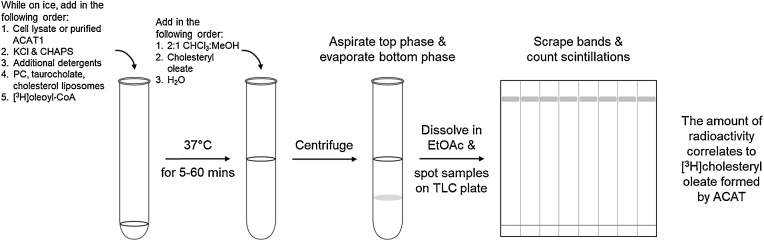
Graphical abstract
Method name: Dissociation of ACAT1 subunits by non-ionic detergents
Keywords: ACAT1, SOAT1, Two-fold dimer, MBOAT, Quaternary structure, Liposomes, Transmembrane, Cholesterol, Enzyme activity, Detergent
Abstract
Acyl-CoA:cholestereol acyltransferase 1 (ACAT1) is a two-fold dimer (homotetramer) and has two distinct dimerization domains. One domain is in an alpha-helical rich region near the cytoplasmic N-terminus. The other is proposed to be near the C-terminus where multiple transmembrane domains promote hydrophobic interactions between two ACAT1 subunits. The truncation of the ACAT1 N-terminal dimerization domain, Δ1-65, creates a dimer which is fully enzymatically active. It is currently not known how the C-terminal dimerization domain contributes to ACAT1 enzymatic activity. Here we describe a simple method that dissociates ACAT1 dimers through the addition of the non-ionic detergents Triton X-100 or octyl glucoside which disrupt the C-terminal dimerization domain. We also document the protocols for a method to exchange Triton X-100 with CHAPS to restore C-terminal dimerization of the ACAT1 protein, and an optimized liposomal assay to assess ACAT enzymatic activity.
-
•
This method can be applied to dissociate ACAT1 subunits by using Triton X-100 or octyl glucoside.
-
•
ACAT1 dimerization can be restored by exchanging Triton X-100 with CHAPS.
-
•
The liposomal ACAT activity assay conditions have been optimized.
Specification Table
| Subject Area: | Biochemistry, Genetics and Molecular Biology |
| More specific subject area: | Analytical Biochemistry, Enzymology |
| Method name: | Dissociation of ACAT1 subunits by non-ionic detergents |
| Name and reference of original method: | B. Neumann, C.C.Y. Chang, T.Y. Chang, Triton X-100 or octyl glucoside inactivates acyl-CoA:cholesterol acyltransferase 1 by dissociating it from a two-fold dimer to a two-fold monomer, Archives of biochemistry and biophysics 671 (2019) 103–110. |
| C. Yu, J. Chen, S. Lin, J. Liu, C.C. Chang, T.Y. Chang, Human acyl-CoA:cholesterol acyltransferase-1 is a homotetrameric enzyme in intact cells and in vitro, The Journal of biological chemistry 274(51) (1999) 36139–45. | |
| C.C. Chang, C.Y. Lee, E.T. Chang, J.C. Cruz, M.C. Levesque, T.Y. Chang, Recombinant acyl-CoA:cholesterol acyltransferase-1 (ACAT-1) purified to essential homogeneity utilizes cholesterol in mixed micelles or in vesicles in a highly cooperative manner, The Journal of biological chemistry 273(52) (1998) 35132–41. | |
| Resource availability: | [3H]oleoyl-CoA, water bath, Organomation N-EVAP Nitrogen Evaporator, Uniplate Analtech Silica gel HL, TLC developing tank, Ecoscint O, scintillation counter, peroxide-free Triton X-100, Beckman L8-70 M ultracentrifuge, SW 60 Ti rotor, Thermo Fisher Scientific HisPur Ni-NTA resin, Corning Costar Spin-X 0.22 μm centrifuge tube filters, USA Scientific 1.5 ml low adhesion microcentrifuge tubes, New Brunswick Scientific Gyrotory Shaker Model G2, Thermo Scientific GelCode Blue Stain Reagent, Pall Corporation 10 K Microsep Advance Centrifugal Device with Omega Membrane |
Method details
Background
Acyl-CoA:cholestereol acyltransferase 1 (ACAT1) is an endoplasmic reticulum resident membrane protein, where it regulates cellular cholesterol homeostasis. In order to study membrane proteins, detergents are required to solubilize cell membranes. Differences in detergents play a crucial role in the assessment of ACAT activity. When solubilized by the non-ionic detergent Triton X-100, ACAT activity is drastically reduced [1,2]. The anionic detergent deoxycholate (DOC) provides a modest recovery of ACAT activity [1]. The use of the zwitterionic detergent 3-[(3-Cholamidopropyl) dimethylammonio]-1-propanesulfonate (CHAPS) with phosphatidylcholine, taurocholate, and cholesterol particles provides the greatest recovery of ACAT enzymatic activity [1]. This finding led to the creation of a reliable ACAT enzymatic activity assay, known as the “mixed micelles assay” [1]. This assay was recently modified to include optimized concentrations of KCl and CHAPS to achieve higher ACAT enzymatic activity [3]. Under the latter condition, the CHAPS, phosphatidylcholine, taurocholate, and cholesterol particles were found to exist mainly as liposomes [3].
ACAT1 is a homotetrameric enzyme which contains two dimerization domains. The N-terminal dimerization domain is not required for proper ACAT1 enzymatic activity [4], whereas, the role of the C-terminal dimerization domain in enzymatic activity is unknown. The following methods can be used to assess how various detergents affect ACAT1 enzymatic activity and its oligomeric state.
Reagents
Peroxide-free Triton X-100, octyl glucoside, KCl, CHAPS, Tris, HCl, egg phosphatidylcholine, sodium taurocholate, cholesterol, EDTA, [3H]oleoyl-CoA, chloroform, methanol, ethyl acetate, fatty acid-free BSA, petroleum ether, diethyl ether, acetic acid, Ecoscint O, sucrose
Solutions
11.2 mM egg PC/9.3 mM taurocholate/1.8 mM cholesterol mixed micelle/liposome mixture, 2 M KCl, 162.6 mM (10%) CHAPS, 1 M Tris−HCl pH 7.8, 30 mg/ml fatty acid-free BSA in 0.25 M Tris−HCl pH 7.8, 75% sucrose
Materials
13 × 100 mm disposable glass tubes, Analtech Uniplate Silica gel HL #P46911, Thermo Fisher Scientific HisPur Ni-NTA resin, Corning Costar Spin-X0.22 μm centrifuge tube filters #8160, USA Scientific 1.5 ml low adhesion microcentrifuge tubes #1415-2600, Beckman 11 × 60 mm Ultra-Clear centrifuge tubes #344062, VWR 96-well PCR Plate #89218-296, VWR PCR sealing film #82018-844, Thermo Scientific GelCode Blue Stain Reagent #24590, Pall Corporation 10 K Microsep Advance Centrifugal Device with Omega Membrane #MCP010C41
Instrumentation/apparatus
Water bath, Organomation N-EVAP Nitrogen Evaporator, TLC developing tank, Scintillation counter, Beckman L8-70M ultracentrifuge, SW 60 Ti rotor, New Brunswick Scientific Gyrotory Shaker Model G2
Protocols
Optimized liposomal ACAT activity assay
The liposomal ACAT activity assay is useful for the determination of ACAT enzymatic activity in cell lysates or of purified ACAT1. This assay is designed so that enzyme activity depends on the amount of cholesterol present in the liposome, rather than the amount of cholesterol present in the cell lysates. Adapted from [1,3] and depicted in Fig. 1.
-
1
Place 13 × 100 mm disposable glass tubes into a tube rack on ice.
-
2
Add 5 μl of cell lysate in 1 M KCl, 40.7 mM (2.5%) CHAPS, 50 mM Tris−HCl pH 7.8 or purified ACAT1 in 0.5 M KCl, 8.1 mM (0.5%) CHAPS, 50 mM Tris−HCl pH 7.8, 10% glycerol to each assay tube.
-
3
Add 5 μl of sample buffer to two additional tubes to determine background levels of [3H].
-
4
Add enough stock solutions of 2 M KCl and 162.6 mM (10%) CHAPS to have a final concentration of 100 mM KCl and 5 mM (0.31%) CHAPS.
-
5
Optional step: add additional detergents (i.e. peroxide-free Triton X-100, octyl glucoside, etc.) in water to the assay tube to assess how they affect ACAT enzymatic activity.
-
6
Add 60 μl of 11.2 mM egg PC, 9.3 mM taurocholate, 1.8 mM cholesterol preformed mixed micelle/liposome mixture [1,3].
-
7
Briefly vortex tubes to mix contents and place on ice for 5 min.
-
8
To one tube at a time, add 10 nmoles of [3H]oleoyl-CoA in 30 mg/ml fatty acid-free BSA in 0.25 M Tris−HCl pH 7.8 [5] and gently vortex to mix.
-
9
Immediately start the assay by placing the tube into a shaking 37 °C water bath for the desired amount of assay time. Normally, between 5–60 min.
-
10
Halt the assay with 3 ml of 2:1 chloroform:methanol solution in the same order tubes were placed into the 37 °C water bath.
-
11
Add 5 μl of 10 mg/ml cholesteryl oleate in chloroform.
-
12
Partition the solvents by adding 1 ml of dH2O and gently vortex for 10 s.
-
13
Centrifuge at 27 × g at room temperature for 10 min.
-
14
Aspirate the top aqueous phase into a radioactive disposal container (the phases will be separated by a thin, white protein layer).
-
15
Blow dry the bottom phase in an Organomation N-EVAP Nitrogen Evaporator.
-
16
The dried lipid samples can be stored at −20 °C for up to 4 weeks.
-
17
Dissolve the dried lipid samples in 100 μl of ethyl acetate.
-
18
Spot the samples onto a Uniplate Analtech Silica gel HL 250 μm thin layer chromatography (TLC) plate on a hot plate.
-
19
Let TLC plate equilibrate to room temperature before placing into TLC development tank.
-
20
Add 36 ml petroleum ether, 4 ml diethyl ether, and 0.4 ml acetic acid (90:10:1) into TLC development tank, mix well, and cover the top of the tank. Let solvent equilibrate for up to 20 min.
-
21
Place TLC plate into the tank and cover. Let the solvent proceed up the plate until the solvent front is halfway or higher from the bottom of the plate.
-
22
Remove the TLC plate from the tank and let dry at room temperature for at least an hour.
-
23
Scrape the cholesteryl oleate band (Rf = 0.9, visible under light as a gray band near the solvent front) into scintillation vials.
-
24
Add 3 ml of Ecoscint O to each vial.
-
25
Cap, invert to mix, place into scintillation counter, and count.
-
26
Convert amount of [3H] present in cholesteryl oleate to nanomoles based on specific radioactivity of [3H]oleoyl-CoA.
Fig. 1.
Schematic diagram of the optimized liposomal ACAT activity assay.
Small-scale exchange of detergents via Ni-NTA bound-ACAT1
Here we describe a method for the exchange of CHAPS with peroxide-free Triton X-100 and vice versa with Ni-NTA bound-ACAT1. This protocol can be used for the exchange of other detergents and scaled up for larger preparations. Adapted from [3].
-
1
Cut the tips of 200 μl pipette tips when working with small volumes of Thermo Fisher Scientific HisPur Ni-NTA resin.
-
2
All centrifugation steps are carried out at 10,000 × g.
-
3
Wash 50 μl of Thermo Fisher Scientific HisPur Ni-NTA resin (100 μl of 50/50 slurry) with 0.5 ml (10 column volumes (CVs)) of dH2O in a Corning Costar Spin-X0.22 μm centrifuge tube filter.
-
4
Supercharge the resin with 0.5 ml (10 CVs) of 100 mM NiSO4.
-
5
Wash with 0.5 ml (10 CVs) of dH2O.
-
6
Wash with 0.5 ml (10 CVs) of 0.5 M KCl, 8.1 mM (0.5%) CHAPS, 50 mM Tris−HCl pH 7.8, 10% glycerol.
-
7
Mix the resin with 50 μl (1 CV) of purified polyhistidine-tagged ACAT1 and transfer to a pre-chilled low adhesion microcentrifuge tube.
-
8
Mix lightly on a New Brunswick Scientific Gyrotory Shaker Model G2 for 1 h at 4 °C.
-
9
Transfer the resin slurry back into the Corning Costar Spin-X0.22 μm centrifuge tube filter, pellet resin, and collect supernatant (flow through) at 4 °C.
-
10
Wash the resin with 0.5 ml (10 CVs) of 1 M KCl, 8.1 mM CHAPS, 50 mM Tris−HCl pH 7.8 at 4 °C.
-
11
Resuspend the resin in 50 μl (1 CV) of 1 M KCl, 8.1 mM CHAPS, 50 mM Tris−HCl pH 7.8 (total volume 100 μl) at 4 °C.
-
12
Aliquot 25 μl of the resin slurry into a pre-chilled low adhesion microcentrifuge tube to measure ACAT activity before exchange with Triton X-100.
-
13
Pellet and wash the resin with 0.375 ml (10 CVs) of 1 M KCl, 1.7 mM peroxide-free Triton X-100, 50 mM Tris−HCl pH 7.8 at 4 °C.
-
14
Resuspend the resin in 37.5 μl (1 CV) of 1 M KCl, 1.7 mM peroxide-free Triton X-100, 50 mM Tris−HCl pH 7.8 (total volume 75 μl) into a pre-chilled low adhesion microcentrifuge tube.
-
15
After the exchange of CHAPS with Triton X-100, the Ni-NTA bound-ACAT1 may be used for other experimentation with ACAT1 bound to the resin or eluted from the resin with 100 mM EDTA added to 1 M KCl, 1.7 mM peroxide-free Triton X-100, 50 mM Tris−HCl pH 7.8.
-
16
Aliquot 25 μl of the resin slurry into a pre-chilled low adhesion microcentrifuge tube to measure ACAT activity after exchange with Triton X-100.
-
17
Use a new or wash the previously used Corning Costar Spin-X0.22 μm centrifuge tube filter with excessive amounts of dH2O.
-
18
Wash the remaining 25 μl of resin with 0.5 ml (20 CVs) of 1 M KCl, 8.1 mM CHAPS, 50 mM Tris−HCl pH 7.8 at 4 °C.
-
19
Resuspend the resin in 25 μl (1 CV) of 1 M KCl, 8.1 mM CHAPS, 50 mM Tris−HCl pH 7.8 (total volume 50 μl) into a pre-chilled low adhesion microcentrifuge tube.
-
20
Perform previously described liposomal ACAT activity assay, in duplicate, with 10 μl of the resin slurry per assay tube.
Sucrose density gradient centrifugation of ACAT1 with various detergents
By using sucrose density gradient centrifugation, we can monitor ACAT1 oligomeric state when exposed to various detergents by comparing its fractionation profile to protein standards. Adapted from [3,6].
-
1Mix 75% w/v sucrose solution into 30, 25, 20, 15, 10, and 5% sucrose with desired amount of KCl, Tris−HCl, and detergent.
-
a1 M KCl, 50 mM Tris−HCl pH 7.8, 0.5% or 8.1 mM CHAPS
-
b1 M KCl, 50 mM Tris−HCl pH 7.8, 0.1% or 1.7 mM peroxide-free Triton X-100
-
c1 M KCl, 50 mM Tris−HCl pH 7.8, 20 mM octyl glucoside
-
a
-
2
Form sucrose gradients in Beckman 11 × 60 mm Ultra-Clear centrifuge tubes #344062 with (from bottom to top) 30, 25, 20, 15, 10, and 5% sucrose. Each layer is 0.6 ml with a total volume of 3.6 ml.
-
3
Freeze gradients in dry-ice/ethanol bath before the addition of each new layer.
-
4
Once completed, the gradients can be thawed at 4 °C to use immediately or can be stored at −80 °C.
-
5
Apply 50 μg of each protein standard (Table 1) to a sucrose gradient to estimate the size of ACAT1 by sucrose density gradient centrifugation. Store proteins as a mix in 0.5 M KCl, 50 mM Tris−HCl pH 7.8, 0.5% CHAPS, 10% glycerol at −80 °C.
-
6Dilute 100 μl of purified ACAT1 or protein standards to a final volume of 250 μl in desired concentrations of KCl, Tris−HCl, and detergent. Additions listed below:
-
a100 μl 2 M KCl, 7.5 μl 1 M Tris−HCl pH 7.8, 7.5 μl 10% CHAPS, 35 μl dH2O
-
b100 μl 2 M KCl, 7.5 μl 1 M Tris−HCl pH 7.8, 25 μl 1% peroxide-free Triton X-100, 17.5 μl dH2O
-
c100 μl 2 M KCl, 7.5 μl 1 M Tris−HCl pH 7.8, 11 μl 460 mM octyl glucoside, 31.5 μl dH2O
-
a
-
7
Centrifuge diluted ACAT1 and protein standards at 21,000 × g at 4 °C for 5 min. Collect the supernatants.
-
8
Apply 200 μl of ACAT1 or 200 μl of protein standards to the top of two tubes with identical gradients.
-
9
Centrifuge in a Beckman SW 60 Ti rotor in a Beckman model L8–70 M ultracentrifuge at 50,000 rpm for 10 h at 4 °C.
-
10
Collect 38 fractions at 100 μl/fraction into a VWR 96-well PCR plate, cover plate with a VWR PCR sealing film, and store at −80 °C.
-
11
For ACAT1 gradients, apply 48 μl of each fraction and 12 μl of 5X loading dye (10% SDS, 250 mM DTT, 50% glycerol, 0.25% bromphenol blue, 50 mM Tris−HCl pH 6.8) to a 10% SDS-PAGE. Transfer to PVDF membrane and visualize ACAT1 with rabbit polyclonal anti-ACAT1 DM102 antibodies or a suitable peptide epitope tag antibody.
-
12
For protein standard gradients, apply 48 μl of each fraction and 12 μl of 5X loading dye of each fraction to a 10% SDS-PAGE. Visualize with Thermo Fisher Scientific GelCode Blue or a similar Coomassie Blue stain.
-
13
Follow the previously mentioned liposomal ACAT activity assay protocol to monitor enzyme activity present in each fraction containing ACAT1 (as determined by Western blot).
Table 1.
Protein standards for sucrose density gradient centrifugation.
| Protein standard | Molecular weight (kDa) | Peak fractionation # | Relative ACAT1 size |
|---|---|---|---|
| BSA | 66.5 | 14 | 1x |
| Aldolase | 144 | 20 | 2x |
| β-Amylase | 210 | 25 | 4x |
| Catalase | 232 | 28 | 4x |
| Thyroglobulin | 660 | 37 | 8x |
Detergent exchange of sucrose density gradient fractions
This method is designed to exchange the non-ionic detergents from specific sucrose density gradient centrifugation fractions with CHAPS to determine if ACAT1 homotetramer can be recovered after exposure to non-ionic detergents. Adapted from [3].
-
1
Reduce non-specific adsorption of the 10 K Pall Microsep Advance Centrifugal Device by soaking the sample reservoir in 10% glycerol overnight at room temperature.
-
2
Remove 10% glycerol by centrifugation at 3200 × g for 10 min.
-
3
Fill the sample reservoir with dH2O and spin at 3200 × g for 10 min. Repeat once.
-
4
Pool fractions #10–20 from a 1 M KCl, 50 mM Tris−HCl pH 7.8, 1.7 mM peroxide-free Triton X-100 or a 20 mM octyl glucoside ACAT1 sucrose density gradient into the conditioned 10 K Pall Microsep Advance Centrifugal Device. Note total volume placed into sample reservoir.
-
5
Spin at 3200 × g at 4 °C for 10 min. or until volume is ∼250 μl.
-
6
Exchange non-ionic detergent with 1 ml of 1 M KCl, 50 mM Tris−HCl pH 7.8, 8.1 mM CHAPS. Spin at 3200 × g at 4 °C for 10 min. or until volume is ∼250 μl. Repeat × 3 until final volume is ∼250 μl.
-
7
Once samples are exchanged, use the previously mentioned sucrose density gradient centrifugation protocol with a 1 M KCl, 50 mM Tris−HCl pH 7.8, 8.1 mM CHAPS sucrose gradient to monitor ACAT1 size.
Acknowledgement
This work was supported by NIH grants RO1 AG037609 and AG063544 to T.Y. Chang and Catherine Chang.
Contributor Information
Bryan Neumann, Email: Bryan.C.Neumann.GR@Dartmouth.edu.
Catherine C.Y. Chang, Email: Catherine.Chang@Dartmouth.edu.
Ta-Yuan Chang, Email: Ta.Yuan.Chang@Dartmouth.edu.
References
- 1.Chang C.C., Lee C.Y., Chang E.T., Cruz J.C., Levesque M.C., Chang T.Y. Recombinant acyl-CoA:cholesterol acyltransferase-1 (ACAT-1) purified to essential homogeneity utilizes cholesterol in mixed micelles or in vesicles in a highly cooperative manner. J. Biol. Chem. 1998;273(52):35132–35141. doi: 10.1074/jbc.273.52.35132. [DOI] [PubMed] [Google Scholar]
- 2.Kaduce T.L., Schmidt R.W., Spector A.A. Acylcoenzyme A:cholesterol acyltransferase activity: solubilization and reconstitution in liposomes. Biochem. Biophys. Res. Commun. 1978;81(2):462–468. doi: 10.1016/0006-291x(78)91556-5. [DOI] [PubMed] [Google Scholar]
- 3.Neumann B., Chang C.C.Y., Chang T.Y. Triton X-100 or octyl glucoside inactivates acyl-CoA:cholesterol acyltransferase 1 by dissociating it from a two-fold dimer to a two-fold monomer. Arch. Biochem. Biophys. 2019;671:103–110. doi: 10.1016/j.abb.2019.06.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Yu C., Zhang Y., Lu X., Chen J., Chang C.C., Chang T.Y. Role of the N-terminal hydrophilic domain of acyl-coenzyme A:cholesterol acyltransferase 1 on the enzyme’s quaternary structure and catalytic efficiency. Biochemistry. 2002;41(11):3762–3769. doi: 10.1021/bi0120188. [DOI] [PubMed] [Google Scholar]
- 5.Bishop J.E., Hajra A.K. A method for the chemical synthesis of 14C-labeled fatty acyl coenzyme A’s of high specific activity. Anal. Biochem. 1980;106(2):344–350. doi: 10.1016/0003-2697(80)90531-x. [DOI] [PubMed] [Google Scholar]
- 6.Yu C., Chen J., Lin S., Liu J., Chang C.C., Chang T.Y. Human acyl-CoA:cholesterol acyltransferase-1 is a homotetrameric enzyme in intact cells and in vitro. J. Biol. Chem. 1999;274(51):36139–36145. doi: 10.1074/jbc.274.51.36139. [DOI] [PubMed] [Google Scholar]