Abstract
The latent cellular reservoir of HIV is recognized as the major barrier to cure from HIV infection. Long non-coding RNAs (lncRNAs) are more tissue and cell type-specific than protein coding genes, and may represent targets of choice for HIV latency reversal. Using two in vitro primary T-cell models, we identified lncRNAs dysregulated in latency. PVT1 and RP11-347C18.3 were up-regulated in common between the two models, and RP11-539L10.2 was down-regulated. The major component of the latent HIV reservoir, memory CD4+ T-cells, had higher expression of these lncRNAs, compared to naïve T-cells. Guilt-by-association analysis demonstrated that lncRNAs dysregulated in latency were associated with several cellular pathways implicated in HIV latency establishment and maintenance: proteasome, spliceosome, p53 signaling, and mammalian target of rapamycin (MTOR). PVT1, RP11-347C18.3, and RP11-539L10.2 were down-regulated by latency reversing agents, suberoylanilide hydroxamic acid and Romidepsin, suggesting that modulation of lncRNAs is a possible secondary mechanism of action of these compounds. These results will facilitate prioritization of lncRNAs for evaluation as targets for HIV latency reversal. Importantly, our study provides insights into regulatory function of lncRNA during latent HIV infection.
Introduction
In the present era of combination anti-retroviral therapy (cART), the latent cellular reservoir of HIV is recognized as the major barrier to a cure [1–3]. Existing latency reversing agents (LRAs) are suboptimal to induce a sustained reduction of the latent reservoir in vivo and suffer from lack of specificity for HIV [4]. Long non-coding RNAs (lncRNAs) may present targets of choice for HIV latency reversal because they are more tissue and cell-type specific than protein coding genes [5] and can be accurately targeted by oligonucleotides.
Though the role of individual lncRNAs in regulation of HIV expression and their possible contribution to HIV latency control has been recognized, the number of lncRNAs that were studied in this setting is limited. For example, siRNA-mediated knockdown or CRISPR-Cas9-induced knockout of Nuclear Paraspeckle Assembly Transcript 1 (NEAT1) resulted in enhanced HIV replication and an increase of unspliced and singly-spliced HIV RNA in the cytoplasmic fraction [6, 7], consistent with a possible role of NEAT1 in HIV RNA nuclear retention. Although these experiments were performed using productively infected cell lines [6], the results from this study suggested a possible role for NEAT1 in post-transcriptional regulation of HIV latency via nuclear retention of HIV transcripts, warranting further investigation in appropriate model systems. Another example is Non-Protein Coding RNA, Repressor of NFAT (NRON), whose function in HIV latency control was demonstrated using an in vitro model of HIV latency and cells from HIV-infected patients receiving cART [8]. NRON promoted HIV latency by recruiting the transactivator protein Tat to the proteasome for degradation; knockdown of NRON resulted in reactivation of the latent provirus [8]. In addition, NRON may also regulate HIV replication via cytoplasmic retention of nuclear factor of activated T-cells (NFAT) [9], which enhances HIV transcription in primary CD4+ T-cells [10]. In contrast, MALAT1 and uc002yug.2 lncRNAs were shown to be positive regulators of HIV replication [11, 12]. MALAT1 sequestered polycomb repressive complex 2 from the HIV LTR, promoting its transcriptionally active state [12]. Uc002yug.2 functioned via up-regulation of Tat and down-regulation of HIV transcriptional repressors Runx1b and Runx1c; overexpression of uc002yug.2 in cells from HIV-infected patients on cART improved HIV reactivation following treatment with phytohemagglutinin M [11]. Because lncRNAs represent a greater fraction of the transcribed human genome than protein coding genes [13], it is plausible to hypothesize that there are more lncRNAs than currently demonstrated that participate in regulation of HIV expression and may contribute to HIV latency.
To-date, very few studies have attempted to explore the complexity of host-HIV interactions in the context of lncRNA expression and function. We and others have previously profiled the entire transcriptome, including lncRNAs, at different time points following HIV infection in the SupT1 cell line [14, 15]. Peng and colleagues demonstrated that early response to HIV infection included changes in expression of many lncRNA, some of which were independent of active HIV replication [15]. We further expanded on this study to explore changes in lncRNA expression during different stages of the HIV replication cycle, including reverse transcription, integration and particle production [14]. Guilt-by-association (GBA) analysis demonstrated that dysregulated lncRNAs were functionally linked to many pathways involved in regulation of T-cell function and anti-viral responses [14, 15], consistent with the idea that lncRNAs represent a part of the host response to HIV infection and may be involved in regulation of induced cellular pathways.
To our knowledge, lncRNAs have not been previously profiled during latent HIV infection in primary CD4+ T cells. In the present study, we sought to identify lncRNA dysregulated in two relevant primary T-cell models of HIV latency using RNA-Seq. The models that use replication-competent wild type HIV virus were selected in order to recapitulate more closely the effects of latency in vivo. Because latency is preferentially established in memory compared to naïve cells, we further aimed to determine the relationship between expression levels of lncRNAs dysregulated in latency and CD4+ T-cell maturation state. To infer the function of dysregulated lncRNA in HIV latency establishment and maintenance, we performed GBA analysis that identifies cellular pathways associated with lncRNAs. Finally, we aimed to determine whether the selected dysregulated lncRNAs are modulated by small molecule LRAs suberoylamilide hydroxamic acid (SAHA) and Romidepsin (RMD), and whether effects of SAHA and RMD on lncRNA may represent novel secondary mechanisms of action of these compounds with respect to HIV reactivation. The main motivation behind the present study was prioritization of candidate lncRNAs for further formal evaluation as targets for therapies aimed at eradication of the latent HIV reservoir.
Results
Models of HIV latency
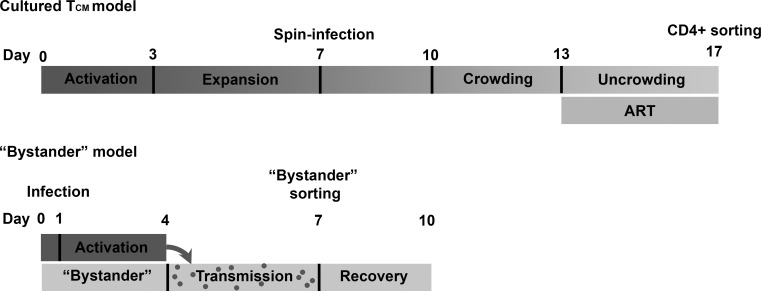
To identify lncRNAs dysregulated in HIV latency, the cultured TCM [16, 17] and the bystander model [18, 19] of HIV latency were used. These models differ by the phenotypic composition of CD4+ T-cells and the route of latency establishment (dividing cells returning to quiescence or resting cells). Cultured TCM model originally developed by Bosque and Planelles [16, 17] is established over 17 days of culture, where initial infection is conducted by spinoculation after 7 days following cell activation and expansion. Further virus replication and transmission is facilitated via cell-to-cell contact during the crowding stage till day 13, when the peak of p24+ expression is observed (21.8 ± 7.8 percent p24+ cells, N = 4). Virus spread is stopped at this point by adding antiretrovirals, and cells are allowed to return to quiescence for another 4 days [20]. Before samples are processed for sequencing, positive magnetic isolation of CD4+ T cells is used to sort out cells that may be productively infected [21]. Integration events in this model were characterized previously using Alu-PCR [21] detecting 33 ± 10.6% cells with HIV provirus on day 17, on average (N = 5), assuming 160 cells generate 1 ng DNA. “Bystander” model, described in [18] and [19], uses a shorter culture period, where a subset of cells is initially stained with a viable dye, e-Fluor 670, infected with HIV and activated, followed by mixing of these cells with resting “bystander” cells. Resting bystander cells are represented by all major phenotypic subsets found in vivo, including naïve, central memory and effector memory cells. Virus transmission from infected proliferating to bystander cells occurs in this co-culture via cell-to-cell contact. Peak of infection (average 37.3 ± 10.4 percent of p24+ proliferating cells, N = 4) is achieved on day 7. At this point, the resting (e-Fluor 670 negative) bystander cells are sorted out from the co-culture by fluorescence activated cell sorting. Following separation from productively infected cells, resting cells are left in culture for another 3 days to complete integration events of virus that entered the cells. Integration events in the end of culture were evaluated previously by droplet digital PCR (ddPCR) [19], detecting 14.9 ± 6% cells with HIV provirus on day 10, on average (N = 4). HIV RNA was detectable in both models at the time of RNA-Seq analysis, and increased following treatments that are used to reactivate HIV out of latency [19, 21]. In addition, a small percentage of cells generated by each of the models, without any reactivation stimuli, is p24+ (average 1.5 ± 1.2 percent and 1.8 ± 1.3 percent for bystander and cultured TCM models, respectively, N = 4). A diagram in Fig 1shows a side-by-side comparison of the key events during the establishment of each model.
Fig 1. Comparison of the time-line of latency establishment between cultured TCM and bystander models.
Cells during activation stage are depicted with darker grey color. Resting cells are presented in light grey. Gradient of color for the cultured TCM model represents gradual return from activation to quiescence. Major steps during model set-up (such as infection and sorting) are indicated above the time-line.
To identify genes dysregulated in latency, both TCM and bystander models were compared to “mock-infected” cells cultured in parallel. These cells underwent the same exposure to activation stimuli, spin-infection, crowding and ART treatment (in case of cultured TCM), or exposure to proliferating cells (in case on bystander model) as their infected counterparts. This design allowed us to control for exposure of cells to cytokines and chemokines in the co-culture. Data from the cultured TCM model was analyzed previously [21] identifying a total of 826 differentially expressed genes between mock-infected cells and the cells from the model of HIV latency. The bystander model was analyzed as part of the present study identifying 618 differentially expressed genes. To test the effect of the exposure to the virus on gene expression, gene expression signatures of latently infected cells were compared with a list of interferon stimulated genes with known antiviral properties [22]. This analysis identified 11 interferon stimulated genes (out of 826 total genes) for the cultured TCM model and was described previously [21]. In the present study, bystander model was assessed in a similar manner, and only one interferon stimulated gene (APOBEC3H) was differentially expressed out of 618 total genes. We also found two upregulated DNA sensors [23], NLR family pyrin domain containing 3 (NLRP3) and pyrin and HIN domain family member 1 (PYHIN1), in cultured TCM model only. Overall, these evaluations suggest that exposure to virus had minimal effect on gene expression at the time of the assay, and it may be more pronounced in the cultured TCM compared to the bystander model.
Identification of lncRNAs dysregulated in the models of HIV latency
Identified differentially expressed genes were annotated as protein coding or non-coding based on their types (e.g. intergenic, antisense, etc.), in both datasets (Tables 1 and S1). Differentially expressed lncRNAs represented 115 (13.9%) and 147 (23.8%) of all differentially expressed genes in the cultured TCM and the bystander models, respectively. The majority of dysregulated lncRNA was up-regulated in both models and originated from intergenic or antisense classes, consistent with genome distribution of different lncRNA types (Fig 2). Intronic lncRNAs were over-represented among lncRNAs dysregulated in latency (cultured TCM model: OR = 2.36, p = 0.02; bystander model: OR = 3.85, p<0.01). Differentially expressed genes and lncRNAs in each of the models of HIV latency were then compared, identifying 92 protein coding genes and 10 lncRNAs that were dysregulated in both models (S1 Fig). All dysregulated lncRNA were consistently either up- or down-regulated in both models (Table 2).
Table 1. Overview of differentially expressed protein coding genes and lncRNAs.
| Cultured TCM model | Bystander model | |
|---|---|---|
| Differential expression (gene level) | ||
| mRNA | ||
| Up | 456 | 310 |
| Down | 226 | 124 |
| lncRNA | ||
| Up | 75 | 136 |
| Down | 40 | 11 |
| Pseudogene | ||
| Up | 19 | 33 |
| Down | 9 | 4 |
| TR_V gene | ||
| Up | 1 | 0 |
| Down | 0 | 0 |
| Total | 826 | 618 |
| Differential expression (lncRNA types) | ||
| Antisense | ||
| Up | 32 | 48 |
| Down | 10 | 5 |
| LincRNA | ||
| Up | 31 | 49 |
| Down | 19 | 3 |
| Intronic | ||
| Up | 4 | 15 |
| Down | 4 | 1 |
| Sense-overlapping | ||
| Up | 1 | 2 |
| Down | 3 | 0 |
| Unclassified (TEC and proctranscript) | ||
| Up | 7 | 22 |
| Down | 4 | 2 |
| Total | 115 | 147 |
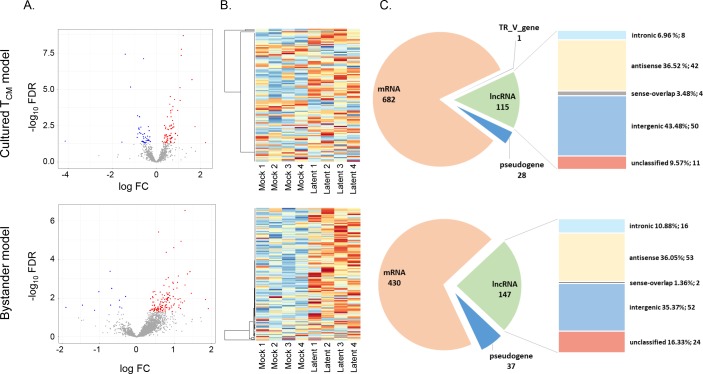
Fig 2. Overview of differentially expressed lncRNAs in the cultured TCM and the bystander models of HIV latency.
(A) Volcano plots showing all lncRNAs (grey dots). Significantly up-regulated lncRNAs (red dots) and down-regulated lncRNAs (blue dots) are highlighted for both latency models. (B) Heatmaps of differentially expressed lncRNAs. (C) Number of differentially expressed mRNAs, pseudogenes, TR_V genes, and lncRNAs, and percentages of lncRNAs of different types out of the total dysregulated lncRNAs.
Table 2. LncRNAs dysregulated in common between the cultured TCM and the bystander model.
| Cultured TCM model | Bystander model | |||||||
|---|---|---|---|---|---|---|---|---|
| ENSEMBL ID | Gene Symbol | lncRNA type | logFC | FC | FDR | logFC | FC | FDR |
| ENSG00000173727 | CMB9-22P13.1 | lincRNA | 0.86 | 1.81 | 0.0115 | 0.52 | 1.43 | 0.0368 |
| ENSG00000232160 | RAP2C-AS1 | antisense | 0.44 | 1.35 | 0.0465 | 0.72 | 1.65 | 0.0021 |
| ENSG00000245532 | NEAT1 | lincRNA | 0.50 | 1.41 | 0.0438 | 0.37 | 1.30 | 0.0413 |
| ENSG00000246526 | RP11-539L10.2 | lincRNA | -0.66 | 0.63 | 0.0377 | -0.67 | 0.63 | 0.0031 |
| ENSG00000249859 | PVT1 | lincRNA | 0.37 | 1.29 | 0.0030 | 0.58 | 1.49 | 0.0000 |
| ENSG00000253878 | RP11-347C18.3 | sense_intronic | 0.73 | 1.66 | 0.0014 | 1.09 | 2.12 | 0.0015 |
| ENSG00000266208 | CTD-2267D19.3 | antisense | 0.41 | 1.32 | 0.0470 | 1.15 | 2.22 | 0.0105 |
| ENSG00000267702 | RP11-53B2.2 | sense_intronic | 0.51 | 1.42 | 0.0253 | 0.56 | 1.48 | 0.0125 |
| ENSG00000271122 | RP11-379H18.1 | antisense | 0.38 | 1.30 | 0.0145 | 0.37 | 1.29 | 0.0107 |
logFC, log2-transformed fold change of counts per million in the RNA-Seq data; FC, fold change; FDR, false discovery rate-corrected p-value.
Validation of expression of lncRNA by ddPCR
Though an overlapping set of dysregulated lncRNAs was identified using RNA-Seq, the difference in expression of lncRNAs between the model of latency and mock-infected cells was frequently less than the effect size of the RNA-Seq experiment (effect size ≈1.5 with N = 4 biological replicates). Small differences in expression were not unexpected because each model of HIV latency is comprised of the mixture of uninfected and latently infected cells, with latently infected cells ranging from 8.8% to 44.3% [19, 21] (defined as percentage of cells with HIV provirus). Therefore, we aimed to use a method independent of RNA-Seq to validate expression of a set of overlapping lncRNAs. DdPCR was chosen for this purpose, because it is the most sensitive technique for detection of small differences in gene expression. Four lncRNAs, PVT1 oncogene (Ensembl ID ENSG00000249859), RP11-347C18.3 (Ensembl ID ENSG00000253878), RP11-539L10.2 (Ensembl ID ENSG00000246526) and NEAT1 (Ensembl ID ENSG00000245532), were selected for validation. Same samples that were sequenced, and three additional sample pairs for the bystander model were analyzed. Up-regulation of PVT1 and RP11-347C18.3 was confirmed both in the cultured TCM and the bystander models (cultured TCM model: PVT1 average fold change 1.8, p = 0.02; RP11-347C18.3 average fold change 2.95, p = 0.01; bystander model: PVT1 average fold change 1.33, p = 0.02; RP11-347C18.3 average fold change 2.12, p = 0.0002) (S2 Fig). Down-regulation of RP11-539L10.2 was confirmed in the bystander model (average fold change -1.28, p = 0.046) (S2 Fig). Up-regulation of NEAT1 was not confirmed in either of the models, neither with the assay that detects both short and long isoforms (N1+N2), nor the long isoform-specific assay (N2) (S2 Fig).
Neither of the RNA-Seq datasets contained the Ensembl identifier for lncRNA NRON (ENSG00000253079), possibly because it was not present in the reference files used for mapping and counting RNA-Seq reads. Because NRON represents the only lncRNA whose function in maintaining HIV in latent state has been demonstrated in a primary T-cell model system and cell from HIV-infected individuals [8], we measured the expression of NRON using ddPCR. Despite the previous report of a relatively high expression of this lncRNA in primary CD4+ T-cells [8], we observed relatively low expression, two orders of magnitude lower than of the other lncRNAs measured. For the cultured TCM model, there was sufficient amount of RNA to measure NRON from only three out of four sequenced sample pairs. In these three donors, NRON was strongly and significantly up-regulated (average fold change 4.2, p = 0.003) in the model of latency compared to mock-infected cells (S2 Fig). Though in some cases NRON was up-regulated in the bystander model, up-regulation was modest (average fold change 1.2 for four donors and 1.1 for 7 donors) and not consistent among the seven replicate experiments (S2 Fig). Thus, though up-regulation of NRON in latency may be used as validation of our approach of identifying lncRNAs that function in latency control, the discrepant results for NRON between the two models are consistent with the idea that route of latency establishment and cell composition may contribute to the mechanisms by which latency is regulated.
LncRNAs dysregulated in the models of HIV latency have higher expression in memory compared to naïve T-cells
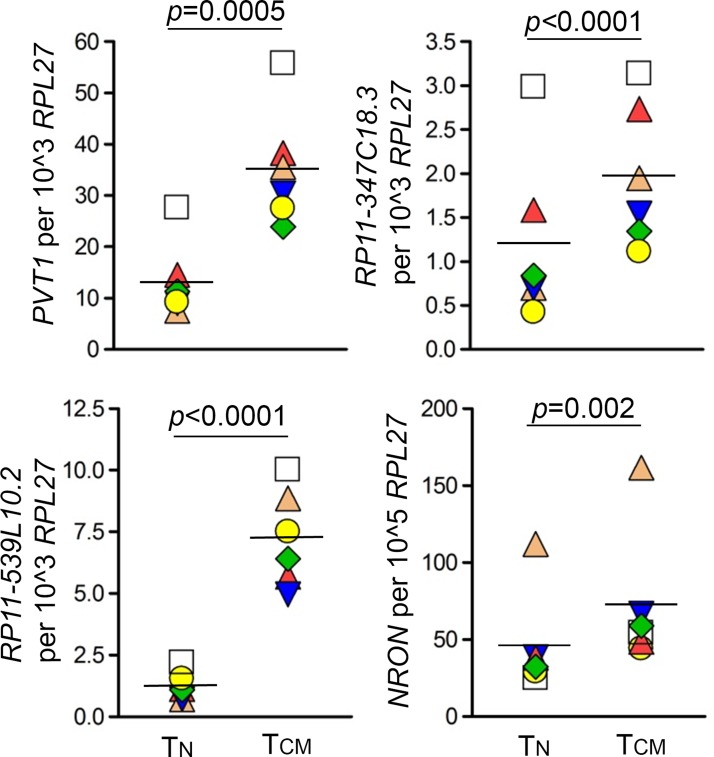
Memory cells are considered to be the major reservoir of latent HIV provirus [24–28]. If a host factor functions to promote HIV latency, its expression would likely be higher in memory than in TN cells. Indeed, an HIV repressor, Blimp-1 (gene symbol PRDM1) was expressed at significantly higher levels in TCM than in TN cells [29]. Therefore, to further assess whether identified lncRNAs likely function in latency control, we measured the expression of PVT1, RP11-347C18.3, RP11-539L10.2 and NRON in TN and memory cell subsets, including TCM and TEM cells. Uninfected cells sorted into major phenotypic subsets were used for this experiment. While PVT1 and RP11-347C18.3 were up-regulated and RP11-539L10.2 was down-regulated in latency, expression of all three lncRNAs was significantly higher in TCM and TEM compared to TN cells (Figs 3 and S3). Specifically, PVT1 expression was on average 3-fold higher in TCM than TN cells (N = 6, p = 0.0005); and on average 2.8-fold higher in TEM than TN cells (N = 4, p<0.0001). RP11-347C18.3 expression was on average 2-fold higher in TCM (N = 6, p<0.0001) or TEM (N = 4, p<0.0001) than TN cells. RP11-539L10.2 expression was on average 6.6-fold higher in TCM than TN cells (N = 6, p<0.0001); and on average 20-fold higher in TEM than TN cells (N = 4, p<0.0001). Consistent with its repressive function for HIV, NRON also had higher expression in TCM and TEM compared to TN cells (Figs 3 and S3) (average 1.7-fold higher in TCM than TN cells (N = 6, p = 0.002); average 4.1-fold higher in TEM than TN cells (N = 3, p<0.001)). Because PVT1 and RP11-347C18.3 were up-regulated in latency and in memory cells, their predicted function would be repression of HIV expression. In contrast, because RP11-539L10.2 was down-regulated in latency and had higher expression in memory cells, it would be predicted to promote HIV expression.
Fig 3. Comparison of lncRNA expression in TN and TCM T-cells.
Expression of PVT1, RP11-347C18.3, RP11-539L10.2 and NRON was measured by ddPCR and normalized to expression of the housekeeping gene RPL27. Six independent experiments were performed using uninfected CD4+ T-cells. Significance was determined using a paired two-sided t-test for log2 transformed data. Data is presented as individual data points (copy numbers normalized to RPL27) symbol-coded by donor, mean of all values is shown. Samples were generated using blood from donors who were different from those who participated in the study that used models of HIV latency. TN, naïve CD4+ T-cells; TCM, central memory CD4+ T-cells.
Identification of cellular pathways associated with lncRNAs dysregulated in the models of HIV latency
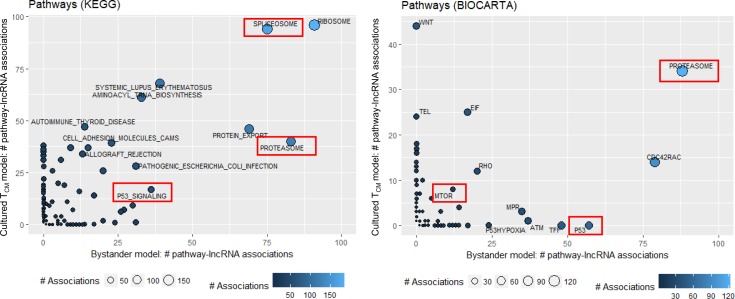
We further inferred function of dysregulated lncRNAs in HIV latency using GBA analysis, which determines correlations between lncRNA and mRNA expression in combination with enrichment strategies to identify cellular pathways associated with lncRNAs. Among the identified pathways, some were found in association with dysregulated lncRNA for both the cultured TCM and the bystander models (S4 and S5 Figs). Specifically, Spliceosome, Ribosome, Proteasome, and Protein export pathways from the Kyoto Encyclopedia of Genes and Genomes (KEGG), and Proteasome pathway from Biocarta database were associated with multiple dysregulated lncRNAs in both models (Fig 4). Of these, proteasome function has been previously implicated in maintaining HIV in latent state, and proteasome inhibitors were proposed as LRAs [30, 31]. Among other pathways with proposed roles in HIV latent reservoir establishment or maintenance, were p53 signaling [21] and mammalian target of rapamycin (MTOR) [32]. KEGG p53 signaling pathway had many lncRNA associations in both models, while lncRNA associations with Biocarta p53 pathway was only found for the bystander model (Fig 4). Fewer lncRNAs were associated to Biocarta MTOR, but for both models of HIV latency (Fig 4). Proper splicing is important for HIV protein production toward viral replication, but also toward antigen presentation to the immune system for killing of infected cells. Therefore, we and others propose that regulation of the spliceosome can critically regulate viral latency through multiple mechanisms [33, 34]. Through down-regulation of singly-spliced and unspliced messages, the virus can remain in a non-replicative state while being invisible to the immune system. Favoring certain doubly-spliced messages, specifically those encoded by Nef, the virus can also down-regulate molecules required for antigen presentation by the infected cell. Thus, it is plausible to hypothesize that spliceosome pathway may also have relevance to regulation of HIV latency. In addition, splicing factors that contain serine/arginine (SR) domains are well known regulators of HIV splicing [35]. Dysregulated lncRNA were found in association with several genes that encode SR protein mRNAs: SRSF2, SRSF3, SRSF6, SRSF7 and SRSF10.
Fig 4. Pathways associated with lncRNA dysregulated in latency, identified by the guilt-by-association (GBA) analysis.
Expression of lncRNAs dysregulated in latency (EdgeR’s FDR corrected p-value<0.05) was correlated with expression of all detected mRNAs. A ranked mRNA gene list was constructed for each lncRNA based on their correlation coefficients and used as input for gene set enrichment analysis (GSEA) with KEGG and Biocarta pathway databases. A lncRNA was considered significantly associated to a pathway when the GSEA FDR corrected p-value was less than 0.05. Number of lncRNA pathway associations is shown on the X-axis for the bystander model and Y-axis for cultured TCM model, color-coded with blue shades and indicated by the size of the circle (lighter blue color and larger circle size correspond to the greater number of associations). Ribosome and spliceosome pathways have the most associations with dysregulated lncRNAs in both models. Pathways that line up along the X axis were identified for the bystander model dataset only; while pathways that line up along the Y axis were identified for the cultured TCM model only. Pathways relevant to HIV latency are highlighted by red boxes.
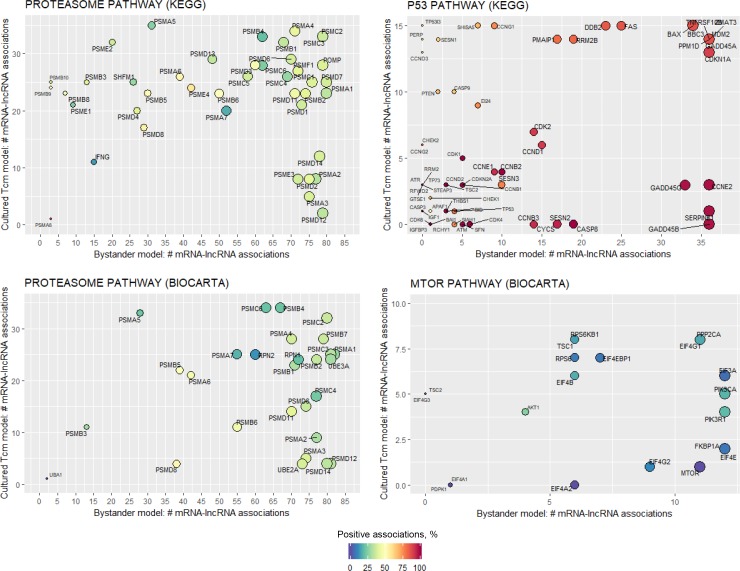
Next, we explored the associations between lncRNA expression and expression of individual members (mRNAs) of pathways implicated in HIV latency. Expression of the individual genes of the KEGG p53 signaling pathway positively correlated with expression of dysregulated lncRNAs, while Biocarta MTOR showed mostly negative associations (Fig 5). Both KEGG and Biocarta proteasome pathway exhibited mostly negative associations (Fig 5). Many mRNAs with highest number of associations to dysregulated lncRNAs were shared for both proteasome and MTOR pathways. In contrast, p53 pathway had a cluster of mRNAs commonly associated with dysregulated lncRNAs for both the cultured TCM and the bystander models (e.g. top right on the top panel in Fig 5), while other mRNAs were model-specific (e.g. a cluster of genes specific for the bystander model of HIV latency at the bottom right on the top panel in Fig 5). Overall, these data are consistent with a possibility that these pathways are regulated via multiple lncRNAs, and that control by lncRNA has both shared and unique components of the same pathway.
Fig 5. Associations between dysregulated lncRNAs and the mRNA components of pathways implicated in HIV latency.
For each lncRNA that was significantly associated with a pathway of interest (red boxes in Fig 4), leading edge mRNA genes were identified that drive the pathway enrichment score (GSEA). Next, the number of lncRNAs associated to pathway-specific mRNAs were plotted (ggplot2 v2.2.1). Number of associations between mRNA and lncRNA is shown on the X-axis for the bystander model and Y-axis for cultured TCM model, and indicated by the size of the circle (larger circle size corresponds to the greater number of associations). Color shows the percentage of positively correlated mRNA/lncRNA pairs (dark red is 100% positive correlations; dark blue–no positive correlations; yellow–equal number of positive and negative correlations).
PVT1 and RP11-347C18.3 are associated with pathways implicated in regulation of HIV latency
We further focused our attention on lncRNAs that were dysregulated in both models of latency and validated by ddPCR, namely PVT1, RP11-347C18.3 and RP11-539L10.2. We aimed to determine whether these lncRNAs were specifically associated with pathways implicated in regulation of HIV expression. Of these, PVT1 exhibited the most associations (Fig 6), including both KEGG and Biocarta p53 signaling pathway, Biocarta MTOR and proteasome pathways, and KEGG spliceosome pathway. Most of these associations were found for the bystander model and not the cultured TCM model; however, spliceosome, and a number of SR splicing factors in particular associated with PVT1 for both models (Fig 6). RP11-347C18.3 was associated with p53 signaling, both for KEGG and Biocarta pathways, and with the KEGG spliceosome pathway (S2 Table). RP11-539L10.2 was not associated with any annotated pathways implicated in HIV latency (S2 Table). The role of RP11-539L10.2 lncRNAs in regulation of HIV latency cannot be excluded based on these results, because it may function via control of a single critical host gene or its protein product, by directly targeting HIV, or via a pathway that is not annotated in the KEGG or Biocarta database. However, roles for PVT1 and RP11-347C18.3 are more likely based on identification of multiple HIV latency-related pathways associated with these lncRNAs.
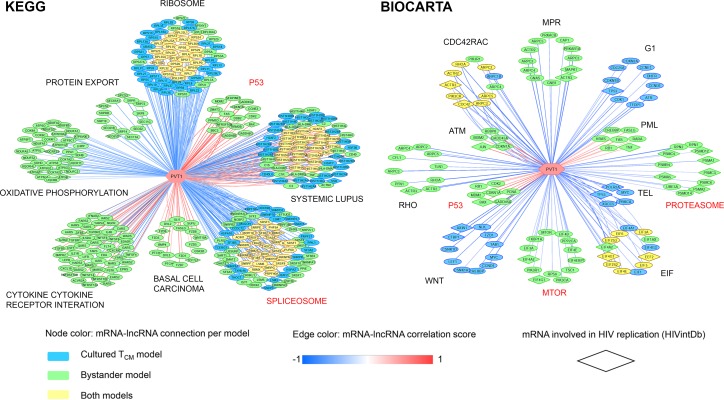
Fig 6. Associations between PVT1 and mRNA components of identified associated KEGG and Biocarta pathways.
Output of the guilt-by-association analysis was filtered for PVT1-specific pathway associations and corresponding leading edge mRNAs (GSEA FDR < 0.05). Next, a matrix with PVT1-specific information on pathway association, HIV latency model, enrichment score and leading edge mRNAs served as input for Cytoscape visualization (v3.4.0). Associations between PVT1 and mRNAs in all pathways (FDR corrected p<0.05) are shown. Nodes represent individual mRNAs associated with PVT1 and are color coded by model of HIV latency: blue, cultured TCM model, green, bystander model, and yellow, common association for both models. Nodes in a shape of a rhombus indicate mRNAs involved in HIV replication (annotated in the NCBI HIV-1 Interaction Database, HIVintDb). The color of the lines connecting PVT1 to mRNAs indicate correlation (red, positive; blue, negative). Names of pathways implicated in HIV latency are indicated in red.
Treatment with SAHA and RMD results in down-regulation of lncRNAs dysregulated in the models of HIV latency
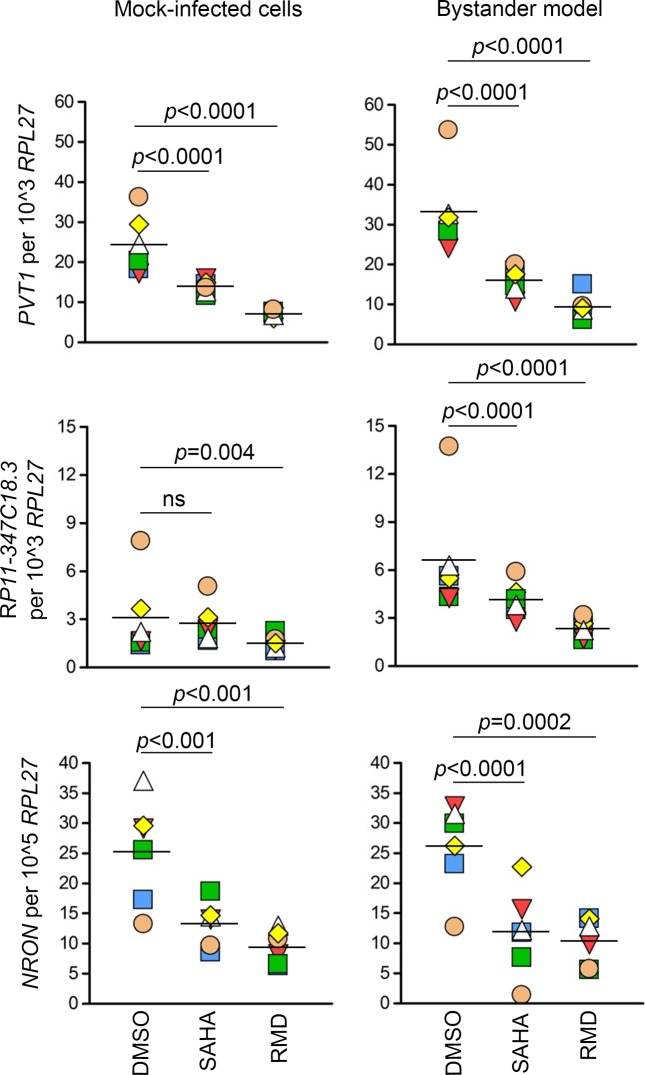
Finally, we have assessed how treatment with clinically tested LRAs, histone deacetylase inhibitors (HDACi) SAHA and RMD, affects expression of dysregulated lncRNAs. Samples from four to six replicates from the bystander model were used for this experiment. PVT1 and NRON were down-regulated by SAHA and RMD in the bystander model and mock-infected cells (PVT1: SAHA mock average fold-change -1.78, RMD mock average fold change -3.48, SAHA model of latency average fold change -2.1, RMD model of latency average fold change -2.8, p<0.0001; NRON: SAHA mock average fold-change -1.9, RMD mock average fold change -2.8, SAHA model of latency average fold change -3.5, RMD model of latency average fold change -2.8, p<0.001) (Fig 7). RP11-347C18.3 was down-regulated in all conditions except by SAHA in mock-infected cells (RMD mock average fold change -2.0, p = 0.004; SAHA model of latency average fold change -1.56, RMD model of latency average fold change -2.7, p<0.0001) (Fig 7). RP11-539L10.2 was down-regulated in all conditions except by RMD in the bystander model of HIV latency (SAHA mock average fold-change -3.81, RMD mock average fold change -3.43, p<0.0001; SAHA model of latency average fold change -2.25, p = 0.002) (S6 Fig). Overall, these results demonstrate that the effects of SAHA and RMD on expression of selected lncRNAs appear to be independent from HIV infection. As the primary mechanism of action of HDACi is chromatin decondensation leading to induction of gene transcription, down-regulation of the tested lncRNAs most likely represents a secondary mechanism of action of these compounds.
Fig 7. Down-regulation of PVT1, RP11-347C18.3 and NRON by SAHA and RMD.
Mock-infected cells and the bystander model of HIV latency were treated with SAHA (1 μM), RMD (15 nM) or their solvent DMSO for 24 hours. Expression of lncRNAs was measured by ddPCR and normalized to expression of the housekeeping gene RPL27. Six replicate experiments were performed. Significance was determined by implementing repeated measures analysis of variance (RM ANOVA) with library nlme in R using log2 transformed data. Data is presented as individual data points (copy numbers normalized to RPL27) symbol-coded by donor, mean of all values is shown. DMSO, dimethyl sulfoxide; SAHA, suberoylanilide hydroxamic acid; RMD, Romidepsin; ns, not significant (p-value > 0.05).
Discussion
The present study aimed to prioritize lncRNAs to be tested as targets for HIV latency reversal. We used several complementary approaches, including the identification of dysregulated lncRNAs in primary T-cell models of HIV latency (Figs 2 and S1 and S2and Tables 1, 2 and S1), GBA analysis (Figs 4–6 and S4 and S5 and S2 Table), evaluation of lncRNA expression in major CD4+ T-cell maturation subsets (Figs 3 and S3) and in CD4+ T-cells treated with clinically tested LRAs (Figs 7 and S6.
Among lncRNAs dysregulated in the models of HIV latency, 10 were in common between the two models (Table 2). In addition, 105 unique lncRNAs were identified for the cultured TCM model, and 137 for the bystander model (S1 Fig). Same protocols were followed to generate and analyze RNA-Seq data for both models (please refer to Materials and Methods); thus, discrepancies in the result caused by differences in experimental and analytical procedures are expected to be minimal. Therefore, it is likely that the observed differences in the identified dysregulated lncRNAs were due to biological differences of the two models. Specifically, T-cell composition and route of latency establishment may contribute to differential dysregulation of lncRNAs in latency. One observation in favor of this conclusion is that NRON was strongly up-regulated in the cultured TCM model that establishes latency in activated cells returning to quiescence, but not in the bystander model that uses direct infection of resting cells (S2 Fig). A prior study that demonstrated the role of NRON in latency maintenance [8] used a model with a similar route for latency establishment as our cultured TCM model.
We further focused our evaluation of possible functions of the four lncRNAs that were dysregulated in both models of latency and had higher expression levels among the ten lncRNA identified, namely PVT1, RP347C18.3, RP539L10.2 and NEAT1. Of these, NEAT1 differential expression was not validated by ddPCR in either model (S2 Fig). The prior studies demonstrating the role of NEAT1 in HIV RNA nuclear retention during productive HIV replication was performed using cell lines [6, 7]. Therefore, it is possible that NEAT1 may not function in the same manner in latently infected primary CD4+ T-cells. Because the effect size with the number of samples (N = 4) available in our RNA-Seq experiment was ≈1.5, and the observed difference for NEAT1 was smaller (1.41-fold in the cultured TCM model and 1.3-fold in the bystander model, Table 2), NEAT1 might represent one of the false positives detected in our RNA-Seq study. The remaining three selected lncRNAs were confirmed by ddPCR and evaluated in CD4+ T-cells of naïve and memory phenotypes. All of them exhibited higher expression in TCM and TEM cells (Figs 3 and S3), which represent the major component of the latent reservoir. PRDM1, a known HIV repressor whose function was evaluated in primary CD4+ T-cells, exhibited a similar pattern of expression in TN and TCM subsets [29]. LncRNA NRON, whose repressive function for HIV expression was previously demonstrated in primary CD4+ T-cells [8], also had higher expression in TCM and TEM compared to TN cells (Figs 3 and S3). Therefore, our observation of elevated expression of selected lncRNAs in memory cells is consistent with their possible function in regulation of HIV expression in latency.
GBA analysis was used to hone into potential functions of dysregulated lncRNAs in HIV latency control by determining whether they are associated to known pathways implicated in establishment or maintenance of HIV latency. GBA analysis identified proteasome, spliceosome, p53 signaling and MTOR among pathways associated with dysregulated lncRNAs (Figs 4 and S4 and S5). Most of the lncRNAs had negative associations with mRNA members of these pathways (Fig 5), consistent with a possibility of inhibitory regulation of gene expression of these pathways by lncRNA. Some of the mechanisms by which this regulation may occur include transcriptional interference, promoter inactivation by binding transcription factors, cytoplasmic retention of transcription factors, and epigenetic silencing [13, 36]. In contrast, the majority of associations in the p53 signaling pathway were positive (Fig 5). Such positive regulation may as well occur through chromatin remodeling and interactions with and recruitment of transcriptional activators [13]. In addition, lncRNAs may function by increasing stability of the mRNAs [37]. Negative [38] as well as positive [39] feedback loops have been reported between lncRNAs and proteins; however, independent studies of interactions between the identified lncRNAs and the members of associated pathways at the protein level would be required to determine whether this may be the case for regulation of HIV latency.
Of the three dysregulated lncRNAs that were validated by ddPCR, PVT1 was found in association with multiple pathways implicated in control of HIV latency (Fig 6). The common pathway identified for the cultured TCM and the bystander models associated with PVT1 was spliceosome (Fig 6). Remarkably, PVT1 was associated with a number of mRNAs encoding SR proteins, in common between the two models (Fig 6). Of these, SRSF2, SRSF6, and SRSF10 have been specifically linked to HIV splicing [40–43]. PVT1 was associated with members of p53 signaling, MTOR and proteasome pathway in the case of the bystander model only (Fig 6), consistent with the idea that pathways that regulate latency may be, at least in part, dependent on the model. PVT1 was found in association with p53 signaling pathway for both KEGG and Biocarta databases (Fig 6). The role of p53 signaling in HIV establishment and maintenance was demonstrated previously [21]. One protein member of this pathway, MDM2 Proto Oncogene (MDM2), was stabilized by PVT1 via its interaction with Enhancer Of Zeste 2 Polycomb Repressive Complex 2 Subunit (EZH2) in the setting of hepatocellular carcinoma [44]. Though the precise mechanism of regulation of p53 signaling pathway by PVT1 in different cell types needs further investigation, the positive regulation of it components was consistent in our study and the study by Guo and colleagues [44]. In addition, EZH2 itself contributes to regulation of HIV latency [45]. PVT1 interacted with and stabilized the EZH2 protein product [44], which could represent a potential mechanism of latency control by PVT1. In addition, PVT1 was shown to act as a sponge for multiple miRNAs [46–51], suggesting the possibility for positive regulation of multiple mRNAs in the p53 signaling pathway that represent miRNA targets. Based on these results, we speculate that PVT1 may function in HIV latency control via modulating mRNA components of p53 signaling pathway and splicing machinery.
Intriguingly, identified associations of PVT1 to cellular pathways demonstrate possible links between inferred mechanisms of regulation of gene expression by PVT1 in HIV latency and other diseases, such as basal cell carcinoma and systemic lupus erythematosus (Fig 6). We speculate that a subset of dysregulated genes that may be regulated by PVT1 is shared between these conditions. For example, TCF7L2 (also known as TCF-4) from the basal cell carcinoma pathway plays a role in HIV latency maintenance [52] and was up-regulated in the bystander model in our study (S1 Table). For the systemic lupus erythematosus pathway, the majority of genes linked to PVT1 are histone-coding genes. Epigenetic alterations, histone modifications in particular, largely contribute to gene expression dysregulation in systemic lupus erythematosus [53]. A number of histone-coding genes were dysregulated at the RNA level in systemic lupus erythematosus [54] and the cultured TCM model in our study (S1 Table).
Lastly, our study investigated the effects of clinically tested LRAs, SAHA and RMD, on expression of lncRNAs dysregulated in latency, PVT1, RP11-347C18.3, RP11-539L10.2, and NRON. SAHA and RMD are HDACi, whose primary mechanism of action is histone hyperacetylation which results in chromatin decondensation and elevated gene expression. However, multiple studies demonstrated that HDACi cause gene down-regulation [55–57], which likely represents secondary mechanisms of action of these compounds. Our prior investigation of these secondary effects on protein coding genes suggested presence of both stimulatory and inhibitory effects of HDACi with respect to HIV reactivation [19, 58]. All tested lncRNAs, PVT1, RP11-347C18.3, RP11-539L10.2, and NRON were down-regulated by SAHA and RMD, independent of the presence of latent infection (Figs 7 and S6). When HIV is present, down-regulation of PVT1, RP11-347C18.3, and NRON, which were up-regulated in latency, may represent stimulatory effects. Further down-regulation of RP11-539L10.2 by SAHA, beyond levels of down-regulation observed in latency compared to mock-infected cells, may represent an inhibitory effect. Thus, both stimulatory and inhibitory effects of HDACi are likely present among non-coding genes. The results from this study provide basis for hypothesis building to identify candidate lncRNAs whose experimental down-regulation (such as via knockdowns) may be synergistic with HDACi treatment for latency reversal.
One limitation of the present study is the low frequency of latently infected cells in a large background of uninfected cells for both models of HIV latency (range 8.8% to 44.3% [19, 21]). This is a common problem for models that use replication-competent wild type HIV virus. Therefore, we conducted a comparison of dysregulated lncRNAs in two different well characterized models [16–19, 21, 59] to attempt to overcome the limitation introduced by high background of uninfected cells. Only lncRNAs found dysregulated in both models were analyzed in more detail. Characterization of single cell transcriptomes using the cultured TCM and bystander models might be beneficial for identification of a greater number of overlapping lncRNAs and for validating the role of T-cell maturation state and prior activation in lncRNA expression signatures of latently infected cells.
The second limitation is a possibility that differential gene expression may result from the exposure of cells to cytokines and chemokines produced by cells during productive infection, or due to exposure to the virus in culture. To control for cytokine and chemokine exposure during T-cell activation, we ensured that the mock-infected cells were treated the same way as infected cells throughout the culture. We have also searched for interferon stimulated genes with known antiviral properties [22] and DNA sensors [23] among genes that were differentially expressed in the latency models compared to mock-infected cells. Some such responses were detected, predominantly in the cultured TCM model. It is also possible that transcriptional profiles could be influenced by virus via interferon-independent cascades [60, 61]. While a good control for these effects might be the use of anti-virals during the stages of latency establishment, or the use of cells exposed to the aldrithiol-2 inactivated virus [62] instead of mock-infected cells, we were limited by analyzing previously published RNA-Seq datasets that did not include these controls. Nonetheless, identification of a subset of genes with known roles in HIV latency supports the potential of this experimental and analytical approach to identify genes specifically affected during HIV latency and not purely by cytokines or exposure to virus. Indeed, expression of p21 (gene symbol CDKN1A) was elevated in the cell line model of latency [63] and in the bystander model in the present study (S1 Table). Its expression negatively correlated with cell-associated HIV RNA and HIV transcriptional activity in HIV-infected virologically suppressed individuals on cART [64], and its protein product inhibited transcription from the integrated provirus via inhibition of Cyclin Dependent Kinase 9 (CDK9), a component of positive transcription elongation factor (P-TEFb) [65]. Another example is PRDM1, up-regulated in both models in the present study (S1 Table), which was previously shown to be a repressor of basal and Tat-dependent HIV transcription [29]. Moreover, expression of the only lncRNA with demonstrated role in HIV repression in primary CD4+ T-cells, NRON [8], was also up-regulated in the cultured TCM model in the present study (S2 Fig). These observations are consistent with the idea that at least a subset of genes and lncRNAs found dysregulated in latency in the present study has demonstrated roles as HIV regulators. Ultimately, identified lncRNAs will need to be validated as HIV regulators using functional studies (e.g. experimental gene expression knockdowns) using relevant model systems and cells from HIV-infected individuals ex vivo.
In summary, the present study used gene expression profiling by RNA-Seq in two primary T-cell models of HIV latency to focus on identification of dysregulated lncRNAs. Despite differences in cell composition and route of latency establishment between the models, common lncRNA signatures associated with latency could be identified. LncRNAs dysregulated in latency had higher expression in memory cells that represent the major HIV reservoir, as compared to TN cells. We further identified pathways associated with dysregulated lncRNA, including pathways previously implicated in HIV latency: proteasome, spliceosome, p53 signaling and MTOR. Of the lncRNAs that were dysregulated in common between the two models, PVT1 had the most associations to these pathways. We have proposed several mechanisms of action for PVT1 to regulate HIV latency, based on evidence for PVT1 function from published literature. Identification of model-specific dysregulated lncRNAs and lncRNA-associated pathways will facilitate better understanding of the mechanisms by which latency is established and maintained in different cell types. The role of the identified lncRNAs in HIV latency and their mechanisms of action warrant further experimental exploration to determine their suitability as targets for antiviral strategies. Our study facilitates prioritization of lncRNAs to be tested in this setting.
Materials and methods
Study participants
Primary CD4+ T-cells from HIV seronegative volunteer blood donors were used for this study. The protocol was approved by the Institutional Review Boards of the University of California San Diego, and the VA San Diego Healthcare System and abides by the Declaration of Helsinki principles. All volunteers gave written informed consent to participate in the study.
In vitro models of HIV latency
To identify lncRNAs dysregulated in HIV latency, two primary T-cell models were used [16–18]. In the cultured TCM model, TN cells are activated and polarized to direct differentiation into TCM cells, which are phenotypically very similar to TCM cells freshly isolated from blood [17]. Latency is established in the presence of IL-2 and cART in cultured TCM cells that are gradually returning to quiescence and becoming resting; positive magnetic isolation of CD4+ T cells is used to sort out cells that remain productively infected [16, 59]. In the bystander model developed by Dr. Celsa Spina [18, 19], latency is established directly in resting CD4+ T-cells, represented by all maturation phenotypes found in vivo. Both the cultured TCM and the bystander models utilized wild type HIVNL4.3 virus. Expression of p24/Gag protein during model-set up was assessed as described previously, using ICp24 antibody conjugated to fluorescein isothiocyanate (FITC) (clone KC57, Coulter) for cultured TCM model [16]; for bystander model, ICp24 antibody conjugated to phycoerythrin (PE) (clone KC57, Coulter) was used. To establish a mock-infected control, a portion of cells was cultured under the same conditions in parallel, but without exposure to the virus.
Identification of lncRNAs dysregulated in HIV latency
For the cultured TCM model, we used a previously published dataset [21] (raw and processed data available at Gene Expression Omnibus (GEO) database, accession number https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE81810) to re-analyze focusing on dysregulated lncRNAs. For the bystander model, we used RNA-Seq data available at GEO under accession number GSE114883. This dataset includes samples for mock-infected cells and latency model treated with HDACi SAHA and RMD and their solvent dimethyl sulfoxide (DMSO). DMSO-treated samples from this dataset were used to perform differential gene expression analysis between mock-infected cells and cells from the model of HIV latency. To minimize variation due to analytical procedures, bystander model data was analyzed in the same manner as previously published TCM model dataset [21]. Briefly, Tophat [66] was used for mapping to human genome version hg38 and HTSeq [67] for read counting against the Gencode gene annotation version 21. Mapping to HIV and synthetic RNA standards from the External RNA Controls Consortium (ERCC) was performed using Bowtie [68]. Differential expression analysis was performed using library EdgeR [69] in Bioconductor R and normalization to ERCC spike-ins with RUVSeq [70]. Genes and lncRNAs with false discovery rate (FDR) corrected p-values less than 0.05 were considered differentially expressed.
Among total differentially expressed genes in both models, lncRNAs and their types (intergenic, intronic, etc.) were determined using Gencode gene annotation version 21 with Ensembl IDs as unique identifiers. Visualization of differential expression included volcano plots and heatmaps that were constructed via in-house bioinformatics scripts and based on the package gplots (v3.0.1) in the R computing environment [71]. Enrichment of lncRNA types in the lists of dysregulated lncRNAs was evaluated via comparison of ratios with the lncRNA type distribution of the total pool of lncRNAs in the Gencode database (Fisher’s exact test).
Isolation of TN, TCM, and TEM CD4+ T-cell subsets
All antibodies were obtained from BD Biosciences, Inc. (San Jose, CA, USA). Antibodies for cell quality control were αCD4 (multiclone SK3 SK4) conjugated to FITC, and αHLA-DR (clone L243) conjugated to PE. Antibodies for isolation of CD4+ T-cell maturation subsets were αCD45RA (clone L48) conjugated to phycoerythrin-cyanine dye (PE-Cy7), and αCD62L (clone DREG-56) conjugated to allophycocyanin (APC). CD4+ T-cells were isolated from whole blood using negative selection (StemCell Technologies, Inc., Vancouver, Canada). All CD4+ T-cell samples had >95% purity and <10% expression of activation marker HLA-DR, as assessed using the Accuri C6 flow cytometer (BD Biosciences, Inc., San Jose, CA, USA). To isolate T-cell maturation subsets, CD4+ T-cells were stained with αCD45RA and αCD62L antibodies, and sorted on a MoFlo XDP Cell Sorter (Beckman Coulter, Brea, CA, USA) to obtain TN (CD45RA+CD62L+), TCM (CD45RA-CD62L+) and TEM (CD45RA-CD62L-) cell populations.
Treatment with SAHA and RMD
Aliquots of SAHA and RMD solubilized in DMSO were provided by Merck Research Laboratories, Inc. (Kenilworth, NJ, USA), and Gilead Sciences, Inc. (Foster City, CA, USA), respectively. Mock-infected cells and the bystander model of HIV latency were treated with 1 μM SAHA, 15 nM RMD or their solvent DMSO for 24 hours.
Isolation of RNA
RNA was isolated from CD4+ T-cells using RNeasy micro kit (Qiagen, Inc., Valencia, CA). RNA concentrations were assessed using Qubit 2.0 fluorometer (Thermo Fisher Scientific, Inc., Waltham, MA, USA).
Droplet digital PCR (ddPCR)
Four lncRNA dysregulated in both models of latency were selected for confirmation by ddPCR. The selection was made based on expression level likely detectable in a single ddPCR reaction (number of reads per base in the RNA-Seq experiment >0.1). Reads per base were calculated by dividing the number of reads mapped to each lncRNA by the average transcript length as determined by GenomicFeatures library [72] in Bioconductor R. The assays to measure lncRNA expression were custom designed and purchased from Integrated DNA Technologies, Inc. (Coralville, IA, USA). The assays were as follows: NEAT1 both short and long isoforms (N1+N2): forward 5’ TTCATGGACCGTGGTTTG 3’, reverse 5’ 56-FAM/CTGCAATGCTAGGACTCAC 3’, probe 5’ TTCCTCATG/ZEN/GCGAGCAGATGGAAC/3IABkFQ 3’; NEAT1 long isoform (N2): 5’ forward ACGTGTTGCATGGTTTCT 3’, reverse: 5’ ATGAGGGCAGTTCTCTGT 3’, probe 5’ 56-FAM/AACAGTAGG/ZEN/GAGATGCCTGGGAGTA/3IABkFQ 3’; PVT1: forward 5’ GAGGGTTGAGATCTCTGTTTAC 3’, reverse 5’ GATGCTTCACCAGGAAGAG 3’, probe 5’ 56-FAM/TCTGCCAAC/ZEN/TTCCTTTGGGTCTCC/3IABkFQ 3’; RP11-347C18.3: forward 5’ AGCTCTCATGTGACCCA 3’, reverse 5’ AATAACCTGGTGAGTTGGC 3’, probe 5’ 56-FAM/ACCTGCAAA/ZEN/TTGTGGGCATTCACG/3IABkFQ 3’; RP11-539L10.2: forward 5’ TTTGGTCCCTGGCTTTG 3’, reverse 5’ CCCTATCTCTGATCATTGTCAC 3’, probe 5’ 56-FAM/TTGTGACCC/ZEN/GAGTGTCAGTTTCCT/3IABkFQ/ 3’. NRON lncRNA, which was not annotated in the RNA-Seq datasets, was measured by ddPCR using assay Hs04274937_s1 from Applied Biosystems (now Thermo Fisher Scientific, Inc., Waltham, MA, USA). Housekeeping gene Ribosomal Protein L27 (RPL27) was used as a normalizer [55, 58, 73]. The assay (HS03044961_g1) to measure RPL27 was purchased from Applied Biosystems (now Thermo Fisher Scientific, Inc., Waltham, MA, USA). RNA was converted to cDNA using qScript (Quanta Bio, Beverly, MA). Concentration of RNA in all cDNA reactions was 1 ng/μl, except for lowly expressed NRON, for which 10 ng/μl was used. DdPCR reactions were set up and run as described previously [58]. LncRNAs whose dysregulation in latency were confirmed (PVT1, RP11-347C18.3, RP11-539L10.2 and NRON) were further measured in CD4+ T-cell maturation subsets and in CD4+ T-cells following treatment with SAHA and RMD.
GBA analysis
The GBA analysis to associate lncRNAs with biological pathways was performed as described previously [14, 74]. Briefly, the GBA pipeline first builds a Spearman’s rank correlation matrix between the differentially expressed lncRNAs and all expressed mRNAs based on the normalized gene expression values of the samples (counts per million). Next, a ranked mRNA gene list was constructed for each lncRNA based on their correlation coefficients and used as input for gene set enrichment analysis (GSEA) [75] together with pathways from the Biocarta [76] and KEGG [77] databases. A lncRNA was considered significantly associated to a pathway when the GSEA FDR corrected p-value was less than 0.05. The GBA was implemented separately for the primary models of HIV latency and results were compared and overlapped afterwards. Visualization of significant pathway- and mRNA-associations was performed with in-house R scripts based on the ggplot2 package (v2.2.1). Visualization of individual lncRNA associations was performed using Cytoscape v3.4.0 [78].
Statistical analyses
All statistical analyses were performed in the R computing environment [71]. Effect size in the RNA-Seq experiment was determined using library RNASeqPower [79]. DdPCR data was expressed as copies of each lncRNA per one thousand copies of RPL27. Because of lower expression of NRON, this lncRNA was expressed as copies per hundred thousand copies of RPL27. These normalized copies were log2 transformed. The equal variance test was performed using function var.test. Based on these results, paired t-tests (function t.test) with equal or unequal variance were used for two-group comparisons. For ddPCR validation experiments, we had prior knowledge about the direction of change of expression in models of latency as compared to mock-infected cells; therefore, one-sided tests were performed. For two-group comparisons without prior knowledge of the direction of change, two-sided tests were conducted. Repeated measures analysis of variance (RM ANOVA) was implemented for multi-group comparisons using library nlme [80] followed by post-hoc Tukey test. For each model, residuals were measured and tested for normality using the shapiro.test in R. Examination of the distribution of residuals and ggplots indicated that there were no large deviations from normality; therefore the requirements of the t-test and ANOVA were satisfied. All graphs were constructed using GraphPad Prism software (GraphPad Software, La Jolla, CA, USA). If RNA from the same donor was used for multiple measurements, the symbol for that donor is consistently shaped and colored throughout.
Supporting information
Venn diagram was constructed using library VennDiagram v1.16.18 in the R computing environment. Dysregulated lncRNA and protein coding genes from S1 Table served as input for construction of the Venn diagram. Area of the diagram is proportional to the number of differentially expressed lncRNAs or protein coding genes.
(PDF)
Same samples that were sequenced (cultured TCM model, N = 4) and three additional bystander model samples (bystander model, N = 7) were subjected to ddPCR with assays to detect selected lncRNA. Expression of PVT1, RP11-347C18.3, RP11-539L10.2, NEAT1 and NRON was measured by ddPCR and normalized to expression of the housekeeping gene RPL27. Significance for ddPCR results was determined using a paired one-sided t-test for log2 transformed data. Data is presented as individual data points (copy numbers normalized to RPL27) symbol-coded by donor, mean of all values is shown. N1, NEAT1 isoform 1 (short); N2, NEAT1 isoform 2 (long); ns, not significant (p-value > 0.05); (*) represents experiments where for the cultured TCM model only three out of four sequenced sample pairs had sufficient RNA for testing.
(PDF)
Expression of PVT1, RP11-347C18.3, RP11-539L10.2 and NRON was measured by ddPCR and normalized to expression of the housekeeping gene RPL27. Out of six experiments shown in Fig 2, a subset of replicates had sufficient number of TEM cells to conduct the assays (N = 4 for PVT1, RP11-347C18.3 and RP11-539L10.2; N = 3 for NRON). Significance was determined by implementing repeated measures analysis of variance (RM ANOVA) with library nlme in R using log2 transformed data. Data is presented as individual data points (copy numbers normalized to RPL27) symbol-coded by donor, mean of all values is shown. TN, naïve CF4+ T-cells; TCM, central memory CD4+ T-cells, TEM, effector memory CD4+ T-cells.
(PDF)
A. Lists of common and unique pathways. B. Venn diagram showing overlap of pathways between the two models.
(PDF)
A. Lists of common and unique pathways. B. Venn diagram showing overlap of pathways between the two models.
(PDF)
Mock-infected cells and the bystander model of HIV latency were treated with SAHA (1μM) and RMD (15 nM) or their solvent DMSO for 24 hours. Expression of RP11-539L10.2 was measured by ddPCR and normalized to expression of the housekeeping gene RPL27. Four replicate experiments were performed. Significance was determined by implementing repeated measures analysis of variance (RM ANOVA) with library nlme in R using log2 transformed data. Data is presented as individual data points (copy numbers normalized to RPL27) symbol-coded by donor, mean of all values is shown. DMSO, dimethyl sulfoxide; SAHA, suberoylanilide hydroxamic acid; RMD, Romidepsin; ns, not significant (p-value > 0.05).
(PDF)
Data is organized in two Excel sheets, showing genes and lncRNAs dysregulated in the cultured TCM and the bystander models of HIV latency. Ensembl_ID, Ensembl gene identifier; HGNC_Gene_Symbol, gene symbol approved by the HUGO gene nomenclature committee (gencode 21); logFC, log2 transformed fold change of counts per million in the RNA-Seq data; FDR, false discovery rate adjusted p-value; gene_type, class of gene according to the nomenclature (protein coding, etc.). Data is sorted by gene_type and then by FDR within each gene type.
(XLSX)
Listed are KEGG and Biocarta pathways that were associated (FDR p<0.05) with RP11-347C18.3 and RP11-539L10.2 in common for the cultured TCM and the bystander models of HIV latency, and unique pathways found for each model. Pathways implicated in HIV latency are highlighted red.
(XLSX)
Acknowledgments
We acknowledge support from the University of San Diego (UCSD) Center for AIDS Research (CFAR; P30 AI036214), especially the Clinical Investigation Core for specimen collection, the Flow Cytometry Core for biohazardous sample sorting, and the Genomics and Sequencing Core for providing access to the server to process the RNA-Seq data and to the instruments for gene expression quantification by ddPCR. We thank Hosiana Abewe and Savitha Deshmukh for assistance with cell isolation for experiments using CD4+ T-cell subsets, and Merck, Inc. and Gilead, Inc. for providing reagents for this study.
Data Availability
Raw and processed data for RNA-Seq datasets are available at the Gene Expression Omnibus (GEO) database (URL: https://www.ncbi.nlm.nih.gov/geo/) under the identifiers GSE81810 (cultured TCM model) and GSE114883 (bystander model). All other relevant data are within the paper and its Supplementary Figures and Tables files.
Funding Statement
This work was supported by funding through a Developmental Grant to NBB from the San Diego CFAR (AI036214), a Career Developmental Award-II (IK2 BX002731) to NBB from the Office of Research and Development, Veterans Health Administration, the National Institutes of Health research grant (AI104282) to CAS, a grant from the Bill & Melinda Gates Foundation through the Grand Challenges Explorations initiative (OPP1045955) to AB, the Research Foundation Flanders (FWO 1.8.020.09.N.00) to LV, and the James B. Pendleton Charitable Trust. The views expressed in this article are those of the authors and do not necessarily reflect the position or policy of the Department of Veterans Affairs or the United States government. The sponsors of this research were not involved in the study design, collection or interpretation of the data, manuscript preparation, or the decision to submit the article for publication.
References
- 1.Richman DD, Margolis DM, Delaney M, Greene WC, Hazuda DJ, Pomerantz RJ. The challenge of finding a cure for HIV infection. Science. 2009;323(5919):1304–7. 10.1126/science.1165706 [DOI] [PubMed] [Google Scholar]
- 2.Ho YC, Shan L, Hosmane NN, Wang J, Laskey SB, Rosenbloom DI, et al. Replication-competent noninduced proviruses in the latent reservoir increase barrier to HIV-1 cure. Cell. 2013;155(3):540–51. 10.1016/j.cell.2013.09.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Castro-Gonzalez S, Colomer-Lluch M, Serra-Moreno R. Barriers for HIV cure: the latent reservoir. AIDS Res Hum Retroviruses. 2018;34(9):739–59. 10.1089/AID.2018.0118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Rasmussen TA, Lewin SR. Shocking HIV out of hiding: where are we with clinical trials of latency reversing agents? Curr Opin HIV AIDS. 2016;11(4):394–401. 10.1097/COH.0000000000000279 [DOI] [PubMed] [Google Scholar]
- 5.Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 2012;22(9):1775–89. 10.1101/gr.132159.111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhang Q, Chen C-Y, Yedavalli VS, Jeang K-T. NEAT1 long noncoding RNA and paraspeckle bodies modulate HIV-1 posttranscriptional expression. mBio. 2013;4(1):e00596–12. 10.1128/mBio.00596-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Liu H, Hu P-W, Couturier J, Lewis DE, Rice AP. HIV-1 replication in CD4+ T cells exploits the down-regulation of antiviral NEAT1 long non-coding RNAs following T cell activation. Virology. 2018;522:193–8. 10.1016/j.virol.2018.07.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Li J, Chen C, Ma X, Geng G, Liu B, Zhang Y, et al. Long noncoding RNA NRON contributes to HIV-1 latency by specifically inducing tat protein degradation. Nat Commun. 2016;7:11730 10.1038/ncomms11730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Imam H, Shahr Bano A, Patel P, Holla P, Jameel S. The lncRNA NRON modulates HIV-1 replication in a NFAT-dependent manner and is differentially regulated by early and late viral proteins. Sci Rep. 2015;5:8639 10.1038/srep08639 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cron RQ, Bartz SR, Clausell A, Bort SJ, Klebanoff SJ, Lewis DB. NFAT1 enhances HIV-1 gene expression in primary human CD4 T cells. Clin Immunol. 2000;94(3):179–91. 10.1006/clim.1999.4831 [DOI] [PubMed] [Google Scholar]
- 11.Huan C, Li Z, Ning S, Wang H, Yu X-F, Zhang W. Long noncoding RNA uc002yug.2 activates HIV-1 latency through regulation of mRNA levels of various RUNX1 isoforms and increased Tat expression. J Virol. 2018;92(9). 10.1128/jvi.01844-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ma L, Sun L, Jin X, Qu D, Sun W-W, Wang J-H, et al. Long noncoding RNA MALAT1 releases epigenetic silencing of HIV-1 replication by displacing the polycomb repressive complex 2 from binding to the LTR promoter. Nucleic Acids Res. 2019;47(6):3013–3027. 10.1093/nar/gkz117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009;136(4):629–41. 10.1016/j.cell.2009.02.006 [DOI] [PubMed] [Google Scholar]
- 14.Trypsteen W, Mohammadi P, Van Hecke C, Mestdagh P, Lefever S, Saeys Y, et al. Differential expression of lncRNAs during the HIV replication cycle: an underestimated layer in the HIV-host interplay. Sci Rep. 2016;6:36111 10.1038/srep36111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Peng X, Sova P, Green RR, Thomas MJ, Korth MJ, Proll S, et al. Deep sequencing of HIV-infected cells: insights into nascent transcription and host-directed therapy. J Virol. 2014;88(16):8768–82. 10.1128/JVI.00768-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Martins LJ, Bonczkowski P, Spivak AM, De Spiegelaere W, Novis CL, DePaula-Silva AB, et al. Modeling HIV-1 latency in primary T cells using a replication-competent virus. AIDS Res Hum Retroviruses. 2016;32(2):187–93. 10.1089/aid.2015.0106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bosque A, Planelles V. Induction of HIV-1 latency and reactivation in primary memory CD4+ T cells. Blood. 2009;113(1):58–65. 10.1182/blood-2008-07-168393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Spina CA, Anderson J, Archin NM, Bosque A, Chan J, Famiglietti M, et al. An in-depth comparison of latent HIV-1 reactivation in multiple cell model systems and resting CD4+ T cells from aviremic patients. PLoS Pathog. 2013;9(12). 10.1371/journal.ppat.1003834 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Beliakova-Bethell N, Mukim A, White CH, Deshmukh S, Abewe H, Richman DD, et al. Histone deacetylase inhibitors induce complex host responses that contribute to differential potencies of these compounds in HIV reactivation. J Biol Chem. 2019;294(14):5576–89. 10.1074/jbc.RA118.005185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bosque A, Famiglietti M, Weyrich AS, Goulston C, Planelles V. Homeostatic proliferation fails to efficiently reactivate HIV-1 latently infected central memory CD4+ T cells. PLoS Pathog. 2011;7(10):e1002288 10.1371/journal.ppat.1002288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.White CH, Moesker B, Beliakova-Bethell N, Martins LJ, Spina CA, Margolis DM, et al. Transcriptomic analysis implicates the p53 signaling pathway in the establishment of HIV-1 latency in central memory CD4 T cells in an In vitro model. PLoS Pathog. 2016;12(11):e1006026 10.1371/journal.ppat.1006026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Schoggins JW, Rice CM. Interferon-stimulated genes and their antiviral effector functions. Curr Opin Virol. 2011;1(6):519–25. 10.1016/j.coviro.2011.10.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Monroe KM, Yang Z, Johnson JR, Geng X, Doitsh G, Krogan NJ, et al. IFI16 DNA sensor is required for death of lymphoid CD4 T cells abortively infected with HIV. Science. 2014;343(6169):428–32. Epub 12/19. 10.1126/science.1243640 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Soriano-Sarabia N, Bateson RE, Dahl NP, Crooks AM, Kuruc JD, Margolis DM, et al. Quantitation of replication-competent HIV-1 in populations of resting CD4+ T cells. J Virol. 2014;88(24):14070–7. 10.1128/JVI.01900-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chun T-W, Carruth L, Finzi D, Shen X, DiGiuseppe JA, Taylor H, et al. Quantification of latent tissue reservoirs and total body viral load in HIV-1 infection. Nature. 1997;387(6629):183–8. 10.1038/387183a0 [DOI] [PubMed] [Google Scholar]
- 26.Siliciano JD, Kajdas J, Finzi D, Quinn TC, Chadwick K, Margolick JB, et al. Long-term follow-up studies confirm the stability of the latent reservoir for HIV-1 in resting CD4+ T cells. Nat Med. 2003;9(6):727–8. 10.1038/nm880 [DOI] [PubMed] [Google Scholar]
- 27.Spina CA, Prince HE, Richman DD. Preferential replication of HIV-1 in the CD45RO memory cell subset of primary CD4 lymphocytes in vitro. J Clin Invest. 1997;99(7):1774–85. 10.1172/JCI119342 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chomont N, El-Far M, Ancuta P, Trautmann L, Procopio FA, Yassine-Diab B, et al. HIV reservoir size and persistence are driven by T cell survival and homeostatic proliferation. Nat Med. 2009;15(8):893–900. 10.1038/nm.1972 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kaczmarek Michaels K, Natarajan M, Euler Z, Alter G, Viglianti G, Henderson AJ. Blimp-1, an intrinsic factor that represses HIV-1 proviral transcription in memory CD4+ T cells. J Immunol. 2015;194(7):3267–74. 10.4049/jimmunol.1402581 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Miller LK, Kobayashi Y, Chen C-C, Russnak TA, Ron Y, Dougherty JP. Proteasome inhibitors act as bifunctional antagonists of human immunodeficiency virus type 1 latency and replication. Retrovirology. 2013;10:120 10.1186/1742-4690-10-120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lin J, Zhang X, Lu W, Xu X, Pan X, Liang T, et al. PR-957, a selective immunoproteasome inhibitor, reactivates latent HIV-1 through p-TEFb activation mediated by HSF-1. Biochem Pharmacol. 2018. 10.1016/j.bcp.2018.08.042 [DOI] [PubMed] [Google Scholar]
- 32.Besnard E, Hakre S, Kampmann M, Lim HW, Hosmane NN, Martin A, et al. The mTOR complex controls HIV latency. Cell Host Microbe. 2016;20(6):785–97. 10.1016/j.chom.2016.11.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lassen KG, Ramyar KX, Bailey JR, Zhou Y, Siliciano RF. Nuclear retention of multiply spliced HIV-1 RNA in resting CD4+ T cells. PLoS Pathog. 2006;2:e68 10.1371/journal.ppat.0020068 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yukl SA, Kaiser P, Kim P, Telwatte S, Joshi SK, Vu M, et al. HIV latency in isolated patient CD4+ T cells may be due to blocks in HIV transcriptional elongation, completion, and splicing. Sci Transl Med. 2018;10(430). 10.1126/scitranslmed.aap9927 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mahiet C, Swanson CM. Control of HIV-1 gene expression by SR proteins. Biochem Soc Trans. 2016;44(5):1417–25. 10.1042/BST20160113 [DOI] [PubMed] [Google Scholar]
- 36.Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Nat Acad Sci U S A. 2009;106(28):11667–72. 10.1073/pnas.0904715106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Zhang R, Ni F, Fu B, Wu Y, Sun R, Tian Z, et al. A long noncoding RNA positively regulates CD56 in human natural killer cells. Oncotarget. 2016;7(45):72546–58. 10.18632/oncotarget.12466 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Zhai N, Xia Y, Yin R, Liu J, Gao F. A negative regulation loop of long noncoding RNA HOTAIR and p53 in non-small-cell lung cancer. OncoTargets Ther. 2016;9:5713–20. 10.2147/OTT.S110219 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Xu M-d, Wang Y, Weng W, Wei P, Qi P, Zhang Q, et al. A positive feedback loop of lncRNA PVT1 and FOXM1 facilitates gastric cancer growth and invasion. Clin Cancer Res. 2017;23(8):2071–80. 10.1158/1078-0432.CCR-16-0742 [DOI] [PubMed] [Google Scholar]
- 40.Erkelenz S, Hillebrand F, Widera M, Theiss S, Fayyaz A, Degrandi D, et al. Balanced splicing at the Tat-specific HIV-1 3'ss A3 is critical for HIV-1 replication. Retrovirology. 2015;12:29 10.1186/s12977-015-0154-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Tranell A, Fenyö EM, Schwartz S. Serine- and arginine-rich proteins 55 and 75 (SRp55 and SRp75) induce production of HIV-1 vpr mRNA by inhibiting the 5'-splice site of exon 3. J Biol Chem. 2010;285(41):31537–47. 10.1074/jbc.M109.077453 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Tranell A, Tingsborg S, Fenyö EM, Schwartz S. Inhibition of splicing by serine-arginine rich protein 55 (SRp55) causes the appearance of partially spliced HIV-1 mRNAs in the cytoplasm. Virus Res. 2011;157(1):82–91. 10.1016/j.virusres.2011.02.010 [DOI] [PubMed] [Google Scholar]
- 43.Shkreta L, Blanchette M, Toutant J, Wilhelm E, Bell B, Story BA, et al. Modulation of the splicing regulatory function of SRSF10 by a novel compound that impairs HIV-1 replication. Nucleic Acids Res. 2017;45(7):4051–67. 10.1093/nar/gkw1223 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Guo J, Hao C, Wang C, Li L. Long noncoding RNA PVT1 modulates hepatocellular carcinoma cell proliferation and apoptosis by recruiting EZH2. Cancer Cell Int. 2018;18:98 10.1186/s12935-018-0582-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tripathy MK, McManamy MEM, Burch BD, Archin NM, Margolis DM. H3K27 demethylation at the proviral promoter sensitizes latent HIV to the effects of vorinostat in ex-vivo cultures of resting CD4+ T cells. J Virol. 2015;89:8392–405. 10.1128/JVI.00572-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Zhao Y, Zhao J, Guo X, She J, Liu Y. Long non-coding RNA PVT1, a molecular sponge for miR-149, contributes aberrant metabolic dysfunction and inflammation in IL-1β-simulated osteoarthritic chondrocytes. Biosci Rep. 2018. 10.1042/bsr20180576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Fu C, Li D, Zhang X, Liu N, Chi G, Jin X. LncRNA PVT1 facilitates tumorigenesis and progression of glioma via regulation of miR-128-3p/GREM1 axis and BMP signaling pathway. Neurotherapeutics. 2018. 10.1007/s13311-018-0649-9 [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 48.He F, Song Z, Chen H, Chen Z, Yang P, Li W, et al. Long noncoding RNA PVT1-214 promotes proliferation and invasion of colorectal cancer by stabilizing Lin28 and interacting with miR-128. Oncogene. 2018. 10.1038/s41388-018-0432-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Huang F, Chen W, Peng J, Li Y, Zhuang Y, Zhu Z, et al. LncRNA PVT1 triggers Cyto-protective autophagy and promotes pancreatic ductal adenocarcinoma development via the miR-20a-5p/ULK1 Axis. Mol Cancer. 2018;17:98 10.1186/s12943-018-0845-6 [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 50.Yang Q, Yu Y, Sun Z, Pan Y. Long non-coding RNA PVT1 promotes cell proliferation and invasion through regulating miR-133a in ovarian cancer. Biomed Pharmacother. 2018;106:61–7. 10.1016/j.biopha.2018.06.112 [DOI] [PubMed] [Google Scholar]
- 51.Chang Z, Cui J, Song Y. Long noncoding RNA PVT1 promotes EMT via mediating microRNA-186 targeting of Twist1 in prostate cancer. Gene. 2018;654:36–42. 10.1016/j.gene.2018.02.036 [DOI] [PubMed] [Google Scholar]
- 52.Henderson LJ, Sharma A, Monaco MCG, Major EO, Al-Harthi L. Human immunodeficiency virus type 1 (HIV-1) transactivator of transcription through its intact core and cysteine-rich domains inhibits Wnt/β-catenin signaling in astrocytes: relevance to HIV neuropathogenesis. J Neurosci. 2012;32(46):16306–13. 10.1523/JNEUROSCI.3145-12.2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Farivar S, Shaabanpour Aghamaleki F. Effects of major epigenetic factors on systemic lupus erythematosus. Iran Biomed J. 2018;22(5):294–302. 10.29252/ibj.22.5.294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Rai R, Chauhan SK, Singh VV, Rai M, Rai G. RNA-seq analysis reveals unique transcriptome signatures in systemic lupus erythematosus patients with distinct autoantibody specificities. PLoS ONE. 2016;11(11):e0166312–e. 10.1371/journal.pone.0166312 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Beliakova-Bethell N, Zhang J, Singhania A, Lee V, Terry V, Richman DD, et al. Suberoylanilide hydroxamic acid induces limited changes in the transcriptome of primary CD4+ T cells. AIDS. 2013;27:29–37. 10.1097/QAD.0b013e32835b3e26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Wozniak MB, Villuendas R, Bischoff JR, Aparicio CB, Martínez Leal JF, de La Cueva P, et al. Vorinostat interferes with the signaling transduction pathway of T-cell receptor and synergizes with phosphoinositide-3 kinase inhibitors in cutaneous T-cell lymphoma. Haematologica. 2010;95(4):613–21. 10.3324/haematol.2009.013870 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.LaBonte M, Wilson P, Fazzone W, Groshen S, Lenz H-J, Ladner R. DNA microarray profiling of genes differentially regulated by the histone deacetylase inhibitors vorinostat and LBH589 in colon cancer cell lines. BMC Med Genomics. 2009;2:67 10.1186/1755-8794-2-67 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.White CH, Johnston HE, Moesker B, Manousopoulou A, Margolis DM, Richman DD, et al. Mixed effects of suberoylanilide hydroxamic acid (SAHA) on the host transcriptome and proteome and their implications for HIV reactivation from latency. Antiviral Res. 2015;123:78–85. 10.1016/j.antiviral.2015.09.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Macedo AB, Resop RS, Martins LJ, Szaniawski MA, Sorensen ES, Spivak AM, et al. Influence of biological sex, age, and HIV status in an in vitro primary cell model of HIV latency using a CXCR4 tropic virus. AIDS Res Hum Retroviruses. 2018;34(9):769–77. 10.1089/AID.2018.0098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Noyce RS, Taylor K, Ciechonska M, Collins SE, Duncan R, Mossman KL. Membrane perturbation elicits an IRF3-dependent, interferon-independent antiviral response. J Virol. 2011;85(20):10926–31. 10.1128/JVI.00862-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Noyce RS, Collins SE, Mossman KL. Identification of a novel pathway essential for the immediate-early, interferon-independent antiviral response to enveloped virions. J Virol. 2006;80(1):226–35. 10.1128/JVI.80.1.226-235.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Rossio JL, Esser MT, Suryanarayana K, Schneider DK, Bess JW Jr., Vasquez GM, et al. Inactivation of human immunodeficiency virus type 1 infectivity with preservation of conformational and functional integrity of virion surface proteins. J Virol. 1998;72:7992–8001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Park J, Lim CH, Ham S, Kim SS, Choi B-S, Roh T-Y. Genome-wide analysis of histone modifications in latently HIV-1 infected T cells. AIDS. 2014;28(12):1719–28. 10.1097/QAD.0000000000000309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Abdel-Mohsen M, Wang C, Strain MC, Lada SM, Deng X, Cockerham LR, et al. Select host restriction factors are associated with HIV persistence during antiretroviral therapy. AIDS. 2015;29(4):411–20. 10.1097/QAD.0000000000000572 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Chen H, Li C, Huang J, Cung T, Seiss K, Beamon J, et al. CD4(+) T cells from elite controllers resist HIV-1 infection by selective upregulation of p21. J Clin Invest. 2011;121(4):1549–60. 10.1172/JCI44539 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25(9):1105–11. 10.1093/bioinformatics/btp120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Anders S, Pyl PT, Huber W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31(2):166–9. 10.1093/bioinformatics/btu638 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10(3):R25–R. 10.1186/gb-2009-10-3-r25 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Robinson MD, McCarthy DJ, Smyth GK. EdgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26(1):139–40. 10.1093/bioinformatics/btp616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Risso D, Ngai J, Speed TP, Dudoit S. Normalization of RNA-seq data using factor analysis of control genes or samples. Nat Biotechnol. 2014;32(9):896–902. 10.1038/nbt.2931 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.R: A language and environment for statistical computing. Vienna, Austria: R Foundation of Statistical Computing; R Core Team, 2017. [Google Scholar]
- 72.Lawrence M, Huber W, Pagès H, Aboyoun P, Carlson M, Gentleman R, et al. Software for computing and annotating genomic ranges. PLoS Comput Biol. 2013;9(8):e1003118 10.1371/journal.pcbi.1003118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Beliakova-Bethell N, Massanella M, White C, Lada SM, Du P, Vaida F, et al. The effect of cell subset isolation method on gene expression in leukocytes. Cytometry Part A. 2014;85A(1):94–104. 10.1002/cyto.a.22352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Lefever S, Anckaert J, Volders P-J, Luypaert M, Vandesompele J, Mestdagh P. decodeRNA—predicting non-coding RNA functions using guilt-by-association. Database (Oxford). 2017;2017:bax042 10.1093/database/bax042 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005;102:15545–50. 10.1073/pnas.0506580102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Nishimura D. BioCarta. Biotech Softw Internet Rep. 2001;2(3):117–20. 10.1089/152791601750294344 [DOI] [Google Scholar]
- 77.Kanehisa M, Goto S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000;28(1):27–30. 10.1093/nar/28.1.27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504. 10.1101/gr.1239303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Therneau T, Hart S, Kocher J-P. Calculating sample size estimates for RNA Seq studies. R package version 1.6.0. [DOI] [PMC free article] [PubMed]
- 80.Pinheiro J, Bates D, DebRoy S, Sarkar D, Team RC. Nlme: linear and nonlinear mixed effects models. R package version 3.1–127. 2016.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Venn diagram was constructed using library VennDiagram v1.16.18 in the R computing environment. Dysregulated lncRNA and protein coding genes from S1 Table served as input for construction of the Venn diagram. Area of the diagram is proportional to the number of differentially expressed lncRNAs or protein coding genes.
(PDF)
Same samples that were sequenced (cultured TCM model, N = 4) and three additional bystander model samples (bystander model, N = 7) were subjected to ddPCR with assays to detect selected lncRNA. Expression of PVT1, RP11-347C18.3, RP11-539L10.2, NEAT1 and NRON was measured by ddPCR and normalized to expression of the housekeeping gene RPL27. Significance for ddPCR results was determined using a paired one-sided t-test for log2 transformed data. Data is presented as individual data points (copy numbers normalized to RPL27) symbol-coded by donor, mean of all values is shown. N1, NEAT1 isoform 1 (short); N2, NEAT1 isoform 2 (long); ns, not significant (p-value > 0.05); (*) represents experiments where for the cultured TCM model only three out of four sequenced sample pairs had sufficient RNA for testing.
(PDF)
Expression of PVT1, RP11-347C18.3, RP11-539L10.2 and NRON was measured by ddPCR and normalized to expression of the housekeeping gene RPL27. Out of six experiments shown in Fig 2, a subset of replicates had sufficient number of TEM cells to conduct the assays (N = 4 for PVT1, RP11-347C18.3 and RP11-539L10.2; N = 3 for NRON). Significance was determined by implementing repeated measures analysis of variance (RM ANOVA) with library nlme in R using log2 transformed data. Data is presented as individual data points (copy numbers normalized to RPL27) symbol-coded by donor, mean of all values is shown. TN, naïve CF4+ T-cells; TCM, central memory CD4+ T-cells, TEM, effector memory CD4+ T-cells.
(PDF)
A. Lists of common and unique pathways. B. Venn diagram showing overlap of pathways between the two models.
(PDF)
A. Lists of common and unique pathways. B. Venn diagram showing overlap of pathways between the two models.
(PDF)
Mock-infected cells and the bystander model of HIV latency were treated with SAHA (1μM) and RMD (15 nM) or their solvent DMSO for 24 hours. Expression of RP11-539L10.2 was measured by ddPCR and normalized to expression of the housekeeping gene RPL27. Four replicate experiments were performed. Significance was determined by implementing repeated measures analysis of variance (RM ANOVA) with library nlme in R using log2 transformed data. Data is presented as individual data points (copy numbers normalized to RPL27) symbol-coded by donor, mean of all values is shown. DMSO, dimethyl sulfoxide; SAHA, suberoylanilide hydroxamic acid; RMD, Romidepsin; ns, not significant (p-value > 0.05).
(PDF)
Data is organized in two Excel sheets, showing genes and lncRNAs dysregulated in the cultured TCM and the bystander models of HIV latency. Ensembl_ID, Ensembl gene identifier; HGNC_Gene_Symbol, gene symbol approved by the HUGO gene nomenclature committee (gencode 21); logFC, log2 transformed fold change of counts per million in the RNA-Seq data; FDR, false discovery rate adjusted p-value; gene_type, class of gene according to the nomenclature (protein coding, etc.). Data is sorted by gene_type and then by FDR within each gene type.
(XLSX)
Listed are KEGG and Biocarta pathways that were associated (FDR p<0.05) with RP11-347C18.3 and RP11-539L10.2 in common for the cultured TCM and the bystander models of HIV latency, and unique pathways found for each model. Pathways implicated in HIV latency are highlighted red.
(XLSX)
Data Availability Statement
Raw and processed data for RNA-Seq datasets are available at the Gene Expression Omnibus (GEO) database (URL: https://www.ncbi.nlm.nih.gov/geo/) under the identifiers GSE81810 (cultured TCM model) and GSE114883 (bystander model). All other relevant data are within the paper and its Supplementary Figures and Tables files.