Abstract
Long non-coding RNAs (lncRNAs) serve critical roles in regulating cellular homeostasis, and their deregulated expression/activity is associated with neoplastic transformation. The maternally expressed gene 3 (MEG3) has been extensively described as a tumor suppressor gene in different types of cancer, including breast cancer. Interestingly, using a panel of seven different breast cancer cell lines, the present study revealed that MEG3 is highly expressed in the triple negative metastatic human Hs578T breast cancer cell line, which is refractory to different therapeutic approaches. Therefore, the present study aimed to investigate the phenotypic impact of MEG3 deletion in this cell line. Using the CRISPR/Cas9 system, complete knockout (KO) of MEG3 was achieved. Deletion was confirmed by genomic PCR and reverse transcription-quantitative PCR. The MEG3_KO cell population displaying the highest efficiency of genomic editing was selected for phenotypic in vitro assays, including wound scratch and Transwell assays, flow cytometry and immunofluorescence. The results demonstrated that MEG3 deletion increased cell proliferation, anchorage-independent cell growth and cell motility, which was consistent with its well-known tumor suppressor function. However, the present study revealed that MEG3_KO also lead to decreased cell invasiveness ability, supporting previous evidence that MEG3 modulates epithelial-to-mesenchymal inducing factors. The present study demonstrated that deletion of MEG3 promoted an increase in transforming growth factor β and N-cadherin protein levels and significant reduction in matrix metallopeptidase 2, zinc-finger E-box binding homeobox 1 and collagen type III α1 chain gene expression levels. Additionally, MEG3_KO cells displayed significant resistance to doxorubicin treatment, demonstrating the role of this lncRNA in cancer cell survival by regulating apoptosis. The present study highlighted the utility of CRISPR/Cas9 for anticancer studies of intergenic lncRNAs and demonstrated that, although Hs578T cells express MEG3 at high levels, these cells display mechanisms to escape the growth suppression effects of this lncRNA. Notably, the detailed pathological mechanisms of MEG3 concerning tumor metastasis remain to be elucidated prior to applying MEG3 expression/activation in future therapeutic approaches for breast cancer treatment.
Keywords: maternally expressed gene 3, long non-coding RNAs, CRISPR/Cas9 system, Hs578T, breast cancer
Introduction
Due to increasingly sophisticated Molecular Biology analytical methods, new groups of non-coding RNAs (ncRNAs) have been described over the past few years (1). These molecules were revealed to be more than simple inert sequences, emerging as functional regulatory components, which play critical roles in modulating chromatin architecture, transcription and RNA splicing, translation and turnover (2). Due to their biological relevance, these ncRNAs were also found to be involved in different human diseases (3), including cancer (4).
Long non-coding RNAs (lncRNAs) are noncoding transcripts of around 200 nucleotides or more (5). The number of annotated sequences greatly improved with third-generation sequencing methods (6), however, the functional characterization of most of these lncRNAs is still lacking (7). Various functional mechanisms have been described for lncRNAs including recruitment of transcription factors for promoter regions, guiding chromatin modifiers to specific genomic loci, allosteric modulation of transcriptional regulatory proteins, alteration of nuclear domains, modulation of translation or mRNA stability and working as a natural competing endogenous RNAs (5,8,9).
The maternally expressed gene 3 (MEG3) was the first lncRNA proposed to have a tumor suppressor function (10,11). Using cDNA-representational difference analysis, MEG3 expression was not detected in either pituitary tumors, when compared to normal human pituitary tissue, nor in several human cancer cell lines (10). Moreover, ectopic expression of MEG3 RNA suppresses cell growth in different tumor cells (12–14), further supporting the tumor suppressor role of this gene.
Despite all the great advances in the field, breast cancer remains to be the leading cause of cancer death among women between 20 to 59 years old (15,16). The most lethal type of breast cancer is the triple negative breast cancer (TNBC), which lacks the expression of cell receptors for estrogen, progesterone and do not show amplification of the human epidermal growth factor receptor 2 (HER2) gene (17). These characteristics prevent the use of conventional drug therapies and account for approximately 15% of all diagnosed breast cancers (18), highlighting the urgent need for well-defined molecular targets for treatment of this type of cancer.
In silico analysis has suggested that MEG3 could be a valuable prognostic factor and a potential therapeutic target for breast cancer patients, with an impact on disease-free survival, relapse-free survival and progression-free survival (19–21). Consistently, functional studies have shown that overexpression of MEG3 decreases breast cancer cell lines growth rate, invasion capacity, and tumor angiogenesis through downregulation of AKT signaling (22) and by enhancing p53 transcriptional activity (23).
The CRISPR/Cas9 system provides a revolutionary genome-editing tool for all areas of Molecular Biology (24–26). Some techniques have been previously applied to achieve lncRNA deletion, however, the CRISPR/Cas9 approach to target lncRNAs has scarcely been explored in the literature (27–29). Similarly to protein-coding genes, Cas9 nuclease may be used to delete the entire lncRNA gene or to introduce RNA-destabilizing elements into their loci, particularly in their promoter region. Here, using a panel of seven breast cancer cell lines, which are representative of tumor progression and aggressiveness in vitro, we found that MEG3 has a discrepant expression in the triple negative metastatic human Hs578T cell line. To better understand the contribution of the lncRNA MEG3 in breast tumorigenesis, we developed a protocol to knockout MEG3 expression by CRISPR/Cas9 and analyzed the phenotypic impact of MEG3_KO using in vitro assays.
Materials and methods
MEG3 expression profiling in breast cancer derived cell lines
Expression profiling was carried out using a panel of breast cancer derived cell lines representing tumor progression, ranging from non-tumorigenic to highly metastatic tumor cells. The following cell lines were obtained from ATCC (American Type Culture Collection): Non-tumoral cell lines MCF10A (CRL-10317; ER-/PR-/AR-/HER2-) and MCF12A (CRL-10782; ER-/PR-/AR+/HER2-); tumoral cell lines estrogen-positive MCF-7 (HTB-22; ER+/PR+/AR+/HER2-), ZR-75-1 (CRL-1500; ER+/PR+/AR+/HER2+); and tumoral cell lines estrogen-negative SK-BR-3 (HTB-30; ER-/PR-/AR+/HER2+), MDA-MB-231 (HTB-26;ER-/PR-/AR+/HER2-) and Hs578T (HTB-126; ER-/PR-/AR+/HER2-). Replicate experiments were carried out with cells at increasing sequential passage number. 293T cells (CRL-3216) were used for the production of the lentiviral particles. All cell lines were maintained at 37°C with 5% CO2 in specific culture mediums following recommendations suggested by the ATCC.
RNA extraction, cDNA synthesis and reverse transcription-quantitative PCR (RT-qPCR)
Total RNA was isolated from cultured cells with Trizol (Life Technologies) and purified with an RNAspin Mini kit (GE Healthcare) according to the manufacturer's instructions, with an extended treatment with DNase I for 1 h. Total RNA was quantified using the ND-1000 (NanoDrop) and its integrity was assessed on a Bioanalyzer (Agilent Technologies). For measuring lncRNAs, reverse transcription was performed with the SuperScript III (Life Technologies) followed by qPCR. PCR with 40 cycles and 1 µg of the resulting purified total RNA (without reverse transcription), using different pairs of primers for tubulin TUBA1C gene and Histone H3 (multiple copy gene) were used to confirm absence genomic DNA in all samples previously. For all genes, oligo-dT20 primed reverse transcription was performed using 1 µg of total RNA in 20 µl of RT reaction with SuperScript III (Life Technologies), followed by qPCR using 5 µl of the 8-fold diluted RT reaction in 20 µl of qPCR (ViiA 7 Real-Time PCR System, Thermo Fisher Scientific). Transcript levels were normalized to HMBS, and represented as relative abundance using the delta Ct method (30). Two controls for the RT step, one without primer (-primer) and the other without reverse transcriptase (-RT) were performed, followed by qPCR with the pair of primers, in order to confirm the absence of RNA self-priming and of genomic DNA contamination in the RT, respectively. Graph design and statistical analyses were carried out using the Graphpad Prism V6 (Graphpad Software). Conditions for PCR reactions were: 40 cycles of 95°C/15 sec, 60°C/1 min, using the following primers: MEG3 forward 5′-TGAAGAACTGCGGATGGAAG-3′ and reverse 5′-CACGTAGGCATCCAGGTGAT-3′; HMBS forward 5′-TGGACCTGGTTGTTCACTCCTT-3′ and reverse 5′-CAACAGCATCATGAGGGTTTTC-3′; TUBA1C forward 5′-TCAACACCTTCTTCAGTGAAACG-3′ and reverse 5′-AGTGCCAGTGCGAACTTCATC-3′; H3 forward 5′-ACGGCTCGTACAAAGCAGAC-3′ and reverse 5′-CCGCTGAAACTTGTTCACTG-3.
Knockout of MEG3 in Hs578T breast cancer cells
Small guide RNAs (sgRNAs) were designed using an online CRISPR Design Tool (http://tools.genome-engineering.org) and then cloned (guide sequences Table I) into the pL-CRISPR.EFS.GFP (Addgene #57818, Massachusetts, USA) or lentiCRISPR-v2 (Addgene #52961, Massachusetts, USA). Viral particles were produced in 293T cells using fourth-generation lentiviral packaging. Hs578T cells were co-transduced in a ratio 1:1 with sgRNAs targeting the 5′ and 3′ ends of the gene locus. Cell populations were first selected with 6 µg/ml puromycin (Life Technologies) for 48 h and after cell expansion, GFP positive cells were sorted by FACS (BD FACSARIA II, BD Bioscience).
Table I.
Primer sequences used to clone the sgRNAs targeting long non-coding RNA MEG3.
| Guide | Forward (5′-3′) | Reverse (5′-3′) |
|---|---|---|
| 5′ sgRNA_MEG3#1 | CACCGTTCGGGTATACGGCAGTCAC | AAACGTGACTGCCGTATACCCGAAC |
| 5′ sgRNA_MEG3#2 | CACCGCACTATTAACGCGATTCTAG | AAACCTAGAATCGCGTTAATAGTGC |
| 3′ sgRNA_MEG3#3 | CACCGTGAACTCTTGTCGGCCAGCG | AAACCGCTGGCCGACAAGAGTTCAC |
| 3′ sgRNA_MEG3#4 | CACCGAAGTGGGCCGAGTCTAAGGT | AAACACCTTAGACTCGGCCCACTTC |
MEG3, maternally expressed gene 3; sgRNA, single guide RNA.
To verify genome-editing efficiency of the different sgRNAs combinations (1+ 3; 1+4; 2+3 and 2+4), transduced cell populations were further expanded for genomic DNA and total RNA extraction. Genomic DNA was extracted with QIAamp DNA Kit (Qiagen, Venlo, Netherlands) and MEG3 deletion was detected by PCR (30 cycles of 94°C/1 min, 60°C/30 sec) using the following primers: forward: 5′-ACCACAGGGTGTTGGTCAT-3′; reverse: 5′-CTCCACCACCACCTCCTC-3′. Total RNA extraction and RT-qPCR were carried out following the same procedure for MEG3 expression profiling characterization. To verify the stability of the MEG3_KO 2+4 cell population, we collected samples from passages 0, 5, 10 and 15 to perform FACS sorting (BD FACSARIA II, BD Bioscience) and RT-qPCR. Hs578T wild type (WT) cells were used as control.
Growth curve and anchorage-independent clonal growth
5×103 cells from the Hs578T_WT and MEG3_KO 2+4 cell population were cultured in 12-well plates in triplicate for each time point. Cells were harvested every 48 h and the total cell number was obtained using the Accuri C6 Plus flow cytometry system (BD Biosciences). Anchorage-independent clonal growth was assessed using the soft-agar assay. Briefly, 104 cells/well were seeded in triplicates in a 24 well plate on top of the 0.6% agarose (Fisher Scientific, Leicestershire, UK) solution in 10% FCS-DMEM. Cells were then allowed to stand for about 10 min before the addition of 500 µl of melted 0.3% agarose in 10% FCS-DMEM. Finally, 500 µl of liquid 10% FCS-DMEM were added. The liquid medium was renewed every two days and total cell colonies were quantified after 14 days using the AMG EVOS FL Inverted Microscope. Statistical analysis was carried out using Graphpad Prism V6 (Graphpad Software).
Migration and invasion assays
For wound-healing scratch assay, 1×105 cells were plated in 24 well plates in triplicates. On the following day, the cell layer was scratched using a 200 µl sterile pipette tip. The wound location was marked and images of the same field were captured to record the wound width at 0,8 and 12 h. The area of migrating cells was measure by ImageJ software. For the transwell assay, cells were washed in phosphate buffered saline (PBS) and resuspended in serum-free medium. A cell suspension containing 104 and 2×104 cells was added to the upper well of transwell migration inserts (pore size: 8 µm, BD Biosciences) or to BD BioCoatTM MatrigelTM invasion chambers (pore size: 8 µm, BD Biosciences), respectively. In the lower well, 700 µl of complete medium were used as chemo-attractant. The cells were maintained for 16 h at 37°C and 5% CO2, followed by fixation in cold methanol for 10 min and staining with Mayer's alum hematoxylin for 20 min. Inserts were mounted in glass slides and six fields per sample were counted, with duplicates for each cell line in each experiment.
Immunofluorescence
For immunofluorescence, cells were seeded onto 13 mm diameter glass coverslips and maintained under usual culture conditions until sub-confluence (less than 80%). Each sample was then fixed in 4% formaldehyde for 10 min, permeabilized with 0.5% Triton X-100 for 10 min and blocked in 1% Bovine Serum Albumin for 60 min, all at room temperature. Primary antibody anti-GFP (1:500, ab6556, Abcam, Cambridge, UK) was incubated overnight at 2–8°C and washed three times with PBS buffer for 10 min. Alexa Fluor 594-phalloidin (1:100, A12381, Thermo Fischer Scientific, Massachusetts, USA) and secondary antibody AlexaFluor 488 goat anti-rabbit IgG (1:400, A27012, Thermo Fischer Scientific) were incubated 1 h at room temperature. Coverslips were mounted using VECTASHIELD Anti-fade Mounting Medium with DAPI (H-1200, Vector Laboratories, CA) and images were acquired with a confocal Zeiss LSM 780-NLO microscope.
Immunofluorescence by high-content screening assays
After seeding the cells in 96-well plates (Greiner Bio-One, 655986), the cultures were washed with PHEM buffer (2 mM HEPES, 10 mM EGTA, 2 mM MgCl2, 60 mM PIPES-pH 6.9) and fixed for 1 h with cold 4% PFA. Cells were permeabilized with 0.1% Triton X-100 for 5 min, blocked with 1% bovine serum albumin (BSA) for 30 min, and then incubated with primary antibody overnight at 4°C. After PHEM-glycine washing (3X), the cells were incubated with the fluorescent dye at room temperature for 1 h and plates were subjected to high-content imaging analysis on MetaXpress High-Content Image Acquisition & Analysis Software (Molecular Devices). The primary antibodies used were mouse anti-N-Cadherin (DAKO, M3613) (1:50) and rabbit anti-TGF-β (Santa Cruz, sc-146) (1:100). After PHEM washing (3X), the cells were incubated with fluorescent dye, namely, FITC-labeled goat anti-rabbit IgG (H + L) (1:100) (A11008 Thermo Fisher Scientific) or AlexaFluor 647 goat anti-mouse IgG (H + L) (1:100) (A21236 Thermo Fisher Scientific) at room temperature for 1 h. Protein expression was determined and quantified through fluorescence signal changes (MFI-Median Fluorescent Intensity) using the Multi Wavelength Cell Scoring module. Cell counts were assessed using Hoechst 33342 (5 µM-Life Technologies, Thermo Fisher Scientific) staining for 1 h, and the stained samples were subjected to high-content imaging analysis. In parallel, the cells were labeled with rabbit anti-GFP (1:1,000) (ABCAM, ab6556) to confirm expression of the reporter protein in MEG3_KO cells. Cytoskeletal F-Actin was marked with AlexaFluor-555 Phalloidin for imaging (1:2,000) (Life Technologies, Thermo Fisher Scientific). The image acquisition and fluorescence intensity measurements were conducted by automatic scanning through the MetaXpress software, using a 40X objective. For each treatment condition and channel, 20 images per well in triplicate, were acquired and analyzed.
Doxorubicin-induced apoptosis
To assess the endogenous modulation of MEG3 expression upon doxorubicin (DXR) treatment (D1515, Sigma-Aldrich, Missouri, USA), 1×105 Hs578T parental cells were plated in six-well plate and treated with DMSO or with a sub-lethal concentration of 0.25 µg/ml DXR for 24 h. Cells were then harvested for RT-qPCR analysis. The chemosensitivity of Hs578T_WT and MEG3_KO cells to DXR was determined by MTT assay. Briefly, 104 cells/well were seeded onto 96-well plates. On the next day, cells were treated with different concentrations of DXR (range, 0–5 µg/ml). At 24 h post-DXR application, cell metabolism was assessed using 0.5 mg/ml MTT (Sigma-Aldrich, St. Louis, MO, USA) solution. Following 4 h incubation at 37°C, the medium was replaced with 150 µl dimethyl sulfoxide (Sigma-Aldrich) and vortexed for 10 min. The absorbance of each well at 490 nm was measured using a microplate reader (SpectraMax Paradigm, Molecular Devices, California, USA). For apoptosis assay, 105 cells/well were seeded onto 6-well plates and incubated overnight under culture conditions. Culture medium was replaced by fresh medium containing DXR (0.5 and 1 µg/ml) and incubated for 24 h. The cells were then harvested, washed with PBS and resuspended in Annexin-V binding buffer (BioVision; #1006). APC conjugated Annexin-V (ThermoFisher Scientific; A35110) was added according to manufacturer's instructions and propidium iodide (ThermoFisher Scientific; P3566) was added at final concentration of 1.0 µg/ml. Samples were kept in the dark for 5 min and the analyses was carried out using FACSARIA II cytometer. For analysis of BAX and BCL2 protein levels, 1×106 cells/ml were trypsinized, centrifuged and fixed in 4% paraformaldehyde. Cells were washed twice with PBS and pelleted at 400 g for 4 min. For permeabilization, cells were incubated for 20 min with 0.05% Triton X-100 in PBS and blocked for 1 h at room temperature in 2% bovine serum albumin. Primary rabbit BAX (#2772S, Cell Signaling) and BCL2 (#2876S, Cell Signaling) antibodies (1:200 dilution) were incubated for 1 h at 4°C. Unbound antibodies were washed out through two cycles of washing with PBS. Rabbit secondary antibody (1:250, Invitrogen AlexaFluor Dye) was incubated for 30 min at 4°C. Cells were washed again and the analyses were performed on a FACS (BD FACSARIA II, BD Bioscience). Data were analyzed using FlowJo 7.6 software and statistical analysis was carried out using the GraphPad Prisma Software.
Statistical analysis
Results are expressed as the mean and standard error of the mean (SEM) of triplicate determinations. Statistical significance was calculated using a two-way ANOVA followed by the Bonferroni test as a post-test or using a two-tailed paired Student t-test. P<0.05 was considered to be significant and statistical results are denoted in the graphs with one asterisk (P<0.05), two asterisks (P<0.01) or three asterisks (P<0.001). All statistical analyses were carried out using Prism 5 (GraphPad Software, La Jolla, CA).
Results
MEG3 is highly expressed in the Hs578T human triple negative mammary cell line
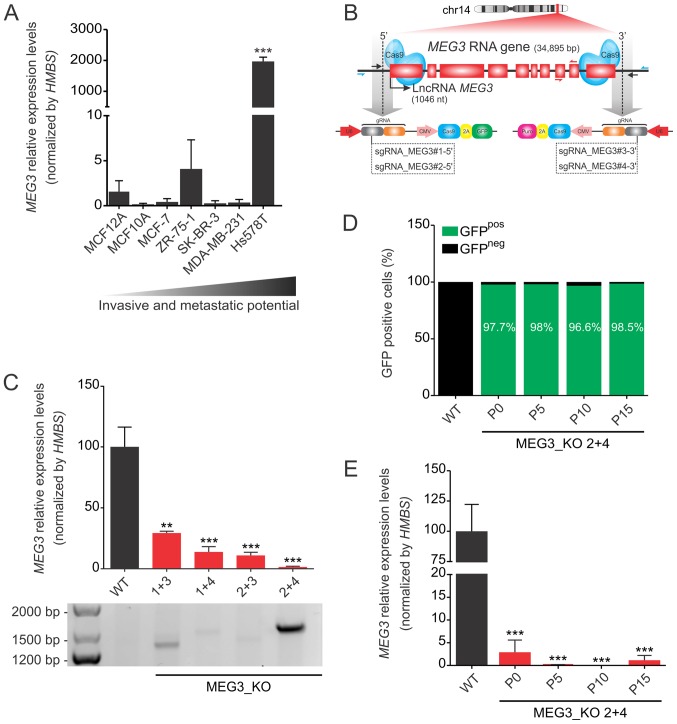
Using a panel of seven breast cancer cell lines, which display different invasive and metastatic potential in vitro, we found that expression of the maternally expressed gene 3 (MEG3) is strikingly increased in the triple negative and highly metastatic Hs578T cell line (average Ct value of 21.5), when compared to the expression levels in the non-tumorigenic MCF12A cell line (average Ct value of 32.9) (Fig. 1A). Interestingly, this gene has been extensively described as a tumor suppressor gene (31). Due to this apparent controversy, we next carried out loss-of-function experiments using the CRISPR/Cas9 knockout system.
Figure 1.
MEG3 is highly expressed in Hs578T cells and its expression can be efficiently deleted using a CRISPR/Cas9 system. (A) Relative mRNA quantification was carried out by RT-qPCR, using a panel of breast cancer cell lines in a scale of increased invasive and metastatic potential. Values are presented as the mean ± SEM (n=3) relative to the expression in the non-tumorigenic MCF12A cell line, after normalization using HMBS as the endogenous control. ***P<0.001 vs. MCF12A. (B) Strategy used to generate MEG3_KO cell population. Hs578T cell line was co-transduced with two sgRNAs, selected with puromycin for 48 h and FACS sorted for GFP-positive cells. The scheme indicates the sgRNA (black arrows) targeting the 5′ (sgRNA_MEG3#1 and #2) and 3′ (sgRNA_MEG3#3 and #4) regions of the MEG3 gene and the primers used for confirmation of genome editing (blue harpoon arrows). Red harpoon arrows represent primers used for RT-qPCR quantifications. (C) Confirmation of knockout cell populations by RT-qPCR and genomic PCR. Values are presented as the mean ± SEM (n=3) relative to the expression in the Hs578T_WT cell line, after normalization using HMBS as an endogenous control. The agarose gel shows the bands amplified when deletion occurred. The primers flank the region outside the sgRNAs target sequence, ranging ~35 kbp. Therefore, the deletion enables fragment amplification, corresponding to 1.500 and 1.300 pb depending on the sgRNA combination. (D) FACS sorting of GFP positive cells in increasing passage numbers (P0, P5, P10 and P15) of the MEG3_KO 2+4 cell population followed by (E) lncRNA MEG3 quantification in the same samples tested in the FACS sorting. Values are presented as the mean ± SEM of three technical replicates for each cell passage relative to the expression in the Hs578T_WT cell line, after normalization using HMBS as an endogenous control. Data were analyzed using one-way ANOVA. **P<0.01; ***P<0.001 vs. WT. FACS, fluorescence-activated cell sorting; GFP, green fluorescent protein; HMBS, hydroxymethylbilane synthase; KO, knockout; lncRNA, long non-coding RNA; MEG3, maternally expressed gene 3; RT-qPCR, reverse transcription-quantitative PCR; sgRNA, single guide RNA; WT, wild-type.
MEG3 CRISPR/Cas9-mediated knockout
To obtain the complete knockout (KO) of the MEG3 gene, we used four sgRNAs: Two targeting the 5′-end (GFP reporter) and two targeting the 3′-end (puromycin resistant cassette) of this gene's locus (Fig. 1B). The cells were then co-transduced with a combination of viral particles coding for the different regions of the gene, thus generating four MEG3_KO cell populations designated according to the sgRNAs used: 1+3, 1+4, 2+3 and 2+4.
We observed that different combinations of sgRNAs result in distinct efficiency of genomic editing, although all KO populations showed significant reduction on lncRNA expression (Fig. 1C). The combination 2+4 displayed a striking reduction of around 98% in MEG3 expression when compared to the Hs578T_WT cells. Likewise, the PCR set up to detect MEG3 deletion, revealed a higher rate of genomic edition in this cell population.
To better test our CRISPR/Cas9 protocol, we next aimed to investigate the knockout stability in this cell population. To that end, we collected samples from sequential passage numbers in order to verify the percentage of GFP-positive cells and MEG3 lncRNA expression levels. FACS sorting confirmed that the number of GFP expressing cells remained above 95% over time (Fig. 1D) while MEG3 expression levels was consistently reduced (Fig. 1E). Therefore, we selected the MEG3_KO 2+4 cell population to investigate the phenotypic effects of MEG3 deletion in the Hs578T cell line.
MEG3 knockout cells exhibit increased proliferation and anchorage-independent cell growth
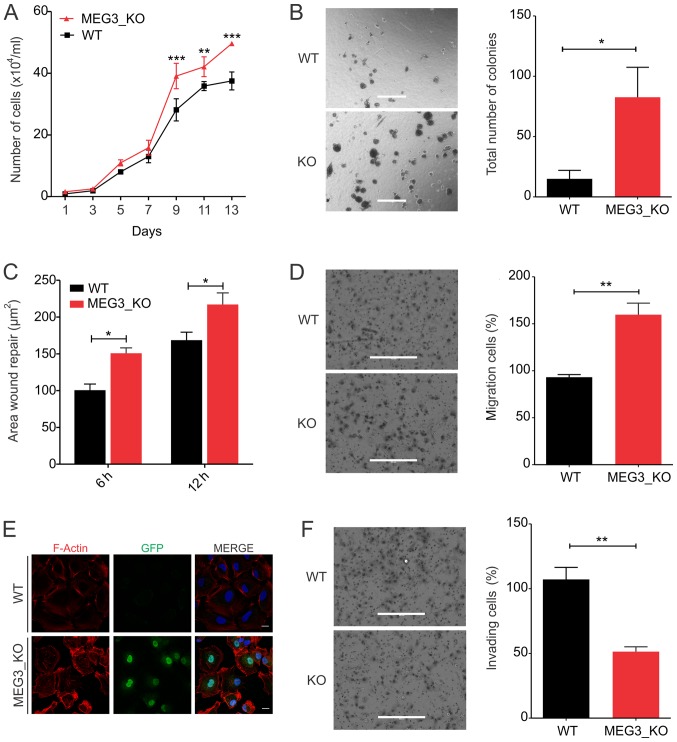
First, we examined the proliferative potential of the MEG3_KO cells, observing that cells display increased proliferative rates upon MEG3 deletion, when compared to the Hs578T_WT cells (Fig. 2A). Next, we evaluated the anchorage-independent cell growth ability in soft agar assay. As shown in Fig. 2B, MEG3_KO cells exhibit significant increase in the ability to form colonies in semi-solid substrate.
Figure 2.
MEG3 knockout leads to increased cell proliferation, anchorage-independent growth and cell migration capacity while cell invasion is reduced. (A) Growth curve of Hs578T_WT and KO cells revealing a significant increase in the proliferation rate of Hs578T cells upon MEG3 deletion by CRISPR/Cas9. **P<0.01; ***P<0.001 vs. WT. (B) Anchorage-independent growth assessment using a soft-agar assay. Representative images of colonies formed in Hs578T_WT and KO cells. Total number of colonies per well was counted. Scale bars, 400 µm. (C) Wound healing assay. The area of migrating cells was measured at different time points (0, 6 and 12 h) using ImageJ software. (D) Transwell migration assay. Representative images showing Hs578T_WT and KO cells that migrated through the membrane. Scale bars, 400 µm. (E) Immunofluorescence of Hs578T_WT and KO cells. GFP (green), F-actin (red) and DAPI (blue). Scale bars, 20 µm. (F) Transwell invasion assay. Representative images showing Hs578T_WT and KO cells that migrated through the membrane. Scale bars, 400 µm. Values are presented as the mean ± SEM for three independent experiments. The data were analyzed using two-way ANOVA or a t-test. *P<0.05 and **P<0.01, as indicated. GFP, green fluorescent protein; KO, knockout; MEG3, maternally expressed gene 3; WT, wild-type.
MEG3 knockout increases cell migration through cytoskeleton changes but reduces the cell invasion capacity
Given the increased anchorage-independent growth capacity of KO_MEG3 cells, we next sought to investigate the effect of MEG3 knockout in cell motility in vitro. Wound healing assays showed that MEG3_KO cells migrate significantly more, when compared to the Hs578T_WT cells (Fig. 2C). We further validated their migration potential using transwell inserts. Consistently, the number of cells which migrated through the membrane was 50% higher in MEG3_KO cells than in Hs578T_WT cells (Fig. 2D). Since cytoskeleton reorganization affects cancer cell motility (32), we investigated whether F-Actin polymerization was changed upon MEG3 deletion. Interestingly, staining of F-Actin showed evident changes in the cytoskeleton organization of KO_MEG3 cells (Fig. 2E).
Next, we used Matrigel-coated inserts to analyze KO_MEG3 cells invasiveness capacity. Surprisingly, we found that these cells invade significantly less, when compared to the WT cells, suggesting a reduced ability of these cells to digest the extracellular matrix and migrate through the membrane (Fig. 2F).
MEG3 regulates the expression of invasiveness factors
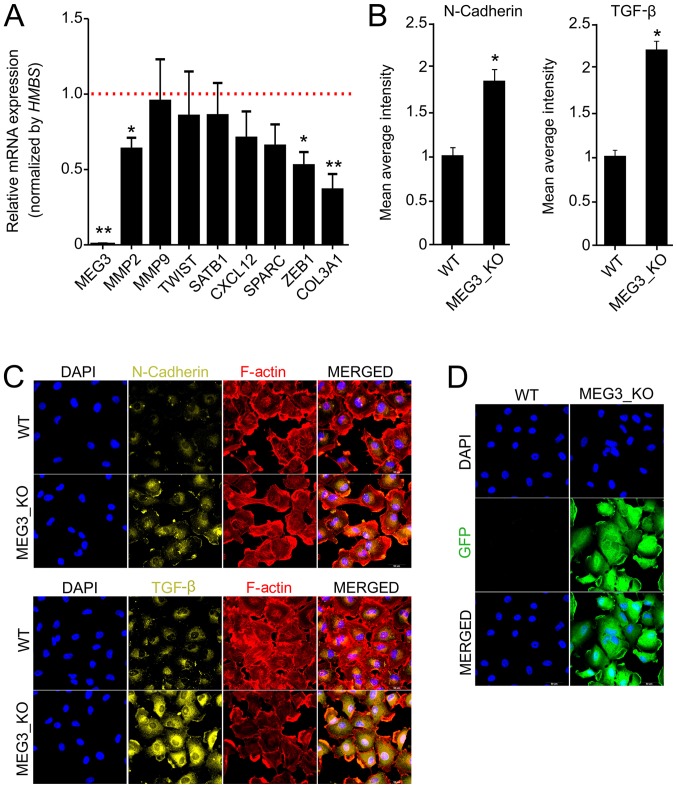
To better understand the effects of MEG3 on cell migration and invasion, we investigated the expression of different markers related to cell motility and invasiveness by RT-qPCR and immunofluorescence. Interestingly, we found significant reduction of MMP2, ZEB1 and COL3A1 in KO cells, when compared to WT (Fig. 3A). Since MEG3 has previously been described to be involved in regulation of EMT factors, we next sought to analyze epithelial-mesenchymal transition (EMT) factors by immunofluorescence (33). We detected an increase, of at least two fold, in TGF-β and N-Cadherin protein expression in KO cells (Fig. 3B-D), but we did not observe significant differences for Vimentin, Fibronectin or E-Cadherin (data not shown).
Figure 3.
MEG3 knockout alters the expression of markers associated with cell invasiveness and epithelial-mesenchymal transition. (A) Relative mRNA quantification was carried out by reverse transcription-quantitative PCR using Hs578T_WT and MEG3_KO (2 + 4) cell lines. Values are presented as the mean ± SEM relative to the expression in the Hs578T_WT cell line, after normalization using HMBS as an endogenous control. The expression levels of the genes in the Hs578T-WT cell line were set as 1, as indicated by the red dotted line. (n=3; test-t). *P<0.05; **P<0.01 vs. Hs578T-WT. (B) Immunofluorescence was performed to quantify the expression of N-cadherin and TGF-β in cells. The fluorescence intensity of each marker was quantified and the data presented are relative to WT cells. A total of 20 images of each well were acquired and the data represent the mean of the images acquired. Values are presented as the mean ± SEM (n=3; test-t) relative to the expression in the Hs578T_WT cell line. *P<0.05 vs. WT. (C) Representative fluorescence microscopy images of each marker. Magnification, ×40. (D) Fluorescence microscopy images revealing GFP expression in MEG3_KO cells. Magnification, ×40. GFP (green), F-actin (red), DAPI (blue), N-cadherin (yellow), TGF-β (yellow). Scale bars, 50 µm. COL3A1, collagen type III αI chain; CXCL12, C-X-C motif chemokine ligand 12; GFP, green fluorescent protein; HMBS, hydroxymethylbilane synthase; KO, knockout; MEG3, maternally expressed gene 3; MMP2, matrix metallopeptidase 2; MMP9, matrix metallopeptidase 9; SATB1, SATB homeobox 1; SPARC, secreted protein acidic and cysteine rich; TGF-β, transforming growth factor β; WT, wild-type; ZEB1, zinc finger E-box binding homeobox 1.
Effect of MEG3 knockout on doxorubicin-induced apoptosis
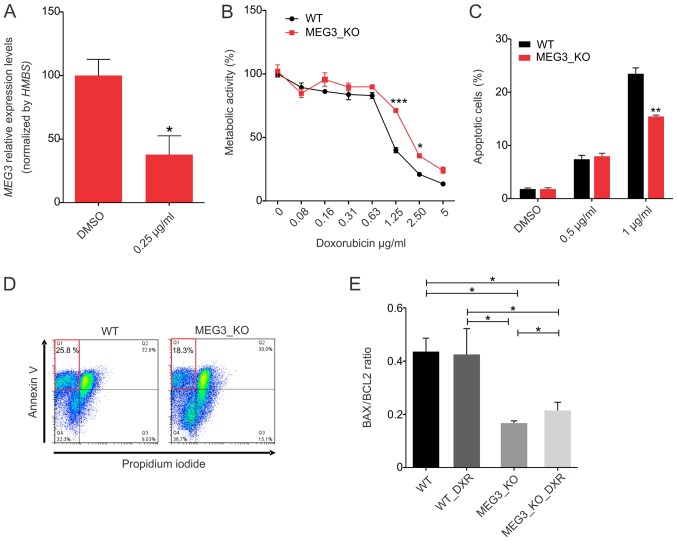
MEG3 has previously been shown to contribute to cisplatin resistance of lung cancer cells (34,35). To verify whether MEG3 is involved in Hs578T cells chemo-resistance, we treated parental cells with a sub-toxic doxorubicin (DXR) concentration (0.25 µg/ml). We found that MEG3 expression is significantly downregulated upon this treatment, suggesting that this lncRNA may be involved with the effect of DXR (Fig. 4A). Moreover, it has been suggested that MEG3 exerts a pro-apoptotic function in different cancer cell lines (12,14,36,37). To extend these findings and determine how MEG3 may affect breast cancer cell sensitivity to chemotherapy, we analyzed the metabolic activity, using the MTT assay, upon cells exposure to different concentrations of DXR for 24 h. We observed that MEG3_KO cells were less sensitive to DXR treatment, when compared to control cells, with significant differences at the concentration of 1.25 µg/ml (Fig. 4B).
Figure 4.
MEG3 knockout decreases the cytotoxicity of DXR in Hs578T cells. (A) Parental Hs578T cells were treated with DMSO or with a sub-lethal concentration of DXR (0.25 µg/ml) for 24 h. Downregulation of endogenous MEG3 expression was demonstrated by reverse transcription-quantitative PCR. Values are presented as the mean ± SEM (n=3) relative to the expression in the DMSO treated cells after normalization using HMBS as an endogenous control (test-t). *P<0.05 vs. DMSO. (B) Hs578T_WT and KO cells were exposed to increasing concentrations of DXR for 24 h. MTT assay demonstrated significant higher metabolic activity in KO cells suggesting increased resistance to drug treatment. Data are presented as the mean ± SEM representative of one representative experiment (two-way ANOVA). (C) Apoptosis assay using PI and Annexin-V analyzed by flow cytometry. The percentage of early apoptotic cells (PI-/Annexin-V+) was quantified and plotted in a bar chart. Data are presented as the mean ± SEM representative of a three technical replicates (test-t). *P<0.05; **P<0.01; ***P<0.001 vs. WT. (D) Representative dot plot graph of Annexin-V/PI stained cells after 24 h treatment with 1 µg/ml doxorubicin. The highlighted red quadrant shows the percentage of apoptotic cells. (E) Flow cytometry quantification of BAX/BCL2 proteins levels in the cells treated and not treated with 1 µg/ml DXR for 24 h. Values are presented as the mean ± SEM of three biological replicates for each cell condition (one-way ANOVA). *P<0.05, as indicated. DXR, doxorubicin; HMBS, hydroxymethylbilane synthase; KO, knockout; MEG3, maternally expressed gene 3; PI, propidium iodide; WT, wild-type.
We further analyzed the rate of appearance of apoptotic cells following treatment with DXR using the Annexin-V binding assay, confirming that this rate is not significantly different when cells are treated for 24 h with 0.5 µg/ml DXR. However, we found a significant resistance in DXR-induced apoptosis when cells were treated with 1 µg/ml (Fig. 4C). The rate of cells undergoing apoptosis was around 28% in Hs578T_WT cells, whereas no more than 18% of MEG3_KO cells were sensitive to this treatment, as represented by the dot plot shown in Fig. 4D.
In parallel, we aimed at investigating the possible relation of BAX (an apoptosis promoter) to BCL2 (an apoptosis inhibitor) ratios with the DXR chemoresistance mediated by MEG3. We found that MEG3_KO cells display a significant reduction in BAX/BCL2 endogenous levels when compared to WT cells, emphasizing the relationship between this lncRNA and the death phenotype (Fig. 4E). We also observed a significant increase of BAX/BCL2 ratio upon treatment in MEG3_KO cell, nevertheless, not sufficient to reach BAX/BCL2 protein levels in WT cells.
Discussion
The versatile role of lncRNAs highlight their relevance and the complexity of the signaling network in which they are involved. Various functional mechanisms have been described for lncRNAs, including recruitment of transcription factors, guiding chromatin modifiers to specific genomic loci, allosteric modulation of transcriptional regulatory proteins, alteration of nuclear domains, modulation of translation and mRNA stability as well as natural competing endogenous RNAs (5,8,9).
MEG3 is a maternally expressed imprinted lncRNA gene, comprised of 10 exons located at chromosome 14q32 (38). Under physiological conditions, it is found to be expressed in many normal tissues and maternal deletion of the MEG3 gene in mice was shown to cause skeletal muscle defects and perinatal death (39). Interestingly, MEG3 gene expression is significantly reduced or completely lost in several human cancers (10,12–14,23) as a result of different events, including promoter hypermethylation, hypermethylation of the intergenic differentially methylated region as well as gene deletion (31).
MEG3 downregulation has been previously suggested to be a potential prognostic factor for breast cancer patients, representing an unfavorable risk factor with significant correlation to patient survival (19–21). Mechanistically, overexpression of MEG3 in MDA-MB-231 and MCF-7 breast cancer cell lines promoted downregulation of AKT signaling, which is pivotal for breast cancer cell growth, invasion and tumor angiogenesis (22). Additionally, MEG3 was shown to act through activation of the p53 pathway, playing a role as a growth suppressor factor (40,41). Importantly, these authors showed that the secondary RNA folding structure of each MEG3 isoform is essential to its function, promoting significant increase in p53 protein levels and stimulating the expression of p53 downstream targets (40).
LncRNAs may regulate cancer cell migration by targeting the Rho/ROCK signaling pathway (42). In our model, we found that MEG3 deletion promoted cell migration, potentially through induction of cytoskeleton rearrangement. Curiously, downregulated MEG3 was previously associated with lymph node metastasis in primary thyroid cancer (43). This study showed that MEG3 suppresses the expression of Rac1 through a specific site in the 3′ UTR. This small signaling GTPase exerts a critical regulatory role driving cell motility by formation of lamellipodia (44), confirming the involvement of MEG3 as a key regulator of F-actin polymerization dynamic.
Here, we used the CRISPR/Cas9 system to analyze the effects of MEG3 deletion in the TNBC Hs578T cell line. Consistently, we found that KO cells display increased proliferation rate and anchorage-independent growth, suggesting greater tumorigenic potential of these cells. In parallel, using Matrigel-coated inserts, we observed a decrease in cell invasion capacity upon MEG3 deletion. Interestingly, integrated analysis has identified MEG3 overexpression and their gene regulation network as an important player in ovarian cancer EMT (45). By genome-wide mapping, it was shown that 73% of MEG3-regulated genes are EMT-linked pathway factors. However, survival analysis showed no significant correlation of MEG3 overexpression with overall patient survival. Another study also described MEG3 as a regulatory RNA for EMT in lung cancer cell lines (33). These authors showed that MEG3 knockdown inhibited EMT by antagonizing TGF-β-dependent changes in the expression of EMT-related genes. Additionally, MEG3 overexpression caused significant changes in the expression of CDH1, E-cadherin, ZEB1, ZEB2, miR-200a, and miR-200c but had no effect in the expression of FN1/fibronectin, vimentin or JARID2. In fact, Mondal and collaborators demonstrated that MEG3 represses the expression of TGF-β pathway genes through formation of RNA-DNA triplex structures (46). Our results show that MEG3 deletion alone leads to increased TGF-β and N-cadherin protein levels and also promotes the reduction of MMP2, ZEB1 and COL3A1 gene expression. These data highlight the role of MEG3 in inhibiting cell migration by regulating the TGF-β pathway however, the exact mechanisms by which MEG3 regulates cell invasion in Hs578T cells and why this effect may differ among different cell lines requires further investigation.
The potential mechanisms involved in chemotherapy drug resistance are largely unclear. Different studies have already suggested that epigenetic alterations, such as histone methylation and acetylation, may play a role in the development of drug resistance (47). In a very elaborate study, Li and collaborators have recently demonstrated that MEG3 promoter methylation correlates with Chronic Myeloid Leukemia tumor stages (48). In that study, MEG3 expression was found to be reduced in advanced phase of the disease, while the expression of methylation related genes, such as DNMT1, DNMT3B, MBD2, MECP2 and HDAC1, was found to be increased, when compared to controls. We speculate that reduced MEG3 expression resulting from DXR treatment may be due to induced promoter hypermethylation. Moreover, it will be interesting to investigate whether the promoter methylation status might be responsible for the discrepant expression levels of MEG3 found in the Hs578T cell line.
MEG3 has been previously implicated in cell death response, regulating intracellular signals triggering apoptosis pathways (12,14,34–37). Thus, MEG3 was shown to inhibit the intrinsic cell survival pathway both in vitro and in vivo by reducing Bcl-2 protein expression, enhancing BAX protein levels and activating caspase-3 in prostate cancer cells (14). Consistently, lung adenocarcinoma patients with lower levels of MEG3 expression displayed worse responses to cisplatin-based chemotherapy (35). MEG3-mediated chemosensitivity enhancement in lung cancer cells was associated with induction of cell-cycle arrest and increased apoptosis, through regulation of p53, β-catenin and survivin, which are target genes of the WNT/β-catenin signaling pathway (34). Here, we demonstrate that Hs578T MEG3_KO cells display reduced ratios of BAX/BCL-2, consistent with a higher resistance to a given apoptotic stimulus. Moreover, MEG3_KO cells showed a slight increase in the ratio of these factors upon treatment with DXR, however, this response is not sufficient to restore the physiological BAX/BCL-2 levels, further supporting a role for MEG3 as an important apoptosis regulator.
In conclusion, lncRNAs are key components involved in diverse biological processes and MEG3 has been previously shown to exert regulatory functions in cell proliferation, apoptosis, migration/invasion and angiogenesis. We confirmed the overall tumorigenic effect of MEG3 deletion by CRISPR/Cas9 system in Hs578T cells, nevertheless, MEG3 was found to be highly expressed in this cell line, suggesting that escape mechanisms are used to counteract its growth suppressor functions, but this need to be further investigated.
Taken together our results indicate that reduced MEG3 expression in breast cancer tissues may contribute to drug resistance in DXR-containing chemotherapy, nevertheless, future therapeutic approaches to promote MEG3 expression/activation should be carefully considered given the ability of MEG3 to modulate EMT factors, which may in turn, promote metastasis. A more conclusive assessment of MEG3 function could benefit from patient derived samples. Also, determining the sensitivity to apoptotic induction and MEG3 expression levels in different models should provide a better understanding of the pathways involved and contribute to decision making regarding patient treatment.
Acknowledgements
The authors would like to thank Dr Ana Marisa Chudzinski-Tavassi (Center of Excellence for New Target Discovery, Butantan Institute, São Paulo, Brazil) for expert advice on the high-content screening assays, and Mr. Eduardo Osório Frare (Center of Excellence for New Target Discovery, Butantan Institute, São Paulo, Brazil) for antibody samples used in the immunofluorescence assays. The authors would also like to thank Ms. Zizi de Mendonça, Ms. Marluce da Cunha Mantovani and Mr. Alan Pereira dos Santos (all Department of Internal Medicine, School of Medicine, University of São Paulo, São Paulo, Brazil) for their technical support.
Funding
The present study was supported by the Brazilian Federal Bank for Social and Economical Development (grant no. 09.2.1066.1), Federal Agency for Higher Education Training (grant no. 88881.068070/2014-01), Brazilian National Council for Research and Development (grant nos. 401430/2013-8, 457601/2013-2, 409960/2013-6, 426896/2016-5 and 465656/2014-5), São Paulo Research State Foundation (grant no. 016/05311-2; Thematic Project), Brazilian Federal Agency for Studies and Projects (grant nos. 01.08.0622.05, 51634-1AD and 01.08.0484.00), Brazilian Science and Technology Ministry, and the Health Ministry. CDP received a post-doctoral fellowship from the Brazilian National Council for Research and Development (grant no. 401828/2012-3). RACM was the recipient of pre-doctoral FAPESP fellowships (nos. 2016/05311-2 and 2013/23271-0). TM received undergraduate training support from the Brazilian National Council for Research and Technology (CNPq; PIBIC no. 147052/2016-5), and MCS is a recipient of a CNPq Productivity Award (no. 3118/2015-6).
Availability of data and materials
The datasets used and/or analyzed during the present study are available from the corresponding author on reasonable request.
Authors' contributions
CDP and RACM developed the concept, designed and performed experiments and wrote the article. TM cloned the CRISPR/Cas9 constructs, generated the KO cell lines and performed functional experiments. HCJF and ACOC performed functional experiments. TFP performed Annexin-V/PI experiments and FACS analysis. MCS contributed to the conception and design of the study, data interpretation, and drafting and critical revision of the manuscript, and approved the manuscript for publication. All authors read and approved the final version of the manuscript.
Ethics approval and consent to participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
References
- 1.Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458:223–227. doi: 10.1038/nature07672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Qureshi IA, Mehler MF. Emerging roles of non-coding RNAs in brain evolution, development, plasticity and disease. Nat Rev Neurosci. 2012;13:528–541. doi: 10.1038/nrn3234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011;12:861–874. doi: 10.1038/nrg3074. [DOI] [PubMed] [Google Scholar]
- 4.Calore F, Lovat F, Garofalo M. Non-coding RNAs and cancer. Int J Mol Sci. 2013;14:17085–17110. doi: 10.3390/ijms140817085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Rinn JL, Chang HY. Genome regulation by long noncoding RNAs. Annu Rev Biochem. 2012;81:145–166. doi: 10.1146/annurev-biochem-051410-092902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhao Y, Li H, Fang S, Kang Y, Wu W, Hao Y, Li Z, Bu D, Sun N, Zhang MQ, Chen R. NONCODE 2016: An informative and valuable data source of long non-coding RNAs. Nucleic Acids Res. 2016;44(D1):D203–D208. doi: 10.1093/nar/gkv1252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Li S, Li B, Zheng Y, Li M, Shi L, Pu X. Exploring functions of long noncoding RNAs across multiple cancers through co-expression network. Sci Rep. 2017;7:754. doi: 10.1038/s41598-017-00856-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Fatica A, Bozzoni I. Long non-coding RNAs: New players in cell differentiation and development. Nat Rev Genet. 2014;15:7–21. doi: 10.1038/nrg3606. [DOI] [PubMed] [Google Scholar]
- 9.Long Y, Wang X, Youmans DT, Cech TR. How do lncRNAs regulate transcription? Sci Adv. 2017;3:eaao2110. doi: 10.1126/sciadv.aao2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhang X, Zhou Y, Mehta KR, Danila DC, Scolavino S, Johnson SR, Klibanski A. A pituitary-derived MEG3 isoform functions as a growth suppressor in tumor cells. J Clin Endocrinol Metab. 2003;88:5119–5126. doi: 10.1210/jc.2003-030222. [DOI] [PubMed] [Google Scholar]
- 11.Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011;10:38. doi: 10.1186/1476-4598-10-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhang J, Yao T, Wang Y, Yu J, Liu Y, Lin Z. Long noncoding RNA MEG3 is downregulated in cervical cancer and affects cell proliferation and apoptosis by regulating miR-21. Cancer Biol Ther. 2016;17:104–113. doi: 10.1080/15384047.2015.1108496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang P, Ren Z, Sun P. Overexpression of the long non-coding RNA MEG3 impairs in vitro glioma cell proliferation. J Cell Biochem. 2012;113:1868–1874. doi: 10.1002/jcb.24055. [DOI] [PubMed] [Google Scholar]
- 14.Luo G, Wang M, Wu X, Tao D, Xiao X, Wang L, Min F, Zeng F, Jiang G. Long non-coding RNA MEG3 inhibits cell proliferation and induces apoptosis in prostate cancer. Cell Physiol Biochem. 2015;37:2209–2220. doi: 10.1159/000438577. [DOI] [PubMed] [Google Scholar]
- 15.Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. doi: 10.3322/caac.21492. [DOI] [PubMed] [Google Scholar]
- 16.Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30. doi: 10.3322/caac.21442. [DOI] [PubMed] [Google Scholar]
- 17.Dai X, Li T, Bai Z, Yang Y, Liu X, Zhan J, Shi B. Breast cancer intrinsic subtype classification, clinical use and future trends. Am J Cancer Res. 2015;5:2929–2943. [PMC free article] [PubMed] [Google Scholar]
- 18.Abramson VG, Lehmann BD, Ballinger TJ, Pietenpol JA. Subtyping of triple-negative breast cancer: Implications for therapy. Cancer. 2015;121:8–16. doi: 10.1002/cncr.28914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang JJ, Guo SH, Jia BQ. Down-regulation of long non-coding RNA MEG3 serves as an unfavorable risk factor for survival of patients with breast cancer. Eur Rev Med Pharmacol Sci. 2016;20:5143–5147. [PubMed] [Google Scholar]
- 20.Tian T, Wang M, Lin S, Guo Y, Dai Z, Liu K, Yang P, Dai C, Zhu Y, Zheng Y, et al. The impact of lncRNA dysregulation on clinicopathology and survival of breast cancer: A systematic review and meta-analysis. Mol Ther Nucleic Acids. 2018;12:359–369. doi: 10.1016/j.omtn.2018.05.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cui X, Yi Q, Jing X, Huang Y, Tian J, Long C, Xiang Z, Liu J, Zhang C, Tan B, et al. Mining prognostic significance of MEG3 in human breast cancer using bioinformatics analysis. Cell Physiol Biochem. 2018;50:41–51. doi: 10.1159/000493956. [DOI] [PubMed] [Google Scholar]
- 22.Zhang CY, Yu MS, Li X, Zhang Z, Han CR, Yan B. Overexpression of long non-coding RNA MEG3 suppresses breast cancer cell proliferation, invasion, and angiogenesis through AKT pathway. Tumour Biol. 2017;39:1010428317701311. doi: 10.1177/1010428317701311. [DOI] [PubMed] [Google Scholar]
- 23.Sun L, Li Y, Yang B. Downregulated long non-coding RNA MEG3 in breast cancer regulates proliferation, migration and invasion by depending on p53's transcriptional activity. Biochem Biophys Res Commun. 2016;478:323–329. doi: 10.1016/j.bbrc.2016.05.031. [DOI] [PubMed] [Google Scholar]
- 24.Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–2308. doi: 10.1038/nprot.2013.143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sánchez-Rivera FJ, Jacks T. Applications of the CRISPR-Cas9 system in cancer biology. Nat Rev Cancer. 2015;15:387–395. doi: 10.1038/nrc3950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Wang H, La Russa M, Qi LS. CRISPR/Cas9 in genome editing and beyond. Annu Rev Biochem. 2016;85:227–264. doi: 10.1146/annurev-biochem-060815-014607. [DOI] [PubMed] [Google Scholar]
- 27.Liu SJ, Horlbeck MA, Cho SW, Birk HS, Malatesta M, He D, Attenello FJ, Villalta JE, Cho MY, Chen Y, et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science. 2017;355(pii):aah7111. doi: 10.1126/science.aah7111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chen W, Zhang G, Li J, Zhang X, Huang S, Xiang S, Hu X, Liu C. CRISPRlnc: A manually curated database of validated sgRNAs for lncRNAs. Nucleic Acids Res. 2019;47:D63–D68. doi: 10.1093/nar/gky904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Goyal A, Myacheva K, Groß M, Klingenberg M, Duran Arqué B, Diederichs S. Challenges of CRISPR/Cas9 applications for long non-coding RNA genes. Nucleic Acids Res. 2017;45:e12. doi: 10.1093/nar/gkw883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001;29:e45. doi: 10.1093/nar/29.9.e45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhou Y, Zhang X, Klibanski A. MEG3 noncoding RNA: A tumor suppressor. J Mol Endocrinol. 2012;48:R45–R53. doi: 10.1530/JME-12-0008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Olson MF, Sahai E. The actin cytoskeleton in cancer cell motility. Clin Exp Metastasis. 2009;26:273–287. doi: 10.1007/s10585-008-9174-2. [DOI] [PubMed] [Google Scholar]
- 33.Terashima M, Tange S, Ishimura A, Suzuki T. MEG3 long noncoding RNA contributes to the epigenetic regulation of epithelial-mesenchymal transition in lung cancer cell lines. J Biol Chem. 2017;292:82–99. doi: 10.1074/jbc.M116.750950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Xia Y, He Z, Liu B, Wang P, Chen Y. Downregulation of Meg3 enhances cisplatin resistance of lung cancer cells through activation of the WNT/β-catenin signaling pathway. Mol Med Rep. 2015;12:4530–4537. doi: 10.3892/mmr.2015.3897. [DOI] [PubMed] [Google Scholar]
- 35.Liu J, Wan L, Lu K, Sun M, Pan X, Zhang P, Lu B, Liu G, Wang Z. The long noncoding RNA MEG3 contributes to cisplatin resistance of human lung adenocarcinoma. PLoS One. 2015;10:e0114586. doi: 10.1371/journal.pone.0114586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhao H, Wang X, Feng X, Li X, Pan L, Liu J, Wang F, Yuan Z, Yang L, Yu J, et al. Long non-coding RNA MEG3 regulates proliferation, apoptosis and autophagy and is associated with prognosis in glioma. J Neurooncol. 2018;140:281–288. doi: 10.1007/s11060-018-2874-9. [DOI] [PubMed] [Google Scholar]
- 37.Shi Y, Lv C, Shi L, Tu G. MEG3 inhibits proliferation and invasion and promotes apoptosis of human osteosarcoma cells. Oncol Lett. 2018;15:1917–1923. doi: 10.3892/ol.2017.7463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Miyoshi N, Wagatsuma H, Wakana S, Shiroishi T, Nomura M, Aisaka K, Kohda T, Surani MA, Kaneko-Ishino T, Ishino F. Identification of an imprinted gene, Meg3/Gtl2 and its human homologue MEG3, first mapped on mouse distal chromosome 12 and human chromosome 14q. Genes Cells. 2000;5:211–220. doi: 10.1046/j.1365-2443.2000.00320.x. [DOI] [PubMed] [Google Scholar]
- 39.Zhou Y, Cheunsuchon P, Nakayama Y, Lawlor MW, Zhong Y, Rice KA, Zhang L, Zhang X, Gordon FE, Lidov HG, et al. Activation of paternally expressed genes and perinatal death caused by deletion of the Gtl2 gene. Development. 2010;137:2643–2652. doi: 10.1242/dev.045724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhou Y, Zhong Y, Wang Y, Zhang X, Batista DL, Gejman R, Ansell PJ, Zhao J, Weng C, Klibanski A. Activation of p53 by MEG3 non-coding RNA. J Biol Chem. 2007;282:24731–24742. doi: 10.1074/jbc.M702029200. [DOI] [PubMed] [Google Scholar]
- 41.Zhang X, Rice K, Wang Y, Chen W, Zhong Y, Nakayama Y, Zhou Y, Klibanski A. Maternally expressed gene 3 (MEG3) noncoding ribonucleic acid: Isoform structure, expression, and functions. Endocrinology. 2010;151:939–947. doi: 10.1210/en.2009-0657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Tang Y, He Y, Zhang P, Wang J, Fan C, Yang L, Xiong F, Zhang S, Gong Z, Nie S, et al. LncRNAs regulate the cytoskeleton and related Rho/ROCK signaling in cancer metastasis. Mol Cancer. 2018;17:77. doi: 10.1186/s12943-018-0825-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wang C, Yan G, Zhang Y, Jia X, Bu P. Long non-coding RNA MEG3 suppresses migration and invasion of thyroid carcinoma by targeting of Rac1. Neoplasma. 2015;62:541–549. doi: 10.4149/neo_2015_065. [DOI] [PubMed] [Google Scholar]
- 44.Parri M, Chiarugi P. Rac and Rho GTPases in cancer cell motility control. Cell Commun Signal. 2010;8:23. doi: 10.1186/1478-811X-8-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Mitra R, Chen X, Greenawalt EJ, Maulik U, Jiang W, Zhao Z, Eischen CM. Decoding critical long non-coding RNA in ovarian cancer epithelial-to-mesenchymal transition. Nat Commun. 2017;8:1604. doi: 10.1038/s41467-017-01781-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Mondal T, Subhash S, Vaid R, Enroth S, Uday S, Reinius B, Mitra S, Mohammed A, James AR, Hoberg E, et al. MEG3 long noncoding RNA regulates the TGF-β pathway genes through formation of RNA-DNA triplex structures. Nat Commun. 2015;6:7743. doi: 10.1038/ncomms8743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Housman G, Byler S, Heerboth S, Lapinska K, Longacre M, Snyder N, Sarkar S. Drug resistance in cancer: An overview. Cancers (Basel) 2014;6:1769–1792. doi: 10.3390/cancers6031769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Li ZY, Yang L, Liu XJ, Wang XZ, Pan YX, Luo JM. The long noncoding RNA MEG3 and its target miR-147 regulate JAK/STAT pathway in advanced chronic myeloid leukemia. EBioMedicine. 2018;34:61–75. doi: 10.1016/j.ebiom.2018.07.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets used and/or analyzed during the present study are available from the corresponding author on reasonable request.