Abstract
Preclinical imaging studies of posttraumatic epileptogenesis (PTE) have largely been proof-of-concept studies with limited animal numbers, and thus lack the statistical power for biomarker discovery. Epilepsy Bioinformatics Study for Antiepileptogenic Therapy (EpiBioS4Rx) is a pioneering multicenter trial investigating preclinical imaging biomarkers of PTE. EpiBios4Rx faced the issue of harmonizing the magnetic resonance imaging (MRI) procedures and imaging data metrics prior to its execution. We present here the harmonization process between three preclinical MRI facilities at the University of Eastern Finland (UEF), the University of Melbourne (Melbourne), and the University of California, Los Angeles (UCLA), and evaluate the uniformity of the obtained MRI data.
Adult, male rats underwent a lateral fluid percussion injury (FPI) and were followed by MRI 2 days, 9 days, 1 month, and 5 months post-injury. Ex vivo scans of fixed brains were conducted 7 months post-injury as an end point follow-up. Four MRI modalities were used: T2-weighted imaging, multi-gradient-echo imaging, diffusion-weighted imaging, and magnetization transfer imaging, and acquisition parameters for each modality were tailored to account for the different field strengths (4.7 T and 7 T) and different MR hardwares used at the three participating centers.
Pilot data collection resulted in comparable image quality across sites. In interim analysis (of data obtained by April 30, 2018), the within-site variation of the quantified signal properties was low, while some differences between sites remained. In T2-weighted images the signal-to-noise ratios were high at each site, being 35 at UEF, 48 at Melbourne, and 32 at UCLA (p < 0.05). The contrast-to-noise ratios were similar between the sites (9, 10, and 8, respectively). Magnetization transfer ratio maps had identical white matter/ gray matter contrast between the sites, with white matter showing 15% higher MTR than gray matter despite different absolute MTR values (MTR both in white and gray matter was 3% lower in Melbourne than at UEF, p < 0.05). Diffusion-weighting yielded different degrees of signal attenuation across sites, being 83% at UEF, 76% in Melbourne, and 80% at UCLA (p < 0.05). Fractional anisotropy values differed as well, being 0.81 at UEF, 0.73 in Melbourne, and 0.84 at UCLA (p < 0.05). The obtained values in sham animals showed low variation within each site and no change over time, suggesting high repeatability of the measurements. Quality control scans with phantoms demonstrated stable hardware performance over time. Timing of post-TBI scans was designed to target specific phases of the dynamic pathology, and the execution at different centers was highly accurate. Besides a few outliers, the 2-day scans were done within an hour from the target time point. At day 9, most animals were scanned within an hour from the target time point, and all but 2 outliers within 24 h from the target. The 1-month post-TBI scans were done within 31 ± 3 days. MRI procedures and animal physiology during scans were similar between the sites.
Taken together, the 10% inter-site difference in FA and 3% difference in MTR values should be included into analysis as a covariate or balanced out in post-processing in order to detect disease-related effects on brain structure at the same scale. However, for a MRI biomarker for post-traumatic epileptogenesis to have realistic chance of being successfully translated to validation in clinical trials, it would need to be a robust TBI-induced structural change which tolerates the inter-site methodological variability described here.
Keywords: Common data element, Diffusion tensor imaging, Magnetization transfer imaging, Multi-site harmonization, Post-traumatic epilepsy, Traumatic brain injury
1. Introduction
Traumatic brain injury (TBI) is induced by an external mechanical force to the brain, which triggers a myriad of impact-related and secondary pathologies (Toth et al., 2016; Pearn et al., 2017; Armstrong et al., 2016; Corrigan et al., 2016). One of the debilitating long-term consequences of TBI, posttraumatic epilepsy (PTE), accounts for up to 10–20% of symptomatic epilepsy in the general population, and the mechanisms for the development of epilepsy are still largely unknown (Herman, 2002; Frey, 2003; Pitkanen and Immonen, 2014). A TBI patient is diagnosed with PTE if she/he presents one unprovoked (late) seizure, that is, an unprovoked seizure after the first post-TBI week (Fisher et al., 2005). To date there are no antiepileptogenic clinical treatments available. Clinical and pre-clinical trials of anti-epileptogenic therapies would be greatly facilitated by the discovery of prognostic biomarkers with high sensitivity and specificity to identify subjects with high risk of developing PTE, enabling patient stratification and making clinical antiepileptogenesis trials practical and affordable (Pitkanen et al., 2018; Engel et al., 2013).
MRI-derived parameters hold great promise for biomarker discovery as MRI is non-invasive, readily available, and provides objective and quantitative indicators of the brain pathophysiology underlying posttraumatic epileptogenesis (van Vliet et al., 2018; Bertoglio et al., 2017; Immonen et al., 2013). However, due to the heterogeneity of TBIs and numerous subject-specific factors such as lifestyle and co-morbidities, biomarker discovery in humans is challenging. Rodent models of TBI provide a tool for rigorous mechanistic biomarker discovery in a time- and cost-sensitive manner (Pitkanen et al., 2018).
A major hurdle in identifying clinically relevant MRI-based biomarkers relates to a lack of standardized procedures in preclinical and clinical biomarker studies. Consequently, comparative and translational analyses of published data are challenging. Moreover, the hardware and pulse sequences used in preclinical studies are much more variable than those in clinical studies, increasing the challenges for harmonization of data acquisition and translation to clinic. The Epilepsy Bioinformatics Study for Antiepileptogenic Therapy (EpiBios4Rx) is a National Institute of Neurological Disorders and Stroke (NINDS) funded Centers-without-Walls study, which aims to find biomarkers and treatments to combat post-traumatic epileptogenesis (https://epibios.loni.usc.edu/). EpiBioS4Rx is a collaborative multi-center, international study conducted in the United States, Europe, and Australia. The preclinical component of the EpiBioS4Rx applies a rat model of PTE induced with lateral fluid-percussion injury (FPI) for biomarker discovery.
To conduct a statistically powered preclinical multi-center MRI biomarker discovery trial, the consortium partners have developed EpiBioS4Rx protocols for the harmonization of pre-clinical MRI methodologies. In the first stage, we generated common data elements and case report forms for all imaging procedures. In the second stage, we tailored the MRI data acquisition sequences and parameters at each center to harmonize the obtained signal between different hardware, software and magnetic field strengths. We did this for all MR modalities included in the study. Signal elements (image elements), that were required to be harmonized, included contrast, signal-to-noise ratio, and modality-specific variables such as fractional anisotropy in diffusion imaging, and magnetization transfer ratio in magnetization transfer imaging. In the third stage, MRI procedures, including timing of imaging relative to injury and the sequence of procedures were harmonized between the centers. Finally, when the MRI datasets have been generated, the data analysis will be harmonized. Herein we describe the optimized MRI acquisition protocols and evaluate the inter-center harmonization of the procedures and the data obtained at the three study sites.
2. Materials and methods
2.1. Microscopic view of the study design
Fig. 1 shows the study design of EpiBioS4Rx Project 1 MRI biomarker discovery. MRI follow-up was done 2 days, 9 days, 1 month and 5 months post-injury capturing the maximal post-injury edema (at day 2), the different phases of the secondary injury (from days 9–30), and the late chronic stage (5 months). After end-stage EEG phenotyping (to determine epileptic versus non-epileptic animals), rats were transcardially perfused, electrodes were removed with the head-set, and the fixed brains were scanned ex vivo.
Fig. 1. Study design.
Longitudinal in vivo MRI follow-up and concurrent blood sampling. The MRT paradigm included four modalities which were imaged at each time point to probe different pathologies developing during the first days to weeks to months after TBI, when the brain becomes epileptogenic. Electrode implantation and 4-wk continuous video-EEG for epilepsy phenotyping were conducted after the last 5-months scan. Ex vivo scans with the same modalities, but modified parameters were executed at the end point.
Four MRI modalities were used: T2-weighted (T2wt), multi-echo gradient echo (MGE), magnetization transfer imaging (MT), and diffusion-weighted imaging (DWI), including diffusion tensor imaging (DTI) and tractography approaches. These modalities were specifically chosen to probe the complex radiological presentations of the post-TBI brain: structural pathologies such as the characteristics, location, severity, and extent of structural lesion, and overall brain atrophy (T2-weighted MRI and MGE); hemorrhages, microbleeds, iron residues, and calcifications (susceptibility-weighted imaging (SWI), T2*); tissue water relaxivity (T2*) and diffusivity (DWI); grey and white matter microstructural alterations and demyelination (DWI, MTR) as well as structural connectivity (DWI tractography). All data were acquired at each time point during a 2-hour scanning session, thus allowing both co-registration between MR modalities and assessment of quantitative changes in individual rats over time.
The consortium includes 4 preclinical NMR facilities at the University of Eastern Finland (Site 1, UEF), University of Melbourne/Florey Institute (Site 2, Melbourne), University of California, Los Angeles (Site 3, UCLA), and Albert Einstein College of Medicine (Site 4, Einstein), hosting magnets of 7T, 4.7 T, 7T, and 9.4 T field strengths. Sites 1–3 participated in this first phase (Project 1). The hardware and software specifications of the site 4 were, however, considered in protocol planning in order to accomplish the future studies at four sites. Volume transmit RF-coils with surface receiver coils were utilized. Table 1 summarizes the hardware at each site.
Table 1.
Instrumentation at each site of the consortium.
| MRI Instrumentation |
||||
|---|---|---|---|---|
| UEF | Melbourne | UCLA | Einstein | |
| Magnet | 7 T/16cm | 4.7 T/33cm | 7 T/30cm | 9.4 T 31 cm |
| Console | Bruker Pharmascan | Bruker Biospec | Bruker Biospec | Agilent Direct Drive |
| Gradients | 600 m T/m risetime 150 μs | 440mT/m risetime 150 μs | 400 mT/m risetime ∼110 μs | 600 m T/m risetime 180 μs |
| Head Rf-coils | Actively decoupled volume transmitter and quadrature surface coil receiver (Bruker) | Actively decoupled volume transmitter and 4-channel surface coil receiver (Bruker) | Actively decoupled volume transmitter and single channel surface coil receiver | Actively decoupled volume transmitter (M2M) and 4-channel surface coil receiver (M2M) |
2.1.1. Animal model
Adult, male Spraque Dawley rats (weight 300–350 g) were used. Animals were housed individually (UEF and Melbourne) or in pairs (UCLA) in controlled environment (temperature 22 °C, humidity 40–70 %, 12-hour light cycle) with ab libitum access to pellet food and water. Lateral fluid percussion injury was induced as previously described by Kharatishvili and colleagues (Kharatishvili et al., 2006). Briefly, a craniotomy of 5 mm in diameter was drilled over the left cortex and fluid-percussion device was used produce fluid-mediated pressure pulse (2.2–3.1 at m) to induce moderate to severe TBI. Dura was left intact. Sham operated rats served as controls. A detailed description of the animal model used, anesthesia, post injury monitoring, and harmonization of animal handling across all sites is presented by Ekolle Ndode-Ekane et al. (2019, in this volume). The predicted number of animals to be included in the study is 192 (64 rats at each of the three sites). The interim analysis presented here has been performed using animals scanned by April 30, 2018 (39 at UEF, 35 in Melbourne, and 26 at UCLA). In addition, 8 sham-operated control rats at each site were scanned to evaluate MRI signal uniformity between the sites, signal variation in the combined cohort, and repeatability over time. Common data elements were recorded for each animal, and the intra-site and inter-site variation in the accuracy of timing of MRI scans were assessed.
At the UEF, all animal procedures were approved by the Animal Health and Welfare committee of the Regional State Administrative Agency and conducted in accordance with the guidelines set by the European Commission Directive 2010/63/EU (UEF). In Melbourne, procedures were approved by the Florey Animal Ethics Committee (ethics number 17–013 UM) and the procedures were performed in accordance with the guidelines of the Australian Code of Practice for the Care and Use of Animals for Scientific Purposes. At UCLA, procedures were approved by the University of California Los Angeles Chancellor’s Animal Research Committee and adhered to the Public Health Service Policy on Humane Care and Use of Laboratory Animals.
2.2. In vivo MRI methods
The four preclinical NMR facilities host magnets of different field strengths, different vendors, and different gradient strengths. MRI techniques were chosen based on their compatibility with each sites’ hardware (limiting factors e.g. the gradient performance), and acquisition parameters were tuned to produce uniform data across different field strengths, e.g., adjusting repetition time and echo time (see Tables 1 and 2).
Table 2. MR techniques, key parameters, and site-specific parameter adjustments (red) at each site involved in harmonization: UEF, Melbourne and UCLA.
Harmonization step 1 was to tailor the sequences executable by each MRI system and account for the different field strengths and gradient performances. Step 2 was to tailor the sequence parameters to unify signal to noise ratio and contrast between different field strengths/sites. Flip angles were calculated to match the Ernst angle, repetition times (TR) to match the T1 relaxation, and echo times (TE) to match the T2 relaxation between sites. Differences in parameters between 7 T and 4.7 T are indicated in red.
| UEF | Melbourne | UCLA | |
|---|---|---|---|
| multi-slice T2 weighted fast spin echo (FSE): edema, anatomy | |||
| Method | TurboRARE | TurboRARE | TurboRARE |
| TR/TEeff | 3400/45 ms | 3400/49ms | 3400/45 ms |
| echoes per excitation | 8 | 8 | 8 |
| resolution/slice thickness | 117 × 117 μm/0.8mm | 117 × 117 μm/0.8mm | 117 × 117 μm/0.8mm |
| 3D multi-echo gradient echo (MGE) 160μm3: SWI, T2* and anatomy | |||
| Method | MGE 3D | MGE 3D | MGE 3D |
| TR/TEmax/flip/SW | 66 ms/43 ms/16°/67kHz | 66 ms/47ms/18°/67kHz | 66 ms/43 ms/16°/67kHz |
| T2* - array: echoes | 13 | 14 | 13 |
| TEmin/ step /TEmax | 2.73/3.1/43 ms | 2.73/3.4/47ms | 2.73/3.1/43 ms |
| Dummys | 120 - > 5640 ms (5*T1) | 120- > 5640 ms (> 5*T1) | 120 - > 5640 ms (5*T1) |
| OVS pulses | 4 OVS | 4 OVS | 4 OVS |
| Fat sat | Yes | Yes | Yes |
| Diffusion-weighted imaging (DWI) 250μm3: DTI and tractography | |||
| Method | EPI 3D spin echo | EPI 3D spin echo | EPI 3D spin echo |
| Segments | 1 | 1 | 1 |
| TR/TE/SW | 1000/26/357kHz | 1000/28/357kHz | 1000/26/357kHz |
| B-val | 2800s/mm2 | 2800s/mm2 | 2800s/mm2 |
| grad duration /separation | 4.2/12ms | 5.0/12ms | 4.2/12ms |
| Amplitude (% of max) | 64.26% | 84.49% | 64.26% |
| Diff orient | 42 + 4 | 42 + 4 | 42 + 4 |
| Magnetization Transfer (MT) 250 μm3 : MTR | |||
| Method | FLASH 3D | FLASH 3D | FLASH 3D |
| TR/TE/flip/SW | 47/2.6/13°/67kHz | 47/2.6/15°/67kHz | 47/2.6/13°/67kHz |
| MT Pulse shape/Power | Gauss/10μT | Gauss/10μT | Gauss/10μT |
| MT Offset | −1500 Hz/5 ppm | −1000Hz/5ppm | −1500 Hz/5 ppm |
| Number of pulses | 3 | 3 | 3 |
| Pulse length | 12ms (228 Hz) | 12ms (228 Hz) | 12ms (228 Hz) |
2.2.1. MRI techniques
Shimming was done using 3D field map-based automated procedure (Mapshim on Bruker) over the entire brain volume, and water linewidth of the shimmed volume was recorded as a quality control.
T2wt multi-slice fast spin echo (RARE) images detecting edema were obtained for optimal lesion characterization (TR 3400 ms, effective TE 45 ms, RARE factor 8, min TE 11ms, 3 averages, field of view (FOV) 30 × 30 mm, 256 × 256 matrix, in-plane resolution 117 μm × 117 μm, 23 coronal (axial) slices, thickness 0.8 mm, gap 0, acquired interleaved, scan time 5 min 26 s). Field dependent adjustments to acquisition parameters are shown in Table 2.
Multi-echo gradient echo (MGE) 3D sequence with 160 μm3 isotropic resolution, TR 66 ms, 13 echoes with TE from 2.7 ms to 43 ms with interval of 3.1ms, flip 16° (close to Ernst angle), FOV 25.6 × 19.5 × 12.8 mm, 160 × 122 × 80 matrix, 1 average, 120 dummy scans to obtain steady state, with outer volume suppression (OVS) and fat suppression, a scan time of 11 min was used. MGE data were used to calculate T2* maps, anatomical mixed contrast images and susceptibility-weighted images (SWI). For high SNR Tl/T2*-weighted mixed-contrast anatomical images the 13 volumes (over the echoes) were summed. SWI were created by applying a phase mask calculated based on the phase shift between the 1 st and 5th echo on the anatomical images.
Diffusion-weighted imaging (DWI) for diffusion tensor imaging (DTI) and tractography was performed by single-shot spin echo 3D echo-planar imaging (EPI), with diffusion gradient duration (δ) = 4.2 ms, diffusion gradient separation (Δ) = 12 ms and b-value = 2800s/mm2 in 42 non-collinear directions with 4 non-diffusion-weighted (b0) images, TR 1000 ms, TE 26 ms (shortest possible with 1 ms added to avoid eddy currents), EPI echo spacing 0.269 ms with 250 μm3 isotropic resolution, FOV 24 × 18 × 12.3 mm, encoding matrix 96 (read) × 54 (PEI) × 49 (PE2), zero-filled to 96 × 72 × 49, spectral width 357 kHz, 1 average, and fat suppression and OVS with 4 saturation bands, resulting in a scan time of 37 min. Reference images (one b0 image) with reversed phase encoding order was acquired for susceptibility-induced off-resonance field correction. Non-respiratory-gated, single-shot approach was adapted to stay within the 2-h time frame for total scan time.
Magnetization transfer (MT) imaging was performed using a fast low angle shot 3D (FLASH-3D) sequence with TR 47 ms, TE 2.6 ms, flip 13° (Ernst angle), 2 averages, 170 dummy scans (5*T1 to achieve steady state), 250 μm3 isotropic resolution, FOV 32 × 26 × 14 mm with a single outer volume saturation (OVS) band, matrix 128 × 104 × 56, and scan time of 7 min. The scan was repeated with and without the MT preparation pulse of −1500 Hz offset, 12 ms Gauss pulse, 228 Hz bandwidth, 3 pulses, inter-pulse delay 0.01 ms and peak power 10 μT (for MT0 images the power was set to zero). Magnetization transfer ratio (MTR) maps were calculated as MTR = ((MT0-MT)/MT0)*100%. To control MT pulse delivery, which is a potential source of error in MTR mapping, B1 map data were obtained using 3D gradient echo EPI (1094 μm3 isotropic resolution, peak power 10 μT, and an array of block pulse lengths from 0 to 3.0 ms with 0.2 ms interval, which after cosine fitting yielded the frequency map across the brain).
Table 2 summarizes the key parameters of each sequence, and field-dependent adjustments are indicated in red. The 3D echo-planar imaging diffusion methodology was the determining factor in planning the imaging protocol due to its demands to the hardware and its long imaging time. A single-shot rather than a multi-shot approach was chosen for the 3D DTI to fit within a 2-h time frame. The use of isoflurane anesthesia and the feasibility of achieving a high-throughput work-flow limited the time/rat in the scanner to a maximum of 2h. Other preconditions were 1) field-of-view to cover the entire brain including the cerebellum, 2) isotropic voxel size for optimal DTI / tractography analysis, and 3) high enough spatial resolution for assessing a variety of sizes of white matter tracts, accurately locating the microbleeds, and distinguishing the glial envelope surrounding the lesion.
Region of interest (ROI) analysis was performed on data obtained from sham-operated rats. Eight shams from each center was included. ROIs were manually drawn by the same investigator (RI) as follows. Cortical ROI at −3.60 mm from bregma, and background noise ROI of the same slice were drawn in T2wt images for signal to noise ratio (SNR) calculation. Cortical grey matter and adjacent corpus callosum as white matter ROI were used to analyze contrast to noise ratio (CNR). In order to quantify the signal attenuation due to the diffusion gradients in DWI, a whole brain ROI in a central horizontal slice were drawn, and the non-diffusion weighted signal intensity was compared to the average of the diffusion weighted signal intensity over all diffusion directions. ROI for fractional anisotropy (FA) analysis was the corpus callosum, and ROIs for grey matter and white matter MTR the cortex and adjacent corpus callosum and external capsule, respectively.
Quality control scans with a phantom were executed at the beginning of each acquisition block in order to monitor hardware stability. These consisted of scout images to monitor the shape of the object as an indicator of gradient performance, pulse calibration to monitor coil performance and its stability over time, and point-resolved spectroscopy (PRESS) to monitor the water peak linewidth and hence the shim quality for obtaining a high field homogeneity.
2.3. Ex vivo sample preparation and MRI methods
At the end of the follow-up after completing the video-EEG monitoring, the rats were transcardially perfused with 0.9% NaCl for 5 min followed by 4% PFA for 20 min, and then immersed into 4% paraformaldehyde for 4 h. Then, the brains were washed in 0.9% NaCl for 24–48 h and immersed in Galden (Solvay) for imaging. Galden is a low-molecular weight perfluoroether liquid with no mobile hydrogen atoms, hence giving no background signal in 1H MRI. Exposure to Galden was kept to a minimum to avoid any compromise in the quality of immunohistochemical stainings to be performed in the biomarker validation phase. That is, brains were not stored in Galden for over 3 days but moved into 2% PFA for any longer storage period. After the scan, brains were flushed in 0.9% NaCl for a minimum of 2 h before cryoprotection (at least 24 h in 20% glycerol made in 0.02 M phosphate buffered saline with potassium), and then, frozen in dry ice. Thus, ex vivo scans were scheduled for 2 days post perfusion. Instruction for ex vivo sample handling and setup were distributed to all centers to ensure identical practices across sites. Importantly, the ex vivo brains were positioned mimicking the brain position in in vivo scans, hence maintaining identical fiber position regarding the B0 and main gradient directions. Misplacement (angle or tilt) of 5° or more required re-positioning (Le. angle < 5° was considered acceptable).
Ex vivo MRI methods were essentially identical to those used for in vivo imaging, including the same coil setup, volume transmitter and surface receiver, as well as the same MR sequences. The site-specific details are shown in Table 1. A longer ex vivo scan time enabled the acquisition of images with a higher resolution in DWI and MT acquisitions (160 μm3 ex vivo vs 250 μm3 in vivo). The faster relaxivity and lower temperature of fixed brain during imaging as compared to that in in vivo imaging were compensated for by adjusting the acquisition parameters as follows. DWI: single-shot EPI 3D, identical 42 diffusion directions with in vivo scan + 4 b0 images, 160 μm3 isotropic resolution, echo train length (ETL) 33.6 ms [T2 in ex vivo (at 7 T) is 40–45 ms], TR 620 ms (T1 saturation corresponding to in vivo conditions), TE minimum 39.5 ms (shortest possible with 1 ms added to avoid eddy currents), 3 averages, b-value was set to 3600 s/mm2 (based on free water diffusivity at 25 °C ex vivo vs 37 °C in vivo). Since fixation decreases the mean diffusivity (in vivo MD 0.61 μm2/ms and ex vivo 0.39 μm2/ms (data not shown), harmonization of the root-mean-square (rms) displacement between in vivo and ex vivo when probing the structures with similar size requires increasing the effective diffusion time in ex vivo imaging. Diffusion gradient separation (Δ) of 18 ms, and duration (δ) 6 ms (preserving the short-pulse approximation Δ ≥38) resulted in rms displacement of 6.0 μm compared to 6.2 μm in vivo. Since the fixation shrinks the tissue, these displacements are comparable. An ex vivo gradient amplitude of 40% was used compared to 64% in vivo and this avoided any site-specific limitations due to gradient performance. Ex vivo scan time for DWI was 1 h 40 min compared to 37 min in in vivo. Ex vivo MGE sequence was the same as in vivo, apart from adjusted FOV, TR (minimum) 48 ms, flip 17° (Ernst), and 13 echoes from 2.73 to 43 ms with 3.1 ms interval. The echo train was the same as used in vivo, and despite the ~5 ms shorter ex vivo T2, the SNR of final echoes was >10 SNR. Scan time was 6 min. Ex vivo MT scans were identical to that used in vivo, except for 160 μm3 isotropic resolution (compared to 250 μm3 in vivo), TR 45 ms, flip angle 17°, and 3 averages. MT preparation pulses were identical to those used in in vivo imaging. Scan time was 22 min as an 11-min scan was repeated with and without the MT preparation pulse. The total duration of each imaging session was about 2h 30 min when the 10-min time for preparations and sample setup were included.
2.4. Statistics
Statistical analyses were performed using SPSS (IBM SPSS Statistics 23). Univariate ANOVA with Bonferroni post hoc test were used to calculate the pairwise group differences in each obtained MRI parameters between the three sites (p-values < 0.05 were considered significant). Repeatability over time was evaluated by related samples Friedman’s 2-way analysis of variance. Data are shown as mean ± standard deviation (std), and standard deviations are also used to describe the variance within site and across all sites.
2.5. Harmonization process
The first version of the MRI protocol was drafted and implemented at UEF in line with the research plan and taking into account the hardware specifications of each site. Drafts of methodological protocols were distributed, and the test runs for the different sequence options were conducted at each site to evaluate if there were any additional limitations imposed by the hardware and practical site-specific scanning conditions. The protocol was then iteratively revised accordingly. Testing was conducted both with brain phantoms (perfused ex vivo rat brain) and with live animals. Importantly, pilot data was also obtained from acute and subacute trauma animals (instead of only controls) 1) to ensure optimal contrast for detecting the TBI-induced pathologies, and 2) to evaluate and optimize the methodologies, for example in the presence of a hemorrhagic lesion, which is a major source of susceptibility artifacts after TBI.
Flow chart of MRI data acquisition harmonization process shows iteration rounds, data sharing, and the establishment of the final common procedures, instructions and quality monitoring paradigms (Fig. 2). Feedback from the partners at the first consortium meeting lead to a modification of the field of view to encompass the cerebellum and changes in image resolution. Test scans of TBI rats at day 2 favored adding one more modality (T2wt) for initial robust lesion characterization. Sharing the data obtained with the 2nd version of the protocol resulted in further modifications to the sequence selection to eliminate site-specific artifacts. Version 3 of the protocol produced artifact free data at each site, and the field-dependent acquisition parameter adjustments were implemented (Table 1). Pilot data was shared to compare the SNR, image quality and potential artifacts. Once images were considered to have the lowest inter-site variability obtainable by using the current settings, final methods and step-by-step operation procedures were distributed to each site.
Fig. 2. Flowchart of the MRI data acquisition harmonization at the three study sites (UEF, yellow; Melbourne, purple; UCLA, green).
MR protocol editing was performed during the first months before the onset of the animal follow-up study. The initial version of the MRI protocol was revised based on the feedback from the consortium and from the MRI experts at each center. Data was shared at each step allowing changes to be made for image artifacts and differences in signal properties by tailoring the sequences. Several optimization rounds among the hands-on MRI experts took place. Finally, pilot data demonstrated comparable image quality between sites, and the experiments on the first animal cohort of the study were started. Monitoring of the consistent data quality during the longitudinal follow-up, repeatability and hardware stability were conducted by interim analysis and additional test measurement pattern with liquid phantom.
Thus, four iteration rounds of optimization and feedback, shared methods, and distributed instructions yielded successful harmonization of MRI data. In addition, frequent quality control phantom scans were established for hardware performance monitoring during the study. Phantom experiments are described in detail in Section 3.5.
3. Results
3.1. Signal harmonization at pilot stage
Step 1 of the harmonization process was to tailor the sequences executable by each system and account for the different field strengths and gradient performances. Sequences were first optimized for probing FPI at 7 T Bruker system at UEF. Thereafter the field dependence of a number of tissue signal parameters were balanced between sites by tuning the acquisition parameters (Table 1). These parameters were: TR that was chosen based upon tissue T1 and T2 relaxivity, flip angle that was calculated to be the Ernst angle, and number of signal averages that was balanced to obtain the same SNR.
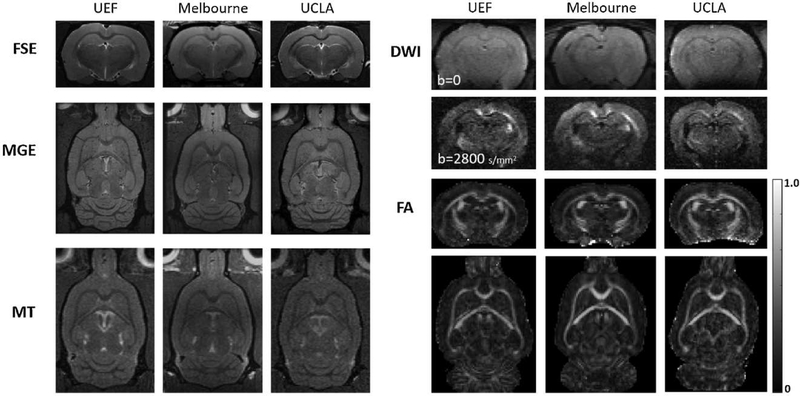
Comparable image quality between sites was achieved as the result of the piloting. SNR, CNR, MTR, diffusion attenuation, and FA varied less than 10% between the sites when representative example cases from each site were compared. Ghosting in echo-planar images was found to be negligible in brain phantoms (with ghost/noise ratio 1.5). Ghosting in naïve animals was limited to the vicinity of the cerebellum. Ghosting observed in pilot FPI animals overlapping the brain was minimized to below 10% (calculated as relative signal increase in the cortex due to the ghost) by minimizing any respiratory related movement. Optimization of OVS scheme and fat suppression scheme resulted in elimination of artefatcs that related to chemical shift and overlapped with the brain tissue. Fig. 3 shows representative data of control rats from each center side by side. Based on the pilot data, systematic MRI data acquisition was started.
Fig. 3. Harmonization of signal and contrast across UEF, Melbourne and UCLA in every MRI modality.
The panel shows representative example images from different sites for each MRI modality: T2-weighted fast spin echo 2D multi-slice (FSE), multi-echo gradient echo images (summed over all 13 echoes) (MGE), magnetization transfer-weighted images (MT), diffusion-weighted imaging (DWI) raw data with b-value 0, and diffusion-weighted with b-value of 2800s/mm2, and fractional anisotropy maps (FA).
3.2. Procedure harmonization
3.2.1. Shared MRI measurement procedures
The finalized methodological protocols and instructions to run the shimming and experimental queue were distributed to all sites and implemented into the common software (ParaVision version 5.1 or 6.0). The site-specific parameters were entered into the measurement protocol by the local researcher in charge. All site-specific modifications were communicated with and agreed on by the coordinating site (UEF).
3.2.2. In vivo procedure harmonization and animal physiology
The animal holders and coil setup at each site resulted in similar animal positioning. At UEF, anesthesia duration for the MRI scan was 1 h 47 min ± 12 min (range 1 h 30 min to 2 h 41 min). Time included both the preceding blood sampling prior to the scanning and the MRI, both done under the same anesthesia. In Melbourne, the anesthesia duration for MRI was 1 h 45 min ± 9 min (ranging from 1 h 26 min to 2 ho 10 min, aborted scans excluded from the calculation). Blood sampling was done under different anesthesia, preceding the MRI imaging. At UCLA, anesthesia duration for MRI was 1 h 45 min ± 5 min (range from 1 h 40 min to 2h 10 min). Blood sampling was done prior to MRI under a different anesthesia. Anesthesia durations did not differ between sites. Anesthesia induction took 4 min (5% isoflurane in induction chamber), blood sample from tail vein (at UEF) took 5–10 min, setup on animal holder 5–10 min, and then the scan was initiated. The scan execution order in written instructions was the same for all sites. Order of sequences was selected based on their sensitivity to anesthesia and temperature: after the MAPSHIM and the 5 min T2wt FSE, the 11 min MGE (T2* mapping) was done next to avoid the effect of putative increase in blood CO2 level towards the end of the long anesthesia, the 37 min DWI was done thereafter, and MT as last. Respiratory rate was monitored with a pressure sensor, and body temperature with a rectal probe utilizing small animal monitoring unit (PC Sam, SA instruments, NY). Animals were kept warm by heated water-circulation of the holder and by planket. Body temperature of TBI rats at the beginning of the scan was low, generally 35–36 °C. Body temperature was raised by adjusting the water circulation, and maintained thereafter at 37 °C. Temperature during the DWI scan was recorded, given the sensitivity of diffusion measurement for the temperature. Isoflurane anesthesia (1.5% isoflurane, 30% O2/70% N2 as carrier gas) level was adjusted according to the breathing rate and monitored through the measurements. Generally, the respiratory rate was stable and similar between sites (maintained between 45–70/min, max range 40–80/min). Table 3 lists the animal physiology related factors that can affect the MRI data. The ranges are comparable across sites. Only 3% of the animals imaged at 2 or 9 d post-TBI had to be withdrawn from the magnet due to respiration-related issues and could not be re-inserted. Consequently, the 2-day or 9-day MRI data in 3 of 35 rats in Melbourne is incomplete due to the aborted scan. In corresponding situation in UEF and UCLA, the rats were re-inserted into the magned and scanning was continued. Only 2% of all rats at all sites died during the scan (2/26 for day 2 scan at UCLA).
Table 3. Factors in animal handling that may affect the obtained MRI data and their variation across sites.
Animal temperature was monitored and adjusted throughout the scan session by adjusting the water circulation heating of the holder. Range of temperatures recorded during the diffusion weighted imaging (DWI) are shown. Respiratory rate was monitored by pressure sensor and maintained constant by adjusting the isoflurane. Respiratory rate during the scan session was generally stable in each animal, but varied between animals. Range of respiratory rates at each site are shown, as are the range isoflurane levels. Carrier gar was identical in all sites (30% O2 / 70% N2). Ranges are comparable across sites. Total anesthesia durations are shown as mean ± std, and they are similar across sites despite the fact that at UEF a blood sample was drawn after the anesthesia induction. Anesthesia duration before the scan onset (anesthesia prior scan) consists of induction (5% isoflurane in chamber), blood sampling, and setting up animal on the holder. Holder setup was similar across sites. Scan execution order was designed to their sensitivity towards effects of blood CO2 levels on T2* relaxation, and effects of body temperature on diffusion. Similar scan execution order was instructed to all sites. Occasional deviations from the order have not been analyzed.
| Animal physiology | UEF | Melbourne | UCLA |
|---|---|---|---|
| Body temperature | 36.5–37.6 °C | 36.0–37.0 °C | 36.0–37.0 °C |
| Respiratory rate: maintained range (brief deviations: min-max) | 48–70bpm (45 – 80bpm) | 50–65bpm (45 – 80bpm) | 45–60bpm (40–80bpm) |
| Anesthesia duration | 1h 47 min ± 12 min | 1h 45 min ± 9 min | 1h 45 min ± 5 min |
| isoflurane level | 1.3–2.0% | 1.5–2.0% | 1.5–2.0% |
| anesthesia duration prior scan induction/blood sampling/setup | 5min/5 – 10 min/5min | 5min/ – /5min | 5min/ – /5min |
3.2.3. Harmonization between ex vivo and in vivo MRI methodologies
Ex vivo MRI serves as a common nominator between the animal cohorts in EpiBioS4Rx, undergoing either chronic MRI follow-up or video-F.F.G monitoring. In particular, ex vivo MRT was conducted for the video-EEG -monitored cohorts as in vivo MRI is not possible due to the recording electrodes and their mounting headset on top of the skull.
Our aim was to tailor the ex vivo imaging to produce data which would carry the same information as the in vivo data (i.e, the same MR modalities) but with a higher spatial resolution. Harmonization between in vivo and ex vivo contrast was maximized by taking into account the altered relaxivity due to the fixation, shrinkage of the brain due to fixation, and lower diffusivity due to the lower temperature of the ex vivo brain.
Methods Section 2.2. Lists the ex vivo parameters. Fig. 4 shows obtained in vivo and ex vivo images in parallel (Fig. 4A–C). A higher resolution (160 μm3) of ex vivo DWI and MT allows more detailed analysis of white matter tracts, for example of the intrahippocampal structures than the in vivo imaging (250 μm3). In post-processed MGE images the contrast ((SIgm-SIwm)/SIgm) was 0.23 in ex vivo and 0.22 in in vivo imaging. Contrast-to-noise ratio (CNR) quantification in vivo was not possible due to the absence of tissue-free ‘noise region’ within the FOV. In those cases, the estimation using the bone as ROI for ‘noise’ yielded ex vivo CNR 15.8 and in vivo CNR 7.4. In the raw diffusion images the SNR of ex vivo b0 images was 28.1, and SNR of ex vivo diffusion-weighted images was 7.9. Since in vivo images lack any background noise region within the FOV, the SNR is estimated with bone ROI as ‘noise’ ROI. The obtained SNR of in vivo b0 images was 16.9 and SNR of in vivo diffusion-weighted images was 3.3. Signal attenuation due to diffusion weighting was 74% ex vivo and 82% in vivo.
Fig. 4. Harmonization between in vivo and ex vivo MRI.
(A–C) Image pairs of the same animal scanned in vivo at 5 months post-injury (left column) and the ex vivo scan of fixed brains at the 7 months post-injury end point (right column). Data from one representative animal imaged at UEF is shown: (A) Mixed-contrast MGE, same resolution 160 μm3 in vivo and ex vivo, (B) FA maps in vivo 250 μm3 (37-min scan time), ex vivo 160 μm3 (1 h 40-min scan time), directionally encoded color map (red left-right, blue up-down, green through plane), (C) MTR maps in vivo 250 μm3 while ex vivo 160 μm3 (note that the shift in MTR magnitude due to fixation cannot be compensated for, and thus the scales differ between in vivo and ex vivo MTR). (D–F) Ex vivo sample positioning mimics the in vivo brain positioning with respect to the main gradient orientations. Fixed brains were immersed in a tube filled with perfluoropolyether (Galden®, Solvay). ‘Nose’ was supported upwards, and any movement in Z-direction or any rotation were eliminated with a support from behind with minimized pressure to the cerebellum (arrowheads in E, center alignment in F). Instructions about ex vivo sample setup were distributed across sites to avoid systematic bias in diffusion measures due to different angle of the brain with respect to the main gradient orientations. Abbreviations: MGE, multi-gradient echo; FA, fractional anisotropy; MTR, magnetization transfer ratio.
In MT imaging, the same preparation pulses were applied as in vivo, recognizing the difficulty to compensate for all possible changes caused by tissue fixation which can influence the mechanisms contributing to MTR. Nevertheless, obtained GM/WM contrast in MTR was close to that of in vivo MT. In WM the ex vivo MTR was 70%, while in vivo it was 67%. In GM the ex vivo MTR was 59%, while in vivo it was 59%.
Similar to the in vivo harmonization pipeline, the ex vivo data acquisition methodologies and procedures were harmonized between the study sites by distributing the detailed instructions of the MR sequence parameters, sample preparation and sample positioning (Fig. 4D–F). Ex vivo experiments were not yet conducted at all sites at the time of the interim analysis, and hence, quantitative data from multi-center harmonization of ex vivo MRI is not included in this report.
3.3. Execution and monitoring of study execution
Study execution was monitored by assessing the timing of the scan and success rate of scans through all animals scanned by April 30, 2018. Variation in signal and contrast between the study sites were evaluated by comparing the data of 8–10 shams from each site at a single time point. Inter-batch variability (reproducibility) was evaluated by analyzing the variation in data of shams from one time point to another.
3.3.1. Signal harmonization
Data obtained from sham-operated experimental controls at different centers was analyzed for SNR, contrast to noise ratio (CNR), signal attenuation due to the diffusion gradients, fractional anisotropy (FA) and MTR. ROIs were manually outlined by the same investigator and sampled across the rostro-caudal extend of the brain to eliminate any effect related to coil positioning. Fig. 5 shows the results and variation between sites. SNR in T2-wt images was 34.9 ± 2.2 at UEF (n = 9), 47.7 ± 2.2 in Melbourne (n = 8), and 32.1 ± 1.7 at UCLA (n = 8) (difference between the sites p < 0.05). Importantly, SNR was 30 or higher in each animal, which can be considered as a good SNR for the visual assessment of lesion location and extent. CNR (contrast between white and grey matter/noise) in T2-wt images were 8.9 ± 1.0 in UEF, 9.6 ± 0.9 in Melbourne, and 8.2 ± 1.2 in UCLA. No difference between the sites was found (p > 0.05). Magnetization transfer ratios (MTR) in grey matter were 58.4 ± 0.5% at UEF, 56.5 ± 0.6% in Melbourne (difference between the sites p < 0.05). MTR in white matter was 67.0 ± 0.7% at UEF and 65.3 ± 0.1% in Melbourne (difference between the sites p < 0.05). However, the WM/GM MTR contrast was comparable between sites, being 1.15 ± 0.01 at UEF and 1.15 ± 0.01 in Melbourne. MT0 data was corrupted for the first animal cohort in UCLA due to a scan execution error, and hence, the MTR from UCLA is missing in this analysis. The error was corrected for the subsequent cohorts of animals. Signal attenuation due to diffusion gradients, i.e. the drop in signal intensity in diffusion-weighted images (average over all diffusion directions) as compared to the b0 was 83.4 ± 0.4% at UEF, 75.8 ± 0.5% in Melbourne, and 79.6 ± 1.1% at UCLA (difference between the sites p < 0.05). Fractional anisotropy (FA) in white matter (corpus callosum) was 0.81 ± 0.01 at UEF, 0.73 ± 0.02 in Melbourne, and 0.84 ± 0.03 at UCLA (p < 0.05). Variation between sites in obtained signal is to be accounted for in post-processing.
Fig. 5. Variability in quantified signal properties of three different signal modalities between the sites.
The success in signal harmonization of T2-weighted (T2wt), diffusion, and magnetization transfer imaging was evaluated by analyzing the data of sham animals 30 days post-operation. (A–B) Signal-to-noise ratio (SNR) in T2wt images differed between sites, but was high in all sites (> 29 in all animals). Contrast was comparable between sites, that is, contrast-to-noise ratio (CNR) between white and grey matter in T2wt images (B) did not differ. (C–D) Attenuation in diffusion weighted images as compared to non-diffusion weighted b0 images (calculated as (b0-DWI)/b0*100) was different between sites as were the calculated fractional anisotropy (FA) values in corpus callosum. (E–F) Magnetization transfer ratio (MTR) white matter/gray matter contrast were equal between sites (E), while MTR in white matter (corpus callosum) did differ between sites (F). Data were obtained from sham-operated rats (n = 8–9 per site). No MTR data from UCLA. Data are shown as mean ± std. Statistical differences: *, p < 0.05 (between the sites, ANOVA with post hoc analysis with Bonferroni).
3.3.2. Reproducibility in data acquisition
Reproducibility of the obtained MRI signal parameters over repeated scans in sham animals were evaluated in UEF cohort. Repeated scans of 9 sham-operated rats during the 5-month follow up at UEF depicted no variation over time. Fig. 6 shows SNR in T2wt images (cortical ROI) as an example of the measures used in the analysis at each time point. As expected, the measurements between day 9 and day 30 yielded similar SNR. Furthermore, there was no reduction in signal level at day 2 after sham-operation even though the distance of the coil from the brain was greater than at later time points due to swelling on top of the head related to sham-surgery -induced swelling on the top of the head and stitches. Neither was there any decrease in SNR at later 5-month follow-up point even though the rats were aged and larger. Low inter-patch variability can be seen also in Fig. 5 where the data of the sham-operated experimental controls acquired over a period of several months shows low variability in SNR, CNR, FA and MTR within each study site, indicating high reproducibility in data acquisition. Pulse powers and linewidths over time did not change, demonstrating hardware stability over time.
Fig. 6. Stability of signal acquisition during the follow-up.
Repeated scans of control animals were used to evaluate the repeatability of MRI measurements between different sessions. The obtained signal-to-noise ratio (SNR) in T2-weighted images in the same sham-operated control rats show no change over time. Note, that SNR is equally good at 2 days and 5 months post-injury as at intermediate time points, even though the postoperative swelling of the tissue on top of the head at day 2, and the large size of the animal, and large size of the skull at 5 months cause the coil to be further away from the brain surface. Statistics: mean ± std, n = 9 (UEF), related samples Friedman’s 2-way analysis of variance; *, p < 0.05.
Patch effect was further studied by comparing the SNR of the T2wt data obtained at 9 days post-injury to that of 30 days. Uniform SNRs were obtained between these repeated scans at UEF and Melbourne (p > 0.05). SNR at UCLA improved from 9 to 30 days post operation (p < 0.05), likely reflecting learning effect on animal and RF-coil positioning. No other patch effect was observed.
3.3.3. Scan success rate
The aim was to have an equal number of samples at each site. Since the collection of MRI data started at different times at different centers, the number of animals scanned varied accordingly. In the interim analysis we monitored the success rate in MRI scans, that is, the success in performing the measurement, and collecting the raw data.
Unsuccessful MRI scans were usually due to respiratory difficulties which caused the animal to be withdrawn from the magnet. Another reason for exclusion related to data corruption by movement. Such instances occurred only in 2day or 9day scans. A rarer reason for imaging failure was temporary hardware failure (one rat at UEF with missing data on day 2). At UEF the success rate for acquisition of different imaging modalities for 39 rats scanned over 3–4 time points was 99% (136/137) for T2wt, 99% (136/137) for MGE, 99% (136/137) for DWI, and 99% (135/137) MTR. That is, a complete data set was acquired for all animals at all time points, except for missing data from one animal at the 2-day time point and one additional failed MT scan. Similarly, at other sites the success rate was close to 100%. At Melbourne, data was missing from 3 rats that were withdrawn from the magnet due to respiratory issues. At UCLA, data was missing from two animals that died during the scan. As noted before, MTR maps were missing at UCLA from cohort 1 due to a sequence execution error.
3.3.4. Timing of imaging relative to the injury induction
For the discovery of imaging biomarkes, the timing of the scan from the injury is a critical factor as the brain damage progresses and varies in type over the time. Timing of scans was comparable across sites (Fig. 7). Even though the nine outliers increased the variability, a high precision in timing was achieved as the 2day measurements and the majority of 9-day measurements were within an hour of the target time. The exact time of 2-day measurement from the injury was 48.0 ± 0.1 h at UEF, 48.1 ± 0.1 h in Melbourne and 55.4 ± 10.9 h at UCLA (difference between the sites, p < 0.05). The time delay from the impact to the 9-day MRI was 216.0 ± 0.12 h at UEF, 212.3 ± 9.0 h in Melbourne, and 228.1 ± 33.7 h at UCLA (difference between the sites, p < 0.05). Importantly, all but 2 animals were scanned within ± 24 h from the scheduled time point. The time of 1-month MRI was 30.0 ± 2.2 d at UEF, 30.1 ± 1.5 d in Melbourne, and 28.7 ± 1.9 d at UCLA (p < 0.05).
Fig. 7. Inter-site variability in timing between the impact and the scan.
Animals scanned by April 30, 2018 were included in the analysis (UEF, green; Melbourne, purple; UCLA, blue). Scatter plots (A–C) show the time of scan for each animal. Group averages and differences between sites (asterisks) are shown on the green box. Despite 9 outliers, all animals are scanned within ± 25 min from the target hour on day 2. Most animals were scanned within an hour from the target at day 9, and all but 2 outliers within 24 h from the target. The range at the 1-month target time point was 31 ± 3 days (with one outlier). Data are shown as mean ± std. Statistical differences: *, p < 0.05 (between the sites, ANOVA with post hoc analysis with Bonferroni); n (UEF) = 39; n(Melboume) = 35; n (UCLA) = 26.
3.4. MRI screening of lesion location and size
MRI plays a critical role in the analysis of the location and extent of the produced lesion. The 2-day MRI scan verifies successful impact and allows immediate elimination of subjects, for example, with non-impact related injury (e.g., craniectomy-related damage, excessive hemorrhage) by predefined exclusion criteria. We also monitored variability in the location and extent of lesion between the study sites. That can be done utilizing either 2-day, 9-day or 1-month post-TBI images. The 1-month post-injury images were optimal for inter-site comparisons as the transient edema had resolved, and the lesions had already progressed significantly, allowing its accurate detection in T2wt imaging. During the early harmonization phase, MRI screening alerted about problems in model production between sites, leading to site-specific corrections in surgical procedures, and yielding eventually to rather uniform variability in the production of lateral FPI model. Fig. 8 depicts the similar lesion location and size range between the three sites.
Fig. 8. MRI screening of the lesion location and size.
T2-weighted coronal images were screened to confirm that (a) the impact given resulted in comparable lateral fluid-percussion injury at each site, (b) the study population did not have additional etiologies for epileptogenesis which would compromise the biomarker analysis. Top row shows the smallest lesions at each site, and bottom row shows the largest lesions from each site (T2wt images). Location (arrow) is uniform across the sites and the cortical lesion extent is comparable.
3.5. Phantom measurements for QC: hardware stability over months / years
Since the EpiBios4Rx biomarker discovery study will continue for several years, the hardware failures and updates had to be accounted for. Thus, the paradigm included periodic phantom quality control (QC) scans every 1–2 months – or whenever a new animal cohort was started. Phantom QC was designed to monitor hardware stability over time: gradient coils (shape distortions), shim coils (linewidths) and Tx/Rx coils (pulse powers). Standardized liquid phantom was used (Vendor’s standard or corresponding 50 ml tube containing 1 g/L CUSO4-5H2O and 3.6 g/L NaCl in double-distilled H20). Phantom scans were executed at each site with variable frequency. Phantom was scanned 7 times/y at UEFand 2 times/y in Melbourne and at UCLA. Fig. 9 depicts the testing pattern and the results at UEF. Fast high-resolution gradient echo images in each dimension show no shape distortions, indicating correct gradient performance (Fig. 9A). Array of excitation pulse lengths depicts the pulse length needed to obtain 90° and 180° flip angles (with constant power) (Fig. 9B). Results of the phantom experiments showed stable excitation pulse powers (attenuation range 16.7–17.2 dB) and shim linewidths in 10 × 10 × 10 mm voxel (range 7.8–15.8 Hz, including one outlier), indicating stable hardware (Fig. 9C). Corresponding values in Melbourne were 10.7–11.9 dB and 9.0–14.5 Hz, and in UCLA 18.5–22.3 dB and 11.0–12.8 Hz, respectively. These observations with phantom were in line with stable pulse calibration and shim parameters obtained in in vivo experiments. Thus, fluctuations in hardware performance over time as an error source in a long-lasting study were unlikely.
Fig. 9. Phantom quality control (QC) experiments.
(A) Gradient performance was assured by screening for any shape distortions in fast high-resolution scout images. None were observed. (B) Global excitation obtained (single pulse) with different pulse lengths, maximum signal intensity corresponding to the 90-degree excitation obtained with 1 ms pulse length. (C) Table of the results of 7 QC phantom scans performed over a period of 1 year at UEF, depicting pulse powers (for 1 ms 90° excitation) over time (shown as attenuation in dB), and the line width obtained (i) globally with single pulse and (ii) with localized iterative shimming to the 10 × 10×10 mm voxel shown in A. Excluding the one outlier (* in C), the linewidths ranges are 25.4–30.3 Hz globally (i) and 7.8–10.8 Hz locally (ii).
3.6. Data transfer and analysis pipelines
To achieve maximal harmonization in data analysis, all data will be transferred to Laboratory of Neuro Imaging (LONI) at USC Stevens Neuroimaging and Informatics Institute. Data will be run through common analysis pipeline built for EpiBioS4Rx and made available to each site. In parallel to this interim analysis, the analysis pipelines are being constructed.
3.6.1. Data transfer to LONI
Data are transferred to LONI via a secure FTP site that was created for data collection and sharing with the goal to have all data modalities (EEG, behavioural, MRI, blood, etc.) on the same database, allowing cross-correlations, grouping, and clustering. The animal imaging data are collected in Bruker format; the secure FTP site is used for the transfer so that LONI can convert the data to NifTI (nii) format, which is then uploaded to LONI’s Image and Data Archive (IDA) along with the other data modalities that are directly uploaded to the site.
3.6.2. Development of data post-processing and analysis pipelines
In conjunction with data storage, indexing, and archiving, LONI will provide data analytic services in collaboration with the imaging centers. This will ensure that image analysis will be performed uniformly regardless of the site of origin; furthermore, this will take advantage of state-of-the-art hardware and software resources for robustly and efficiently designing, documenting, and executing data analysis workflows.
LONI’s hardware infrastructure uses a fault-tolerant, high-availability systems design to ensure 24/7 functionality. The primary storage cluster is 51 EMC Isilon nodes with 5.3 usable petabytes of highly available, high performance storage. The compute infrastructure within the datacenter boasts 4096 cores and 38 terabytes of aggregate memory space. This highly available, redundant system is designed for such demanding big data applications. To take advantage of these compute resources, LONI uses a batch-queuing system for parallel processing, and in-house software has been developed to facilitate its use in neuroimage analysis.
The LONI Pipeline (Dinov et al., 2010) is software for designing computational workflows using a graphical data flow model, which provides an intuitive way to orchestrate complex image analysis tasks that are performed on large datasets. This will enable both local and remote users to submit jobs to process imaging data and to maintain data provenance to ensure that uniform processing can be verified. The data analysis will be implemented in the LONI Pipeline using components from the Quantitative Imaging Toolkit (QIT) (Cabeen et al., 2018) and other 3rd party packages, such as FSL (Jenkinson et al., 2012). QIT is developed at the institute to provide a uniform interface to state-of-the-art image processing algorithms and visualization techniques, such as diffusion modeling and tractography based analysis. It will be used within the LONI Pipeline for analyzing data, and it will also provide remote collaborators with a tool to explore anatomical models and statistical results derived from the study data.
4. Discussion
Harmonization of MRI procedures, acquisition methods and data quality between different experimental sites of a consortium is a crucial step for the success of multi-site imaging studies, both preclinical and clinical. Here we present the first effort to rigorously achieve a harmonization in preclinical MRI imaging at three pre-clinical MR imaging centers, including the University of Eastern Finland (UEF), The University of Melbourne/Florey Institute (Melbourne, Australia) and the University of California Los Angeles (UCLA, USA). In addition to harmonization procedures, we analyzed the success of harmonization by conducting an interim analysis of the procedural accuracy to guide the planning for future pre-clinical multicenter imaging studies and data analysis.
Harmonization process
Harmonization process between 3 preclinical imaging centers with different MRI systems using different gradients, RF-coils and field strengths required several iteration rounds to fine-tune methodologies. The process required active communication, method sharing, data sharing, feedback and documentation of procedures. Pilot experiments started about 6 months prior to the imaging of experimental animals. Piloting was done by first imaging ex vivo brains from naive rats and rats with lateral FPI, and then moving on to in vivo imaging. Importantly, we also included rats with acute FPI and hemorrhagic lesions in the method optimization phase to determine whether the tailored sequences produced artefact-free data also in the presence of hemorrhages. Protocol changes resulting from the iterations included field of view adaptation to include cerebellum, decision to have isotropic voxel size in all 3D modalities, resolution modifications in DWI, MT and MGE, and adding the T2wt scan for optimal edema detection. In order to stay within the 2-h imaging time frame, the segmented EPI was changed to a single-shot approach and tested for any additional artifacts in animals with acute TBI. Moreover, B1 field mapping was added to the protocol as a reference for MTR, and reversed phase encoded b0 scan was added to the protocol as a reference for DWI.
Role of MRI in validation of injury quality
MR imaging of the lesion was found to be of paramount importance to ensure that the lesion location and extent were comparable between the study sites. Further, T2wt images were analyzed to ensure that the study population did not have additional epileptogenic injuries that would influence the biomarker analysis and data interpretation.
Reproducible MR signal quality was obtained over time
Animal weight at 5 months post-injury (i.e., 8-month-old rats) was approximately 500 g. To fit them inside the animal holder for MRI scan was an initial concern. However, the selection of the animal vendor and diet were successful (details in Ekolle Ndode-Ekane et al. 2019, in this volume), and the 5-month scans were doable with no deviation in signal properties as compared to smaller size rats. Also, movement artifacts due to respiration as well as the extra distance from the surface coil due to swelling around the sutured surgical incision were concerns for the 2-day scan. However, they occurred less frequently than anticipated, and no systematic effects on the signal properties were observed. Over all, the repeatability of MR signal quality, e.g. the SNR, was excellent.
Hardware stability was monitored and confirmed over time
Hardware stability was excellent as depicted by the consistent shim linewidths, pulse powers both in in vivo scans and QC phantom scans. QC phantom scans also ruled out the occurrence of any gradient malfunction, which could be deducted by the lack of any shape distortions in the images of the phantom.
MRI signal properties of each MR modality were harmonized
MR signal properties were harmonized to the maximum obtainable degree, and the variation within each site was low. Despite rigorous harmonization efforts, some significant inter-site differences remained such as 4–15 % differences in FA and 3% difference in MTR. Consequently, post-processing approaches to level the signals between the sites are considered, for example, a method utilizing rotation-invariant spherical harmonics to account for voxel-wise differences in raw diffusion data between sites (Mirzaalian et al., 2018). Another option is to use a normalization scheme and include the site as a covariate in statistical analysis. The objective is to pool data from all sites to achieve high enough subject numbers for a statistically powered biomarker discovery for post-traumatic epileptogenesis.
The expected signal changes due to the TBI pathology are greater or of the same magnitude than the site-to-site variability. For example, in the body of the corpus callosum the FA differed between sites up to 15%. Pathological changes of FA observed in the animal models of TBI range from 7 to 8 % (Laitinen et al., 2015) to 24–29 % (Harris et al., 2016). In patients FA has been reported to drop 15–20 % (Kumar et al., 2009; Xu et al., 2007). However, the magnitude of the signal changes attributable to the epileptogenesis are unknown. In addition to analyzing the spatio-temporal changes in FA, our analysis strategy for DWI data will utilize tractography and tract-based statistics that are shown to be sensitive to TBI pathology (Harris et al., 2016; Xu et al., 2007; Wright et al., 2017). We hypothesize that the changes in structural connectivity will reveal epileptogenesis related axonal degeneration and reorganization.
Scan procedures were similar at all sites
Animal handling and animal physiology may affect the observed MRI signal. Furthermore, for biomarker study we need to pay particular attention to some imaging-related procedural details that can affect other biomarker modalities (e.g., duration of anesthesia affecting plasma/EEG biomarkers). Procedures were harmonized and yielded no site-specific error sources. Minor variation in success rate arose from the aborted scans due to the occurrence of respiratory problems during scanning of injured animals on day 2. Care was taken to keep the body temperature to 36–37 degrees Celsius, since TBI animals (in acute time points, in particular) tend to get mildly hypothermic. Without appropriate heating pad and blanket the temperature could drop to 34 degrees Celsius inside the cold magnet bore. This may result from the post-injury disturbances in hypothalamic and pituitary endocrine function and hence impaired body temperature regulation during first days post-injury (Heidelbaugh, 2016; Molaie and Maguire, 2018). Lower temperature would decrease the observed diffusion (mean diffusivity) of water molecules in the tissue, and thus, change the sensitivity of the DWI, and the magnitude of DWI based biomarker candidates. Moreover, hypothermia is considered as an additional therapeutic intervention with direct effects on the disease progression after head trauma or status epilepticus (Phillips et al., 2018; Eroglu et al., 2017). Effect of anesthesia was also considered. Anesthesia durations were carefully recorded, and they did not differ between sites. Isoflurane was selected as an anesthetics as it was easy to adapt to be used in all centers, and it is preferred by Animal Licence committees. Elevated blood carbon dioxide levels towards the end of the long anesthesia have an affect on the T2* relaxation. The scan execution order was designed accordingly, acquiring the MGE scan within the first 30 min.
Timing of the scan from the impact
Since evolution of post-TBI secondary pathology is highly dynamic, expression of any imaging biomarker depends on the timing of the scan from the injury (Pitkanen et al., 2018). The variation in timing was monitored within sites and across all sites, and it was found very low, with only a few outliers. Upon inclusion of the outliers the deviations in timing of the acute scans are to be taken into account as a covariate. However, the biomarkers discovered have to be robust enough to allow some shifts in timing, since clinical translation inevitably will introduce greater dispersion in timing.
Data analysis harmonization
This report focuses on the harmonization pipeline of the preclinical MRI data acquisition. The crucial next step is to conduct the data analysis in a harmonized manner for which a common platform is essential. Analysis modules are to be tailored specifically to this animal model to overcome the challenges in utilizing common template approaches in highly deformed trauma brain with highly variable diffeomorphism required across the subjects. Moreover, findings at the group level are to be projected back to the subject level to identify the changes specifically related to the development of post-traumatic epilepsy. Once the best and most robust imaging biomarkers for posttraumatic epileptogenesis are in place, the next step is to use them to select the population for the preclinical therapy trial, and monitor the treatment effects.
Acknowledgements
This research was supported by the National Institute of Neurological Disorders and Stroke (NINDS) Center without Walls of the National Institutes of Health (NIH) under Award Number U54NS100064 (EpiBioS4Rx).
Abbreviations
- CNR
contrast-to-noise ratio
- DWI
diffusion weighted imaging
- FA
fractional anisotropy
- MGE
multi-echo gradient echo
- MT
magnetization transfer imaging
- MTR
magnetization transfer ratio
- SNR
signal-to-noise ratio
Footnotes
Appendix A. Supplementary data
Supplementary material related to this article can be found, in the online version, at doi:https://doi.org/10.1016/j.eplepsyres.2019.01.001.
This article is part of a Virtual Special Issue ‘Discovery of diagnostic biomarkers for post-traumatic epileptogenesis – an interim analysis of procedures in a preclinical multicenter trial EpiBios4Rx’.
References
- Armstrong RC, Mierzwa AJ, Sullivan GM, Sanchez MA, 2016. Myelin and oligodendrocyte lineage cells in white matter pathology and plasticity after traumatic brain injury. Neuropharmacology 110, 654–659. [DOI] [PubMed] [Google Scholar]
- Bertoglio D, Verhaeghe J, Dedeurwaerdere S, Grohn O, 2017. Neuroimaging in animal models of epilepsy. Neuroscience 358, 277–299. [DOI] [PubMed] [Google Scholar]
- Cabeen R, Laidlaw D, Toga A, 2018. Quantitative imaging toolkit: software for interactive 3D visualization, data exploration, and computational analysis of neuroimaging datasets. Proc International Society for Magnetic Resonance in Medicine (ISMRM). [Google Scholar]
- Corrigan F, Mander KA, Leonard AV, Vink R, 2016. Neurogenic inflammation after traumatic brain injury and its potentiation of classical inflammation. J. Neuroinflammation 13 264–016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dinov I, Lozev K, Petrosyan P, Liu Z, Eggert P, Pierce J, et al. , 2010. Neuroimaging study designs, computational analyses and data provenance using the LONI pipeline. PLoS One 5 10.1371/journal.pone.0013070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engel J, Pitkanen A, Loeb JA, Dudek FE, Bertram EH, Cole AJ, et al. , 2013. Epilepsy biomarkers. Epilepsia 54 (Suppl. 4), 61–69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eroglu O, Deniz T, Kisa U, Atasoy P, Aydinuraz K, 2017. Effect of hypothermia on apoptosis in traumatic brain injury and hemorrhagic shock model. Injury 48, 2675–2682. [DOI] [PubMed] [Google Scholar]
- Fisher RS, van Emde Boas W, Blume W, Eiger C, Genton P, Lee P, et al. , 2005. Epileptic seizures and epilepsy: definitions proposed by the International League Against Epilepsy (ILAE) and the International Bureau for Epilepsy (IBE). Epilepsia 46, 470–472. [DOI] [PubMed] [Google Scholar]
- Frey LC, 2003. Epidemiology of posttraumatic epilepsy: a critical review. Epilepsia 44, 11–17. [DOI] [PubMed] [Google Scholar]
- Harris NG, Verley DR, Gutman BA, Sutton RL, 2016. Bi-directional changes in fractional anisotropy after experiment TBI: Disorganization and reorganization? Neuroimage 133, 129–143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heidelbaugh JJ, 2016. Endocrinology update: hypopituitarism. FP Essent. 451, 25–30. [PubMed] [Google Scholar]
- Herman ST, 2002. Epilepsy after brain insult: targeting epileptogenesis. Neurology 59, S21–6. [DOI] [PubMed] [Google Scholar]
- Immonen R, Kharatishvili I, Grohn O, Pitkanen A, 2013. MRI biomarkers for post-traumatic epileptogenesis. J. Neurotrauma 30, 1305–1309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jenkinson M, Beckmann CF, Behrens TE, Woolrich MW, Fsl, Smith SM, 2012. Neuroimage 62, 782–790. [DOI] [PubMed] [Google Scholar]
- Kharatishvili L, Nissinen JP, McIntosh TK, Pitkanen A, 2006. A model of post-traumatic epilepsy induced by lateral fluid-percussion brain injury in rats. Neuroscience 140, 685–697. [DOI] [PubMed] [Google Scholar]
- Kumar R, Husain M, Gupta RK, Hasan KM, Haris M, Agarwal AK, et al. , 2009. Serial changes in the white matter diffusion tensor imaging metrics in moderate traumatic brain injury and correlation with neuro-cognitive function. J. Neurotrauma 26, 481–495. [DOI] [PubMed] [Google Scholar]
- Laitinen T, Sierra A, Bolkvadze T, Pitkanen A, Grohn O, 2015. Diffusion tensor imaging detects chronic microstructural changes in white and gray matter after traumatic brain injury in rat. Front. Neurosci 9, 128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirzaalian H, Ning L, Savadjiev P, Pasternak O, Bouix S, Michailovich O, et al. , 2018. Multi-site harmonization of diffusion MRI data in a registration framework. Brain Imaging Behav. 12, 284–295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molaie AM, Maguire J, 2018. Neuroendocrine abnormalities following traumatic brain injury: an important contributor to neuropsychiatric sequelae. Front. Endocrinol. (Lausanne) 9, 176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pearn ML, Niesman IR, Egawa J, Sawada A, Almenar-Queralt A, Shah SB, et al. , 2017. Pathophysiology associated with traumatic brain injury: current treatments and potential novel therapeutics. Cell. Mol. Neurobiol 37, 571–585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phillips KF, Deshpande LS, DeLorenzo RJ, 2018. Hypothermia reduces mortality, prevents the calcium plateau, and is neuroprotective following status epilepticus in rats. Front. Neurol 9, 438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pitkanen A, Immonen R, 2014. Epilepsy related to traumatic brain injury. Neurotherapeutics 11, 286–296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pitkanen A, Ekolle Ndode-Ekane X, Lapinlampi N, Puhakka N, 2018. Epilepsy biomarkers - toward etiology and pathology specificity. Neurobiol. Dis 10.1016/j.nbd.2018.05.007 2018 May 18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toth P, Szarka N, Farkas E, Ezer E, Czeiter E, Amrein K, et al. , 2016. Traumatic brain injury-induced autoregulatory dysfunction and spreading depression-related neurovascular uncoupling: pathomechanisms, perspectives, and therapeutic implications. Am. J. Physiol. Heart Circ. Physiol 311, H1118–H1131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Vliet EA, Aronica E, Vezzani A, Ravizza T, 2018. Neuroinflammatory pathways as treatment targets and biomarker candidates in epilepsy: emerging evidence from preclinical and clinical studies. Neuropathol. Appl. Neurobiol 44 (February (1)), 91–111. [DOI] [PubMed] [Google Scholar]
- Wright DK, Johnston LA, Kershaw J, Ordidge R, O’Brien TJ, Shultz SR, 2017. Changes in apparent Fiber density and track-weighted imaging metrics in White Matter following experimental traumatic brain injury. J. Neurotrauma 34, 2109–2118. [DOI] [PubMed] [Google Scholar]
- Xu J, Rasmussen IA, Lagopoulos J, Haberg A, 2007. Diffuse axonal injury in severe traumatic brain injury visualized using high-resolution diffusion tensor imaging. J. Neurotrauma 24, 753–765. [DOI] [PubMed] [Google Scholar]