Abstract
Modern society is hugely dependent on finite oil reserves for the supply of fuels and chemicals. Moving our dependence away from these unsustainable oil-based feedstocks to renewable ones is, therefore, a critical factor towards the development of a low carbon bioeconomy. Lignin derived from biomass feedstocks offers great potential as a renewable source of aromatic compounds if methods for its effective valorization can be developed. Synthetic biology and metabolic engineering offer the potential to synergistically enable the development of cell factories with novel biosynthetic routes to valuable chemicals from these sustainable sources. Pathway design and optimization is, however, a major bottleneck due to the lack of high-throughput methods capable of screening large libraries of genetic variants and the metabolic burden associated with bioproduction. Genetically encoded biosensors can provide a solution by transducing the target metabolite concentration into detectable signals to provide high-throughput phenotypic read-outs and allow dynamic pathway regulation. The development and application of biosensors in the discovery and engineering of efficient biocatalytic processes for the degradation, conversion, and valorization of lignin are paving the way towards a sustainable and economically viable biorefinery.
Keywords: Genetically encoded biosensors, Sustainable chemical production, Lignocellulose valorization, Biorefinery
Background
Modern society is largely dependent on natural geological supplies, including crude oil, coal, and natural gas. Our reliance on these fossil fuels is, however, responsible for many causes of global concern, particularly greenhouse gas emissions and the depletion of natural resources [1, 2]. These concerns require a shift towards a sustainable economy with alternative routes to chemicals, fuels, and energy from renewable feedstocks, for which plant biomass constitutes an attractive raw material [3, 4]. During the last few decades, the concept of bioeconomy has become increasingly adopted by governments and industries, owing to its many potential environmental and economic advantages. Indeed, policies and legislations have been introduced to stimulate demand for bio-based products and biofuels derived from sustainable biomass. In the US, the bioeconomy initiative has been introduced to promote the development of new conversion technologies to upgrade biomass feedstocks [5–7]. Targets for chemical production from biomass in Europe are expected to replace at least 30% of oil-based chemicals by 2030, in accordance with the EU Bioeconomy Strategy [8].
There is a particular global socioeconomic interest in leveraging aromatic chemicals derived from lignocellulosic biomass [6, 8–10]. The major fraction of these plant aromatics is contained in the recalcitrant polymer lignin, which is found in quantities of up to 30% w/w, making it the most abundant aromatic heteropolymer on Earth [11, 12]. Thus, lignin has tremendous potential as a renewable feedstock to satisfy the current chemical and energy demands for a carbon neutral bioeconomy [10]. Lignocellulosic materials are mainly derived from non-edible residues from agriculture, dedicated energy crops and wood deposits [7, 13, 14]. Particularly high amounts of lignin are likely to be generated from the numerous cellulosic bioethanol plants which are being commissioned worldwide [15–17]. However, the current biomass utilization processes in these industries fail to recognize lignin as a valuable waste stream. Second-generation lignocellulosic biorefineries focus solely on the valorization of sugars produced from saccharification and enzymatic treatment, which are fermented to bioethanol and other biofuels, while the remaining lignin produced is used for power and heat generation [18]. Instead, efficient lignocellulosic biorefineries must strive to maximise the utilization and valorization of all the distinct biomass fractions, including lignin [19].
Enzymatic cocktails are extensively used in second-generation bioethanol production from cellulose [20]; thus, the extension of this enzymatic approach to the valorization of lignin could provide an expanded framework to a sustainable biorefinery approach. However, direct biological degradation and conversion of lignin into high value aromatic chemicals has not yet been realized at an industrial scale, owing to the intrinsic difficulties with its heterogeneity and recalcitrance. The discovery and development of efficient biocatalytic processes for the degradation, conversion, and valorization of lignin will be essential to achieve a novel, sustainable and economically viable valorization process for the long-term success of lignocellulosic biorefineries. This review focuses on the use of genetically encoded biosensors as tools to achieve these goals, and describes future avenues for technological advancement in the field. In particular, we review genetically encoded biosensors that have been developed and optimized to date to sense biomass-derived aromatic monomers and sugars towards lignin valorization applications.
The screening bottleneck—metabolic pathway engineering and optimization
Metabolic engineering and synthetic biology present an emerging opportunity to develop direct routes to value-added chemicals from lignin. Aromatic precursors can be fed into functionally reconstructed biotransformation routes or modified to allow their introduction into de novo biosynthetic pathways. Lignin monomers can consequently be upgraded into fine chemicals such as flavouring agents, fragrances, and pharmaceuticals. Such approaches have enabled the production of numerous aromatic acids, alcohols, and aldehydes [21], and of flavonoids [22], stilbenes [23], and coumarins [24]. However, incorporation of heterologous or de novo biosynthetic pathways into a host often leads to an imbalanced metabolic flux. This imbalance can lead to toxic accumulation of intermediates that inhibit cell growth and decrease process performance. In addition, the lack of effective transport systems in these hosts can also result in reduced productivity, as these proteins are responsible for importing aromatic substrates and exporting products [25]. Pathway design and optimization for balanced enzymatic activity is, therefore, central to reach an optimal product yield and titer [26–28].
Strategies for pathway optimization and protein engineering of biocatalysts are generally based on the generation of large genetic variant libraries with the final aim of yielding highly producing strains and biocatalysts. Millions of variants are designed and obtained via rational or random mutagenesis techniques and often aided by computational predictions [29]. However, the screening and selection of target phenotypes as well as monitoring and quantifying the synthesis of chemical products constitute a major bottleneck in the design-build-test cycle, as conventional methods for metabolite measurement are low-throughput and laborious [30, 31]. Without high-throughput screening and selection methods, the ideal target strains and biocatalysts could remain unrealised. Genetically encoded biosensors provide a solution, as they can be deployed to translate the target metabolite concentration into useful detectable signals amenable to high-throughput analysis.
Genetically encoded biosensors
In nature, cells have evolved sophisticated genetic regulatory systems to perceive changes in environmental conditions and detect intra- and extracellular metabolites [32]. In these systems, sensing of the target signal or metabolite (input) is coupled to an actuator response, which leads to the alteration of protein or gene activity within the cell (output). Such systems can be harnessed to construct genetically encoded biosensors suitable to measure the concentration of target metabolites and translate the input into a useful output (e.g., fluorescent protein and biosynthetic pathway expression). As such, biosensors provide a method to detect, monitor and regulate cellular metabolic productivity [32–35].
Several molecular systems have been used to create biosensors, including RNA riboswitches and several ligand-dependent proteins and receptors such as histidine kinases (HK) and allosteric transcription factors (aTFs) [33, 36, 37]. Extensive reviews have recently been published that cover each mechanism relating to the construction of biosensors [33, 34, 36, 38–40]. Here, we focus on protein–based systems, particularly transcriptional biosensors and their application towards biomass-derived metabolite detection.
Transcription factors are proteins capable of regulating gene transcription by binding to specific DNA operator sequences within promoters, in turn controlling RNA polymerase activity. These regulators may activate (activators) or repress (repressors) the expression of actuator genes and regulators [41]. Allosteric TFs (aTFs) undergo a change in their conformation when activated by an effector molecule, thus altering their DNA-binding activity [42]. Allosteric transcriptional regulation has become a particularly popular method for metabolite sensing due to the myriad of native effector molecule specificities [33] including numerous aromatic compounds (Table 1), and have been constructed as tools for distinct applications and outputs (Fig. 1).
Table 1.
Genetically encoded biosensors developed and applied to sense aromatic monomers and sugars towards lignin valorization
| Target molecule(s) | Sensing element | Output element | Host organism | References |
|---|---|---|---|---|
| Aromatic monomers | ||||
| Protocatechuic acid | PcaU | Engineered PpcaU, P3B5B | Escherichia coli | [69, 101] |
| PcaU | Engineered PpcaU | Pseudomonas putida | [70] | |
| PobR variant | Ppob | Escherichia coli | [75] | |
| PcaV | Engineered PPV | Escherichia coli | [76] | |
| Vanillin | EmrR | PemrR | Escherichia coli | [68] |
| Yap1/Msn2 | Padh7 | Escherichia coli | [89] | |
| QacR variant | PqacA | Escherichia coli | [74] | |
| nd | PyeiW | Escherichia coli | [64] | |
| EmrR | Engineered PemrR | Escherichia coli | [67] | |
| EmrR | PemrR | Escherichia coli | [79] | |
| YqhC | PyqhD | Escherichia coli | [102] | |
| PcaV variant | Engineered PPV | Escherichia coli | [76] | |
| VanR | Engineered PVanO | Escherichia coli | [103] | |
| nd | PLPD00563 | Rhodococcus opacus | [104] | |
| Vanillic acid | VanR | PvanCC | Escherichia coli | [101] |
| EmrR | PemrR | Escherichia coli | [67] | |
| EmrR | PemrR | Escherichia coli | [79] | |
| p-Coumaric acid | EmrR | Engineered PemrR | Escherichia coli | [67] |
| EmrR | PemrR | Escherichia coli | [79] | |
| PadR | PpadC | Corynebacterium glutamicum | [48] | |
| Ferulic acid and related compounds | FerC | Engineered PLC and PPC | Escherichia coli | [66] |
| FerC | Pech | Escherichia coli | [88] | |
| Cinnamaldehyde | BldR | PSso2536adh | Escherichia coli | [105] |
| Benzoic acid | EmrR | PemrR | Escherichia coli | [79] |
| NahR | Psal | Escherichia coli | [85] | |
| BenR | PbenA | Escherichia coli | [80] | |
| nd | PLPD06580 | Rhodococcus opacus | [104] | |
| 4-Hydroxybenzoic acid | EmrR | PemrR | Escherichia coli | [79] |
| PobR variant | Ppob | Escherichia coli, Pseudomonas putida | [75, 86] | |
| nd | PLPD06764 | Rhodococcus opacus | [104] | |
| 2-Hydroxybenzoic acid | NahR | Psal | Escherichia coli | [85] |
| Benzaldehyde | BldR | PSso2536adh | Escherichia coli | [105] |
| Benzoic acid derivatives | XylS | Pm | Escherichia coli | [71] |
| Salicylic acid | EmrR | PemrR | Escherichia coli | [79] |
| Engineered AraC | PBAD | Escherichia coli | [106] | |
| Salicylaldehyde | BldR | PSso2536adh | Escherichia coli | [105] |
| NahR | Psal | Escherichia coli | [85] | |
| Salicylic acid derivatives | XylS, NahR | Pm, Psal | Escherichia coli | [71] |
| Phenol | EmrR | PemrR | Escherichia coli | [79] |
| Phenol derivatives | DmpR | Po | Escherichia coli | [71] |
| Syringaldehyde | EmrR | PemrR | Escherichia coli | [68] |
| BTX | XylR | Engineered Po’ | Escherichia coli | [65] |
| Other aromatic aldehydes | BldR | PSso2536adh | Escherichia coli | [105] |
| PcaV variant | Engineered PPV | Escherichia coli | [76] | |
| C5/C6 sugars | ||||
| Xylose | XBP−XynA | PxylF | Escherichia coli | [107] |
| XylR | Engineered PGAL1/XylR | Saccharomyces cerevisiae | [91] | |
| XylR–XBP | PxylF | Escherichia coli | [95] | |
| d-Arabinose | araF | FRET | Escherichia coli | [94] |
| AraC variant | PBAD | Escherichia coli | [90] | |
| l-Rhamnose | RhaS | PrhaBAD | Escherichia coli | [96] |
| l-Mannose | RhaS | PrhaBAD | Escherichia coli | [96] |
| Cellulase | CelR | PTRC | Escherichia coli | [92] |
| Maltose | AraF | FRET | Escherichia coli | [94] |
| Glucose, glutamate | MglB, YbeJ | FRET | Escherichia coli | [93] |
nd not determined
Fig. 1.
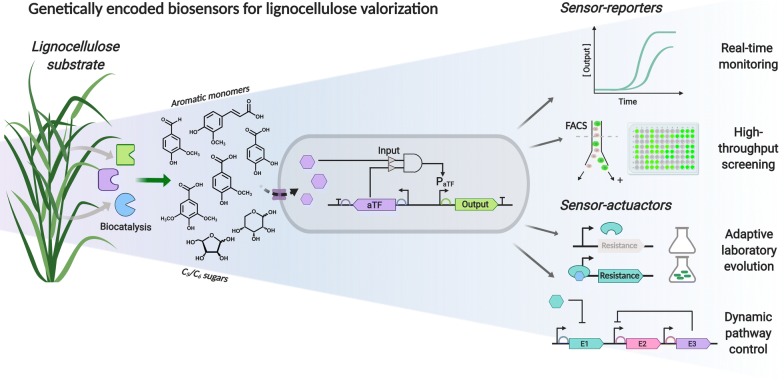
Applications of genetically encoded biosensors in lignin valorization. Transcriptional biosensors allow the coupling of aromatic monomer or sugar release detection (input) with the expression of a distinct measurable output. Reporter-based outputs can be applied in real-time metabolite monitoring and high-throughput screening of large mutant libraries using FACS. Inputs coupled to an actuator output can afford the dynamic control of enzymatic pathways according to the input level or provide an adaptive advantage under selective conditions through adaptive laboratory evolution, allowing the enrichment of high-producing strains
Sensor reporters—biosensors for phenotype screening
Genetically encoded biosensors are well suited for phenotype screening when the detection of a target metabolite is coupled to fluorescent protein expression [36]. These systems generate an output signal that is proportional to the final product yield, allowing the biosensor to be used in applications such as the real-time monitoring of metabolite bioproduction (Fig. 1). Owing to the correlation between fluorescence output and enzyme or pathway efficiency, substrate utilization and final product production can be monitored to facilitate optimization [43].
When coupled to sorting techniques such as fluorescence-activated cell sorting (FACS) or microfluidics, biosensors permit the high-throughput screening and sorting of large mutant genetic libraries for the selection of high-producing phenotypes and biocatalysts, useful in directed evolution applications (Fig. 1) [27, 35]. When assessing variant libraries, conventional screening techniques such as high-performance liquid chromatography (HPLC) or mass spectrometry (MS) are restricted to only 103 clones per day, and are time-consuming and laborious. In contrast, biosensors and FACS-mediated screening can enable the high-throughput screening of up to 107 clones per day [44]. Transcriptional biosensors have been particularly successful in the sorting and selection of improved or novel enzymes and pathways using FACS [31, 33, 43, 45]. For instance, a biosensor based on the dimethyl phenol regulatory protein (DmpR) was utilized in the high-throughput screening of 6 × 106 mutants from a tryptophan-indole lyase library to select improved enzymes using this sorting technique [44]. Other genetically encoded biosensors have also been employed for the analysis of metabolic production of mevalonate [46], lysine [43], and triacetic acid lactones [47]. In addition, when biosensors are coupled to microfluidics, library sizes of more than 108 clones can be screened per day [48]. In general, biosensor-based techniques provide optimal sensitivity and high performance, closing the gap between variant generation and the subsequent selection from these libraries.
Sensor actuators—biosensors for dynamic pathway control and adaptive evolution
Besides providing information about the intracellular presence of target metabolites, biosensors can alternatively be employed as actuators when effector binding is coupled to expression of genes encoding enzymes with activity towards the effector (Fig. 1). Heterologous bioproduction can lead to toxic intermediate accumulation and metabolic stress which retards cell growth, rendering the process ineffective (see “The screening bottleneck—metabolic pathway engineering and optimization” section) [26]. Utilizing biosensors as regulatory systems for dynamic pathway control can alleviate these problems by allowing temporal and on-demand gene and pathway expression, reducing the metabolic burden of toxic intermediates and improving the efficient utilisation of cellular energy and resources [28]. For instance, Zhang et al. [49] developed a sensor-actuator system based on the fatty acid-responsive FadR aTF, which allowed cells to sense sufficient accumulation of the intermediate fatty acyl-CoA and regulate a pathway for its conversion into fatty acid ethyl ester. This strategy achieved balanced cell growth rate and increased biodiesel production by threefold [49]. Similarly, Xu et al. [50] constructed a sensor-actuator system based on the malonyl-CoA responsive FapR aTF to dynamically regulate a fatty acid (FA) biosynthesis pathway upon malonyl-CoA levels. In this way, the system achieved a balanced metabolism with a 2.1-fold increase in FA production compared to the uncontrolled pathway [50, 51].
Moreover, controlling evolution with biosensors by linking sensing and cell fitness (i.e., through regulation of an antibiotic resistance gene) enables the enrichment of mutant libraries with improved phenotypes and productivities through adaptive laboratory evolution (ALE). Adaptive evolution approaches have proven a valuable tool to improve enzymatic activity and metabolic productivity under desired selective conditions [52]. This approach has been demonstrated for the increased production of numerous metabolites, including tyrosine [53], l-valine [54] and muconic acid [55]. This approach also allows control of cell-to-cell variation in metabolic engineering applications, allowing the fine-tuning of synthetic pathways for optimized production [56, 57]. Xiao et al. [58] developed a method to exploit cell-to-cell variation for increased biosynthesis of free fatty acid (FFA) and tyrosine. The method is based on the coupling of an FFA biosensor with the expression of a tetracycline resistance gene, allowing the continuous enrichment of high-producing strains under tetracycline selective conditions. As a result, the approach achieved a threefold increase in FFA and tyrosine production [58].
Development of biosensors for lignin-derived monomers
Owing to the distinct genetic configurations and applicability of genetically encoded biosensors, these systems constitute ideal technologies for lignocellulose valorization. To date, numerous biosensors have been developed to sense aromatic monomers, including phenolic compounds obtained through direct lignin depolymerisation and other potential intermediates relevant to biomass valorization (Table 1). The variety of developed aromatic monomer biosensors provide the opportunity to facilitate the regulation, evolution, and selection of highly productive biocatalysts, pathways, and strains useful in lignocellulose degradation and valorization through diverse biosensor-based applications.
Development and optimization of biosensors based on available regulatory systems
The majority of the aromatic-responsive biosensors developed to date are based on the adaptation of characterized regulatory elements from aromatic degradation pathways and tolerance systems from soil microorganisms. These elements predominately consist of aromatic-responsive transcription factors and their cognate promoters/operators [59–61], and have been prominently derived from Pseudomonas spp., Acinetobacter spp. and the well-characterized lignin-degrading Sphingobium sp. strain SYK-6. Other species have also been identified as efficient lignin-degrading microorganisms, such as Rhodococcus jostii RHA1 [60, 62, 63] that could thus constitute an ideal host and source of responsive elements for the development of biosensors. The discovery and characterization of these aromatic-responsive elements have been partly accomplished by different strategies involving gene regulation in response to the target aromatic. For instance, Sana et al. [64] utilized an RNA-seq analysis of E. coli cultures treated with vanillin to identify upregulated genes. The identified PyeiW promoter upstream from these genes was thus used to construct a vanillin biosensor by placing it upstream of a reporter gene.
Once the responsive genetic elements have been identified, fine-tuning is often required to obtain optimal performance. Different strategies have been utilized in the design and optimization of biosensors, predominantly targeting different genetic elements affecting the dose–response curve (Fig. 2). A common strategy has been based on the engineering of the cognate promoter sequences and operator sites that bind to the aromatic-responsive aTFs (Fig. 2) [38]. Kim et al. [65] initially described an optimized BTX-responsive biosensor built from the previously characterized xylose regulator (XylR) aTF from Pseudomonas spp., where the cognate promoter was first upstream from the luciferase reporter gene, and then was exchanged with an engineered promoter of higher strength that lacked certain XylR upstream activating sequences. The exchange resulted in increased fold change of more than 3000-fold in response to o-Xylene [65].
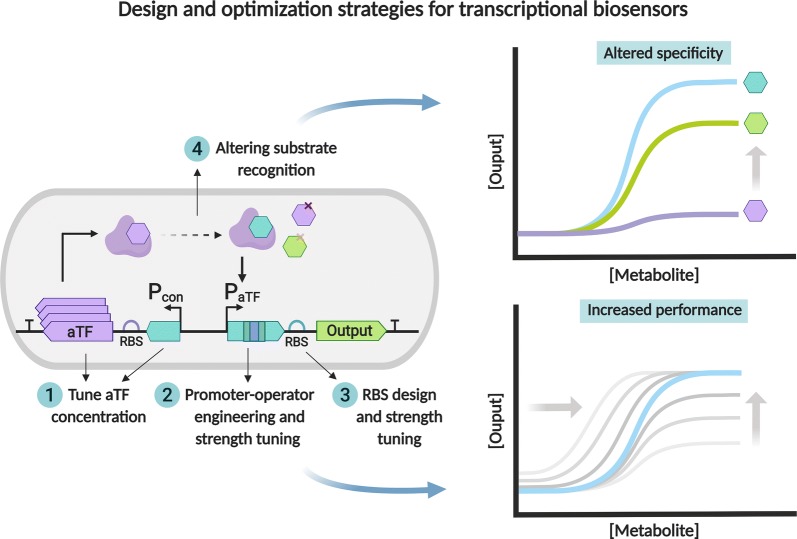
Fig. 2.
Design and optimization strategies for transcriptional biosensors. The performance of transcriptional biosensors can be altered by fine-tuning the expression levels of the allosteric transcription factor (1), engineering the cognate promoter and operator sequences (2), and/or ribosome binding site (3). The biosensor substrate recognition scope can be altered by engineering the specificity of the transcription factor (4)
The binding of the aTF to its cognate promoter sequence can also be rationally modulated by developing chimeric promoter–operator sequences. Machado and Dixon [66] developed a biosensor based on the FerC repressor, a MarR-type regulator protein and its corresponding transcription factor binding (operator) site from Sphingobium sp. SYK-6. The reporter output signal was amplified by engineering chimeric promoter–operators, composed of the native operator sites and phage-derived promoter elements. The developed biosensor afforded detection of various hydroxycinnamic acids, including ferulic acid and p-coumaric acid with more than 20-fold response [66]. A similar hybrid promoter engineering approach was also recently used to develop promoters of higher strength induced by phenolic compounds present in lignocellulosic hydrolysates, including vanillin and vanillic acid. The strategy was based on the reconstruction of the spacer sequence of the native phenolic-inducible PemrR promoter from Escherichia coli, which resulted in up to 4.6- and 9.2-fold increase in output signal in response to vanillin and vanillic acid, respectively [67]. These engineered promoters and their corresponding aTFs will thus be useful towards the development of biosensors for lignin valorization applications.
An alternative approach employed to enhance biosensor performance has been to adjust the intracellular concentration of the aromatic-responsive aTF (Fig. 2). Ho et al. [68] utilized a library of constitutive promoters of varying strength to regulate the expression of the vanillin responsive EmrR aTF. Increased aTF concentration with optimal promoter strength permitted an improved performance in response to vanillin and a broader specificity range towards other aromatics [68]. In addition, tuning and development of aromatic transcriptional biosensors by ribosome-binding site (RBS) selection and modification have been used to improve biosensor performance (Fig. 2). Siedler et al. [48] developed a p-coumaric acid-responsive biosensor based on the PadR aTF. The initial biosensor design resulted in the overexpression of PadR, which led to the inhibition of its cognate promoter. Modifying the downstream RBS, reduced aTF expression and consequently optimized fold change (130-fold) in response to p-coumaric acid was achieved [48].
Biosensors have also been optimized by targeting both the promoter–operator sequences and their RBS. A protocatechuic acid (PCA) biosensor was developed in E. coli based on the characterized PcaU aTF from Acinetobacter, an IcIR-type TF involved in the catabolism of PCA [69]. Combined engineering of the PcaU-responsive promoter sequence and the downstream RBS afforded an increase in reporter output signal and improved fold change (up to 14-fold) in response to PCA [69]. In follow-up studies, the PcaU-derived biosensor was transferred into Pseudomonas putida, achieved by promoter engineering and protein directed evolution. These approaches allowed the optimization of aTF–DNA interaction and aTF dimerization, which ultimately afforded a > 12-fold output signal at 30 mM PCA concentration [70].
Xue et al. [71] developed and optimized a set of biosensors based on an array of previously identified transcription factors and cognate promoter sequences from Pseudomonas (DmpR, HbpR, NahR, and XylS) [71]. These biosensors jointly responded to 20 different aromatic compounds, some of which can be derived as products of lignocellulose degradation and/or valorization, including several benzoate, salicylate, and phenol derivatives [71]. In these biosensors, the expression of the aTF was fine-tuned using a constitutive promoter library, and expression of reporter gene was optimized by RBS engineering [71].
These examples provide an indication of the promising array of lignocellulose-derived substrates and intermediates that can be detected using available genetically encoded biosensors (Table 1). The development of additional and novel substrate specificities would, however, be useful to increase the scope of lignocellulose-derived products that can be detected and to, therefore, enhance the utility of genetically encoded biosensors.
Design and selection of aromatic-responsive biosensors with altered substrate recognition
Protein engineering and directed evolution techniques constitute valuable approaches to extend the effector recognition capabilities of transcriptional biosensors, which is useful for the detection of molecules, where natural biosensor systems have yet to be discovered or characterized (Fig. 2) [42, 72]. In particular, unique aromatic-responsive parts with novel metabolite specificities have been engineered through random and site-directed mutagenesis techniques or computational design. Computational methods offer investigation of wider mutagenic arrays, supporting the de novo design of customized specificities within particular protein structures, such as ligand and DNA-binding domains [42, 73]. De los Santos et al. [74] utilized a rational, computational structure-based homology modelling approach to develop a library of QacR variants, a transcriptional repressor from the TetR family, which resulted in the selection of variants with altered specificity towards the monomeric aromatic vanillin [74]. Jha et al. [75] engineered novel specificity towards PCA through the Rosetta homology-based redesign of the ligand-binding domain of the PobR transcription factor, which is natively responsive to 4-hydroxybenzoic acid (4HB). This modelling approach allowed for the in silico prediction of an aTF variant library from a homology model. FACS screening was then utilized to select improved clones, yielding a variant aTF with high transcriptional induction in response to PCA and increased performance with the native 4HB inducer [75]. More recently, the MarR-type repressor pcaV, which is natively responsive to PCA, was reengineered using a biosensor-based FACS selection method to alter its substrate specificity. The directed evolution approach resulted in the identification of a pcaV variant (Van2) that permitted the in vivo detection of vanillin and other aromatic aldehydes, and was used to construct a vanillin biosensor [76].
Application of biosensors in lignocellulose valorization
Functional metagenomic screening
Release of unmodified aromatic monomers from lignin following harsh chemical pre-treatment has proven challenging [77]. In contrast, bacterial lignolytic activities can afford the release of unmodified low molecular weight aromatics from lignin-rich waste streams. Several bacterial enzymes are known to degrade and transform lignocellulosic biomass into monomeric units [78]. Due to the heterogeneous and highly cross-linked nature of lignin, no enzymatic process has been demonstrated to achieve this conversion with high levels of efficiency. As such, further enzyme development and discovery would be a valuable step towards an integrated lignocellulose biorefinery.
High-throughput screens using genetically encoded biosensors allow for the detection and selection of novel biocatalysts from environmental metagenomic samples, which constitute a rich source of genetic information for the discovery of biocatalysts, transporters, and regulatory elements (Fig. 3a) [79, 80]. This approach was applied by Strachan et al. [79], where a predicted multicopper oxidase (CopA) was identified from a community-wide metagenomic scaffold library of bacterial DNA samples isolated from a coal bed, using a genetically encoded biosensor. The biosensor was based on the EmrR aTF and its cognate promoter, which were found to directly respond to mono-aromatic lignin degradation products, including vanillin and vanillic acid (Fig. 3a). The identified enzyme was then employed in the degradation of industrially processed lignin, demonstrating lignin transformation activity [79].
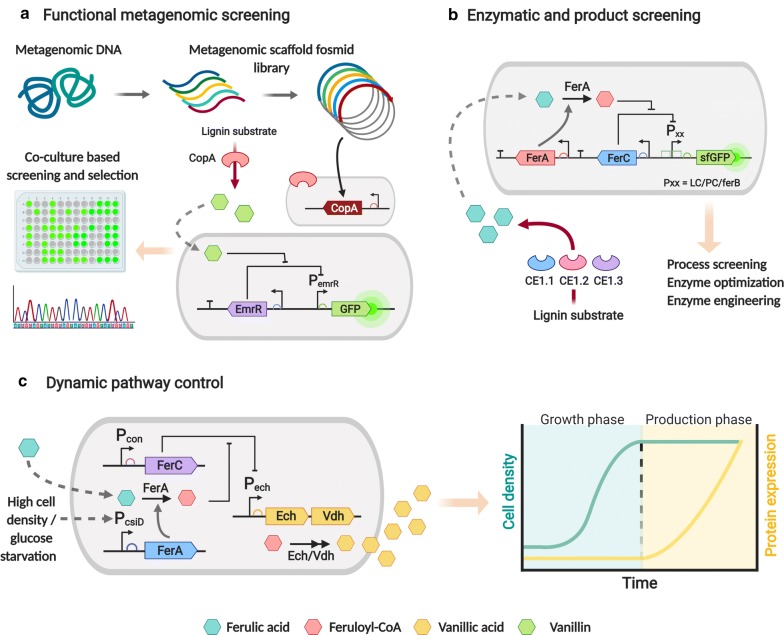
Fig. 3.
Genetically encoded biosensors and their applications in lignin valorization: a functional metagenomic screening method based on an EmrR-based biosensor for identifying lignin-degrading and transforming enzymes from metagenomic samples. A multicopper oxidase (CopA) enzyme capable of degrading lignin to vanillin and syringaldehyde was identified from a recovered metagenomic fosmid DNA library using this system [67]. b Transcriptional biosensors as screening methods to allow enzymatic characterization and evolution. A FerC-based biosensor was applied to identify optimal feruloyl esterase (CE1) activity by detecting ferulic acid release [69]. c Dynamic pathway regulation system for vanillic acid production. A ferulic acid biosensor was utilized to activate and regulate the biosynthetic pathway only when high cell density was reached and enough substrate was present, diminishing metabolic stress and increasing productivity [68]
When genetically encoded biosensors are integrated within other screening systems such as FACS, methods can be developed that offer a powerful system for metagenomic sample screening. One such system includes the substrate-induced gene expression (SIGEX) system, a high-throughput promoter-trap method can that can activate expression of reporter genes upon target aromatic induction, and subsequently uses FACS to isolate clones from metagenomic DNA libraries [81]. SIGEX has been employed to identify several catabolic genes and transcription factors that regulate reporter gene expression in response to aromatic compounds, including salicylate and several catechol-derived chemicals such as 4-chlorocatechol [82]. The SIGEX system was also employed by Meier et al. [83], where several aromatic hydrocarbon degrading enzymes and their respective regulators were discovered from aromatic hydrocarbon-contaminated soil samples. The technique subsequently aided the discovery and mapping of operons involved in aromatic hydrocarbon metabolism [83].
Enzyme, pathway, and strain production screening
Determination of substrate and product concentrations is a major bottleneck for enzymatic optimization and directed evolution processes (see “The screening bottleneck—metabolic pathway engineering and optimization” section). Genetically encoded biosensors have been applied as screening tools to identify active and optimal enzymes and biocatalytic pathways regarding lignin-degrading and transforming enzymes (Fig. 3b) [33, 44, 84]. For instance, the developed FerC repressor-based biosensor described earlier was applied to screen for optimal feruloyl esterase enzyme (CE1) activity against different biomass sources by detecting ferulic acid release (Fig. 3b) [66]. Moreover, the optimized PCA-responsive biosensor based on the pcaU transcription factor from Acinetobacter sp. ADP1 was used in the enzymatic screening and selection of dehydroshikimate dehydratase enzymes (AsbF) with PCA-producing activity. Using FACS, individual biosensor cells carrying the desired AsbF could be isolated from an inactive AsbF population [69]. van Sint Fiet et al. [85] utilized a mutated variant of the aTF NahR from Pseudomonas to detect benzoic acid and 2-hydroxybenzoic acid and regulate expression of lacZ and the tetracycline resistance gene tetA. The system was successful in the selection of biocatalytically active cells from an enzymatic library encoding a benzaldehyde dehydrogenase (xylC), as these active mutants allowed benzaldehyde-dependant cell growth [85].
Applying biosensors for enzymatic screening can also facilitate the optimization of productivity of aromatic-transforming enzymes and pathways. Recently, Jha et al. [86] developed a 4HB biosensor in P. putida to optimize enzymatic activity and alleviate product inhibition in chorismate pyruvate-lyase (UbiC). Along with the engineered 4HB biosensor–enzyme coupling, benzoic acid was utilized as a product surrogate that inhibited UbiC activity. As a result, they generated an UbiC variant library with reduced product inhibition and isolated a clone that eventually allowed a twofold yield increase in the bioproduction of muconic acid from glucose. Moreover, the p-coumaric acid-responsive biosensor developed by Siedler et al. [48] based on the PadR repressor was used in the screening of yeast p-coumaric acid production after co-encapsulation within microfluidic droplets. This system was subsequently used in the screening and sorting of high-producing strains from a yeast producer mutant library, indicating that the system could be used to enrich p-coumaric acid producing strains [48].
Dynamic pathway control
Several biosensors have also been developed and integrated into biosynthetic pathways for metabolic engineering applications [41, 87]. Balanced utilization of cellular resources for enhanced chemical production from lignocellulose has been accomplished through the use of biosensors that dynamically regulate intermediate metabolic fluxes and synthetic pathways. Dynamic regulation permits cells to adapt to environmental changes and regulate timely synthesis at each step in a pathway and improve resource utilisation efficiency and productivity (Fig. 3c).
Lo et al. [88] achieved the decoupling of cell growth from metabolite production by dynamically upregulating enzymatic pathways upon nutrient depletion using a biosensor module. The two-layered system allowed separation of the growth and production phases, and selective utilization of target substrates once the growth nutrients had been depleted (Fig. 3c). By applying this system with a p-coumaric acid sensing module, metabolic stress was lowered by twofold, while bioproductivity was improved fivefold [88].
Furthermore, an autoregulatory lignin valorization system was developed to diminish aromatic toxicity and circumvent required addition of exogenous inducers [89]. The system was based on the ADH7 vanillin-inducible promoter, which dynamically regulated a catechol biosynthesis pathway only in the presence of vanillin, alleviating metabolic burden and reducing toxicity.
Other responsive elements available
Sugar responsive elements
Besides utilizing genetically encoded biosensors to respond to lignin degradation products and intermediates, it is important to develop and implement regulatory systems that can respond to sugars derived from lignocellulose to establish a fully integrated biorefinery process. Hemicelluloses in lignocellulose comprise a very heterogeneous group of polysaccharides, including xylans, arabinoxylans, and cellulose [12]. The hydrolysis and saccharification of these polysaccharides result in the release of sugars, such as xylose, arabinose, and glucose. Constructing biosensors that respond to sugars other than cellulose could help to discover and improve the activity of hydrolysing enzymes towards the development of sugar-fermentative strains for biomass-derived high value production and in second-generation bioethanol production. In addition, these sugars constitute an ideal source of inducers that could be utilized in the dynamic regulation of phenolic and aromatic degrading enzymes for their upgrading into high value chemicals and pharmaceuticals.
To date, several regulation systems have been developed that focus on the detection of these lignocellulose-derived sugars (Table 1). Transcriptional biosensors have been constructed for xylose, cellobiose, and d-arabinose based on the naturally occurring XylR and CelR and on an engineered AraC transcription factor, respectively [90–92]. The latter was constructed by engineering the AraC aTF through random mutagenesis and FACS-based screening to select novel ligand-binding specificity towards d-arabinose [90]. FRET biosensors have also been designed and optimized for the detection of d-arabinose, glucose, and glutamate, using the AraF, MglB, and YbeJ bacterial periplasmic-binding proteins (PBPs), respectively [93, 94]. Moreover, Ribeiro et al. [95] developed a xylose biosensor based on the xylose transcriptional activator XylR and the xylose-binding protein (XBP). The integration of XBP increased the activity of the PxylF promoter by 40%, resulting in increased GFP signal output in the presence of xylose.
These sugar-inducible regulatory systems constitute valuable mechanisms that could be utilized for the simultaneous degradation and biotransformation of hemicellulose and lignin fractions while maintaining reduced product inhibition and increased productivity. Furthermore, the scope of sugars and their analogues that can be detected using biosensors should also be extended. Kelly et al. [96] developed a biosensor based on the l-rhamnose inducible rhaBAD expression system to identify alternative orthogonal effectors. Among these, they found that the sugar analogue l-mannose provided the greatest response fold change [96].
Conclusion and perspectives
Owing to their importance in the chemical, energy, and pharmaceutical industries, lignocellulosic feedstocks continue to be an attractive source of aromatic compounds. To this end, biomass-derived bioproduction has been modest, achieving low productivity when compared to commercial-scale demands [21]. Effective lignocellulose-derived high value chemical production will likely be accomplished through the discovery and directed evolution of novel biosynthetic pathways and biocatalysts. Emerging biosensor tools for the rapid evolution and high-throughput screening of optimal strains are powerful tools to increase the rate of development and optimisation of such pathways [43]. Biosensor-aided discovery of novel lignin-degrading enzymes will continue to be a crucial tool for the effective detection of lignin degradation capabilities against a wide range of metabolites.
The application of these biosensors is particularly interesting for sampling environmental metagenomes [79], as these constitute a rich and untapped source of enzymatic diversity from uncultivable microbes [97]. Moreover, despite their importance in aromatic transport and metabolic balance, many aromatic transporters and their functions are still unknown or poorly characterized. Biosensors could thus be applied in the identification, validation, and evolution of new substrate-specific transporter proteins [25, 98, 99].
Furthermore, bioproduction of aromatics will need to be incorporated with regulated synthetic pathways to achieve fully integrated dynamic control and production of metabolites to achieve an overall improved bioproduction process. Dynamic pathway control mediated by sugar or aromatic-based regulatory systems will provide a beneficial bioproduction approach by allowing on-demand expression of lignocellulose-transforming biosynthetic pathways when they are most required. To accelerate these biosensor-mediated processes, the current selection of products and intermediate metabolites that can be detected will need to be expanded. Development of novel genetically encoded biosensors will be aided by developments in protein engineering techniques and computational design to engineer novel ligand specificities [42]. In addition, the knowledge gained from advancements in this area can also translate into the development of biosensors with specificity towards complex aromatic molecules, including flavonoids and vitamins [100].
Going forward, the increased adoption of evolutionary strategies is anticipated to aid chemical bioproduction and metabolic engineering developments. Overcoming these current productivity limitations will offer great opportunities for the use of genetically encoded biosensors in pathway engineering for lignocellulose valorization. Advancements in these areas will help diversify the array of products that can be obtained from lignin and provide efficient biocatalysts for their integration into economically viable biorefinery processes.
Acknowledgements
Not applicable.
Abbreviations
- BTX
benzene, toluene, xylene
- FACS
fluorescence-activated cell sorting
- HK
histidine kinases
- PBP
periplasmic-binding protein
- RR
response regulator
- SIGEX
substrate-induced gene expression
Authors’ contributions
GA and ND wrote and reviewed the main manuscript text, and GA prepared figures. Both authors read and approved the final manuscript.
Funding
N.D. is supported by the BBSRC David Phillips Fellowship (BB/K014773/1), and the BBSRC grant (BB/P01738X/1). G.A. is supported by the BBSRC DTP grant (Grant BB/M011208/1).
Availability of data and materials
Not applicable.
Ethics approval and consent to participate
Not applicable.
Consent for publication
All authors consent to publication.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Bugg TDH, Rahmanpour R. Enzymatic conversion of lignin into renewable chemicals. Curr Opin Chem Biol. 2015;29:10–17. doi: 10.1016/j.cbpa.2015.06.009. [DOI] [PubMed] [Google Scholar]
- 2.Sindhu R, Binod P, Pandey A. Biological pretreatment of lignocellulosic biomass—an overview. Bioresour Technol. 2016;199:76–82. doi: 10.1016/j.biortech.2015.08.030. [DOI] [PubMed] [Google Scholar]
- 3.Kumar R, Singh S, Singh OV. Bioconversion of lignocellulosic biomass: biochemical and molecular perspectives. J Ind Microbiol Biotechnol. 2008;35:377–391. doi: 10.1007/s10295-008-0327-8. [DOI] [PubMed] [Google Scholar]
- 4.Zakzeski J, Bruijnincx PCA, Jongerius AL, Weckhuysen BM. The catalytic valorization of lignin for the production of renewable chemicals. Chem Rev. 2010;110:3552–3599. doi: 10.1021/cr900354u. [DOI] [PubMed] [Google Scholar]
- 5.Biomass Research & Development Board. The billion-ton bioeconomy initiative: challenges and opportunities. 2016. https://biomassboard.gov/pdfs/the_bioeconomy_initiative.pdf. Accessed 27 June 2019.
- 6.Biomass Research & Development Board. The bioeconomy initiative: implementation framework. 2018. https://biomassboard.gov/pdfs/Bioeconomy_Initiative_Implementation_Framework_FINAL.pdf. Accessed 27 June 2019.
- 7.Langholtz M, Stokes B, Eaton L. 2016 Billion-ton report: advancing domestic resources for a thriving bioeconomy. U.S. Department of Energy. 2016. 10.2172/1271651. http://energy.gov/eere/bioenergy/2016-billion-ton-report. Accessed 27 June 2019.
- 8.European Commission. A sustainable bioeconomy for Europe: strengthening the connection between economy, society and the environment. 2018. 10.2777/478385. http://europa.eu. Accessed 27 June 2019.
- 9.Perlack RD. Biomass as feedstock for a bioenergy and bioproducts industry: the technical feasibility of a billion-ton annual supply. 2005. 10.2172/885984http://www.osti.gov/servlets/purl/885984-pH60fJ/. Accessed 27 June 2019.
- 10.Holladay JE, White JF, Bozell JJ, Johnson D. Top value-added chemicals from biomass—volume II—Results of screening for potential candidates from biorefinery lignin. Pacific Northwest National Laboratory. 2007. 10.2172/921839http://www.osti.gov/servlets/purl/921839/. Accessed 27 June 2019.
- 11.Galkin MV, Samec JSM. Lignin valorization through catalytic lignocellulose fractionation: a fundamental platform for the future biorefinery. Chemsuschem. 2016;9:1544–1558. doi: 10.1002/cssc.201600237. [DOI] [PubMed] [Google Scholar]
- 12.Scheller HV, Ulvskov P. Hemicelluloses. Annu Rev Plant Biol. 2010;61:263–289. doi: 10.1146/annurev-arplant-042809-112315. [DOI] [PubMed] [Google Scholar]
- 13.Vennestrøm PNR, Osmundsen CM, Christensen CH, Taarning E. Beyond petrochemicals: the renewable chemicals industry. Angew Chemie Int Ed. 2011;50:10502–10509. doi: 10.1002/anie.201102117. [DOI] [PubMed] [Google Scholar]
- 14.Tuck CO, Perez E, Horvath IT, Sheldon RA, Poliakoff M. Valorization of biomass: deriving more value from waste. Science. 2012;337:695–699. doi: 10.1126/science.1218930. [DOI] [PubMed] [Google Scholar]
- 15.Beckham GT, Johnson CW, Karp EM, Salvachúa D, Vardon DR. Opportunities and challenges in biological lignin valorization. Curr Opin Biotechnol. 2016;42:40–53. doi: 10.1016/j.copbio.2016.02.030. [DOI] [PubMed] [Google Scholar]
- 16.Strassberger Z, Tanase S, Rothenberg G. The pros and cons of lignin valorisation in an integrated biorefinery. RSC Adv. 2014;4:25310–25318. doi: 10.1039/C4RA04747H. [DOI] [Google Scholar]
- 17.Bruijnincx PCA, Rinaldi R, Weckhuysen BM. Unlocking the potential of a sleeping giant: lignins as sustainable raw materials for renewable fuels, chemicals and materials. Green Chem. 2015;17:4860–4861. doi: 10.1039/c5gc90055g. [DOI] [Google Scholar]
- 18.Abdelaziz OY, Brink DP, Prothmann J, Ravi K, Sun M, García-Hidalgo J, Sandahl M, Hulteberg CP, Turner C, Lidén G, Gorwa-Grauslund MF. Biological valorization of low molecular weight lignin. Biotechnol Adv. 2016;34:1318–1346. doi: 10.1016/j.biotechadv.2016.10.001. [DOI] [PubMed] [Google Scholar]
- 19.Esposito D, Antonietti M. Redefining biorefinery: the search for unconventional building blocks for materials. Chem Soc Rev. 2015;44:5821–5835. doi: 10.1039/C4CS00368C. [DOI] [PubMed] [Google Scholar]
- 20.Lopes AM, Ferreira Filho EX, Moreira LRS. An update on enzymatic cocktails for lignocellulose breakdown. J Appl Microbiol. 2018;125:632–645. doi: 10.1111/jam.13923. [DOI] [PubMed] [Google Scholar]
- 21.Machas M, Kurgan G, Jha AK, Flores A, Schneider A, Coyle S, Varman AM, Wang X, Nielsen DR. Emerging tools, enabling technologies, and future opportunities for the bioproduction of aromatic chemicals. J Chem Technol Biotechnol. 2018;94:38–52. doi: 10.1002/jctb.5762. [DOI] [Google Scholar]
- 22.Wang Y, Chen S, Yu O. Metabolic engineering of flavonoids in plants and microorganisms. Appl Microbiol Biotechnol. 2011;91:949–956. doi: 10.1007/s00253-011-3449-2. [DOI] [PubMed] [Google Scholar]
- 23.Katsuyama Y, Funa N, Miyahisa I, Horinouchi S. Synthesis of unnatural flavonoids and stilbenes by exploiting the plant biosynthetic pathway in Escherichia coli. Chem Biol. 2007;14:613–621. doi: 10.1016/j.chembiol.2007.05.004. [DOI] [PubMed] [Google Scholar]
- 24.Yang SM, Shim GY, Kim BG, Ahn JH. Biological synthesis of coumarins in Escherichia coli. Microb Cell Fact. 2015 doi: 10.1186/s12934-015-0248-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kell DB, Swainston N, Pir P, Oliver SG. Membrane transporter engineering in industrial biotechnology and whole cell biocatalysis. Trends Biotechnol. 2015;33:237–246. doi: 10.1016/j.tibtech.2015.02.001. [DOI] [PubMed] [Google Scholar]
- 26.Nielsen J, Keasling JD. Engineering cellular metabolism. Cell. 2016;164:1185–1197. doi: 10.1016/j.cell.2016.02.004. [DOI] [PubMed] [Google Scholar]
- 27.Rogers JK, Taylor ND, Church GM. Biosensor-based engineering of biosynthetic pathways. Curr Opin Biotechnol. 2016;42:84–91. doi: 10.1016/j.copbio.2016.03.005. [DOI] [PubMed] [Google Scholar]
- 28.Holtz WJ, Keasling JD. Engineering static and dynamic control of synthetic pathways. Cell. 2010;140:19–23. doi: 10.1016/j.cell.2009.12.029. [DOI] [PubMed] [Google Scholar]
- 29.Raman S, Rogers JK, Taylor ND, Church GM. Evolution-guided optimization of biosynthetic pathways. Proc Natl Acad Sci. 2014;111:17803–17808. doi: 10.1073/pnas.1409523111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Rogers JK, Church GM. Genetically encoded sensors enable real-time observation of metabolite production. Proc Natl Acad Sci. 2016;113:2388–2393. doi: 10.1073/pnas.1600375113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Dietrich JA, McKee AE, Keasling JD. High-throughput metabolic engineering: advances in small-molecule screening and selection. Annu Rev Biochem. 2010;79:563–590. doi: 10.1146/annurev-biochem-062608-095938. [DOI] [PubMed] [Google Scholar]
- 32.Liu D, Evans T, Zhang F. Applications and advances of metabolite biosensors for metabolic engineering. Metab Eng. 2015;31:15–22. doi: 10.1016/j.ymben.2015.06.008. [DOI] [PubMed] [Google Scholar]
- 33.De Paepe B, Peters G, Coussement P, Maertens J, De Mey M. Tailor-made transcriptional biosensors for optimizing microbial cell factories. J Ind Microbiol Biotechnol. 2017;44:623–645. doi: 10.1007/s10295-016-1862-3. [DOI] [PubMed] [Google Scholar]
- 34.Michener JK, Thodey K, Liang JC, Smolke CD. Applications of genetically-encoded biosensors for the construction and control of biosynthetic pathways. Metab Eng. 2012;14:212–222. doi: 10.1016/j.ymben.2011.09.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Rogers JK, Guzman CD, Taylor ND, Raman S, Anderson K, Church GM. Synthetic biosensors for precise gene control and real-time monitoring of metabolites. Nucleic Acids Res. 2015;43:7648–7660. doi: 10.1093/nar/gkv616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Shi S, Ang EL, Zhao H. In vivo biosensors: mechanisms, development, and applications. J Ind Microbiol Biotechnol. 2018;45:491–516. doi: 10.1007/s10295-018-2004-x. [DOI] [PubMed] [Google Scholar]
- 37.Ravikumar S, Baylon MG, Park SJ, Choi J. Engineered microbial biosensors based on bacterial two-component systems as synthetic biotechnology platforms in bioremediation and biorefinery. Microb Cell Fact. 2017;16:62. doi: 10.1186/s12934-017-0675-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lim HG, Jang S, Jang S, Seo SW, Jung GY. Design and optimization of genetically encoded biosensors for high-throughput screening of chemicals. Curr Opin Biotechnol. 2018;54:18–25. doi: 10.1016/j.copbio.2018.01.011. [DOI] [PubMed] [Google Scholar]
- 39.Liu Y, Liu Y, Wang M. Design, optimization and application of small molecule biosensor in metabolic engineering. Front Microbiol. 2017;8:2012. doi: 10.3389/fmicb.2017.02012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Koch M, Pandi A, Borkowski O, Cardoso Batista A, Faulon J. Custom-made transcriptional biosensors for metabolic engineering. Curr Opin Biotechnol. 2019;59:78–84. doi: 10.1016/j.copbio.2019.02.016. [DOI] [PubMed] [Google Scholar]
- 41.Mahr R, Frunzke J. Transcription factor-based biosensors in biotechnology: current state and future prospects. Appl Microbiol Biotechnol. 2016;100:79–90. doi: 10.1007/s00253-015-7090-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Taylor ND, Garruss AS, Moretti R, Chan S, Arbing MA, Cascio D, Rogers JK, Isaacs FJ, Kosuri S, Baker D, Fields S, Church GM, Raman S. Engineering an allosteric transcription factor to respond to new ligands. Nat Methods. 2016;13:177–183. doi: 10.1038/nmeth.3696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Binder S, Schendzielorz G, Stäbler N, Krumbach K, Hoffmann K, Bott M, Eggeling L. A high-throughput approach to identify genomic variants of bacterial metabolite producers at the single-cell level. Genome Biol. 2012;13:R40. doi: 10.1186/gb-2012-13-5-r40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Kwon KK, Lee D-H, Kim SJ, Choi S-L, Rha E, Yeom S-J, Subhadra B, Lee J, Jeong KJ, Lee S-G. Evolution of enzymes with new specificity by high-throughput screening using DmpR-based genetic circuits and multiple flow cytometry rounds. Sci Rep. 2018;8:2659. doi: 10.1038/s41598-018-20943-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Schendzielorz G, Dippong M, Grünberger A, Kohlheyer D, Yoshida A, Binder S, Nishiyama C, Nishiyama M, Bott M, Eggeling L. Taking control over control: use of product sensing in single cells to remove flux control at key enzymes in biosynthesis pathways. ACS Synth Biol. 2014;3:21–29. doi: 10.1021/sb400059y. [DOI] [PubMed] [Google Scholar]
- 46.Tang S-Y, Cirino PC. Design and application of a mevalonate-responsive regulatory protein. Angew Chemie Int Ed. 2011;50:1084–1086. doi: 10.1002/anie.201006083. [DOI] [PubMed] [Google Scholar]
- 47.Tang S-Y, Qian S, Akinterinwa O, Frei CS, Gredell JA, Cirino PC. Screening for enhanced triacetic acid lactone production by recombinant Escherichia coli expressing a designed triacetic acid lactone reporter. J Am Chem Soc. 2013;135:10099–10103. doi: 10.1021/ja402654z. [DOI] [PubMed] [Google Scholar]
- 48.Siedler S, Khatri NK, Zsohár A, Kjærbølling I, Vogt M, Hammar P, Nielsen CF, Marienhagen J, Sommer MOA, Joensson HN. Development of a bacterial biosensor for rapid screening of yeast p-coumaric acid production. ACS Synth Biol. 2017;6:1860–1869. doi: 10.1021/acssynbio.7b00009. [DOI] [PubMed] [Google Scholar]
- 49.Zhang F, Ouellet M, Batth TS, Adams PD, Petzold CJ, Mukhopadhyay A, Keasling JD. Enhancing fatty acid production by the expression of the regulatory transcription factor FadR. Metab Eng. 2012;14:653–660. doi: 10.1016/j.ymben.2012.08.009. [DOI] [PubMed] [Google Scholar]
- 50.Xu P, Li L, Zhang F, Stephanopoulos G, Koffas M. Improving fatty acids production by engineering dynamic pathway regulation and metabolic control. Proc Natl Acad Sci. 2014;111:11299–11304. doi: 10.1073/pnas.1406401111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Johnson AO, Gonzalez-Villanueva M, Wong L, Steinbüchel A, Tee KL, Xu P, Wong TS. Design and application of genetically-encoded malonyl-CoA biosensors for metabolic engineering of microbial cell factories. Metab Eng. 2017;44:253–264. doi: 10.1016/j.ymben.2017.10.011. [DOI] [PubMed] [Google Scholar]
- 52.Williams TC, Pretorius IS, Paulsen IT. Synthetic evolution of metabolic productivity using biosensors. Trends Biotechnol. 2016;34:371–381. doi: 10.1016/j.tibtech.2016.02.002. [DOI] [PubMed] [Google Scholar]
- 53.Chou HH, Keasling JD. Programming adaptive control to evolve increased metabolite production. Nat Commun. 2013 doi: 10.1038/ncomms3595. [DOI] [PubMed] [Google Scholar]
- 54.Mahr R, Gätgens C, Gätgens J, Polen T, Kalinowski J, Frunzke J. Biosensor-driven adaptive laboratory evolution of l-valine production in Corynebacterium glutamicum. Metab Eng. 2015;32:184–194. doi: 10.1016/j.ymben.2015.09.017. [DOI] [PubMed] [Google Scholar]
- 55.Leavitt JM, Wagner JM, Tu CC, Tong A, Liu Y, Alper HS. Biosensor-enabled directed evolution to improve muconic acid production in Saccharomyces cerevisiae. Biotechnol J. 2017;12:1600687. doi: 10.1002/biot.201600687. [DOI] [PubMed] [Google Scholar]
- 56.Wang T, Dunlop MJ. Controlling and exploiting cell-to-cell variation in metabolic engineering. Curr Opin Biotechnol. 2019;57:10–16. doi: 10.1016/j.copbio.2018.08.013. [DOI] [PubMed] [Google Scholar]
- 57.Lv Y, Qian S, Du G, Chen J, Zhou J, Xu P. Coupling feedback genetic circuits with growth phenotype for dynamic population control and intelligent bioproduction. Metab Eng. 2019;54:109–116. doi: 10.1016/j.ymben.2019.03.009. [DOI] [PubMed] [Google Scholar]
- 58.Xiao Y, Bowen CH, Liu D, Zhang F. Exploiting nongenetic cell-to-cell variation for enhanced biosynthesis. Nat Chem Biol. 2016;12:339–344. doi: 10.1038/nchembio.2046. [DOI] [PubMed] [Google Scholar]
- 59.Tropel D, van der Meer JR. Bacterial transcriptional regulators for degradation pathways of aromatic compounds. Microbiol Mol Biol Rev. 2004;68:474–500. doi: 10.1128/MMBR.68.3.474-500.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Kamimura N, Takahashi K, Mori K, Araki T, Fujita M, Higuchi Y, Masai E. Bacterial catabolism of lignin-derived aromatics: new findings in a recent decade: update on bacterial lignin catabolism. Environ Microbiol Rep. 2017;9:679–705. doi: 10.1111/1758-2229.12597. [DOI] [PubMed] [Google Scholar]
- 61.Díaz E, Prieto MA. Bacterial promoters triggering biodegradation of aromatic pollutants. Curr Opin Biotechnol. 2000;11:467–475. doi: 10.1016/s0958-1669(00)00126-9. [DOI] [PubMed] [Google Scholar]
- 62.Ahmad M, Roberts JN, Hardiman EM, Singh R, Eltis LD, Bugg TDH. Identification of DypB from Rhodococcus jostii RHA1 as a lignin peroxidase. Biochemistry. 2011;50:5096–5107. doi: 10.1021/bi101892z. [DOI] [PubMed] [Google Scholar]
- 63.Roberts JN, Singh R, Grigg JC, Murphy MEP, Bugg TDH, Eltis LD. Characterization of dye-decolorizing peroxidases from Rhodococcus jostii RHA1. Biochemistry. 2011;50:5108–5119. doi: 10.1021/bi200427h. [DOI] [PubMed] [Google Scholar]
- 64.Sana B, Chia KHB, Raghavan SS, Ramalingam B, Nagarajan N, Seayad J, Ghadessy FJ. Development of a genetically programed vanillin-sensing bacterium for high-throughput screening of lignin-degrading enzyme libraries. Biotechnol Biofuels. 2017;10:32. doi: 10.1186/s13068-017-0720-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Kim MN, Park HH, Lim WK, Shin HJ. Construction and comparison of Escherichia coli whole-cell biosensors capable of detecting aromatic compounds. J Microbiol Methods. 2005;60:235–245. doi: 10.1016/j.mimet.2004.09.018. [DOI] [PubMed] [Google Scholar]
- 66.Machado LFM, Dixon N. Development and substrate specificity screening of an in vivo biosensor for the detection of biomass derived aromatic chemical building blocks. Chem Commun. 2016;52:11402–11405. doi: 10.1039/C6CC04559F. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Varman AM, Follenfant R, Liu F, Davis RW, Lin YK, Singh S. Hybrid phenolic-inducible promoters towards construction of self-inducible systems for microbial lignin valorization. Biotechnol Biofuels. 2018;11:1–13. doi: 10.1186/s13068-018-1179-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Ho JCH, Pawar SV, Hallam SJ, Yadav VG. An improved whole-cell biosensor for the discovery of lignin-transforming enzymes in functional metagenomic screens. ACS Synth Biol. 2018;7:392–398. doi: 10.1021/acssynbio.7b00412. [DOI] [PubMed] [Google Scholar]
- 69.Jha RK, Kern TL, Fox DT, M Strauss CE. Engineering an Acinetobacter regulon for biosensing and high-throughput enzyme screening in E. coli via flow cytometry. Nucleic Acids Res. 2014;42:8150–8160. doi: 10.1093/nar/gku444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Jha RK, Bingen JM, Johnson CW, Kern TL, Khanna P, Trettel DS, Strauss CEM, Beckham GT, Dale T. A protocatechuate biosensor for Pseudomonas putida KT2440 via promoter and protein evolution. Metab Eng Commun. 2018;6:33–38. doi: 10.1016/j.meteno.2018.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Xue H, Shi H, Yu Z, He S, Liu S, Hou Y, Pan X, Wang H, Zheng P, Cui C, Viets H, Liang J, Zhang Y, Chen S, Zhang HM, Ouyang Q. Design, construction, and characterization of a set of biosensors for aromatic compounds. ACS Synth Biol. 2014;3:1011–1014. doi: 10.1021/sb500023f. [DOI] [PubMed] [Google Scholar]
- 72.Ellefson JW, Ledbetter MP, Ellington AD. Directed evolution of a synthetic phylogeny of programmable Trp repressors article. Nat Chem Biol. 2018;14:361–367. doi: 10.1038/s41589-018-0006-7. [DOI] [PubMed] [Google Scholar]
- 73.Raman S, Taylor N, Genuth N, Fields S, Church GM. Engineering allostery. Trends Genet. 2014;30:521–528. doi: 10.1016/j.tig.2014.09.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.de los Santos ELC, Meyerowitz JT, Mayo SL, Murray RM. Engineering transcriptional regulator effector specificity using computational design and in vitro rapid prototyping: developing a vanillin sensor. ACS Synth Biol. 2016;5:287–295. doi: 10.1021/acssynbio.5b00090. [DOI] [PubMed] [Google Scholar]
- 75.Jha RK, Chakraborti S, Kern TL, Fox DT, Strauss CEM. Rosetta comparative modeling for library design: engineering alternative inducer specificity in a transcription factor. Proteins Struct Funct Bioinform. 2015;83:1327–1340. doi: 10.1002/prot.24828. [DOI] [PubMed] [Google Scholar]
- 76.Machado L, Currin A, Dixon N. Directed evolution of the PcaV allosteric transcription factor to generate a biosensor for aromatic aldehydes. bioRxiv. 2019 doi: 10.1101/689232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Ragauskas AJ, Beckham GT, Biddy MJ, Chandra R, Chen F, Davis MF, Davison BH, Dixon RA, Gilna P, Keller M, Langan P, Naskar AK, Saddler JN, Tschaplinski TJ, Tuskan GA, Wyman CE. Lignin valorization: improving lignin processing in the biorefinery. Science. 2014;344:1246843. doi: 10.1126/science.1246843. [DOI] [PubMed] [Google Scholar]
- 78.Brown ME, Chang MC. Exploring bacterial lignin degradation. Curr Opin Chem Biol. 2014;19:1–7. doi: 10.1016/J.CBPA.2013.11.015. [DOI] [PubMed] [Google Scholar]
- 79.Strachan CR, Singh R, VanInsberghe D, Ievdokymenko K, Budwill K, Mohn WW, Eltis LD, Hallam SJ. Metagenomic scaffolds enable combinatorial lignin transformation. Proc Natl Acad Sci. 2014;111:10143–10148. doi: 10.1073/pnas.1401631111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Uchiyama T, Miyazaki K. Product-induced gene expression, a product-responsive reporter assay used to screen metagenomic libraries for enzyme-encoding genes. Appl Environ Microbiol. 2010;76:7029–7035. doi: 10.1128/AEM.00464-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Uchiyama T, Abe T, Ikemura T, Watanabe K. Substrate-induced gene-expression screening of environmental metagenome libraries for isolation of catabolic genes. Nat Biotechnol. 2005;23:88–93. doi: 10.1038/nbt1048. [DOI] [PubMed] [Google Scholar]
- 82.Uchiyama T, Miyazaki K. Metagenomic screening for aromatic compound-responsive transcriptional regulators. PLoS ONE. 2013;8:e75795. doi: 10.1371/journal.pone.0075795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Meier MJ, Paterson ES, Lambert IB. Use of substrate-induced gene expression in metagenomic analysis of an aromatic hydrocarbon-contaminated soil. Appl Environ Microbiol. 2016;82:897–909. doi: 10.1128/AEM.03306-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Choi SL, Rha E, Lee SJ, Kim H, Kwon K, Jeong YS, Rhee YH, Song JJ, Kim HS, Lee SG. Toward a generalized and high-throughput enzyme screening system based on artificial genetic circuits. ACS Synth Biol. 2014;3:163–171. doi: 10.1021/sb400112u. [DOI] [PubMed] [Google Scholar]
- 85.van Sint Fiet S, van Beilen JB, Witholt B. Selection of biocatalysts for chemical synthesis. Proc Natl Acad Sci. 2006;103:1693–1698. doi: 10.1073/pnas.0504733102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Jha RK, Narayanan N, Pandey N, Bingen JM, Kern TL, Johnson CW, Strauss CEM, Beckham GT, Hennelly SP, Dale T. Sensor-enabled alleviation of product inhibition in chorismate pyruvate-lyase. ACS Synth Biol. 2019;8:775–786. doi: 10.1021/acssynbio.8b00465. [DOI] [PubMed] [Google Scholar]
- 87.Zhang J, Jensen MK, Keasling JD. Development of biosensors and their application in metabolic engineering. Curr Opin Chem Biol. 2015;28:1–8. doi: 10.1016/j.cbpa.2015.05.013. [DOI] [PubMed] [Google Scholar]
- 88.Lo T-M, Chng SH, Teo WS, Cho H-S, Chang MW. A two-layer gene circuit for decoupling cell growth from metabolite production. Cell Syst. 2016;3:133–143. doi: 10.1016/j.cels.2016.07.012. [DOI] [PubMed] [Google Scholar]
- 89.Wu W, Liu F, Singh S. Toward engineering E. coli with an autoregulatory system for lignin valorization. Proc Natl Acad Sci. 2018;115:2970–2975. doi: 10.1073/pnas.1720129115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Tang SY, Fazelinia H, Cirino PC. AraC regulatory protein mutants with altered effector specificity. J Am Chem Soc. 2008;130:5267–5271. doi: 10.1021/ja7109053. [DOI] [PubMed] [Google Scholar]
- 91.Teo WS, Chang MW. Bacterial XylRs and synthetic promoters function as genetically encoded xylose biosensors in Saccharomyces cerevisiae. Biotechnol J. 2015;10:315–322. doi: 10.1002/biot.201400159. [DOI] [PubMed] [Google Scholar]
- 92.Kwon KK, Yeom S-J, Lee D-H, Jeong KJ, Lee S-G. Development of a novel cellulase biosensor that detects crystalline cellulose hydrolysis using a transcriptional regulator. Biochem Biophys Res Commun. 2018;495:1328–1334. doi: 10.1016/j.bbrc.2017.11.157. [DOI] [PubMed] [Google Scholar]
- 93.Deuschle K, Okumoto S, Fehr M, Looger LL, Kozhukh L, Frommer WB. Construction and optimization of a family of genetically encoded metabolite sensors by semirational protein engineering. Protein Sci. 2005;14:2304–2314. doi: 10.1110/ps.051508105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Kaper T, Lager I, Looger LL, Chermak D, Frommer WB. Fluorescence resonance energy transfer sensors for quantitative monitoring of pentose and disaccharide accumulation in bacteria. Biotechnol Biofuels. 2008;1:11. doi: 10.1186/1754-6834-1-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Ribeiro LF, Bressan F, Furtado GP, Meireles F, Ward RJ. d-Xylose detection in Escherichia coli by a xylose binding protein-dependent response. J Biotechnol. 2013;168:440–445. doi: 10.1016/j.jbiotec.2013.10.019. [DOI] [PubMed] [Google Scholar]
- 96.Kelly CL, Liu Z, Yoshihara A, Jenkinson SF, Wormald MR, Otero J, Estévez A, Kato A, Marqvorsen MHS, Fleet GWJ, Estévez RJ, Izumori K, Heap JT. Synthetic chemical inducers and genetic decoupling enable orthogonal control of the rhaBAD promoter. ACS Synth Biol. 2016;5:1136–1145. doi: 10.1021/acssynbio.6b00030. [DOI] [PubMed] [Google Scholar]
- 97.Hug LA, Baker BJ, Anantharaman K, Brown CT, Probst AJ, Castelle CJ, Butterfield CN, Hernsdorf AW, Amano Y, Ise K, Suzuki Y, Dudek N, Relman DA, Finstad KM, Amundson R, Thomas BC, Banfield JF. A new view of the tree of life. Nat Microbiol. 2016;1:16048. doi: 10.1038/nmicrobiol.2016.48. [DOI] [PubMed] [Google Scholar]
- 98.Bali AP, Genee HJ, Sommer MOA. Directed evolution of membrane transport using synthetic selections. ACS Synth Biol. 2018;7:789–793. doi: 10.1021/acssynbio.7b00407. [DOI] [PubMed] [Google Scholar]
- 99.Genee HJ, Bali AP, Petersen SD, Siedler S, Bonde MT, Gronenberg LS, Kristensen M, Harrison SJ, Sommer MOA. Functional mining of transporters using synthetic selections. Nat Chem Biol. 2016;12:1015–1022. doi: 10.1038/nchembio.2189. [DOI] [PubMed] [Google Scholar]
- 100.Siedler S, Stahlhut SG, Malla S, Maury JÔ, Neves AR. Novel biosensors based on flavonoid-responsive transcriptional regulators introduced into Escherichia coli. Metab Eng. 2014;21:2–8. doi: 10.1016/j.ymben.2013.10.011. [DOI] [PubMed] [Google Scholar]
- 101.Meyer AJ, Segall-Shapiro TH, Glassey E, Zhang J, Voigt CA. Escherichia coli “Marionette” strains with 12 highly optimized small-molecule sensors. Nat Chem Biol. 2019;15:196–204. doi: 10.1038/s41589-018-0168-3. [DOI] [PubMed] [Google Scholar]
- 102.Frazão CR, Maton V, François JM, Walther T. Development of a metabolite sensor for high-throughput detection of aldehydes in Escherichia coli. Front Bioeng Biotechnol. 2018;6:118. doi: 10.3389/fbioe.2018.00118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Kunjapur AM, Prather KLJ. Development of a vanillate biosensor for the vanillin biosynthesis pathway in E. coli. ACS Synth Biol. 2019;8:1958–1967. doi: 10.1021/acssynbio.9b00071. [DOI] [PubMed] [Google Scholar]
- 104.DeLorenzo D, Moon TS. Construction of genetic logic gates based on the T7 RNA polymerase expression system in Rhodococcus opacus PD630. ACS Synth Biol. 2019 doi: 10.1021/acssynbio.9b00213. [DOI] [PubMed] [Google Scholar]
- 105.Fiorentino G, Ronca R, Bartolucci S. A novel E. coli biosensor for detecting aromatic aldehydes based on a responsive inducible archaeal promoter fused to the green fluorescent protein. Appl Microbiol Biotechnol. 2009;82:67–77. doi: 10.1007/s00253-008-1771-0. [DOI] [PubMed] [Google Scholar]
- 106.Qian S, Li Y, Cirino PC. Biosensor-guided improvements in salicylate production by recombinant Escherichia coli. Microb Cell Fact. 2019 doi: 10.1186/s12934-019-1069-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Ribeiro LF, Tullman J, Nicholes N, Silva SRB, Vieira DS, Ostermeier M, Ward RJ. A xylose-stimulated xylanase–xylose binding protein chimera created by random nonhomologous recombination. Biotechnol Biofuels. 2016;9:119. doi: 10.1186/s13068-016-0529-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.