQ recognition in the pss background (Figure 3D), these results suggest that an EDS1-SAG101b complex is necessary and sufficient for resistance signaling downstream of XopQ in N. benthamiana. Despite its upregulation during infection (Figure 1B), PAD4 appears to not contribute to Roq1-mediated immune signaling in N. benthamiana.
EDS1-SAG101b Functions in Diverse TNL-Mediated Responses
The finding that EDS1 and SAG101b are required for XopQ-induced resistance responses in N. benthamiana was surprising because major resistance-signaling functions reside in EDS1-PAD4 in Arabidopsis (Feys et al., 2005; Wagner et al., 2013; Cui et al., 2017, 2018). We tested additional inducers of presumed EDS1-dependent defense responses in our set of mutant lines to analyze whether EDS1-SAG101b are generally required for immune signaling in N. benthamiana or whether this is specific for the TNL Roq1 recognizing XopQ (Schultink et al., 2017). Expression of a TIR domain fragment of the TNL DM2h (Stuttmann et al., 2016), a variant of a TIR fragment of RPS4 (RPS41-234_E111K; Swiderski et al., 2009; Williams et al., 2014), and coexpression of the Tobacco mosaic virus helicase protein p50 together with the tobacco TNL receptor N (Burch-Smith et al., 2007) induced HR-like cell death on wild-type but not eds1 or pss mutant plants (Figure 4). As with XopQ, cell-death induction could be restored in pss plants by coexpression of SlSAG101b but not SlSAG101a or SlPAD4 (Figure 4A). These results indicate that EDS1-SAG101b are generally required for TNL-induced defenses in N. benthamiana, whereas PAD4 and SAG101a cannot functionally replace SAG101b, even when expressed under control of the strong 35S promoter in transient assays.
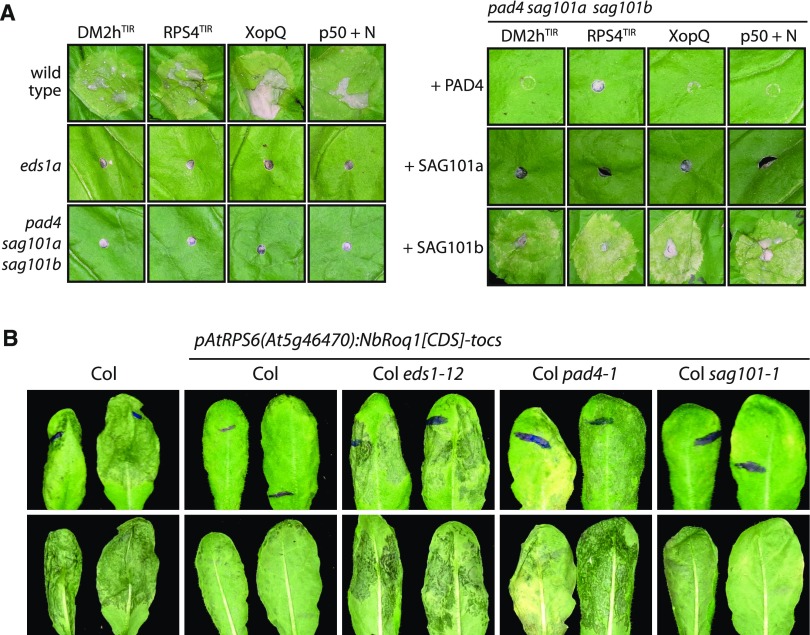
Figure 4.
Genetic Dependencies of TNL-Type Immune Receptors in N. benthamiana and Arabidopsis.
(A) EDS1-dependent cell-death induction requires SAG101b in N. benthamiana. Inducers of presumably EDS1-dependent cell death (DM2hTIR – DM2h(1-279); RPS4TIR – RPS4(1-234)_E111K [Swiderski et al., 2009]; XopQ – XopQ-myc; p50 + N – p50-Cerulean + N-Citrine [Burch-Smith et al., 2007]) were expressed in different N. benthamiana lines, as indicated (left), or coexpressed with PAD4, SAG101a or SAG101b (from tomato and fused to a 4xmyc-TwinStrep tag) in the pss mutant line (right). Phenotypes were documented 5 dpi.
(B) Functionality and genetic dependency of Roq1 in Arabidopsis. A T-DNA construct coding for Roq1 under control of an RPS6 promoter fragment and an ocs terminator was transformed into the indicated Arabidopsis lines. Four-week-old control and T1 plants were infected with Pst DC3000 bacteria (syringe infiltration, OD600 = 0.001). Symptom development was documented 3 dpi. At least eight independent T1 plants were tested for each genotype per replicate, and the experiment was conducted three times with similar results.
In a complementary approach, the Roq1 gene was transferred into Arabidopsis wild-type and eds1, sag101, and pad4 mutant lines. Roq1 was expressed under control of the Arabidopsis RPS6 promoter and the octopine synthase terminator (ocs, Agrobacterium tumefaciens). RPS6 encodes a TNL receptor recognizing HopA1 from Pseudomonas syringae pv syringae strain 61 (Kim et al., 2009), and its promoter was chosen for Roq1 expression to potentially avoid dominant negative effects often arising from overexpression of immune receptors (e.g., Wirthmueller et al., 2007). T1 transgenic seeds from transformation of the Roq1 expression constructs into Col-0 wild type and eds1-12, pad4-1 and sag101-1 mutant lines were selected by FAST seed coat fluorescence (Shimada et al., 2010), respective plants grown in soil and infected with Pst DC3000 bacteria. Strain DC3000 contains the effector HopQ1 homologous to XopQ, which is also recognized by Roq1 (Adlung et al., 2016; Schultink et al., 2017; Zembek et al., 2018).
If Roq1 can function in Arabidopsis, we expected to generate resistance to Pst DC3000, which is an aggressive pathogen in accession Col-0. Indeed, severe tissue collapse was observed in Col-0 plants infected with DC3000 at 3 d post inoculation (dpi), whereas plants containing the Roq1 transgene were mostly asymptomatic (Figure 4B). As a TNL receptor, we expected Roq1 to function in an EDS1-dependent manner in Arabidopsis. In agreement, tissue collapse similar to that of Col-0 plants was observed in eds1-12 pRPS6:Roq1 plants. The pad4-1 transgenics containing the pRPS6:Roq1 transgene behaved similar to eds1-12 transgenics and Col-0, whereas transgenic lines in the sag101 background were as resistant as wild-type plants expressing Roq1 (Figure 4B). Similar results were obtained when Roq1 was expressed under the control of a Ubiquitin 10 promoter fragment. Roq1 is thus functionally dependent on EDS1-SAG101b in N. benthamiana but requires EDS1-PAD4 to mediate resistance in Arabidopsis. We conclude that it is not the TNL receptors but rather the differences within the EDS1 protein family of respective plant species that determine which EDS1 heterocomplexes function in TNL signaling.
EDS1 Complexes Are Not Sufficient for TNL Signaling, but Additional Factors Divergent between Individual Species Are Required
Considering that EDS1-PAD4 and EDS1-SAG101b are required for immune signaling in Arabidopsis and N. benthamiana, respectively, these complexes might have identical functions, albeit different evolutionary origin. Alternatively, functional recruitment to immune signaling might occur by different mechanisms in these species. We sought to analyze these aspects by transferring EDS1 family genes from Arabidopsis into N. benthamiana and vice versa. We first attempted to restore XopQ-induced cell death in eds1 or pss mutant N. benthamiana plants (Figure 5; Supplemental Figure 6). Arabidopsis EDS1 family genes were expressed, in different combinations and with or without an epitope tag, from a single T-DNA, and XopQ was coexpressed (Supplemental Figure 6A). As controls, SlEDS1-HA and SlSAG101b-myc were coexpressed with XopQ. Arabidopsis and tomato orthologs were expressed to similar levels (Supplemental Fig. 6B). However, XopQ-induced cell death was efficiently restored by coexpression of SlEDS1 and SlSAG101b (in eds1 and pss mutant plants, respectively), but not by coexpression of the Arabidopsis EDS1 family proteins, in any given combination (Figure 5A). Thus, Arabidopsis EDS1 complexes fail to function in Roq1 signaling in N. benthamiana.
Figure 5.
Cross-Species Transfer of EDS1 Family Genes.
(A) Arabidopsis EDS1-PAD4-SAG101 proteins cannot functionally replace EDS1-SAG101b in N. benthamiana. Indicated proteins were expressed (by agroinfiltration) either in eds1 or pss mutant lines, and phenotypes were documented 7 dpi. Arabidopsis proteins were expressed with or without an epitope tag, and images originate from untagged proteins (Supplemental Figure 6 shows details on T-DNA constructs and protein detection).
(B) Tomato EDS1-PAD4 can function in TNL signaling in Arabidopsis. Col eds1-2 pad4-1 double mutant was transformed with constructs for expression of EDS1 and PAD4, either from Arabidopsis or tomato and with or without an epitope tag (Supplemental Figure 6C) and under control of Arabidopsis promoter fragments. Segregating T2 populations were selected with BASTA, and 3-week-old plants infected with H. arabidopsidis isolate Cala2. True leaves were used for Trypan Blue staining 7 dpi. At least four independent T2 populations were tested for each construct with similar results. Lines expressing untagged proteins were used for infection assays. Lines expressing epitope-tagged proteins were used for immunodetection (Supplemental Figure 6D). fh, free hyphae; hr, hypersensitive response.
Reciprocally, the Arabidopsis eds1-2 pad4-1 double mutant line was transformed with constructs encoding EDS1 and PAD4 from Arabidopsis or tomato and under control of the corresponding native promoter elements from Arabidopsis (Supplemental Figure 6C). For each transformation, several independent T2 populations were tested for complementation of the eds1-2 pad4-1 immunity defects by infection with Hpa isolate Cala2. Cala2 is recognized via the TNL RPP2 in Col-0 (Sinapidou et al., 2004) but is highly virulent on eds1 pad4 plants (Figure 5B). Transformants expressing EDS1-PAD4 from tomato were as resistant to Hpa Cala2 as transformants expressing the Arabidopsis homologs and indistinguishable from wild-type Col-0 (Figure 5B). In simultaneously generated transgenics expressing epitope-tagged variants, EDS1 and PAD4 from Arabidopsis and tomato accumulated to similar levels as assessed by immunodetection (Supplemental Figure 6D). An additional set of transgenic plants was generated in the eds1-2 pad4-1 sag101-1 triple mutant background that expressed SlEDS1 and different combinations of SlPAD4 and/or SlSAG101 isoforms (Supplemental Figure 6E). Transgenic plants expressing SlEDS1 together with SlPAD4 and SlSAG101 isoforms were resistant to Hpa isolate Cala2, although HR-associated cell death appeared less confined than in Col-0 or control plants expressing Arabidopsis EDS1-PAD4 (Supplemental Figure 6E). By contrast, transgenics expressing SlEDS1 and a SlSAG101 isoform, but not SlPAD4, were susceptible. Again, all proteins were detected in transgenic plants expressing epitope-tagged variants (Supplemental Figure 6F).
Hence, SlEDS1-SlSAG101b are sufficient for all tested immune responses in N. benthamiana but fail to function in Arabidopsis. By contrast, SlEDS1-SlPAD4 can function in RPP2-mediated resistance in Arabidopsis but not in any tested TNL-mediated response in N. benthamiana. Thus, proteins of the PAD4 phylogenetic clade appear to operate, together with EDS1, in TNL signaling in Arabidopsis, and these functions are executed by EDS1-SAG101 in N. benthamiana. Taken together with the observation that AtEDS1-AtSAG101-AtPAD4 fail to mediate TNL signaling N. benthamiana, we conclude that EDS1 complexes do not form a complete functional module in TNL signaling by themselves. We propose that additional factors, divergent between species as a result of coevolution with EDS1 complexes, are required by these heterocomplexes to mediate immune responses.
Rapid Analyses of EDS1 Complexes in the N. benthamiana System
One rationale for genetic dissection of the EDS1 gene family in N. benthamiana was the establishment of an experimental system that allows rapid analysis of EDS1 complexes and their immune competence. Previous experiments showed that XopQ-induced cell death can be restored in the eds1 and pss mutant lines by Agrobacterium-mediated coexpression of EDS1 and SAG101b, respectively (Figure 4A; Adlung et al., 2016; Qi et al., 2018). The HR-like cell death provoked by XopQ expression is relatively mild in wild-type N. benthamiana plants and further delayed and dampened in transient complementation assays but highly reproducible under our conditions.
A crystal structure of the Arabidopsis EDS1-SAG101 complex and an experimentally validated homology model of the EDS1-PAD4 complex were reported (Wagner et al., 2013). Because Arabidopsis EDS1 complexes were not functional in N. benthamiana, homology models of EDS1-based heterodimers from tomato were generated (Figure 6). A structure similar to that of Arabidopsis EDS1-SAG101 was predicted for the immune-competent tomato EDS1-SAG101b complex, and most surface-exposed, conserved residues mapped to the heterocomplex interface (Figures 6A and 6B).
Figure 6.
EDS1-SAG101b Heterocomplexes Are the Functional Modules in N. benthamiana TNL Signaling.
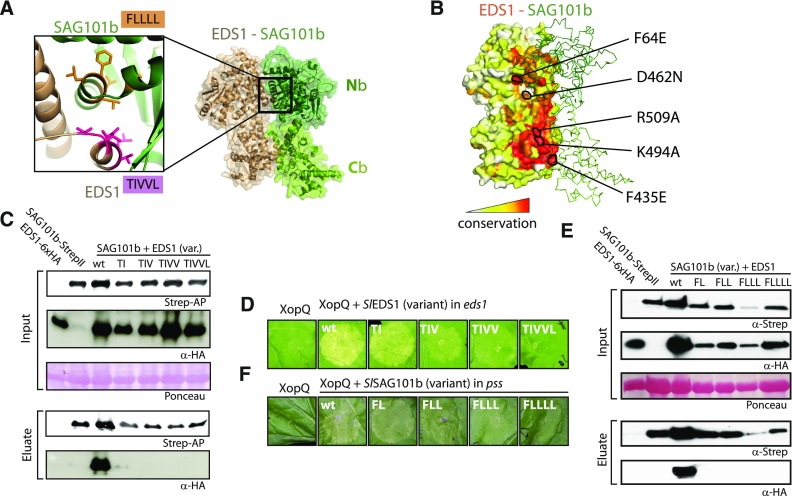
(A) Homology model of the tomato EDS1-SAG101b complex used for transient reconstitution of TNL-signaling in mutant N. benthamiana tissues. The N-terminal (Nb) and C-terminal (Cb) domains of SAG101b are depicted in dark and light green, respectively. The inset shows the symmetrically arranged helices of EDS1 and SAG101b forming the N-terminal interaction interface. Amino acids targeted by mutagenesis are shown as sticks and are highlighted in pink (EDS1) and orange (SAG101b), respectively.
(B) Conservation of surface-exposed amino acids in SlEDS1. SlSAG101b is shown in ribbon presentation (green). EDS1 residues functionally interrogated by mutagenesis are marked.
(C) Interaction of EDS1 variants with SAG101b. Indicated proteins were (co)expressed in N. benthamiana by agroinfiltration and tissues used for StrepII purification at 3 dpi.
(D) Functionality of EDS1 variants affected in heterocomplex formation. Indicated variants (as in [C], with C-terminal 6xHA) were coexpressed with XopQ-myc in eds1 mutant plants, and plant reactions were documented 7 dpi.
(E) Interaction of SAG101b variants with EDS1. As in [C], but SAG101b-StrepII variants were coexpressed with EDS1.
(F) Functionality of SAG101b variants affected in heterocomplex formation. SAG101b-StrepII variants were coexpressed with XopQ-myc in pss mutant plants, and plant reactions were documented 7 dpi.
To validate the structural models and the N. benthamiana system, we decided to disrupt EDS1-SAG101b complex formation by mutagenesis of key amino acids within the N-terminal interaction interface (Wagner et al., 2013). The N-terminal interface is formed mainly by hydrophobic interactions between a protruding helix of EDS1 accommodated in a corresponding pocket on SAG101b (Figure 6A). Residues within the EDS1 helix were sequentially mutated: T264F and I268E (TI), followed by V265E (TIV), V269E (TIVV) and L261E (TIVVL). All variants accumulated to comparable levels in planta, and TIV or higher order mutants did not copurify in detectable amounts with StrepII-tagged SAG101b (Figure 6C). When tested by yeast two hybrid, interaction of the EDS1 variants with SAG101b and also PAD4 and SAG101a gradually declined and was still detectable for the TIVV quadruple mutant variant (Supplemental Figure 7). In accordance with complex formation being progressively impaired, SlEDS1 variants also lost their activity for restoring XopQ-induced cell death when coexpressed in eds1 mutant N. benthamiana plants (Figure 6D). Only the quintuple TIVVL variant was completely nonfunctional. Similarly, mutations were serially introduced into SAG101b: F17S and L22S (FL), L13S (FLL), L16S (FLLL) and L18S (FLLLL). In copurification assays, interaction with EDS1 was detectable only for wild-type SAG101b, and protein accumulation of SAG101 variants was mildly affected (Figure 6E). SAG101b variants were tested for functionality by appearance of HR-like cell death upon coexpression with XopQ in pss mutant plants (Figure 6F). Cell death was reduced for SAG101b-FLL and abolished for the quadruple and quintuple mutant variants. These data suggest that heterocomplex formation is required for immune functions of EDS1 and SAG101b and thus support previous findings from analysis of EDS1 in the Arabidopsis system (Wagner et al., 2013).
Next, we set out to identify additional functionally relevant features of EDS1-SAG101b complexes (a summary of tested variants is shown in Supplemental Table 1). We first focused on several positively charged residues lining an assumed cavity on the heterodimer surface (Wagner et al., 2013) and recently reported as required for immune signaling in Arabidopsis (Bhandari et al., 2019). The residues within SlEDS1 (R509, K494), homologous to those reported in Arabidopsis (R493, K478), were targeted by mutagenesis, and respective variants tested for functionality (Supplemental Figure 8). All variants restored XopQ-induced HR-like cell death as efficiently as wild type SlEDS1. We propose that functional relevance of the positively charged residues might be masked by overexpression in the N. benthamiana system or might not be conserved across different species.
We also introduced F64E, F435E, and D462N exchanges into SlEDS1 (Figure 7). F64 is a single, conserved residue exposed on the N-terminal lipase-like domain of EDS1 and framing the assumed cavity (Figure 6B). F435 is exposed in the monomeric EDS1 but fully buried by the association with SAG101 and was used to probe the importance of the C-terminal interaction surface. D462 connects the N- and C-terminal domains and might mediate crosstalk at the domain interface. All variants retained interaction with SAG101b, as tested by copurification, although stability of D462N was impaired (Figure 7A). When coexpressed with XopQ in eds1 plants, immune activities were reduced for F64E and D462N variants and fully abolished for F435E. We also tested F435D and F435A variants. While F435D failed to restore immune capacity in eds1 plants, F435A was functional (Supplemental Figure 8). We assume that disruption of the apolar patch at the C-terminal interface and introduction of a charged residue in F435E/D dislocates the EP domains within the EDS1-SAG101b heterocomplex against each other without disturbing overall complex assembly mainly driven by the N-terminal interaction surface. This finding supports a role for the heterodimeric EP domain surface in TNL signaling.
Figure 7.
Identification of Nonfunctional EDS1-SAG101b Complex Variants.
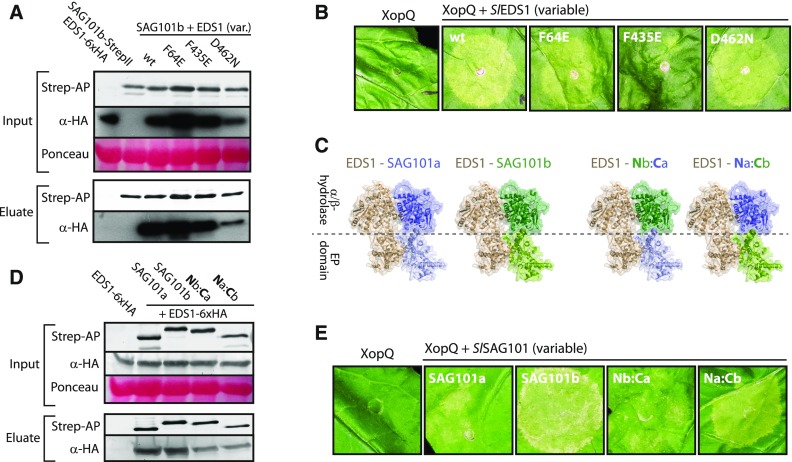
(A) Interaction of EDS1 variants with SAG101b. Indicated proteins were (co)expressed in N. benthamiana by agroinfiltration. Tissues were used 3 dpi for StrepII purification.
(B) Immune activities of EDS1 variants. Indicated EDS1 variants (with C-terminal 6xHA tag) were transiently coexpressed with XopQ-myc in eds1 mutant plants by agroinfiltration. Plant reactions were documented 7 dpi.
(C) Structural basis for SlSAG101a-SlSAG101b chimeric proteins. EDS1 and SAG101 both contain an N-terminal hydrolase-like and a C-terminal EP domain. In the heterodimer, an N-terminal interface is formed by the hydrolase-like domains, and a C-terminal interface is formed by the EP domains. For chimeras, the N terminus of SAG101b (aa 1-322) or SAG101a (aa 1-339) was fused with the C terminus of SAG101a (aa 340-581) or SAG101b (aa 323-567), respectively.
(D) Heterocomplex formation by SAG101 chimeric proteins. SAG101 chimeras and native SAG101 isoforms (with a C-terminal StrepII tag) were coexpressed with EDS1-6xHA by agroinfiltration. Tissues were used 3 dpi for StrepII-purification.
(E) Functionality of SAG101 chimeric proteins. Indicated proteins were expressed together with XopQ-myc in pss mutant plants by agroinfiltration. Plant reactions were documented 7 dpi.
We constructed chimeric proteins from nonfunctional SAG101a and functional SAG101b to further analyze this aspect (Figure 7C). Although differences between SAG101a and SAG101b isoforms remain unclear, we hypothesized that if the EP domain surface is crucial for immune functions, only the chimeric protein carrying the C terminus of SAG101b might be functional, whereas both chimeras should engage in heterocomplexes with EDS1. SAG101a/b chimeras accumulated to levels comparable with the native isoforms when expressed by agroinfiltration and also formed complexes with EDS1 (Figure 7D). Chimeras and native SAG101 isoforms were coexpressed with XopQ in pss mutant plants to test for functionality. SAG101a and the Nb-Ca chimeric protein did not show any activity. In contrast, the Na-Cb chimeric protein was able to restore XopQ-induced cell death, albeit to a lesser extent than SAG101b (Figure 7E). These results suggest that the main differences discriminating SAG101a and SAG101b and their immune competence reside in the C-terminal EP domain and further support the EP domain surface as crucial for the function of EDS1 heterodimers in TNL signaling (Wagner et al., 2013; Bhandari et al., 2019). As the most probable mode of action of EDS1 complexes in immune signaling, we propose that further interaction partners might be recruited via the heterodimeric EP domain surface.
DISCUSSION
EDS1 is essential for signaling downstream of TNL-type immune receptors and forms mutually exclusive heterodimeric complexes with PAD4 and SAG101 (Hu et al., 2005; Rietz et al., 2011; Wagner et al., 2013; Schultink et al., 2017). Based on analyses in Arabidopsis, immune functions were so far mainly accounted to EDS1-PAD4 (Feys et al., 2005; Wagner et al., 2013; Cui et al., 2017, 2018). In this study, we show that an EDS1-SAG101 complex is necessary and sufficient for all tested TNL-dependent immune responses in N. benthamiana, whereas PAD4 does not appear to contribute to immunity (Figures 3 and 4). A role for NbSAG101 in effector-triggered immunity was also reported in previous virus-induced gene-silencing-based analyses (Pombo et al., 2014).
Several lines of evidence suggest that EDS1-SAG101 complexes, rather than EDS1-PAD4, might also fulfill immune signaling functions in other species containing TNL-type immune receptors outside the Brassicaceae family. First, there is strict co-occurrence of SAG101 and TNL-type immune receptors in genomes, strongly suggesting a functional link (Figure 1A; Wagner et al., 2013). Second, Brassicaceae PAD4 orthologs lack an insertion within the lipase-like domain, which is present in non-Brassicaceae PAD4 orthologs and EDS1 orthologs but absent in SAG101 orthologs (Wagner et al., 2013). Therefore, PAD4 might have evolved by a unique path or mechanism in Brassicaceae, supporting the hypothesis that PAD4 immune-signaling functions in this family might represent a notable exception. It is interesting to note that NbPAD4 is induced concomitantly with NbEDS1 under infection conditions, although it does not contribute to immune signaling in our experiments. In contrast, only minor transcript changes were detected for the signaling-competent NbSAG101b (Figure 1B). A possible explanation might be that upregulation of PAD4 serves to inactivate EDS1-SAG101b–mediated immune signaling, by titration of SAG101b from EDS1. Competition of interactors for the EDS1 moiety for complex formation is supported by enhanced XopQ-induced cell death observed in reconstitution assays in pss mutant plants in comparison to eds1 mutant plants, but overexpression of SlEDS1 together with SlSAG101b without XopQ was not sufficient for cell-death induction. Future reverse genetic studies are needed to clarify which EDS1-containing heterocomplexes function predominantly in TNL signaling. Furthermore, it remains to be determined which functions are exerted by EDS1 complexes that are not involved in immune signaling or EDS1 complexes (with PAD4) in organisms lacking TNLs.
Genome analysis revealed a duplication of SAG101 in most Solanaceae (Figure 1B). Absence of the SAG101a isoform in C. annuum might indicate reduced selective forces toward its preservation. However, although both the NbSAG101a2 and NbSAG101b2 genes showed signs of pseudogenization in allotetraploid N. benthamiana, this was not observed for SAG101a in any of the remaining analyzed Solanaceae genomes. In addition, differential subcellular localization patterns were detected for SlSAG101 isoforms (Figure 2; Supplemental Figure 3). This finding argues against SAG101a representing merely a duplicated gene but rather supports distinct functions of individual isoforms. An additional EDS1-SAG101 complex might provide fine-tuning of EDS1 activities in many Solanaceae, but no evidence shows a contribution to TNL-mediated immune responses. It is conceivable that NbSAG101a may participate in signaling downstream of yet uncharacterized N. benthamiana TNLs, as some Arabidopsis TNLs were also reported to depend on EDS1-SAG101 rather than EDS1-PAD4 (Zhu et al., 2011; Xu et al., 2015). We did not analyze functional relevance of EDS1 subcellular distribution in N. benthamiana, but nuclear-localized EDS1 was required and sufficient for several tested immune responses in Arabidopsis (García et al., 2010; Stuttmann et al., 2016). It is worth noting that both AtEDS1-AtSAG101 and SlEDS1-SlSAG101a are confined to nuclei and have only minor or no functions in immunity, whereas complexes required for TNL signaling (AtEDS1-AtPAD4, SlEDS1-SlSAG101b) also distributed to the cytoplasm.
In interactions of Arabidopsis with P. syringae bacteria or filamentous pathogens Hpa and Golovinomyces orontii, eds1 mutant plants are significantly impaired in basal resistance (Falk et al., 1999; Lipka et al., 2005; Rietz et al., 2011; Schön et al., 2013). Similarly, an eds1mutant tomato line (sun1-1) was more susceptible to several virulent pathogens, including Xcv (Hu et al., 2005). By contrast, we did not observe any basal resistance phenotype when comparing in planta growth of avirulent (Xcv 85-10) and virulent (Xcv ΔxopQ) bacteria in wild-type, eds1, pss, and roq1 N. benthamiana lines (Figure 3; Adlung et al., 2016; Schultink et al., 2017; Qi et al., 2018). Moreover, basal resistance to several viral or other bacterial isolates was not impaired in EDS1-silenced N. benthamiana plants (Peart et al., 2002). These results do not support a general role of EDS1 complexes in basal immunity. We therefore assume that any contribution of EDS1 to basal resistance results from loss of TNL-mediated ETI and thus does not represent an independent function. This is in line with expression of basal resistance by the term "PTI/MTI + weak ETI − effector-triggered susceptibility" (Jones and Dangl, 2006), and the TNL-mediated component of "weak ETI" being abolished in eds1 lines. One explanation why TNL-mediated ETI may have less importance in N. benthamiana is the the relatively small complement of TNLs in this species. Automated R gene identification previously identified only 17 TIR domain-encoding genes in N. benthamiana (Hofberger et al., 2014). Although incomplete N. benthamiana genomes and flawed annotations likely lead to underestimation of TNL diversity, TNLs represent the predominant class of NLRs in Arabidopsis with ∼70 to 100 TNLs per genome (Meyers et al., 2003; Peele et al., 2014; Van de Weyer et al., 2019). By contrast, CNLs appear to be the predominant NLR class present in N. benthamiana.
The N. benthamiana system features the key advantage of rapid and robust transient protein expression by agroinfiltration. This was exploited to design transient complementation assays for EDS1-SAG101b functional analyses based on induction of HR-like cell death by XopQ (Adlung et al., 2016; Schultink et al., 2017; Qi et al., 2018). We confirmed significance of results obtained in this highly simplified system by disrupting EDS1-SAG101b complex formation (Figure 6). We mutagenized key residues within the interface and showed that higher order mutants containing multiple amino acid exchanges fail to function in immune signaling. Notably, several EDS1 and SAG101 variants for which interaction was undetectable by copurification (e.g., EDS1-TIVV, SAG101-FLL) still functioned, at least partially, in cell-death induction, indicating that low-level complex formation is sufficient for immune responses. Similarly, a previously described Arabidopsis PAD4-MLF variant (Wagner et al., 2013) deficient in complex formation complemented immune deficiency of a pad4-1 sag101-3 double mutant line when tested (J. Stuttmann and J. Parker, unpublished data). Consequently, extension of the interface analysis from EDS1 to its interaction partner SAG101 lends important support to the previous notion that complex formation is a prerequisite for immune signaling.
Beyond the N-terminal interaction surface, five highly conserved positions distributed all along the surface of EDS1 in the vicinity of the interface with SAG101b were tested for functional significance in the N. benthamiana system (Figure 7). R509 and K494 are located in a large conserved cavity formed in EDS1 heterodimeric assemblies, and homologous positions are crucial for full EDS1 immune activities in Arabidopsis (Bhandari et al., 2019). However, we did not observe reduced immune functions for corresponding SlEDS1 variants. D462 was targeted to probe potential domain crosstalk within EDS1. The SlEDS1D462N variant was considerably less competent in restoration of XopQ-induced HR, but this might also be explained by reduced stability (Figures 7A and 7B). In contrast, F64E and F435E/D, which delimit upper and lower boundaries of the assumed cavity, were not impaired in stability but had reduced or abolished immune activities, respectively. F64 might act as conserved gatekeeper, whereas perturbation of the C-terminal interface by F435E/D is expected to have more profound effects on overall topology of the EP domain assembly (Figure 7). The F435 variants suggest a crucial role of the C-terminal EP domains for immune signaling (Wagner et al., 2013; Bhandari et al., 2019), which is further supported by partial restoration of immune functions in SAG101a by grafting of the SAG101b C terminus in SAG101Na-Cb protein chimeras (Figure 7). Nonetheless, functional impairment of SlEDS1F64E shows that also the N-terminal α/β-hydrolase domain contributes to immune signaling, and roles are thus not limited to alignment of C-terminal EP domains in the heterodimer.
A most plausible hypothesis is that the large surface on SlEDS1 delimited by F435 and F64 is required for interactor recruitment to mediate immune signaling. Recruitment of protein interactors is also supported by results of our cross-species transfer of Roq1 and EDS1 family genes. In line with previous reports showing that even immune receptors from evolutionarily distant species maintain functionality when introduced into new plant lineages (Maekawa et al., 2012), transgenic expression of NbRoq1 in Arabidopsis conferred resistance to Pst DC3000 bacteria, presumably through recognition of HopQ (Figure 4B). In contrast, AtEDS1-AtPAD4-AtSAG101 failed to restore immune signaling in N. benthamiana mutant plants (Figure 5A). Similarly, SlEDS1-SlPAD4 can fulfill immune functions in Arabidopsis but not N. benthamiana, whereas the opposite was observed for SlEDS1-SlSAG101b (Figure 5B; Supplemental Figure 6). Consequently, NLR functions are generally conserved, but EDS1 complexes functionally diverged in different species, which suggests coevolution of signaling-competent heterodimeric assemblies with additional factors, most likely protein interactors. One expectation would be that mutant lines deficient in interactors required for EDS1 immune signaling are impaired in TNL-mediated resistance responses. Although a number of proteins were reported to interact with EDS1 complexes (Arabidopsis Interactome Mapping Consortium., 2011; Bhattacharjee et al., 2011; Heidrich et al., 2011; Kim et al., 2012; Cui et al., 2018), mutant lines deficient in respective genes are not generally TNL signaling deficient.
Importantly, the atypical CNL receptor NRG1 was recently identified as a key component required for signaling by TNL receptors (Brendolise et al., 2018; Qi et al., 2018; Castel et al., 2019; Wu et al., 2019). N. benthamiana nrg1 mutant plants still retain some competence to detect XopQ (via Roq1), and it was hypothesized that residual TNL signaling in these plants might be mediated by the Activated Disease Resistance1 (ADR1) class of helper CNLs (Bonardi et al., 2011; Dong et al., 2016; Schultink et al., 2017; Qi et al., 2018). Indeed, differential requirement of TNLs for helper CNLs of the NRG1 and ADR1 classes and partial redundancy of helper functions were reported in Arabidopsis: NRG1 proteins are critical for function of most TNLs, but others require helpers of the ADR1 class or can signal via both pathways (Dong et al., 2016; Castel et al., 2019; Wu et al., 2019). CNLs of the NRG1 family are limited to those genomes containing TNLs (Collier et al., 2011), as also observed for SAG101, and physical association of NbNRG1 with NbEDS1 was reported (Qi et al., 2018). Thus, proteins of the NRG1 class of helper CNLs may represent plausible candidate interaction partners, which might be recruited by EDS1 complexes to form a functional signaling module. Indeed, Lapin et al (2019) show that coexpression of AtNRG1 together with AtEDS1-AtSAG101 can restore XopQ-induced cell death and Roq1-mediated bacterial growth restriction in N. benthamiana plants deficient in EDS1 family genes, thus strongly supporting this hypothesis (Lapin et al., 2019). However, physical association of NRG1 proteins with EDS1, as reported by Qi et al. (2018), will require further analysis. A similar interaction between AtEDS1 and AtNRG1.1 could be detected by Wu et al. (2019) only when using EDS1 as bait in coimmunoprecipitation assays (co-IPs) but not in reciprocal experiments and might result from stickiness of EDS1 in co-IP experiments (Wu et al., 2019). It should be noted that co-IPs in the report by Qi et al. (2018) also show formation of NbEDS1 homodimers. We could not detect homodimerization of the highly similar SlEDS1 in co-IPs, FRET-based interaction assays (Figure 2; Supplemental Figure 3), or by yeast two hybrid. The mechanisms underlying the functional relationships between TNLs, EDS1 complexes, and helper CNLs thus remain a major question to pursue in future analyses.
METHODS
Plant Material, Growth Conditions, Bacterial Strains and Infection Assays
Nicotiana benthamiana wild-type plants and the published eds1a-1 and pad4-1 single and eds1a-1 pad4-1 double mutant lines were used (Ordon et al., 2017). N. benthamiana plants were cultivated in a greenhouse with a 16-h light period (sunlight and/or IP65 lamps (Philips) equipped with Agro 400 W bulbs (SON-T); 130–150 µE/m2⋅s; switchpoint ∼100 µE/m2⋅s), 60% relative humidity at 24/20°C (day/night). Arabidopsis (Arabidopsis thaliana) wild-type accession Columbia and the previously published eds1-2 pad4-1 double, eds1-2 pad4-1 sag101-1 triple mutant (Feys et al., 2005; Wagner et al., 2013), and eds1-12 single mutant (Ordon et al., 2017) lines were used. Arabidopsis plants were grown under short day conditions at 23/21°C and with 60% relative humidity or in a greenhouse under long day conditions for seed set.
For bacterial growth assays, the Xcv strain 85-10 (Thieme et al., 2005) and the ΔxopQ mutant (Adlung et al., 2016) were syringe-infiltrated at an OD600 = 0.0004, leaf discs were harvested with a cork borer at different time points and disrupted in 10 mM MgCl2 using a bead mill, and bacterial titers were determined by plating dilution series. For each time point and strain, samples were taken from at least four independent leaves and treated as biological replicates. Bacterial growth assays were repeated at least three times with similar results.
For type III secretion system-dependent protein translocation via Pseudomonas fluorescens, a previously described derivative of the “EtHAn” strain (Thomas et al., 2009) containing a plasmid for translocation of XopQ fused to a secretion signal of AvrRpt2, was used (Gantner et al., 2018). Hyaloperonospora arabidopsidis isolate Cala2 was used for infection of Arabidopsis plants, and infections were done as described previously by Stuttmann et al., 2011. True leaves were stained with Trypan Blue 7 dpi, and representative micrographs are shown.
Phylogenetic Analyses
Genomes as indicated in Supplemental Data Set 1 were mined for EDS1 family genes by tBLASTn using tomato (Solanum lycopersicum) proteins as query. Gene models were examined or assigned using fgenesh+ (Solovyev, 2007) and multiple sequence alignments. For the verification of N. benthamiana EDS1 family gene models, RNA-seq data (accession number GSE83618; Legay et al., 2016) was downloaded from the National Center for Biotechnology Information Gene Expression Omnibus website and mapped to the N. benthamiana genome (v1.01; Bombarely et al., 2012) using the CLC genomics workbench (version 7.5.5; Qiagen). Read mappings were inspected manually and supported gene models depicted in Supplemental Figure 1 for NbEDS1, NbPAD4 and NbSAG101a1. For NbSAG101a2 and NbSAG101b2, the Niben101Scf03969g06010 and Niben101Scf09577g01001 locus annotation (encoding for truncated SAG101 proteins), not the fgenesh+-predicted gene models, were supported by inspection of RNA-seq data. The NbSAG101b1 gene model was verified by cloning of the cDNA. For phylogenetic tree construction, sequences as provided in Supplemental Data Set 1 were aligned using the MAFFT G-INS-I algorithm (Katoh and Standley, 2013). A phylogenetic tree was calculated using a procedure derived from the SeaView graphical user interface (Gouy et al., 2010). Site selection was performed using Gblocks (Castresana, 2000) to trim the alignments from highly gapped columns with options −b4 = 5, −b5 = h, −b2 = 18 (50% of number of species +1) for less stringent selection. Then, the tree was computed with the PhyML program (version 3.1; Guindon et al., 2010) using the LG substitution model (with optimized across-site rate variation using four substitution rate categories), empirical amino acid equilibrium frequencies, and an optimized fraction of invariable sites. The tree topology search was performed using the best of NNI (Nearest Neighbor Interchange) and SPR (Subtree Pruning and Regrafting) strategy. Branch support for the final tree was computed with PhyML using 1,000 bootstrap replicates. The final tree was visualized with iTol (Letunic and Bork, 2019). The associated multiple sequence alignment was visualized with Jalview (Waterhouse et al., 2009) and is provided as Supplemental Data Set 2.
Agrobacterium-Mediated Expression, StrepII Purification, and Immunodetection
For transient Agrobacterium-mediated expression of proteins in N. benthamiana (agroinfiltration), plate-grown bacteria were resuspended in Agrobacterium infiltration medium (10 mM MES, pH 5.8, 10 mM MgCl2). Single strains were infiltrated at an OD600 = 0.6. For coexpression, OD600 = 0.4 for each strain was used. All constructs for expression of proteins in N. benthamiana contained the 35S promoter. EDS1, PAD4, SAG101, or variants were coexpressed with XopQ-myc in reconstitution assays, and phenotypes were documented 5 to 7 dpi. Phenotypic assays were conducted at least 4 times, and representative results are shown. For immunodetection of proteins in support of reconstitution assays and for copurification, proteins were expressed without XopQ to avoid interference due to the negative effect of XopQ recognition on Agrobacterium-mediated protein expression (Adlung and Bonas, 2017) but using the same N. benthamiana genetic background as in respective reconstitution experiments. Leaf tissue was ground in liquid nitrogen, powder was resuspended in Laemmli buffer, and proteins were denatured by boiling before SDS-PAGE for immunodetection. For StrepII purifications, 1 g of leaf tissue was ground in liquid nitrogen, and the leaf powder was resuspended in 2.5 mL of extraction buffer (50 mM Tris pH 8, 150 mM NaCl, 5 mm EDTA, 5mM EGTA, 10 mM DTT, 0,1% (v/v) Triton X-100). Suspensions were cleared by centrifugation and supernatants passed through a 0.45 µm syringe filter. 2 mL of cleared extracts were incubated with 120 μL of Strep-Tactin high-capacity matrix (IBA) for 20 min at 4°C on a rotary wheel. The matrix was washed several times with extraction buffer before elution of proteins by boiling with 100 μL of Laemmli buffer. Proteins were resolved by SDS-PAGE and transferred to a nitrocellulose membrane (GE Healthcare). StrepII-tagged proteins were detected using Strep-Tactin alkaline phosphatase conjugate (IBA) or a mouse monoclonal StrepII antibody (Sigma-Aldrich). Further primary antibodies used were α-mCherry (Abcam, ab167453); mouse monoclonal α-GFP and α-c-myc, rat α-HA (all from Roche); and α-FLAG (Sigma-Aldrich). Secondary antibodies were coupled to horseradish peroxidase (GE Healthcare) or alkaline phosphatase (Sigma-Aldrich). Protein gel blots were used for verification of protein expression in support of reconstitution assays, and protein copurification assays were repeated at least three times with similar results.
Plant Transformation and Genome Editing
Arabidopsis plants were transformed as previously described (Logemann et al., 2006). For transformation of N. benthamiana, leaves of greenhouse-grown plants were surface sterilized, cut, and cocultivated with Agrobacteria containing Cas9/sgRNA constructs. Explants were surface-sterilized, and transgenic plants were regenerated. A detailed protocol is provided as an online resource (dx.doi.org/10.17504/protocols.io.sbaeaie). Details on constructs, target sites for editing of N. benthamiana SAG101 isoforms and Roq1, and generated mutant alleles are provided in Supplemental Figure 4. T0 plants were tested phenotypically by challenge inoculation with XopQ-translocating P. fluorescens bacteria and screened by PCR. A transgene-free pad4-1 sag101a-1 sag101b-1 (pss) triple mutant was isolated from a segregating T1 population by PCR screening and crossed to wild type for isolation of the pad4-1 sag101b-1 double mutant line. Homozygous, nontransgenic seed lots were used for experiments. For the roq1 mutant line, a T1 population segregating for two different disruptive alleles (roq1-3 and roq1-4; Supplemental Figure 4) was used for infection assays.
Live-Cell Imaging and Analysis by Förster Resonance Energy Transfer and Acceptor Photobleaching
Images were taken on a LSM780 laser scanning microscope (Zeiss). For imaging of GFP and mCherry, fluorophores were excited with 488 nm and 561 nm laser lines, and emission detected at 493-556 nm and 597-636 nm, respectively. For localization studies, at least three independent experiments were conducted, and multiple images were taken for each replicate and each construct. Similar results were obtained in all experiments. For intensity-based Förster resonance energy transfer (FRET; FRET and acceptor photobleaching), mCherry was bleached using the 561 nm laser at 100% intensity, and GFP fluorescence was measured before and after bleach. Nuclei were selected for measurements. FRET efficiency was calculated by the formula EfD = 1 −FD1/FD2 (EfD, FRET-efficiency donor; FD1, intensity donor before bleach, FD2, intensity donor after bleach) using the FRET module of ZEN software (Zeiss). At least 15 measurements per donor/acceptor combination were done per experiment, and data were reproduced in four independent repetitions.
Molecular Cloning and Yeast Two Hybrid Interaction Assays
Constructs were generated by Golden Gate (Engler et al., 2008) and Gateway (Thermo Fisher Scientific; according to manufacturer’s instructions) cloning. Golden Gate reactions with either BsaI or BpiI were performed using 20 to 40 fmol of each DNA module and cycling between 37°C and 16°C, as described previously by Weber et al., 2011. DNA modules of the MoClo Plant Toolkit, Plant Parts I (Engler et al., 2014), and Plant Parts II (Gantner et al., 2018) collections were used. Novel level 0 modules were generated as described previously, and restriction sites were eliminated by site-directed mutagenesis or overlapping PCR products (Engler et al., 2008, 2014). Details on generated constructs and oligonucleotides used for cloning are provided in Supplemental Tables 2 and 3, respectively. Previously described (Gantner et al., 2018) Golden Gate-compatible or Gateway-converted derivatives of pGADT7 and pGBKT7 (Clontech) were used for yeast two hybrid assays. Respective constructs were transformed in yeast strain PJ69-4a by standard procedures (Gietz and Schiestl, 2007). Plate-selected cotransformants were cultivated in liquid Synthetic Drop-Out media for 48 h, and dilution series were prepared and plated on selective media using a multipipette. Extraction of proteins for immunodetection was performed as described previously by Kushnirov (2000).
Gene Expression Analysis
Tomato RNA sequencing data was accessed using the TomExpress portal (http://tomexpress.toulouse.inra.fr/; Zouine et al., 2017). Data was visualized as a normalized expression heatmap using Spearman representation. Expression values for different conditions were added manually. For gene expression analyses in N. benthamiana, plants were syringe infiltrated with Xcv bacteria at an OD600 = 0.02 in 10 mM MgCl2 or mock infiltrated. RNA was extracted by a standard protocol using TRIzol reagent (ambion; Thermo Fisher Scientific). Briefly, two leaf discs (diameter 9 mm) were frozen in liquid nitrogen, and tissues were disrupted using Zirkonia beads (N039.1; Carl Roth) and a bead mill. RNA was extracted with 1 mL TRIzol; 100 μL of bromochloropropane was added for phase separation; and RNA was precipitated, washed, dried, and resuspended in 40 μL water. The Reverse Transcriptase Core Kit was used for cDNA synthesis, and the Takyon No ROX SYBR 2X MasterMix Blue dTTP qPCR Kit (both Eurogentech) was used for quantitative real-time PCR using a CFX96 detection system (Bio-Rad). The previously described reference genes Protein Phosphatase 2A (PP2A) and Elongation Factor 1-α (EF1α) were used for data normalization (Liu et al., 2012) with similar results, and data from normalization to PP2A is shown. Primers used for quantitative real-time PCR are listed in Supplemental Table 4. All primers had efficiencies of 90% to 105%, as evaluated by dilution series.
Protein Modeling
Structural models of S. lycopersicum EDS1-SAG101a and EDS1-SAG101b complexes were modeled using the structure of the Arabidopsis EDS1-SAG101 complex as a template (PDB:4NFU; Wagner et al., 2013). Sequences of SlSAG101a and SlSAG101b share 38% and 36% sequence identity with AtSAG101, respectively, whereas SlEDS1 and AtEDS1 share 40%. All three sequence-template pairs could thus be confidently aligned using the hhpred algorithm (Zimmermann et al., 2018), and structural models were generated and relaxed based on these alignments using rosettaCM (Song et al., 2013) with limited need for manual realignment in the regions with insertions. Sequence conservation was calculated using the rate4site algorithm (Pupko et al., 2002) and mapped at the surface of structural models using PyMOL (PyMOL Molecular Graphics System, version 2.0; Schrödinger). Structural models and analysis are provided in Supplemental Data Set 3.
Accession Numbers
Sequence data from this article can be found in The Arabidopsis Information Resource, GenBank, and/or Solgenomics databases or the QUT N. benthamiana genome database (http://benthgenome.qut.edu.au/) under the following accession numbers: AtEDS1-At3g48090 (Gene ID: 823964); AtPAD4-At3g52430 (Gene ID: 824408); AtSAG101-At5g14930 (Gene ID: 831345); SlEDS1-Solyc06g071280.2; SlPAD4-Solyc02g032850.2; SlSAG101a-Solyc02g069400.2; SlSAG101b-Solyc02g067660.2; NbEDS1-Niben101Scf06720g01024.1 (QUT: Nbv6.1trP77101); NbPAD4-Niben101Scf02544g01012.1 (QUT: Nbv6.1trP15293); NbSAG101a-Niben101Scf00271g02011.1 (QUT: Nbv6.1trP10532); NbSAG101b-Niben101Scf01300g01009.1 (QUT: Nbv6.1trP73488); NbRoq1-GenBank: MF773579.1. Additional sequence information is provided in Supplemental Data Set 1.
Supplemental Data
Supplemental Figure 1. EDS1 family gene models from Arabidopsis, S. lycopersicum, and N. benthamiana. Supports Figure 1.
Supplemental Figure 2. Expression of tomato EDS1 family genes. Supports Figure 1.
Supplemental Figure 3. Localization and complex formation of tomato EDS1 proteins. Supports Figure 2.
Supplemental Figure 4. Generation of mutant lines by genome editing. Supports Figure 3.
Supplemental Figure 5. Functional comparison of EDS1 and SAG101b from N. benthamiana and S. lycopersicum for XopQ-induced cell death. Supports Figure 3.
Supplemental Figure 6. Cross-species transfer of EDS1-family genes. Supports Figure 5.
Supplemental Figure 7. Heterocomplex formation by EDS1 variants in a yeast two hybrid system. Supports Figure 6.
Supplemental Figure 8. Immune competence of further EDS1 variants. Supports Figure 7.
Supplemental Table 1. SlEDS1 and SlSAG101 variants functionally interrogated in this study.
Supplemental Table 2. Plasmids used in this study.
Supplemental Table 3. Oligonucleotides used in this study.
Supplemental Table 4. Oligonucleotides used for quantitative RT-PCR.
Supplemental Data Set 1. Protein sequences used for phylogenetic analyses and associated gene models.
Supplemental Data Set 2. Multiple sequence alignment underlying phylogenetic analyses. Extended data supporting Figure 1.
Supplemental Data Set 3. Structural models and analysis of tomato EDS1, PAD4, SAG101a, and SAG101b.
Dive Curated Terms
The following phenotypic, genotypic, and functional terms are of significance to the work described in this paper:
Acknowledgments
We thank Bianca Rosinsky for taking care of plant growth facilities and growing plants; Ulla Bonas for generous support, discussions, and critical reading of the article; Martin Schattat for assistance with FRET experiments; Magdalena Krzymowska for providing N and p50 expression constructs; and Jessica Andreani for assistance with phylogenetic analyses. This work was funded by the Deutsche Forschungsgemeinschaft (DFG) (GRC grant STU 642-1/1 to J.S.) and seed funding by the CRC 648 (DFG) to J.S., and by the French Infrastructure for Integrated Structural Biology (FRISBI) (grant ANR-10-INSB-05-01 to R.G.).
AUTHOR CONTRIBUTIONS
J.G. and J.O. performed most of the experiments and analyzed data. C.K. performed additional experiments. J.S. designed the study, supervised the research, and analyzed data. R.G. performed phylogenetic analyses and homology modeling and provided structural insights. J.S. wrote the article with contributions from J.G. and R.G.
Footnotes
Articles can be viewed without a subscription.
References
- Aarts N., Metz M., Holub E., Staskawicz B.J., Daniels M.J., Parker J.E. (1998). Different requirements for EDS1 and NDR1 by disease resistance genes define at least two R gene-mediated signaling pathways in Arabidopsis. Proc. Natl. Acad. Sci. USA 95: 10306–10311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adlung N., Bonas U. (2017). Dissecting virulence function from recognition: Cell death suppression in Nicotiana benthamiana by XopQ/HopQ1-family effectors relies on EDS1-dependent immunity. Plant J. 91: 430–442. [DOI] [PubMed] [Google Scholar]
- Adlung N., Prochaska H., Thieme S., Banik A., Blüher D., John P., Nagel O., Schulze S., Gantner J., Delker C., Stuttmann J., Bonas U. (2016). Non-host resistance induced by the Xanthomonas effector XopQ is widespread within the genus Nicotiana and functionally depends on EDS1. Front. Plant Sci. 7: 1796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arabidopsis Interactome Mapping Consortium. (2011). Evidence for network evolution in an Arabidopsis interactome map. Science 333: 601–607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bent A.F., Kunkel B.N., Dahlbeck D., Brown K.L., Schmidt R., Giraudat J., Leung J., Staskawicz B.J. (1994). RPS2 of Arabidopsis thaliana: a leucine-rich repeat class of plant disease resistance genes. Science 265: 1856–1860. [DOI] [PubMed] [Google Scholar]
- Bentham A., Burdett H., Anderson P.A., Williams S.J., Kobe B. (2017). Animal NLRs provide structural insights into plant NLR function. Ann. Bot. 119: 698–702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernoux M., Ve T., Williams S., Warren C., Hatters D., Valkov E., Zhang X., Ellis J.G., Kobe B., Dodds P.N. (2011). Structural and functional analysis of a plant resistance protein TIR domain reveals interfaces for self-association, signaling, and autoregulation. Cell Host Microbe 9: 200–211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernoux M., Burdett H., Williams S.J., Zhang X., Chen C., Newell K., Lawrence G.J., Kobe B., Ellis J.G., Anderson P.A., Dodds P.N. (2016). Comparative analysis of the flax immune receptors L6 and L7 suggests an equilibrium-based switch activation model. Plant Cell 28: 146–159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhandari D., Lapin D., Kracher B., vonBorn P., Bautor J., Niefind K., Parker J (2019). An EDS1 heterodimer signalling surface enforces timely reprogramming of immunity genes in Arabidopsis. Nat. Commun. 10: 772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhattacharjee S., Halane M.K., Kim S.H., Gassmann W. (2011). Pathogen effectors target Arabidopsis EDS1 and alter its interactions with immune regulators. Science 334: 1405–1408. [DOI] [PubMed] [Google Scholar]
- Bombarely A., Rosli H.G., Vrebalov J., Moffett P., Mueller L.A., Martin G.B. (2012). A draft genome sequence of Nicotiana benthamiana to enhance molecular plant-microbe biology research. Mol. Plant Microbe Interact. 25: 1523–1530. [DOI] [PubMed] [Google Scholar]
- Bonardi V., Tang S., Stallmann A., Roberts M., Cherkis K., Dangl J.L. (2011). Expanded functions for a family of plant intracellular immune receptors beyond specific recognition of pathogen effectors. Proc. Natl. Acad. Sci. USA 108: 16463–16468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brendolise C., Martinez-Sanchez M., Morel A., Chen R., Dinis R., Deroles S., Peeters N., Rikkerink E., Montefiori M. (2018). NRG1-mediated recognition of HopQ1 reveals a link between PAMP and effector-triggered immunity. https://www.biorxiv.org/content/10.1101/293050v1. [Google Scholar]
- Burch-Smith T.M., Schiff M., Caplan J.L., Tsao J., Czymmek K., Dinesh-Kumar S.P. (2007). A novel role for the TIR domain in association with pathogen-derived elicitors. PLoS Biol. 5: e68. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Büttner D. (2016). Behind the lines-actions of bacterial type III effector proteins in plant cells. FEMS Microbiol. Rev. 40: 894–937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castel B., Ngou P.M., Cevik V., Redkar A., Kim D.S., Yang Y., Ding P.,Jones J.D.G (2019). Diverse NLR immune receptors activate defence via the RPW8-NLR NRG1. New Phytol. 222: 966-980 [DOI] [PubMed] [Google Scholar]
- Castresana J. (2000). Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17: 540–552. [DOI] [PubMed] [Google Scholar]
- Chakravarthy S., Velásquez A.C., Ekengren S.K., Collmer A., Martin G.B. (2010). Identification of Nicotiana benthamiana genes involved in pathogen-associated molecular pattern-triggered immunity. Mol. Plant Microbe Interact. 23: 715–726. [DOI] [PubMed] [Google Scholar]
- Chen G., Wei B., Li G., Gong C., Fan R., Zhang X. (2018). TaEDS1 genes positively regulate resistance to powdery mildew in wheat. Plant Mol. Biol. 96: 607–625. [DOI] [PubMed] [Google Scholar]
- Collier S.M., Hamel L.P., Moffett P. (2011). Cell death mediated by the N-terminal domains of a unique and highly conserved class of NB-LRR protein. Mol. Plant Microbe Interact. 24: 918–931. [DOI] [PubMed] [Google Scholar]
- Cui H., Tsuda K., Parker J.E. (2015). Effector-triggered immunity: From pathogen perception to robust defense. Annu. Rev. Plant Biol. 66: 487–511. [DOI] [PubMed] [Google Scholar]
- Cui H., Gobbato E., Kracher B., Qiu J., Bautor J., Parker J.E. (2017). A core function of EDS1 with PAD4 is to protect the salicylic acid defense sector in Arabidopsis immunity. New Phytol. 213: 1802–1817. [DOI] [PubMed] [Google Scholar]
- Cui H., Qiu J., Zhou Y., Bhandari D.D., Zhao C., Bautor J., Parker J.E. (2018). Antagonism of transcription factor MYC2 by EDS1/PAD4 complexes bolsters salicylic acid defense in Arabidopsis effector-triggered immunity. Mol. Plant 11: 1053–1066. [DOI] [PubMed] [Google Scholar]
- Dong O.X., Tong M., Bonardi V., El Kasmi F., Woloshen V., Wünsch L.K., Dangl J.L., Li X. (2016). TNL-mediated immunity in Arabidopsis requires complex regulation of the redundant ADR1 gene family. New Phytol. 210: 960–973. [DOI] [PubMed] [Google Scholar]
- Engler C., Kandzia R., Marillonnet S. (2008). A one pot, one step, precision cloning method with high throughput capability. PLoS One 3: e3647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engler C., Youles M., Gruetzner R., Ehnert T.M., Werner S., Jones J.D., Patron N.J., Marillonnet S. (2014). A Golden Gate modular cloning toolbox for plants. ACS Synth. Biol. 3: 839–843. [DOI] [PubMed] [Google Scholar]
- Falk A., Feys B.J., Frost L.N., Jones J.D., Daniels M.J., Parker J.E. (1999). EDS1, an essential component of R gene-mediated disease resistance in Arabidopsis has homology to eukaryotic lipases. Proc. Natl. Acad. Sci. USA 96: 3292–3297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feys B.J., Moisan L.J., Newman M.A., Parker J.E. (2001). Direct interaction between the Arabidopsis disease resistance signaling proteins, EDS1 and PAD4. EMBO J. 20: 5400–5411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feys B.J., Wiermer M., Bhat R.A., Moisan L.J., Medina-Escobar N., Neu C., Cabral A., Parker J.E. (2005). Arabidopsis SENESCENCE-ASSOCIATED GENE101 stabilizes and signals within an ENHANCED DISEASE SUSCEPTIBILITY1 complex in plant innate immunity. Plant Cell 17: 2601–2613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gantner J., Ordon J., Ilse T., Kretschmer C., Gruetzner R., Löfke C., Dagdas Y., Bürstenbinder K., Marillonnet S., Stuttmann J. (2018). Peripheral infrastructure vectors and an extended set of plant parts for the modular cloning system. PLoS One 13: e0197185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- García A.V., Blanvillain-Baufumé S., Huibers R.P., Wiermer M., Li G., Gobbato E., Rietz S., Parker J.E. (2010). Balanced nuclear and cytoplasmic activities of EDS1 are required for a complete plant innate immune response. PLoS Pathog. 6: e1000970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gietz R.D., Schiestl R.H. (2007). Frozen competent yeast cells that can be transformed with high efficiency using the LiAc/SS carrier DNA/PEG method. Nat. Protoc. 2: 1–4. [DOI] [PubMed] [Google Scholar]
- Goodin M.M., Zaitlin D., Naidu R.A., Lommel S.A. (2008). Nicotiana benthamiana: Its history and future as a model for plant-pathogen interactions. Mol. Plant Microbe Interact. 21: 1015–1026. [DOI] [PubMed] [Google Scholar]
- Gouy M., Guindon S., Gascuel O. (2010). SeaView version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 27: 221–224. [DOI] [PubMed] [Google Scholar]
- Guindon S., Dufayard J.F., Lefort V., Anisimova M., Hordijk W., Gascuel O. (2010). New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 59: 307–321. [DOI] [PubMed] [Google Scholar]
- Hecker A., Wallmeroth N., Peter S., Blatt M.R., Harter K., Grefen C. (2015). Binary 2in1 Vectors Improve in Planta (Co)localization and Dynamic Protein Interaction Studies. Plant Physiol. 168: 776–787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heidrich K., Wirthmueller L., Tasset C., Pouzet C., Deslandes L., Parker J.E. (2011). Arabidopsis EDS1 connects pathogen effector recognition to cell compartment-specific immune responses. Science 334: 1401–1404. [DOI] [PubMed] [Google Scholar]
- Hofberger J.A., Zhou B., Tang H., Jones J.D., Schranz M.E. (2014). A novel approach for multi-domain and multi-gene family identification provides insights into evolutionary dynamics of disease resistance genes in core eudicot plants. BMC Genomics 15: 966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu G., deHart A.K., Li Y., Ustach C., Handley V., Navarre R., Hwang C.F., Aegerter B.J., Williamson V.M., Baker B. (2005). EDS1 in tomato is required for resistance mediated by TIR-class R genes and the receptor-like R gene Ve. Plant J. 42: 376–391. [DOI] [PubMed] [Google Scholar]
- Jacob F., Vernaldi S., Maekawa T. (2013). Evolution and conservation of plant NLR functions. Front. Immunol. 4: 297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones J.D., Dangl J.L. (2006). The plant immune system. Nature 444: 323–329. [DOI] [PubMed] [Google Scholar]
- Jones J.D., Vance R.E., Dangl J.L. (2016). Intracellular innate immune surveillance devices in plants and animals. Science 354: aaf6395. [DOI] [PubMed] [Google Scholar]
- Katoh K., Standley D.M. (2013). MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30: 772–780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khan M., Subramaniam R., Desveaux D. (2016). Of guards, decoys, baits and traps: Pathogen perception in plants by type III effector sensors. Curr. Opin. Microbiol. 29: 49–55. [DOI] [PubMed] [Google Scholar]
- Kim S.H., Kwon S.I., Saha D., Anyanwu N.C., Gassmann W. (2009). Resistance to the Pseudomonas syringae effector HopA1 is governed by the TIR-NBS-LRR protein RPS6 and is enhanced by mutations in SRFR1. Plant Physiol. 150: 1723–1732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim T.-H., Kunz H.-H., Bhattacharjee S., Hauser F., Park J., Engineer C., Liu A., Ha T., Parker J.E., Gassmann W., Schroeder J.I. (2012). Natural variation in small molecule-induced TIR-NB-LRR signaling induces root growth arrest via EDS1- and PAD4-complexed R protein VICTR in Arabidopsis. Plant Cell 24: 5177–5192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krasileva K.V., Zheng C., Leonelli L., Goritschnig S., Dahlbeck D., Staskawicz B.J. (2011). Global analysis of Arabidopsis/downy mildew interactions reveals prevalence of incomplete resistance and rapid evolution of pathogen recognition. PLoS One 6: e28765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kushalappa A.C., Yogendra K.N., Karre S. (2016). Plant innate immune response: Qualitative and quantitative resistance. Crit. Rev. Plant Sci. 35: 38–55. [Google Scholar]
- Kushnirov V.V. (2000). Rapid and reliable protein extraction from yeast. Yeast 16: 857–860. [DOI] [PubMed] [Google Scholar]
- Lapin D., Kovacova V., Sun X., Dongus J.A., Bhandari D.D., von Born P., Bautor J., Guarneri N., Rzemieniewski J., Stuttmann J., Beyer A., Parker J.E. (2019). A coevolved EDS1-SAG101-NRG1 module mediates cell death signaling by TIR-domain immune receptors. Plant Cell tpc.00118.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Legay S., Guerriero G., André C., Guignard C., Cocco E., Charton S., Boutry M., Rowland O., Hausman J.F. (2016). MdMyb93 is a regulator of suberin deposition in russeted apple fruit skins. New Phytol. 212: 977–991. [DOI] [PubMed] [Google Scholar]
- Letunic I., Bork P. (2019). Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 47: 256-259 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lipka V., et al. (2005). Pre- and postinvasion defenses both contribute to nonhost resistance in Arabidopsis. Science 310: 1180–1183. [DOI] [PubMed] [Google Scholar]
- Liu D., Shi L., Han C., Yu J., Li D., Zhang Y. (2012). Validation of reference genes for gene expression studies in virus-infected Nicotiana benthamiana using quantitative real-time PCR. PLoS One 7: e46451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Logemann E., Birkenbihl R.P., Ülker B., Somssich I.E. (2006). An improved method for preparing Agrobacterium cells that simplifies the Arabidopsis transformation protocol. Plant Methods 2: 16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Macho A.P., Zipfel C. (2014). Plant PRRs and the activation of innate immune signaling. Mol. Cell 54: 263–272. [DOI] [PubMed] [Google Scholar]
- Macho A.P., Zipfel C. (2015). Targeting of plant pattern recognition receptor-triggered immunity by bacterial type-III secretion system effectors. Curr. Opin. Microbiol. 23: 14–22. [DOI] [PubMed] [Google Scholar]
- Maekawa T., et al. (2011b). Coiled-coil domain-dependent homodimerization of intracellular barley immune receptors defines a minimal functional module for triggering cell death. Cell Host Microbe 9: 187–199. [DOI] [PubMed] [Google Scholar]
- Maekawa T., Kufer T.A., Schulze-Lefert P. (2011a). NLR functions in plant and animal immune systems: So far and yet so close. Nat. Immunol. 12: 817–826. [DOI] [PubMed] [Google Scholar]
- Maekawa T., Kracher B., Vernaldi S., Ver Loren van Themaat E., Schulze-Lefert P. (2012). Conservation of NLR-triggered immunity across plant lineages. Proc. Natl. Acad. Sci. USA 109: 20119–20123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyers B.C., Kozik A., Griego A., Kuang H., Michelmore R.W. (2003). Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 15: 809–834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mindrinos M., Katagiri F., Yu G.-L., Ausubel F.M. (1994). The A. thaliana disease resistance gene RPS2 encodes a protein containing a nucleotide-binding site and leucine-rich repeats. Cell 78: 1089–1099. [DOI] [PubMed] [Google Scholar]
- Monteiro F., Nishimura M.T. (2018). Structural, functional, and genomic diversity of plant NLR proteins: An evolved resource for rational engineering of plant immunity. Annu. Rev. Phytopathol. 56: 243–267. [DOI] [PubMed] [Google Scholar]
- Naim F., Nakasugi K., Crowhurst R.N., Hilario E., Zwart A.B., Hellens R.P., Taylor J.M., Waterhouse P.M., Wood C.C. (2012). Advanced engineering of lipid metabolism in Nicotiana benthamiana using a draft genome and the V2 viral silencing-suppressor protein. PLoS One 7: e52717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ordon J., Gantner J., Kemna J., Schwalgun L., Reschke M., Streubel J., Boch J., Stuttmann J. (2017). Generation of chromosomal deletions in dicotyledonous plants employing a user-friendly genome editing toolkit. Plant J. 89: 155–168. [DOI] [PubMed] [Google Scholar]
- Parker J.E., Holub E.B., Frost L.N., Falk A., Gunn N.D., Daniels M.J. (1996). Characterization of eds1, a mutation in Arabidopsis suppressing resistance to Peronospora parasitica specified by several different RPP genes. Plant Cell 8: 2033–2046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peart J.R., Cook G., Feys B.J., Parker J.E., Baulcombe D.C. (2002). An EDS1 orthologue is required for N-mediated resistance against tobacco mosaic virus. Plant J. 29: 569–579. [DOI] [PubMed] [Google Scholar]
- Peele H.M., Guan N., Fogelqvist J., Dixelius C. (2014). Loss and retention of resistance genes in five species of the Brassicaceae family. BMC Plant Biol. 14: 298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poland J.A., Balint-Kurti P.J., Wisser R.J., Pratt R.C., Nelson R.J. (2009). Shades of gray: The world of quantitative disease resistance. Trends Plant Sci. 14: 21–29. [DOI] [PubMed] [Google Scholar]
- Pombo M.A., Zheng Y., Fernandez-Pozo N., Dunham D.M., Fei Z., Martin G.B. (2014). Transcriptomic analysis reveals tomato genes whose expression is induced specifically during effector-triggered immunity and identifies the Epk1 protein kinase which is required for the host response to three bacterial effector proteins. Genome Biol. 15: 492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pupko T., Bell R.E., Mayrose I., Glaser F., Ben-Tal N. (2002). Rate4Site: An algorithmic tool for the identification of functional regions in proteins by surface mapping of evolutionary determinants within their homologues. Bioinformatics 18: S71–S77. [DOI] [PubMed] [Google Scholar]
- Qi T., Seong K., Thomazella D.P.T., Kim J.R., Pham J., Seo E., Cho M.-J., Schultink A., Staskawicz B.J. (2018). NRG1 functions downstream of EDS1 to regulate TIR-NLR-mediated plant immunity in Nicotiana benthamiana. Proc. Natl. Acad. Sci. USA. 115: E10979-E10987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rietz S., Stamm A., Malonek S., Wagner S., Becker D., Medina-Escobar N., Vlot A.C., Feys B.J., Niefind K., Parker J.E. (2011). Different roles of Enhanced Disease Susceptibility1 (EDS1) bound to and dissociated from Phytoalexin Deficient4 (PAD4) in Arabidopsis immunity. New Phytol. 191: 107–119. [DOI] [PubMed] [Google Scholar]
- Rosli H.G., Zheng Y., Pombo M.A., Zhong S., Bombarely A., Fei Z., Collmer A., Martin G.B. (2013). Transcriptomics-based screen for genes induced by flagellin and repressed by pathogen effectors identifies a cell wall-associated kinase involved in plant immunity. Genome Biol. 14: R139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schön M., Töller A., Diezel C., Roth C., Westphal L., Wiermer M., Somssich I.E. (2013). Analyses of wrky18 wrky40 plants reveal critical roles of SA/EDS1 signaling and indole-glucosinolate biosynthesis for Golovinomyces orontii resistance and a loss-of resistance towards Pseudomonas syringae pv. tomato AvrRPS4. Mol. Plant Microbe Interact. 26: 758–767. [DOI] [PubMed] [Google Scholar]
- Schultink A., Qi T., Lee A., Steinbrenner A., Staskawicz B (2017). Roq1 mediates recognition of the Xanthomonas and Pseudomonas effector proteins XopQ and HopQ1. Plant J. 92: 787-795 [DOI] [PubMed] [Google Scholar]
- Shen C., Lu A., Xie W.J., Ruan J., Negro R., Egelman E.H., Fu T.-M., Wu H. (2019). Molecular mechanism for NLRP6 inflammasome assembly and activation. Proc. Natl. Acad. Sci. USA. 116: 2052-2057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimada T.L., Shimada T., Hara-Nishimura I. (2010). A rapid and non-destructive screenable marker, FAST, for identifying transformed seeds of Arabidopsis thaliana. Plant J. 61: 519–528. [DOI] [PubMed] [Google Scholar]
- Sinapidou E., Williams K., Nott L., Bahkt S., Tör M., Crute I., Bittner-Eddy P., Beynon J. (2004). Two TIR:NB:LRR genes are required to specify resistance to Peronospora parasitica isolate Cala2 in Arabidopsis. Plant J. 38: 898–909. [DOI] [PubMed] [Google Scholar]
- Solovyev V.V. (2007). Statistical approaches in Eukaryotic gene prediction. In Handbook of Statistical Genetics. Balding D.J., Bishop M., Cannings C. eds. (New Jersey: John Wiley & Sons; ) pp. 97-159. [Google Scholar]
- Song Y., DiMaio F., Wang R.Y., Kim D., Miles C., Brunette T., Thompson J., Baker D. (2013). High-resolution comparative modeling with RosettaCM. Structure 21: 1735–1742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spoel S.H., Dong X. (2012). How do plants achieve immunity? Defence without specialized immune cells. Nat. Rev. Immunol. 12: 89–100. [DOI] [PubMed] [Google Scholar]
- Stuttmann J., Hubberten H.M., Rietz S., Kaur J., Muskett P., Guerois R., Bednarek P., Hoefgen R., Parker J.E. (2011). Perturbation of Arabidopsis amino acid metabolism causes incompatibility with the adapted biotrophic pathogen Hyaloperonospora arabidopsidis. Plant Cell 23: 2788–2803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stuttmann J., Peine N., Garcia A.V., Wagner C., Choudhury S.R., Wang Y., James G.V., Griebel T., Alcázar R., Tsuda K., Schneeberger K., Parker J.E. (2016). Arabidopsis thaliana DM2h (R8) within the Landsberg RPP1-like resistance locus underlies three different cases of EDS1-conditioned autoimmunity. PLoS Genet. 12: e1005990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swiderski M.R., Birker D., Jones J.D. (2009). The TIR domain of TIR-NB-LRR resistance proteins is a signaling domain involved in cell death induction. Mol. Plant Microbe Interact. 22: 157–165. [DOI] [PubMed] [Google Scholar]
- Takken F.L.W., Goverse A. (2012). How to build a pathogen detector: Structural basis of NB-LRR function. Curr. Opin. Plant Biol. 15: 375–384. [DOI] [PubMed] [Google Scholar]
- Thieme F., et al. (2005). Insights into genome plasticity and pathogenicity of the plant pathogenic bacterium Xanthomonas campestris pv. vesicatoria revealed by the complete genome sequence. J. Bacteriol. 187: 7254–7266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas W.J., Thireault C.A., Kimbrel J.A., Chang J.H. (2009). Recombineering and stable integration of the Pseudomonas syringae pv. syringae 61 hrp/hrc cluster into the genome of the soil bacterium Pseudomonas fluorescens Pf0-1. Plant J. 60: 919–928. [DOI] [PubMed] [Google Scholar]
- Van de Weyer A-L, Monteiro F, Furzer OJ, Nishimura MT, Cevik V, Witek K, Jones D.G.J., Dangl J.L., Weigel D., Bemm F (2019). The Arabidopsis thaliana pan-NLRome. https://www.biorxiv.org/content/10.1101/537001v1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Venugopal S.C., Jeong R.-D., Mandal M.K., Zhu S., Chandra-Shekara A.C., Xia Y., Hersh M., Stromberg A.J., Navarre D., Kachroo A., Kachroo P. (2009). Enhanced disease susceptibility 1 and salicylic acid act redundantly to regulate resistance gene-mediated signaling. PLoS Genet. 5: e1000545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vlot A.C., Dempsey D.A., Klessig D.F. (2009). Salicylic acid, a multifaceted hormone to combat disease. Annu. Rev. Phytopathol. 47: 177–206. [DOI] [PubMed] [Google Scholar]
- Wagner S., Stuttmann J., Rietz S., Guerois R., Brunstein E., Bautor J., Niefind K., Parker J.E. (2013). Structural basis for signaling by exclusive EDS1 heteromeric complexes with SAG101 or PAD4 in plant innate immunity. Cell Host Microbe 14: 619–630. [DOI] [PubMed] [Google Scholar]
- Wang J., Hu M., Wang J., Qi J., Han Z., Wang G., Chai J. (2019a). Reconstitution and structure of a plant NLR resistosome conferring immunity. Science. 364: 5870. [DOI] [PubMed] [Google Scholar]
- Wang J., Wang J., Hu M., Wu S., Qi J., Wang G., Chai J. (2019b). Ligand-triggered allosteric ADP release primes a plant NLR complex. Science 364. [DOI] [PubMed] [Google Scholar]
- Waterhouse A.M., Procter J.B., Martin D.M., Clamp M., Barton G.J. (2009). Jalview version 2: A multiple sequence alignment editor and analysis workbench. Bioinformatics 25: 1189–1191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weber E., Engler C., Gruetzner R., Werner S., Marillonnet S. (2011). A modular cloning system for standardized assembly of multigene constructs. PLoS One 6: e16765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei H.L., Zhang W., Collmer A. (2018). Modular study of the type III effector repertoire in Pseudomonas syringae pv. tomato DC3000 reveals a matrix of effector interplay in pathogenesis. Cell Reports 23: 1630–1638. [DOI] [PubMed] [Google Scholar]
- Wiermer M., Feys B.J., Parker J.E. (2005). Plant immunity: The EDS1 regulatory node. Curr. Opin. Plant Biol. 8: 383–389. [DOI] [PubMed] [Google Scholar]
- Williams S.J., et al. (2014). Structural basis for assembly and function of a heterodimeric plant immune receptor. Science 344: 299–303. [DOI] [PubMed] [Google Scholar]
- Wirthmueller L., Zhang Y., Jones J.D., Parker J.E. (2007). Nuclear accumulation of the Arabidopsis immune receptor RPS4 is necessary for triggering EDS1-dependent defense. Curr. Biol. 17: 2023–2029. [DOI] [PubMed] [Google Scholar]
- Wu Z., Li M., Dong O.X., Xia S., Liang W., Bao Y., Wasteneys G., Li X. (2019). Differential regulation of TNL-mediated immune signaling by redundant helper CNLs. New Phytol. 222: 938–953. [DOI] [PubMed] [Google Scholar]
- Xu F., Zhu C., Cevik V., Johnson K., Liu Y., Sohn K., Jones J.D., Holub E.B., Li X. (2015). Autoimmunity conferred by chs3-2D relies on CSA1, its adjacent TNL-encoding neighbour. Sci. Rep. 5: 8792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu X., Feng B., He P., Shan L. (2017). From chaos to harmony: Responses and signaling upon microbial pattern recognition. Annu. Rev. Phytopathol. 55: 109–137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zembek P., et al. (2018). Two strategies of Pseudomonas syringae to avoid recognition of the HopQ1 effector in Nicotiana species. Front. Plant Sci. 9: 978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu S., Jeong R.D., Venugopal S.C., Lapchyk L., Navarre D., Kachroo A., Kachroo P. (2011). SAG101 forms a ternary complex with EDS1 and PAD4 and is required for resistance signaling against turnip crinkle virus. PLoS Pathog. 7: e1002318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zimmermann L., Stephens A., Nam S.Z., Rau D., Kübler J., Lozajic M., Gabler F., Söding J., Lupas A.N., Alva V. (2018). A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430: 2237–2243. [DOI] [PubMed] [Google Scholar]
- Zouine M., Maza E., Djari A., Lauvernier M., Frasse P., Smouni A., Pirrello J., Bouzayen M. (2017). TomExpress, a unified tomato RNA-Seq platform for visualization of expression data, clustering and correlation networks. Plant J. 92: 727–735. [DOI] [PubMed] [Google Scholar]