Abstract
The aim of this study was to identify for the first time single nucleotide polymorphisms (SNPs) associated with Haemonchus contortus resistance in Florida Native sheep, using a targeted sequencing approach. One hundred and fifty-three lambs were evaluated in this study. At the start of the trial, phenotypic records for fecal egg count (FEC), FAMACHA score, body condition score (BCS), and weight were recorded and deworming of sheep with levamisole (18 mg/kg of body weight) was performed. Ten days post-deworming (baseline) and 28 d post-baseline, a full hematogram of each sheep was obtained and FEC, FAMACHA score, BCS, and weight were assessed. Average daily gain was calculated at the end of the trial. Out of 153 animals, 100 sheep were selected for genotyping using a targeted sequencing approach. Targeted sequencing panel included 100 candidate genes for immune response against H. contortus. SNPs were discarded if call rate <95% and minor allele frequency ≤0.05. A mixed model was used to analyze the response variables and included the identity by state matrix to control for population structure. A contemporary group (age, group, and sex) was included as fixed effect. Bonferroni correction was used to control for multiple testing. Eighteen SNPs on chromosomes 1, 2, 3, 4, 6, 7, 11, 15, 18, 20, 24, and 26 were significant for different traits. Our results suggest that loci related to Th17, Treg, and Th2 responses play an important role in the expression of resistant phenotypes. Several genes including ITGA4, MUC15, TLR3, PCDH7, CFI, CXCL10, TNF, CCL26, STAT3, GPX2, IL2RB, and STAT6 were identified as potential markers for resistance to natural H. contortus exposure. This is the first study that evaluates potential genetic markers for H. contortus resistance in Florida Native sheep.
Keywords: Florida Native sheep, genome-wide association study, Haemonchus contortus, immune response, targeted sequencing
Introduction
Gastrointestinal infections are the main health problem for grazing sheep in the southern United States. The humid environmental conditions in this region are ideal for survival and growth of Haemonchus contortus and other gastrointestinal nematodes (GINs) of sheep (Miller et al., 1998). The predominant strategy for controlling GIN is anthelmintic treatment. However, anthelmintic resistance has reached epidemic proportions in small ruminant populations, and many cases the anthelmintic resistance has been reported to almost all available drugs (Miller and Craig, 1996; Zajac and Gipson, 2000; Kaplan, 2004; Howell et al., 2008; Goolsby et al., 2017). Breeding for parasite resistance offers many benefits to long-term sustainable sheep production. The main benefits of using selection to enhance parasite resistance in sustainable sheep breeding programs are that genetic change is permanent, the animal’s performance is improved, and infectivity of pastures is decreased, leading to additional benefits for susceptible and young sheep grazing within the same flock (Bishop, 2012). Traditionally, selection of resistant sheep to GIN has focused on estimated breeding values calculated from phenotypic and pedigree information (Goddard and Hayes, 2009). Breeding values for resistance to GIN based on fecal egg count (FEC) are now available for sheep breeds in Australia (www.sheepgenetics.org.au) and New Zealand (www.sil.co.nz). Phenotypic recording for selection for parasite resistance can be complemented with other indicator traits such as FAMACHA® score, packed cell volume (PCV), or hematocrit (HCT), eosinophil count (EC), and determination of IgA, IgM, and IgG levels (Bishop, 2012). With current technologies, incorporation of genotype information, using genetic markers in sheep breeding programs aimed to select individuals for enhanced resistance to GIN, is a promising faster approach that reduces the generation interval in the selection process and avoids the requirement for animals to be challenged with gastrointestinal parasites.
In the United States, several sheep breeds have been studied and appear to have naturally developed enhanced resistance to GIN based on FEC. These breeds include St. Croix, Katahdin, Barbados Black Belly, Gulf Coast Native, and Florida Native (Zajac et al., 1988; Amarante et al., 1999a,b; Baker and Gray, 2004; Estrada-Reyes et al., 2018). Florida Native sheep is a multipurpose (wool and meat), landrace breed locally adapted to the warm humid environmental conditions from Florida. It is believed that this breed was developed from sheep (Churra) introduced by Spanish when they founded St. Agustin in 1565. Sheep became feral for several decades until Florida ended the open range at the end of World War II (https://livestockconservancy.org/index.php/heritage/internal/cracker-sheep). According to the American Livestock Breeds Conservancy, this heritage breed is now on the endangered list of sheep breeds (http://livestockconservancy.org/). To ensure genetic diversity and long-term survival of the breed, protection of the current genetic stock is critical and conservation efforts are required to promote its breeding and production. Natural parasite resistance is a common feature of Florida Native sheep, and good health and reproductive performance have persisted for many generations. The genetics underlying resistance to GIN in Florida Native sheep have been studied without identifying the genes or gene variants controlling variation, with selection based on phenotypic measures such as FEC and PCV (Zajac et al., 1988; Amarante et al., 1999a,b).
Genome-wide association studies have successfully identified genomic variants associated with complex traits and biological pathways responsible for genetic variation with moderate to large effects in some livestock populations (Karim et al., 2000; Grisart et al., 2002). However, resistance to GNI is quantitative in nature with polygenic effects, and several studies aimed to identify the genetic variation controlling gastrointestinal parasite suggested a scattering of effects across the sheep genome (Kemper et al., 2011; McRae et al., 2014; Benavides et al., 2016; Sweeney et al., 2016). Many of the genomic regions identified from these studies are related to the sheep immune response against GNI.
The implementation of genomic markers associated to resistance to GIN could improve the selection of animals early in life and benefit the production costs. To date, results from several studies with other sheep breeds suggest that genetic architecture that underlines resistance to GIN relies on DNA polymorphisms within many genes. These studies suggest that resistance to GIN is quantitative in nature and polygenic (controlled by many genes with small effects) (McRae et al., 2014; Benavides et al., 2016; Sweeney et al., 2016). Exploration of genetic variation for resistance to GIN can be performed either with SNP-based genome scans or more specifically in candidate regions involved with innate and adaptive immune pathways related to GIN. Identification of variants such as SNPs in these candidate regions may help to identify a set of genetic markers significantly associated with parasite resistance. Thus, the objectives of this study were: 1) to use a targeted sequencing approach to identify SNPs in 100 genes related to the immune response during natural infections with H. contortus in Florida Native sheep; 2) to perform a SNP-based association analysis for FEC, FAMACHA score, HCT, red blood cell count (RBC), hemoglobin level (HGB), white blood cell count (WBC), neutrophil count (NEU), lymphocyte count (LC), monocyte count (MC), basophil count (BC), and EC; and 3) to identify the potential candidate genes associated to resistance to natural H. contortus infections in Florida Native sheep.
Materials and Methods
Sheep Population and Phenotypic Data
The research protocol was approved by the University of Florida Institutional Animal Care and Use Committee (approval number 201810108). All animals used in this study were from a commercial Florida Native sheep farm from Ocala, Florida. The selection flock was established in 1998 from an initial group of 80 rams ranked for H. contortus FEC, with high and low FEC rams mated with approximately 100 foundation dams. Since then, animals were selected for resistance to GIN solely on the basis of H. contortus FEC phenotyping and parentage recording. One hundred and fifty-three female and male sheep were tested for GINs under natural grazing conditions. Animals were born between December 2017 and February 2018 and naturally exposed to parasites since birth. They were approximately 3 and 5 mo old on average and were grouped based on age. At the beginning of the study, initial FEC, FAMACHA score, weight, and body condition score (BCS) were measured and animals were drenched with levamisole (18 mg/kg of body weight) the same day of initial screening. Then, animals returned to grazing conditions. Ten days post-deworming, FEC reduction was confirmed and FAMACHA score, weight, BCS, and a full hematogram of each sheep were performed. This date was used as baseline (0 d) for future phenotype collection dates. Finally, 28 d post-baseline (28 d), the same parasitological and hematological parameters were measured and ADG was determined based on weight and days under study. Identification of GIN genus was carried out from fresh fecal samples collected directly from the rectum of evaluated sheep as described by Roberts and O’Sullivan (1950) at start of the trial (initial FEC) and during baseline (0 d) and 28 d post-baseline (28 d). However, due to low and relatively constant numbers for the other species, only eggs of Haemonchus spp. were recorded and analyzed.
The Shapiro–Wilk test was used to test all variables for normality. Box–Cox transformation was performed for all the traits recorded to obtain a normal distribution of values. The R “car” library was used to estimate the power parameter λ and carry out square root transformation for FEC (Estrada-Reyes et al., 2018) and logarithmic transformation (log10) for RBC, HGB, WBC, NEU, LC, MC, BC, and EC. For all the traits, the different time points evaluated during the study (initial screening, 0 d, and 28 d) and the difference between 28 d post-baseline and baseline (28 d minus 0 d) were included in the analysis. Spearman phenotypic correlations among all the parameters evaluated were obtained using R software (https://cran.r-project.org/).
Genotyping Using Capture Sequencing
Genomic DNA from blood samples collected from the jugular vein using vacutainer tubes with anticoagulant EDTA at day 0 post-infection was isolated using DNeasy Blood & Tissue Kit (Qiagen, Valencia, CA) according the manufacturer’s instructions and stored at −20 °C. The DNA yield was calculated from a spectrophotometric measurement at 260 nm (NanoDrop‐1000, Thermo Scientific), and the purity was assessed using a ratio 260/280 nm.
DNA samples (250 ng/μL per sample) from 100 selected sheep samples with the highest (>1,000 eggs/g of feces, n = 52) and lowest (<900 eggs/g of feces, n = 48) FEC (Supplementary Figure 1) were sent to RAPiD Genomics (Gainesville, FL) for sequencing of 100 genes using DNA Capture sequencing. Briefly, this approach uses a hybrid capture-based enrichment sequencing in which regions of interest are captured by sequence-specific hybridization probes. A focused panel that contained a set of 100 genes was used and custom designed to include genomic regions of interest. Selection of the gene panel was performed based on results from previous studies in sheep (Araujo et al., 2009; MacKinnon et al., 2009; Silva et al., 2012; Venturina et al., 2013; McRae et al., 2014; Benavides et al., 2015, 2016; Sweeney et al., 2016; Berton et al., 2017; Estrada-Reyes et al., 2018). Also, genes related to the immune response against H. contortus and other GIN were considered as candidates for sequencing. The sheep genome assembly from Ovis aries (Oar_v4.0) available at the National Center for Biotechnology Information (NCBI) genome browser was used to design the biotinylated 120-mer probes used for sequencing. Probes were designed to capture sequences at each target gene.
For library preparation, Illumina’s guidelines were followed and Nextera tagmentation, which converts DNA into adapter-tagged libraries, was used. Libraries were denatured and biotin-labeled probes were used for hybridization. Biotinylated probes were captured by streptavidin-coated beads. Then, DNA fragments bound to the streptavidin-coated beads were magnetically pulled down and enriched DNA fragments were eluted. After library preparation, libraries were captured by surface-bound oligos complimentary to library adapters. Then, each library fragment was amplified into distinct, clonal clusters through bridge amplification. After cluster generation, templates were ready for sequencing. Illumina sequencing by synthesis chemistry was used for DNA Capture sequencing and samples were then sequenced using a depth of 500–1,000×. Data from sequencing was demultiplexed, cleaned, and trimmed. The 3′ ends were trimmed by removing low quality bases with <20 quality score reads. Clean reads were aligned to genome with MOSAIK software (Lee et al., 2014). For SNP calling, Freebayes software was used and VCFtools (Danecek et al., 2011) was used to generate VCF files. Supplementary Table 1 presents a summary of the genes evaluated in our study.
Quality Control and Association Analysis
Quality control of SNP data set was performed in JMP Genomics 6.0 software (SAS Institute Inc., Cary, NC). A total of 1,546 SNPs were available for analysis. Markers were removed based on genotype call rates per marker (<95%) and minor allele frequency ≤0.05, and the final marker dataset consisted of 1,277 SNPs. For the association study, a mixed model was used to evaluate the H. contortus FEC, FAMACHA score, HCT, RBC, HGB, WBC, NEU, LC, MC, BC, and EC. The GenABEL package (Aulchenko et al., 2007) from R software was used to perform the association analysis. Fixed effects included SNP and a contemporary group conformed by age of the animal, sex, and group. To control for population structure, the genomic relationship matrix was included in the analysis as random effect and it was calculated from SNP information. The mixed model used for the association analysis was as follows:
where y represents the vector of the phenotypic observation for each animal; b is the vector of fixed effects, u is the vector of random animal polygenic effect, u ∼ N (0, Gσ u2), being G the additive genomic relationship matrix; a represents the fixed SNP additive effect, and e represents the residual vector, e ∼ N (0, Iσ e2). The matrices X, Z, and W represent the incidence matrices for b, u, and a, respectively. SNP markers were tested one at a time, and a Bonferroni correction was used to control for multiple testing as follows:
Bioinformatics Analysis of Significant SNPs
Information regarding the identity of the genes underlying the significant SNPs associated with different traits was obtained from the sheep genome assembly (Oar_v4.0) available at the NCBI genome browser. Then, significant associated SNPs were screened for putative function to evaluate the possible changes in the free energy and the secondary structure of the ovine mRNA. The mRNA sequences from Oar_v4.0 available at the NCBI genome browser were downloaded for the genes with significant SNPs. For each mRNA sequence, a segment of 500 base pairs harboring the SNP in the middle of the sequence was used for prediction of mRNA secondary structures and molecular stability (free energy, G) using the RNA fold web server (Lorenz et al., 2011).
From each association analysis, genes with significant SNPs associated with the evaluated traits were analyzed using Gene Ontology software (Ashburner et al., 2000) to identify molecular function and immunological pathways. These genes were also used in the construction of a gene network related to H. contortus infection. The Cytoscape v3.7.1 software was used for gene network graphics (Shannon et al., 2003).
Results and Discussion
Phenotypic Data
The GIN genus identified from eggs were the following: Haemonchus spp, Moniezia spp, Coccidia spp, and Strongyloides spp. Table 1 shows the descriptive statistics for parasitological and hematological traits available for this study. Table 2 presents the phenotypic correlations between H. contortus FEC and other traits. Significant moderate negative correlations were observed between H. contortus FEC (initial and 28 d) and RBC, and HGB (P < 0.01). For HCT at 28 d, a significant moderate negative correlation was observed with initial H. contortus FEC (P = 0.00005), and with H. contortus FEC at 28 d (P = 0.00009). Significant moderate positive correlated responses were observed between initial H. contortus FEC and WBC (0 and 28 d) (P = 0.0001 and P = 0.0008, respectively). Also, H. contortus FEC at 28 d was positively correlated with WBC at 0 and 28 d (P = 0.0003 and P = 0.0002, respectively). Finally, significant moderate positive correlations were observed between ADG and H. contortus FEC (initial and 28 d) (P = 0.03 and P = 0.013, respectively). For FAMACHA, BC, and EC, no significant correlations were observed with H. contortus FEC (initial and 28 d). A significant strong positive correlation (0.89) (P = 2.2 × 10−16) between HGB at 28 d and RBC at 28 d was observed. Supplementary Table 1 shows the phenotypic correlated responses between all the traits under study.
Table 1.
Descriptive statistics for H. contortus fecal egg count (FEC, eggs/gram) FAMACHA score, average daily gain (ADG), body condition score (BCS), hematocrit (HCT, %), red blood cell count (RBC, M/μL), hemoglobin level (HGB, g/dL), white blood cell count (WBC, K/μL), neutrophil count (NEU, K/μL), lymphocyte count (LC, K/μL), monocyte count (MC, K/μL), basophil count (BC, K/μL), and eosinophil count (EC, K/μL) at −10 d baseline, 0 d (baseline), and 28 d post-baseline in Florida Native sheep
| Trait | Time | n | Mean | SD | Min | Max |
|---|---|---|---|---|---|---|
| FEC, eggs/gram | Initial FEC (−10 d) | 100 | 1,926.00 | 312.6 | 0.00 | 15,800.00 |
| FEC (0 d) | 100 | 21.00 | 7.10 | 0.00 | 500.0 | |
| FEC (28 d) | 100 | 1,593.00 | 270.5 | 0.00 | 19,300.00 | |
| FAMACHA, score | Initial FAMACHA (−10 d) | 100 | 2.93 | 0.72 | 2.00 | 4.00 |
| FAMACHA (0 d) | 100 | 2.87 | 0.63 | 2.00 | 5.00 | |
| FAMACHA (28 d) | 100 | 2.75 | 0.68 | 1.00 | 4.00 | |
| ADG, g | 100 | 167.83 | 58.97 | 49.89 | 344.73 | |
| BCS, score | Initial BCS (−10 d) | 100 | 2.32 | 0.26 | 1.75 | 3.00 |
| BCS (0 d) | 100 | 2.42 | 0.28 | 2.00 | 3.25 | |
| BCS (28 d) | 100 | 2.37 | 0.26 | 1.75 | 3.00 | |
| HCT, % | HCT (0 d) | 100 | 41.00 | 0.40 | 24.00 | 52.00 |
| HCT (28 d) | 100 | 40.00 | 0.50 | 22.00 | 50.00 | |
| RBC, M/μL | RBC (0 d) | 100 | 13.11 | 1.22 | 10.64 | 16.40 |
| RBC (28 d) | 100 | 13.42 | 1.80 | 7.63 | 18.17 | |
| HGB, g/dL | HGB (0 d) | 100 | 11.82 | 1.01 | 8.60 | 14.00 |
| HGB (28 d) | 100 | 11.99 | 1.37 | 7.10 | 15.10 | |
| WBC, K/μL | WBC (0 d) | 100 | 10.30 | 2.78 | 5.22 | 21.68 |
| WBC (28 d) | 100 | 10.24 | 2.24 | 4.60 | 15.06 | |
| NEU, K/μL | NEU (0 d) | 100 | 2.73 | 1.66 | 0.55 | 12.26 |
| NEU (28 d) | 100 | 2.40 | 0.79 | 1.00 | 4.36 | |
| LC, K/μL | LC (0 d) | 100 | 5.48 | 1.90 | 1.63 | 9.62 |
| LC (28 d) | 100 | 5.86 | 1.97 | 1.28 | 10.27 | |
| MC, K/μL | MC (0 d) | 100 | 1.42 | 0.72 | 0.21 | 3.33 |
| MC (28 d) | 100 | 1.65 | 0.74 | 0.24 | 4.20 | |
| BC, K/μL | BC (0 d) | 100 | 0.07 | 0.10 | 0.01 | 0.61 |
| BC (28 d) | 100 | 0.05 | 0.06 | 0.01 | 0.43 | |
| EC, K/μL | EC (0 d) | 100 | 0.48 | 0.47 | 0.01 | 3.15 |
| EC (28 d) | 100 | 0.36 | 0.24 | 0.02 | 1.19 |
Table 2.
Phenotypic correlations between initial (−10 d) or 28 d H. contortus fecal egg count (FEC) and neutrophil count (NEU) FAMACHA (28 d), hematocrit (HCT), red blood cell count (RBC, 28 d), hemoglobin level (HGB, 28 d), white blood cell count (WBC, 0 and 28 d), basophil count (BC 0 d), and eosinophil count (EC 28 d)1
| FAMACHA | HCT | RBC | HGB | WBC | NEU | BC | EC | |||
|---|---|---|---|---|---|---|---|---|---|---|
| 28 d | 28 d | 28 d | 28 d | 0 d | 28 d | 0 d | 0 d | 28 d | ||
| FEC | Initial (−10 d) | −0.004 | −0.4 | −0.36 | −0.36 | 0.37 | 0.32 | 0.29 | 0.03 | 0.02 |
| 28 d | −0.14 | −0.38 | −0.27 | −0.27 | 0.34 | 0.14 | 0.2 | 0.04 | 0.17 |
1Values in bold indicate significance at P < 0.05.
Genomic Regions Identified by the Association Analysis
A total of 23 SNPs covering 14 genes were associated with H. contortus FEC, FAMACHA, RBC, HGB, WBC, NEU, BC, and EC. No significant variants were detected for HCT, LC, and MC. A summary of the significant SNPs associated with these traits is presented in Table 3. Overall, associated SNPs identified in this study were located on intergenic and untranslated regions (UTRs) of 14 genes in OAR1, 2, 3, 4, 6, 7, 11, 15, 18, 19, 20, 24, and 26 (Fig. 1). The proportion of variance explained by majority of the significant SNPs associated with FEC, FAMACHA score, WBC, RBC, BC, and EC was less than 0.2. Thus, based on the current and previous studies, it is possible that many genes from the immune response (with small effects) are contributing to parasite resistance/susceptibility in Florida Native sheep populations.
Table 3.
Significant SNPs associated with H. contortus fecal egg count (FEC, eggs/gram), FAMACHA score, red blood cell count (RBC, M/μL), hemoglobin level (HGB, g/dL), white blood cell count (WBC, K/μL), neutrophil count (NEU, K/μL), lymphocyte count (LC, K/μL), monocyte count (MC, K/μL), basophil count (BC, K/μL), and eosinophil count (EC, K/μL) in Florida Native sheep1
| Trait | OAR | Position (bp) | Gene name | SNP | −log10 P-values | Estimate | Proportion of variance explained | MAF | Mutation |
|---|---|---|---|---|---|---|---|---|---|
| FEC (initial −10 d) | 2 | 127,159,353 | Exon 24 ITGA4 | G/A | 4.108 | 21.58 | 0.14 | 0.16 | Synonymous |
| FEC (initial −10 d) | 11 | 41,860,577 | Intron 13 STAT3 | G/C | 4.052 | 15.94 | 0.10 | 0.11 | Intronic |
| FEC (initial −10 d) | 15 | 55,310,482 | 5′UTR MUC15 | T/C | 3.773 | 24.22 | 0.11 | 0.12 | UTR |
| FEC (28 d) | 3 | 180,166,632 | 5′UTR IL12RB2 | G/A | 4.065 | 21.58 | 0.14 | 0.11 | UTR |
| FEC (28 d) | 3 | 180,167,554 | 5′UTR IL12RB2 | C/G | 3.531 | 15.94 | 0.13 | 0.12 | UTR |
| FEC (28 d) | 6 | 15,681,195 | Exon 6 CFI | G/A | 4.009 | 24.22 | 0.15 | 0.11 | Synonymous |
| FAMACHA (28 d) | 3 | 180,148,777 | Exon 12 IL12RB2 | G/A | 3.616 | −0.33 | 0.13 | 0.40 | UTR |
| FAMACHA (28 d) | 7 | 73,930,804 | 5′UTR GPX2 | A/G | 3.531 | −0.36 | 0.03 | 0.49 | Synonymous |
| RBC (28 d) | 18 | 26,116,868 | Exon 18 IL16 | C/T | 4.558 | −0.08 | 0.16 | 0.11 | Synonymous |
| HGB (28 d) | 18 | 26,116,868 | Exon 18 IL16 | C/T | 3.563 | −1.60 | 0.42 | 0.11 | Synonymous |
| WBC (0 d) | 3 | 162,039,038 | 3′UTR STAT6 | G/A | 3.42 | 0.06 | 0.02 | 0.46 | UTR |
| WBC (28 d) | 6 | 49,768,053 | 5′UTR PCDH7 | G/T | 4.781 | −0.008 | 0.17 | 0.11 | UTR |
| WBC (28 d) | 6 | 49,768,057 | 5′UTR PCDH7 | G/T | 3.61 | −0.006 | 0.12 | 0.11 | UTR |
| WBC (28 d) | 15 | 55,310,748 | 5′UTR MUC15 | C/T | 3.999 | 0.0005 | 0.14 | 0.38 | UTR |
| NEU (0 d) | 6 | 90,454,870 | 3′UTR CXCL10 | A/G | 3.809 | 0.46 | 0.14 | 0.12 | UTR |
| NEU (0 d) | 20 | 26,766,451 | 3′UTR TNF | C/T | 3.668 | 0.67 | 0.13 | 0.30 | UTR |
| BC (0 d) | 24 | 33,935,613 | 3′UTR CCL26 | C/T | 4.335 | −1.65 | 0.16 | 0.46 | UTR |
| EC (28 d) | 2 | 127,200,458 | Intron 10 ITGA4 | A/G | 3.687 | 0.25 | 0.13 | 0.12 | Intronic |
| EC (28 d) | 2 | 14,799,731 | Exon 2 TLR3 | T/C | 4.094 | 0.26 | 0.15 | 0.11 | Synonymous |
1For each SNP, the chromosome (OAR), chromosomal position (bp), gene name and region, base pair substitution (SNP), estimate of SNP effect, proportion of variance explained by SNP, minor allele frequency (MAF), and mutation type (UTR, synonymous or intronic) are presented.
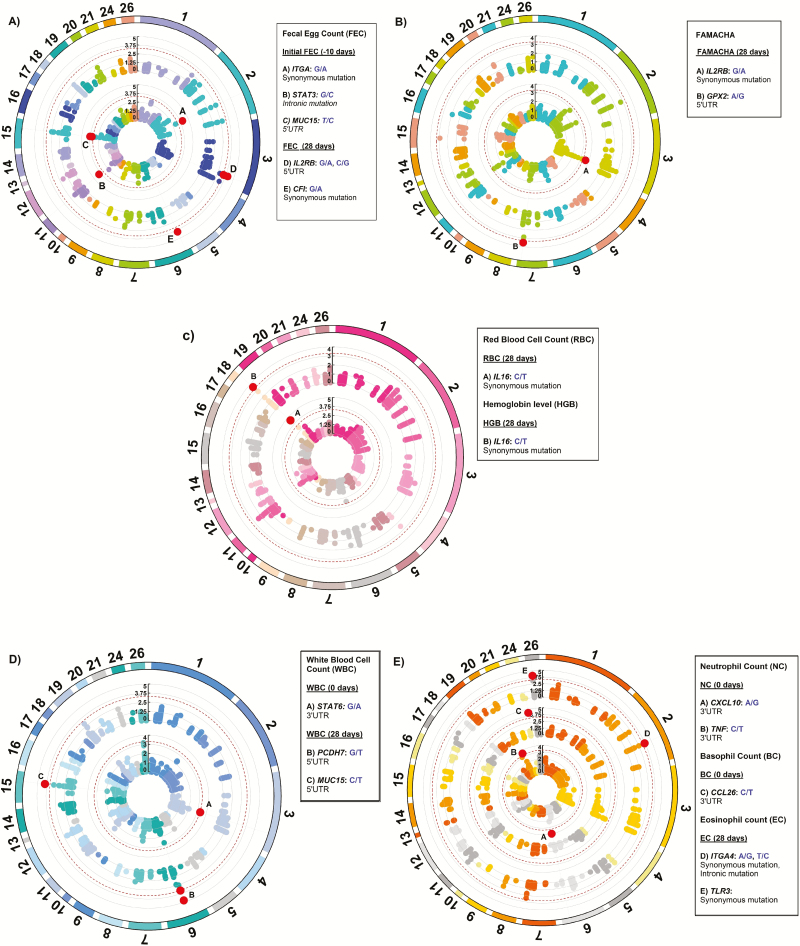
Figure 1.
Manhattan plots for natural H. contortus fecal egg count (A, FEC), FAMACHA score (B), red blood cell count (C, RBC), hemoglobin level (C, HGB), white blood cell count (D, WBC), neutrophil count (E, NEU), basophil count (E, BC), and eosinophil count (E, EC). The 3.3 -log10 P-value threshold was indicated with the red dotted line. The red dots indicate the significant SNPs identified above the threshold for each trait.
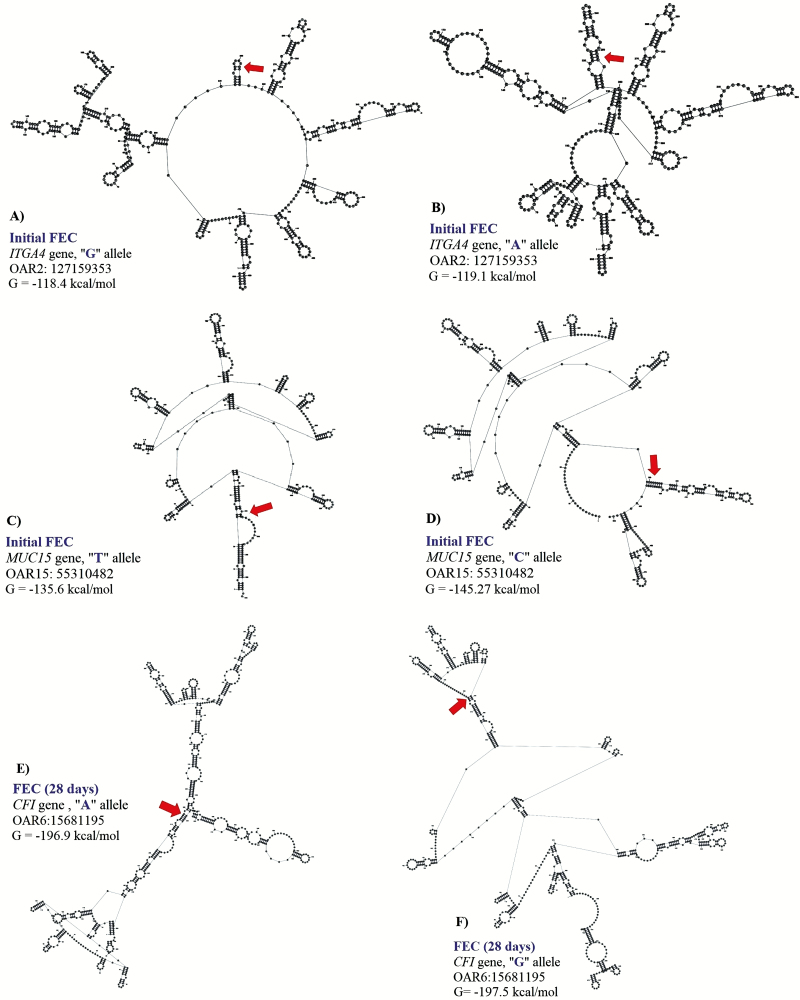
For initial H. contortus FEC, SNPs located in exon 24 of ITGA4 (OAR2:127159353), intron 13 of STAT3 (OAR11:41860577), and the 5′ UTR of MUC15 (OAR15:55310482) genes were significant. These proportions of variance explained by these SNPs was 0.14, 0.10 and 0.11, respectively, for initial H. contortus FEC. The OAR2:127159353 is a G to A substitution which codes for a synonymous codon (Val), where the GG genotype (1,179.4 ± 938.37 eggs/g of feces) had lower FEC when compared with the AG (3,239.94 ± 2,094.891 eggs/g of feces) and AA (6,890.1 ± 4,099.9 eggs/g of feces) genotypes. For this polymorphism, moderate changes in mRNA stability were observed for the “G” allele (−118.4 kcal/mol) when compared with the “A” allele (−119.1 kcal/mol), with great changes in mRNA secondary structure.
The OAR11:41860577 generates a G to C substitution, and OAR15:55310482 confers a base pair change from T to C. For OAR15:55310482, only two genotypes where observed, and the TT genotype (945.6 ± 804.1 eggs/g of feces) had lower FEC when compared to the CT genotype (4,444.4 ± 2,300.3 eggs/g of feces). For this SNP, the “T” allele had greater mRNA stability (−135.6 kcal/mol) when compared with the “C” allele (−145.27 kcal/mol) and strong changes in mRNA structure were observed (Fig. 2).
Figure 2.
Change in the free energy of the thermodynamic ensemble for the different mRNA variants observed for H. contortus fecal egg count (FEC) in Florida Native sheep. Predicted secondary structure of mRNA of 500 bp segment of ITGA4 (A: G allele and B: A allele), MUC15 (C: T allele and D: C allele), and CFI (E: A allele and F: G allele) genes.
Integrin subunit alpha 4 (ITGA4) gene codes for a cell adhesion molecule related to eosinophil mobilization (Rothenberg et al., 2001). Therefore, the protein complex co-formed by ITGA4 protein and beta 1 or beta 7 subunits regulates cell motility and migration. Thus, it is possible that eosinophils use this protein complex to be mobilized to the area of infection through IL-5 independent or dependent mechanisms. Eosinophils are produced in the bone marrow and are commonly found in almost all the sheep gastrointestinal tract. Sheep eosinophilia is related to reduction in H. contortus larvae establishment (Balic et al., 2006; Terefe et al., 2007). In calves exposed to Ostertagia, Cooperia, and Nematodirus spp., higher levels of ITGA4 expression is observed in the mesenteric lymph nodes of resistant animals. In peripheral blood from Chinese indigenous white goats exposed to H. contortus, ITGA4 gene was identified as upregulated in resistant individuals with lower FEC when compared with susceptible animals (Bhuiyan et al., 2017).
The STAT3 gene belongs to the STAT family genes known as signal transducer and activator of transcription. STAT3 protein can be activated by phosphorylation in response to several key activators such as IL-6, IL-10, IL-23, IL-21, and IL-11 cytokines to initiate Th17 signaling and IL17 production. Stronger Th17 response is observed in resistant Caribbean hair sheep at 3 d post-challenge with H. contortus (MacKinnon et al., 2009). Therefore, excretory and secretory proteins from H. contortus larvae are linked to increased Th17 activation and IL17 production (Gadahi et al., 2016). Thus, after initial mechanisms of parasite rejection and complement activation, Th17 response is activated to regulate pathogen clearance and tissue inflammation.
Mucin 15 (MUC15) gene produces a high-molecular-weight membrane-bound glycoprotein consisting of a mucin peptide backbone and O-linked oligosaccharides. Epithelial surfaces of the gastrointestinal tract are composed of these types of proteins and other mucins to protect against GINs and represent the first line of defense against pathogens (Moncada et al., 2003). It is possible that some of these types of mucins are ligands for GIN adhesins (McGuckin et al., 2011). In Dorper × Red Maasai sheep, MUC15 was identified as potential candidate marker for FEC (Benavides et al., 2015). Thus, ITGA4 and MUC15 could play an important role during the initial establishment and rejection of H. contortus in Florida Native sheep.
For H. contortus FEC at 28 d and the difference between 28 and 0 d, the same significant SNPs were identified within the 5′UTR of IL2RB (OAR3:180166632 and OAR3: 180167554) and exon 6 of CFI (OAR6: 15681195) genes. The proportion of variance explained by these polymorphisms was 0.14, 0.13, and 0.15, respectively, for FEC at 28 d. These SNPs are substitutions from G to A, C to G, and G to A, respectively. No significant changes were observed in the stability and secondary structure of the mRNA for OAR3:180166632 in the IL2RB gene. For OAR3: 180167554, the CC genotype (3,451.9 ± 1,725.2 eggs/g of feces) presented more FEC when compared with the GC (2,397.3 ± 1,793.3 eggs/g of feces) and GG (677.2 ± 539.3 eggs/g of feces) genotypes, with no significant changes in stability and secondary structure of the mRNA (Fig. 2).
The OAR6: 15681195 is a synonymous mutation with a minor change in mRNA stability of 0.6 kcal/mol but strong impact on the mRNA secondary structure (Fig. 2). For this polymorphism, the GG genotype (249.9 ± 124.9 eggs/g of feces) had lower FEC when compared with the AG (2,370.7 ± 1,775.2 eggs/g of feces) and AA (1,142.1 ± 909.8 eggs/g of feces) genotypes.
Complement factor I (CFI) gene encodes a serine protease implicated in regulation of the complement cascade. In hair sheep from the Canary Islands, high expression of CFI gene was observed in resistant animals (Guo et al., 2016). In our study, a genetic variant within this gene was associated with FEC at 28 d suggesting that complement activation is one of the earliest events in the Florida Native sheep immune responses to protect against H. contortus.
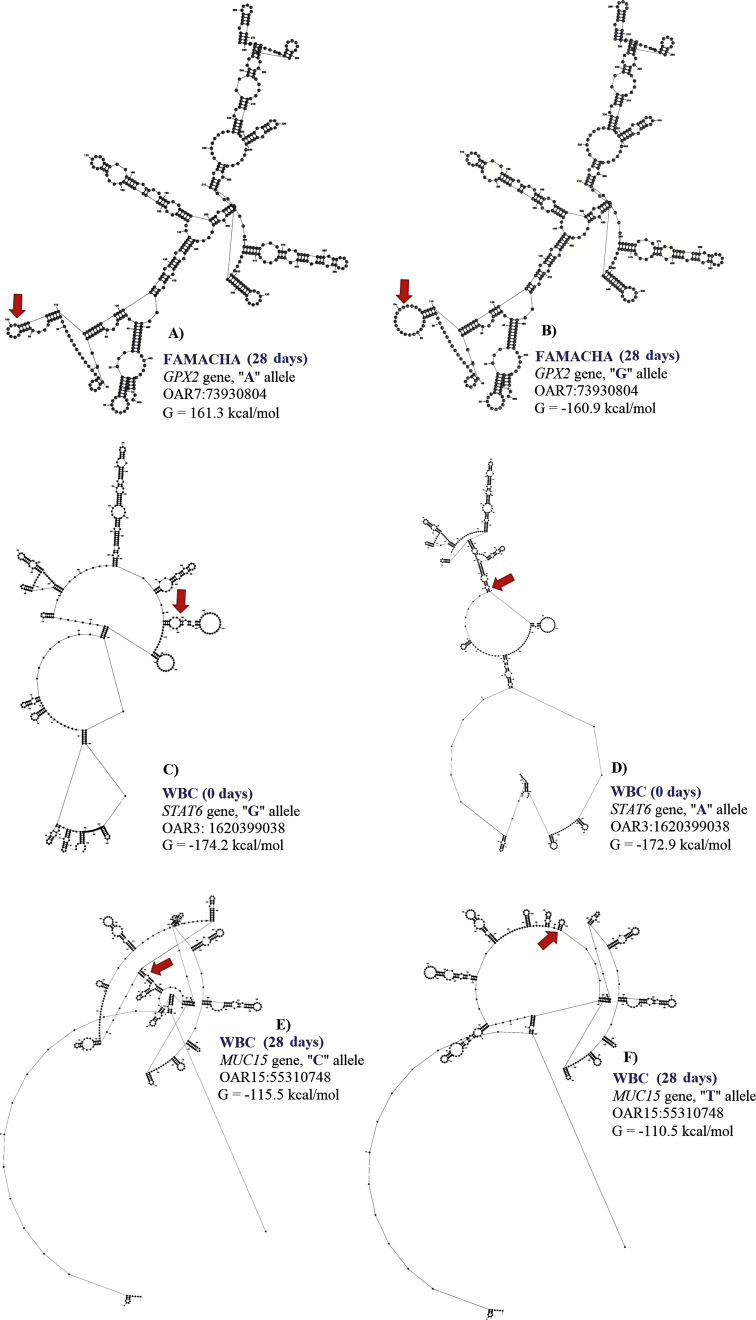
For FAMACHA at 28 d, a significant SNP (OAR3:180148777) in exon 12 of IL2RB associated with this trait. The proportion of variance explained by OAR3:180148777 was 0.13. This SNP (G/A) is part of a synonymous codon and no significant changes in stability and secondary structure of the mRNA were observed. For this locus, the GG genotype (2.11 ± 0.64) had lower FAMACHA score when compared with AG (2.85 ± 0.67) and AA (2.95 ± 0.58) genotypes. Also, for FAMACHA at 28 d, one SNP (OAR7:73930804) in the 3′UTR of GPX2 gene was significantly associated with this trait and explained 0.03 of the proportion of variance observed. The OAR7:73930804 generates an A to G substitution which results in minor changes in mRNA stability (a difference of −0.4 kcal/mol between variants) and mRNA secondary structure (Fig. 3).
Figure 3.
Change in the free energy of the thermodynamic ensemble for the different mRNA variants observed for FAMACHA score and white blood cell count (WBC, 0 and 28 d) in Florida Native sheep. Predicted secondary structure of mRNA of 500 bp segment of GPX2 (A: G allele and B: A allele), STAT6 (C: G allele and D: A allele), and MUC15 (E: C allele and F: T allele) genes.
Similar findings have been observed in Santa Inȇs sheep exposed to natural H. contortus infections, where genetic variants within IL2RB gene are associated to HCT (Berton et al., 2017). IL2RB protein is part of the IL-2 receptor complex (IL2R) co-formed by IL2RA, IL2RB, and IL2Rγ subunits, and it is activated by IL2 cytokine. This cytokine activates Treg cells, a subpopulation of T cells responsible for maintenance of immunological self-tolerance and homeostasis through immune suppression. Long-term infection with GINs is linked to activation of Treg populations (Smits and Yazdanbakhsh, 2007). The exact role of Treg response during H. contortus infections in Florida Native sheep remains unclear but a faster switch from Th1 to Th2/Treg response is observed in resistant Suffolk sheep exposed to Teladorsagia circumcincta (Hassan et al., 2011).
The GPX2 gene encodes the glutathione peroxidase 2, which catalyzes the reduction of peroxides and protects against oxidative damage in the epithelium of the gastrointestinal tract. The generation of host oxidants such as reactive oxygen and nitrogen species is important for parasite control (Ingham et al., 2008; Patel et al., 2009). In Perendale sheep, expression of GPX2 was upregulated in the duodenum of susceptible individuals when compared to resistant animals (Keane et al., 2006). In contrast, expression of GPX2 gene was upregulated in resistant sheep infected with H. contortus (Menzies et al., 2010) during initial days of infection. Similar findings were observed in resistant Australian sheep, where an increase in the expression of GPX2 gene is related to H. contortus challenge during the first days of infection (Lees et al., 2011). Results from these studies suggest that increase of GPX2 expression is essential during larval expulsion in the first days of infection.
For RBC and HGB at 28 d, the same SNP located in IL16 gene (OAR18: 26116868, C/T) was associated with both traits. The proportion of variance explained by OAR18: 26116868 was 0.16 and 0.12 for these traits, respectively. This SNP codes for a synonymous codon (Ser) which did not have significant changes in stability and secondary structure of the mRNA (Fig. 3). For OAR18: 26116868 only two genotypes were observed, and the CC genotype presented higher RBC (13.66 ± 1.58 M/μL) and HGB (12.17 ± 1.21 g/dL) when compared with the CT genotype (11.48 ± 1.75 M/μL and 10.57 ± 1.67 g/dL, respectively).
IL16 cytokine is related to inflammatory response and its synthesis is induced by T lymphocytes upon antigen exposure. Also, IL-16 promotes CD4+ T-cell production and it is commonly known as lymphocyte chemoattractant factor (O’Shea et al., 2013). In susceptible Perendale sheep selected for high FEC, expression of IL16 gene was downregulated in duodenum tissue (Diez-Tascón et al., 2005). In contrast, in Angus yearlings infected with gastrointestinal parasites, IL16 gene was highly expressed in the mesenteric lymph nodes of susceptible individuals (Araujo et al., 2009).
For WBC at 0 d, one SNP (OAR3: 162039038) in the 3′UTR of STAT6 gene was associated with this trait and explained 0.12 of the proportion of variance observed. For OAR3: 162039038, the G to A substitution results in moderate changes in mRNA stability (−1.3 kcal/mol) but great impact on mRNA secondary structure (Fig. 4).
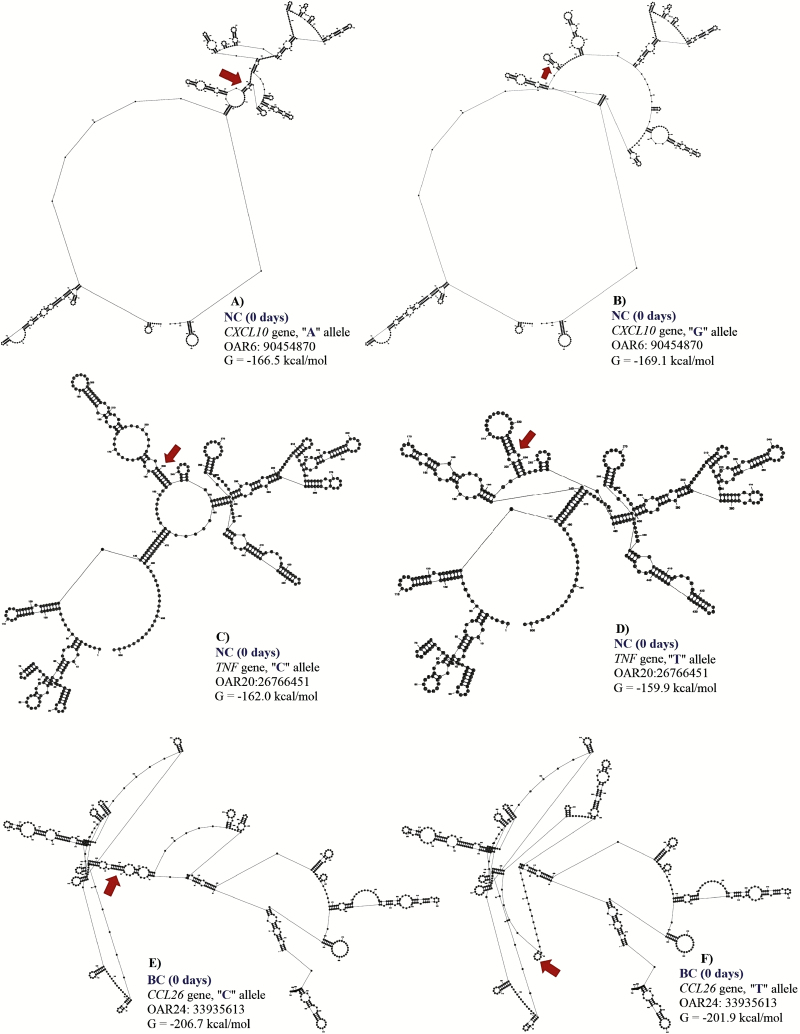
Figure 4.
Change in the free energy of the thermodynamic ensemble for the different mRNA variants observed for neutrophil count (NEU, 0 d) and basophil count (BC, 0 d) in Florida Native sheep. Predicted secondary structure of mRNA of 500 bp segment of CXCL10 (A: A allele and B: G allele), TNF (C: C allele and D: T allele), and CCL26 (E: C allele and F: T allele) genes.
Several studies have suggested that mechanisms related to STAT6 activation via IL4 receptor are required for the expulsion of gastrointestinal parasites in sheep (Urban et al., 1998; Venturina et al., 2013). Signal transducer and activator of transcription 6 (STAT6) gene encodes a protein that mediates Th2 signaling. Th2 response is linked to resistance to GIN in sheep (Pernthaner et al., 2005). Commonly features of Th2 response include IL4, IL5, and IL13 cytokine production, eosinophil infiltration, and activation mediated by IL5, and IgE production. IgE antibodies bind to antigens expressed in the membrane of gastrointestinal parasites. Then, Fc receptors (FceRI and II) located on the surface of eosinophils recognize IgE antibodies, leading to degranulation and releasing of cytotoxic proteins that damage the parasite membrane (Meeusen and Balic, 2000; McRae et al., 2015). In wool sheep infected with H. contortus, stronger Th2 response is observed at 27 d post-infection, with upregulation of IL4 and downregulation of IFNγRB expression.
Two SNPs (OAR6:49768053 and OAR6:49768057) in the 5′UTR region of PCDH7 gene were associated to WBC at 28 d. The proportion of variance explained by both SNPs was 0.17 and 0.12, respectively. Both polymorphisms generate G to T substitutions with no significant changes in stability and secondary structure of the mRNA. For OAR6:49768053, the GG genotype (10.59 ± 2.34 K/μL) presented higher WBC at 28 d when compared with the GT (10.06 ± 2.17 K/μL) and the TT (6.97 ± 0.40 K/μL) genotypes. For OAR6:49768057, the GT genotype (10.96 ± 2.36 K/μL) presented higher WBC when compared with the GG (10.59± 2.34 K/ μL) and the TT (9.65 ± 2.22 K/μL) genotypes.
Protocadherin 7 (PCDH7) gene encodes a protein from the cadherin family. Cadherins are cell adhesion molecules responsible for cell interactions and maintenance of tissue structure and morphogenesis (Li and Gasbarre, 2010). Studies in Angus yearlings observed upregulation of PCDH7 expression in susceptible animals infected with GINs. However, the exact role of PCDH7 protein during H. contortus infections in Florida Native sheep remains unknown.
One SNP (OAR15: 55310748) in the 5′UTR of MUC15 gene was also associated with WBC at 28 d, which results in a base pair change from C to T. The OAR15: 55310748 explained 0.14 of the proportion of variance observed for this trait. This SNP is a putative functional SNP with great changes in mRNA stability (5 kcal/mol) and mRNA secondary structure (Fig. 4). For this locus, the CC genotype (10.72 ± 2.50 K/μL) presented higher number of WBC when compared with the CT (10.01 ± 2.09 K/μL) and TT (10.37 ± 2.25 K/μL) genotypes.
For NEU at 0 d, two SNPs (OAR6:90454870 and OAR20:26766451) in the 3′ UTR of CXCL10 and TNF genes were associated with this trait. These SNPs explained 0.14 and 0.13 of the proportion of variance observed. The OAR6:90454870 is an A to G substitution, and OAR20: 26766451 is a C to T substitution. Both polymorphisms are putative functional SNPs with moderate changes in mRNA stability (a difference of 2.6 kcal/mol and −2.1 kcal/mol between variants, respectively) but great impact on mRNA structure. For the OAR6:90454870, the “A” allele presented higher mRNA stability (−166.5 kcal/mol) than the “G” allele (−169.1 kcal/mol) (Fig. 5). The AA genotype (2.58 ± 1.62 K/μL) had lower NEU when compared with the AG (3.19 ± 0.68 K/μL) and the GG (6.82 ± 0.72 K/μL) genotypes.
Figure 5.
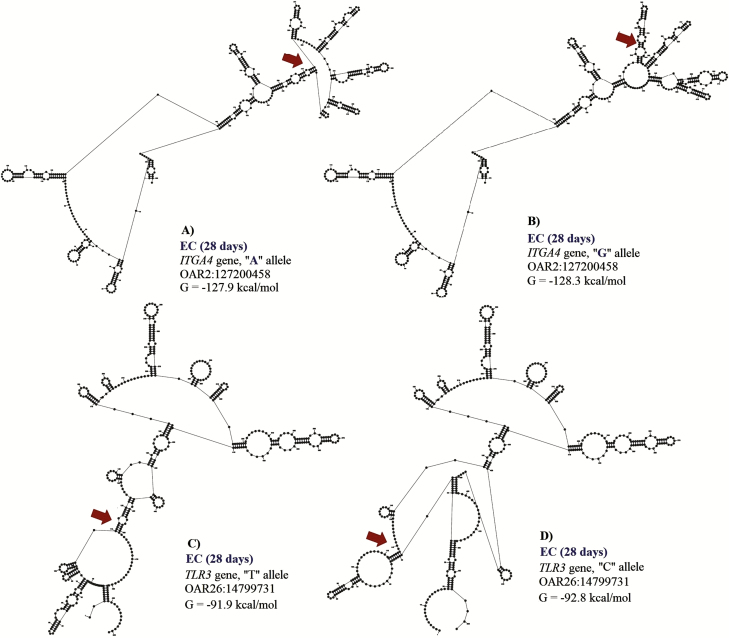
Change in the free energy of the thermodynamic ensemble for the different mRNA variants observed for eosinophil count (EC, 28 d) in Florida Native sheep. Predicted secondary structure of mRNA of 500 bp segment of ITGA4 (A: A allele and B: G allele), and TLR3 (C: T allele and D: C allele) genes.
For OAR20:26766451 located on TNF gene, the CC genotype (3.20 ± 1.98 K/μL) had higher NEU when compared to the CT (2.27 ± 0.91 K/ μL) and the TT (1.94 ± 0.61 K/μL) genotypes, with minor changes in mRNA stability but moderate impact on mRNA secondary structure (Fig. 5).
The TNF and CXCL10 genes encode proteins related to inflammatory response. TNF protein is commonly secreted by macrophages. CXCL10 protein is produced by monocytes and its production is induced by interferon gamma (IFNγ). In Santa Inȇs sheep, polymorphisms within CXCL10 gene have been associated with platelet count. Thus, it is possible that TNF and CXCL10 genes are related to inflammatory responses in Florida Native sheep exposed to natural H. contortus infections.
For BC at 0 d, one SNP (OAR24:33935613) in the 3′UTR region of CCL26 gene was associated with this trait. This SNP explained 0.16 of the proportion of variance observed. The C to T substitution from OAR24: 33935613 results in a great change in mRNA stability of −4.8 kcal/mol and has a great impact on mRNA secondary structure. The “T” allele (−201.9 kcal/mol) had greater mRNA stability than the “C” allele (−206.7 kcal/mol) (Fig. 5). The TT genotype (0.14 ± 0.13 K/μL) presented higher BC when compared to the CT (0.07 ± 0.06 K/μL) and the CC (0.05 ± 0.04 K/μL) genotypes.
The CCL26 protein is a member of the eotaxin family. Mature eosinophils are activated and migrate to the site of infection in response to various chemoattractants, such as IL-5, CCL11, CCL24, and CCL26 (Rosenberg et al., 2013). Upregulation of CCL26 gene has been observed in draining lymph node tissue from Martinik × Romane back-cross resistant sheep exposed to H. contortus (Sallé et al., 2014). Based on these results, eosinophil migration could play an important role to control H. contortus in Florida Native sheep.
Two SNPs (OAR2:127200458 and OAR26: 14799731) in ITGA4 gene and exon 2 of TLR3 were associated with EC at 28 d. These polymorphisms explained 0.13 and 0.15 of the proportion of variance observed for EC at 28 d. The A to G substitution from OAR2:127199427 results in a minor change of −0.4 kcal/mol in mRNA stability but a moderate change in mRNA secondary structure (Fig. 6). Minor changes in mRNA stability were observed for the “A” allele (−127.9 kcal/mol) when compared to the “G” allele (−128.3 kcal/mol). For this SNP, the GG genotype (0.39 ± 0.24 K/μL) had greater EC when compared with GA (0.27 ± 0.23 K/μL) and AA (0.19 ± 0.18 K/μL) genotypes.
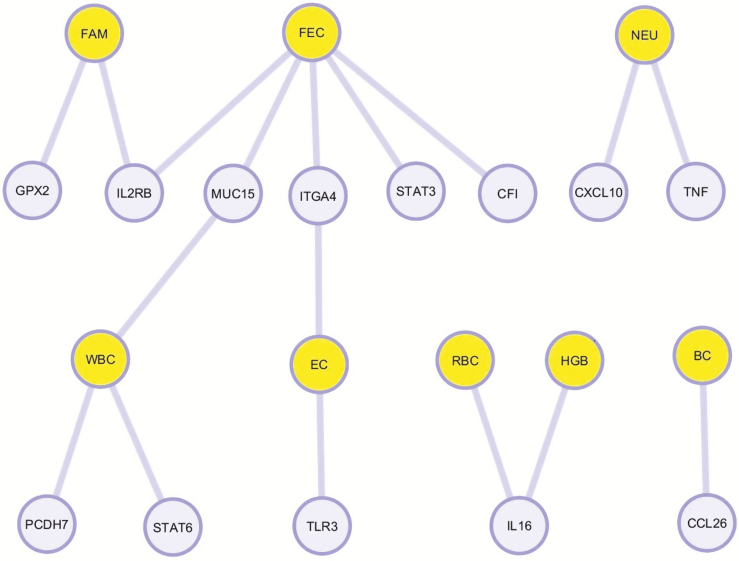
Figure 6.
Gene network for the genes with significant SNPs associated with H. contortus fecal egg count (FEC), FAMACHA score (FAM), red blood cell count (RBC), hemoglobin level (HGB), white blood cell count (WBC), neutrophil count (NEU), basophil count (BC), and eosinophil count (EC). The traits FEC, FAMACHA, WBC, and EC shared more genes in common with significant SNPs than RBC, HGB, NEU, and BC.
The T to C substitution from OAR26: 14799731 results in a minor change of 0.9 kcal/mol but has a strong impact on mRNA secondary structure (Fig. 6). The “T” allele (−91.9 kcal/mol) presented greater mRNA stability than the “C” allele (−92.8 kcal/mol), and the TT genotype (0.39 ± 0.37 K/μL) had greater EC than the CT (0.16 ± 0.15 K/μL) and CC (0.24 ± 0.23 K/μL) genotypes.
Our results could suggest that genetic variation within ITGA4 gene could be essential for controlling H. contortus infections in Florida Native sheep, due to its association with FEC and EC. Toll-like receptors (TLRs) are commonly expressed in antigen-presenting cells and other immune cells. TLR genes are found upregulated in the gut mucosa of resistant sheep infected with H. contortus and Trichostrongylus colubriformis (Ingham et al., 2008). TLR3 signaling could be one of the first immune response mechanisms activated by H. contortus in Florida Native sheep.
Overall, significant SNPs associated with FEC, FAMACHA score, WBC, NEU, BC, and EC in Florida Native sheep exposed to natural H. contortus infection are putative functional SNPs that affect the mRNA secondary structure and stability. Some of the genes (ITGA4, STAT3, MUC15, IL2RB, and CFI) containing the significant SNPs for FEC were previously highlighted as potential candidate markers for gastrointestinal parasite resistance in sheep populations (Periasamy et al., 2014, Benavides et al., 2015), and identified as differentially expressed in Chinese goats and Angus cattle during H. contortus infection and GIN (Araujo et al., 2009; Bhuiyan et al., 2017).
Results presented in this study confirm that stability of mRNA and conformational changes in the secondary structure of mRNA are important and possible causal mechanisms of gene expression variability. The 5′ UTR region is known to influence mRNA translation efficiency and secondary structures within this region could inhibit translational mechanisms (Leppek et al., 2018). Also, polymorphisms within the UTR regions confer more deleterious effects than polymorphisms in coding regions (Johnson et al., 2011).
Immunological Pathways Related to H. contortus Infection
Five pathways were related to significant SNPs associated with H. contortus FEC. These pathways include JAK-STAT signaling pathway, CD4+ T-cell commitment, Th17 signaling, endothelial cell proliferation, and leptin signaling and feeding behavior (Table 4). Leptin is commonly secreted by adipocytes, the placenta, and gastric epithelial cells. High concentrations of leptin in the stomach showed to induce inappetence in rats (Bado et al., 1998). Some studies suggested that increase in leptin concentration is responsible for immune mediated reduction in feed intake in sheep infected with T. colubriformis and T. circumcincta (Stear et al., 2003; Greer et al., 2009). Thus, inappetence could be used by Florida Native sheep as a protective response to minimize feed intake while the damaged mucosa by H. contortus is being repaired.
Table 4.
Immunological pathways related to H. contortus FEC, FAMACHA score, red blood cell count (RBC), hemoglobin level (HGB), white blood cell count (WBC), neutrophil count (NEU), basophil count (BC), and eosinophil count (EC) in Florida Native sheep; Gene = gene name; GO term = gene ontology pathway name; Number of genes in gene list = number of genes from list identified in the pathway; Number of genes in GO term = total number of genes included in the GO term
| Trait | Gene | GO term | GO term name | Number of genes in gene list | Number of genes in GO term |
|---|---|---|---|---|---|
| FEC | ITGA4, STAT3, MUC15, IL2RB, CFI | GO:0060397 | JAK-STAT cascade involved in growth hormone signaling pathway | 2 | 6 |
| GO:0043373 | CD4-positive, alpha-beta T-cell lineage commitment | 2 | 8 | ||
| GO:0002295 | T-helper cell lineage commitment | 2 | 8 | ||
| GO:0070102 | Interleukin-6-mediated signaling pathway | 2 | 9 | ||
| GO:0072540 | T-helper 17 cell lineage commitment | 2 | 6 | ||
| GO:0072539 | T-helper 17 cell differentiation | 2 | 6 | ||
| GO:0072538 | T-helper 17 type immune response | 2 | 6 | ||
| GO:1905562 | Regulation of vascular endothelial cell proliferation | 2 | 11 | ||
| GO:1905564 | Positive regulation of vascular endothelial cell proliferation | 2 | 7 | ||
| GO:0060259 | Regulation of feeding behavior | 2 | |||
| GO:0033210 | Leptin-mediated signaling pathway | 2 | 9 | ||
| GO:0044320 | Cellular response to leptin stimulus | 2 | 12 | ||
| FAMACHA score | IL2RB, GPX2 | GO:0098869 | Cellular oxidant detoxification | 1 | 86 |
| GO:0019221 | Cytokine-mediated signaling pathway | 1 | 91 | ||
| RBC, HGB | IL16 | GO:0005125 | Cytokine activity | 1 | 173 |
| WBC | STAT6, PCDH7, MUC15 | GO:0042093 | T-helper cell differentiation | 1 | 17 |
| GO:0002295 | T-helper cell lineage commitment | 1 | 8 | ||
| GO:0002294 | CD4-positive, alpha-beta T-cell differentiation involved in immune response | 1 | 17 | ||
| GO:0035771 | Interleukin-4-mediated signaling pathway | 1 | 2 | ||
| GO:0048295 | Positive regulation of isotype switching to IgE isotypes | 1 | 2 | ||
| NEU | CXCL10, TNF | GO:0002439 | Chronic inflammatory response to antigenic stimulus | 1 | 1 |
| GO:1901740 | Negative regulation of myoblast fusion | 1 | 1 | ||
| BC | CCL26 | GO:0031728 | CCR3 chemokine receptor binding | 1 | 4 |
| EC | ITGA4, TLR3 | GO:0034138 | Toll-like receptor 3 signaling pathway | 2 | 4 |
For FAMACHA, two pathways related to cellular oxidant detoxification and cytokine-mediated signaling were identified. For ADG, and RBC/HGB, pathways related to positive regulation of T-cell proliferation and cytokine activity were observed. The related pathways to WBC included T-helper differentiation and commitment, CD4+ T-cell differentiation, IL4-mediated signaling, and positive regulation of isotype switching to IgE. For NEU, pathways related to inflammatory response were identified. Finally, for BC and EC, pathways such as CCR3 chemokine receptor signaling and toll-like receptor 3 signaling were associated with these traits.
Immune response pathways associated with genes containing significant SNPs for WBC were related to CD4+ T-cell differentiation and Th2 response (IL4 signaling and isotype switching to favor IgE antibody production). From these genes, polymorphisms in MUC15 gene were also associated with FEC in our study. Although different SNPs were associated with both traits, the 5′UTR of MUC15 gene seems to play an essential role for H. contortus protection. Thus, initial establishment and rejection of H. contortus could be mediated by MUC15 gene and other molecules in Florida Native sheep.
From our gene network diagram, FEC, FAMACHA, WBC, and EC had more shared genes with significant SNPs when compared with NEU, HGB, RBC, and BC (Fig. 6). Also, FEC was the trait with more associated SNPs in our population. Some of these traits such as FEC and FAMACHA could be easily incorporated into future breeding programs to control GIN in Florida Native sheep.
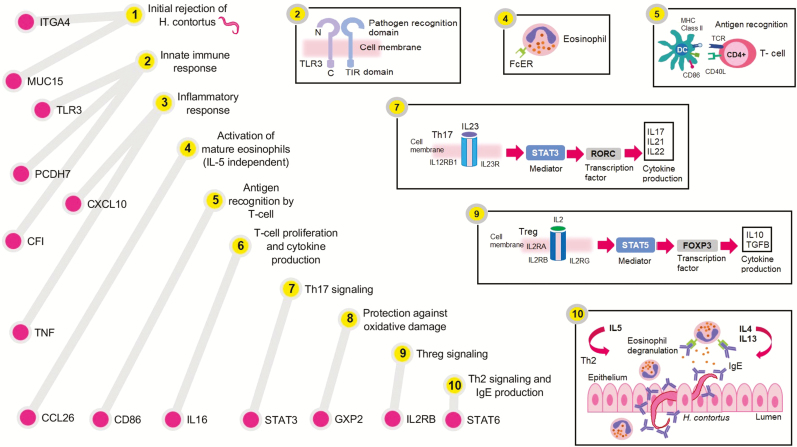
Overall, significant polymorphisms associated with these traits highlighted potential immune response mechanisms related to: initial nematode rejection, innate immune response (inflammatory response and complement activation), activation of eosinophils (IL5 independent), antigen recognition by T cells, T-cell proliferation and cytokine production, Th17 signaling, protection against oxidative damage, Treg signaling, and Th2 signaling and IgE production. These mechanisms could be representative of the potential immune mechanisms used by Florida Native sheep to control natural H. contortus infections (Fig. 7).
Figure 7.
Immune response mechanisms associated with natural H. contortus infection. . Significant polymorphisms in ITGA4 and MUC15 genes are related to initial rejection of the larvae. For innate immune response, significant SNPs were identified in TLR3 and PCDH7 genes. For the inflammatory response, significant polymorphisms in CXCL10 and TNF genes were observed. Activation of mature eosinophils, antigen recognition by T cells, T-cell proliferation, Th17 signaling, protection against oxidative damage, Treg signaling, and Th2 response were related to significant SNPs within CCL26, IL16, STAT3, GPX2, IL2RB, and STAT6 genes, respectively.
Implications
This study is the first report of potential candidate markers for parasite resistance in Florida Native sheep. However, future studies using monospecific infection with H. contortus, genome-wide scans, and additional animals would be required to validate our findings before these markers could be used with confidence in selection programs.
SNPs within 14 genes were significantly associated with FEC, FAMACHA score, WBC, NEU, BC, and EC. These genes were related to JAK-STAT signaling, CD4+ T-cell differentiation and commitment, Th17 signaling, leptin signaling and regulation of feeding behavior, cytokine-mediated signaling, Th2 response, TLR3 signaling, and CCR3 chemokine receptor binding. It is possible that immune response against natural H. contortus infections is mediated by these pathways in Florida Native sheep. Our results indicate that ITGA4, MUC15, TLR3, PCDH7, CFI, CXCL10, TNF, CCL26, IL16, STAT3, GPX2, IL2RB, and STAT6 genes could be potential markers for resistance to H. contortus exposure in Florida Native sheep. Future studies will need to validate these markers in other Florida Native sheep populations before they can be applied.
Supplementary Material
Footnotes
This research was supported Southern SARE (GS17-173). ZMER was partially supported by Consejo Nacional de Ciencia y Tecnologia. We gratefully acknowledge Dr. Francisco Peñagaricano, Assistant Professor at the University of Florida, for his guidance and support, and Eduardo Rodriguez and Kaitlyn Sarlo for their help during data analysis and sample processing.
Literature Cited
- Amarante A. F. T., Craig T. M., Ramsey W. S., Davis S. K., and Bazer F. W.. . 1999a. Nematode burdens and cellular responses in the abomasal mucosa and blood of Florida Native, Rambouillet and crossbreed lambs. Vet. Parasitol. 80:311–324. doi:10.1016/S0304-4017(98)00229-5 [DOI] [PubMed] [Google Scholar]
- Amarante A. F., Craig T. M., Ramsey W. S., El-Sayed N. M., Desouki A. Y., and Bazer F. W.. . 1999b. Comparison of naturally acquired parasite burdens among Florida native, Rambouillet and crossbreed ewes. Vet. Parasitol. 85:61–69. doi: 10.1016/s0304-4017(99)00103-x [DOI] [PubMed] [Google Scholar]
- Araujo R. N., Padilha T., Zarlenga D., Sonstegard T., Connor E. E., Van Tassel C., Lima W. S., Nascimento E., and Gasbarre L. C.. . 2009. Use of a candidate gene array to delineate gene expression patterns in cattle selected for resistance or susceptibility to intestinal nematodes. Vet. Parasitol. 162:106–115. doi: 10.1016/j.vetpar.2008.12.017 [DOI] [PubMed] [Google Scholar]
- Ashburner M., Ball C. A., Blake J. A., Botstein D., Butler H., Cherry J. M., Davis A. P., Dolinski K., Dwight S. S., Eppig J. T., Harris M. A., Hill D. P., Issel-Tarver L., Kasarskis A., Lewis S., Matese J. C., Richardson J. E., Ringwald M., Rubin G. M., and Sherlock G.. . 2000. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25:25–29. doi: 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aulchenko Y. S., Ripke S., Isaacs A., and van Duijn C. M.. . 2007. GenABEL: an R library for genome-wide association analysis. Bioinformatics 23:1294–1296. doi: 10.1093/bioinformatics/btm108 [DOI] [PubMed] [Google Scholar]
- Bado A., Levasseur S., Attoub S., Kermorgant S., Laigneau J. P., Bortoluzzi M. N., Moizo L., Lehy T., Guerre-Millo M., Le Marchand-Brustel Y., and Lewin M. J. M.. . 1998. The stomach is a source of leptin. Nature 394:790–793. doi: 10.1038/29547 [DOI] [PubMed] [Google Scholar]
- Baker R. L., and Gray G. D.. . 2004. Appropriate breeds and breeding schemes for sheep and goats in the tropics. In: Sani R. A., Gray G. D., and Baker R. L., editors, Worm control for small ruminants in tropical Asia. Australian Center for International Agricultural Research, Canberra, Australia: p. 63–95. [Google Scholar]
- Balic A., Cunningham C. P., and Meeusen E. N.. . 2006. Eosinophil interactions with Haemonchus contortus larvae in the ovine gastrointestinal tract. Parasite Immunol. 28:107–115. doi: 10.1111/j.1365-3024.2006.00816.x [DOI] [PubMed] [Google Scholar]
- Benavides M. V., Sonstegard T. S., Kemp S., Mugambi J. M., Gibson J. P., Baker R. L., Hanotte O., Marshall K., and Van Tassell C.. . 2015. Identification of novel loci associated with gastrointestinal parasite resistance in a Red Maasai x Dorper backcross population. PLoS ONE 10:e0122797. doi: 10.1371/journal.pone.0122797 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benavides M. V., Sonstegard T. S., and Van Tassell C.. . 2016. Genomic regions associated with sheep resistance to gastrointestinal nematodes. Trends Parasitol. 32:470–480. doi: 10.1016/j.pt.2016.03.007 [DOI] [PubMed] [Google Scholar]
- Berton M. P., de Oliveira Silva R. M., Peripolli E., Stafuzza N. B., Martin J. F., Álvarez M. S., Gavinã B. V., Toro M. A., Banchero G., Oliveira P. S., Pereira Eler J., Baldi F., and Sterman Ferraz J. B.. . 2017. Genomic regions and pathways associated with gastrointestinal parasites resistance in Santa Inês breed adapted to tropical climate. J. Anim. Sci. Biotechnol. 8:73. doi: 10.1186/s40104-017-0190-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhuiyan A. A., Li J., Wu Z., Ni P., Adetula A. A., Wang H., Zhang C., Tang X., Bhuyan A. A., Zhao S., and Du X.. . 2017. Exploring the genetic resistance to gastrointestinal nematodes infection in goat using RNA-sequencing. Int J Mol Sci. 18:751. doi: 10.3390/ijms18040751 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bishop S. C. 2012. Possibilities to breed for resistance to nematode parasite infections in small ruminants in tropical production systems. Animal 6:741–747. doi: 10.1017/S1751731111000681 [DOI] [PubMed] [Google Scholar]
- Danecek P., Auton A., Abecasis G., Albers C. A., Banks E., DePristo M. A., Handsaker R. E., Lunter G., Marth G. T., Sherry S. T., Mcvean G., and Durbin R.; 1000 Genomes Project Analysis Group 2011. The variant call format and VCFtools. Bioinformatics 27:2156–2158. doi: 10.1093/bioinformatics/btr330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diez-Tascón C., Keane O. M., Wilson T., Zadissa A., Hyndman D. L., Baird D. B., McEwan J. C., and Crawford A. M.. . 2005. Microarray analysis of selection lines from outbred populations to identify genes involved with nematode parasite resistance in sheep. Physiol. Genomics 21:59–69. doi: 10.1152/physiolgenomics.00257.2004 [DOI] [PubMed] [Google Scholar]
- Estrada‐Reyes Z. M., Tsukahara Y., Goetsch A. L., Gipson T. A., Sahlu T., Puchala R., Wang Z., Hart S. P., and Mateescu R. G.. . 2018. Effect of Ovar‐DRA and Ovar‐DRB1 genotype in small ruminants with haemonchosis. Parasite Immunol. 40:e12534. doi: 10.1111/pim.12534 [DOI] [PubMed] [Google Scholar]
- Gadahi J. A., Yongqian B., Ehsan M., Zhang Z. C., Wang S., Yan R. F., Song X. K., Xu L. X., and Li X. R.. . 2016. Haemonchus contortus excretory and secretory proteins (HcESPs) suppress functions of goat PBMCs in vitro. Oncotarget 7:35670–35679. doi: 10.18632/oncotarget.9589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goddard M. E., and Hayes B. J.. . 2009. Mapping genes for complex traits in domestic animals and their use in breeding programmes. Nat. Rev. Genet. 10:381–391. doi: 10.1038/nrg2575 [DOI] [PubMed] [Google Scholar]
- Goolsby M. K., Leite-Browning M. L., and Browning R. Jr. 2017. Evaluation of parasite resistance to commonly used commercial anthelmintics in meat goats on humid subtropical pasture. Small Rumin. Res. 146:37–40. doi:10.1016/j.smallrumres.2016.11.022 [Google Scholar]
- Greer, A. W., Y. R. Boisclair, M. Stankiewicz, and R. W. McAnulty. 2009. Leptin concentrations and the immune-mediated reduction of feed intake in sheep infected with the nematode Trichostrongylus columbriformis. Brit. J. Nutr. 102:954–957. [DOI] [PubMed] [Google Scholar]
- Grisart B., Coppieters W., Farnir F., Karim L., Ford C., Berzi P., Cambisano N., Mni M., Reid S., Simon P., Spelman R., Georges M., and Snell R.. . 2002. Positional candidate cloning of a QTL in dairy cattle: identification of a missense mutation in the bovine DGAT1 gene with major effect on milk yield and composition. Genome Res. 12:222–231. doi: 10.1101/gr.224202 [DOI] [PubMed] [Google Scholar]
- Guo Z., González J. F., Hernandez J. N., McNeilly T. M., Corripio-Miyar Y., Frew D., Morrison T., Yu P., and Li R. W.. . 2016. Possible mechanisms of host resistance to Haemonchus contortus infection in sheep breeds native to the Canary Islands. Sci. Rep. 6:1–14. doi: 10.1038/srep26200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hassan M., Hanrahan J. P., Good B., Mulcahy G., and Sweeney T.. . 2011. A differential interplay between the expression of Th1/Th2/Treg related cytokine genes in Teladorsagia circumcincta infected DRB1*1101 carrier lambs. Vet Res. 42(1):45. doi: 10.1186/1297-9716-42-45 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Howell S. B., Burke J. M., Miller J. E., Terrill T. H., Valencia E., Williams M. J., Williamson L. H., Zajac A. M., and Kaplan R. M.. . 2008. Prevalence of anthelmintic resistance on sheep and goat farms in the southeastern United States. J. Am. Vet. Med. Assoc. 233:1913–1919. doi: 10.2460/javma.233.12.1913 [DOI] [PubMed] [Google Scholar]
- Ingham A., Reverter A., Windon R., Hunt P., and Menzies M.. . 2008. Gastrointestinal nematode challenge induces some conserved gene expression changes in the gut mucosa of genetically resistant sheep. Int. J. Parasitol. 38:431–442. doi: 10.1016/j.ijpara.2007.07.012 [DOI] [PubMed] [Google Scholar]
- Johnson A., Trumbower H., and Sadee W.. . 2011. RNA structures affected by single nucleotide polymorphisms in transcribed regions of the human genome. Bioinformatics 2:WMC001600. [Google Scholar]
- Kaplan R. M. 2004. Drug resistance in nematodes of veterinary importance: a status report. Trends Parasitol. 20:477–481. doi: 10.1016/j.pt.2004.08.001 [DOI] [PubMed] [Google Scholar]
- Karim L., Coppieters W., Grobet L., Valentini A., and Georges M.. . 2000. Convenient genotyping of six myostatin mutations causing double-muscling in cattle using a multiplex oligonucleotide ligation assay. Anim. Genet. 31:396–399. doi: 10.1046/j.1365-2052.2000.00684.x [DOI] [PubMed] [Google Scholar]
- Keane O. M., Zadissa A., Wilson T., Hyndman D. L., Greer G. J., Baird D. B., McCulloch A. F., Crawford A. M., and McEwan J. C.. . 2006. Gene expression profiling of naïve sheep genetically resistant and susceptible to gastrointestinal nematodes. BMC Genomics 7:42. doi: 10.1186/1471-2164-7-42 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kemper K. E., Emery D. L., Bishop S. C., Oddy H., Hayes B. J., Dominik S., Henshall J. M., and Goddard M. E.. . 2011. The distribution of SNP marker effects for faecal worm egg count in sheep, and the feasibility of using these markers to predict genetic merit for resistance to worm infections. Genet. Res. (Camb). 93:203–219. doi: 10.1017/S0016672311000097 [DOI] [PubMed] [Google Scholar]
- Lee W. P., Stromberg M. P., Ward A., Stewart C., Garrison E. P., and Marth G. T.. . 2014. MOSAIK: a hash-based algorithm for accurate next-generation sequencing short-read mapping. PLoS ONE 9:e90581. doi: 10.1371/journal.pone.0090581 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lees M. S., Robinson N. A., Ingham A. B., Kotze A. C., and Piedrafita D. M.. . 2011. Dual oxidase 2 and glutathione peroxidase gene expression are elevated in hyperimmunized sheep challenged with Haemonchus contortus. Vet. Parasitol. 179:113–122. doi: 10.1016/j.vetpar.2011.02.013 [DOI] [PubMed] [Google Scholar]
- Leppek K., Das R., and Barna M.. . 2018. Functional 5’ UTR mRNA structures in eukaryotic translation regulation and how to find them. Nat. Rev. Mol. Cell Biol. 19:158–174. doi: 10.1038/nrm.2017.103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li R. W., and Gasbarre L. C.. . 2010. Gene expression in the bovine gastrointestinal tract during nematode infection. In: LaMann G., editor. Vet parasitology. Nova Biomedical Press, New York: p. 157–178. [Google Scholar]
- Lorenz R., Bernhart S. H., Höner Zu Siederdissen C., Tafer H., Flamm C., Stadler P. F., and Hofacker I. L.. . 2011. ViennaRNA package 2.0. Algorithms Mol. Biol. 6:26. doi: 10.1186/1748-7188-6-26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacKinnon K. M., Burton J. L., Zajac A. M., and Notter D. R.. . 2009. Microarray analysis reveals difference in gene expression profiles of hair and wool sheep infected with Haemonchus contortus. Vet. Immunol. Immunopathol. 130:210–220. doi: 10.1016/j.vetimm.2009.02.013 [DOI] [PubMed] [Google Scholar]
- McGuckin M. A., Lindén S. K., Sutton P., and Florin T. H.. . 2011. Mucin dynamics and enteric pathogens. Nat. Rev. Microbiol. 9:265–278. doi: 10.1038/nrmicro2538 [DOI] [PubMed] [Google Scholar]
- McRae K. M., McEwan J. C., Dodds K. G., and Gemmell N. J.. . 2014. Signatures of selection in sheep bred for resistance or susceptibility to gastrointestinal nematodes. BMC Genomics 15:637. doi: 10.1186/1471-2164-15-637 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McRae K. M., Stear M. J., Good B., and Keane O. M.. . 2015. The host immune response to gastrointestinal nematode infection in sheep. Parasite Immunol. 10:605–613. doi: 10.1111/pim.12290 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meeusen E. N., and Balic A.. . 2000. Do eosinophils have a role in the killing of helminth parasites? Parasitol. Today 16:95–101. doi: 10.1016/s0169-4758(99)01607-5 [DOI] [PubMed] [Google Scholar]
- Menzies M., Reverter A., Andronicos N., Hunt P., Windon R., and Ingham A.. . 2010. Nematode challenge induces differential expression of oxidant, antioxidant and mucous genes down the longitudinal axis of the sheep gut. Parasite Immunol. 32:36–46. doi: 10.1111/j.1365-3024.2009.01156.x [DOI] [PubMed] [Google Scholar]
- Miller J. E., Bahirathan M., Lemarie S. L., Hembry F. G., Kearney M. T., and Barras S. R.. . 1998. Epidemiology of gastrointestinal nematode parasitism in Suffolk and Gulf Coast Native sheep with special emphasis on relative susceptibility to Haemonchus contortus infection. Vet. Parasitol. 74:55–74. doi: 10.1016/s0304-4017(97)00094-0 [DOI] [PubMed] [Google Scholar]
- Miller D. K., andCraig T. M.. . 1996. Use of anthelmintic combinations against multiple resistant Haemonchus contortus in Angora goats. Small Rumin. Res. 19:281–283. [Google Scholar]
- Moncada D. M., Kammanadiminti S. J., and Chadee K.. . 2003. Mucin and Toll-like receptors in host defense against intestinal parasites. Trends Parasitol. 19:305–311. [DOI] [PubMed] [Google Scholar]
- O’Shea J. J., Gadina M., and Siegel R.. . 2013. Cytokines and cytokine receptors. In: Rich R. R., Fleisher T. A., Shearer W. T., Schroeder H. W., Frew A. J., and Weyand C. M., editors, Clinical immunology. 4th ed London: p. 108–135. doi: 10.1016/B978-0-7234-3691-1.00033-7 [DOI] [Google Scholar]
- Patel N., Kreider T., Urban J. F. Jr, and Gause W. C.. . 2009. Characterisation of effector mechanisms at the host:parasite interface during the immune response to tissue-dwelling intestinal nematode parasites. Int. J. Parasitol. 39:13–21. doi: 10.1016/j.ijpara.2008.08.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Periasamy K., Pichler R., Poli M., Cristel S., Cetrá B., Medus D., Basar M., Thiruvenkadan A. K., Ramasamy S., Ellahi M. B., Mohammed F., Teneva A., Shamsuddin M., Garcia Podesta M., and Diallo A.. . 2014. Candidate gene approach for parasite resistance in sheep—variation in immune pathway genes and association with fecal egg count. PLoS ONE. 9:e88337. doi: 10.1371/journal.pone.0088337 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pernthaner A., Cole S. A., Morrison L., and Hein W. R.. . 2005. Increased expression of interleukin-5 (IL-5), IL-13, and tumor necrosis factor alpha genes in intestinal lymph cells of sheep selected for enhanced resistance to nematodes during infection with Trichostrongylus colubriformis. Infect. Immunol. 73: 2175–2183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts F. H. S., and O’Sullivan P. J.. . 1950. Methods for egg counts and larval cultures for strongyles infesting the gastro-intestinal tract of cattle. Aust J Agric Res. 1(1):99–102. doi: 10.1071/AR9500099 [DOI] [Google Scholar]
- Rosenberg H. F., Dyer K. D., and Foster P. S.. . 2013. Eosinophils: changing perspectives in health and disease. Nat. Rev. Immunol. 13:9–22. doi: 10.1038/nri3341 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothenberg M. E., Mishra A., Brandt E. B., and Hogan S. P.. . 2001. Gastrointestinal eosinophils. Immunol. Rev. 179:139–155. doi: 10.1034/j.1600-065x.2001.790114.x [DOI] [PubMed] [Google Scholar]
- Sallé G., Moreno C., Boitard S., Ruesche J., Tircazes-Secula A., Bouvier F., Aletru M., Weisbecker J. L., Prévot F., Bergeaud J. P., Trumel C., Grisez C., Liénard E., and Jacquiet P.. . 2014. Functional investigation of a QTL affecting resistance to Haemonchus contortus in sheep. Vet. Res. 45:68. doi: 10.1186/1297-9716-45-68 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shannon P., Markiel A., Ozier O., Baliga N. S., Wang J. T., Ramage D., Amin N., Schwikowski B., and Ideker T.. . 2003. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13:2498–2504. doi: 10.1101/gr.1239303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silva M. V., Sonstegard T. S., Hanotte O., Mugambi J. M., Garcia J. F., Nagda S., Gibson J. P., Iraqi F. A., McClintock A. E., Kemp S. J., Boettcher P. J., Malek M., Van Tassell C. P., and Baker R. L.. . 2012. Identification of quantitative trait loci affecting resistance to gastrointestinal parasites in a double backcross population of Red Maasai and Dorper sheep. Anim. Genet. 43:63–71. doi: 10.1111/j.1365-2052.2011.02202.x [DOI] [PubMed] [Google Scholar]
- Smits H. H., and Yazdanbakhsh M.. . 2007. Chronic helminth infections modulate allergen-specific immune responses: protection against development of allergic disorders? Ann. Med. 39:428–439. doi: 10.1080/07853890701436765 [DOI] [PubMed] [Google Scholar]
- Stear M. J., Bishop S. C., Henderson N. G., and Scott I.. . 2003. A key mechanism of pathogenesis in sheep infected with the nematode Teladorsagia circumcincta. Anim. Health Res. Rev. 4:45–52. doi: 10.1079/AHRR200351 [DOI] [PubMed] [Google Scholar]
- Sweeney T., Hanrahan J. P., Ryan M. T., and Good B.. . 2016. Immunogenomics of gastrointestinal nematode infection in ruminants—breeding for resistance to produce food sustainably and safely. Parasite Immunol. 38:569–586. doi: 10.1111/pim.12347 [DOI] [PubMed] [Google Scholar]
- Terefe G., Grisez C., Prevot F., Bergeaud J. P., Dorchies P., Brunel J. C., François D., Fourquaux I., and Jacquiet P.. . 2007. In vitro pre-exposure of Haemonchus contortus L3 to blood eosinophils reduces their establishment potential in sheep. Vet. Res. 38:647–654. doi:10.1051/vetres:2007019 [DOI] [PubMed] [Google Scholar]
- Urban J. F. Jr, Noben-Trauth N., Donaldson D. D., Madden K. B., Morris S. C., Collins M., and Finkelman F. D.. . 1998. IL-13, IL-4Ralpha, and Stat6 are required for the expulsion of the gastrointestinal nematode parasite Nippostrongylus brasiliensis. Immunity 8:255–264. doi: 10.1016/s1074-7613(00)80477-x [DOI] [PubMed] [Google Scholar]
- Venturina V. M., Gossner A. G., and Hopkins J.. . 2013. The immunology and genetics of resistance of sheep to Teladorsagia circumcincta. Vet. Res. Commun. 37(2):171–181. doi: 10.1007/s11259-013-9559-9 [DOI] [PubMed] [Google Scholar]
- Zajac A. M., and Gipson T. A.. . 2000. Multiple anthelmintic resistance in a goat herd. Vet. Parasitol. 87:163–172. doi: 10.1016/s0304-4017(99)00174-0 [DOI] [PubMed] [Google Scholar]
- Zajac A. M., Herd R. P., and McClure K. E.. . 1988. Trichostrongylid parasite populations in pregnant or lactating and unmated Florida Native and Dorset/Rambouillet ewes. Int. J. Parasitol. 18:981–985. doi: 10.1016/0020-7519(88)90181-6 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.