Abstract
Background
Shiga toxin-producing Escherichia coli (STEC) is a leading cause of worldwide food-borne and waterborne infections. Despite an increase in the number of STEC outbreaks, there is a lack of data on prevalence of STEC at the farm level, distribution of serogroups, and virulence factors.
Results
In the present study, a total of 91 (6.16%) STEC strains were isolated from 1477 samples including pig intestines, pig feces, cattle feces, milk, and water from dairy farms. The isolation rates of STEC strains from pig intestines, pig feces, and cattle feces were 7.41% (32/432), 4.38% (21/480), and 9.57% (38/397), respectively. No STEC was isolated from the fresh milk and water samples. By O-serotyping methods, a total of 30 types of O-antigens were determined, and the main types were O100, O97, O91, O149, O26, O92, O102, O157, and O34. Detection of selected virulence genes (stx1, stx2, eae, ehxA, saa) revealed that over 94.51% (86/91) of the isolates carried more than two types of virulence associated genes, and approximately 71.43% (65/91) of the isolates carried both stx1 and stx2, simultaneously. Antimicrobial susceptibility tests showed that most of the STEC isolates were susceptible to ofloxacin and norfloxacin, but showed resistance to tetracycline, kanamycin, trimethoprim-sulfamethoxazole, streptomycin, amoxicillin, and ampicillin. MLST determined 13 categories of sequence types (STs), and ST297 (31.87%; 29/91) was the most dominant clone. This clone displayed a close relationship to virulent strains STEC ST678 (O104: H4). The prevalence of ST297 clones should receive more attentions.
Conclusions
Our preliminary data revealed that a heterogeneous group of STEC is present, but the non-O157 serogroups and some ST clones such as ST297 should receive more attentions.
Keywords: Shiga toxin-producing Escherichia coli, O-serogroups, Virulence genes, Antimicrobial susceptibility, MLST genotypes
Background
Shiga toxin-producing Escherichia coli (STEC) is a significant foodborne pathogen that is capable of causing watery or bloody diarrhea, hemorrhagic colitis, and hemolytic uremic syndrome [1–3]. O (somatic) polysaccharides and H (flagellar) surface antigens form the basis for the serological determination of STEC strains [4, 5]. There are currently more than 100 types of O antigens having been determined from STEC isolates, and several serogroups such as O157, O26, O104, O45, O103, O111, O121, and O145 are commonly associated with severe illness in humans worldwide [2, 4, 6–9]. In China, the first ever severe outbreak of E. coli O157:H7 occurred in Xuzhou, Jiangsu Province, in 1999, which caused the death of 177 people [10]. While limited data on STEC in humans in China are available, both STEC O157 and non-O157 STEC including some predominant serogroups associated with human disease, such as O26, O45, O103, O111, and O121, have been detected and isolated from domestic and wild animals as well as raw meats in different regions [11–14]. A recent study has revealed that the overall prevalence of STEC O157:H7 was 41.3% along the production and supply chain of pork around Hubei Province in Central China, and the prevalence found in slaughter houses, wet- and super-markets were 86.25% (69/80), 53.3% (32/60), and 28.3% (17/60), respectively [13]. These data suggest a big threat to the food safety and even human health in this region.
There are many virulence factors associated with the fitness and pathogenesis of STEC, but Shiga toxin (Stx, also called Vero toxin) is regarded as the most important one [1, 15]. STEC strains mainly produce two Stx types, Stx1 and Stx2, which are further classified into three subtypes for Stx1 (Stx1a, Stx1c, Stx1d) and seven subtypes for Stx2 (Stx2a, Stx2b, Stx2c, Stx2d, Stx2e, Stx2f, Stx2g) [16]. In addition to Stx, the STEC strains also possess many other virulence determinants, including the locus of enterocyte effacement (LEE), hemolysin, STEC autoagglutinating adhesion (Saa), lipopolysaccharide (LPS), outer membrane proteins (OMPs), fimbrial, and peroxidase [15, 17–22].
It is proposed that food-producing animals such as cattle, pigs, chickens are major reservoirs for STEC [23]; and many STEC outbreaks are associated with consumption of meat and other products of food-producing animals contaminated with STEC strains, and/or water contaminated with feces of food-producing animals [24, 25]. Despite an increase in the number of STEC outbreaks, there is a lack of data on prevalence of STEC at the farm level, distribution of serogroups, and virulence factors [2]. Since pork and milk are the common daily food for the Chinese people and Central China, including Hubei, Anhui, Hunan and Henan provinces, is one of main pig rearing and pork producing regions in China, in this study, we performed an isolation, identification and characterization of STEC strains from pigs, cattle, milk and water samples collected from pig and cattle farms in Central China.
Results
Isolation of STEC
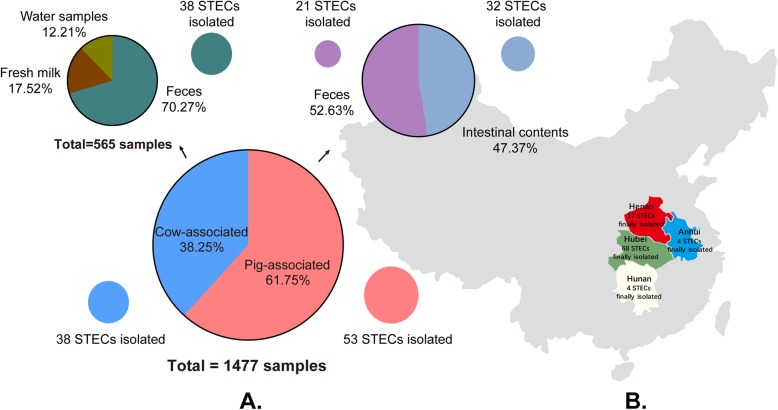
A total of 1477 samples, including 432 samples of intestinal contents from pigs with diarrhea, 480 fecal samples from pigs with diarrhea, 397 fecal samples from cows with diarrhea, 99 samples of fresh milk and 69 water samples from dairy farms, were collected from four provinces of Central China (Hubei, Anhui, Hunan, Henan) for PCR detection of Shiga toxin encoding genes (stx) and STEC isolation. Of the 1477 samples detected, 119 (8.06%) samples were positive for stx1 and/or stx2. STEC strains were isolated from 91 (76.47%) of the 119 stx-positive samples. The isolation rates of STEC strains from pig intestines, pig feces, and cattle feces were 7.41% (32/432), 4.38% (21/480), and 9.57% (38/397), respectively (Fig. 1a, Table 1). However, there were no STEC strains being isolated from the fresh milk and water samples collected (Fig. 1a, Table 1). Biochemical tests showed that all isolates were capable of fermenting glucose, maltose, lactose, and xylose, raffinose, lysine, and ornithine, but were unable to use gluconate, phenylalanine, and citrate.
Fig. 1.
Isolation of STEC from samples collected from Hubei, Henan, Hunan, and Anhui in China. Panel a displays the number of each type of the samples collected for STEC isolation and the number of STEC strains isolated. Panel b shows the places of samples collected and the number of STEC isolated from different provinces in Central China. The authors sincerely acknowledge the Ministry of Natural Resources of the People’s Republic of China for providing the map of China free for public use
Table 1.
Serogroups, virulence factors and sequence types (STs) of the 91 STEC isolates
| ST | No. of isolates | Serogroup | Stx1 | Stx2 | eae | ehxA | saa |
|---|---|---|---|---|---|---|---|
| ST10 | 5 | O34 | + | + | – | + | – |
| O92 | – | + | – | – | – | ||
| O97 | – | + | – | + | – | ||
| O98 | + | + | – | – | + | ||
| O149 | – | + | – | + | – | ||
| ST26 | 2 | O69 | + | + | – | + | + |
| O100 | – | + | – | + | + | ||
| ST29 | 11 | O76 | + | + | – | + | + |
| O92 | + | + | + | – | – | ||
| O97 | + | + | – | + | – | ||
| O97 | + | + | – | + | – | ||
| O100 | + | + | – | + | + | ||
| O100 | + | – | – | + | + | ||
| O100 | + | + | – | + | – | ||
| O102 | – | + | + | – | – | ||
| O102 | – | + | + | – | – | ||
| O102 | + | + | – | + | – | ||
| Nontypable | + | + | – | + | – | ||
| ST101 | 7 | O26 | + | + | – | – | – |
| O97 | + | + | – | + | – | ||
| O100 | – | + | – | – | – | ||
| O100 | + | + | – | + | + | ||
| O100 | + | + | – | + | – | ||
| O102 | + | + | – | + | – | ||
| O149 | – | + | – | + | – | ||
| ST156 | 1 | O64 | + | + | – | + | – |
| ST297 | 29 | O5 | + | + | – | + | – |
| O6 | + | + | – | + | – | ||
| O21 | – | + | – | + | – | ||
| O22 | + | + | – | + | + | ||
| O26 | + | + | – | + | – | ||
| O26 | + | + | – | + | – | ||
| O26 | + | + | – | + | – | ||
| O26 | + | + | – | + | – | ||
| O39 | – | + | – | + | – | ||
| O54 | + | + | – | + | – | ||
| O55 | + | + | – | + | – | ||
| O75 | + | + | – | + | – | ||
| O91 | + | + | – | + | – | ||
| O91 | + | + | – | + | + | ||
| O97 | + | + | – | + | – | ||
| O97 | + | + | – | + | + | ||
| O97 | + | + | – | + | + | ||
| O97 | + | + | – | + | – | ||
| O97 | + | + | – | + | – | ||
| O97 | + | + | – | + | – | ||
| O97 | + | + | – | + | – | ||
| O100 | + | + | – | + | – | ||
| O100 | + | – | – | + | + | ||
| O100 | + | – | – | + | – | ||
| O100 | + | – | – | + | – | ||
| O145 | + | + | – | + | – | ||
| O149 | – | + | – | + | – | ||
| O173 | + | + | – | + | – | ||
| O173 | + | + | – | + | – | ||
| ST542 | 2 | O157 | + | + | + | + | – |
| O157 | + | + | + | + | – | ||
| ST602 | 13 | O34 | + | + | – | + | – |
| O34 | + | + | – | + | – | ||
| O55 | + | + | – | + | – | ||
| O75 | + | + | – | + | – | ||
| O91 | + | – | – | + | + | ||
| O91 | + | – | – | + | – | ||
| O91 | + | – | – | + | – | ||
| O91 | + | – | – | + | – | ||
| O91 | + | – | – | + | + | ||
| O97 | + | + | – | + | – | ||
| O118 | + | + | – | + | + | ||
| O149 | + | + | – | + | – | ||
| Nontypable | + | + | – | + | – | ||
| ST793 | 1 | O3 | – | + | + | – | – |
| ST813 | 4 | O34 | + | + | – | – | – |
| O92 | + | + | – | + | – | ||
| O97 | + | + | – | + | – | ||
| O100 | + | + | – | + | – | ||
| ST1294 | 8 | O91 | + | – | – | + | + |
| O100 | + | + | – | + | + | ||
| O110 | + | + | – | + | + | ||
| O149 | + | + | – | + | – | ||
| Nontypable | + | + | – | + | – | ||
| Nontypable | + | + | – | + | – | ||
| Autoagglutination | – | + | – | – | – | ||
| Autoagglutination | – | + | – | – | – | ||
| ST1623 | 6 | O42 | + | + | – | + | – |
| O54 | + | + | – | + | – | ||
| O78 | – | + | – | + | + | ||
| O92 | + | + | – | + | + | ||
| O92 | + | + | – | + | + | ||
| O149 | – | + | – | – | – | ||
| ST1721 | 2 | O9 | + | + | – | + | + |
| O167 | + | + | – | + | + |
“+”: Positive; “-”: Negative
Serogroups and virulence genotypes
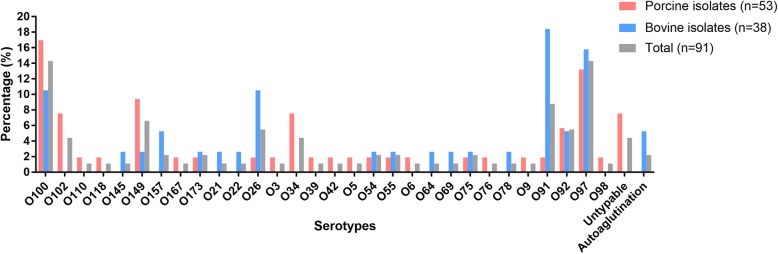
By O-serotyping methods, a total of 30 categories of serogroups were determined for the 93 STEC isolates, and O100, O97, O91, O149, O26, O92, O102, O157, and O34 were the main serogroups (Fig. 2). There were 17 categories of serogroups identified among the bovine isolates (isolates from cow-associated samples), and 25 categories of serogroups among the porcine isolates (isolates from pig-associated samples) (Fig. 2, Table 1). Main serogroups among the porcine isolates were O100, O97, O149, O102, and O34. For bovine isolates, prevalent serogroups were O91, O97, O100, O157, and O26 (Fig. 2). In particularly, serogroup O157 was only detected in STEC strains originated from cows.
Fig. 2.
Distribution of the O-antigens among the STEC isolates. Columns in pink displays the percentage of STECs isolated from the feces and intestinal contents of pigs; Columns in sky blue displays the percentage of STECs isolated from the feces of cows; Columns in gray displays the percentage of the total STECs isolated herein (STECs from pigs plus STECs from cows)
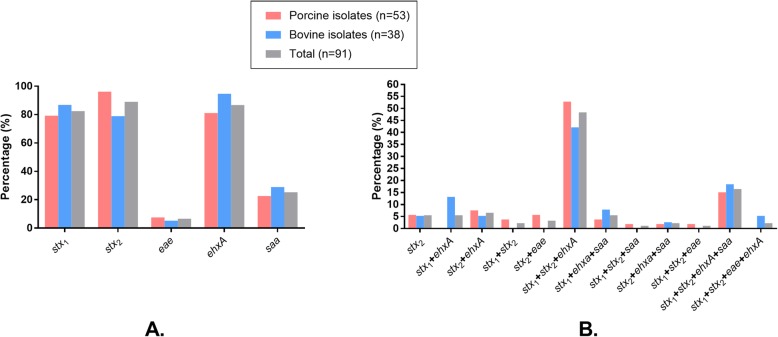
The positive rate of the four virulence associated genes (stx1, stx2, eae, ehxA, saa) among the 91 STEC isolates ranged from 6.59% (eae, 6/91) to 89.01% (stx2, 81/91) (Fig. 3a). The detection rates of the two Stx encoding genes stx1 and stx2 were 82.42% (75/91) and 89.01% (81/91), respectively. Among the stx1-positive isolates, stx1a was the most predominant subtype (78.67%, 59/75), followed by stx1c (17.33%, 13/75) and stx1d (4.00%, 3/75). For the stx2-positive isolates, stx2e was the most predominant subtype (56.79%, 46/81), followed by stx2b (17.28%, 14/81), stx2d (9.88%, 8/81), stx2a (7.14%, 6/81), stx2c (6.17%, 5/81) and stx2g (2.47%, 2/81). Stx1a (100%, 33/33) and stx2b (46.67%, 14/30) were the most predominant stx1 and stx2 subtypes for the bovine isolates while stx1a (61.90%, 26/42) and stx2e (91.20%, 46/51) were the most predominant stx1 and stx2 subtypes for the porcine isolates. Over 94.51% (86/91) of the isolates carried more than two types of virulence associated genes, and approximately 71.43% (65/91) of the isolates carried both stx1 and stx2, simultaneously (Fig. 3b). The percentages of the isolates carrying four types, three types, two types, and one type of the virulence genes detected were 18.68% (17/91), 58.24% (53/91), 17.58% (16/91), and 5.49% (5/91), respectively. Approximately 10.99% (10/91) of the isolates only carried stx1, and 17.58% (16/91) of the isolates only carried stx2.
Fig. 3.
Distribution of main virulence genes (a) and their groups (b) among the STEC isolates. Panel a shows the distribution of main virulence genes while panel b shows the distribution of different groups of virulence genes. Columns in pink displays the percentage of STECs isolated from the feces and intestinal contents of pigs; Columns in sky blue displays the percentage of STECs isolated from the feces of cows; Columns in gray displays the percentage of the total STECs isolated herein (STECs from pigs plus STECs from cows)
In combination the serogroups with the virulence genes, isolates with different serogroups except O157 carried at least one Stx encoding gene (Table 1). In addition, all O100, O149, O26, O34, O91, and O97 isolates (the number of isolates with these serogroups is more than three) were negative to eae, while all O102, O149, O157, O26, and O34 isolates were PCR-negative for the presence of saa (Table 1).
Cytotoxicity
Cytotoxicity tests showed that all isolates positive to stx1 and/or stx2 were capable of making Vero cells rounding and exfoliation. However, no cytopathic effect was observed in the cells inoculated with the isolates negative to both stx1 and stx2.
Antimicrobial susceptibility
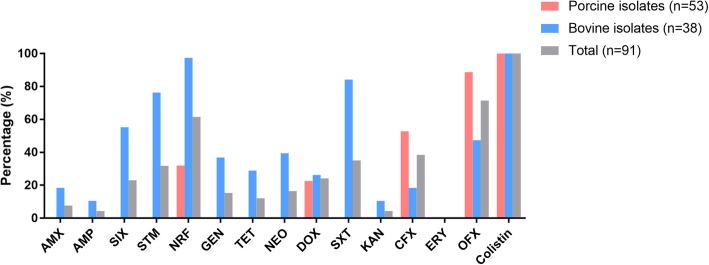
Antimicrobial susceptibility testing results showed that more than 50% of the STEC isolates were sensitive to ofloxacin (71.43%; 65/91), and norfloxacin (61.54%; 56/91). However, less than 10% of the isolates were sensitive to amoxicillin (7.69%; 7/91), ampicillin (4.40% 4/91), and kanamycin (4.40% 4/91). In particularly, all isolates were resistant to erythromycin (100%, 91/91) (Fig. 4). Most of the isolates from cattle feces were sensitive to norfloxacin (97.37%; 37/38), trimethoprim-sulfamethoxazole (84.21%, 32/38), and streptomycin (76.32%; 29/38). All isolates tested herein were sensitive to colistin; the MIC values were determined as ≤1 μg/ml. Approximately half of the bovine isolates were sensitive to sulfafurazole (55.26%; 21/38) and ofloxacin (47.37%; 18/38). For isolates from pigs, 88.68% (47/53) of the isolates were sensitive to ofloxacin. However, there were no isolates from pig intestines and/or feces sensitive to trimethoprim-sulfamethoxazole, streptomycin, sulfafurazole, neomycin, gentamicin, tetracycline, amoxicillin, ampicillin, kanamycin, and cefotaxime (Fig. 4).
Fig. 4.
Antimicrobial susceptibility of the 91 STEC isolates. AMX: amoxicillin; AMP: ampicillin; SIX: sulfafurazole; STM: streptomycin; NRF: norfloxacin; GEN: gentamicin; TET: tetracycline; NEO: neomycin; DOX: doxycycline; SXT: trimethoprim-sulfamethoxazole; KAN: kanamycin; CFX: cefotaxime; ERY: erythromycin; OFX: ofloxacin. Columns in pink displays the percentage of STECs isolated from the feces and intestinal contents of pigs; Columns in sky blue displays the percentage of STECs isolated from the feces of cows; Columns in gray displays the percentage of the total STECs isolated herein (STECs from pigs plus STECs from cows)
MLST genotypes
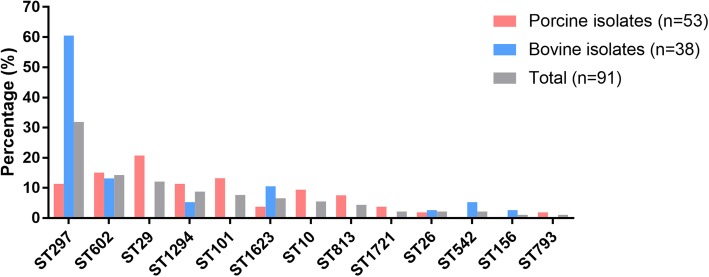
A total of 13 categories of sequence types (STs) were determined among the 91 STEC isolates using the MLST method (Fig. 5). Among these STs, ST297 (31.87%; 29/91) was the most frequent, followed by ST602 (14.29%; 13/91). The other determined STs included ST29 (12.09%; 11/91), ST1294 (8.79%; 8/91), ST101 (7.69%; 7/91), ST1623 (6.59%; 6/91), ST10 (5.49%; 5/91), ST813 (4.39%; 4/91), ST542 (2.20%; 2/91), ST1721 (2.20%; 2/91), ST26 (2.20%; 2/91), ST156 (1.10%; 1/91), and ST793 (1.10%; 1/91). For the 53 porcine isolates, a total of 11 types of STs were determined, and ST29 (20.75%; 11/53), ST602 (15.09%; 8/53), ST101 (13.21%; 7/53), ST297 (11.32%; 6/53), and ST1294 (11.32%; 6/53) were the common STs (Fig. 5). For the 38 bovine isolates, seven types of STs were identified, and ST297 (60.53%; 23/38), ST602 (13.16%; 5/38), ST542 (5.26%; 2/38), and ST1623 (10.53%; 4/38) were commonly present (Fig. 5).
Fig. 5.
Distribution of the sequence types (STs) among the STEC isolates. Columns in pink displays the percentage of STECs isolated from the feces and intestinal contents of pigs; Columns in sky blue displays the percentage of STECs isolated from the feces of cows; Columns in gray displays the percentage of the total STECs isolated herein (STECs from pigs plus STECs from cows)
Phylogenetic analysis showed that the MLST genotypes ST297, ST602, ST101, and ST26 displayed a relationship, and they also showed a close relatedness to the epidemic MLST genotypes ST678 (O104: H4) and ST17 (O45: H2) (Fig. 6). In addition, genotype ST29 was closely related to ST16 (O111: H8), ST21 (O26: H11; O145: H+), and ST723 (O103: H11) (Fig. 6).
Fig. 6.
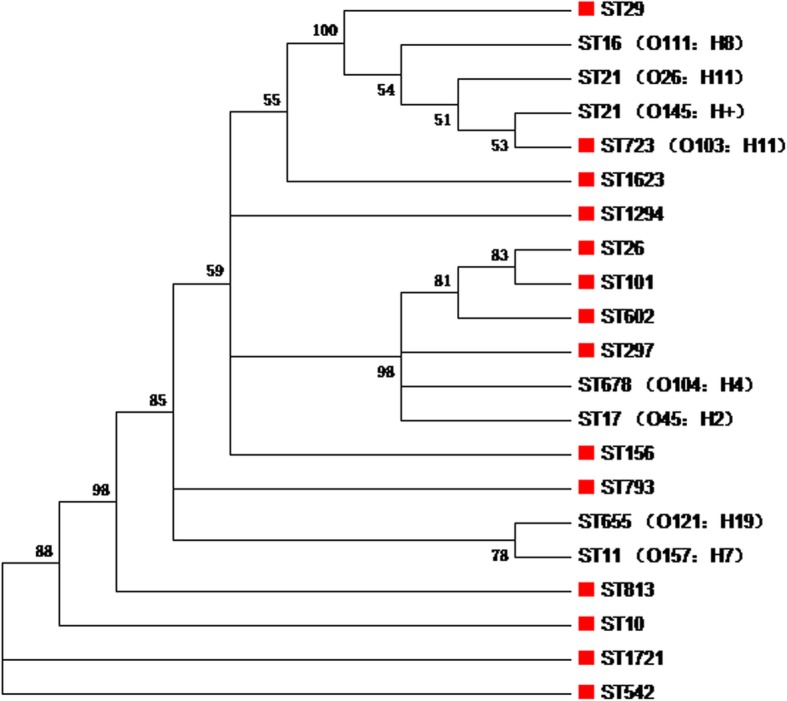
Phylogenetic analysis on different STEC sequence types clones. The tree was constructed based on the MLST data by MEGAX [26], using neighbor-joining algorithm with 1000 bootstrapping
Discussion
STEC is a leading cause of foodborne and waterborne infections worldwide, food-producing animals such as cattle, and pigs are major reservoirs for STEC [23]. Among different kinds of food producing animals, cattle and other ruminants are considered to be the major reservoirs for STEC [23, 27, 28]. STEC strains are more frequently isolated from cattle and other ruminants than from other animals such as pigs, cats, and dogs [23, 29]. In agreement with these suggestions, the rate of STEC isolation from cattle (9.57%) was higher than that from pigs (5.81%). However, the isolate rate of STEC from cattle feces (9.57%) in present study is different from the reports from the other countries [2, 29]. These differences might be explained by differences in feed, seasonal peak, age, or detecting methods [2].
A total of 30 types of O-antigen were determined for the 91 STEC isolates by O-serotyping methods, with the exception of 4 isolates which were not typable (Fig. 2). This might be because there are only 50 types of O antisera are available, and these four strains do not react with the available antisera. Among these 30 categories of O-serogroups, the most frequently occurring serogroups were O100, O97, O91, O149, O26, O92, O102, O157, and O34 (Fig. 2). These serogroups have been isolated from pigs, cattle, sheep, and water in both China and the other countries [30–37]. It has been known that O157 is the most common serogroup that causes human illness in most parts of the world [4]. It has been also reported that cattle are the most common reservoir of E. coli O157 [38]. Corresponding to this suggestion, the O157 serogroup was only determined within the isolates from cattle in the present study (Fig. 2). Although there was no STEC O157 being isolated from pigs in Central China in the present study, a recent study has revealed that the overall prevalence of E. coli O157:H7 in pig farms around Hubei, a province located in Central China, is approximately 12.8% (16/125) [13]. These findings suggest that the prevalence of E. coli O157 in this region is still a problem. In addition to O157, O26 also displayed a high proportion of identification (Fig. 2). It is worthy of note that this type of O-antigen has been declared by the U.S. Department of Agriculture (USDA) as one of the “Big 6” (O26, O45, O103, O111, O121, and O145) non-O157 serogroups that are most commonly associated with severe illness in humans [4]. STEC O26 has been detected and isolated from diarrheal patient in China [11]. It should be noted that another member of the “Big 6”, the O145, was also identified in the present study (Fig. 2). In addition, STEC O149 has been also detected and isolated from diarrheal patient in China [11]. The determination of these non-O157 serogroups represents a great risk on public health and should also receive more attentions.
Virulence genotyping based on the detection of six virulence genes (stx1, stx2, ehxA, eae, and saa) showed that the detection rates of stx1 (82.42%), stx2 (89.01%), and ehxA (86.81%) were higher than those of the other virulence genes (Fig. 3a); most of the STEC isolates possess stx1, stx2, and ehxA simultaneously (Fig. 3b). It is known that both stx1 and stx2 are responsible for encoding the Shiga toxin, which is the most important and common virulence factors of STEC [15]. In particular, the detection rate of stx2 (89.01%) was higher than stx1 (82.42%), and a small proportion of isolates (5.49%, 5/91) only carried stx2 (Fig. 3a and b). It has been reported that stx2 is more often associated with severe disease [39]. Therefore, those strains might be more harmful. Both Stx1 and Stx2 have several subtypes, and some subtypes are more frequently associated with human disease [4]. It has been widely documented that STEC isolates from pigs normally harbor Stx2e subtype [40–42], and in agreement with these studies [40–42], approximately 91.20% of the porcine isolates positive to stx2 determined in the present study harbored this subtype (Stx2e). STEC producing Stx2e is known to be closely associated with edema disease in pigs [43], the high proportion of stx2e detection in STEC isolates from pig intestines and/or feces in this study suggest a big threat to the pig health. Although Stx2e-producing STEC strains are still not proposed as pathogens for humans [43], active actions are still required to control and decrease the prevalence of such strains in pigs in a One Health perspective. In the present study, we also identified several other Stx-subtypes such as Stx1a, Stx1c, Stx1d, Stx2b, Stx2d, Stx2a, Stx2c, and Stx2g. Among these subtypes, stx2a and stx2c are proposed to be associated with high virulence and the ability to cause hemolytic-uremic syndrome (HUS), while stx2d, stx2e, stx1a, and stx1c occurred in milder or asymptomatic infections [43, 44]. The detection of those subtypes in STEC strains from food producing animals such as pigs and cows we detected in this study represents a high risk on public health. It is worthy of note that stx1d, stx2b, stx2g have been also detected in STEC strains from patients in Demark, however, HUS does not develop in these patients [45–47].
In addition to stx1 and stx2, the prevalence of ehxA (86.81%) was also very high, showing a good agreement with previously studies [48–51]. It is worthy of note that ehxA is generally used as a diagnostic indicator because the presence of ehxA is frequently correlated with the Shiga toxin [49, 51]. In agreement with this conclusion, ehxA displayed a high detection rate from the stx-positive STEC strains in the present study (Fig. 3b). In contrast to these genes which have high rates of detection, the detection rates of eae, and saa were relatively low. These results are similar to previously studies [50, 51], suggesting that these virulence genes are not common. However, their presence in particularly the detection of eae should be given a concern. It has been reported that the combination of eae and stx2 has an especial association with the development of HUS and bloody diarrhea [46, 47, 52]. In the present study, all eae-positive STEC strains isolated in Central China were detected to be positive for stx2 (Table 1). The determination of such strains represents a high risk on public health in this region.
The antimicrobial resistance (AMR) of STEC is also a serious problem that the world is now facing. It has been reported that STEC isolates from both humans and food-producing animals displayed resistance most often to tetracycline, kanamycin, trimethoprim-sulfamethoxazole, streptomycin, amoxicillin, and ampicillin [36, 53–56]. In agreement with these studies, a low proportion of STEC isolates from the present study was susceptible to those types of antimicrobials (Fig. 4). These findings suggest a serious profile of AMR in STEC in food-producing animals. While there is a number of articles reporting the colistin resistance prevalence in E. coli [57–59], it is worthy of note that all STEC isolates were sensitive to colistin in the present study.
MLST is also a strategy commonly used for STEC surveillance [36, 60]. In this study, 13 types of STs were determined for the 91 STEC isolates. In particularly, many isolates belonging to different STs possessed the same serogroups (Table 1). These findings are consistent with the findings of other publications [61, 62], suggesting that STEC isolates with the same serogroups might have genotypical diversity. Among the determined STs, ST297 possesses the highest rate of isolation (31.87%) compared to the remaining identified STs (Fig. 5). Interestingly, ST297 is rarely reported in STEC. A previous study determined five ST297 from 75 STEC food strains, with a detection rate of 6.67% [63]. In another study, the detection rate of ST297 among STEC isolates from cattle in Korea was only 4.69% (3/64) [64]. Our results are quite different from these studies, suggesting that the prevalence of the ST in different regions of the world might be different. The ST297 isolates harbored many types of O-antigens, including O26 and O145, the important members of the “Big 6” declared by the USDA [4]. In particular, all O26 isolates recovered in the present study are ST297 clone (Table 1). It has been reported that the STs of STEC O26 associated with a broad spectrum of diseases in Europe are ST29 and/or ST21 [65–70]. In addition, the ST297 clones isolated in this study displayed a close relationship to STEC ST678 (O104: H4) (Fig. 6). It should be noted that the STEC ST678 (O104: H4) isolates have caused the outbreak of human gastroenteritis and human hemolytic-uremic syndrome in Europe [8, 71]. In the next step, we intend to do follow up study to determine the genetic and phenotypical characteristics of these ST297 clones. In addition, the sequence types of the two STEC O157 were determined as ST542 (Table 1). Although the sequence type of STEC O157 is normally determined as ST11 [61, 72], O157 isolates determined as non-ST11 have been also documented elsewhere. For instance, four O157 isolates from the US and/or UK are determined as ST1804 [62]. These findings suggest there might be other STs for STEC O157. In the next step, we will do follow up study to determine the genetic and phenotypical characteristics of these two isolates.
Conclusions
In conclusion, the present study performed an isolation and a characteristic analysis of STEC from pigs and cattle. Our preliminary data revealed that a heterogeneous group of STEC is present, but the non-O157 serogroups and some ST clones such as ST297 should receive more attentions. In the next step, we intend to do a follow up study to correlate the pathogenicity of these STEC with the stx-subtypes as well as the ST clones.
Methods
Sample collection and bacterial isolation
A total of 1477 samples were tested in this study. These samples included intestinal contents from pigs with diarrhea (432 samples), fecal samples from pigs (480 samples) and cows (397 samples) with diarrhea, fresh milk (99 samples), and water samples from dairy farms (69 samples) (Fig. 1a). The 912 pig-associated samples (feces and intestinal contents) were collected from 323 pig farms in Central China (Hubei, Anhui, Hunan, Henan) between 2016 and 2017, while the 565 cow-associated samples (feces, milk, and water samples from dairy farms) were from three dairy farms in different regions Hubei Province in 2017 (Fig. 1a and b). Bacterial isolation was performed following a previously described protocol with some modifications [2]. In brief, each of the samples were mixed in sterilized 0.9% normal saline by vortexing. After a centrifugation at 500×g for 1 min, 500 μL of the supernatant of the mixture was inoculated into 5 mL modified E. coli broth (Nissui, Tokyo, Japan) and incubated at 37 °C for 18~24 h.
After that, genomic DNA was extracted from the cultures by boiling 100 μL aliquot of each incubated broth directly, as described previously [2]. The extracted DNA was evaluated by electrophoresis on a 1% agarose gel and/or using a Nanodrop2000 (Thermo Scientific, Waltham, USA). Presence of the Stx encoding gene stx1 and/or stx2 was determined by PCR assays using the genomic DNA extracted herein as the template and the primers listed in Table 2. PCR reaction was performed in a 25 μL mixture containing 2 μL of the template DNA, 2.5 μL of 10× PCR Buffer (TAKARA, Japan), 2 μL of dNTP (TAKARA, Japan), 0.5 μL of rTaq (TAKARA, Japan), each of the forward and reverse primers 0.5 μL, and 17.0 μL of nucleotide-free water (TAKARA, Japan). Thermocycler conditions used for PCR were 95 °C for 5 min, followed by 30 cycles of denaturation at 94 °C for 30 s, annealing at different temperatures listed in Table 2 for 40 s, and extension at 72 °C for 1 min, with a final extension at 72 °C for 10 min before storage at 4 °C. DNA from STEC O157:H7 strain EDL933 and nucleotide-free water were included as positive and blank controls, respectively. The PCR product was visualized using 1% agarose gel electrophoresis under ultraviolet light.
Table 2.
Primers used in this study
| Primers | Sequences (5′-3′) | Annealing Temp. (°C) | Product size (bp) | Function | References |
|---|---|---|---|---|---|
| Bacterial identification and virulence genotyping | |||||
| stx1-F | ACACTGGATGATCTCAGTGG | 60 | 614 | Amplifying stx1 | Botteldoorn et al., 2003b |
| stx1-R | CTGAATCCCCCTCCATTATG | ||||
| stx2-F | GGCACTGTCTGAAACTGCTCC | 64 | 255 | Amplifying stx2 | Leung et al., 2001d |
| stx2-R | TCGCCAGTTATCTGACATTCTG | ||||
| 16S-F | ATGGCTCAGATTGAACGC | 50 | 1505 | Amplifying 16 SrRNA | REN et al., 2012g |
| 16S-R | CAGGTTCCCCTACGGTTA | ||||
| eae-F | GTGGCGAATACTGGCGAGACT | 64 | 890 | Amplifying eae | Nielsen et al., 2003e |
| eae-R | CCCCATTCTTTTTCACCGTCG | ||||
| ehxA-F | GCATCATCAAGCGTACGTTCC | 60 | 534 | Amplifying ehxA | Bandyopadhyay et al., 2011a |
| ehxA-R | AATGAGCCAAGCTGGTTAAGCT | ||||
| saa-F | CCTCACATCTTCTGCAAATACC | 60 | 1688 | Amplifying saa | Paton et al., 2001f |
| saa-R | GTTGTCGTTCATATTTTACCATCCAATGGACATG | ||||
| MLST genotyping | |||||
| Adk-F1 | TCATCATCTGCACTTTCCGC | 54 | 583 | Amplifying adk | Ding et al., 2012c |
| Adk-R1 | CCAGATCAGCGCGAACTTCA | ||||
| FumC-F1 | TCACAGGTCGCCAGCGCTTC | 54 | 806 | Amplifying fumC | |
| FumC-R1 | GTACGCAGCGAAAAAGATTC | ||||
| GyrB-F1 | TCGGCGACACGGATGACGGC | 60 | 911 | Amplifying gyrB | |
| GyrB-R1 | ATCAGGCCTTCACGCGCATC | ||||
| Icd-F1 | ATGGAAAGTAAAGTAGTTGTT CCGGCACA | 54 | 878 | Amplifying icd | |
| Icd-R1 | GGACGCAGCAGGATCTGTT | ||||
| Mdh-F1 | ATGAAAGTCGCAGTCCTCGGC GCTGCTGGCGG | 60 | 932 | Amplifying mdh | |
| Mdh-R1 | TTAACGAACTCCTGCCCCAGAGCGATATCTTTCTT | ||||
| PurA-F1 | TCGGTAACGGTGTTGTGCTG | 54 | 816 | Amplifying purA | |
| PurA-R1 | CATACGGTAAGCCACGCA GA | ||||
| RecA-F1 | CGCATTCGCTTTACCCTGACC | 58 | 780 | Amplifying recA | |
| RecA-R1 | TCGTCGAAATCTACGGACCGGA | ||||
| Adk-F2 | TCATCATCTGCACTTTCCGC | – | – | adk Sequencing | |
| Adk-R2 | CCAGATCAGCGCGAACTTCA | ||||
| FumC-F2 | TCACAGGTCGCCAGCGCTTC | – | – | fumC Sequencing | |
| FumC-R2 | TCCCGGCAGATAAGCTGTGG | ||||
| GyrB-F2 | TCGGCGACACGGATGACGGC | – | – | gyrB Sequencing | |
| GyrB-R2 | GTCCATGTAGGCGTTCAGGG | ||||
| Icd-F2 | ATGGAAAGTAAAGTAGTTGTTCCGGCACA | – | – | icd Sequencing | |
| Icd-R2 | GGACGCAGCAGGATCTGTT | ||||
| Mdh-F2 | AGCGCGTTCTGTTCAAATGC | – | – | mdh Sequencing | |
| Mdh-R2 | CAGGTTCAGAACTCTCTCTGT | ||||
| PurA-F2 | CGCGCTGATGAAAGAGATGA | – | – | purA Sequencing | |
| PurA-R2 | CATACGGTAAGCCACGCAGA | ||||
| RecA-F2 | ACCTTTGTAGCTGTACCACG | – | – | recA Sequencing | |
| RecA-R2 | TCGTCGAAATCTACGGACCGGA | ||||
aBandyopadhyay S, Mahanti A, Samanta I, Dutta TK, Ghosh MK, Bera AK, Bandyopadhyay S, Bhattacharya D. Virulence repertoire of Shiga toxin-producing Escherichia coli (STEC) and enterotoxigenic Escherichia coli (ETEC) from diarrhoeic lambs of Arunachal Pradesh, India. Trop Anim Health Prod. 2011;43(3):705-10
bBotteldoorn N, Heyndrickx M, Rijpens N, Herman L. Detection and characterization of verotoxigenic Escherichia coli by a VTEC/EHEC multiplex PCR in porcine faeces and pig carcass swabs. Res Microbiol. 2003;154(2):97-104
cDing Y, Tang X, Lu P, Wu B, Xu Z, Liu W, Zhang R, Bei W, Chen H, Tan C. Clonal analysis and virulent traits of pathogenic extraintestinal Escherichia coli isolates from swine in China. BMC Vet Res. 2012;8:140
dLeung PH, Yam WC, Ng WW, Peiris JS. The prevalence and characterization of verotoxin-producing Escherichia coli isolated from cattle and pigs in an abattoir in Hong Kong. Epidemiol Infect. 2001;126(2):173-9
eNielsen EM, Andersen MT. Detection and characterization of verocytotoxin-producing Escherichia coli by automated 5' nuclease PCR assay. J Clin Microbiol. 2003;41(7):2884-93
fPaton AW, Srimanote P, Woodrow MC, Paton JC. Characterization of Saa, a novel autoagglutinating adhesin produced by locus of enterocyte effacement-negative Shiga-toxigenic Escherichia coli strains that are virulent for humans. Infect Immun. 2001;69(11):6999-7009
gREN L, YU X, SONG D, ZHEN K, QIN Y, WANG Y. Isolation, identification and phylogenetic analysis of E. coli from Yaks. China Animal Husbandry & Veterinary Medicine. 2012;39(1): 168-171
In the next step, bacterial cultures positive to at least one of stx1 and stx2 were streak-plated onto sorbitol MacConkey agar (Hangzhou Microbial Reagent CO., LTD, Hangzhou, China), and incubated at 37 °C for 18~24 h. After this stage, the isolates were purified and cultured following the standard methods used for bacterial identification [73]. Presumptive isolates of E. coli were finally confirmed via Galanz staining, biochemical testing, and 16S rRNA amplification and sequencing.
Serotyping and virulence genotyping
O-polysaccharide antigens serogroups of STEC isolates were determined by Slide agglutination test based on the reaction of the bacterial strains against the 50 kinds of O antisera purchased from China Institute of Veterinary Drug Control (Beijing, China). STEC O157:H7 strain EDL933 was used as positive control.
Virulence genotyping was performed by PCR assays amplifying another three virulence associated genes eae, ehxA, and saa of with primers listed in Table 2. The PCR volume and procedure were the same as that used for determining the Stx encoding genes. Positive and blank control samples were included in each set of reactions. The PCR product was visualized using 1% agarose gel electrophoresis under ultraviolet light. The stx subtypes (stx1a, stx1c, stx1d, stx2a, stx2b, stx2c, stx2d, stx2e, stx2f, stx2g) were also determined by PCR assays with primers and reaction procedures described previously [16].
Cytotoxicity
Vero cells (purchased from ATCC) were used to test the cytotoxicity of the STEC strains isolated herein. In brief, isolates were inoculated in Luria-Bertani (LB) broth (Sigma-Aldrich, MO) and shaken at 37 °C for 18~24 h. Bacterial culture were then centrifuged at 20000×g for 40 min, followed by a filtration through a 0.22 μm membrane. Filtrate was inoculated into Vero cells and the cells were incubated at 37 °C for 18~24 h to observe the morphology. Filtrates collected from STEC O157:H7 strain EDL933, E. coli DH5α, cell medium were included as controls.
Antimicrobial susceptibility tests
Antimicrobial susceptibility of the STEC isolates was determined by using the disc diffusion method, following the protocols recommended by Clinical and Laboratory Standards Institute [74]. A total of 14 types of antibiotics including amoxicillin (AMX), ampicillin (AMP), sulfafurazole (SIX), streptomycin (STM), norfloxacin (NRF), gentamicin (GEN), tetracycline (TET), neomycin (NEO), doxycycline (DOX), trimethoprim-sulfamethoxazole (SXT), kanamycin (KAN), cefotaxime (CFX), erythromycin (ERY), and ofloxacin (OFX) were tested. Results were interpreted using the CLSI breakpoints, when available. Resistance to colistin was also tested using broth microdilution method, as recommended by CLSI [74]. Colistin with final concentrations of 1 μg/mL, 2 μg/mL, and 4 μg/mL was made in a 96-well plate in pre-reduced supplemented Mueller-Hinton (MH) broth (Hopebio, Qingdao, China). Interpretation of testing results was based on EUCAST breakpoint (> 2 μg/mL), as the CLSI document (VET01S) does not provide a breakpoint for interpretation of colistin. Each antibiotic was tested with three duplicates. E. coli ATCCR 25922 was used as quality control.
Multilocus sequence typing
Multilocus sequence typing (MLST) was performed using the previously described protocols [60]. Nucleotide sequences of seven housekeeping genes (adk, fumC, gyrB, icd, mdh, purA, and recA) were amplified and sequenced using the primers listed in Table 2. PCR reaction was performed in a 50-μl reaction mixture containing 2 μl of the template DNA, 2 μL of dNTP mixture (TAKARA, Japan), 5 μL of 10 × PCR buffer (TAKARA, Japan), 0.5 μL of rTaq polymerase (TAKARA, Japan), each of the forward and reverse primer 1 μL, and 38.5 μL of nuclease-free water. The reaction was performed under the following standard cycling procedure: an initial denaturation at 95 °C for 5 min, followed by 30 cycles of denaturation at 94 °C for 30 s, annealing at 54–60 °C for 45 s (see Table 2), extension at 72 °C for 1 min, and a final extension at 72 °C for 10 min. The PCR products were initially analyzed by electrophoresis on a 1% agarose gel. Products with the correct size were sequenced at Sangon (Shanghai, China). Nucleotide sequences of the housekeeping genes were submitted to the Escherichia coli MLST Database (http://mlst.warwick.ac.uk/mlst/dbs/Ecoli) to determine the sequence types automatically. Phylogenetic tree was generated based on the MLST data by MEGAX [26], using neighbor-joining algorithm with 1000 bootstrapping.
Acknowledgements
We acknowledge Dr. Shaowen Li at Huazhong Agricultural University College of Animal Science and Veterinary Medicine, Wuhan, China for kindly providing STEC O157:H7 reference strain EDL933.
Abbreviations
- AMP
Ampicillin
- CFX
Cefotaxime
- DOX
Doxycycline
- ERY
Erythromycin
- GEN
Gentamicin
- KAN
Kanamycin
- LEE
The locus of enterocyte effacement
- LPS
Lipopolysaccharide
- MLST
Multilocus sequence typing
- NEO
Neomycin
- NRF
Norfloxacin
- OFX
Ofloxacin
- OMPs
Outer membrane proteins
- Saa
STEC autoagglutinating adhesion
- SIX
Sulfafurazole
- ST
Sequence type
- STEC
Shiga toxin-producing Escherichia coli
- STM
Streptomycin
- Stx
Shiga toxin
- stx
Shiga toxin encoding genes
- SXT
Trimethoprim-sulfamethoxazole
- TET
Tetracycline
Authors’ contributions
ZP, XW, and BW contributed to the conception and design of this work; ZP, WL, ZH, XL, RG, LH, XT, and CT participated in the sample collection, PCR detection and bacterial isolation as well as the laboratory work with the bacterial isolates and the antimicrobial susceptibility testing; ZP, HC, XW, and BW drafted and revised the manuscript. All authors read and approved the final manuscript.
Funding
This work was supported by the National Key Research and Development Program of China (Grant numbers: 2017YFC1600101 and 2017YFC1600103) from the Ministry of Science and Technology of the People’s Republic of China, and the Agricultural Science and Technology Innovation Program of Hubei Province (Grant numbers: 2018skjcx05). Zhong Peng was supported in part by China Postdoctoral Science Foundation (grant number: 2018 M640719). The funders have no role in the study design, data collection and interpretation, or the decision to submit the work for publication.
Availability of data and materials
Not applicable.
Ethics approval and consent to participate
Not applicable. All of the clinical samples used in this study were submitted by veterinarians/or the farm owners to the Huazhong Agricultural University Veterinary Diagnostic Laboratory for routine testing.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Xiangru Wang, Email: wangxr228@mail.hzau.edu.cn.
Bin Wu, Email: wub@mail.hzau.edu.cn.
References
- 1.Kaper JB, Nataro JP, Mobley HL. Pathogenic Escherichia coli. Nat Rev Microbiol. 2004;2:123–140. doi: 10.1038/nrmicro818. [DOI] [PubMed] [Google Scholar]
- 2.Mekata H, Iguchi A, Kawano K, Kirino Y, Kobayashi I, Misawa N. Identification of O serotypes, genotypes, and virulotypes of Shiga toxin-producing Escherichia coli isolates, including non-O157 from beef cattle in Japan. J Food Prot. 2014;77:1269–1274. doi: 10.4315/0362-028X.JFP-13-506. [DOI] [PubMed] [Google Scholar]
- 3.Newell DG, La Ragione RM. Enterohaemorrhagic and other Shiga toxin-producing Escherichia coli (STEC): where are we now regarding diagnostics and control strategies? Transbound Emerg Dis. 2018;65(Suppl 1):49–71. doi: 10.1111/tbed.12789. [DOI] [PubMed] [Google Scholar]
- 4.Croxen MA, Law RJ, Scholz R, Keeney KM, Wlodarska M, Finlay BB. Recent advances in understanding enteric pathogenic Escherichia coli. Clin Microbiol Rev. 2013;26:822–880. doi: 10.1128/CMR.00022-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fratamico PM, DebRoy C, Liu Y, Needleman DS, Baranzoni GM, Feng P. Advances in molecular serotyping and subtyping of Escherichia coli. Front Microbiol. 2016;7:644. doi: 10.3389/fmicb.2016.00644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bielaszewska M, Mellmann A, Zhang W, Kock R, Fruth A, Bauwens A, Peters G, Karch H. Characterisation of the Escherichia coli strain associated with an outbreak of haemolytic uraemic syndrome in Germany, 2011: a microbiological study. Lancet Infect Dis. 2011;11:671–676. doi: 10.1016/S1473-3099(11)70165-7. [DOI] [PubMed] [Google Scholar]
- 7.Corogeanu D, Willmes R, Wolke M, Plum G, Utermohlen O, Kronke M. Therapeutic concentrations of antibiotics inhibit Shiga toxin release from enterohemorrhagic E. coli O104:H4 from the 2011 German outbreak. BMC Microbiol. 2012;12:160. doi: 10.1186/1471-2180-12-160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Frank C, Werber D, Cramer JP, Askar M, Faber M, An der Heiden M, Bernard H, Fruth A, Prager R, Spode A, Wadl M, Zoufaly A, Jordan S, Kemper MJ, Follin P, Muller L, King LA, Rosner B, Buchholz U, Stark K, Krause G. Epidemic profile of Shiga-toxin-producing Escherichia coli O104:H4 outbreak in Germany. N Engl J Med. 2011;365:1771–1780. doi: 10.1056/NEJMoa1106483. [DOI] [PubMed] [Google Scholar]
- 9.Wasey A, Salen P. StatPearls. Treasure Island: StatPearls Publishing StatPearls Publishing LLC; 2018. Escherichia coli (E coli 0157 H7) [PubMed] [Google Scholar]
- 10.Xiong Y, Wang P, Lan R, Ye C, Wang H, Ren J, Jing H, Wang Y, Zhou Z, Bai X, Cui Z, Luo X, Zhao A, Wang Y, Zhang S, Sun H, Wang L, Xu J. A novel Escherichia coli O157:H7 clone causing a major hemolytic uremic syndrome outbreak in China. PLoS One. 2012;7:e36144. doi: 10.1371/journal.pone.0036144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bai X, Hu B, Xu Y, Sun H, Zhao A, Ba P, Fu S, Fan R, Jin Y, Wang H, Guo Q, Xu X, Lu S, Xiong Y. Molecular and phylogenetic characterization of non-O157 Shiga toxin-producing Escherichia coli strains in China. Front Cell Infect Microbiol. 2016;6:143. doi: 10.3389/fcimb.2016.00143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hu B, Kou ZQ, Shao CC, Yin HY, Liu ZD, Xu XH, Fang M, Chen BL, Wei CY, Li GF, Bi ZW. Characteristics and drug resistance of non-O157 Shiga toxin-producing E. coli in animal feces, from Shandong Province. Zhonghua Yu Fang Yi Xue Za Zhi. 2018;52:271–276. doi: 10.3760/cma.j.issn.0253-9624.2018.03.010. [DOI] [PubMed] [Google Scholar]
- 13.Khan SB, Zou G, Xiao R, Cheng Y, Rehman ZU, Ali S, Memon AM, Fahad S, Ahmad I, Zhou R. Prevalence, quantification and isolation of pathogenic Shiga toxin Escherichia coli O157:H7 along the production and supply chain of pork around Hubei Province of China. Microb Pathog. 2018;115:93–99. doi: 10.1016/j.micpath.2017.12.019. [DOI] [PubMed] [Google Scholar]
- 14.Shao CC, Hu B, Bi ZW, Kou ZQ, Fang M, Chen BL, Bi ZQ. Serotype identification and antibiotic susceptibility of Shiga toxin-producing Escherichia coli in the Weishan area in Shandong Province, China. Zhonghua Yu Fang Yi Xue Za Zhi. 2017;51:70–75. doi: 10.3760/cma.j.issn.0253-9624.2017.01.014. [DOI] [PubMed] [Google Scholar]
- 15.Stevens MP, Frankel GM. The locus of enterocyte effacement and associated virulence factors of Enterohemorrhagic Escherichia coli. Microbiol Spectr. 2014;2:Ehec-0007-2013. doi: 10.1128/microbiolspec.EHEC-0007-2013. [DOI] [PubMed] [Google Scholar]
- 16.Scheutz F, Teel LD, Beutin L, Pierard D, Buvens G, Karch H, Mellmann A, Caprioli A, Tozzoli R, Morabito S, Strockbine NA, Melton-Celsa AR, Sanchez M, Persson S, O’Brien AD. Multicenter evaluation of a sequence-based protocol for subtyping Shiga toxins and standardizing Stx nomenclature. J Clin Microbiol. 2012;50:2951–2963. doi: 10.1128/JCM.00860-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Brunder W, Schmidt H, Karch H. KatP, a novel catalase-peroxidase encoded by the large plasmid of enterohaemorrhagic Escherichia coli O157:H7. Microbiology. 1996;142(Pt 11):3305–3315. doi: 10.1099/13500872-142-11-3305. [DOI] [PubMed] [Google Scholar]
- 18.Karch H, Heesemann J, Laufs R, O’Brien AD, Tacket CO, Levine MM. A plasmid of enterohemorrhagic Escherichia coli O157:H7 is required for expression of a new fimbrial antigen and for adhesion to epithelial cells. Infect Immun. 1987;55:455–461. doi: 10.1128/iai.55.2.455-461.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Paton AW, Woodrow MC, Doyle RM, Lanser JA, Paton JC. Molecular characterization of a Shiga toxigenic Escherichia coli O113:H21 strain lacking eae responsible for a cluster of cases of hemolytic-uremic syndrome. J Clin Microbiol. 1999;37:3357–3361. doi: 10.1128/jcm.37.10.3357-3361.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Schmidt H, Beutin L, Karch H. Molecular analysis of the plasmid-encoded hemolysin of Escherichia coli O157:H7 strain EDL 933. Infect Immun. 1995;63:1055–1061. doi: 10.1128/iai.63.3.1055-1061.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Scotland SM, Willshaw GA, Smith HR, Rowe B. Properties of strains of Escherichia coli O26:H11 in relation to their enteropathogenic or enterohemorrhagic classification. J Infect Dis. 1990;162:1069–1074. doi: 10.1093/infdis/162.5.1069. [DOI] [PubMed] [Google Scholar]
- 22.Sherman P, Cockerill F, 3rd, Soni R, Brunton J. Outer membranes are competitive inhibitors of Escherichia coli O157:H7 adherence to epithelial cells. Infect Immun. 1991;59:890–899. doi: 10.1128/iai.59.3.890-899.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Heredia N, Garcia S. Animals as sources of food-borne pathogens: a review. Anim Nutr. 2018;4:250–255. doi: 10.1016/j.aninu.2018.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hilborn ED, Mermin JH, Mshar PA, Hadler JL, Voetsch A, Wojtkunski C, Swartz M, Mshar R, Lambert-Fair MA, Farrar JA, Glynn MK, Slutsker L. A multistate outbreak of Escherichia coli O157:H7 infections associated with consumption of mesclun lettuce. Arch Intern Med. 1999;159:1758–1764. doi: 10.1001/archinte.159.15.1758. [DOI] [PubMed] [Google Scholar]
- 25.Olsen SJ, Miller G, Breuer T, Kennedy M, Higgins C, Walford J, McKee G, Fox K, Bibb W, Mead P. A waterborne outbreak of Escherichia coli O157:H7 infections and hemolytic uremic syndrome: implications for rural water systems. Emerg Infect Dis. 2002;8:370–375. doi: 10.3201/eid0804.000218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35:1547–1549. doi: 10.1093/molbev/msy096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gonzalez Garcia EA. Animal health and foodborne pathogens: enterohaemorrhagic O157:H7 strains and other pathogenic Escherichia coli virotypes (EPEC, ETEC, EIEC, EHEC) Pol J Vet Sci. 2002;5:103–115. [PubMed] [Google Scholar]
- 28.Heuvelink AE, van den Biggelaar FL, de Boer E, Herbes RG, Melchers WJ, Huis JH, Monnens LA. Isolation and characterization of verocytotoxin-producing Escherichia coli O157 strains from Dutch cattle and sheep. J Clin Microbiol. 1998;36:878–882. doi: 10.1128/jcm.36.4.878-882.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Beutin L, Geier D, Steinruck H, Zimmermann S, Scheutz F. Prevalence and some properties of verotoxin (Shiga-like toxin)-producing Escherichia coli in seven different species of healthy domestic animals. J Clin Microbiol. 1993;31:2483–2488. doi: 10.1128/jcm.31.9.2483-2488.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Beutin L, Miko A, Krause G, Pries K, Haby S, Steege K, Albrecht N. Identification of human-pathogenic strains of Shiga toxin-producing Escherichia coli from food by a combination of serotyping and molecular typing of Shiga toxin genes. Appl Environ Microbiol. 2007;73:4769–4775. doi: 10.1128/AEM.00873-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Grant MA, Mogler MA, Harris DL. Comparison of enrichment procedures for Shiga toxin-producing Escherichia coli in wastes from commercial swine farms. J Food Prot. 2009;72:1982–1986. doi: 10.4315/0362-028X-72.9.1982. [DOI] [PubMed] [Google Scholar]
- 32.Houser BA, Donaldson SC, Padte R, Sawant AA, DebRoy C, Jayarao BM. Assessment of phenotypic and genotypic diversity of Escherichia coli shed by healthy lactating dairy cattle. Foodborne Pathog Dis. 2008;5:41–51. doi: 10.1089/fpd.2007.0036. [DOI] [PubMed] [Google Scholar]
- 33.Kaufmann M, Zweifel C, Blanco M, Blanco JE, Blanco J, Beutin L, Stephan R. Escherichia coli O157 and non-O157 Shiga toxin-producing Escherichia coli in fecal samples of finished pigs at slaughter in Switzerland. J Food Prot. 2006;69:260–266. doi: 10.4315/0362-028X-69.2.260. [DOI] [PubMed] [Google Scholar]
- 34.Kobayashi H, Shimada J, Nakazawa M, Morozumi T, Pohjanvirta T, Pelkonen S, Yamamoto K. Prevalence and characteristics of Shiga toxin-producing Escherichia coli from healthy cattle in Japan. Appl Environ Microbiol. 2001;67:484–489. doi: 10.1128/AEM.67.1.484-489.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lienemann T, Pitkanen T, Antikainen J, Molsa E, Miettinen I, Haukka K, Vaara M, Siitonen A. Shiga toxin-producing Escherichia coli O100:H(−): stx2e in drinking water contaminated by waste water in Finland. Curr Microbiol. 2011;62:1239–1244. doi: 10.1007/s00284-010-9832-x. [DOI] [PubMed] [Google Scholar]
- 36.Meng Q, Bai X, Zhao A, Lan R, Du H, Wang T, Shi C, Yuan X, Bai X, Ji S, Jin D, Yu B, Wang Y, Sun H, Liu K, Xu J, Xiong Y. Characterization of Shiga toxin-producing Escherichia coli isolated from healthy pigs in China. BMC Microbiol. 2014;14:5. doi: 10.1186/1471-2180-14-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Sanchez S, Garcia-Sanchez A, Martinez R, Blanco J, Blanco JE, Blanco M, Dahbi G, Mora A, Hermoso de Mendoza J, Alonso JM, Rey J. Detection and characterisation of Shiga toxin-producing Escherichia coli other than Escherichia coli O157:H7 in wild ruminants. Vet J. 2009;180:384–388. doi: 10.1016/j.tvjl.2008.01.011. [DOI] [PubMed] [Google Scholar]
- 38.Heiman KE, Mody RK, Johnson SD, Griffin PM, Gould LH. Escherichia coli O157 outbreaks in the United States, 2003-2012. Emerg Infect Dis. 2015;21:1293–1301. doi: 10.3201/eid2108.141364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Boerlin P, McEwen SA, Boerlin-Petzold F, Wilson JB, Johnson RP, Gyles CL. Associations between virulence factors of Shiga toxin-producing Escherichia coli and disease in humans. J Clin Microbiol. 1999;37:497–503. doi: 10.1128/jcm.37.3.497-503.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Baranzoni GM, Fratamico PM, Gangiredla J, Patel I, Bagi LK, Delannoy S, Fach P, Boccia F, Anastasio A, Pepe T. Characterization of Shiga toxin subtypes and virulence genes in porcine Shiga toxin-producing Escherichia coli. Front Microbiol. 2016;7:574. doi: 10.3389/fmicb.2016.00574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Cha W, Fratamico PM, Ruth LE, Bowman AS, Nolting JM, Manning SD, Funk JA. Prevalence and characteristics of Shiga toxin-producing Escherichia coli in finishing pigs: implications on public health. Int J Food Microbiol. 2018;264:8–15. doi: 10.1016/j.ijfoodmicro.2017.10.017. [DOI] [PubMed] [Google Scholar]
- 42.Tseng M, Fratamico PM, Bagi L, Manzinger D, Funk JA. Shiga toxin-producing E. coli (STEC) in swine: prevalence over the finishing period and characteristics of the STEC isolates. Epidemiol Infect. 2015;143:505–514. doi: 10.1017/S0950268814001095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Scheutz F. Taxonomy meets public health: the case of Shiga toxin-producing Escherichia coli. Microbiol Spectr. 2014;2:1-15. [DOI] [PubMed]
- 44.Orth D, Grif K, Khan AB, Naim A, Dierich MP, Wurzner R. The Shiga toxin genotype rather than the amount of Shiga toxin or the cytotoxicity of Shiga toxin in vitro correlates with the appearance of the hemolytic uremic syndrome. Diagn Microbiol Infect Dis. 2007;59:235–242. doi: 10.1016/j.diagmicrobio.2007.04.013. [DOI] [PubMed] [Google Scholar]
- 45.Bielaszewska M, Friedrich AW, Aldick T, Schurk-Bulgrin R, Karch H. Shiga toxin activatable by intestinal mucus in Escherichia coli isolated from humans: predictor for a severe clinical outcome. Clin Infect Dis. 2006;43:1160–1167. doi: 10.1086/508195. [DOI] [PubMed] [Google Scholar]
- 46.Friedrich AW, Bielaszewska M, Zhang WL, Pulz M, Kuczius T, Ammon A, Karch H. Escherichia coli harboring Shiga toxin 2 gene variants: frequency and association with clinical symptoms. J Infect Dis. 2002;185:74–84. doi: 10.1086/338115. [DOI] [PubMed] [Google Scholar]
- 47.Persson S, Olsen KE, Ethelberg S, Scheutz F. Subtyping method for Escherichia coli Shiga toxin (verocytotoxin) 2 variants and correlations to clinical manifestations. J Clin Microbiol. 2007;45:2020–2024. doi: 10.1128/JCM.02591-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Aidar-Ugrinovich L, Blanco J, Blanco M, Blanco JE, Leomil L, Dahbi G, Mora A, Onuma DL, Silveira WD, Pestana de Castro AF. Serotypes, virulence genes, and intimin types of Shiga toxin-producing Escherichia coli (STEC) and enteropathogenic E. coli (EPEC) isolated from calves in Sao Paulo, Brazil. Int J Food Microbiol. 2007;115:297–306. doi: 10.1016/j.ijfoodmicro.2006.10.046. [DOI] [PubMed] [Google Scholar]
- 49.Beutin L, Montenegro MA, Orskov I, Orskov F, Prada J, Zimmermann S, Stephan R. Close association of verotoxin (Shiga-like toxin) production with enterohemolysin production in strains of Escherichia coli. J Clin Microbiol. 1989;27:2559–2564. doi: 10.1128/jcm.27.11.2559-2564.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Jajarmi M, Imani Fooladi AA, Badouei MA, Ahmadi A. Virulence genes, Shiga toxin subtypes, major O-serogroups, and phylogenetic background of Shiga toxin-producing Escherichia coli strains isolated from cattle in Iran. Microb Pathog. 2017;109:274–279. doi: 10.1016/j.micpath.2017.05.041. [DOI] [PubMed] [Google Scholar]
- 51.Shin SW, Byun JW, Jung M, Shin MK, Yoo HS. Antimicrobial resistance, virulence genes and PFGE-profiling of Escherichia coli isolates from South Korean cattle farms. J Microbiol. 2014;52:785–793. doi: 10.1007/s12275-014-4166-1. [DOI] [PubMed] [Google Scholar]
- 52.Cookson AL, Bennett J, Thomson-Carter F, Attwood GT. Molecular subtyping and genetic analysis of the enterohemolysin gene (ehxA) from Shiga toxin-producing escherichia coli and atypical enteropathogenic E. coli. Appl Environ Microbiol. 2007;73:6360–6369. doi: 10.1128/AEM.00316-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Fratamico PM, Bhagwat AA, Injaian L, Fedorka-Cray PJ. Characterization of Shiga toxin-producing Escherichia coli strains isolated from swine feces. Foodborne Pathog Dis. 2008;5:827–838. doi: 10.1089/fpd.2008.0147. [DOI] [PubMed] [Google Scholar]
- 54.Uemura R, Sueyoshi M, Nagayoshi M, Nagatomo H. Antimicrobial susceptibilities of Shiga toxin-producing Escherichia coli isolates from pigs with edema disease in Japan. Microbiol Immunol. 2003;47:57–61. doi: 10.1111/j.1348-0421.2003.tb02786.x. [DOI] [PubMed] [Google Scholar]
- 55.Wang XM, Liao XP, Liu SG, Zhang WJ, Jiang HX, Zhang MJ, Zhu HQ, Sun Y, Sun J, Li AX, Liu YH. Serotypes, virulence genes, and antimicrobial susceptibility of Escherichia coli isolates from pigs. Foodborne Pathog Dis. 2011;8:687–692. doi: 10.1089/fpd.2010.0739. [DOI] [PubMed] [Google Scholar]
- 56.Zhao S, White DG, Ge B, Ayers S, Friedman S, English L, Wagner D, Gaines S, Meng J. Identification and characterization of integron-mediated antibiotic resistance among Shiga toxin-producing Escherichia coli isolates. Appl Environ Microbiol. 2001;67:1558–1564. doi: 10.1128/AEM.67.4.1558-1564.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Cointe A, Birgy A, Mariani-Kurkdjian P, Liguori S, Courroux C, Blanco J, Delannoy S, Fach P, Loukiadis E, Bidet P, Bonacorsi S. Emerging multidrug-resistant hybrid Pathotype Shiga toxin-producing Escherichia coli O80 and related strains of clonal complex 165, Europe. Emerg Infect Dis. 2018;24:2262–2269. doi: 10.3201/eid2412.180272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Denisuik AJ, Garbutt LA, Golden AR, Adam HJ, Baxter M, Nichol KA, Lagace-Wiens P, Walkty AJ, Karlowsky JA, Hoban DJ, Mulvey MR, Zhanel GG. Antimicrobial-resistant pathogens in Canadian ICUs: results of the CANWARD 2007 to 2016 study. J Antimicrob Chemother. 2018;74:645–653. doi: 10.1093/jac/dky477. [DOI] [PubMed] [Google Scholar]
- 59.Zhang X, Zhang B, Guo Y, Wang J, Zhao P, Liu J, He K. Colistin resistance prevalence in Escherichia coli from domestic animals in intensive breeding farms of Jiangsu Province. Int J Food Microbiol. 2018;291:87–90. doi: 10.1016/j.ijfoodmicro.2018.11.013. [DOI] [PubMed] [Google Scholar]
- 60.Ding Y, Tang X, Lu P, Wu B, Xu Z, Liu W, Zhang R, Bei W, Chen H, Tan C. Clonal analysis and virulent traits of pathogenic extraintestinal Escherichia coli isolates from swine in China. BMC Vet Res. 2012;8:140. doi: 10.1186/1746-6148-8-140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Fierz L, Cernela N, Hauser E, Nuesch-Inderbinen M, Stephan R. Characteristics of Shigatoxin-producing Escherichia coli strains isolated during 2010-2014 from human infections in Switzerland. Front Microbiol. 2017;8:1471. doi: 10.3389/fmicb.2017.01471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Oprea M, Ciontea AS, Militaru M, Dinu S, Cristea D, Usein CR. Molecular typing of Escherichia coli O157 isolates from Romanian human cases. Jpn J Infect Dis. 2018;71:455–461. doi: 10.7883/yoken.JJID.2018.129. [DOI] [PubMed] [Google Scholar]
- 63.Hauser E, Mellmann A, Semmler T, Stoeber H, Wieler LH, Karch H, Kuebler N, Fruth A, Harmsen D, Weniger T, Tietze E, Schmidt H. Phylogenetic and molecular analysis of food-borne Shiga toxin-producing Escherichia coli. Appl Environ Microbiol. 2013;79:2731–2740. doi: 10.1128/AEM.03552-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kang E, Hwang SY, Kwon KH, Kim KY, Kim JH, Park YH. Prevalence and characteristics of Shiga toxin-producing Escherichia coli (STEC) from cattle in Korea between 2010 and 2011. J Vet Sci. 2014;15:369–379. doi: 10.4142/jvs.2014.15.3.369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Bielaszewska M, Mellmann A, Bletz S, Zhang W, Kock R, Kossow A, Prager R, Fruth A, Orth-Holler D, Marejkova M, Morabito S, Caprioli A, Pierard D, Smith G, Jenkins C, Curova K, Karch H. Enterohemorrhagic Escherichia coli O26:H11/H-: a new virulent clone emerges in Europe. Clin Infect Dis. 2013;56:1373–1381. doi: 10.1093/cid/cit055. [DOI] [PubMed] [Google Scholar]
- 66.Chase-Topping ME, Rosser T, Allison LJ, Courcier E, Evans J, McKendrick IJ, Pearce MC, Handel I, Caprioli A, Karch H, Hanson MF, Pollock KG, Locking ME, Woolhouse ME, Matthews L, Low JC, Gally DL. Pathogenic potential to humans of bovine Escherichia coli O26, Scotland. Emerg Infect Dis. 2012;18:439–448. doi: 10.3201/eid1803.111236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Januszkiewicz A, Wolkowicz T, Chrost A, Szych J. Characterization of the Shiga toxin-producing Escherichia coli O26 isolated from human in Poland between 1996 and 2014. Lett Appl Microbiol. 2015;60:605–608. doi: 10.1111/lam.12413. [DOI] [PubMed] [Google Scholar]
- 68.Marejkova M, Blahova K, Janda J, Fruth A, Petras P. Enterohemorrhagic Escherichia coli as causes of hemolytic uremic syndrome in the Czech Republic. PLoS One. 2013;8:e73927. doi: 10.1371/journal.pone.0073927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Zhang WL, Bielaszewska M, Liesegang A, Tschape H, Schmidt H, Bitzan M, Karch H. Molecular characteristics and epidemiological significance of Shiga toxin-producing Escherichia coli O26 strains. J Clin Microbiol. 2000;38:2134–2140. doi: 10.1128/jcm.38.6.2134-2140.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Zweifel C, Cernela N, Stephan R. Detection of the emerging Shiga toxin-producing Escherichia coli O26:H11/H- sequence type 29 (ST29) clone in human patients and healthy cattle in Switzerland. Appl Environ Microbiol. 2013;79:5411–5413. doi: 10.1128/AEM.01728-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Stalb S, Barth SA, Sobotta K, Liebler-Tenorio E, Geue L, Menge C. Pro-inflammatory capacity of Escherichia coli O104:H4 outbreak strain during colonization of intestinal epithelial cells from human and cattle. Int J Med Microbiol. 2018;308:899–911. doi: 10.1016/j.ijmm.2018.06.003. [DOI] [PubMed] [Google Scholar]
- 72.Lee JB, Han D, Lee HT, Wi SM, Park JH, Jo JW, Cho YJ, Hahn TW, Lee S, Kang B, Kwak HS, Kim J, Yoon JW. Pathogenic and phylogenetic characteristics of non-O157 Shiga toxin-producing Escherichia coli isolates from retail meats in South Korea. J Vet Sci. 2018;19:251–259. doi: 10.4142/jvs.2018.19.2.251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.James V, Funke G, Jorgensen JH, Landry ML, Warnock DW. Manual of clinical microbiology. 10. Washington DC: ASM press; 2011. [Google Scholar]
- 74.CLSI. Performance standards for antimicrobial disk and dilution susceptibility tests for bacteria isolated from animals, 3rd ed. CLSI suplement VET01S. Clinical and Laboratory Standards Institute, Wayne 2015.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.