Abstract
Cotranslational protein targeting is a conserved process for membrane protein biogenesis. In Escherichia coli, the essential ATPase SecA was found to cotranslationally target a subset of nascent membrane proteins to the SecYEG translocase at the plasma membrane. The molecular mechanism of this pathway remains unclear. Here we use biochemical and cryoelectron microscopy analyses to show that the N-terminal amphipathic helix of SecA and the ribosomal protein uL23 form a composite binding site for the transmembrane domain (TMD) on the nascent protein. This binding mode further enables recognition of charged residues flanking the nascent TMD and thus explains the specificity of SecA recognition. Finally, we show that membrane-embedded SecYEG promotes handover of the translating ribosome from SecA to the translocase via a concerted mechanism. Our work provides a molecular description of the SecA-mediated cotranslational targeting pathway and demonstrates an unprecedented role of the ribosome in shielding nascent TMDs.
Introduction
Membrane protein biogenesis is crucial for cell viability, due to the abundance (~30% of proteome) of membrane proteins and their participation in numerous essential cellular functions such as energy generation, molecular transport, and cell-cell communication1–3. The localization, insertion and folding of transmembrane domains (TMDs) are energetically costly4 and kinetically demanding5. To overcome these challenges, a strategy widely used by cells is to recruit molecular chaperones when the TMD on a nascent polypeptide emerges from the ribosomal exit tunnel. These chaperones protect the nascent TMD from aggregation and also act as or in collaboration with dedicated targeting machinery to cotranslationally deliver the nascent membrane protein to the translocation machinery on the target membrane6.
Diverse membrane protein targeting pathways have been discovered. The most well-studied route is mediated by the signal recognition particle (SRP)3,7, which is cotranslationally recruited to ribosomes8,9 and shields TMDs or hydrophobic signal sequences adjacent to the N-terminus of the nascent protein10. The interaction of SRP with the SRP receptor delivers the ribosome•nascent chain complex (RNC) to the Sec61p translocase at the eukaryotic endoplasmic reticulum (ER), or the SecYEG translocase at the bacterial plasma membrane10,11. In eukaryotic cells, the SRP-independent targeting (SND) components help in the delivery and insertion of a subclass of membrane proteins harboring internal TMDs to the ER, possibly before the nascent protein finishes its synthesis12,13. The ER membrane protein complex (EMC) could insert a subset of nascent TMDs into the membrane both co- and post-translationally14–16. The diversity of membrane protein targeting and translocation machineries are suggested to accommodate different properties of membrane proteins, such as the hydrophobicity13,15,16, location12, and topology of the TMDs13,14,16.
SecA is another emerging bacterial protein biogenesis factor that can mediate the cotranslational targeting and translocation of some of the membrane proteins17–19. SecA binds to the ribosome near uL23 in proximity to the exit tunnel17,19 and could be recruited to many membrane proteins during translation20. The most well-characterized membrane protein substrate for cotranslational delivery by SecA is RodZ18,21, a single pass type II membrane protein essential for cell division. SecA is necessary and sufficient for the targeting of RodZ to the SecYEG translocon in a strictly cotranslational mechanism in vitro and vivo18. SecA binds to RNCs bearing the RodZ nascent chain with high affinity (Kd ≤ 1 nM), and this binding survives the competition from other ribosome-associated protein biogenesis factors such as SRP and trigger factor (TF). The RodZ TMD is flanked by basic residues at the N-terminus and acidic residues at the C-terminus (net charge of −4), both of which are important for high affinity binding of SecA in preference over SRP18. However, little is known about how SecA protects hydrophobic TMDs emerging from the ribosome exit tunnel, nor the molecular basis of its charge preferences during this recognition.
SecA was known to be an essential ATPase that drives the post-translational translocation of secretory proteins harboring less hydrophobic signal sequences across SecYEG22, often in collaboration with the chaperone SecB. In this post-translational mode, SecA binds the signal sequence via a hydrophobic groove in the pre-protein crosslinking (PPXD) domain23,24. Another surface on SecA, Patch A, provides additional contact sites for hydrophobic segments in the mature regions of secretory proteins25. It is unclear whether SecA uses the same preprotein binding sites to recognize nascent TMDs emerging from the ribosome in its recently described cotranslational mode of targeting. Furthermore, SecA binds with high affinity to anionic phospholipids and to SecYEG26,27, and previous structures28,29 suggested that SecA and the ribosome share partially overlapping binding sites on SecYEG. Biochemical data also indicate that SecA and the 70S ribosome compete for binding to SecYEG30. How the cotranslational recognition by SecA leads to the efficient delivery of nascent membrane proteins to SecYEG18,21 remains an outstanding puzzle.
To address these questions, we combined biochemical and structural analyses to study the molecular mechanism of this pathway. Site-specific crosslinking showed that the ribosome induces a distinct mode of nascent protein recognition by SecA. A cryoEM structure of SecA bound to RNCRodZ showed that the nascent TMD is sandwiched in a composite binding site formed by the N-terminal amphipathic helix of SecA and a hydrophobic groove on uL23 of the ribosome, and revealed the molecular basis for the charge preference during nascent protein recognition by SecA. Finally, quantitative kinetic analyses demonstrate that SecYEG remodels the RNC-bound SecA to facilitate nascent protein transfer to SecYEG, and the transfer process is further facilitated by the elongation of the nascent polypeptide.
Results
The ribosome promotes nascent protein interaction with SecA N-terminal amphipathic helix
As a model cotranslational SecA substrate, we used the inner membrane protein RodZ21, which is cotranslationally targeted and translocated by SecA in vitro and in vivo18. To systematically probe how SecA interacts with the nascent RodZ TMD, we used thio-specific crosslinking with bismaleimidohexane (BMH) to test the proximity between a single cysteine (C111) one residue upstream of the RodZ TMD and individual cysteines engineered at various positions on the SecA surface. All the single cysteine variants of SecA are functional in mediating post-translational translocation of proOmpA and the cotranslational translocation of RodZ (Supplementary Fig. 1a–b). We probed the cotranslational nascent chain interactions of SecA using purified RNCs18,31 bearing residues 104–160 of the RodZ nascent chain. The RNCs are stalled using the SecM arrest peptide (SecM residues 133–170) fused to the C-terminus of the RodZ nascent chain, so that the RodZ TMD (residues 112–132) is exposed outside the ribosome exit tunnel (Supplementary Fig.1c). SecA recognizes stalled RNCRodZ with subnanomolar affinity and high specificity18; the in vitro binding data with stalled RNCs are consistent with the ability of SecA to drive the targeting and insertion of RodZ into the bacterial inner membrane in vitro and the dispensability of SRP for RodZ targeting in vivo and in vitro18. Hence, the stalled RNCs can faithfully report on nascent chain interaction with and selection by SecA under physiological conditions. The Mreb-binding domain (MBD; residues 1–103) of RodZ upstream of its TMD is not essential for SecA recruitment18 nor for the SecA-dependent membrane targeting and integration of RodZ (Supplementary Fig. 1d–f), and was therefore removed in this work.
The results of this cysteine scan revealed strong crosslinks between the RodZ nascent chain and the cysteines engineered at SecA residues 7, 10 and 12 (Fig. 1a,c,e), located in the conserved amphipathic helix32 at the N-terminus of SecA (termed “helix N1”). Weaker but detectable crosslinks were also observed with the cysteines in nucleotide binding domain-I (NBD-I; residues 34, 56, 402 and 403) and helical scaffold domain (HSD; residue 636) of SecA, all of which are within ~30 Å of helix N1 (Fig. 1a and Supplementary Fig. 1j). These crosslinks are dependent on the presence of crosslinker, RNCRodZ and SecA (Supplementary Fig. 1h). In contrast, no crosslinks were detected with the cysteines engineered in the SecA PPXD domain (residues 232, 235, and 306) or in Patch A (residue 193), which are known to bind hydrophobic segments on preproteins post-translationally23,24. The SecA clamp region (residue 369)27 and the two-helix finger loop (THF; residue 797)33 were previously reported to contact the translocating polypeptide, but also failed to crosslink to the RodZ nascent chain. SecA endogenous cysteines, 3 of which located in the C-terminal zinc-finger domain, also did not crosslink to the RodZ nascent chain (Supplementary Fig. 1g).
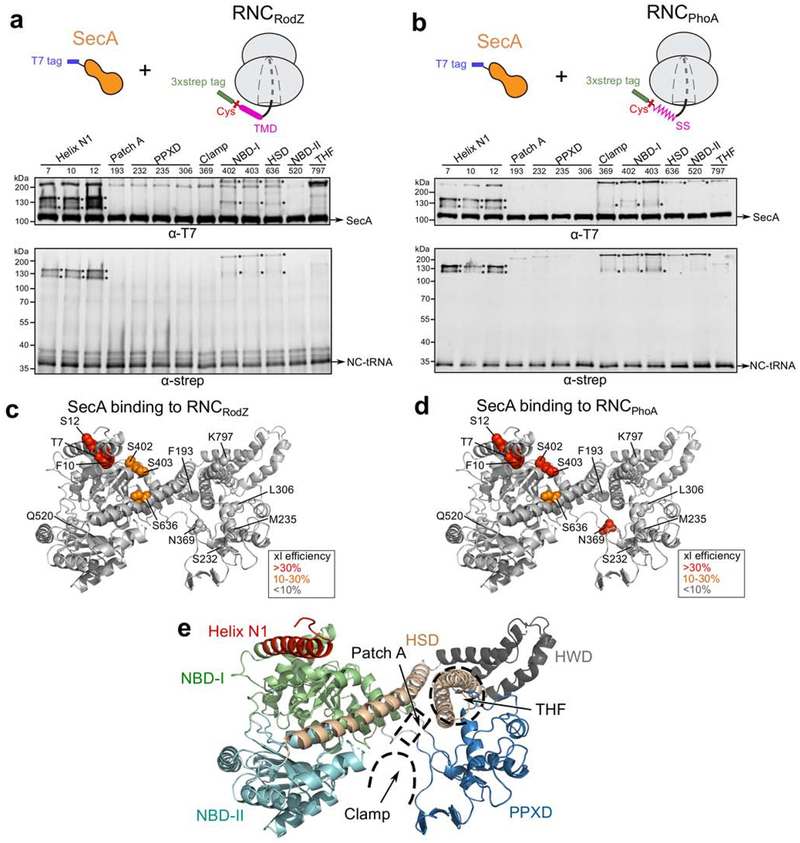
Fig. 1. Thio-specific crosslinking to map SecA surface residues that contact hydrophobic sequences of nascent proteins on the ribosome.
a-b, Engineered single cysteines at indicated positions of SecA were tested for crosslinking by BMH to the TMD of RNCRodZ (a) or the signal sequence of RNCphoA (b). SecA is T7-tagged at the flexible C-terminus. The RodZ (residues 104–160) or phoA (residues 1–52) nascent chain was fused to a C-terminal SecM stall sequence and an N-terminal 3xstrep tag, and contains a single cysteine upstream of the RodZ TMD (C111) or the phoA signal sequence (C4). Crosslinking reactions contained 1 μM SecA and 500 nM RNC. Asterisks indicate crosslinking products detected by both the anti-T7 and anti-strep antibodies. ‘NC’, nascent chain. The additional bands in the anti-T7 blots are potentially SecA dimer induced by cysteine-cysteine crosslinking (online Methods).c-d, Crosslinking efficiency from the data in panels (a) and (b) are mapped onto the structure of SecA generated in this work. Crosslinking efficiencies were calculated by dividing the amount of crosslinked nascent chain over the total amount of nascent chain, based on western-blots against the strep tag. Residues are shown in spheres and colored based on crosslinking efficiency as indicated. e, The individual domains are highlighted in SecA structure. The following color scheme was used: helix N1, red; NBD-I, green; NBD-II, cyan; PPXD, blue; HSD, wheat; HWD (helical wing domain), gray. Patch A, clamp and THF regions are highlighted by dashed lines.
The following observations corroborated these interaction patterns of SecA during its cotranslational recognition of nascent protein. First, another thio-specific crosslinker bismaleimidoethane (BMOE), which has a shorter spacer length than BMH (8.0 Å vs. 13.0 Å, respectively), generated a similar crosslinking pattern between the RodZ nascent chain and SecA, albeit with slightly lower efficiency (Supplementary Fig. 1j). Moreover, RNC exposing a less hydrophobic signal sequence (SS) from the secretory protein, PhoA, exhibited a SecA crosslinking pattern similar to that of RNCRodZ (Fig. 1b,d,e), arguing against the notion that the observed crosslinking pattern is specific to the RodZ TMD. Finally, as an orthogonal approach to detect the distance between SecA and nascent chain on the ribosome, we incorporated a fluorescent amino acid, 7-hydroxycoumaryl ethylglycin (Cm), immediately upstream of the RodZ TMD (residue 111) or the PhoA SS (residue 4) on the RNC. We monitored the Förster resonance energy transfer (FRET) between or and an acceptor dye, BODIPY-FL (BDP), incorporated at various positions on SecA (Supplementary Fig. 2a). The highest FRET efficiency was observed with BDP labeled at helix N1 of SecA, whereas BDP labeled at sites away from the N-terminus of SecA, such as PPXD and Patch A, exhibited low FRET efficiency with the Cm dye on the nascent chains (Supplementary Fig. 2a–c). Thus, helix N1 of SecA is the primary binding site for TMDs or signal sequences as the nascent polypeptide emerges from the ribosome.
These results are surprising, as previous work identified PPXD or Patch A as the sites used by SecA to interact with hydrophobic sequences on preprotein substrates. To test if the ribosome is responsible for this difference, we purified SUMO-RodZ and SUMO-PhoA fusion proteins, in which the RodZ TMD or the PhoA SS was C-terminally fused to the SUMO protein, and probed the post-translational interactions of these proteins with SecA using thio-specific crosslinking. Both substrates crosslinked efficiently to the cysteines in SecA PPXD and/or Patch A, as well as multiple sites across all the domains of SecA (Supplementary Fig. 2d–g), consistent with previous observations using PhoA as the model post-translational substrate23,25,34. Compared to the observations with RNC, SecA helix N1 was a less dominant binding site in the absence of the ribosome. These results strongly suggest that the ribosome induces a distinct mode of substrate recognition by SecA and confines nascent protein interactions to its N-terminus.
Structure of SecA bound to nascent RodZ on the ribosome
High-resolution structural information on a complex formed between the translating ribosome and SecA is still lacking. To better understand the interaction between the nascent chain and SecA during the cotranslational targeting pathway, we set out to determine the cryo-EM structure of the RNCRodZ•SecA complex. Initial efforts to obtain a stable complex for cryo-EM did not yield high resolution information, suggesting that the binding of SecA on the ribosome is flexible. To increase the stability of this complex, BMH crosslinking was used to stabilize the contact between SecA (C12) and a specific cysteine engineered upstream (C111) or downstream (C146) of the RodZ TMD (Supplementary Fig. 2h). RNCRodZ (C146), which gave the most efficient (45%) crosslink to SecA, was used for the cryo-EM studies.
Despite the high crosslinking efficiency, our initial structural analysis showed a low occupancy of SecA on the ribosome, which underscores the sensitivity of this complex under cryo-EM freezing conditions. We therefore collected a large dataset and employed an extensive focused 3D classification and refinement scheme, which resulted in a cryo-EM structure of the RNCRodZ•SecA complex at an overall resolution of 3.1 Å (Fig. 2a, Supplementary Fig. 3, and methods). The contact points between SecA and the ribosome as well as the RodZ TMD were resolved to side chain resolution (3.1 – 3.5 Å), which allowed us to build these regions de novo and to assign the registry and directionality of the RodZ TMD (Fig. 2b and Supplementary Fig. 4). The local resolution of SecA was further improved using a 3D refinement scheme that focused on the SecA region and masked out the rest of the ribosome. This strategy yielded a SecA structure at a local resolution of 5.7 Å, where secondary structural elements can be clearly resolved (Fig. 2c and Supplementary Fig. 4). This also allowed us to manually adjust the α-helices and place the PPXD of SecA, which is known from previous structural and biochemical studies to adopt multiple conformations26,35,36, as a rigid body into the EM density.
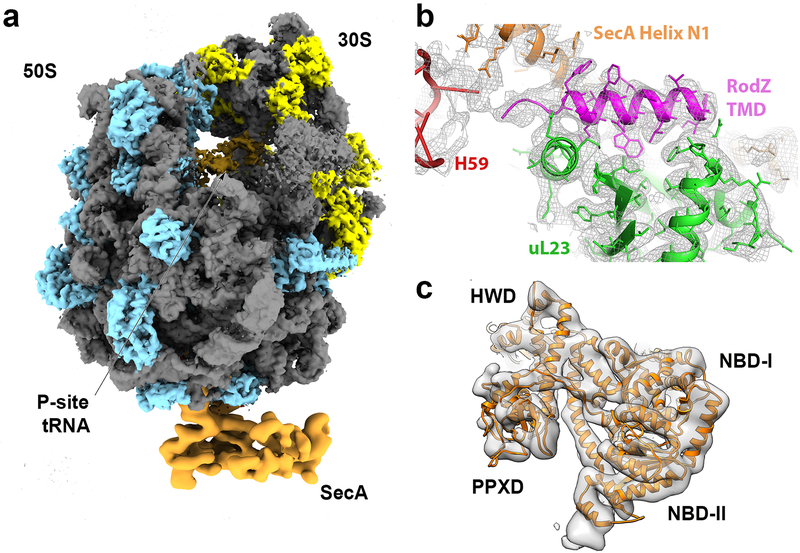
Fig. 2. Cryo-EM structure of the SecA•RNCRodZ.
a, Overview of the cryo-EM model of the SecA•RNCRodZ complex. Ribosomal RNA is in grey, ribosomal proteins in the large and small subunits are in blue and yellow, respectively, the P-site tRNA is in beige, and EM-densities of the ribosome and SecA are from the global refinement and focused refinement, respectively, and are filtered to the respective resolutions of 3.1 and 5.7 Å. b, Representative fit of the RodZ TMD and the contact points between the ribosome and SecA with the overlaid EM-density (grey mesh). The ribosomal protein uL23 and the RodZ TMD are colored in green and magenta, respectively. c, Representative fit of the SecA model with the overlaid EM-density obtained from the focused refinement approach.
The density of SecA covers the ribosome exit tunnel and lies parallel to the ribosome surface, with NBD-I most proximal to the exit tunnel, whereas PPXD and HWD points away (Fig. 3a,b). Interactions between SecA and the ribosome are mediated exclusively through contacts with regions of the 23S rRNA in the vicinity of ribosomal proteins uL23, uL24, and uL29, consistent with previous observations17,19. Specifically, rRNA H59 contacts basic amino acids (R16, R19, R20) on the positively charged face of the amphipathic helix N1 of SecA, which extends down from NBD-I towards the ribosomal exit tunnel (Fig. 3a,e, Fig. 4a and Supplementary Fig. 5a). In addition, 23S rRNA H7 contacts basic residues (R602, K609) on NBD-II of SecA (Fig. 4a, blue).
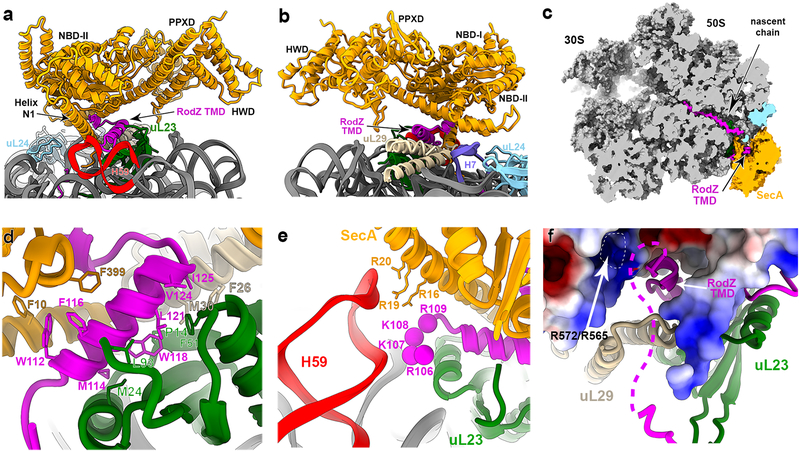
Fig. 3. Snapshots of the SecA•RNCRodZ model.
a-b, Close-ups of the contact points between SecA and the surface of the ribosome. The two views are rotated relative to each other by 180° along the vertical axis of the model. c, Surface representation of the SecA•RNCRodZ model with a cross-section of the ribosome tunnel region where the nascent chain and the TMD of RodZ can be visualized. d, Close-up view of the composite TMD binding pocket formed by uL23 and SecA. uL29 may also contribute to this binding pocket. Hydrophobic residues that contact the RodZ TMD are highlighted in sticks. e, 23S rRNA H59 contacts basic residues (spheres) preceding the RodZ TMD and basic residues (sticks) on the hydrophilic side of SecA helix N1. f, Sequence downstream of the RodZ TMD (dashed line) may be positioned in the vicinity of a basic patch on SecA, which is shown as electrostatic surface. The basic surface provided by R572 and R565 is highlighted. The following color scheme was used: uL23, green; uL24, cyan; uL29, wheat; SecA, orange; RodZ TMD, magenta; H59, red; H7, deep blue.
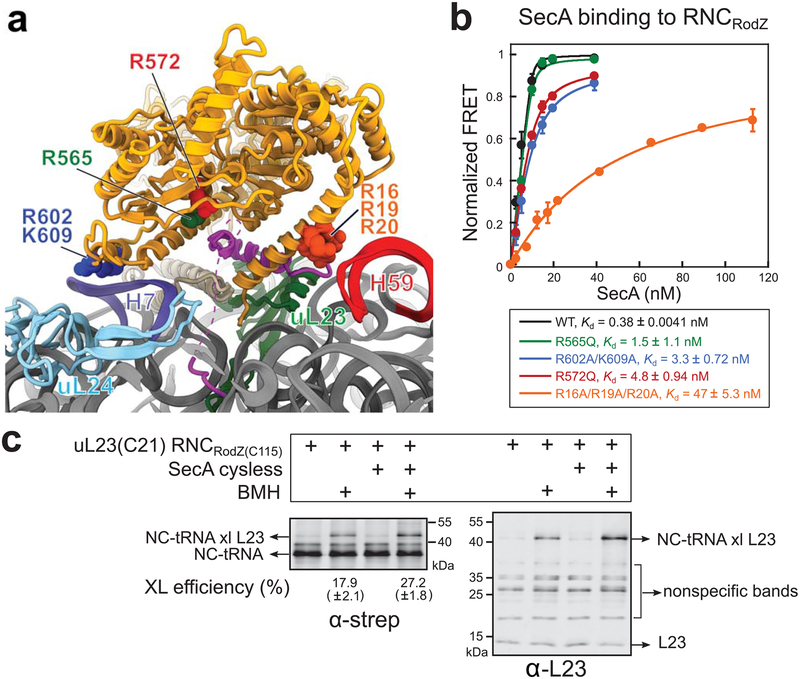
Fig. 4. Validation of the interaction sites between RNCRodZ and SecA.
a, SecA mutations tested biochemically are highlighted in spheres in the SecA•RNCRodZ model. Residues are shown in spheres and colored as in (b). The remainder of the structure is colored as in Fig. 3. b, Equilibrium titrations to measure the Kd values for the binding of RNCRodZ to wild type (WT) SecA and indicated SecA mutants. Cm was incorporated at residue 111 upstream of the RodZ TMD on RNC. All SecA variants were labeled by BDP at residue 12. Lines are fits of the data to Eq. 2 (Supplementary Note 1), and the obtained Kd values are summarized. All values are reported as mean ± s.d., with n = 2–3 independent measurements. c, Crosslinking between a pair of engineered cysteines at residue 115 in the RodZ TMD and residue 21 of uL23 in RNCRodZ. Crosslinking was induced by BMH, and the crosslinked product was detected by western blot with anti-strep and anti-L23 antibodies. The numbers underneath the α-strep blot indicate crosslinking efficiency, calculated from the ratio of the intensity of crosslinked bands relative to the total intensity of tRNA-linked nascent chain (NC-tRNA). XL, crosslinking. All values are reported as mean ± s.d., with n = 2 independent measurements.
The RodZ nascent chain is resolved within the ribosome tunnel from the CAA end of the P-site tRNA to the tunnel exit (Fig. 3c and Supplementary Fig. 5e). The RodZ TMD is bound within a composite pocket contributed by: (1) residues (P14, M24, F51, L93) from ribosomal protein uL23, (2) the hydrophobic face (L2, I3, L5–6, F10) of the amphipathic helix N1 of SecA, (3) residue F399 in NBD-I of SecA, and (4) residue F639 in HSD of SecA (Fig. 3d, Supplementary Fig. 5a–d). Residues M30 and F26 of uL29 may also contribute to part of the binding pocket. These interactions shield the nascent TMD from the aqueous cytosolic environment prior to membrane insertion. The RodZ TMD is preceded by six consecutive positively charged residues, four of which are resolved and contact the 23S rRNA at H59 (Fig. 3e) in a mode similar to previous observations with an SRP-bound signal sequence on the RNC10. This observation implicates a potential role of H59 in multiple cotranslational targeting pathways.
To test the role of the SecA-ribosome contacts observed in the structure, we measured the equilibrium dissociation constant (Kd) of RNCRodZ bound to various SecA mutants based on FRET between Cm-labeled RNCRodZ and BDP-labeled SecA described above (Supplementary Fig. 2a,b). Mutation of the basic residues (R16, R19 and R20) on SecA that contact 23S rRNA H59 reduced its binding affinity for RNCRodZ over 100-fold (Fig. 4a,b, orange), indicating the essential role of this contact in stabilizing the SecA-RNC interaction. Mutation of R602 and K609 in SecA, which contact 23S rRNA H7, had a more modest effect, ~10-fold (Fig. 4a,b, blue), suggesting this to be an ancillary ribosome contact site.
We previously showed that the hydrophobic TMD of RodZ as well as enrichment of basic and acidic residues N- and C-terminal to the TMD, respectively, are important for high affinity binding between SecA and RNCRodZ18. While the basic residues N-terminal to the RodZ TMD were resolved in the structure and contact rRNA H59, the region C-terminal to the RodZ TMD was not resolved. Nevertheless, the C-terminus of the RodZ TMD points towards a SecA surface rich in positively charged residues that could provide a contact site for the acidic sequence C-terminal to the RodZ TMD (Fig. 3f). In support of this model, conservative mutation of two Arg residues in this surface (565 or 572) to Gln each caused a 4–12 fold weakened binding of SecA to RNCRodZ (Fig. 4a,b, green and red). These results provide a structural basis to explain the charge preferences of SecA during its cotranslational nascent chain recognition. Finally, the resolved density also suggests a looped conformation of the RodZ nascent chain that positions sequences C-terminal of the RodZ-TMD close to residue 12 of SecA (Supplementary Fig. 5f), explaining the efficient crosslink of SecA(C12) to cysteines both upstream (C111) and downstream (C146) of the RodZ TMD (Supplementary Fig. 2h).
The composite TMD binding pocket formed by both uL23 and SecA observed in the structure is in good agreement with our crosslinking data that SecA residues 7, 10, 12, 402, 403 and 636 are in close proximity to the RodZ TMD on the ribosome. To test whether the hydrophobic cleft on uL23 provides a potential docking site for the nascent TMD, we deleted genomic uL23 and complemented cells with a mutant uL23 harboring an engineered single cysteine, C21, in the hydrophobic groove that contact the RodZ TMD (Fig. 3d). We purified RNCRodZ harboring a cysteine at uL23 (C21) and a cysteine in the RodZ TMD (C115). Addition of BMH induced a significant crosslink between uL23 and the RodZ nascent chain, and the presence of SecA further increased the crosslinking efficiency ~50% (Fig. 4c). These results suggest that the nascent TMD has an intrinsic preference to dock at uL23, and this interaction is further stabilized by SecA.
SecYEG and nascent chain elongation facilitate RNC handover from SecA to SecYEG
The results above provide the molecular basis for the initial recognition of RNCRodZ by SecA, which subsequently targets the RodZ nascent chain to SecYEG for membrane integration in a strictly cotranslational pathway18,21. Intriguingly, our structure suggested that the RNC-binding surface of SecA heavily overlaps with its anionic phospholipid interaction surface32 and was on the same face as its SecYEG docking site37 (Supplementary Fig. 6a). This raises questions as to how the SecA-bound RNCRodZ is delivered to SecYEG.
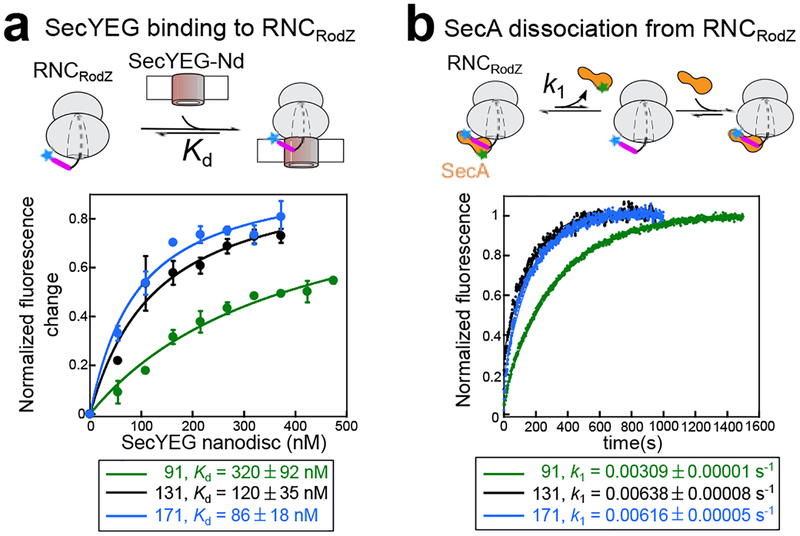
To address this question, we first asked whether the nascent chain length affects the interaction of RNCRodZ with SecYEG embedded in the phospholipid bilayer. We prepared fluorescently labeled with three nascent chain lengths: RNCRodZ91 (the same construct characterized above), RNCRodZ131, and RNCRodZ171. We reconstituted purified SecYEG complex in large lipid nanodiscs (Nd) formed by the ApoE422K scaffold protein (Supplementary Fig. 6b–e)32. Interaction with SecYEG-Nd leads to quenching of the Cm fluorescence on (Supplementary Fig. 6f, lanes 1 and 4), which was used to measure RNC-SecYEG binding. SecYEG-Nd binds RNCRodZ91 with modest affinity, and this binding affinity increased 3–4 fold with RNCRodZ131 and RNCRodZ171 (Fig. 5a). To further test if elongation of the nascent chain impacts the interaction of SecA with RNCRodZ, we measured the rate constant of SecA dissociation from RNCRodZ (k1) based on the loss of FRET from a preformed •SecABDP complex upon chase with excess unlabeled SecA (Fig. 5b and Supplementary Fig. 6f, lanes 2 and 3). SecA dissociation from RNCRodZ91 was slow, with a k1 of 0.0031 s−1, and was accelerated ~2-fold with longer nascent chain (Fig. 5b). Thus, elongation of the nascent chain modestly enhances the binding of RNC to membrane-embedded SecYEG and reduces the kinetic stability of the RNCRodZ•SecA complex, which could potentiate the nascent chain for handover from SecA to SecYEG.
Fig. 5. Effects of nascent chain length on SecYEG binding to and SecA dissociation from RNCRodZ.
a, Equilibrium titrations to measure the Kd value of the SecYEG•RNCRodZ complex. Top panel, scheme of the assay: RNCRodZ labeled with Cm (blue star) at RodZ residue 111 upstream of its TMD (magenta) was incubated with indicated concentrations of SecYEG-Nd, prepared and quantified as described in the methods. Bottom panel, representative titration curves of SecYEG binding to RNCRodZ at different nascent chain lengths. Complex formation was monitored by quenching of the Cm fluorescence by SecYEG-Nd (Supplementary Fig. 6f, lanes 1 and 4). The lines are fits of the data to Eq. 2 (Supplementary Note 1), and the obtained Kd values are summarized below. All values represent mean ± s.d., with n = 2–3 independent measurements. b, Measurement of the dissociation rate constant (k1) of SecA from RNCRodZ. Top panel, scheme of the assay: RNCRodZ labeled as in (a) was pre-incubated with SecA labeled with BDP (green star) at residue 12, and excess (300 nM) unlabeled SecA was added to chase the FRET signal (Supplementary Fig. 6f, lanes 2 and 3). Bottom panel, representative time courses of the chase reactions for RNCRodZ with indicated nascent chain lengths. The data were fit to Eq. 3 (Supplementary Note 1), and the obtained k1 values are summarized below. All values represent mean ± s.d., with n = 2–3 independent measurements.
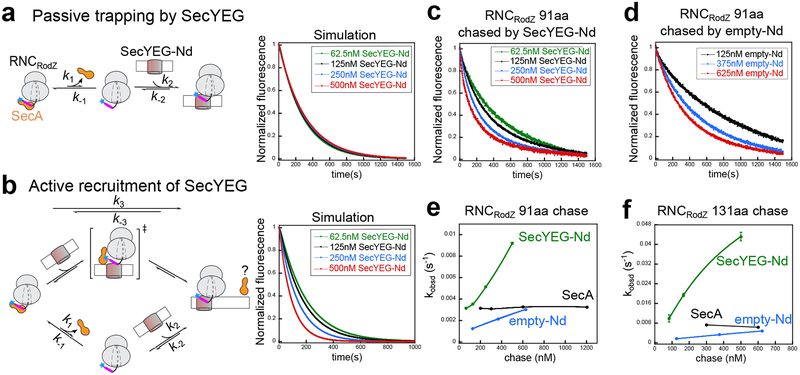
The slow SecA dissociation from RNCRodZ (t1/2 = 108–220 s) raised questions as to how RodZ targeting occurs with kinetic competence; for comparison, cotranslational targeting mediated by SRP occurs in less than 5 s3. We therefore tested alternative models for how RNC pre-bound by SecA could be delivered to SecYEG. If RNCRodZ must dissociate from SecA before it binds SecYEG (Fig. 6a, “passive” model on the left), the rate of RNCRodZ engagement with SecYEG-Nd would be limited by the slow dissociation of SecA from RNCRodZ and independent of the concentration of SecYEG-Nd (Fig. 6a, simulation results on the right and Supplementary Fig. 7c–d). In contrast, if SecYEG directly associates with SecA•RNCRodZ and alters its conformation to facilitate the handover (Fig. 6b, “active recruitment” model on the left), the transfer reaction will be accelerated by increasing concentrations of SecYEG-Nd and significantly faster than spontaneous SecA dissociation from RNCRodZ (Fig. 6b, simulations on the right and Supplementary Fig. 7e–f).
Fig. 6. Measurements to distinguish different mechanisms of RNCRodZ delivery to SecYEG by SecA.
a, Scheme (left) and kinetic simulations (right) for the model in which SecYEG passively binds RNCRodZ that has dissociated from SecA. RodZ TMD is colored in magenta. b, Scheme (left) and kinetic simulations (right) for the model in which SecYEG forms a ternary complex with SecA-bound RNCRodZ to actively displace SecA (upper route). The passive pathway was included in the simulation for completeness (lower route). Question mark denotes that it is unclear whether SecA is completely displaced after the transfer, or substantially repositioned relative to its initial mode of interaction with the RNC. c, Representative fluorescence time traces to measure the transfer of RNCRodZ at a chain length of 91aa. RNCRodZ labeled with Cm (blue star) at RodZ residue 111 was pre-incubated with unlabeled SecA, and challenged with indicated concentrations of SecYEG-Nd. Docking of RNCRodZ onto SecYEG-Nd was monitored by quenching of Cm fluorescence (Supplementary Fig. 6f, lanes 5 and 7). Each trace is the average of 6–8 measurements. The data were fit to Eq. 3 (Supplementary Note 1) to obtain the apparent rate constant (kobsd) of transfer. Note that the time traces are biphasic, and control experiment indicated that the slow phase was due to dye bleaching (Supplementary Note 1). d, Same as in (c), except that empty nanodisc was used instead of SecYEG-Nd. e-f, Summary of the observed rate constants for transfer of SecA-bound RNCRodZ to SecYEG-Nd (green) or to empty nanodiscs (blue), and for spontaneous dissociation of SecA from RNCRodZ (black) at RodZ nascent chain lengths of 91 aa (e) and 131 aa (f). Error bars are shown but may not be visible. All values represent mean ± s.d., with n = 2–3 independent measurements.
To distinguish between these models, we pre-formed a complex of with unlabeled SecA and challenged the complex with varying concentrations of SecYEG-Nd. As the fluorescence intensity of was unaffected by unlabeled SecA but was quenched when bound to SecYEG-Nd (Supplementary Fig. 6f, lanes 1, 5 and 7), transfer of to SecYEG was monitored by quenching of Cm fluorescence (Fig. 6c). At the end of the transfer reaction, quenching of the Cm fluorescence on was the same, within error, as that obtained after direct binding of to SecYEG-Nd (Supplementary Fig. 6f, lanes 4 vs. 7). In addition, the Cm fluorescence intensity at the end of transfer was similar regardless of whether was prebound to SecABDP or unlabeled SecA (Supplementary Fig. 6f, lanes 6 vs. 7), indicating that FRET between and SecABDP was lost. These observations strongly suggest that at the end of the transfer reaction, the Cm dye on was embedded in an environment dominated by SecYEG-Nd, whereas SecA was displaced from its initial position on RNCRodZ.
With RNCRodZ91, the observed transfer rate constant (kobsd) was accelerated by increasing concentrations of SecYEG-Nd (Fig. 6c,e) and became significantly faster than spontaneous SecA dissociation at SecYEG-Nd concentrations above 300 nM (Fig. 6e, green vs black), indicating that the active model became the dominant pathway for the delivery and transfer of RNCRodZ at moderate SecYEG concentrations. Control reactions using empty nanodiscs were significantly slower than those using SecYEG-Nd, indicating a role of SecYEG in the accelerated transfer (Fig. 6d,e and Supplementary Fig. 6f, lanes 5 and 8). As another negative control, spontaneous dissociation of SecA from RNCRodZ was independent of chase concentration (Fig. 6e,f, black; Supplementary Fig. 7h), as would be expected for a unimolecular reaction (Supplementary Fig. 7g). Analogous results were observed for the transfer of RNCRodZ131 to SecYEG-Nd (Fig. 6f and Supplementary Fig. 7i–k). Compared to RNCRodZ91, the transfer reactions with RNCRodZ131 were 3–4 fold faster at comparable SecYEG-Nd concentrations, whereas the reactions with empty nanodisc were similarly slow regardless of nascent chain length (Fig. 6 e vs f). As the reconstituted SecYEG-Nd contains ~0.55 copy of SecYEG per copy of nanodisc (Supplementary Fig. 6c), the observed transfer kinetics was a lower estimate of the efficiency at which SecYEG stimulates RNC transfer. Finally, consideration of the reaction equilibrium indicated that the RNCRodZ•SecYEG complexes generated during the transfer reaction were much more stable than those obtained from binding of free RNCRodZ to SecYEG (Supplementary Note 2), providing additional evidence that transfer occurred via an active mechanism that bypasses the formation of free RNCRodZ. Together, these results showed that SecYEG is recruited to SecA-bound RNCRodZ and actively promotes transfer of the nascent protein from SecA to this translocase, and this process is further facilitated by elongation of the nascent polypeptide.
Discussion
Emerging data indicate that nascent membrane proteins are cotranslationally delivered to and inserted into their membrane destinations via diverse pathways in both prokaryotic and eukaryotic organisms7,12,15. Besides the well-studied SRP pathway, little is known about the molecular mechanism of the alternative cotranslational targeting pathways. This work elucidates the molecular basis of cotranslational nascent membrane protein recognition and delivery by SecA in bacteria. Using a recently identified cotranslational SecA substrate, RodZ, we show that the hydrophobic TMD on the nascent polypeptide emerging from the ribosome is sandwiched in a composite binding site formed by both SecA and the ribosomal protein uL23, and explain the structural basis for the charge preference of SecA during its cotranslational recruitment. Furthermore, the SecYEG complex in the membrane can associate with and actively remodel the SecA-bound RNC, which together with elongation of the nascent polypeptide facilitates handover of the RNC to the membrane translocon.
The structure here provides a precedent for active participation of the ribosome exit site in forming a shared nascent TMD docking site with a ribosome-associated protein biogenesis factor (RPB). This is distinct from previous observations where the ribosome simply provides a docking site for an RPB, such as SRP or TF, which is responsible for mediating interaction with the nascent polypeptide10,38,39. The position of the nascent TMD was also distinct from that in the RNC•SRP structure (Supplementary Fig. 8). The observation of significant crosslink between the RodZ nascent chain and the hydrophobic groove of uL23 suggests that uL23 provides a transient early binding site to shield hydrophobic sequences on the nascent polypeptide before the latter engages with an RPB. This interaction could be further regulated by the RPBs, as was observed here for SecA. In addition, H59 of the 23S rRNA provides a contact site for basic residues upstream of a hydrophobic TMD (this work) or signal sequence10,11, raising the possibility that H59 acts as a hub to select for enrichment of basic residues in the targeting sequences of membrane and secretory proteins.
The biochemical and structural work here also demonstrate that the ribosome induces a distinct mode of nascent protein recognition by SecA. On SecA-bound RNC, the substrate recognition site is confined to the N-terminal region of SecA, whereas the previously identified preprotein binding sites on SecA, PPXD and Patch A, are not involved in recognition. In contrast, the same TMD or signal sequence can crosslink to PPXD, Patch A and multiple other sites on SecA in the absence of the ribosome (Fig. 1 and Supplementary Fig. 2). These observations strongly suggest that SecA can alternate between two modes of substrate recognition: the post-translational mode defined previously, and the cotranslational mode described in this work. In addition to the ribosome, multiple determinants, such as hydrophobicity of the TMD and enrichment of charges flanking the TMD, could bias the relative energetics and hence the selection of SecA’s recognition modes. The complete repertoire of nascent proteins that are cotranslationally recognized and targeted by SecA remains to be determined.
While SecA is sufficient for the cotranslational targeting and insertion of RodZ in biochemical reconstitutions18, the cotranslational interaction of SecA with the RodZ nascent chain could be further regulated by SRP. Although the in vivo translocation of RodZ showed no18 or partial21 dependence on SRP and no dependence on FtsY21, selective ribosome profiling experiments showed an SRP enrichment on the RodZ nascent chain after 154 residues9. Consistent with this, biochemical work showed that SRP binds strongly to RNCRodZ even with SecA present (Kd = 24 nM), and that SecA allosterically regulates SRP binding to RNCRodZ18. These data suggest a model in which SRP and SecA cobind on RNCRodZ. Possibly, SRP could assist SecA in the targeting of RodZ, or the two pathways could provide alternative targeting routes for RodZ.
More generally, the ribosome exit tunnel is a crowded environment where multiple RPBs in addition to SecA can dock and access the nascent polypeptide40. Although SecA can provide a docking site for both TMDs and signal sequences emerging from the ribosome, a variety of factors, such as the affinity of SecA for the RNC and regulation by other RPBs, likely dictate which substrates enter the SecA-mediated cotranslational targeting pathway in vivo. We previously showed that, although both SRP and SecA can recognize the RodZ TMD, the MreB binding domain of RodZ preceding its TMD weakens SRP binding18. Likewise, although SecA by itself binds with reasonable affinity to RNCs exposing the TMD of FtsQ, an SRP substrate, this interaction did not withstand competition from SRP. In addition, RNC bearing the nascent chain of PhoA, a post-translational SecA substrate, binds 200-fold more weakly to SecA than RNCRodZ, likely due to the lower hydrophobicity of its signal sequence and the lack of C-terminal acidic residues. These and other observations7,41,42 suggest that multiple sequence and structural elements on the nascent polypeptide, the preferential recognition of each element by individual RPBs, and competition or regulation by other RPBs together dictate the selection of nascent proteins into distinct biogenesis pathways. Reciprocally, the diversity of protein targeting factors and the dual mode of substrate recognition by SecA could accommodate the targeting needs of diverse nascent proteins with different hydrophobicity, charge distribution, and TMD location or topology.
The available structures show that SecA- and SecYEG-bound RNCs share multiple overlapping binding sites. Helix N1 of SecA, which provides an important TMD- and ribosome binding site, is also vital for its interaction with anionic phospholipids. There are also extensive overlaps in the SecA and ribosome docking sites on SecYEG as well as the SecA and SecYEG binding sites on the ribosome (Supplementary Fig. 8). Given these overlaps, it is puzzling how SecA-bound RNCs are targeted to the SecYEG translocon. Our data here provide kinetic evidence that SecYEG can directly associate with SecA-bound RNCs to generate a transient ternary intermediate in which the SecA-RNC interaction is weakened, allowing facilitated transfer of the RNC to SecYEG in a concerted pathway. Such a mechanism may be enabled by the multiple interaction sites of SecA on the ribosome. We showed here that basic residues near the N-terminus (R16/R19/R20) and in the HSD (R602/K609) of SecA both contribute to ribosome binding; a previous work suggested that additional basic residues in the SecA HSD (K625/R633) also provide a ribosome contact site17. The multi-dentate, electrostatically driven interaction could allow SecYEG to ‘invade’ part of the SecA-ribosome interaction surface without waiting for complete SecA dissociation, thus generating an accelerated path for RNC handover. These observations resonate with those during SRP-dependent cotranslational protein targeting, in which RNCs pre-bound to SRP and SR are transferred to SecYEG in a concerted mechanism involving a major rearrangement of the SRP•SR complex on the ribosome10,43. A concerted cargo handover mechanism minimizes the loss of RNC while allowing a kinetically more facile path for the transfer, and could be envisioned for other membrane protein biogenesis pathways.
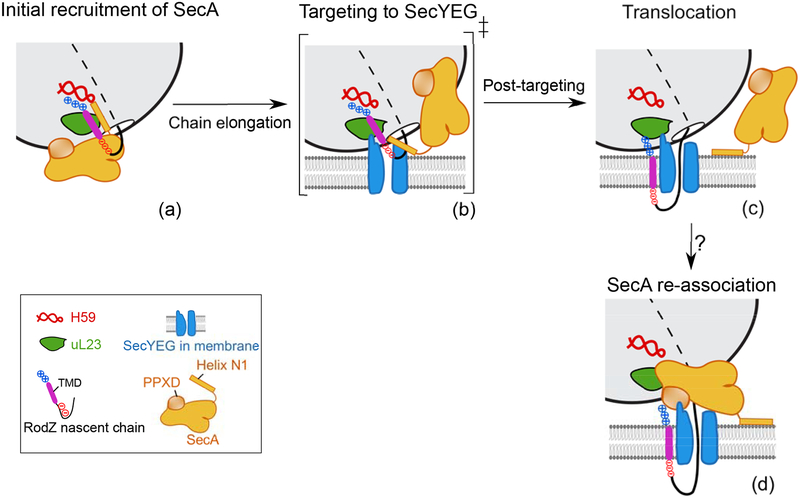
We propose the following model for SecA-mediated cotranslational protein targeting (Fig. 7). SecA is recruited cotranslationally to nascent proteins emerging from the ribosome via multiple interactions, including recognition of the TMD via the composite binding site formed by SecA helix N1 and the hydrophobic groove on uL23, and recognition of charged residues flanking the TMD via H59 of 23S rRNA and the basic SecA surface near helix N1 (Fig. 7a). SecYEG associates with and actively remodels the RNC•SecA complex, generating a transient intermediate in which SecA is repositioned on the RNC with weakened contacts (Fig. 7b), thus facilitating handover of the TMD from SecA to SecYEG (Fig. 7c). The hydrophobic groove of uL23 and H59 could help stabilize the nascent TMD during the handover, and the interaction of SecA helix N1 with anionic phospholipids might promote its repositioning in this intermediate (Fig. 7b). After the nascent polypeptide docks onto SecYEG and initiates translocation, SecA may remain bound at the membrane via helix N132 (Fig. 7c). As the ribosome-translocon junction can be transiently disrupted during membrane protein integration44, and as SecA is required for the translocation of membrane proteins containing large periplasmic loops45–47, it is plausible that SecA could reassociate with the translocation complex as periplasmic domains emerge from the ribosome and use its ATPase cycle to drive translocation (Fig. 7d).
Fig. 7. Model of SecA-mediated cotranslational recognition and targeting of membrane proteins.
a, Nascent TMD emerges from exit tunnel and recruits SecA. b, SecYEG invades the SecA-bound RNC, generating a transient ternary complex in which SecA is repositioned and its contacts with the RNC are weakened. c, The nascent TMD is released from SecA and docks onto SecYEG, which initiates translocation. SecA could remain bound to the membrane surface via helix N1. d, As the nascent chain further elongates, SecA could also re-associate with the translocation complex and use its ATPase cycle to drive translocation of the periplasmic domain. Question mark denotes that it is unclear whether step (d) occurs, nor what the molecular signals are that trigger SecA reassociation.
Methods
Protein expression and purification
N-terminally His6-tagged E. coli SecA contains a mutation (C98S) to remove the surface exposed cysteine, as described before18. For SecA used in crosslinking and cryoEM studies, the three cysteines at the non-essential C-terminus of SecA was also removed (‘ΔZFD’, residues 885–896 deleted). SecA variants used for crosslinking and fluorescent analyses were expressed and purified as described before18. SecA used for cryoEM was further purified by size exclusion chromatography on Superdex 200 10/300 GL (GE healthcare) in buffer containing 50 mM KHEPES, pH 7.5, 150 mM KOAc, 10 mM Mg(OAC)2, 2 mM DTT.
SecYEG containing N-terminally His6-tagged SecY was expressed and purified as described before49 with slight modifications. Cells were induced at log phase by 0.5 mM IPTG for 3 hrs at 37 °C. Harvested cells were resuspended in KC300G buffer (50 mM KHEPES pH 7.5, 300 mM NaCl, 10% glycerol) and lysed by sonication. Lysate was clarified by centrifugation at 12,000 g, 4 °C for 20 min in JA 20 rotor (Beckman Coulter). The supernatant was ultracentrifuged at 42,000 rpm, 4 °C in Ti70 rotor (Beckman Coulter) for 50 min. The membrane fraction was resuspended in KS200G buffer (50 mM KHEPES pH 7.5, 200 mM NaCl, 10% glycerol) by dounce homogenizer (Wheaton). N-Dodecyl-β-D-Maltopyranoside (DDM) and KS200G buffer were added to the membrane suspension to adjust the total protein concentration to 10 mg/ml and DDM concentration to 10% (w/w). The suspension was clarified by ultracetrifugation at 42,000 rpm for 50 min in Ti 70 rotor, and purified by Ni-NTA agarose. Protein was loaded and washed in SecYEG buffer 1 (50 mM KHEPES pH 7.5, 200 mM NaCl, 10% glycerol, 10 mM imidazole, 0.02% DDM), and eluted in SecYEG buffer 2 (50 mM KHEPES pH 7.5, 200 mM NaCl, 10% glycerol, 300 mM imidazole, 0.02% DDM). Eluted protein was dialyzed into KS50G (50 mM KHEPES pH 7.5, 50 mM NaCl, 10% glycerol, 0.02% DDM), loaded onto SP sepharose Fast Flow (GE healthcare) in KS50G, and eluted using a gradient of 50–1000 mM NaCl. Protein fractions were pooled and concentrated to ~75 μM using Amicon, 50K MWCO centrifugal filter unit (MilliporeSigma).
SecB, SRP, FtsY, trigger factor, and ApoE422K were expressed and purified as described18,32. SUMO fusions to the TMD of RodZ (residue 104–133) or signal sequence of PhoA (residues 1–21) were expressed and purified as described previously18.
RNC preparation
Stalled RNCs were generated by in vitro translation in S30 extract as described previously18. 7-hydroxycoumaryl ethylglycine (Cm)-labeled RNCs were prepared similarly, except that the translation reaction was supplemented with Cm (Bachem), tRNACm, and Cm tRNA synthetase, as described before18,50. RNC containing an engineered cysteine at uL23 was prepared using S30 extracts from strain KC624 harboring uL23(S21C) (KC6 ΔrplW::kan pL23S21C). KC624 was made from E. coli strain KC651, by transforming with the plasmid pEK20 containing single cysteine mutant uL23, and subsequent knocking out genomic uL23 via lambda-red recombination52. RNC for cryoEM study was further purified by sucrose gradient (10–50 %) to enrich monosome.
Crosslinking
SecA, RNCs, and SUMO fusion proteins were buffer exchanged into labeling buffer A (50 mM KHEPES, pH 7.0, 150 mM KOAc, 10 mM Mg(OAC)2, 5 mM TCEP). Crosslinking reactions between SecA and RNC were performed in labeling buffer A and contained 1 μM SecA, 0.5 μM RNC, and 0.2 mM BMH (or BMOE). Crosslinking reaction between SecA and SUMO fusion proteins were performed in labeling buffer B (20 mM KHEPES, pH 7.0, 50 mM KOAc, 2 mM Mg(OAC)2, 5 mM TCEP) and contained 1 μM SecA, 8.3 μM SUMO fusions, and 2 mM BMH. All crosslinking reactions were carried out at room temperature for 45 min, and quenched by addition of DTT to 100 mM. Samples were analyzed by SDS-PAGE and western blotting using the indicated antibodies.
Cryo-EM data collection
RNCs (500 nM) were mixed with SecA (1 μM) in the presence of 0.2 mM bismaleimidohexane (BMH) for 50 minutes protected from light and at room temperature. The final reaction size was 10 μl in buffer C (50 mM HEPES-KOH pH 7.4, 100 mM KOAc, 10 mM Mg(OAC)2. After quenching the reaction with 100 mM Dithiothreitol (DTT) for 5 minutes, reaction was diluted to 60 μl in buffer C to a final concentration of 80 nM of RNCs and cooled on ice for 15 minutes. The sample (5 μl) was then applied to Quantifoil grids (R2/2) freshly coated with a thin layer of carbon, incubated for one minute before plunge frozen into a liquid ethane/propane mix cooled to liquid nitrogen temperature using a Vitrobot Mark IV at 95% relative humidity and previously cooled to 4 °C. Cryo-EM data collection was performed using a Titan Krios electron microscope (ThermoFisher Scientific) operated at 300 KV and equipped with Falcon 3EC direct electron detector. Micrographs were recorded in integrating mode as movie stacks with exposure time of 1.66 seconds and a total of 33 frames were stored per movie stack. The defocus range applied was −1.5 to −2.8 μm. The calibrated magnification of the data acquisition was 100,719x, which resulted in a pixel size of 1.39 Å per pixel and an electron dose of 40 e−/Å2 was applied. The EPU software was used as a setup for the automatic data collection and a total of 15,162 movie stacks were collected at a rate of 100 images per hour.
Data processing and map calculation
Motioncorr253 was used for performing drift collection and dose weighting on the movie stacks. Contrast transfer function (CTF) was first calculated using GCTF54 for aligned and non-dose weighted frames. CTF was then carefully inspected for drift and only images that extend beyond 5 Å were retained. A total of 2,613,025 particle-images were picked from the dose-weighted frames with BATCHBOXER implemented in EMAN55 and using projections of an empty 70S ribosome filtered to 40 Å resolution as a reference. After 25 iterations of two dimensional (2D) image classification in RELION356 on binned images (5.56 Å per pixel), a total of 2,477,544 particle-images were selected and further refined following the 3D refine approach in RELION3 and using a bacterial ribosome a reference filtered to 60 Å resolution. Images were then subjected to 3D focused classification without alignments by applying a circular mask onto the ribosome tunnel exit site. The 3D classification yielded two classes, one of which contained a density of the SecA protein bound to the ribosome (329,048 particle-images). The remaining classes were either of an empty ribosome or contained weak density on the exit tunnel and thus were discarded. To further improve the density of SecA, a second round of 3D classification was performed using 2-fold binned images (2.78 Å per pixel), and the 3D class displaying high resolution features of SecA was retained (140,665 particle-images). A final round of 3D classification was performed by adjusting the tau value (T=10) in RELION and this yielded a 3D class where secondary structural elements of SecA can be resolved. The selected particles (37,334 particles) were refined at a full pixel size without binning (1.39 Å per pixel; box size is 320 × 320 pixels) resulting an overall average reconstruction of 3.1 Å resolution (Map1) using the 3D refinement approach in cryoSPARC57. Although the contact points between the ribosome and SecA were resolved to side-chain resolution (< 3.5 Å), the outer shell of SecA was around 8 Å resolution. To improve the local resolution of SecA, a focused refinement approach was performed by masking out the ribosome of the refined map, then re-centering the picked particle images around SecA density. Local angular searches were then applied in addition to small angular increments (1.8 degrees), which yielded a final reconstruction of the SecA protein at a local resolution of 5.7 Å (Map2). Local resolution and gold standard FSC plots using FSC=0.143 criterion were calculated as implemented in RELION3. Final maps were sharpened either in RELION3 or with the auto-sharpen option implemented in PHENIX58.
Model building
For the model building of the RNC•SecA complex, coordinates of the 50S (PDB ID:5GAG) and of SecA (PDB ID: 2FSF) and the coordinates of SecA PPXD (PDB ID: 2VDA) were docked as rigid body elements into the cryo-EM map using USCF CHIMERA59. Manual adjustments of the protein α-helices of SecA were done using COOT60 first into Map2 as the secondary structural elements of SecA were resolved. The contact sites between the ribosome and SecA in Map1 were resolved to 3.1–3.5 Å and allowed us to build these regions de novo, in particular, the amphipathic helix at the N-terminal end of SecA and the nascent chain within the ribosome tunnel. H59 of the ribosomal RNA and uL23 were manually adjusted to better fit the EM density. The density of the transmembrane domain (TMD) of RodZ was resolved and assigned based on the side chain density of this region, which allowed to establish the registry and directionality of the TMD. The rest of the nascent chain region in the ribosome tunnel was traced as poly-alanine chains. The resulting model was then refined into the corresponding EM densities and subjected to five cycles of real space refinements using phenix.real_space_refine61, during which protein secondary structure, Ramachandran and side chain rotamer restraints, RNA base pair restraints were applied. The fit of the EM map was validated using the real space correlation coefficients (CCmask) between the model and the versus map Fourier Shell Correlation (FSC) at FSC=0.5 as a cut-off criterion and resulted in similar resolution as the half-set map FSC using FSC=0.143 criterion. Numbering of the TMD residues of RodZ corresponds to the numbering of residues in the full protein sequence from E. coli. In the deposited model, residues numbering of the nascent chain construct starts from the first methionine as residue number 1. Images were prepared in either Chimera, ChimeraX, or PyMOL.
Western blot
Rabbit anti-uL23 antibody was customized from GenScript using CGKVKRHGQRIGRRS as the epitope. Anti-T7 antibody was purchased from Abcam. Anti-strep, anti-HA and anti-SUMO antibody were purchased from GenScript. Primary antibodies were incubated with IRDye® 800CW secondary antibodies (LI-COR) for detection. Protein band intensity was quantified by the Odyssey® CLx imaging system. The additional bands in the anti-T7 blots are potentially SecA dimer induced by cysteine-cysteine crosslinking, because: (i) their apparent size (slightly above 200kDa) is consistent with a SecA dimer (204kDa); (ii) their appearance depends on the presence of the crosslinker; (iii) their intensity depends on the position of engineered cysteine on SecA (Fig. 1a and new Supplementary Fig. 1g), with the strongest band at residue 797 (Fig. 1a) near a reported SecA dimer interface48.
Nanodisc reconstitution
Reconstitution was carried out as described previously32 with slight modifications. A lipid mixture containing DOPC:DOPG:DOPE at a molar ratio of 4:3:3 (Avanti) was dried under nitrogen gas and then in a vacuum desiccator overnight, and resuspended in lipid buffer (20 mM HEPES pH 7.5, 112 mM KCl, 0.4 mM TCEP, 46 mM Na-cholate) at a total lipid concentration of 22 mM. Reconstitution reactions were performed in nanodisc buffer (25 mM HEPES pH 7.5, 140 mM KCl, 0.5 mM TCEP, 46 mM Na-cholate) and contained 13.4 μM SecYEG, 133.8 μM ApoE422K and 12.2 mM lipid mixture for SecYEG nanodisc; and 133.8 μM ApoE422K and 12.2 mM lipid mixture for empty nanodisc. The mixtures were incubated at 4 °C for 1 hr on a rotary shaker. Bio-bead SM-2 resin (Bio-rad) was washed by methanol, ddH2O and bead buffer (25 mM HEPES pH 7.5, 140 mM KCl, 0.5 mM TCEP). Remove extra washed bio-bead so that the remaining bead volume is equal to the volume of the reconstitution reaction mixture. The reconstitution reaction mixture was added to the remaining bead and incubated at 4 °C for overnight on rotary shaker. The mixture was filtered to remove bio-bead and pelleted at 77,000 rpm for 25 min in TLA 120.2 (Beckman Coulter) to remove aggregates. The supernatant was concentrated in Amicon, 30K MWCO centrifugal filter unit (MilliporeSigma). The concentration of nanodisc was calculated as follows: the concentration of ApoE422K in nanodisc was quantified by SDS-PAGE using known concentrations of purified ApoE422K as standards. As each large nanodisc (~40 nm) contains ~8 copies of ApoE422K on average32,62, the concentration of ApoE422K was divided by 8 to obtain the concentration of nanodisc. The concentration of SecYEG in nanonisc was determined by SDS-PAGE using known concentrations of purified SecYEG as standards.
Negative stain electron microscopy
10 nM nanodiscs were applied onto a glow discharged ultrathin C film on holey carbon support film, 400 mesh, Cu grids (Ted Pella, Inc.). Samples were stained with 3% uranyl acetate. Data were collected using a FEI Tecnai T12 transmission electron microscope at 120 keV on a Gatan Ultrascan 2k × 2k CCD detector. Images were acquired using a 1 s exposure time at a nominal magnification of 42,000x at 2–3um defocus, resulting in 2.5 Å per pixel.
Fluorescence labeling of SecA
The single cysteine mutant of SecA (C98S/S12C) and its derivatives were reduced with 2 mM DTT at 4 °C for 30 min followed by dialysis in Labeling buffer (20 mM KHEPES, pH 7.0, 300 mM KCl, 10% glycerol, 2 mM TCEP) to remove DTT. 40 μM SecA (C98S/S12C) was mixed with a 20-fold excess of BODIPY-FL maleimide on a rotary shaker at 4 °C for 4 hr. After quenching with 10 mM DTT, free dye was removed by chromatography on Sephadex G-25 column (Sigma-Aldrich) in SRP buffer (50 mM KHEPES, pH 7.5, 150 mM KOAc, 10 mM Mg(OAc)2, 2 mM DTT, 10% glycerol). Labeling efficiencies was ~90 %, determined using the adsorption coefficient of ε = 73,000 M−1cm−1 for BODIPY-FL maleimide in aqueous buffer18. The cysteines in the zinc-finger domain of SecA are coordinated by Zn2+ and were not labeled (data not shown).
In vitro translation-translocation in PURE system
Translation was performed at 30 °C using PURExpress® in vitro protein synthesis kit (NEB) supplemented with 35S-Methionine (1.5 mCi/ml, PerkinElmer) and the indicated concentrations of cytosolic factors (SecA, SecB, Ffh, FtsY or TF). Unless otherwise indicated, 0.5 mg/ml Urea-washed IMV18 was added 5 min after initiation of translation. The reaction was continued for 85 min at 30 °C, after which it was split equally into two samples, one of which was digested with 0.5 mg/ml proteinase K for 30 min at 25 °C. Digestion was stopped by addition of 5 mM PMSF, after which the sample was incubated on ice for 10 min. Samples with and without proteinase K treatment were analyzed by SDS-PAGE and autoradiography.
Statistics and Reproducibility
All crosslinking experiments and biophysical measurements were repeated independent at least twice with similar results. Statistics are reported for all quantitative measurements.
Reporting Summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data Availability
Cryo-EM maps are deposited in the EMDB with accession codes EMD-10073 (RNCRodZ•SecA) and EMD-10074 (SecA, local refinement), and model coordinates are deposited in the wwPDB with accession code PDB 6S0K. Other data are available from corresponding authors upon reasonable request.
Supplementary Material
Table 1.
Cryo-EM data collection, refinement and validation statistics
| RNCRodZ•SecA (EMDB-10073, PDB 6S0K) | SecA (local refinement) (EMDB-10074) | |
|---|---|---|
| Data collection and processing | ||
| Magnification | 100,719 | 100,719 |
| Voltage (kV) | 300 | 300 |
| Electron exposure (e–/Å2) | 40 | 40 |
| Defocus range (μm) | 1.5 – 2.8 | 1.5 – 2.8 |
| Pixel size (Å) | 1.39 | 1.39 |
| Symmetry imposed | C1 | C1 |
| Initial particle images (no.) | 2,613,025 | 2,613,025 |
| Final particle images (no.) | 37,334 | 37,334 |
| Map resolution (Å) | 3.1 | 5.7 |
| FSC threshold | 0.143 | 0.143 |
| Map resolution range (Å) | 2.9 – 9.5 | 5.5 – 9.5 |
| Refinement | ||
| Initial model used (PDB code) | 5GAG, 2FSF, 2VDA | |
| Model resolution (Å) | 3.3 | |
| FSC threshold | 0.5 | |
| Model resolution range (Å) | – | |
| Map sharpening B factor (Å2) | –130.53 | |
| Model composition | ||
| Non-hydrogen atoms | 98,708 | |
| Protein residues | 4,384 | |
| Nucleic acid residues | 3,006 | |
| Ligands | 69 | |
| B factors (Å2) | ||
| Protein | 148.63 | |
| Nucleotide | 95.06 | |
| Ligand | 60.40 | |
| R.m.s. deviations | ||
| Bond lengths (Å) | 0.005 | |
| Bond angles (°) | 0.776 | |
| Validation | ||
| MolProbity score | 1.52 | |
| Clashscore | 4.21 | |
| Poor rotamers (%) | 0.73 | |
| Ramachandran plot | ||
| Favored (%) | 95.44 | |
| Allowed (%) | 4.28 | |
| Disallowed (%) | 0.28 |
Acknowledgements
We thank A. McDowall and H. Wang for assistance with negative stain electron microscopy data collection, J. Rothman for sharing plasmid ApoE422k, D. Boehringer and A. Scaiola for the support with EM data collection and processing, M. Leibundgut and M. Saurer for the support with model building, and members of the Shan and Ban groups for discussions and comments on the manuscript. We also thank T. Miller, M. Zimmer and F. Huber for helpful discussions. Cryo-EM data was collected at the Scientific Center for Optical and Electron Microscopy at the ETH Zurich (ScopeM). We gratefully acknowledge the support of NVIDIA Corporation for the Titan Xp GPU used in this research through a GPU Grant program awarded to A.J.; M.J. was supported by the internal research grant of the ETH to N.B. (ETH-40 16–2). This work was supported by National Institutes of Health grant GM107368A and the Gordon and Betty Moore Foundation through grant GBMF2939 to S.-o. Shan and by the Swiss National Science Foundation (SNSF) (grant number 310030B_163478), National Center of Excellence in Research (NCCR) RNA & Disease Program of the SNSF (grant number 51NF40_141735) to N.B..
Footnotes
Competing Interests Statement
The authors declare no competing interests.
References
- 1.Shao S & Hegde RS Target Selection during Protein Quality Control. Trends Biochem Sci 41, 124–137 (2015). [DOI] [PubMed] [Google Scholar]
- 2.Brandman O & Hegde RS Ribosome-associated protein quality control. Nat Struct Mol Biol 23, 7–15 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Zhang X & Shan SO Fidelity of cotranslational protein targeting by the signal recognition particle. Annu Rev Biophys 43, 381–408 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Cymer F, von Heijne G & White SH Mechanisms of integral membrane protein insertion and folding. J Mol Biol 427, 999–1022 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Elvekrog MM & Walter P Dynamics of co-translational protein targeting. Curr Opin Chem Biol 29, 79–86 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Nyathi Y, Wilkinson BM & Pool MR Co-translational targeting and translocation of proteins to the endoplasmic reticulum. Biochim Biophys Acta 1833, 2392–402 (2013). [DOI] [PubMed] [Google Scholar]
- 7.Zhang X, Rashid R, Wang K & Shan SO Sequential checkpoints govern substrate selection during cotranslational protein targeting. Science 328, 757–760 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chartron JW, Hunt KC & Frydman J Cotranslational signal-independent SRP preloading during membrane targeting. Nature 536, 224–8 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Schibich D et al. Global profiling of SRP interaction with nascent polypeptides. Nature 536, 219–223 (2016). [DOI] [PubMed] [Google Scholar]
- 10.Jomaa A, Boehringer D, Leibundgut M & Ban N Structures of the E. coli translating ribosome with SRP and its receptor and with the translocon. Nat Commun 7, 10471 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Jomaa A et al. Structure of the quaternary complex between SRP, SR, and translocon bound to the translating ribosome. Nat Commun 8, 15470 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Aviram N et al. The SND proteins constitute an alternative targeting route to the endoplasmic reticulum. Nature 540, 134–138 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ast T, Cohen G & Schuldiner M A network of cytosolic factors targets SRP-independent proteins to the endoplasmic reticulum. Cell 152, 1134–1145 (2013). [DOI] [PubMed] [Google Scholar]
- 14.Chitwood PJ, Juszkiewicz S, Guna A, Shao S & Hegde RS EMC Is Required to Initiate Accurate Membrane Protein Topogenesis. Cell 175, 1507–1519.e16 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Guna A, Volkmar N, Christianson JC & Hegde RS The ER membrane protein complex is a transmembrane domain insertase. Science 359, 470–473 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shurtleff MJ et al. The ER membrane protein complex interacts cotranslationally to enable biogenesis of multipass membrane proteins. eLife 7, 1–23 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Huber D et al. SecA Interacts with Ribosomes in Order to Facilitate Posttranslational Translocation in Bacteria. Mol Cell 41, 343–353 (2011). [DOI] [PubMed] [Google Scholar]
- 18.Wang S, Yang CI & Shan SO SecA mediates cotranslational targeting and translocation of an inner membrane protein. J Cell Biol 216, 3639–3653 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Singh R et al. Cryo-electron microscopic structure of SecA bound to the 70S ribosome. J Biol Chem 289, 7190–7199 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Huber D et al. SecA cotranslationally interacts with nascent substrate proteins in vivo. J Bacteriol 199, e00622–16 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rawat S, Zhu L, Lindner E, Dalbey RE & White SH SecA drives transmembrane insertion of RodZ, an unusual single-span membrane protein. J Mol Biol 427, 1023–1037 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hartl FU, Lecker S, Schiebel E, Hundrick JP & Wickner W The binding cascade of SecB to SecA to SecY/E mediates preprotein targeting to the E. coli plasma membrane. Cell 63, 269–279 (1990). [DOI] [PubMed] [Google Scholar]
- 23.Kimura E, Akita M, Matsuyaman S & Mizushima S Determination of a region in SecA that interacts with presecretory proteins in Escherichia coli. J Biol Chem 266, 6600–6606 (1991). [PubMed] [Google Scholar]
- 24.Gelis I et al. Structural basis for signal-sequence recognition by the translocase motor SecA as determined by NMR. Cell 131, 756–769 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chatzi KE et al. Preprotein mature domains contain translocase targeting signals that are essential for secretion. J Cell Biol 216, 1357–1369 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zimmer J, Nam Y & Rapoport TA Structure of a complex of the ATPase SecA and the protein-translocation channel. Nature 455, 936–943 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bauer BW & Rapoport TA Mapping polypeptide interactions of the SecA ATPase during translocation. Proc Natl Acad Sci U S A 106, 20800–20805 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Frauenfeld J et al. Cryo-EM structure of the ribosome-SecYE complex in the membrane environment. Nat Struct Mol Biol 18, 614–21 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Park E et al. Structure of the SecY channel during initiation of protein translocation. Nature 506, 102–6 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wu ZC, De Keyzer J, Kedrov A & Driessen AJM Competitive binding of the SecA ATPase and ribosomes to the SecYEG translocon. J Biol Chem 287, 7885–7895 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nakatogawa H & Ito K The ribosomal exit tunnel functions as a discriminating gate. Cell 108, 629–636 (2002). [DOI] [PubMed] [Google Scholar]
- 32.Koch S et al. Lipids activate SecA for high affinity binding to the SecYEG complex. J Biol Chem 291, 22534–22543 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Erlandson KJ et al. A role for the two-helix finger of the SecA ATPase in protein translocation. Nature 455, 984–7 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Or E, Navon A & Rapoport T Dissociation of the dimeric SecA ATPase during protein translocation across the bacterial membrane. EMBO J 21, 4470–4479 (2002). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Osborne AR, Clemons WM Jr. & Rapoport TA Monomer closed PPXD; dimer open PPXD. Proc Natl Acad Sci U S A 101, 10937–10942 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sharma V et al. Crystal structure of Mycobacterium tuberculosis SecA, a preprotein translocating ATPase. Proc Natl Acad Sci U S A 100, 2243–8 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Li L et al. Crystal structure of a substrate-engaged SecY protein-translocation channel. Nature 531, 395–399 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Knorr AG et al. Ribosome-NatA architecture reveals that rRNA expansion segments coordinate N-terminal acetylation. Nat Struct Mol Biol 26, 35–39 (2019). [DOI] [PubMed] [Google Scholar]
- 39.Ferbitz L et al. Trigger factor in complex with the ribosome forms a molecular cradle for nascent proteins. Nature 431, 590–596 (2004). [DOI] [PubMed] [Google Scholar]
- 40.Akopian D, Shen K, Zhang X & Shan SO Signal recognition particle: an essential protein-targeting machine. Annu Rev Biochem 82, 693–721 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ariosa A, Lee JH, Wang S, Saraogi I & Shan S egulation by a chaperone improves substrate selectivity during cotranslational protein targeting. Proc Natl Acad Sci U S A 112, E3169–78 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ranjan A, Mercier E, Bhatt A & Wintermeyer W Signal recognition particle prevents N-terminal processing of bacterial membrane proteins. Nat Commun 8, 15562 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Saraogi I, Akopian D & Shan SO Regulation of cargo recognition, commitment, and unloading drives cotranslational protein targeting. J Cell Biol 205, 693–706 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Devaraneni PK et al. Stepwise insertion and inversion of a type II signal anchor sequence in the ribosome-Sec61 translocon complex. Cell 146, 134–147 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Traxler B & Murphy C Insertion of the polytopic membrane protein MalF is dependent on the bacterial secretion machinery. J Biol Chem 271, 12394–12400 (1996). [DOI] [PubMed] [Google Scholar]
- 46.Sääf A, Andersson H, Gafvelin G & von Heijne G SecA-dependence of the translocation of a large periplasmic loop in the Escherichia coli MalF inner membrane protein is a function of sequence context. Mol Membr Biol 12, 209–215 (1995). [DOI] [PubMed] [Google Scholar]
- 47.Gebert JF, Overhoff B, Manson MD & Boos W The Tsr chemosensory transducer of Escherichia coli assembles into the cytoplasmic membrane via a SecA-dependent process. J Biol Chem 263, 16652–16660 (1988). [PubMed] [Google Scholar]
Method-only References
- 48.Yu D, Wowor AJ, Cole JL & Kendall DA Defining the Escherichia coli SecA dimer interface residues through in vivo site-specific photo-cross-linking. Journal of Bacteriology 195, 2817–2825 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Akopian D, Dalal K, Shen K, Duong F & Shan SO SecYEG activates GTPases to drive the completion of cotranslational protein targeting. J Cell Biol 200, 397–405 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Saraogi I, Zhang D, Chandrasekaran S & Shan SO Site-specific fluorescent labeling of nascent proteins on the translating ribosome. J Am Chem Soc 133, 14936–14939 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Calhoun KA & Swartz JR Total amino acid stabilization during cell-free protein synthesis reactions. J Biotechnol 123, 193–203 (2006). [DOI] [PubMed] [Google Scholar]
- 52.Datsenko KA & Wanner BL One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. U.S.A 97, 6040–6045 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Zheng SQ et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat Methods 14, 331–332 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zhang K Gctf: Real-time CTF determination and correction. J Struct Biol 193, 1–12 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Ludtke SJ, Baldwin PR & Chiu W EMAN: semiautomated software for high-resolution single-particle reconstructions. J Struct Biol 128, 82–97 (1999). [DOI] [PubMed] [Google Scholar]
- 56.Zivanov J et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7(2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Punjani A, Rubinstein JL, Fleet DJ & Brubaker MA cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat Methods 14, 290–296 (2017). [DOI] [PubMed] [Google Scholar]
- 58.Afonine PV et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr D Struct Biol 74, 814–840 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Pettersen EF et al. UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 25, 1605–12 (2004). [DOI] [PubMed] [Google Scholar]
- 60.Emsley P & Cowtan K Coot: model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr 60, 2126–32 (2004). [DOI] [PubMed] [Google Scholar]
- 61.Afonine PV et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr D Struct Biol 74, 531–544 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Blanchette CD et al. Quantifying size distributions of nanolipoprotein particles with single-particle analysis and molecular dynamic simulations. J Lipid Res 49, 1420–1430 (2008). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Cryo-EM maps are deposited in the EMDB with accession codes EMD-10073 (RNCRodZ•SecA) and EMD-10074 (SecA, local refinement), and model coordinates are deposited in the wwPDB with accession code PDB 6S0K. Other data are available from corresponding authors upon reasonable request.