Abstract
Nucleic acid-sensing Toll-like receptors (TLRs) are subject to complex regulation to facilitate recognition of microbial DNA and RNA while limiting recognition of self-nucleic acids1. Failure to properly regulate these TLRs can lead to autoimmune and autoinflammatory disease2–6. Intracellular localization of these receptors is thought to be critical for self vs. non-self discrimination7, yet the molecular mechanisms that reinforce compartmentalized activation of intracellular TLRs remain poorly understood. Here we describe a new mechanism that prevents TLR9 activation from locations other than endosomes. This control is achieved through the regulated release of the receptor from its trafficking chaperone Unc93b1, which only occurs within endosomes and is required for ligand binding and signal transduction. Preventing TLR9 release from Unc93b1, either through mutations in Unc93b1 that increase affinity for TLR9 or through an artificial tether that impairs release, results in defective signaling. While TLR9 and TLR3 release from Unc93b1, TLR7 does not dissociate from Unc93b1 in endosomes and is regulated via distinct mechanisms. This work defines a new checkpoint that reinforces compartmentalized activation of TLR9 and provides a mechanism by which activation of individual endosomal TLRs may be distinctly regulated.
The multi-pass transmembrane protein Unc93b1 contributes to the compartmentalized activation of nucleic acid-sensing TLRs by mediating their trafficking from the ER to endosomes8. Mutations in Unc93b1 that result in aberrant TLR trafficking can lead to autoimmune disease3,9. It has been suggested that Unc93b1 may regulate additional sorting steps that are distinct between individual TLRs10; however, the mechanisms by which Unc93b1 contributes to the proper compartmentalization of endosomal TLRs and differentially regulates their responses remain unclear.
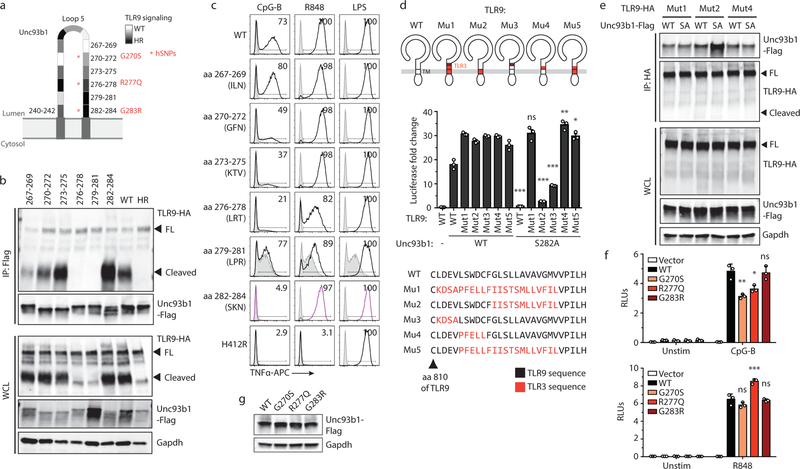
To dissect the mechanisms by which Unc93b1 regulates TLR signaling, we performed a scanning-alanine mutagenesis screen of Unc93b1 in RAW264.7 macrophages (for further information see Methods). The screen identified an Unc93b1 mutation (SKN to AAA at residues 282–284 in loop 5, hereafter referred to as Unc93b1SKN) that abolished signaling of TLR9, but did not affect TLR7 or TLR3 (Fig. 1a,b). Surprisingly, TLR9 trafficking appeared unaltered, as receptor cleavage, which occurs within endosomes, was unaffected (Fig. 1c). Also, Unc93b1SKN itself trafficked normally to Lamp1+ endosomes (Extended Data Fig. 1a). Disruption of TLR9 signaling without abrogating trafficking to endosomes was unexpected and suggested a new trafficking-independent function for Unc93b1.
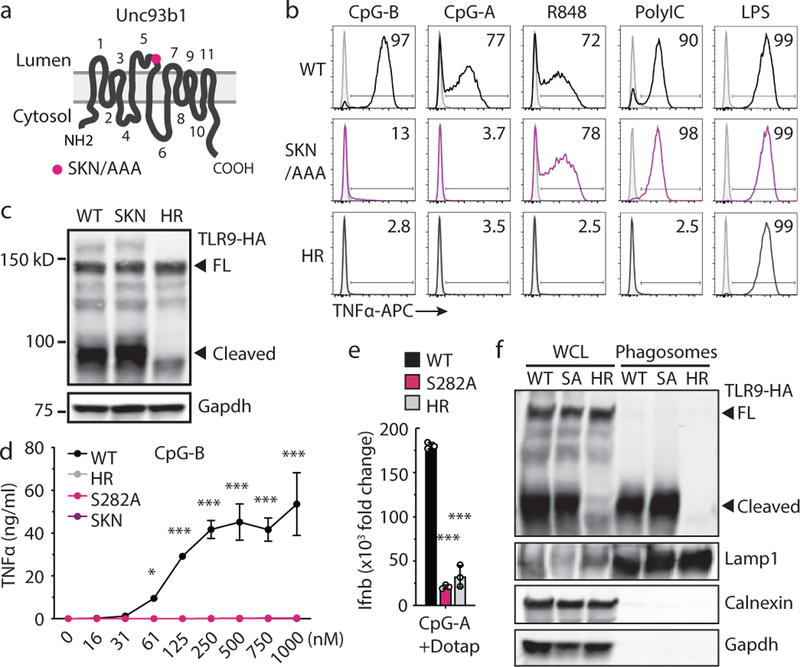
Fig. 1: An Unc93b1 mutation results in defective TLR9 signaling despite normal trafficking.
(a) Schematic of Unc93b1 topology with the SKN/AAA mutation indicated. (b) Intracellular cytokine staining of TNFα in Unc93b1-deficient RAW macrophage lines expressing the indicated Unc93b1 alleles (WT, SKN, and non-functional HR) after stimulation with CpG-B or CpG-A (150nM), R848 (25ng/ml), PolyIC (500ng/ml), or LPS (10ng/ml). Shaded histograms show unstimulated controls. (c) Immunoblot of TLR9-HA from the RAW macrophage lines shown in (b). FL: full-length. (d) TNFα production of the indicated RAW macrophage lines after 8h stimulation with increasing concentrations of CpG. n=2 biological replicates, representative of two independent experiments. (e) Quantitative RT-PCR analysis of Ifnb expression in the indicated RAW macrophage lines after 8h stimulation with CpG-A/DOTAP (1µM); n=3 biological replicates, representative of two independent experiments. (f) Whole cell (WCL) or phagosome lysates of the indicated RAW macrophage lines probed for TLR9-HA, Lamp-1, Calnexin, and Gapdh. All data are mean ± s.d.; p-values in (d) determined by two-way ANOVA followed by a Tukey’s post-test, WT vs S282A: * indicates p=0.0208, *** indicates p<0.0001; p-values in (e) determined by unpaired two-tailed Student’s t-test, *** indicates p<0.0001. All data are representative of at least three independent experiments, unless otherwise noted.
We individually mutated each amino acid within the SKN motif to alanine and determined that the S282A (SA) mutation was sufficient to abolish TLR9 signaling, whereas K283A and N284A were not (Extended Data Fig. 1b). Similar to the Unc93b1SKN mutant, Unc93b1S282A-expressing macrophages failed to respond to TLR9, while TLR3, TLR7, TLR5, and TLR13 responses were unaffected (Fig. 1d and Extended Data Fig. 1c–f). Induction of IFNß after stimulation with CpG-A/Dotap was also abrogated (Fig. 1e). Again, levels of cleaved TLR9 in phagosomes were similar compared to wildtype, indicating normal trafficking (Fig. 1f).
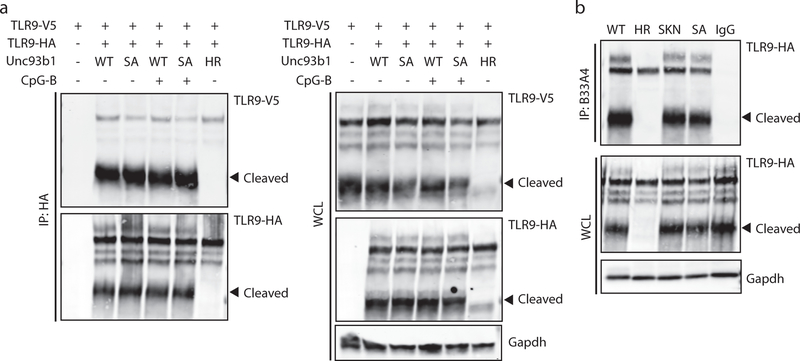
Based on the lumenal position of the Unc93b1S282A mutation, we reasoned that ligand binding might be affected. Indeed, CpG binding to TLR9 was reduced in mutant cells (Fig. 2a). To rule out differences in DNA uptake or sampling, we verified that all cell lines were equally capable of endocytosing CpG-B (Extended Data Fig. 2a) and that the ligands were effectively delivered to Lamp1+ endolysosomes containing TLR9 (Extended Data Fig. 2b,c). We also ruled out defects in TLR9 dimerization11,12 and defects in the association between N-terminal and C-terminal fragments after receptor cleavage13 (Extended Data Fig. 3a,b). Altogether, these results suggested a reduced ability of TLR9 to bind ligand in Unc93b1S282A-expressing cells unrelated to known mechanisms of receptor regulation.
Figure 2. Unc93b1S282A attenuates ligand binding and increases association with TLR9.
(a) Biotin-CpG-bound TLR9 complexes were precipitated from lysates of RAW macrophage lines expressing TLR9-HA and the indicated Unc93b1 alleles after stimulation with Biotin-CpG (1µM) for 4h, followed by immunoblot for TLR9-HA. Bars show mean ± s.d. of CpG-bound TLR9 relative to total TLR9; n=3 (WT), n=4 (SKN or S282A), and n=2 (HR), each dot represents an independent experiment. (b) Immunoprecipitation of Unc93b1-Flag from the RAW macrophage lines in (a) followed by immunoblot of TLR9-HA. TLR9 levels in whole cell lysates (WCL) are also shown. Representative of six independent experiments. Bars show mean ± s.d. of TLR9 bound to Unc93b1; n=6 (WT) n=7 (SKN or S282A); each dot represents an independent experiment. (c) Proximity ligation assay to analyze interaction between TLR9-HA and the indicated Unc93b1 alleles in RAW macrophages. Representative images are shown. Control staining was performed by omitting primary anti-HA antibodies. Quantification of PLA signals per cell volume is shown below images. Each dot represents an individual cell. Data are pooled from two independent experiments. Scale bar = 10 µm (d) Immunoprecipitation of TLR9-HA from phagosome preparations of RAW macrophage lines in (a) followed by immunoblot for Unc93b1-Flag. TLR9 and Unc93b1 levels in phagosomes and whole cell lysates are also shown. Representative of two independent experiments. All p-values determined by unpaired two-tailed Student’s t-test.
We considered the possibility that a loss of interaction between Unc93b1S282A and TLR9 during trafficking might impair ligand binding. On the contrary, TLR9 showed greater interaction with the Unc93b1 SKN and S282A mutants (Fig. 2b and Extended Data Fig. 4a). Interaction with TLR7 was unaffected (Extended Data Fig. 4b). Furthermore, the increased interaction with TLR9 was not observed when the neighboring Unc93b1 residues K283 or N284 were mutated (Extended Data Fig. 4c). We confirmed the increased interaction between TLR9 and Unc93b1S282A by proximity ligation assay (Fig. 2c) and in purified phagosomes (Fig. 2d).
Examination of additional Unc93b1 mutants within loop 5 for their impact on TLR9 signaling and binding (Extended Data Fig. 5a) identified a larger region within Unc93b1 (spanning amino acids 270–284 and containing Ser282) in which mutations either abolished or increased the interaction with TLR9 (Extended Data Fig. 5b). Irrespective of the direction, all mutations consistently reduced TLR9 signaling (Extended Data Fig. 5c), highlighting the importance of an optimal binding affinity between Unc93b1 and TLR9 for proper TLR9 function. Most of the mutations had little or no effect on TLR7 function (Extended Data Fig. 5c). To identify the reciprocal binding region in TLR9 that engages with the loop 5 region of Unc93b1, we used a set of TLR9 chimeric variants in which varying segments of the juxtamembrane and transmembrane domains had been replaced with the corresponding sequences of TLR34. As TLR3 is unaffected by Unc93b1S282A (Extended Data Fig. 1c), we reasoned that this strategy could identify critical residues within TLR9 that mediate interaction with Unc93b1. The analysis identified a short motif of five amino acids (LSWDC) in the juxtamembrane region of TLR9 that when reverted to the corresponding TLR3 sequence (PFELL) restored function and normal interaction in the presence of Unc93b1S282A (Extended Data Fig. 5d,e). We conclude that the LSWDC motif within TLR9 engages with loop 5 of Unc93b1 and that mutations within this interaction surface can alter TLR9-Unc93b1 association and impact TLR9 function.
We next explored the importance of loop 5 for function of human Unc93b1. Two out of three reported SNPs within the respective Unc93b1 region (from the NCBI Single Nucleotide Polymorphism Database, with Minor Allele Frequencies between 8.5 – 9.1E-06) reduced TLR9 signaling (Extended Data Fig. 5f). The reduction in signaling was specific to TLR9, as none of the alleles reduced TLR7 signaling (Extended Data Fig. 5f). These results suggest that human genetic variants in this region of Unc93b1 may affect TLR9 function.
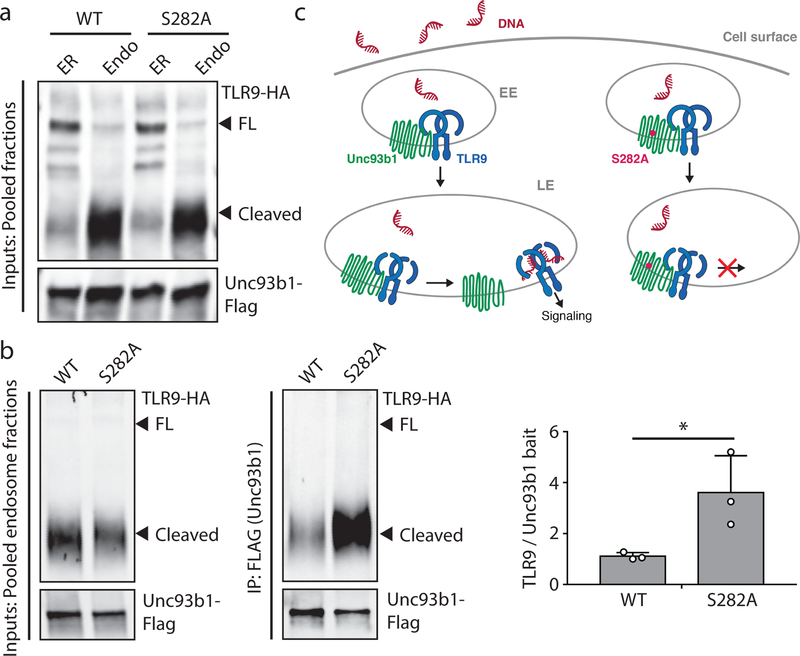
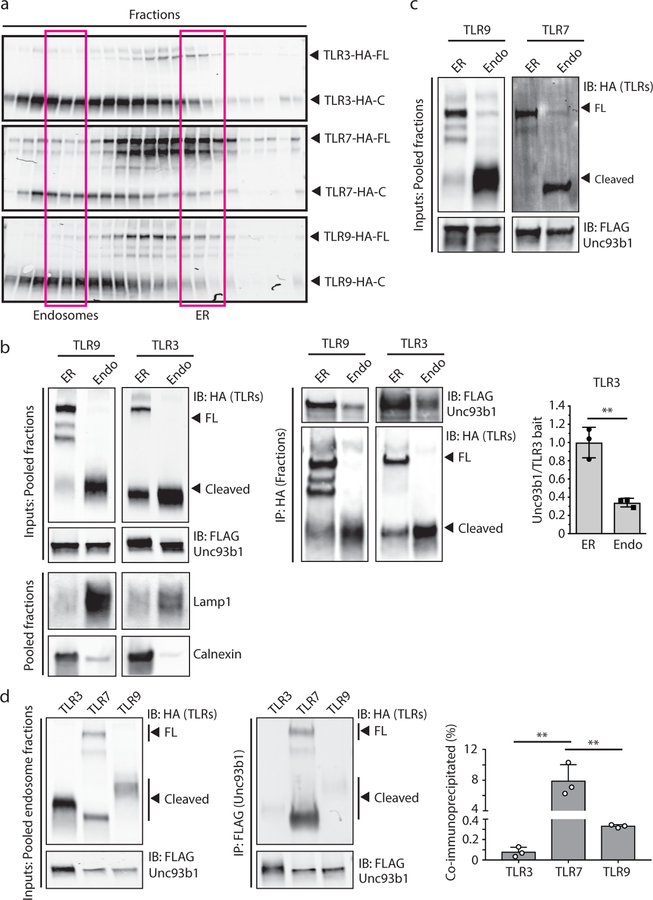
Based on the finding that an enhanced association with Unc93b1 is detrimental to TLR9 function, we reasoned that TLR9 might require release from Unc93b1 in endosomes prior to ligand binding and activation. We investigated this possibility by using cellular fractionation to separate ER and endosomes and measured the extent of TLR9-Unc93b1 interaction in each organelle preparation (Fig. 3a and Extended Data Fig. 9a). The selected endosomal fractions contained high levels of CpG-B-biotin ligand (when fed to cells), indicating that we had enriched the relevant signaling endosomes for TLR9 (Extended Data Fig. 6). As predicted, the association of Unc93b1 and TLR9 was weaker in endosomes than in the ER (Fig. 3b and Extended Data Fig. 7a). Furthermore, Unc93b1S282A showed an overall stronger association with TLR9, both in the ER and in endosomes (Fig. 3b and Extended Data Fig. 7b), suggesting that the altered chemistry of the Unc93b1 mutant might have an overall “sticky” effect that prevents efficient release of TLR9 in endosomes.
Fig. 3: TLR9, but not TLR7, must release from Unc93b1 for signaling.
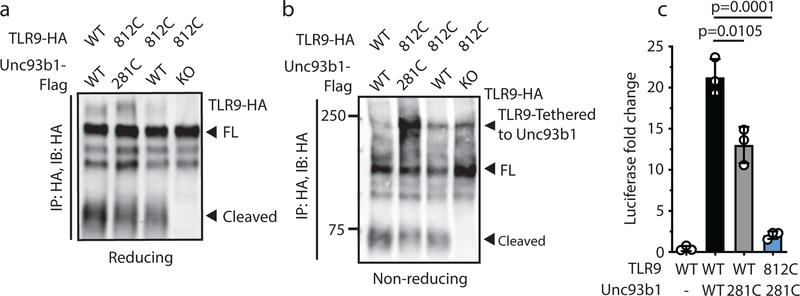
(a) Subcellular fractionation of RAW macrophage lines showing the distributions of TLR9, Unc93b1, and the indicated organelle markers. The pooled fractions, enriched for endosomes or ER, used for subsequent co-IP experiments are boxed. (b) Immunoprecipitation of TLR9-HA from pooled ER or endosome (Endo) fractions as shown in (a), followed by immunoblots for Unc93b1-Flag and TLR9-HA. Total TLR9-HA levels were normalized across fractions. Input controls are shown in Extended Data Fig. 7a. Bar graph shows mean ± s.d. of Unc93b1 bound to TLR9 across fractions. Each dot represents data from an independent experiment (n=3). p-values determined by paired two-tailed Student’s t-test. (c) Immunoprecipitation of TLR9-HA under non-reducing conditions from RAW macrophage lines expressing the indicated TLR9-HA and Unc93b1-Flag cysteine mutants, followed by immunoblot for Unc93b1. The non-reducing condition visualizes disulfide-bond formation between Unc93b1 and TLR9, which disappears after treatment with dithiothreitol (reducing condition). Bar graph below shows an NF-κB luciferase assay in HEK293T cells expressing the indicated cysteine mutant combinations and stimulated with CpG-B (1µM) for 16h. Data are normalized to Renilla expression and shown as relative luciferase units (RLUs); n=4 biological replicates. *** indicates p<0.0001 (unpaired two-tailed Student’s t-test.). (d) Immunoprecipitation of TLR9- and TLR7-HA from pooled ER or endosome fractions of macrophage lines co-expressing Unc93b1-Flag, followed by Flag and HA immunoblots. Input controls are shown in Extended Data Fig. 9c. Bar graph shows mean ± s.d. of Unc93b1 bound to TLR9 or TLR7 across ER and endosome fractions. Each dot represents data from an independent experiment (n=3). p-values determined by paired two-tailed Student’s t-test.
Based on these results we considered a release model, whereby TLR9 must dissociate from Unc93b1 for efficient ligand binding and signaling. Unc93b1 might interfere with DNA recognition by keeping the receptor in an unfavorable conformation for ligand binding, which would explain the attenuated ligand binding of TLR9 when forcefully bound to Unc93b1S282A (Extended Data Fig. 7c). To test this model independently and more directly, we engineered a cysteine bridge between TLR9 and Unc93b1 to tether the two proteins together and prevent release. We thereby focused on the previously identified loop 5 binding region of Unc93b1 and the juxtamembrane region of TLR9, (Fig. 3c). We identified a pair of Unc93b1 and TLR9 cysteine mutants (Unc93b1281C and TLR9812C) that trafficked to endosomes (Extended Data Fig. 8a) yet remained attached through an intermolecular disulfide bond (Fig. 3c, top left, and Extended Data Fig. 8b). Preventing release from Unc93b1 through this tether completely abrogated TLR9 signaling (Fig. 3c, bottom left). Importantly, Unc93b1281C and TLR9812C were functional when expressed alone, ruling out the possibility that the cysteine mutations simply created non-functional proteins (Fig. 3c, bottom left, and Extended Data Fig. 8c).
Next, we asked if other endosomal TLRs also require release from Unc93b1 for signaling. Similar to TLR9, TLR3 released from Unc93b1 in endosomes (Extended Data Fig. 9b); however the insensitivity of TLR3 to Unc93b1S282A suggests that interaction with Unc93b1 is mediated through a distinct interaction surface. Surprisingly, the interaction between TLR7 and Unc93b1 did not decrease in endosomes (Fig. 3d and Extended Data Fig. 9c,d), suggesting that TLR7 can bind ligand and signal while associated with Unc93b1. In fact, in an accompanying manuscript, we describe how the continued association of TLR7 and Unc93b1 in endosomes is critical for a distinct regulatory mechanism which prevents TLR7-dependent autoimmunity14. Thus, Unc93b1 utilizes different mechanisms to regulate activation of nucleic acid-sensing TLRs in endosomes.
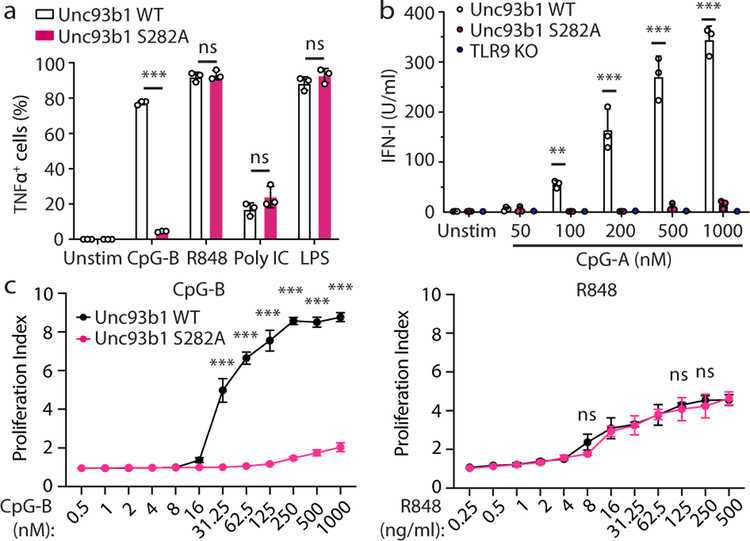
Finally, we tested the importance of TLR9 release from Unc93b1 in vivo. We introduced the Unc93b1S282A mutation into the germline of mice using Cas9 genome editing (Extended Data Fig. 10a). BMMs from these mice showed a selective loss of TLR9 responses, whereas TLR7, TLR3, and TLR4 responses were normal (Fig. 4a). Type I IFN production by pDCs in response to TLR9 stimulation was similarly ablated (Fig. 4b). Lastly, B cells from Unc93b1S282A mice did not proliferate in response to CpG-B, but showed normal proliferation in response to R848 and LPS (Fig. 4c and Extended Data Fig. 10b). These data support, across multiple cell types and with endogenous protein levels, a model in which TLR9 must be released from Unc93b1 in endosomes to function.
Fig. 4: Release from Unc93b1 is required for TLR9 function in vivo.
(a-c) Immune cells from Unc93b1S282A mice show a selective loss of TLR9 signaling. (a) Intracellular cytokine staining of TNFα in bone marrow-derived macrophages from Unc93b1WT/- and Unc93b1S282A/- mice after stimulation with CpG-B (0.5µM), R848 (25 ng/ml), Poly IC (20µg/ml), or LPS (50ng/ml). Data are mean ± s.d.; n=3 biological replicates. *** indicates p<0.0001, ns = not significant (unpaired two-tailed Student’s t-test). Representative of three independent experiments. (b) IFN-I production by pDCs after 16h stimulation with the indicated concentrations of CpG-A. Each data point represents one mouse (n=3). Data are mean ± s.d.; p-values determined by two-way ANOVA followed by a Tukey’s post-test. For WT compared to S282A, ** indicates p=0.0064 and *** indicates p<0.0001. Representative of three independent experiments. (c) Proliferation of CFSE-labeled splenic B cells from the indicated mice after stimulation for 3d with CpG-B or R848. The proliferation index is defined as geometric fluorescent intensity (gMFI) CSFEUnstim: gMFI CFSESample. Each curve shows the dose response of cells from three separate mice. Data are mean ± s.d.; p-values determined by two-way ANOVA followed by a Sidak’s post-test. *** indicates p<0.0001; ns = not significant.
The differential regulation of TLR9 and TLR7 that we describe may explain the enigmatic observation of TLR9 and TLR7 contributing distinctly to the pathology of certain autoimmune diseases15,16. Inhibition of TLR9 function is strictly linked to proper trafficking and localization, both through Unc93b1 association, as we describe here, and through the requirement for ectodomain proteolysis and ligand processing in endolysosomes17–20. Accordingly, overexpression of TLR9 does not induce disease21. In contrast, TLR7 appears subject to more ‘tunable’ regulation that dampens but does not eliminate signaling (see our accompanying paper14), and overexpression of TLR7 is sufficient to break tolerance and drive autoimmunity2,5,6. Why distinct mechanisms of regulation have evolved for such functionally similar innate receptors remains unclear. One possibility is that differences in the trafficking of TLR7 and TLR9 influence the likelihood that self RNA or DNA will be encountered; indeed, TLR9 traffics to endosomes via the plasma membrane while TLR7 is thought to bypass the plasma membrane10. Alternatively, the nature of the ligands recognized by each receptor may require differing degrees of tunability. Recent work has revealed that TLR7 and TLR8 ligands are quite simple (e.g., TLR7 recognizes the purine nucleoside guanosine together with a 3-mer uridine-containing ssRNA)22–24. In this case, avoiding self-recognition may require more subtle modulation of signaling than is necessary for TLR9. Regardless of any teleological rationale, dissecting the mechanisms that underlie differential regulation of these TLRs should reveal new avenues for therapeutic manipulation of TLR activation.
Methods
Antibodies and Reagents
The following antibodies were used for immunoblots, immunoprecipitations, or flow cytometry: rat anti-HA as purified antibody or matrix (3F10, Roche), mouse anti-FLAG as purified antibody or matrix (M2, Sigma-Aldrich), anti-V5 (rabbit polyclonal, AB3792, Millipore), anti-mLamp-1 (goat polyclonal, AF4320, R&D Systems), anti-Calnexin (rabbit polyclonal, ADI-SPA-860-F, Enzo Life Sciences), rabbit anti-Rab5 (C8B1, Cell Signaling), anti-EEA1 (rabbit polyclonal, 2411S, Cell Signaling), mouse anti-GM130 (610822, BD), anti-TLR9 (B33A4, kind gift from Kensuke Miyake), anti-Gapdh (GT239, GeneTex), anti-TNFα-APC (MP6-XT22, eBioscience), CD19 (6D5, BioLegend), purified anti-CD16/32 Fc Block (2.4G2, BD), goat anti-mouse IgG-AlexaFluor680 (Invitrogen), goat anti-rat IgG-AlexaFluor680 (Invitrogen), rabbit anti-goat IgG-AlexaFluor680 (Invitrogen), goat anti-mouse IRDye 800CW (Licor), donkey anti-rabbit IRDye 680RD (Licor), goat anti-rat IRDye 800CW (Licor). For immunofluorescence: rat anti-HA (3F10, Roche), rabbit anti-Lamp1 (ab24170, Abcam), donkey anti-rat AlexaFluor488 (Jackson Immunoresearch), donkey anti-rabbit AlexaFluor647 (Jackson Immunoresearch). Cells were mounted in Vectashield Hard Set Mounting Medium for Fluorescence (Vector Laboratories). For PLA: rabbit anti-HA (ab9110, Abcam), mouse anti-FLAG (M2, Sigma-Aldrich). For ELISA: anti-mouse TNFα purified (1F3F3D4, eBioscience), anti-mouse TNFα Biotin (XT3/XT22, eBioscience), Streptavidin HRP (BD Pharmingen).
The following TLR ligands were used: CpG-B (ODN1668: TCCATGACGTTCCTGATGCT, all phosphorothioate linkages) and CpG-A (ODN1585: G*G*GGTCAACGTTGAG*G*G*G*G*G, asterix indicate phosphorothioate linkages) were synthesized by Integrated DNA Technologies (Cy3 or biotin was attached to the 5-prime end for imaging or biochemistry experiments), R848 (Invivogen), PolyIC HMW (Invivogen), ultrapure Flagellin-ST (Invivogen), ORN Sa19 (Invivogen), and LPS (Invivogen). Human IL-1b was purchased from Invitrogen. NP-40 (Igepal CA-630) was purchased from Sigma-Aldrich. Saponin, was purchased from Acros. Lipofectamine-LTX reagent (Invitrogen) was used for transient transfection of plasmid DNA. DOTAP liposomal transfection reagent (Roche) was used for transfection of CpG-A. OptiMEM-I (Invitrogen) was used as media to form nucleic acid complexes for transient transfections. Streptavidin Magnetic Beads for biotin pull downs and Protein G agarose were purchased from Pierce. Violet fluorescent reactive dye for live dead staining of cells for flow cytometry was purchased from Invitrogen. ProMag 1 Series-COOH Surfactant free magnetic beads (#25029) for phagosome preparations were purchased from Polysciences. For renilla luciferase assays we used Coelenterazine native (Biotum). For firefly luciferase assays we used Luciferin (Biosynth). Assays were performed in Passive Lysis Buffer (Promega).
Unc93b1 library design and mutagenesis screen
The Unc93b1 mutagenesis library has been generated by Invitrogen. Briefly, the mouse Unc93b1 gene was optimized for the codon bias of Mus musculus and regions of very high (>80%) and very low (<30%) GC content have been avoided. The codon-optimized mouse Unc93b1 gene was C-terminally tagged with 3xFLAG (DYKDHDGDYKDHDIDYKDDDDK) and subjected to a triple-alanine scanning mutagenesis spanning sequences corresponding to tail and loop regions of the protein. The individual mutant constructs were cloned into a custom-made MSCV-based retroviral vector carrying an IRES-driven PuromycinR-T2A-mCherry double-selection. The library was provided as 204 individual plasmids.
Each Unc93b1 mutant was stably expressed in a RAW macrophage cell line in which both endogenous Unc93b1 alleles were disrupted by Cas9 genome editing. As expected, deletion of endogenous Unc93b1 led to lack of responses to nucleic acids and failure of TLR7 to traffic to endosomes (data not shown). To evaluate TLR function in cells expressing each mutant, we stimulated each line with ligands for TLR3, TLR7, and TLR9 (Unc93b1-dependent TLRs) and TLR4 (an Unc93b1-independent TLR) and measured TNFα production. RAW macrophages used for PolyIC and Flagellin stimulations were retrovirally transduced to express either TLR3-HA or TLR5-HA, respectively.
Plasmid constructs
For additional site-directed mutagenesis, AccuPrime Pfx DNA polymerase (Invitrogen) was used following the QuickChange II Site-directed Mutagenesis protocol from Agilent Technologies. The following MSCV-based retroviral vectors were used to express TLR7 and TLR9 in cell lines: MSCV2.2 (IRES-GFP), MSCV-Thy1.1 (IRES-Thy1.1), or MIGR2 (IRES-hCD2). TLR7 and TLR9 were fused to HA (YPYDVPDYA) at the C-terminal end. TLR7 sequence was synthesized after codon optimization by Invitrogen’s GeneArt Gene Synthesis service, as previously described10.
Mice
Mice were housed under specific-pathogen-free conditions at the University of California, Berkeley. All mouse experiments were performed in accordance with the guidelines of the Animal Care and Use Committee at UC Berkeley. C57BL/6J mice were from the Jackson Laboratory. Unc93b1S282A mice were generated using Cas9 genome editing. The guide RNA used was: TGCTGCGCGGCAGCGTCCGAAGG. The single-stranded oligo template contained a 91bp homology arm on the PAM proximal site and a 39bp homology arm on the PAM distal site. Four phosphothioate linkages were incorporated into the ends of the oligo template (one at the 5’ and three at the 3’ end of the oligo). Briefly, female C57BL/6J mice at 4 weeks of age were superovulated and mated overnight with C57BL/6J male mice (>8 weeks old). Zygotes were harvested from superovulated females and were placed in KSOM medium (MR106D, Millipore) before use. Embryos were transferred to 10µl Opti-MEM reduced serum media (31985062, Thermo Fisher Scientific) and then mixed with 10µl CRISPR reagent (i.e. Cas9/gRNA/ss oligo). This 20µl mixture (embryos in RNP solution) was undergone electroporation in a 1mm electroporation cuvette using a Gene Pulser XCell electroporator (1652660, Bio-Rad). After electroporation, 96 embryos were transferred to four CD1 recipients via oviduct transfer. Offspring were genotyped by sequencing for the correct targeted allele and further bred to ensure germline transmission. For experiments we used Unc93b1S282A/- mice and Unc93b1WT/- mice as littermate controls. Unless noted, mice were analyzed at 5–8 weeks of age. In total, a minimum of 8 mice per group were analyzed, divided between three experiments.
Type I interferon production by pDCs
Bone marrow was flushed and red blood cells were lysed using Ammonium-Chloride-Potassium (ACK) buffer (Gibco). 105 cells per well were seeded into 96-well flat-bottom plates. Cells were stimulated with various concentrations of CpG-A for 16h. The next day, supernatants were transferred onto L-292 ISRE-luciferase reporter cells to determine the amount of released type I IFN. Recombinant mouse IFN-β (pbl interferon source) was used for the standard curve. Reporter cells were incubated in bone marrow supernatants for 8h, lysed by passive lysis (Promega) and luciferase activity was measured on an LMaxII-384 luminometer (Molecular Devices).
B cell proliferation assay
Spleens were digested with collagenase 8 (Sigma) and DNase-I for 45 min and red blood cells were lysed using ACK buffer (Gibco). Splenocytes were labeled with 12.5µg/ml CFSE (Invitrogen) for 10min at 37°C and immediately underlayed with 3ml FCS to spin out CSFE. Cells were taken up in media (RPMI/10%FCS/L-glutamine/Pen-Strep/HEPES/Sodium pyruvate/ β-mercaptoethanol), counted, and seeded at 200,000 cells per well in round-bottom 96-well plates. Cells were incubated in media with various concentrations of CpG-B, R848, or LPS for 72h. Flow cytometry was used to analyze stimulated cells. Live, singlet cells were pre-gated on CD19+ and cell proliferation was determined by the geometric mean fluorescence intensity (gMFI) of CFSE (Extended Data Fig. 10c). For the quantification, a proliferation index was determined by dividing the gMFI CSFE of the unstimulated control by the gMFI CSFE of the stimulated sample (CSFEUnstim:CFSESample) and plotted along the ligand titration.
Plasmid constructs
AccuPrime Pfx DNA polymerase (Invitrogen) was used for site directed mutagenesis using the QuikChange II Site-directed Mutagenesis protocol from Agilent Technologies. The following murine stem cell virus (MSCV)-based retroviral vectors were used to express Unc93b1, TLR9, TLR7, and TLR3 in cell lines: MSCV-PuromCherry (IRES-PuromycinR-T2A-mCherry), MSCV2.2 (IRES-GFP), MSCV-Thy1.1 (IRES-Thy1.1), and MIGR2 (IRES-hCD2). 3×FLAG (DYKDHDGDYKDHDIDYKDDDDK) was fused to the C-terminus of Unc93b1. TLR9, TLR7, and TLR3 were fused to HA (YPYDVPDYA) at the C-terminal end. TLR7 sequence was synthesized after codon optimization by Invitrogen’s GeneArt Gene Synthesis service as previously described10. TLR9 chimeras of the juxtamembrane and transmembrane regions were previously described4.
Cells and tissue culture conditions
HEK293T cells were obtained from American Type Culture Collection (ATCC). GP2–293 packaging cell lines were obtained from Clontech. The above cell lines were cultured in DMEM complete media supplemented with 10% (vol/vol) FCS, L-glutamine, penicillin-streptomycin, sodium pyruvate, and HEPES (pH 7.2, Invitrogen). RAW264.7 macrophage cell lines (ATCC) were cultured in RPMI 1640 (same supplements as above).
BMMs were differentiated for seven days in RPMI complete media (same supplements as above plus 0.00034% (vol/vol) beta-mercaptoethanol) and supplemented with M-CSF containing supernatant from 3T3-CSF cells.
To generate HEK293T Unc93b1−/− cells, guide RNAs were designed and synthesized as gBlocks as previously described25 and subcloned into pUC19 (guide RNA: CTCACCTACGGCGTCTACC). Humanized Cas9–2xNLS-GFP was a gift from the Doudna laboratory, University of California, Berkeley, CA. HEK293T cells were transfected using Lipofectamine LTX with equal amounts of the guide RNA plasmid and Cas9 plasmid. Seven days post transfection cells were plated in a limiting-dilution to obtain single cells. Correct targeting was verified by PCR analysis and loss of response to TLR9 and TLR7 stimulation in an NF-κB luciferase assay. Unc93b1−/− RAW macrophages were generated with the Cas9(D10A)-GFP nickase (guide RNAs: 1) GGCGCTTGCGGCGGTAGTAGCGG, 2) CGGAGTGGTCAAGAACGTGCTGG, 3) TTCGGAATGCGCGGCTGCCGCGG, 4) AGTCCGCGGCTACCGCTACCTGG). Macrophages were transfected with cas9(D10A) and all four guide RNAs using Lipofectamine LTX and Plus reagent and single cell-sorted on cas9-GFP two days later. Correct targeting was verified by loss of response to TLR7 stimulation and sequencing of the targeted region after TOPO cloning.
Retroviral transduction
For retroviral transduction of RAW macrophages, VSV-G-pseudotyped retrovirus was made in GP2–293 packaging cells (Clontech). GP2–293 cells were transfected with retroviral vectors and pVSV-G using Lipofectamine LTX reagent. 24h post-transfection, cells were incubated at 32°C. 48h post-transfection viral supernatant (with polybrene at final 5μg/ml) was used to infect target cells overnight at 32°C and protein expression was checked 48h later. Target cells were sorted on a BD FACSAria Fusion Sorter to match expression or drug-selected with Puromycin, starting 48h after transduction. Efficiency of drug selection was verified by equal mCherry expression of target cells.
For retroviral transduction of bone marrow derived macrophages, bone marrow was harvested and cultured in M-CSF containing RPMI for two days. Progenitor cells were transduced with viral supernatant (produced as above) on two successive days by spinfection for 90min at 32°C. 48h after the second transduction cells were put on Puromycin selection and cultured in M-CSF containing RPMI media until harvested on day 8.
Luciferase assays
Activation of NF-κB in HEK293T cells was performed as previously described18. Briefly, transfections were performed in OptiMEM-I (Invitrogen) with LTX transfection reagent (Invitrogen) according to manufacturer’s guidelines. Cells were stimulated with CpG-B (200nM-1µM), R848 (100–200ng/ml), or human IL-1b (20ng/ml) after 24h and lysed by passive lysis after an additional 12–16h. Luciferase activity was measured on a LMaxII-384 luminometer (Molecular Devices).
Immunoprecipitation and western blot analysis
Cells or purified phagosomes were lysed in NP-40 buffer (50mM Tris-HCl [pH 7.4], 150mM NaCl, 1% NP-40, 5mM EDTA and supplemented with EDTA-free complete protease inhibitor cocktail (Roche) and 1mM PMSF). After incubation at 4°C on a rotator, lysates were cleared of insoluble material by centrifugation. For immunoprecipitations, lysates were incubated with anti-HA matrix or anti-FLAG matrix (pre-blocked with 1% BSA-PBS) overnight, and washed four times in lysis buffer the next day. For immunoprecipitations with anti-TLR9 (B33A4), cells were lysed in lysis buffer containing 20mM Tris-HCl [pH 7.4], 150mM NaCl, 1mM CaCl2, 10% glycerol, 1mM DTT, 0.5% NP-40, supplemented with EDTA-free complete protease inhibitor cocktail (Roche) and 1mM PMSF. Cleared lysates were incubated with 8µg/ml anti-TLR9 (B33A4) overnight. The next day, 50µl Protein G agarose (pre-blocked with 1% BSA-PBS) was added to lysates for 1–2h and washed four times in lysis buffer. Precipitated proteins were denatured in SDS-PAGE buffer at room temperature for 1h. Proteins were separated by SDS-PAGE (Bio-Rad TGX precast gels) and transferred to Immobilon PVDF membranes (Millipore) in a Trans-Blot Turbo transfer system (Bio-Rad). Membranes were probed with the indicated antibodies and developed using the Licor Odyssey Blot Imager. Relative band intensities were quantified using Fiji (ImageJ)26. For co-immunoprecipitation efficiencies in Extended Data Fig. 9d, the total amount of each TLR in the pooled endosomal fractions was calculated by multiplying the band intensities by the dilution factor (based on the volume loaded and the pooled fraction volume). The relative proportion of Unc93b1-bound TLR was calculated as the ratio of the quantified Unc93b1-bound TLR over the total TLR amount (converted to percentage for Extended Data Fig. 9d). To normalize across different experiments, equal amounts of protein standards were loaded in each gel and band intensities were used as a normalization reference.
Streptavidin pull-downs were performed on cells fed biotin-CpG-B for 4h, lysed in NP-40 buffer (same as above) and cleared of insoluble debris. Lysates were incubated for 2h with streptavidin magnetic beads (pre-blocked with 1% BSA-PBS), rotated at 4°C, and washed four times in lysis buffer. Precipitates were boiled in SDS buffer, separated by SDS–PAGE, and probed by anti-HA immunoblot.
For visualizing disulfide bond formation in RAW macrophages, cells were lysed in buffer containing 10% DDM/CHS detergent for 2h at 4°C. After removing insoluble material, lysates were incubated with HA matrix for 2–4h and washed four times in buffer containing 0.25% DDM/CHS. Protein was eluted in lysis buffer containing 10% DDM/CHS and 300µg/ml HA peptide for 1h at room temperature. Eluates were divided in half and denatured in either reducing (+DTT) or non-reducing (-DTT) SDS buffer for 1h at room temperature.
Flow cytometry
Cells were seeded into non-treated tissue culture 24-well plates or round-bottom 96-well plates. The next day cells were stimulated with the indicated TLR ligands. To measure TNFα production, BrefeldinA (BD GolgiPlug) was added to cells 30min after stimulation, and cells were collected after an additional 5.5h. Cells were stained for intracellular TNFα with a Fixation & Permeabilization kit according to manufacturer’s instructions (eBioscience).
For measuring DNA uptake, cells were fed Cy3-fluorescent CpG-B for the indicated amounts of time (control cells for no uptake were pre-chilled and stimulated on ice). Cells were washed 3 times with ice-cold PBS, fixed in 1% PFA/2%FCS/PBS, and analyzed on an BD LSR Fortessa flow cytometer.
Enzyme-linked immunosorbent assay (ELISA)
Cells were seeded into tissue culture-treated flat-bottom 96-well plates. The next day cells were stimulated with the indicated TLR ligands. For TNFα ELISAs NUNC Maxisorp plates were coated with anti-TNFα at 1.5µg/ml overnight at 4°C. Plates were then blocked with PBS + 1% BSA (w/v) at 37°C for 1h before cell supernatants diluted in PBS + 1% BSA (w/v) were added and incubated at room temperature for 2h. Secondary anti-TNFα biotin was used at 1µg/ml followed by Streptavidin-HRP. Plates were developed with 1mg/ml OPD in Citrate Buffer (PBS with 0.05M NaH2PO4 and 0.02M Citric acid, pH 5.0) with HCl acid stop.
Quantitative PCR
Cells were lysed in RNAzol (Molecular Research Center) and RNA was purified using the Direct-zol RNA MiniPrep Plus kit (Zymo Research) according to manufacturer’s instructions. RNA was treated with RQ1 RNase-free DNase (Promega) and concentrated using the RNA clean and concentrator-5 Kit (Zymo Research). cDNA was prepared from 100–500ng RNA with iScript cDNA synthesis kit, and quantitative PCR was performed with SYBR green on a StepOnePlus thermocycler (Applied Biosystems). Primers were synthesized by Integrated DNA Technologies. Primer sequences (PrimerBank): Ifnb1 F: AGCTCCAAGAAAGGACGAACA; Ifnb1 R: GCCCTGTAGGTGAGGTTGAT; Tnfa F: CAGGCGGTGCCTATGTCTC; Tnfa R: CGATCACCCCGAAGTTCAGTAG; Gapdh F: AGGTCGGTGTGAACGGATTTG; Gapdh R: GGGGTCGTTGATGGCAACA
Microscopy
Coverslips (High-performance, 18×18mm, thickness no. 1½ Zeiss) were acid-washed in 3M HCl, washed extensively in water, dipped in 70% EtOH and allowed to air-dry. Cells were plated onto coverslips and allowed to settle overnight. The next day, cells were incubated with Cy3-CpG-A for 2h at 37°C. Coverslips were washed with PBS, fixed with 4% PFA/PBS for 15min, and permeabilized with 0.5% saponin/PBS for 5min. To quench PFA autofluorescence, coverslips were treated with sodium borohydride/0.1% saponin/PBS for 10min. After washing 3x with PBS, cells were blocked in 1% BSA/0.1% saponin/PBS for 1h. Slides were stained in blocking buffer with anti-HA, and anti-Lamp1 (see antibodies above), washed with PBS and incubated for 45min with secondary antibodies. Cells were washed 3x in PBS and mounted in VectaShield Hard Set without Dapi. Cells were imaged on a Zeiss Elyra PS.1 with a 100x/1.46 oil immersion objective in Immersol 518F / 30°C 1004 QS (Zeiss). Z-Sections were acquired, with three grid rotations at each Z-position. The resulting dataset was SIM processed and Channel Aligned using Zeiss default settings in Zen. The completed super-resolution Z-Series was visualized and analyzed using Fiji (ImageJ). To compare the degree of colocalization of two proteins a single section from the middle of the Z-Series was selected and analyzed using a customized pipeline for object-based colocalization in Cell Profiler27. Briefly, primary objects (Unc93b1 vs Lamp1, or CpG-A vs TLR9) were identified and related to each other to determine the degree of overlap between objects. Data are expressed as % of object 1 colocalized with object 2. For some colocalization experiments, pixel intensities of two different fluorophores were measured by using the Plot Profile tool in Fiji to create a plot of intensity values along a line scan in the image.
Proximity Ligation assay
Coverslips (High-performance, 18×18mm, thickness no. 1½ Zeiss) were acid-washed in 3M HCl, washed extensively in water, dipped in 70% EtOH and allowed to air-dry. Cells were plated onto coverslips and allowed to settle overnight. Coverslips were washed with PBS, fixed with 4% PFA/PBS for 15min, and permeabilized with 0.5% saponin/PBS for 5min. To quench PFA autofluorescence, coverslips were treated with sodium borohydride/0.1% saponin/PBS for 10min. After washing 3x with PBS, cells were subjected to the PLA staining protocol. All steps, including blocking, incubations with primary antibodies (rabbit anti-HA (ab9110) and mouse anti-Flag (M2)) and Duolink PLA probes, ligase and polymerase were followed exactly according to the manufacturer’s instructions. Cells were mounted in Duolink In Situ mounting medium plus DAPI and imaged on a Zeiss Elyra PS.1 with a 63x/1.46 oil immersion objective (also see Microscopy section). The post-processed super-resolution Z-series were analyzed in Imaris x64 9.2.1. Briefly, surface volumes for each cell were generated using the dispersion of the PLA signal as a proxy for the cell boundary. Surface volumes were masked and the numbers of PLA spots within the masked areas were quantified using the spot finder tool. Spot size was set to 0.25µm and threshold to 2100.
Phagosome isolation
Cells in a confluent 15cm dish were incubated with ∼108 magnetic beads (1μm size, Polysciences) for 4h. After rigorous washing in PBS, cells were scraped into 10ml sucrose homogenization buffer (SHB: 250μM sucrose, 3mM imidazole pH 7.4) and pelleted by centrifugation. Cells were resuspended in 2ml SHB plus protease inhibitor cocktail with EDTA (Roche) and 1mM PMSF and disrupted by 25 strokes in a steel dounce homogenizer. The disrupted cells were gently rocked for 10min on ice to free endosomes. Beads were collected with a magnet (Dynal) and washed 4x with SHB plus protease inhibitor. After the final wash, phagosome preparations were denatured in 2x SDS buffer at room temperature for 1h and analyzed by western blot.
Cell fractionation by sucrose density-centrifugation
Cells in four confluent 15cm dishes were washed in ice-cold PBS, scraped in 10ml sucrose homogenization buffer (SHB: 250μM sucrose, 3mM imidazole pH 7.4) and pelleted by centrifugation. Cells were resuspended in 2ml SHB plus protease inhibitor cocktail with EDTA (Roche) and 1mM PMSF and disrupted by 25 strokes in a steel dounce homogenizer. The disrupted cells were centrifuged for 10min at 1,000g to remove nuclei. Supernatants were loaded onto continuous sucrose gradients (percent iodixanol: 0, 10, 20, 30) and ultracentrifuged in an SW41 rotor at 25,800rpm for 2h (Optima L-90K Ultracentrifuge, Beckman Coulter). 22 fractions of 420µl were collected from top to bottom. 100µl of each fraction were denatured in SDS buffer for western blot analysis. For immunoprecipitations, three fractions corresponding to ER or endosomes were combined and lysed for 1h after addition of protease inhibitor cocktail and NP-40 to a final concentration of 1%. Co-immunoprecipitation with anti-HA or anti-FLAG matrix was performed as described above.
CpG-B biotin dot blot
Cells in four confluent 15cm dishes were fed 1µM CpG-B-biotin for 4h and cell fractionation was performed as described in “Cell fractionation by sucrose density-centrifugation”. The individual fractions were lysed in 0.5% NP40 and 3µl lysate per fraction was spotted onto an Ambion BrightStar Plus Nylon membrane. The membrane was air-dried and DNA crosslinked in a Stratalinker on the auto-crosslink setting (1200 (x100) mJ/cm2). The membrane was washed in TTBS containing 0.1% SDS for 1h at room temperature to reduce background, followed by blocking in Odyssey blocking buffer for 1h at room temperature. The membrane was incubated with Streptavidin-IR 680 (1:5000) for 30min at room temperature, washed 3x in TTBS and imaged on a Licor Odyssey scanner.
β-Hexosaminidase activity assay
10µl of each gradient fraction was mixed with 50µl of substrate solution (6mM 4-methylumbelliferyl-N-acetyl-β-D-glucosaminidine (Sigma) dissolved in assay buffer (50mM citrate-phosphate buffer, pH 4.5; containing 0.1% Triton-X 100) and incubated for 20min at 37°C. Enzymatic reaction was stopped by adding 100µl Stop buffer (200mM Na2CO3, 110mM glycine) and fluorescence was read at ex 365/em 450nm in black assay plates.
Quantification and Statistical Analysis
Statistical parameters, including the number of independent repeats, the number of biological replicates per representative experiment, and statistical significance, are reported in the Figures and Figure Legends. Representative images have been repeated at least three times, unless otherwise stated in the figure legend. Data is judged to be statistically significant when p<0.05 by two-tailed Student’s t-test. To compare the means of different groups, a one-way ANOVA followed by a Tukey’s posttest was used. To compare means of different groups across a dose response, a two-way ANOVA followed by a Tukey’s or Sidak’s posttest was used. In figures, asterisks denote statistical significance (∗p<0.05, ∗∗p<0.01, ∗∗∗p<0.001). Statistical analysis was performed in GraphPad PRISM 7 (Graph Pad Software Inc.).
Extended Data
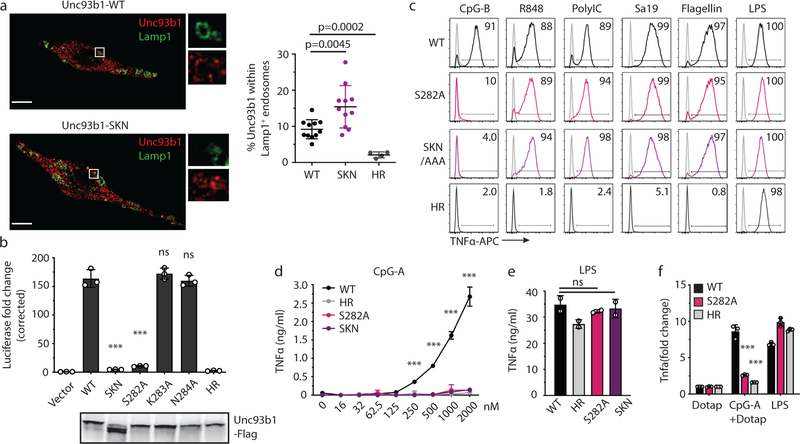
Extended Data Fig. 1: A luminal Unc93b1 mutation results in defective TLR9 signaling despite normal trafficking.
(a) Colocalization of Unc93b1 and Lamp1 in macrophages expressing the indicated Unc93b1 alleles using superresolution structured illumination microscopy. Shown are representative images: Unc93b1 (red) and Lamp1 (green). Boxed areas are magnified. The plot shows quantification of the percentage of total Unc93b1 within Lamp1+ endosomes. Each dot represents an individual cell. p values determined by unpaired two-tailed Student’s t-test. Data are from a single experiment. Scale bars: 10µm. (b) Unc93b1S282A is sufficient for the TLR9 signaling defect. NF-κB luciferase assay in HEK293T cells stimulated with CpG-B (1µM) for 16h. Data are normalized to Unc93b1-independent hIL-1b responses and expressed as luciferase fold change over unstimulated controls. n=3 biological replicates. *** indicates p<0.0001, ns = not significant (determined by unpaired two-tailed Student’s t-test). Blot below shows Unc93b1 expression levels. Representative of two independent experiments. (c) Intracellular cytokine staining of TNFα in macrophage lines expressing the indicated Unc93b1 alleles after stimulation with CpG-B (1 µM), R848 (100 ng/ml), PolyIC (100 ng/ml), Sa19 (200 ng/ml), Flagellin (100 ng/ml), and LPS (10 ng/ml). Gray histograms show unstimulated controls. Representative of two independent experiments. (d,e) TNFα production of the indicated macrophage lines after 8h stimulation with increasing concentrations of CpG-A (d), or LPS (50ng/ml) (e). n=2 biological replicates; *** indicates p<0.0001, ns = not significant, p values determined by two-way ANOVA followed by a Tukey’s posttest in (d) or by one-way ANOVA followed by a Tukey’s posttest in (e). Representative of two independent experiments. (f) Quantitative RT-PCR analysis of Tnfa expression in the indicated macrophage lines 8h after stimulation with CpG-A/DOTAP (1µM) or LPS (10ng/ml). n=3 biological replicates, *** indicates p=0.0003 for S282A vs WT) and p=0.0002 for HR vs WT, determined by unpaired two-tailed Student’s t-test. Representative experiment of two independent repeats. All data are shown as mean ± s.d.
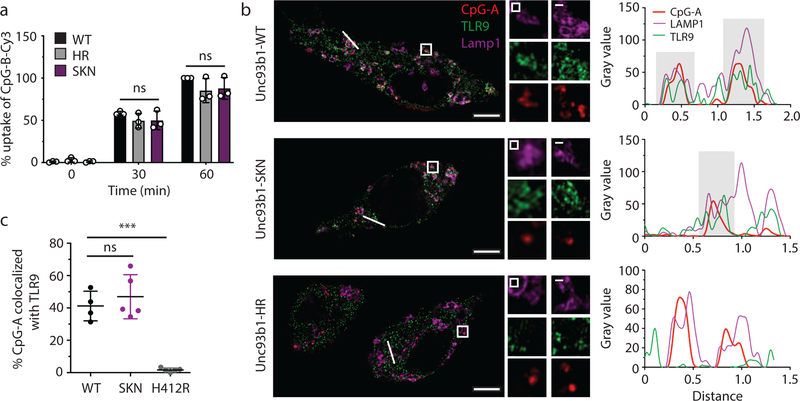
Extended Data Fig. 2: Unc93b1S282A does not affect DNA delivery to TLR9-containing endosomes.
(a) Uptake of Cy3-labeled CpG-B (1µM) of macrophage lines expressing the indicated alleles of Unc93b1-Flag. Data are presented as mean ± s.d. relative uptake compared to WT at 60 min. n=3 biological replicates. p-values determined by two-way ANOVA followed by a Tukey’s posttest when comparing WT vs SKN, ns = not significant. (b) Colocalization of CpG-A, TLR9-HA, and Lamp1 in macrophage lines shown in (a) after incubation with Cy3-labeled CpG-A (1µM) for 2h using superresolution structured illumination microscopy. Shown are representative images: TLR9 (green), Lamp1 (magenta), and CpG-A (red). Boxed areas and areas containing white lines are magnified. The histograms display fluorescent intensity plots of pixels along the white lines. Shaded areas highlight regions of colocalization of CpG-B, TLR9, and Lamp1. (c) Quantification of the percentage of CpG-A colocalized with TLR9. Each dot represents an individual cell, n=4 (WT), n=5 (SKN) and n=5 (HR). Data are mean ± s.d., *** indicates p<0.0001, ns = not significant. p values determined by unpaired two-tailed Student’s t-test. Scale bars: 5µm. Data are from a single experiment.
Extended Data Fig. 3: Unc93b1S282A does not affect TLR9 dimerization or the association between N-terminal and C-terminal cleavage products of TLR9.
(a) Macrophage lines co-expressing TLR9-HA and TLR9-V5 together with the indicated Unc93b1-Flag alleles were subjected to HA-immunoprecipitation followed by V5-immunoblot. TLR9 levels in whole cell lysates (WCL) are also shown. (b) Macrophage lines expressing TLR9-HA and the indicated Unc93b1-Flag alleles were subjected to immunoprecipitation with an anti-TLR9 antibody specific to the N-terminal cleavage fragment (B33A4), followed by immunoblot of the C-terminal TLR9 fragment with anti-HA. Data are representative of at least two independent experiments.
Extended Data Fig. 4: Unc93b1S282A shows a stronger interaction with TLR9, but not TLR7.
(a) Unc93b1SKN and Unc93b1S282A display a stronger association with TLR9. Immunoprecipitation of TLR9-HA from macrophage lines expressing the indicated Unc93b1 alleles, followed by immunoblot of Unc93b1-Flag. (b) Unc93b1SKN and Unc93b1S282A do not affect the interaction with TLR7. Immunoprecipitation of Unc93b1-Flag from macrophage lines expressing TLR7-HA and the indicated Unc93b1 alleles, followed by immunoblot of TLR7-HA. (c) Immunoprecipitation of Unc93b1-Flag from macrophage lines expressing the indicated Unc93b1 alleles, followed by immunoblot of TLR9-HA. All blots are representative of at least two independent experiments. SA: S282A. KA: K283A. NA: N284A. WCL: whole cell lysate. FL: full-length.
Extended Data Fig. 5: Identification of residues within loop 5 of Unc93b1 that mediate interaction with TLR9.
(a) Schematic of the tested loop 5 mutants of Unc93b1 and the relative TLR9 responses indicated in shades of gray: white indicates a response equivalent to WT while black indicates no response. Asterisks show human Unc93b1 SNPs that have been tested in (f). (b) A larger region in loop 5 of Unc93b1 mediates binding to TLR9. Immunoprecipitation of Unc93b1-Flag from macrophage lines expressing the indicated Unc93b1 mutants (spanning amino acids 267–284, and non-functional HR) followed by immunoblot of TLR9-HA. Representative of two independent experiments. (c) Intracellular cytokine staining of TNFα in macrophage lines shown in (b) after stimulation with CpG-B (25 nM), R848 (100 ng/ml), and LPS (10 ng/ml). Shaded histograms show unstimulated controls. Representative of three independent experiments. (d) Schematics showing relative positions and a sequence alignment (bottom) of swapped regions within the TLR9/3 chimeras. Colored regions indicate TLR3 sequences. NF-κB luciferase assay in HEK293T cells transiently transfected with the indicated TLR9 and Unc93b1 mutants and stimulated with CpG-B (200nM) for 16h. Data are normalized to Unc93b1-independent hIL-1b responses and expressed as luciferase fold change over unstimulated controls. Data are mean ± s.d., n=3 biological replicates. p-values determined by two-way ANOVA followed by a Sidak’s posttest comparing each TLR9 allele coexpressed with Unc93b1 WT vs S282A (TLR9WT: p<0.0001, TLR9Mut1: p>0.9999, TLR9Mut2: p<0.0001, TLR9Mut3: p<0.0001, TLR9Mut4: p=0.0020, TLR9Mut5: p=0.0171. Representative experiment of two independent repeats. (e) TLR9 mutants that rescue signaling in the presence of Unc93b1S282A also show normal binding to Unc93b1S282A. HA immunoprecipitation of the indicated TLR9 mutants transiently expressed in HEK293T cells that stably express the indicated Unc93b1-Flag alleles, followed by immunoblot of Unc93b1-Flag. Representative of three independent experiments. (f) Human Unc93b1 variants with SNPs in loop 5 show decreased TLR9 signaling. NF-κB luciferase assay in HEK293T cells expressing TLR9 or TLR7 and the indicated human Unc93b1-Flag variants and stimulated with CpG-B (250 nM) or R848 (250 ng/ml) for 16h, respectively. Data are normalized to Renilla expression and expressed as relative luciferase units (RLUs). Data are mean ± s.d., n=3 biological replicates. p-values are determined by one-way ANOVA followed by a Tukey’s posttest. For CpG-B stimulations: p=0.0048 (WT vs G270S), p=0.0113 (WT vs R277Q), p=0.9994 (WT vs G283R). Representative of four independent experiments. For R848 stimulations: p=0.2001 (WT vs G270S), p=0.0002 (WT vs R277Q), p=0.9933 (WT vs G283R). Representative of three independent repeats experiment. (g) Expression levels of the Unc93b1 mutants used in (f). *p< 0.05, **p< 0.01, ***p< 0.001, ns: not significant. WCL: whole cell lysate. FL: full-length.
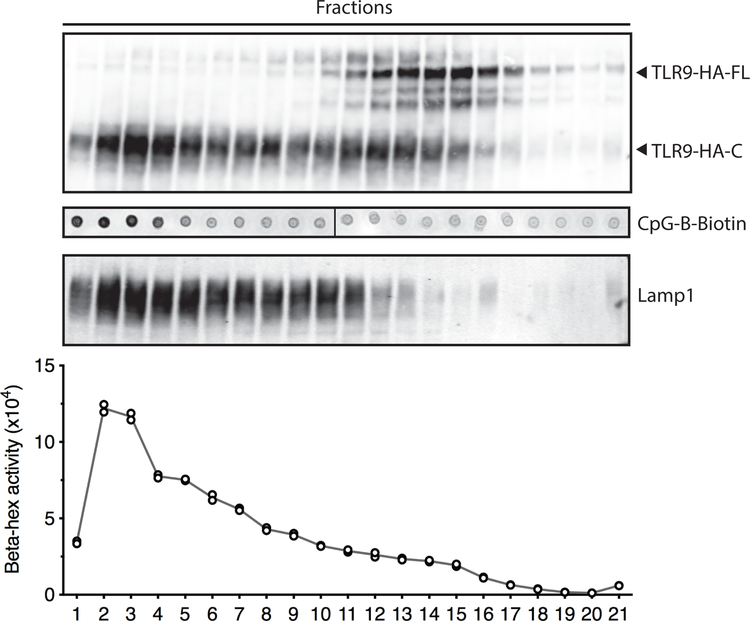
Extended Data Fig. 6: Cellular fractionation showing the distribution profiles for CpG-B-biotin ligand and β-hexosaminidase.
Macrophages were stimulated for 4h with biotinylated CpG-B (1µM) and subjected to sub-cellular fractionation by density-gradient centrifugation. The distributions of TLR9-HA, CpG-B, Lamp1, and β-hexosaminidase activity are shown. Representative of two independent experiments.
Extended Data Fig. 7: The TLR9-Unc93b1 association is reduced in endosomes compared to the ER.
(a) Input controls of TLR9 and Unc93b1 (relates to Fig. 3b). TLR9 and Unc93b1 levels in pooled ER or endosome fractions from macrophage lines expressing TLR9-HA and the indicated Unc93b1 alleles. FL: full-length. Representative of three independent experiments. (b) Increased interaction between TLR9 and Unc93b1S282A in endosomes. Immunoprecipitation of Unc93b1-Flag from pooled endosome fractions followed by immunoblot for TLR9-HA. Input controls are also shown. Representative of three independent experiments. Bar graph shows the quantification of TLR9 bound to Unc93b1 in pooled endosome fractions, normalized by Unc93b1-Flag levels in endosome fractions. Data are mean ± s.d., each dot represents data from an independent experiment (n=3). * indicates p=0.0413 (unpaired two-tailed Student’s t-test). (c) Release model of TLR9.
Extended Data Fig. 8: Tethering of TLR9 and Unc93b1.
(a) Cysteine mutants of TLR9 and Unc93b1 do not affect trafficking of TLR9 to endosomes. Immunoblot of TLR9-HA from macrophage lines expressing the indicated TLR9-HA and Unc93b1-Flag cysteine mutants. Representative of two independent experiments. (b) TLR9-HA immunoblot under non-reducing conditions after immunoprecipitation of TLR9-HA from macrophage lines shown in (a). The high molecular weight band indicates disulfide-bond formation between Unc93b1 and TLR9. Representative of two independent experiments. (c) Unc93b1-tethered TLR9 is unable to signal. NF-κB luciferase assay in HEK293T cells expressing the indicated cysteine mutant combinations and stimulated with CpG-B (1µM) for 16h. Data are normalized to Renilla expression and expressed as luciferase fold change over unstimulated controls. Data are mean ± s.d., n=3 biological replicates. p-values determined by unpaired two-tailed Student’s t-test. Representative of three independent experiments. FL: full-length.
Extended Data Fig. 9: TLR3 but not TLR7 releases from Unc93b1 in endosomes.
(a) Subcellular fractionation of macrophages lines showing the distributions of TLR3, TLR7, and TLR9 across fractions. The pooled endosome and ER fractions for subsequent coimmunoprecipitations are highlighted. (b) Immunoprecipitation of TLR9- and TLR3-HA from pooled ER or endosome fractions of macrophage lines expressing wildtype Unc93b1. Immunoprecipitated TLR-HA levels were normalized across fractions and probed for levels of Unc93b1-Flag. Bar graph shows the quantification of Unc93b1 bound to TLR3 between ER and endosome fractions. Data are mean ± s.d., each dot represents data from an independent experiment (n=3). ** indicates p=0.0039 (paired two-tailed Student’s t-test). (c) Input controls of TLR9, TLR7 and Unc93b1 in pooled ER and endosome fractions (relates to Fig. 3d). (d) Less TLR3 or TLR9 is associated with Unc93b1WT in pooled endosome fractions compared to TLR7. Immunoprecipitation of Unc93b1-Flag from pooled endosome fractions (as shown in (a)) followed by immunoblot for TLR3-, TLR7-, or TLR9-HA. Bar graph shows the calculated relative proportion of Unc93b1-bound TLR compared to total amount of the same TLR in the pooled endosome fractions. Data represent the mean ± s.d., each dot represents data from an independent experiment (n=3). ** indicates p=0.0032 (TLR3 vs TLR7) and p=0.0036 (TLR7 vs TLR9), determined by unpaired two-tailed Student’s t-test. All immunoblots are representative of three independent experiments. FL: full-length.
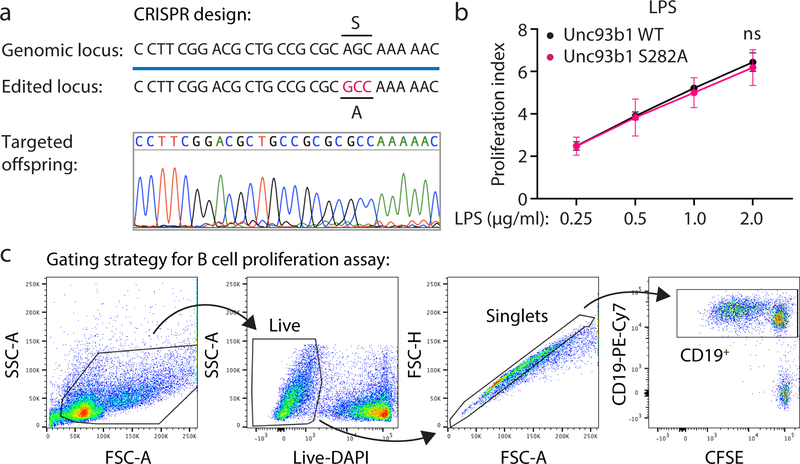
Extended Data Fig. 10. Generation of Unc93b1S282A knock-in mice and B-cell stimulation with LPS.
(a) CRISPR/Cas-9 strategy to generate Unc93b1S282A knock-in mice. Blue line indicates the guide sequence. Red bases indicated the edited codon. A representative sequencing trace of genomic DNA from an edited founder mouse is shown. (b) B cell proliferation in CSFE-labeled splenocyte cultures of the indicated mouse genotypes after stimulation for 3 days with increasing doses of LPS. The proliferation index is defined as geometric fluorescent intensity (gMFI) CSFEUnstim: gMFI CFSESample. Each curve shows the dose response of cells from three mice. Data are mean ± s.d. p values determined by two-way ANOVA followed by a Sidak’s posttest, ns = not significant. (c) Gating strategy for B cell stimulation assay.
Supplementary Material
Acknowledgments:
We thank members of the Barton and Vance Lab for helpful discussions and critical reading of the manuscript. We thank Baobin Li and Stephen Brohawn for supplying the DDM/CHS detergent and for technical advice, Angus Yiu-fai Lee and the Gene Targeting Facility of the Cancer Research Lab at UC Berkeley for generating the Unc93b1S282A/- knock-in mice, Hector Nolla and Alma Valeros for assistance with cell sorting at the Flow Cytometry Facility of the Cancer Research Laboratory at UC Berkeley, Steven Ruzin and Denise Schichnes for assistance with microscopy on the Zeiss Elyra PS.1 at the Biological Imaging Center at UC Berkeley. This work was supported by the NIH (AI072429, AI105184 and AI063302 to G.M.B.) and by the Lupus Research Institute (Distinguished Innovator Award to G.M.B.). O.M. was supported by an Erwin Schrödinger (J 3415-B22) and CRI Irvington postdoctoral fellowship. B.L. was supported by the UC Berkeley Tang Distinguished Scholars Program. B.J.W. was supported by a summer undergraduate research fellowship from UC Berkeley. Research reported in this publication was supported in part by the National Institutes of Health S10 program under award number 1S10OD018136–01.
Footnotes
Author Information: The authors declare no competing financial interest. Readers are welcome to comment on the online version of the paper.
Data availability: The datasets generated during and/or analyzed during the current study are either included within the manuscript or are available from the corresponding author on reasonable request. Source Data for Fig. 4 and Extended Data Fig. 10 are included in the online version of the paper. Gel source data can be found in Supplementary Fig. 1.
Supplementary Information is linked to the online version of the paper at www.nature.com/nature.
References:
- 1.Majer O, Liu B & Barton GM Nucleic acid-sensing TLRs: trafficking and regulation. Curr Opin Immunol 44, 26–33, doi: 10.1016/j.coi.2016.10.003 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Deane JA et al. Control of toll-like receptor 7 expression is essential to restrict autoimmunity and dendritic cell proliferation. Immunity 27, 801–810, doi: 10.1016/j.immuni.2007.09.009 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Fukui R et al. Unc93B1 restricts systemic lethal inflammation by orchestrating Toll-like receptor 7 and 9 trafficking. Immunity 35, 69–81, doi: 10.1016/j.immuni.2011.05.010 (2011). [DOI] [PubMed] [Google Scholar]
- 4.Mouchess ML et al. Transmembrane mutations in Toll-like receptor 9 bypass the requirement for ectodomain proteolysis and induce fatal inflammation. Immunity 35, 721–732, doi: 10.1016/j.immuni.2011.10.009 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Pisitkun P et al. Autoreactive B cell responses to RNA-related antigens due to TLR7 gene duplication. Science 312, 1669–1672, doi: 10.1126/science.1124978 (2006). [DOI] [PubMed] [Google Scholar]
- 6.Subramanian S et al. A Tlr7 translocation accelerates systemic autoimmunity in murine lupus. Proc Natl Acad Sci U S A 103, 9970–9975, doi: 10.1073/pnas.0603912103 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Barton GM, Kagan JC & Medzhitov R Intracellular localization of Toll-like receptor 9 prevents recognition of self DNA but facilitates access to viral DNA. Nat Immunol 7, 49–56, doi: 10.1038/ni1280 (2006). [DOI] [PubMed] [Google Scholar]
- 8.Kim YM, Brinkmann MM, Paquet ME & Ploegh HL UNC93B1 delivers nucleotide-sensing toll-like receptors to endolysosomes. Nature 452, 234–238, doi: 10.1038/nature06726 (2008). [DOI] [PubMed] [Google Scholar]
- 9.Fukui R et al. Unc93B1 biases Toll-like receptor responses to nucleic acid in dendritic cells toward DNA- but against RNA-sensing. J Exp Med 206, 1339–1350, doi: 10.1084/jem.20082316 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Lee BL et al. UNC93B1 mediates differential trafficking of endosomal TLRs. Elife 2, e00291, doi: 10.7554/eLife.00291 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Latz E et al. Ligand-induced conformational changes allosterically activate Toll-like receptor 9. Nat Immunol 8, 772–779, doi: 10.1038/ni1479 (2007). [DOI] [PubMed] [Google Scholar]
- 12.Ohto U et al. Structural basis of CpG and inhibitory DNA recognition by Toll-like receptor 9. Nature 520, 702–705, doi: 10.1038/nature14138 (2015). [DOI] [PubMed] [Google Scholar]
- 13.Onji M et al. An essential role for the N-terminal fragment of Toll-like receptor 9 in DNA sensing. Nat Commun 4, 1949, doi: 10.1038/ncomms2949 (2013). [DOI] [PubMed] [Google Scholar]
- 14.Majer O, Liu B, Kreuk LSM, Krogan N & Barton GM Unc93b1 recruits Syntenin-1 to dampen TLR7 signaling and prevent autoimmunity. Accepted at Nature. [DOI] [PMC free article] [PubMed]
- 15.Christensen SR et al. Toll-like receptor 7 and TLR9 dictate autoantibody specificity and have opposing inflammatory and regulatory roles in a murine model of lupus. Immunity 25, 417–428, doi: 10.1016/j.immuni.2006.07.013 (2006). [DOI] [PubMed] [Google Scholar]
- 16.Nickerson KM et al. TLR9 regulates TLR7- and MyD88-dependent autoantibody production and disease in a murine model of lupus. J Immunol 184, 1840–1848, doi: 10.4049/jimmunol.0902592 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Chan MP et al. DNase II-dependent DNA digestion is required for DNA sensing by TLR9. Nat Commun 6, 5853, doi: 10.1038/ncomms6853 (2015). [DOI] [PubMed] [Google Scholar]
- 18.Ewald SE et al. The ectodomain of Toll-like receptor 9 is cleaved to generate a functional receptor. Nature 456, 658–662, doi: 10.1038/nature07405 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Park B et al. Proteolytic cleavage in an endolysosomal compartment is required for activation of Toll-like receptor 9. Nat Immunol 9, 1407–1414, doi: 10.1038/ni.1669 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ewald SE et al. Nucleic acid recognition by Toll-like receptors is coupled to stepwise processing by cathepsins and asparagine endopeptidase. J Exp Med 208, 643–651, doi: 10.1084/jem.20100682 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Roberts AW et al. Tissue-Resident Macrophages Are Locally Programmed for Silent Clearance of Apoptotic Cells. Immunity 47, 913–927 e916, doi: 10.1016/j.immuni.2017.10.006 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shibata T et al. Guanosine and its modified derivatives are endogenous ligands for TLR7. Int Immunol 28, 211–222, doi: 10.1093/intimm/dxv062 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tanji H et al. Toll-like receptor 8 senses degradation products of single-stranded RNA. Nat Struct Mol Biol 22, 109–115, doi: 10.1038/nsmb.2943 (2015). [DOI] [PubMed] [Google Scholar]
- 24.Zhang Z et al. Structural Analysis Reveals that Toll-like Receptor 7 Is a Dual Receptor for Guanosine and Single-Stranded RNA. Immunity 45, 737–748, doi: 10.1016/j.immuni.2016.09.011 (2016). [DOI] [PubMed] [Google Scholar]
- 25.Mali P et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826, doi: 10.1126/science.1232033 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Schindelin J et al. Fiji: an open-source platform for biological-image analysis. Nat Methods 9, 676–682, doi: 10.1038/nmeth.2019 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Carpenter AE et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol 7, R100, doi: 10.1186/gb-2006-7-10-r100 (2006). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.