Abstract
We developed a loop-mediated isothermal amplification (LAMP)–fluorescent loop primer (FLP) assay for genotyping the A/G2254 single nucleotide polymorphism (SNP) in the viral DNA polymerase gene of species Equid alphaherpesvirus 1 (EHV-1), which is associated with the neuropathogenic potential of this virus. In addition to the use of regular LAMP primers to amplify the target region, a 5’-FAM–labeled backward loop primer (FLB) and 3’-dabcyl–labeled quencher probe (QP) were designed for annealing curve analysis of the amplification product. The QP, which contacts the FLB, is located at the SNP site and has the A2254 allele. LAMP reactions were performed at 63°C for 40 min, and the subsequent annealing curve analyses were accomplished within 20 min. The LAMP-FLP assay could clearly differentiate A2254 and G2254 genotypes according to the difference in the annealing temperature of the QP between the 2 genotypes. Good agreement between the LAMP-FLP and the real-time PCR for genotyping of this SNP was observed in the detection of EHV-1 in equine clinical samples. The newly developed assay is a simple and rapid method for detecting and differentiating EHV-1 strains with A2254 and G2254 polymorphisms and would be suitable for clinical use.
Keywords: equid herpesvirus 1, genotyping techniques, polymorphism, single nucleotide
Species Equid alphaherpesvirus 1 (EHV-1) is a member of genus Varicellovirus of the subfamily Alphaherpesvirinae (family Herpesviridae, order Herpesvirales). EHV-1 is present in horse populations worldwide and causes economically significant diseases including respiratory disease in young horses, abortion in pregnant mares, death in neonatal foals, and myeloencephalopathy (equine herpesvirus myeloencephalopathy; EHM).2,11 Several host factors (e.g., breed, sex, age, and immunologic status) are considered to be associated with the development of EHM.1,5 On the other hand, a single non-synonymous nucleotide substitution of guanine (G) for adenine (A) at position 2254 in the viral DNA polymerase gene (encoded by open reading frame [ORF]30) of EHV-1 has been significantly associated with neuropathogenic potential in strains of this virus,16 although other possible single amino acid substitutions have been identified in the same gene (e.g., tyrosine to serine at position 753, and histidine to arginine at position 250).4,18 An increase in EHM outbreaks among horses in the United States and Europe, caused by G2254 strains, is of major concern to the equine veterinary community because affected animals often have a poor prognosis.11 In the face of an EHV-1 outbreak, early diagnosis is crucial to the successful containment of virus spread, and differentiation between the 2 genotypes helps in planning quarantine protocols. However, current genotyping of the A/G2254 single nucleotide polymorphism (SNP) in ORF30 of EHV-1 relies on sequencing or real-time PCR (rtPCR) methods,17 which are not normally available for on-site use.
Loop-mediated isothermal amplification (LAMP) is a simple and rapid nucleic acid amplification method that has several advantages over conventional PCR methods.15 Because the reaction is carried out at a constant temperature, there is no need for a thermocycler, and the high amplification rate of LAMP provides rapid detection of the target gene. In addition, nucleic acid purification from samples is not necessarily required for the reaction, given that the LAMP method is less affected by inhibitors contained in clinical materials.6 Therefore, LAMP-based assays have been widely used for field detection of various pathogens.12 The LAMP–fluorescent loop primer (FLP) method, which was invented by Nippon Gene (Tokyo, Japan), is a SNP genotyping method based on LAMP with an FLP.7,8,14 A SNP is genotyped by annealing curve analysis of fluorescent-labeled amplification products of the target gene. Its entire process can be implemented on a portable fluorescence measuring device (LF-8 plus; Nippon Gene) for analysis of isothermal nucleic acid amplification products such as Genie II and III (OptiGene, Horsham, West Sussex, UK). Given that these devices have been developed for point-of-care testing, they are lightweight, small, and user-friendly.
The general LAMP method uses 4 types of basic primers (forward and backward outer primers [F3 and B3], forward and backward inner primers [FIP and BIP]) and 2 additional primers called loop primer F and B, which accelerate the reaction.13,15 In the LAMP-FLP method, the 5’-end of the loop primer F or B is labeled with a fluorescent molecule. In addition, a quencher probe (QP), which is an oligonucleotide labeled with a quenching molecule at the 3’-end, is also used in the method. The position of the QP is determined in order to include the target SNP and contact the 5’-end of the fluorescent-labeled loop primer. Annealing of the probe to its target sequence results in quenching of fluorescence, and the annealing temperature between the template and probe changes depending on which allele is present at the locus. The annealing temperature in the case of the mismatch is lower than that in the case of the full match. In the LAMP-FLP method, SNP genotyping is achieved by identification of the annealing temperature of the QP, which is specific for each genotype. We applied the LAMP-FLP method to the assay for genotyping of A/G2254 SNP in ORF30 of EHV-1.
The specificity of the assay was determined by 8 Japanese strains of EHV-1, which consisted of 4 A2254 strains and 4 G2254 strains (Table 1). In addition, the other 6 common equine viruses, including equid alphaherpesviruses 3 and 4 (EHV-3, -4), equid gammaherpesviruses 2 and 5 (EHV-2, -5), alphaarterivirus equid (formerly equine arteritis virus; EAV), and influenza A virus (equine influenza virus of the H3N8 subtype; EIV), were tested (Table 1).
Table 1.
Viruses used and results of the LAMP-FLP assay for equid alphaherpesvirus 1.
| Species | Strain | Clinical sign | LAMP-FLP |
|---|---|---|---|
| Equid alphaherpesvirus 1 | 89C105 (A2254) | EHM | + |
| 90C16 (A2254) | Respiratory disease | + | |
| 00C19 (A2254) | Respiratory disease | + | |
| 06-I-68 (A2254) | Abortion | + | |
| 93C2 (G2254) | EHM | + | |
| 01C1 (G2254) | EHM | + | |
| 06-I-80 (G2254) | Abortion | + | |
| 14-I-323 (G2254) | Abortion | + | |
| Equid alphaherpesvirus 3 | ECE-V-T2-FCS 2/IX’75* | NIA | − |
| Equid alphaherpesvirus 4 | TH20p* | Respiratory disease | − |
| Equid gammaherpesvirus 2 | 08C3 | Subclinical infection | − |
| Equid gammaherpesvirus 5 | 11-I-790 | EMPF | − |
| Alphaarterivirus equid | Modified Bucyrus† | Vaccine strain | − |
| Influenza A virus | A/equine/Ibaraki/1/07 | Respiratory disease | − |
Genotypes of ORF30 of equid alphaherpesvirus 1 are indicated in parentheses. EHM = equine herpesvirus myeloencephalopathy; EMPF = equine multinodular pulmonary fibrosis; LAMP-FLP = loop-mediated isothermal amplification assay using a fluorescent loop primer; NIA = no information available; + = positive; − = negative.
Kindly provided by the National Institute of Animal Health, Ibaraki, Japan.
Kindly provided by the Animal Quarantine Service, Ministry of Agriculture, Forestry and Fisheries, Yokohama, Japan.
Nasal swabs were collected from 89 febrile (≥38.5°C) Thoroughbred racehorses reared at the Miho Training Center of the Japan Racing Association from 2017 to 2018, none of which had neurologic signs. Nasal samples were obtained using 1 × 1.5 cm absorbent cotton swabs, which were immersed in 2.5 mL of transport medium (PBS supplemented with 0.06% [w/v] tryptose phosphate broth, 500 units/mL penicillin, 500 µg/mL streptomycin, and 1.25 µg/mL amphotericin B). The samples were stored at −80°C until use.
Viral DNA was extracted from 200 μL of culture supernatants of the DNA viruses and 400 μL of nasal swabs (QIAamp DNA blood mini kit; Qiagen, Hilden, Germany). Viral RNA was extracted from 140 μL of culture supernatants of the RNA viruses (QIAamp viral RNA mini kit; Qiagen). All extracted nucleic acids were stored at −80°C until use.
Real-time PCR (rtPCR) was performed with the primers and probes (Table 2) specific for the detection and A/G2254 genotyping of EHV-1.17 The reaction mixture contained QuantiTect multiplex PCR master mix (Qiagen), 0.4 μM of primers and 0.2 μM of probes, and 4 μL of template DNA, with a total volume of 20 μL. Using the StepOnePlus real-time PCR system (Applied Biosystems, Foster City, CA), the rtPCR condition was as follows: initial denaturation at 95°C for 15 min, followed by 40 cycles at 95°C for 15 s, and 58°C for 1 min. If the amplification plot of a given sample did not cross the threshold by cycle 39, the sample was considered negative. All samples were run in triplicate, and if ≥2 of 3 replicates of the samples showed a positive reaction, it was considered positive for virus detection.
Table 2.
Equid alphaherpesvirus 1 ORF30-specific primer and probe sets used in LAMP-FLP and real-time PCR assays.
| Assay/Primer or probe | Location (nt)* | Sequence (5’–3’) |
|---|---|---|
| LAMP-FLP | ||
| F3 | 2114–2132 | CCGTATCCGGCTTTCATGT |
| B3 | 2334–2317 | CAAGATGCCAAGCAGGCT |
| FIP (F1c-F2) | F1c: 2213–2193 | GCCAGGGTGGTGAAACAGAGG GGACCCCGTGGTTGTGT |
| F2: 2133–2149 | ||
| BIP (B1c-B2) | B1c: 2216–2237 | TCGATGAAGTGGATCTGGCCGG GCGAATATGGGCGTGGACAA |
| B2: 2313–2294 | ||
| LF | 2172–2150 | TGGGTATAAGCTAGCGAAGTCAA |
| FLB | 2263–2281 | FAM-ACGTTCGAGGTGGGTGACC |
| QP | 2244–2262 | ACCATCCGTCaACTACTCG-Dabcyl |
| Real-time PCR† | ||
| E1Fwd | 2229–2245 | TCTGGCCGGGCTTCAAC |
| E1Rev | 2284–2266 | TTTGGTCACCCACCTCGAA |
| E1PrA2254 | 2247–2262 | HEX-ATCCGTCaACTACTCG-BHQ2 |
| E1PrG2254 | 2247–2262 | FAM-ATCCGTCgACTACTCG-BHQ1 |
Single nucleotide polymorphism at position 2254 in ORF30 of equid alphaherpesvirus 1 is indicated with a lowercase letter. FLB = fluorescent backward loop primer; LAMP-FLP = loop-mediated isothermal amplification assay using a fluorescent loop primer; LF = forward loop primer; nt = nucleotide; ORF30 = open reading frame 30; QP = quencher probe.
Nucleotides were numbered according to GenBank accession KC924819.
Reference 17.
The primers and probe for the LAMP-FLP assay were generated by a design and synthesis service (Nippon Gene; Table 2). The QP has an adenine at position 2254; therefore, the probe matches the sequence of the A2254 genotype. The reaction was conducted with a DNA amplification kit for LAMP-FLP (Nippon Gene) according to the manufacturer’s instructions. Briefly, 25 μL of reaction mixture contained 2.5 μL of 10× reaction buffer, 1.4 mM each of 4 deoxyribonucleotide triphosphate, 0.2 μM each of 2 outer primers (F3 and B3), 1.6 μM each of 2 inner primers (FIP and BIP), 0.8 μM of the forward loop primer (LF), 0.2 μM of fluorescent backward loop primer (FLB), 0.5 μM of the QP, 8 units of Bst DNA polymerase (Nippon Gene), and 4 μL of template DNA. The entire process of the LAMP-FLP assay including isothermal amplification and annealing curve analysis was performed using LF-8 Plus (Nippon Gene). The reaction mixture was incubated at 63°C for 40 min for isothermal amplification. The temperature was then elevated and maintained at 98°C for 5 min to denature the double-stranded DNA into a single strand. In the last step, the change of fluorescence intensity was measured by lowering the temperature from 98°C to 40°C at the rate of 0.05°C/s, which took ~20 min. The fluorescence data were analyzed by the LF-8 Analyzer software (Nippon Gene) with optimized settings (measure period of 4 s; smoothing count of 80). The fluorescence intensity changing rate was calculated by primary differentiation of a curve obtained by plotting the fluorescence intensity against the temperature. The primary differentiation values formed a peak at the annealing temperature of the QP, which was automatically detected by the LF-8 Analyzer software. The threshold of an absolute value of the peak was set to 2000 to exclude the small peaks generated by the fluorescent changes unrelated to probe annealing.
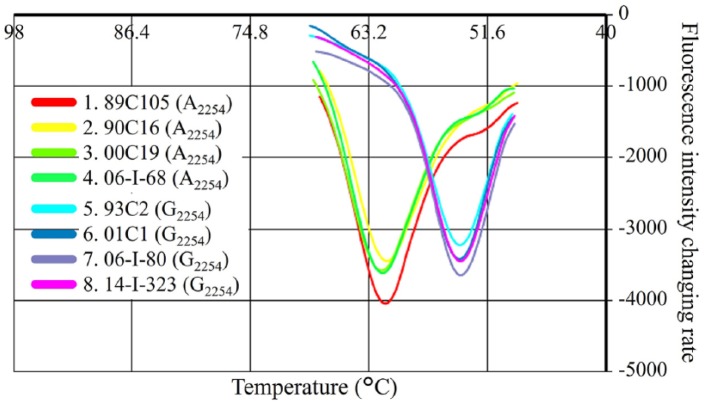
The LAMP-FLP assay could detect all tested EHV-1 strains (Table 1, Fig. 1). The mean and standard deviation of annealing temperatures of the A2254 and G2254 genotypes were 61.1 ± 0.2°C (range: 60.9–61.5°C) and 53.8 ± 0.1°C (53.7–53.9°C), respectively. The other 6 viruses, including EHV-2, -3, -4, -5, EIV and EAV, were not detected by the assay.
Figure 1.
The results of annealing curve analysis of the loop-mediated isothermal amplification assay using a fluorescent loop primer (LAMP-FLP) of equid alphaherpesvirus 1 strains. The A2254 and G2254 genotypes were clearly differentiated according to their annealing temperature.
The analytical sensitivity of the LAMP-FLP assay for each genotype was determined and compared with the rtPCR assay using 2-fold serial dilutions of a plasmid DNA containing each target sequence. Each 2-fold dilution of plasmid DNA was amplified in 16 replicates; the limit of detection with 95% probability (LOD95%) was calculated by Logit analysis (JMP software v.13; SAS Institute, Cary, NC). The LOD95% of the LAMP-FLP assays for the A2254 and G2254 genotypes were 20.3 and 20.3 copies per reaction, respectively (Table 3). The LOD95% of the rtPCR assay for the A2254 and G2254 genotypes were 2.5 and 7.8 copies per reaction, respectively.
Table 3.
Analytical sensitivity comparison between LAMP-FLP and real-time PCR assay using 2-fold serial dilutions of a plasmid DNA containing each target sequence.
| Copy number of plasmid DNA/reaction | No. of positive replicates | |||
|---|---|---|---|---|
| LAMP-FLP |
Real-time PCR |
|||
| A2254 | G2254 | A2254 | G2254 | |
| 40 | 16 (100) | 16 (100) | NT | NT |
| 20 | 15 (94) | 15 (94) | NT | NT |
| 10 | 13 (81) | 13 (81) | NT | NT |
| 5 | 7 (44) | 9 (56) | 16 (100) | 16 (100) |
| 2.5 | 3 (19) | 4 (25) | 15 (94) | 12 (75) |
| 1.25 | NT | NT | 12 (75) | 8 (50) |
| 0.625 | NT | NT | 9 (56) | 6 (38) |
| 0.3125 | NT | NT | 2 (13) | 4 (25) |
| LOD95% [copies/reaction] | 20.3 | 20.3 | 2.5 | 7.8 |
| 95% CI | 12.9–57.3 | 12.4–68.1 | 1.6–7.2 | 3.8–48.4 |
CI = confidence intervals; LAMP-FLP = loop-mediated isothermal amplification assay using a fluorescent loop primer; LOD95% = limit of detection with 95% probability; NT = not tested. Numbers in parentheses are % of positives per 16 replicates.
The detection of EHV-1 in clinical samples was compared between the LAMP-FLP and the rtPCR assay. Agreement between the 2 assays was evaluated by the kappa value, which was calculated using JMP software v.13 (SAS Institute). The rtPCR detected 14 positive samples, 12 of which were also positive by the LAMP-FLP assay (Table 4). The genotyping results of both assays were coincident with each other and indicated that all positive samples contained the A2254 strain (range of the annealing temperature in the LAMP-FLP assay: 60.5–61.6°C). All 75 of the rtPCR-negative samples were also negative by the LAMP-FLP assay. The positive and negative agreement values for the LAMP-FLP assay compared to rtPCR were 85.7% (95% confidence intervals [CI]; 57.2–98.2%) and 100% (95.2–100%), respectively. The kappa value was 0.91 (0.79–1), which indicates almost perfect agreement.9
Table 4.
Comparison of LAMP-FLP and real-time PCR detection of equid alphaherpesvirus 1 in clinical samples.
| LAMP-FLP | Real-time PCR | ||
|---|---|---|---|
| Positive | Negative | Total | |
| Positive | 12 | 0 | 12 |
| Negative | 2 | 75 | 77 |
| Total | 14 | 75 | 89 |
All positive samples contained the A2254 genotype strain of equid alphaherpesvirus 1. LAMP-FLP = loop-mediated isothermal amplification assay using a fluorescent loop primer.
We successfully developed the LAMP-FLP assay for genotyping of A/G2254 SNP in ORF30 of EHV-1. Several SNP genotyping methods based on LAMP have been developed for the detection and differentiation of animal or plant pathogens.3,10 One method uses inner primers possessing a specific 5’-end nucleotide that matches with the corresponding allele, which inhibit amplification of the template DNA possessing the mismatched allele.10 The method can be performed without a special device for a fluorescent analysis. However, it is difficult to design primers that effectively suppress amplification of the mismatched allele.19 In addition, the method needs to set up multiple reactions using primer sets specific for each allele to detect all genotypes. The guanine quenching probe (QProbe; Nippon Steel and Sumikin Eco-Tech, Tokyo, Japan), which is an oligonucleotide 3’- or 5’-end labeled with a fluorescent molecule quenched by contact with guanine, has been applied to the LAMP-based SNP genotyping method.3 Although this method uses the same theory as LAMP-FLP, the design of the probe has the limitation that the fluorescent-labeled end of the probe must be located adjacent to a guanine residue on the target sequence. The LAMP-FLP method only needs a single reaction for genotyping, and its probe can be designed more flexibly than the QProbe.
Our LAMP-FLP assay can be accomplished within 70 min and could clearly differentiate the A2254 and G2254 genotypes according to the major difference in the annealing temperature of the QP between the 2 genotypes. These features are suitable for point-of-need detection, given that rapid turnaround time and output of clear results are important for detection tests at a clinical site. The LAMP-FLP assay was less sensitive than the rtPCR assay, which is one of the most sensitive methods for the detection and discrimination of A/G2254 SNP in ORF30 of EHV-1. The sensitivity of the LAMP-FLP assay would be, however, sufficient for detection purposes because good agreement was observed between the 2 assays for EHV-1 detection in equine clinical samples. Therefore, we consider our LAMP-FLP assay to be a good candidate for an on-site test of EHV-1 infection. In future studies, we will evaluate the suitability of several simple DNA extraction methods for the LAMP-FLP assay.
Acknowledgments
We thank Akira Kokubun, Akiko Kasagawa, Akiko Suganuma, and Kaoru Makabe for their invaluable technical assistance.
Footnotes
Declaration of conflicting interests: The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding: The authors received no financial support for the research, authorship, and/or publication of this article.
References
- 1. Allen GP. Risk factors for development of neurologic disease after experimental exposure to equine herpesvirus-1 in horses. Am J Vet Res 2008;69:1595–1600. [DOI] [PubMed] [Google Scholar]
- 2. Allen GP, et al. Equid herpesvirus 1 and equid herpesvirus 4 infections. In: Coetzer J AW, et al., eds. Infectious Diseases of Livestock. Cape Town: Oxford University Press, 2004:829–859. [Google Scholar]
- 3. Ayukawa Y, et al. Novel loop-mediated isothermal amplification (LAMP) assay with a universal QProbe can detect SNPs determining races in plant pathogenic fungi. Sci Rep 2017;7:4253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Bryant NA, et al. Genetic diversity of equine herpesvirus 1 isolated from neurological, abortigenic and respiratory disease outbreaks. Transbound Emerg Dis 2018;65:817–832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Goehring LS, et al. Equine herpesvirus type 1-associated myeloencephalopathy in The Netherlands: a four-year retrospective study (1999–2003). J Vet Intern Med 2006;20:601–607. [DOI] [PubMed] [Google Scholar]
- 6. Kaneko H, et al. Tolerance of loop-mediated isothermal amplification to a culture medium and biological substances. J Biochem Biophys Methods 2007;70:499–501. [DOI] [PubMed] [Google Scholar]
- 7. Kitani M, inventor; Nippon Gene Co., Ltd., assignee. Primer and probe set used in identifying gene polymorphism, and use thereof. Japan patent WO2014/157377. 2014 Oct 2. [Google Scholar]
- 8. Komura R, et al. Simultaneous detection of benzimidazole-resistant strains of Fusarium head blight using the loop-mediated isothermal amplification-fluorescent loop primer method. J Gen Plant Pathol 2018;84:247–253. [Google Scholar]
- 9. Landis JR, Koch GG. The measurement of observer agreement for categorical data. Biometrics 1977;33:159–174. [PubMed] [Google Scholar]
- 10. Lin P, et al. Loop-mediated isothermal amplification-single nucleotide polymorphism analysis for detection and differentiation of wild-type and vaccine strains of mink enteritis virus. Sci Rep 2018;8:8393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Lunn DP, et al. Equine herpesvirus-1 consensus statement. J Vet Intern Med 2009;23:450–461. [DOI] [PubMed] [Google Scholar]
- 12. Mori Y, et al. Loop-mediated isothermal amplification (LAMP): recent progress in research and development. J Infect Chemother 2013;19:404–411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Nagamine K, et al. Accelerated reaction by loop-mediated isothermal amplification using loop primers. Mol Cell Probes 2002;16:223–229. [DOI] [PubMed] [Google Scholar]
- 14. Nishina K, et al. Polymorphism of the oxytocin receptor gene modulates behavioral and attitudinal trust among men but not women. PLoS One 2015;10:e0137089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Notomi T, et al. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res 2000;28:E63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Nugent J, et al. Analysis of equid herpesvirus 1 strain variation reveals a point mutation of the DNA polymerase strongly associated with neuropathogenic versus nonneuropathogenic disease outbreaks. J Virol 2006;80:4047–4060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Smith KL, et al. New real-time PCR assay using allelic discrimination for detection and differentiation of equine herpesvirus-1 strains with A2254 and G2254 polymorphisms. J Clin Microbiol 2012;50:1981–1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Smith KL, et al. The increased prevalence of neuropathogenic strains of EHV-1 in equine abortions. Vet Microbiol 2010;141:5–11. [DOI] [PubMed] [Google Scholar]
- 19. Xie S, et al. Development of an electrochemical method for ochratoxin A detection based on aptamer and loop-mediated isothermal amplification. Biosens Bioelectron 2014;55:324–329. [DOI] [PubMed] [Google Scholar]