Abstract
Previous genome-scale phylogenetic analyses of Fungi have under sampled taxa from Zoopagales; this order contains many predacious or parasitic genera, and most have never been grown in pure culture. We sequenced the genomes of 4 zoopagalean taxa that are predators of amoebae, nematodes, or rotifers and the genome of one taxon that is a parasite of amoebae using single cell sequencing methods with whole genome amplification. Each genome was a metagenome, which was assembled and binned using multiple techniques to identify the target genomes. We inferred phylogenies with both super matrix and coalescent approaches using 192 conserved proteins mined from the target genomes and performed ancestral state reconstructions to determine the ancestral trophic lifestyle of the clade. Our results indicate that Zoopagales is monophyletic. Ancestral state reconstructions provide moderate support for mycoparasitism being the ancestral state of the clade.
Keywords: Acaulopage tetraceros, Cochlonema odontosperma, Single cell genomics, Stylopage hadra, Zoopage, Zoophagus insidians
1. Introduction
Fungi are an evolutionary lineage within Opisthokonta with remarkable morphological and ecological diversity. Fungi range from single-celled organisms, such as yeasts and chytrids, to multicellular organisms with differentiated tissues and complex life-cycles, such as morels, mushrooms, and rusts. Saprotrophic, mutualistic, predacious, and parasitic lifestyles have arisen multiple times within Fungi. Thus, Fungi represent an ideal group to study the evolution of complex traits and interactions, such as the predacious lifestyle. However, the deep nodes within Fungi are under-sampled and poorly resolved, which limits our ability to reconstruct ancestral states and determine homologies of lifestyles within Fungi (Spatafora et al., 2016; Berbee et al., 2017).
Predation and parasitism are both forms of exploitation (Molles, 2008). Some authors do not distinguish between predator and parasite when discussing fungi (e.g. Duddington, 1956), which is natural because the two categories often overlap and combining them has advantages from an ecological point of view (Raffel et al., 2008). The disadvantage to lumping predators and parasites together is that the different lifestyles present different selective forces on an organism (Lafferty and Kurtis, 2002). For example, selection on a predator will involve means of physically immobilizing prey while selection on a parasite will involve mechanisms of avoiding host immune defenses. Additionally, some authors use nematophagous (e.g., de Freitas Soares et al., 2018), amoebophagous (e.g., Corsaro et al., 2018), or carnivorous (e.g., Thorn and Barron, 1984; Yang et al., 2012) instead of “predator” or “parasite”, but this approach also lumps predators and parasites together. For the purposes of this study, we define predatory fungi to include those taxa that exploit more than one individual as a food source during a life-history stage (Lafferty and Kurtis, 2002). This definition of predatory fungi includes trapping along undifferentiated hyphae as is seen in Stylopage hadra (Drechsler, 1935b), trapping with specialized structures as is seen in Zoophagus insidians (Prowse, 1954), penetration of eggs as is seen in Rhopalomyces elegans (Ellis and Hesseltine, 1962), and/or secretion of immobilizing poisons as is seen in Pleurotus ostreatus (Thorn and Barron, 1984). We define parasitic fungi to include those taxa that exploit a single individual during a life-history stage (Lafferty and Kurtis, 2002). This definition of parasitic fungi includes endoparasites such as Cochlonema odontosperma, ectoparasites such as Amoebophilus simplex, and mycoparasites such as Piptocephalis cylindrospora. Predation has evolved multiple times in Fungi, though the exact number of times it has evolved remains to be determined. Predation has evolved in Ascomycota (Yang et al., 2012; Baral et al., 2018), Basidiomycota (Dreschler, 1935c; Barron and Dierkes, 1977; Saikawa et al., 1994; Thorn et al., 2000; Luo et al., 2004; Zouhar et al., 2013), and Zoopagomycota (White et al., 2006; Corsaro et al., 2018).
Zoopagomycota are an under sampled lineage from the former phylum Zygomycota (hereafter referred to as zygomycetes). Zygomycetes represent the earliest fungal forms that transitioned from zoosporic reproduction (e.g., Chytridiomycota and Blastocladiomycota) to nonflagellated taxa with a primarily terrestrial habit and aerial dispersal (Berbee et al., 2017). Spatafora et al. (2016) re-classified the zygomycetes into two phyla and six subphyla. The main phyla are Mucoromycota, which mainly associate with plants and their byproducts (e.g. mostly plant symbionts and saprotrophs on plant debris), and Zoopagomycota, which are associated with heterotrophic eukaryotes (e.g., mostly animal symbionts, microbe symbionts, mycoparasites, and saprotrophs on animal debris). Zoopagomycota contains three subphyla: the Entomophthoramycotina, the Kickxellomycotina, and the Zoopagomycotina. Among the lineages of Zoopagomycota, Zoopagomycotina is the least studied group. Zoopagomycotina contains one order (Zoopagales) of mycoparasites and taxa that parasitize or prey on amoebae, nematodes, and rotifers. Zoopagales is of great interest because it contains the highest diversity of predacious fungal genera (Kirk et al., 2008). In previous genome-scale phylogenetic analyses, Entomophthoromycotina was represented by six species, Kickxellomycotina was represented by four species, and Zoopagomycotina was only represented by one species (mycoparasite Piptocephalis cylindrospora). Sparse taxon sampling may explain the weak support for relationships among the subphyla within Zoopagomycota and increased sampling may help clarify relationships in this group (Spatafora et al., 2016).
To resolve the phylogeny of Zoopagales and examine the evolution of the predacious lifestyle, we used single cell methods to generate partial genomes for phylogenetic analyses. In contrast to methods that obtain DNA from bulk samples for whole genome or metagenome sequencing, single cell methods obtain DNA from lysed cells with multiple displacement amplification (MDA; Gawad et al., 2016). Multiple displacement amplification uses isothermal priming and extension with [PHI]29 polymerase that has high processivity and a lower error rate than taq (Binga et al., 2008; Gawad et al., 2016), though it can generate chimeras (Gawad et al., 2016). The use of single cell techniques has been developed and used extensively to study prokaryotic microbes (Woyke et al., 2017). These techniques were used to obtain genomes for the microsporidian fungi Amphiamblys sp. and Metchnikovella incurvate with subsequent placement in molecular phylogenies using markers from across the genomes (Mikhailov et al., 2016; Galindo et al., 2018). Ahrendt et al. (2018) developed a high-throughput single cell pipeline targeting unculturable fungi. They showed that combining multiple single cell amplified genomes recovers a larger portion of the genome and that enough markers are recovered to infer robust phylogenies (Ahrendt et al., 2018).
We previously showed that DNA sequence analysis of single to multiple cells can be used to generate rDNA data for members of Zoopagales, but deep relationships were poorly resolved with rDNA markers (Davis et al., in press). For this study, we generated whole genome sequences for Acaulopage tetraceros, Cochlonema odontosperma, Stylopage hadra, Zoopage sp. and Zoophagus insidians using single cell methods with whole genome amplification from a few cells. Using a set of 192 conserved orthologous markers (James et al., 2013; Spatafora et al., 2016), we retrieved amino acid sequences from the annotated genomes and inferred a phylogeny using supermatrix and coalescent approaches. Based on morphology-based taxonomy and previous rDNA phylogenies (Corsaro et al., 2018; Davis et al., in press), we hypothesized that Zoopagales would be monophyletic. We sought to determine the relationships among the sampled genera in Zoopagales, to produce a robust hypothesis of how many times predation has evolved in the clade, and to determine the ancestral trophic state of the clade. Based on their phylogeny, Corsaro et al. (2018) hypothesized the ancestral state of the clade to be predacious (=zooparasitic). Based on our rDNA phylogeny, we hypothesized the ancestral state of the clade to be either mycoparasitism or predation (Davis et al., in press).
2. Materials and methods
2.1. Isolates
Acaulopage tetraceros (Fig. 1A–B) was isolated from soil collected from a farm on Foster Loop Road, Hale County, Alabama, USA (33.087807, −87.632033) and incubated on a 0.1% sea salt water agar plate (0.1 g sea salt, 1L water, 15 g agar; Michel et al., 2015) for 15 days. Cochlonema odontosperma (Fig. 1E–F) was isolated from a specimen of Auricularia auricula collected from a tree stump on Manchester Road, Summit County, Ohio, USA (40.941174, −81.568891) March 2016; thinly sliced pieces were moistened with sterile water and incubated on 0.1% sea salt agar for 1 week. Stylopage hadra (Fig. 1C–D) was isolated from forest soil collected from the E. S. George Reserve, Livingston Co., MI (42.4600, −84.0231) October 2015 and incubated on ¼ strength corn meal agar (0.5 g corn meal infusion solids, 1L water, 15 g agar) inoculated with wild type N2 Caenorhabditis elegans. Zoopage sp. (Fig. 1H) was isolated from soil collected beneath leaf litter in a Populus stand on Chenoweth Dr., Summit County, Ohio, USA (41.000930504, −81.500723362) August 2016, and incubated on a 0.1% sea salt water agar plate for 1 month. Zoophagus insidians (Fig. 1G) was isolated from water collected from Bryant’s Bog, Cheboygan County, Michigan, USA (45.558653, −84.6775977) July 2016 and incubated in a sterile Petri dish with a sesame seed for one week. Voucher slides of C. odontosperma and A. tetraceros were deposited in The University of Michigan Herbarium (MICH 231390 and MICH 231389 respectively).
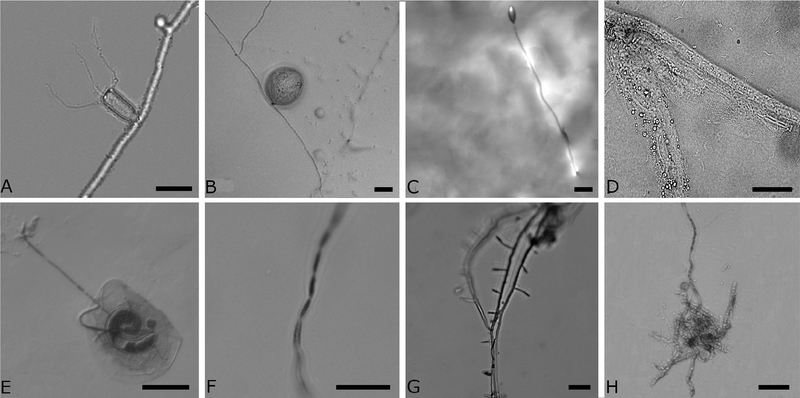
Fig. 1.
Micrographs of the fungi from Zoopagales sequenced. (A) Conidium of Acaulopage tetraceros imaged at 40× magnification. Scale bar is 25 μm. (B) Amoeba trapped on a hypha of A. tetraceros imaged at 20× magnification. Scale bar is 25 μm. (C) Conidium and conidiophore of Stylopage hadra imaged with a stereoscope. Scale bar is 25 μm. (D) Partially digested nematode captured on a S. hadra hypha. Scale bar is 50 μm. (E) Coiled thallus of Cochlonema odontosperma inside of an amoeba with a chain of conidia emerging. Imaged with differential interference contrast microscopy at 40× magnification. Scale bar is 50 μm. (F) Conidia of Cochlonema odontosperma. Scale bar is 25 μm. (G) Hypha of Zoophagus insidians with peg traps imaged at 20×. Scale bar is 25 μm. (H) Partially digested amoeba trapped on a hypha of Zoopage sp imaged at 40×. Scale bar is 25 μm.
2.2. Multiple displacement amplification & polymerase chain reaction
We used a single cell approach (Gawad et al., 2016) to obtain genomes for these fungi by lysing and performing whole genome amplification of target taxa using the Qiagen REPLI-g Single Cell kit (Qiagen, Maryland, USA). We used the manufacturer’s protocol except with half-volume reactions. The laminar flow hood and workstations were sterilized with a 70% ethanol/50% bleach solution and exposure to UV light for 15mins prior to use. PBS solution (provided in the kit), sterile water, pipette tips, pipettes, 1.5 mL microfuge tubes, 0.2 mL PCR tubes, and racks were sterilized with exposure to UV light for 2hrs in a UV Stratalinker 2400. Under laminar flow, 2 μL PBS was aliquoted into PCR tubes. A portion of each fungus (1 hypha of Zoophagus insidians; 10 spores of Acaulopage tetraceros; 5 spore chains of Zoopage sp; 10 spores of Stylopage hadra; and 2 thalli of Cochlonema odontosperma) was transferred into the 2 μL PBS solution using a dental file (Henry Schein Inc., Melville, New York, USA). We then added 1.5 μL of the lysis buffer and the tubes were incubated in a thermocycler at 65C for 10mins. To neutralize the reaction, 1.5 μL of the stop buffer was added. To each tube, 20 μL of the MDA master mix was added, and tubes were incubated on a thermocycler at 30C for 6 hrs. To maximize the amount of the genome recovered, two to six replicates were made for each fungus (Ahrendt et al., 2018). Successful amplification was confirmed via gel electrophoresis.
Successful MDA reactions were diluted 1:100 for quality control PCR. To confirm that the target fungus was amplified in each reaction, the eukaryotic small subunit of the ribosomal DNA (18S) was amplified, sequenced, and assembled as previously described (Davis et al., in press). To check levels of bacterial contamination, the prokaryotic small subunit of the ribosomal DNA (16S) was amplified and sequenced using the BSF/BSR primer pair (Weisburg et al., 1991) under the same PCR conditions as the 18S sequences. To check the integrity of the target genome, the single copy gene EF1-α was amplified and sequenced using the EF1a1F/EF1a1R primer pair (James et al., 2006).
2.3. Library preparation & sequencing
Each MDA reaction was prepared as a separate library with an Illumina Nextera XT Library Prep kit according to the manufacturer’s instructions, except using 2.5 ng of input DNA instead of 1 ng. For Acaulopage tetraceros, Stylopage hadra, Zoopage sp., and Zoophagus insidians, libraries were sequenced on an Illumina HiSeq-2500 whereas Cochlonema odontosperma libraries were sequenced on an Illumina NextSeq 500.
2.4. Assembly, contig binning, and protein prediction
Preliminary quality control of the raw reads was performed with fastqc 0.11.5 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/), which revealed the presence of Nextera transposase contamination in some reads. Raw reads were trimmed with trimmomatic 0.36 (Bolger et al., 2014) with palindrome parameters and passed through fastqc a second time to ensure the transposase sequences were removed. The libraries for each individual species were co-assembled in spades 3.10.0 (Bankevich et al., 2012). Contigs less than 1000 bp were removed using a publicly available script (https://github.com/elyons/BioInfoUtility).
Because the assemblies were likely to be metagenomes, we used three approaches to metagenomic contig binning to sort the target genome from potential contamination. Each assembly was binned using ESOM 1.1 (http://databionic-esom.sourceforge.net/), GC-coverage blob plots (Kumar et al., 2013), and the combination of relative synonymous codon usage (RSCU) and ClaMS (Pati et al., 2011; Mikhailov et al., 2016). The final genome only included contigs if at least two methods identified them as part of the target genome. This selection process has since been developed into a stand-alone pipeline (https://github.com/amsesk/scgid). Briefly, for the ESOM analysis, each assembly was segmented into 3000 bp windows and tetramer frequencies calculated for each window using the print_tetramer_freqs_deg_filter_-esom.pl perl script (Dick et al., 2009). The frequencies were used to train ESOM to generate a topological map of the genomes present in the assembly. We overlaid mega-blast-assigned taxonomy and then manually selected the area corresponding to the target genome. The resulting file was used to obtain the corresponding contigs from the assembly using the getClassFasta.pl perl script (Dick et al., 2009). In order to generate GC-coverage blob plots, proteins on each contig in the assembly were predicted using Augustus 3.2.3 (Stanke and Waack, 2003; Keller et al., 2011). Taxonomic identity was assigned to each contig based on the top blastp hit of its predicted proteins against an amended version of the Uniprot swiss-prot database, which was expanded with a set of draft zygomycete and chytrid proteomes, at a relaxed e-value threshold of 1e−5. Next GC content was plotted against coverage for each contig in R 3.4.1 (https://www.r-project.org/), and these points annotated with taxonomy. Contigs with a blastp-assigned taxonomy were either included or excluded based on that taxonomy (e.g., include Fungi and other eukaryotes but exclude Bacteria). To make decisions about the inclusion/exclusion of unidentified contigs, contigs identified as Fungi were used to draw GC-coverage windows encompassing a best estimate of target-genome contig spread in GC-coverage space (Table S1). Then all unidentified contigs that fell within the selection window were indiscriminately included in the blob-derived final assembly. To select contigs based on RSCU (Mikhailov et al., 2016), the RSCU index for each contig was calculated using the software package GCUA (McInerney, 1998). The resulting RSCU values were then used to construct a distance matrix and infer neighbor-joining trees in R. Taxonomy according to blastn searches of the NCBI nt database was overlaid on leaves and high-confidence target clades were selected for each assembly. These clades were used to train ClaMS (Pati et al., 2011). Using 2-mer frequencies and a Pearson’s distance cutoff of 0.1, ClaMS determined whether each contig in the assembly had similar sequence signature to those in the training set. These similar contigs constituted the final RSCU-derived genome. As noted above, the consensus assembly determined by majority rule: the draft genome of each fungus contained only those contigs identified by two or more of the methods. Statistics for the consensus genomes were determined with assemblathon_stats.pl (Earl et al., 2011). Completeness was estimated with cegma 2.5 (Parra et al., 2007). Proteins were predicted with Maker 2.3.8 (Holt and Yandell, 2011) with Rhizopus oryzae as the augustus model and keep_preds set to 1.
2.5. Genome-scale phylogenetic analyses
We retrieved protein sequences for additional taxa from NCBI and the MycoCosm portal of the Joint Genome Institute (JGI) for use in the genome-scale phylogenetic analyses (Table 1). We followed the Spatafora et al. (2016) pipeline, using their hmmer models for 192 markers and several custom scripts from their GitHub repository (https://github.com/zygolife/Phylogenomics). Hmmer 3.1b2 (hmmer.org) was used to screen the genomes for the corresponding markers, and custom scripts from Spatafora et al. (2016) were used to assemble the identified sequences into fasta files. We used hmmalign to align the markers and a combination of default settings in easel and trimAl 1.2 (Capella-Gutierrez et al., 2009) with settings determined automatically to trim the alignments. Gene trees were inferred with RAxML 8.2.8 (Stamatakis, 2006; Stamatakis et al., 2008) with the model of evolution determined automatically and bootstrapped with 100 replicates. Individual gene trees were screened manually for paralogs and contamination. Paralogs were suspected if ingroup RAxML taxa were on long branches and/or grouped with the outgroup. Sequences suspected of paralogy were checked against NCBI’s nr database with blastp; if the top match was to a non-orthologous gene, then the suspect amino acid sequence was considered a paralog. Orthologs were identified by manually screening the hmmer output for potential orthologs and comparing their blastp identification to those of the outgroup taxa Capsaspora owczarzaki and Drosophila melanogaster. Contamination was suspected if a taxon was misplaced (e.g. Pandora formicae or Zoophagus insidians grouping with Drosophila melanogaster) with 100% bootstrap support. Contaminated sequences were replaced when possible with the correct ortholog identified with the same procedure described above for paralogs. Once all paralogs and contaminants were removed, new gene trees were inferred and screened. The markers were concatenated using custom scripts from Spatafora et al. (2016), and the final tree was inferred with RAxML 8.2.8 using 20 searches starting from distinct randomized maximum parsimony trees (Zhou et al., 2018) with the model of evolution determined automatically and bootstrapped using 500 pseudoreplicates. A greedy consensus tree was constructed with ASTRAL 5.6.1 (Mirarab et al., 2014; Zhang et al., 2017) using the individual gene trees under default settings. Individual gene trees were also used to calculate Internode Certainty (IC/ICA) and Tree Certainty (TC/TCA) for both the ML and ASTRAL trees using the -f i flag in RAxML (Salichos et al., 2014). Since the gene tree set contained both comprehensive and partial trees, the probabilistic adjustment scheme was used (Kobert et al., 2016). Since our gene tree set contained mostly partial gene trees, we used QuartetScores (Zhou et al., 2017) to calculate Quadripartition Internode Certainty (QP-IC), Lowest Quartet Internode Certainty (LQ-IC), and Extended Quadripartition Internode Certainty (EQP-IC) for both the ASTRAL and ML trees.
Table 1.
List of taxa and genome sources.
| Species | GenBank accession/web portal |
|---|---|
| Acaulopage tetraceros | QZWU00000000 |
| Allomyces macrogynus | ACDU00000000 |
| Arthrobotrys oligospora | ADOT00000000 |
| Basidiobolus heterosporus | JNE000000000 |
| Basidiobolus meristosporus | http://genome.jgi.doe.gov/Basme2finSC |
| Capsaspora owczarzaki | ACFS00000000 |
| Catenaria anguillulae | http://genome.jgi.doe.gov/Catan1 |
| Cochlonema odontosperma | QZWT00000000 |
| Coemansia reverse | JZJC00000000 |
| Conidiobolus coronatum | JXYT00000000 |
| Conidiobolus thromboides | http://genome.jgi.doe.gov/Conthl |
| Coprinopsis cincerea | AACS00000000 |
| Dimargaris cristalligena | https://genome.jgi.doe.gov/DimcrSCl/DimcrSCl |
| Drosophilia melanogaster | http://flybase.org |
| Gonopodya prolifera | LSZK00000000 |
| Hesseltinella vesiculosa | http://genome.jgi.doe.gov/Hesve2finisherSC |
| Lichtheimia hyalospora | http://genome.jgi.doe.gov/Lichyl |
| Linderina pennispora | http://genome.jgi.doe.gov/Linpel |
| Martensiomyces pterosporus | http://genome.jgi.doe.gov/Marptl |
| Monosiga brevicolis | ABFJ00000000 |
| Mortierella elongata | http://genome.jgi.doe.gov/Morel2 |
| Neurospora crassa | AABX00000000 |
| Pandora formicae | GCRV00000000 |
| Phycomyces blakesleeanus | http://genome.jgi.doe.gov/Phybl2 |
| Piptocephalis cylindrospora | http://genome.jgi.doe.gov/Pipcy2/Pipcy2.home.html |
| Ramicandelaber brevisporus | http://genome.jgi.doe.gov/Rambrl |
| Rhizoclosmatium globosum | https://genome.jgi.doe.gov/Rhihy1/Rhihy1 |
| Rhizophagus irregularis | JARB00000000 |
| Rhizopus delemar | AACW00000000 |
| Rozella allomycis | ATJD00000000 |
| Saccharomyces cerevisiae | http://yeastgenome.org |
| Schizosaccharomyces pombe | http://www.pombase.org |
| Smittium culicis | GCA_001970855 |
| Smittium mucronatum | GCA_001953115 |
| Spizellomyces punctatus | ACOE00000000 |
| Stylopage hadra | |
| Syncephalis fuscata | https://genome.jgi.doe.gov/Synfusl |
| Syncephalis plumigaleata | https://genome.jgi.doe.gov/Synplul |
| Syncephalis pseudoplumigaleata | https://genome.jgi.doe.gov/Synpsl |
| Thamnocephalis spaerospora | https://genome.jgi.doe.gov/Thasp1 |
| Umbelopsis ramanniana | http://genome.jgi.doe.gov/Umbra1 |
| Ustilago maydis | AACP00000000 |
| Zancudomyces culisetae | GCA_001969505 |
| Zoopage sp. | QZWS00000000 |
| Zoophagus insidians | QZWR00000000 |
| Zoophthora radicans | http://genome.jgi.doe.gov/ZooradStandDraft.FD/ |
To reconstruct ancestral trophic lifestyles, the ML and ASTRAL trees were analyzed in Mesquite 3.40 (Maddison and Maddison, 2018). Taxa were coded according to their trophic lifestyle: saprobe, multitrophic, parasite of microorganisms, predator of microorganisms, mycoparasite, insect parasite, insect commensal, and plant mutualist. Definitions of these trophic lifestyles were generated based on descriptions in Alexopoulos et al. (1996). We considered a taxon a saprobe if it relied on dead and/or decaying biomass for energy and nutrients. A taxon was considered a parasite of microorganisms if an individual infected a host through spores and grew internally (i.e., endoparasite) or externally (i.e., ectoparasite). A taxon was considered a predator of microorganisms if it used multiple individuals as a food source. Mycoparasites were considered those fungi that parasitize other fungi. Insect parasites were those fungi that grow in or on insect hosts and have a negative effect to the host, and insect commensals were those fungi that grow in or on insect hosts and have no apparent effect on the host. Finally, multitrophic taxa were those taxa that are capable of getting nutrients and energy through two or more trophic lifestyles. Examples of multitrophic fungi are Conidiobolus coronatus and Catenaria anguillulae: both of these fungi are parasites that can be grown as saprobes in axenic culture. Ancestral states were reconstructed using both parsimony and maximum likelihood. For the maximum likelihood analysis, the stochchar module (Maddison and Maddison, 2006) was used with the one-parameter Markov k-state model, a generalized Jukes-Cantor model (Lewis, 2001). The judgement of the best character state(s) was made using a decision threshold of 2 log likelihoods with the character state(s) with the lower likelihood being rejected (Maddison and Maddison, 2006; Ekman et al., 2008). For the ancestor of the Zoopagales, we report the negative log likelihood values and the probability (proportional relative likelihood) for the character states passing the decision threshold (Maddison and Maddison, 2006; Reyes et al., 2018). For ease of interpretation, following Reyes et al. (2018), we classify the proportional relative likelihoods into categories of support. Character states with proportional relative likelihood values ≥ 0.90 were considered to receive strong support; values between 0.61 and 0.89 were considered as moderately supported; and, values ≤ 0.60 were considered weakly supported (Reyes et al., 2018). The ancestral state for a node was only considered resolved if a single character state both passed the decision threshold and received strong support.
2.6. Data accessibility
Assemblies and raw reads were deposited in NCBI’s databases under Bioproject PRJNA451036. Scripts and representative output files are available on GitHub (https://github.com/wjdavis90/ZyGoLife/tree/master/phylogenomics). The concatenated alignment, ML tree, ASTRAL tree, and trophic character matrix are deposited in TreeBase (http://purl.org/phylo/treebase/phylows/study/TB2:S22698).
3. Results
3.1. Quality control of MDA genomic libraries
Three libraries of Acaulopage tetraceros (Fig. 1A–B) were successfully amplified by MDA. Sanger sequencing of 18S (Genbank: MG920179) and EF1-α sequences (GenBank: MH923231) confirmed the presence of the fungus in each library. 16S sequences (GenBank: MH791043) were clean and identified as Chitinophaga sp. Three libraries of Cochlonema odontosperma (Fig. 1E–F) were successfully amplified. 18S (GenBank: MG920177) and EF1-α (GenBank: MH923229) confirmed the presence of the target fungus in two libraries; in the third, the sequences were identified as Mortierella sp., and the library was discarded. 16S did not return clean sequences. Five libraries of Stylopage hadra (Fig. 1C–D) were successfully amplified, and the presence of the fungus was confirmed with 18S (GenBank: MG920178) and EF1-α (GenBank: MH708607). Four out of five 16S sequences from the S. hadra libraries were clean and identified as Mycoavidus sp. (GenBank: MH536851). Three libraries of Zoopage sp. (Fig. 1H) were successfully amplified, 18S (GenBank: MG920182) and EF1-α (GenBank: MH923232) were identified as the target fungus, and 16S did not return clean sequences. Two libraries of Zoophagus insidians (Fig. 1G) were successfully amplified (18S GenBank: MG920183). 18S sequence of one indicated the presence of contaminating rotifer DNA, but EF1-α (GenBank: MH923230) indicated the fungus was present in both libraries. No clean 16S sequences could be obtained from either library.
3.2. Assembly, binning, & protein prediction
Each of the three genome binning methods were differentially successful with different genome assemblies, demonstrating the utility of the consensus method (Table S2). For example, the Acaulopage tetraceros ESOM map indicated the presence of two main genomes with one being fungal and the other being bacterial (Fig. 2A); however, this method did not resolve a clear map of Zoophagus insidians (Fig. 2B). The consensus genomes ranged in size from 10.20 Mbp to 55.96 Mbp and an estimated completeness that ranged from 72% to 91% (Table 2).
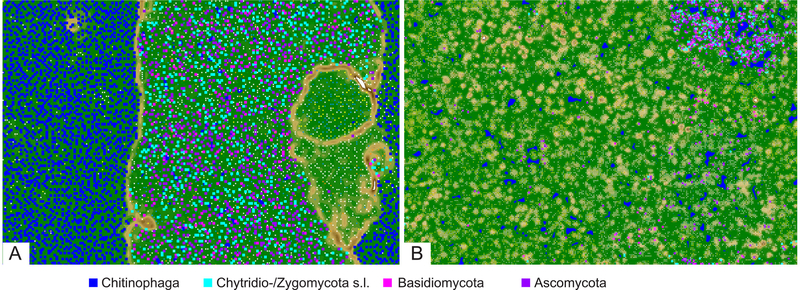
Fig. 2.
ESOM profiles for A. tetraceros (A) and Z. insidians (B) with blue/green areas indicating genomic space of high similarity and brown/white areas indicating genomic space of low similarity. In both, dark blue pixels represent contigs taxonomically identified as Chitinophaga sp. Aqua pixels represent contigs taxonomically identified as Chytridiomycota or Zygomycota s.l. Fuchsia pixels represent contigs taxonomically identified as Basidiomycota, and purple pixels represent contigs taxonomically identified as Ascomycota. Note the presence of brown and white ridges delineating the area corresponding to fungal contigs in (A) the A. tetraceros ESOM and their absence in (B) the Z. insidians ESOM.
Table 2.
Number of MDA libraries sequenced and assembly statistics for the taxa sequenced in this study.
| Taxon | Number of libraries sequenced & co-assembled | Number of scaffolds | Total size (Mbp) | Longest scaffold (bp) | N50 (bp) | GC content (%) | Cegma estimated completeness (%) | Number of conserved loci retrieved |
|---|---|---|---|---|---|---|---|---|
| Acaulopage tetraceros | 3 | 472 | 10.2 | 251,313 | 49,495 | 44 | 83.06 | 184 |
| Cochlonema odontosperma | 2 | 1819 | 16.84 | 147,303 | 21,635 | 54 | 89.52 | 174 |
| Stylopage hadra | 5 | 20,112 | 55.96 | 35,094 | 3564 | 36 | 77.42 | 110 |
| Zoopage sp. | 3 | 1958 | 13.92 | 64,126 | 14,452 | 54 | 71.77 | 163 |
| Zoophagus insidians | 2 | 2432 | 21.01 | 161,997 | 23,511 | 40 | 90.73 | 183 |
3.3. Phylogenomics
The concatenated alignment contained 51,571 distinct patterns. The best ML tree from the concatenated alignment (−ln = 1912187; Fig. 3; Fig. S1) was inferred with the LG model of protein evolution. With few exceptions, nodes across the tree had ≥90% bootstrap support. Zoopagomycota was recovered as monophyletic with 97% bootstrap support. Basidiobolus was placed sister to the Zoopagales with56% bootstrap support, and Entomophthoromycotina and Kickxellomycotina were placed sister to each other with 62% bootstrap support. Zoopagales (Zoopagomycotina) was recovered as monophyletic with 100% bootstrap support and all internal nodes had 100% bootstrap support. Zoophagus insidians and Stylopage hadra were sister taxa. Acaulopage tetraceros was sister to the Zoophagus + Stylopage clade. Cochlonema odontosperma and Zoopage sp. were sister taxa and were sister to the Zoophagus + Stylopage + Acaulopage clade, forming a clade containing all of the sampled predators and parasites. Piptocephalis cylindrospora was sister to this clade, while the remaining mycoparasites formed a clade comprised of Thamnocephalis sphaerospora and Syncephalis spp.
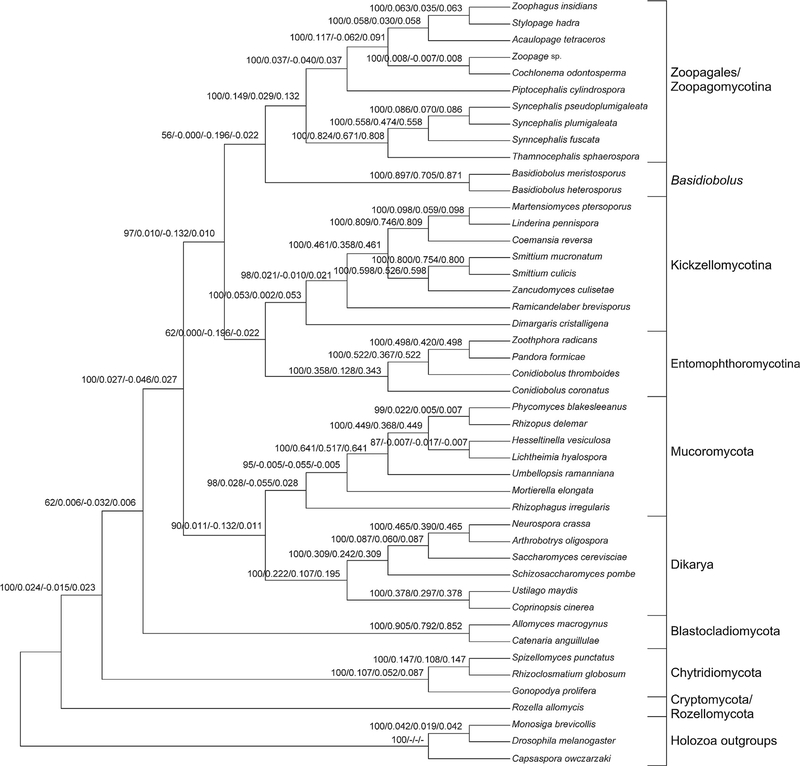
Fig. 3.
Best ML tree (−ln = 1912187) from the concatenated alignment. Node labels are bootstrap support/QP-IC/LQ-IC/EQP-IC. The TC/TCA scores for the tree were 12.732/13.715.
The ASTRAL tree (Fig. 4) was congruent with the ML when considering relationships within Zoopagales and most other relationships across the kingdom. However, there were three cases of incongruence. First, the ASTRAL tree placed Hesseltinella vesiculosa sister to the Phycomyces blakesleeanus + Rhizopus oryzae clade. Second, in the ASTRAL tree, Basidiobolus was sister to the Entomophthorales, which is consistent with its current classification based primarily on morphological features. Third, in the ASTRAL tree, Kickxellomycotina was sister to the Zoopagales.
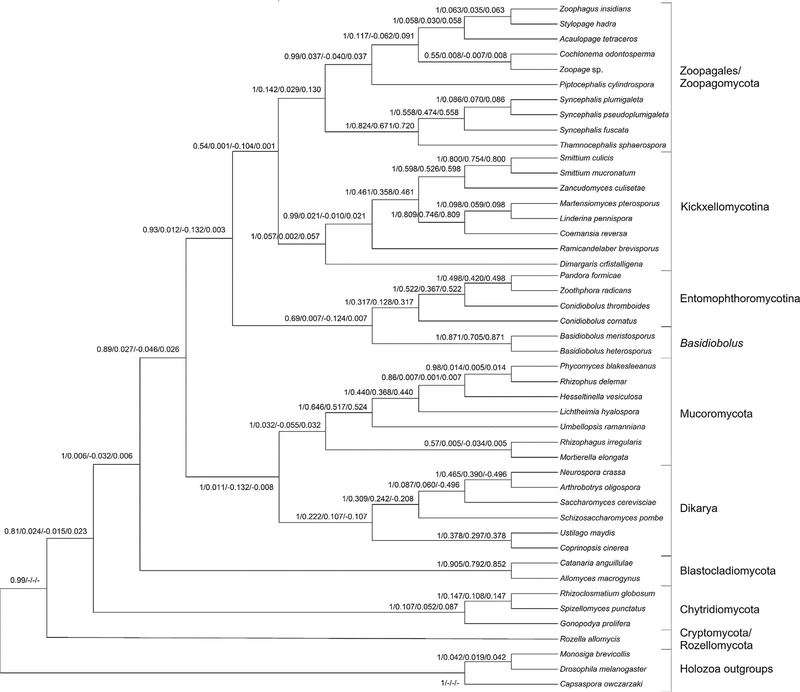
Fig. 4.
ASTRAL consensus tree. Node labels are consensus frequency/QP-IC/LQ-IC/EQP-IC. The TC/TCA scores for the tree were 12.798/13.655.
Internode certainty scores were generally low (≤0.50) across the ML and ASTRAL trees (Figs. 3 and 4). The internode representing Zoopagales received QP-IC/LQ-IC/EQP-IC scores of 0.142/0.029/0.130 in the ASTRAL tree and 0.149/0.029/0.132 in the ML tree. In the Zoopagales, the most uncertain internode was Cochlonema + Zoopage with QP-IC/LQ-IC/EQP-IC scores of 0.008/−0.007/0.008 in the ASTRAL tree and 0.008/−0.007/0.007 in the ML tree. The most certain internode in the Zoopagales was the Thamnocephalis + Syncephalis clade with QP-IC/LQ-IC/EQP-IC scores of 0.824/0.671/0.720 in the ASTRAL tree and 0.824/0.671/0.808 in the ML tree. Concerning the three cases of incongruence between the ASTRAL and ML trees, the ASTRAL placement of Basidiobolus, Hesseltinella and Kickxellomycotina received QP-IC/LQ-IC/EQP-IC scores of 0.007/−0.124/0.007, 0.007/0.001/0.007, and 0.001/−0.104/0.001 respectively. In the ML tree, placement of Basidiobolus, Hesseltinella and Kickxellomycotina received QP-IC/LQ-IC/EQP-IC scores of −0.0001/−0.196/−0.022, −0.007/−0.017/−0.007, and 0.0004/−0.196/−0.022 respectively. The TC/TCA scores for the ML tree were 12.732/13.715 and were 12.798/13.655 for the ASTRAL tree.
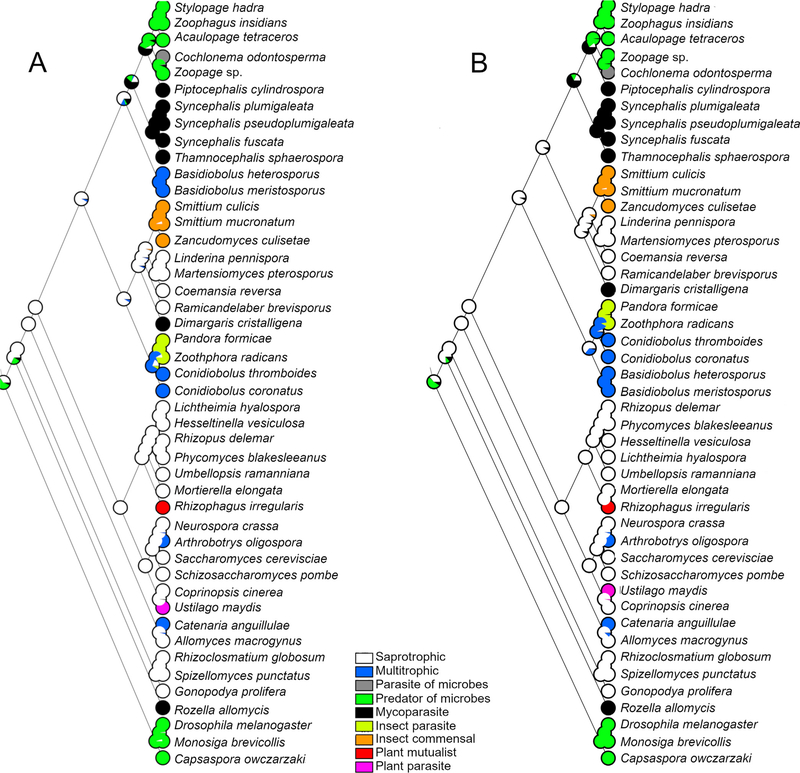
Maximum parsimony ancestral state reconstruction of the ML and ASTRAL trees indicated mycoparasitism was the ancestral trophic lifestyle of Zoopagales. Maximum likelihood ancestral state reconstructions of the ML and ASTRAL trees yielded similar results (Fig. 5A, B). For the ML tree (Fig. 5A), the character states that passed the decision threshold at the node representing Zoopagales were mycoparasitism (−lnL = 66.73) with moderate support (0.61), saprotrophic (−lnL = 67.95) with weak support (0.18), predacious (−lnL = 68.00) with weak support (0.17), and multitrophic (−lnL = 69.64) with weak support (0.03). For the ASTRAL tree (Fig. 5B), the character states at the Zoopagales node were mycoparasitism (−lnL = 66.83) with moderate support (0.66), saprotrophic (−lnL = 68.12) with weak support (0.18), and predacious (−lnL = 68.41) with weak (0.14) support.
Fig. 5.
Ancestral state reconstructions of trophic state on the best ML tree (A) and the ASTRAL consensus tree (B) using maximum likelihood. Pie graphs at nodes show the proportional likelihood of each character state.
3.4. Bacterial endosymbionts
As mentioned in Section 3.1, the MDA libraries of Acaulopage tetraceros and Stylopage hadra yielded clean 16S sequences. The top BLAST hit for the 16S sequence from the A. tetraceros libraries was to Chitinophaga sp. NR 2–02 (KM253107) with 99% similarity across the full length of the sequence. In the taxonomy report of the A. tetraceros genome, 48 contigs were identified as Chitinophaga. These contigs had an average coverage of 500× with a range of 6×–3300×. In the ESOM analysis of the A. tetraceros genome, there was a clear genomic region corresponding to Chitinophaga (the contigs highlighted in blue in Fig. 2A).
Clean, identical 16S sequences were present in four of the five Stylopage hadra libraries. The top BLAST hit was to Mycoavidus cysteinexigens strain B1-EB (AP018150) with 99% similarity across the full length of the sequence. We identified 89 contigs with a cumulative sequence length of approximately 1.92 Mbp corresponding the Mycoavidus genome. These contigs had an average coverage of 66× ranging from 3× to 1160×. ESOM analysis revealed a similarly clear genomic region encompassing the contigs identified as Mycoavidus (data not shown).
4. Discussion
Fungal lifestyles vary greatly across the tree of life, and different nutritional habits have arisen multiple times. Understanding the genomic basis of the evolution of these lifestyles can be facilitated by comparing taxa that have the same lifestyle but are of phylogenetically diverse origins. While such comparative studies can take place at the tips, e.g. among species of Tolypocladium (Quandt et al., 2018), a comprehensive comparison across the fungal tree is hampered by the lack of sampling and resolution at the deeper nodes. To help lay the groundwork for resolving these deeper nodes and to produce the first genomic sequences of the largest predacious fungal group, we sequenced the genomes of 5 predacious/parasitic taxa of Zoopagales.
In this study, we use single cell genomic approaches to generate genome-scale data for five uncultured zoopagalean fungi that interact antagonistically with other microorganisms. A side effect of the nonspecific amplification steps required by these methods is that sequencing libraries are especially prone to contamination from the environment and evidence uneven sequencing coverage across the resulting metagenomes (Pinard et al., 2006). Successful identification of assembly scaffolds belonging to our target fungi was further hindered by underrepresentation of zoopagalean fungi in sequence databases. To combat these problems, we leveraged scaffold binning approaches that hinge on sequence characteristics other than taxonomic annotation (Mikhailov et al., 2016). Despite the strong effects of contaminating sequence reads on library purity, we are able to resolve phylogenetic trees that describe relationships among zoopagalean fungi with high confidence. This bodes well for future genomic studies in Zoopagomycota as well as other uncultured lineages across the tree of life.
Given our methods, there are a few potential sources of error worth mentioning. First, the addition of the MDA step is likely a source of additional chimeras and mutations compared to a bulk extraction and direct sequencing of DNA. However, the error rates of the [PHI]29 are lower than that of taq (Binga et al., 2008; Gawad et al., 2016), though the formation of chimeras is still a concern. Second, since we were working with metagenomes, there is the possibility of contamination from other genomes. In fact, we did observe this with the genomes of Pandora formicae (from GenBank), Cochlonema odontosperma, and Zoophagus insidians, which were contaminated with genes from ants, Mortierella, and rotifers, respectively. These contaminating genes were identified using the gene trees and removed and/or replaced with correct orthologue when possible. Thus, we are reasonably confident that these sources of error have been accounted for and that those errors that were not detected had a minimal effect on our results.
Our inferred tree confirms that Zoopagales is a monophyletic lineage, but the relationship of Zoopagales to the other orders and subphyla remains unresolved. Early molecular studies based on 18S data challenged the monophyly of Zoopagales due to Zoophagus insidians grouping with Kickxellales (Tanabe et al., 2000; White et al., 2006). More recent studies relying on 18S sequences showed that with increased taxon sampling, Zoopagales is recovered as a monophyletic lineage, though internal nodes often have < 80% bootstrap support (Corsaro et al., 2018; Davis et al., in press). In agreement with previous studies (i.e., Corsaro et al., 2018; Davis et al., in press) we resolve a monophyletic Zoopagales but do so with strong bootstrap support for all internal nodes. We recover a Zoophagus insidians + Stylopage hadra clade and a Syncephalis + Thamnocephalis clade. In stark contrast to previous studies, our results indicate Zoophagus insidians + Stylopage harda is a relatively recent diverging lineage with strong bootstrap support rather than an early diverging lineage. In our analysis, the Zoophagus + Stylopage clade is sister to Acaulopage rather than Piptocephalis as observed in 18S phylogenies. In addition, the Syncephalis + Thamnocephalis clade is a separate lineage from the other taxa rather than being sister to Acaulopage and Cochlonema. This shows that additional sampling of 18S is not sufficient to resolve relationships within Zoopagales. Instead, it will take a combination of additional taxa and loci to accurately explore the relationships within this group.
Our results affirm the need for reclassification of genera in Zoopagales, which has been noted in previous studies. Currently, the mycoparasitic genera Piptocephalis and Syncephalis are classified together in the Piptocephalidaceae (Benny et al., 2016a), but previous 18S rRNA molecular phylogenies placed these taxa in separate clades (e.g. White et al., 2006; Corsaro et al., 2018) making the family Piptocephlidaceae paraphyletic. Our results confirm that the family Piptocephalidaceae is paraphyletic. The amoeba predators were also paraphyletic in our tree, and our results suggest Cochlonemataceae should not have been separated from Zoopagaceae (Benny et al., 2016b). However, additional sampling of taxa from the other families, Helicocephalidaceae and Sigmoideomycetaceae, is needed before formal revisions can be made.
Corsaro et al. (2018) hypothesized the ancestral state of Zoopagales was “zooparasitic” (i.e., predacious) due to the position of Zoophagus insidians. Based on our rDNA phylogeny, we hypothesized the ancestral state to be mycoparasitism or predation (Davis et al., in press). Our results could not resolve the ancestral trophic state of Zoopagales with mycoparasitism, predation, and saprobe as possible candidates. In both the ASTRAL and ML tree reconstructions, mycoparasitism had the highest probability among the character states passing the decision threshold, which could be interpreted to mean it is the most likely ancestral state (Schluter et al., 1997). An alternative interpretation is that because mycoparasitism has only moderate support as the ancestral character state, the ancestral state remains unresolved (Reyes et al., 2018). Even so, it is our opinion that mycoparasitism is the best candidate for the ancestral state of the Zoopagales. We base this claim on the fact that reconstructions of both the ML and ASTRAL trees gave the highest probability to mycoparasitism, the fact that the other candidate character states were weakly supported and the difference in probability between mycopararsitism and saprotrophy was ~0.46, the fact that mycoparasites and animal parasites have more similar genomic signatures compared to each other than to saprobes (Ahrendt et al., 2018), and the observation that the ancestor to the clade likely did not form specialized traps as, based on the position of Zoophagus insidians, specialized traps are a derived trait. A strong argument can be made that mycoparasitism of the Zoopagalean ancestor evolved from saprotrophy as it is the ancestral state of the three nodes preceding Zoopagalean divergence (i.e., only the saprobic trophic state passed the decision threshold), the fact that saprobes generally have a large complement of enzymes that can then be co-opted for other lifestyles (e.g. Kohler et al., 2015; Quandt et al., 2015), and that the fact that transitions from saprobe to other trophic states are more common than transitions back to saprobe (Tedersoo et al., 2010; Spatafora et al., 2012; Kohler et al., 2015).
Both Corsaro et al. (2018) and Davis et al. (in review) recover amoebophagous fungi as a monophyletic lineage suggesting the ability to utilize amoeba evolved once. Our results do not recover a monophyletic lineage of amoebophagous fungi, but they do not discount a single evolution of amoebophagy. Rather, they suggest that amoeba may have been the ancestral prey with later transitions to nematodes and rotifers in Stylopage + Zoophagus and endoparasitism in Cochlonema. However, our inferences are limited by the fact most of the diversity in Zoopagales is still not represented. For example, the placement of ectoparasites such as Amoebophilus remains unknown. Increased sampling of zoopagalean taxa and a more fine-scale ancestral state analysis is required to determine the ancestral invertebrates that were utilized and also to determine the frequency of host switching within the order.
It is tempting to hypothesize that the evolution of predation evolved a single time in the Zoopagales based on our results. However, in previous phylogenetic analyses of the order (e.g., Tanabe et al., 2000; White et al., 2006; Corsaro et al., 2018; Davis et al., in press), Rhopalomyces elegans was consistently placed sister to Syncephalis and Thamnocephalis with high bootstrap support. Rhopalomyces elegans preys on nematode eggs and adult nematodes (Ellis and Hesseltine, 1962; Barron, 1973), and other members of the genus prey on nematodes or rotifers (Barron, 1980). Considering that the Syncephalis + Thamnocephalis clade received strong support in both the ML and ASTRAL trees and was the most certain internode in the Zoopagales, we hypothesize that a genome-wide phylogeny would also place Rhopalomyces sister to Syncephalis and Thamnocephalis. Furthermore, there is no molecular data for several other predacious taxa (e.g., Cystopage, Brachymyces, and Verrucomyces). Thus, until the placement of these taxa is known, it is unclear how many times the predacious lifestyle evolved in Zoopagales.
Contrary to Spatafora et al. (2016), we cannot confidently determine the relationships among Entomophthoromycotina, Kickxellomycotina, and Zoopagomycotina. The ML tree and ASTRAL tree from Spatafora et al. (2016) were based on the same 192 conserved proteins and placed a monophyletic Entomophthoromycotina sister to a clade containing the Zoopagomycotina and Kickxellomycotina. Our ASTRAL consensus tree is congruent with the topology recovered by Spatafora et al. (2016), but in our ML tree, the Entomophthoromycotina was not monophyletic because Basidiobolus was sister to Zoopagales. As well, our ML tree place Kixellomycotina sister to Entomophthorales instead of Zoopagales. Our Internode Certainty and Tree Certainty analyses suggest that the ASTRAL topology is the correct one: the TC scores for the ASTRAL tree were slightly higher than those for the ML tree, though the TCA score was higher for the ML tree; and, the quartet IC scores for the ASTRAL tree placement of Basidiobolus and Kickxellomycotina were higher than the ML placement of those taxa. However, additional taxon and marker sampling coupled with a detailed analysis of conflicting signal and noise among the markers and/or an analysis of non-bifurcating evolution is needed to determine the correct topology.
There is a growing body of research on bacterial endosymbionts in fungi, and it is proposed that the symbiosis is a major factor in fungal ecology and evolution (Araldi-Brondolo et al., 2017). These endosymbionts are mainly known from plant-associated fungi, but also appear to be frequent in Mucoromycota (Araldi-Brondolo et al., 2017). However, at this time, there are no reports of endosymbionts in predacious fungi. An interesting observation made in the course of this study was the potential presence of endosymbiotic bacteria in Stylopage hadra and Acaulopage tetraceros. The 16S sequence from S. hadra was identified as Mycoavidus sp, a genus known to be an endosymbiont of Mortierellomycotina (Araldi-Brondolo et al., 2017). Given that this bacterial taxon is a known endosymbiont and that S. hadra spores are aerial (Drechsler, 1935b), it seems unlikely this is a contaminant. The 16S sequence from A. tetraceros was identified as Chitinophaga, a genus known to be an endosymbiont of at least one strain of Fusarium (Shaffer et al., 2017). However, most members of Chitinophaga are soil saprobes and potentially degrade fungal mycelia (Brabcová et al., 2016). Also, A. tetraceros spores are sessile on the hyphae (Dreschler, 1935a). Thus, it is harder to make the case that Chitinophaga is an endosymbiont of A. tetraceros. Screening additional isolates from a broad geographic range both molecularly and microscopically will be needed to distinguish between the bacterial endosymbiont and contaminant hypotheses.
5. Conclusions
We sequenced the genomes of several predacious taxa in Zoopagales in an effort to help resolve the deep nodes in the fungal phylogeny. Our results confirm a monophyletic Zoopagales but are unable to resolve the ancestral trophic state of the zoopagalean ancestor. The relationships among the subphyla of Zoopagomycota varied depending on method of analysis, which suggests that greater taxonomic and genomic sampling is required to resolve these deep nodes.
Supplementary Material
Acknowledgements
Miranda R. Guinaldo digitally edited rough hand drawings to produce the graphical abstract, for which we are grateful. Elsevier’s Illustration Services were used to make Figs. 3–4, for which we are grateful. We are grateful to Jim & Vickie Davis and Elizabeth Rhodes for the samples yielding Cocholonema odontosperma and Zoopage sp. We are grateful to Jason Martin for the soil sample yielding Acaulopage tetraceros. We thank Doina Ciobanu for advice on single cell methods. We thank Alisha Quandt, Rob Powers and Rabern Simmons for bioinformatic advice. We also acknowledge Katy Lazarus for help in producing genomes of Syncephalis species. We are grateful to the Joint Genome Institute for allowing us to use their data. We appreciate the work of the reviewers and the editors of Molecular Phylogenetics and Evolution. This project was funded by NSF grants DEB 1441677 and DEB 1441604.
Footnotes
Appendix A. Supplementary material
Supplementary data to this article can be found online at https://doi.org/10.1016/j.ympev.2019.01.006.
References
- Ahrendt SR, Quandt CA, Ciobanu D, Clum A, Salamov A, Andreopoulos B, Cheng J-F, Woyke T, Pelin A, Henrissat B, Reynolds NK, Benny GL, Smith ME, James TY, Grigoriev IV, 2018. Leveraging single-cell genomics to expand the fungal tree of life. Nat. Microbiol 10.1038/s41564-018-0261-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alexopoulos CJ, Mims CW, Blackwell M, 1996. Introductory Mycology, fourth ed. John Wiley and Sons Inc. [Google Scholar]
- Araldi-Brondolo SJ, Spraker J, Shaffer JP, Woytenko EH, Baltrus DA, Gallery RE, Arnold AE, 2017. Bacterial endosymbionts: Master modulators of fungal phenotypes. Microbiol. Spectrum 5 10.1128/microbiolspec.FUNK-0056-2016. [DOI] [PubMed] [Google Scholar]
- Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA, 2012. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol 19, 455–477. 10.1089/cmb.2012.0021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baral H-O, Weber E, Gams W, Hagedorn G, Liu B, Liu X, Marson G, Marvanová L, Stadler M, Weiß M, 2018. Generic names in the Orbiliaceae (Orbiliomycetes) and recommendations on which names should be protected or suppressed. Mycol. Prog 17, 5–31. 10.1007/s11557-017-1300-6. [DOI] [Google Scholar]
- Barron GL, 1973. Nematophagous fungi: Rhopalomyces elegans. Can. J. Bot 51, 2505–2507. [Google Scholar]
- Barron GL, 1980. The biological role of Rhopalomyces magnus. Mycologia 72, 427–430. [Google Scholar]
- Barron GL, Dierkes Y, 1977. Nematophagous fungi: Hohenbuehelia, the perfect state of Nematoctonus. Can. J. Bot 55, 3054–3062. [Google Scholar]
- Benny GL, Ho H-M, Lazarus KL, Smith ME, 2016a. Five new species of the obligate mycoparasite Syncephalis (Zoopagales, Zoopagomycotina) from soil. Mycologia 108, 1114–1129. [DOI] [PubMed] [Google Scholar]
- Benny GL, Smith ME, Kirk PK, Tretter ED, White MM, 2016b. Challenges and future perspectives in the systematics of Kickxellomycotina, Mortierellamycotina, Mucoromycotina, and Zoopagomycotina In: Li D–W (Ed.), Biology of Microfungi. Springer International Publishing, Switzerland, pp. 65–126. [Google Scholar]
- Berbee ML, James TY, Strullu-Derrien C, 2017. Early diverging Fungi: Diversity and impact at the dawn of terrestrial life. Annu. Rev. Microbiol 71, 41–60. [DOI] [PubMed] [Google Scholar]
- Binga EK, Lasken RS, Neufeld JD, 2008. Something from (almost) nothing: the impact of multiple displacement amplification on microbial ecology. The ISME J. 2, 233. [DOI] [PubMed] [Google Scholar]
- Bolger AM, Lohse M, Usadel B, 2014. Trimmomatic: A flexible trimmer for illumina sequence data. Bioinformatics btu170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brabcová V, Nováková M, Davidová A, Baldrian P, 2016. Dead fungal mycelium in forest soil represents a decomposition hotspot and a habitat for a specific microbial community. New Phytol. 210, 1369–1381. [DOI] [PubMed] [Google Scholar]
- Capella-Gutierrez S, Silla-Martinez JM, Gabaldon T, 2009. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corsaro D, Köhler M, Wylezich C, Venditti D, Walochnik J, Michel R, 2018. New insights from molecular phylogenetics of amoebophagous fungi (Zoopagomycota, Zoopagales). Parasitol. Res 117, 157–167. 10.1007/s00436-017-5685-6. [DOI] [PubMed] [Google Scholar]
- Davis WJ, Amses KR, James ES, James TY, in press. A new 18S rRNA phylogeny of uncultured predacious fungi (Zoopagales). Mycologia. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dick GJ, Andersson A, Baker BJ, Simmons SS, Thomas BC, Yelton AP, Banfield JF, 2009. Community-wide analysis of microbial genome sequence signatures. Genome Biol. 10, R85 10.1186/gb-2009-10-8-r85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drechsler C, 1935a. Some non-catenulate conidial phycomycetes preying on terricolous amoebae. Mycologia 27, 176–205. [Google Scholar]
- Drechsler C, 1935b. A new species of conidial phycomyete preying on nematodes.Mycologia 27, 206–215. [Google Scholar]
- Drechsler C, 1935c. A new mucendinaceous fungus capturing and consuming Amoeba verrucose. Mycologia 27, 216–223. [Google Scholar]
- Duddington CL, 1956. The predacious fungi: Zoopagales and Moniliales. Biol. Rev 31,152–193. [Google Scholar]
- Ekman S, Andersen HL, Wedin M, 2008. The limitations of ancestral state reconstruction and the evolution of the ascus in the Lecanorales (lichenized Ascomycota). Syst. Biol 57, 141–156. [DOI] [PubMed] [Google Scholar]
- Earl D, Bradnam K, St. John J, Darling A, Lin D, Fass J, Yu HOK, Buffalo V, Zerbino DR, Diekhans M, Nguyen N, Ariyarante PN, Sung WK, Ning Z, Haimel M, Simpson JT, Fonseca NA, Birol I, Docking TR, H IY, Rokhsar DS, Chikhi R, Lavenier D, Chapuis G, Naquin D, Maillet N, Schatz MC, Kelley DR, Phillippy AM, Koren S, Yang SP, Wu W, Chou WC, Srivastava A, Shaw TI, Ruby JG, Skewes-Cox P, Betegon M, Dimon MT, Solovyev V, Seledtsov I, Kosarey P, Vorobyey D, Ramirez-Gonzalez R, Leggett R, MacLean D, Xia F, Luo R, Li Z, Xie Y, Liu B, Gnerre S, MacCallum I, Przybylski D, Ribeiro FJ, Yin S, Sharpe T, Hall G, Kersey PJ, Durbin R, Jackman SD, Chapman JA, Huang X, DeRisi JL, Caccamo M, Li Y, Jaffe DB, Green RE, Haussler D, Korf I, Paten B, 2011. Assemblathon 1: A competitive assessment of de novo short read assembly methods. Genome Res. 21, 224–2241. 10.1101/gr.126599.111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ellis JJ, Hesseltine CW, 1962. Rhopalomyces and Spinellus in pure culture and the parasitism of Rhopalomyces on nematode eggs. Nature 193, 699–700. [Google Scholar]
- Galindo LJ, Torruella G, Moreira D, Timpano H, Paskerova G, Smirnov A,Nassonova E, López-García P, 2018. Evolutionary genomics of Metchnikovella incurvate (Metchnikovellidae): an early branching Microsporidium. Genome Biol. Evol 10, 2736–2748. 10.1093/gbe/evy205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gawad C, Koh W, Quake SR, 2016. Single-cell genome sequencing: current state of the science. Nat. Rev. Genet 17, 175–188. [DOI] [PubMed] [Google Scholar]
- de Freitas Soares FE, Sufiate BL, de Queiroz JH, 2018. Nematophagous fungi: Far beyond the endoparasite, predator, and ovicidal groups. Agric. Nat. Resour 52, 1–8. 10.1016/j.anres.2018.05.010. [DOI] [Google Scholar]
- Holt C, Yandell M, 2011. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinf. 12, 491 10.1186/1471-2105-12-491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- James TY, Kauff F, Schoch CL, Matheny PB, Hofstetter V, Cox CJ, Celio G, Gueidan C, Fraker E, Miadlikowska J, Lumbsch HT, Raunhut A, Reeb V, Arnold AE, Amtoft A, Stajich JE, Hosaka K, Sung G-H, Johnson D, O’Rourke B, Crockett M, Binder M, Curtis JM, Slot JC, Wang Z, Wilson AW, Schüßler A, Longcore JE, O’Donnell K, Mozley-Standridge S, Porter D, Letcher PM, Powell MJ, Taylor JW, White MM, Griffith GW, Davies DR, Humber RA, Morton JB, Sugiyama J, Rossman AY, Rogers JD, Pfister DH, Hewitt D, Hansen K, Hambleton S, Shoemaker RA, Kohlmeyer J, Volkman-Kolhmeyer B, Spotts RA, Serdani M, Crous PW, Hughes KW, Matasuura K, Langer E, Langer G, Untereiner WA, Lücking R, Büdel B, Geiser DM, Aptroot A, Diederich P, Schmitt I, Schultz M, Yahr R, Hibbett DS, Lutzoni F, McLaughlin DJ, Spatafora JW, Vilgalys R, 2006. Reconstructing the early evolution of Fungi using a six-gene phylogeny. Nature 443, 818–822. 10.1038/nature05110. [DOI] [PubMed] [Google Scholar]
- James TY, Pelin A, Bonen L, Ahrendt S, Sain D, Corradi N, Stajich JE, 2013. Shared signatures of parasitism and phylogenomics unite Cryptomycota and microsporidia. Curr. Biol 23, 1548–1553. 10.1016/j.cub.2013.06.057. [DOI] [PubMed] [Google Scholar]
- Keller O, Kollmar M, Stanke M, Waack, 2011. A novel hybrid gene prediction method employing protein multiple sequence alignments. Bioinformatics 10.1093/bioinformatics/btr010. [DOI] [PubMed] [Google Scholar]
- Kirk PM, Cannon PF, Minter DW, Stalpers JA, 2008. Dictionary of the Fungi. 10th ed. CABI. [Google Scholar]
- Kobert K, Salichos L, Rokas A, Stamatakis A, 2016. Computing the Internode Certainty and related measures from partial gene trees. Mol. Biol. Evol 33, 1606–1617. 10.1093/molbev/msw040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohler A, Kuo A, Nagy LG, Morin E, Barry KW, Buscot F, Canbäck B, Choi C, Cichocki N, Clum A, Copaert J, Copeland A, Cost MD, Doré J, Floudas D, Gay G, Girlanda M, Henrissat B, Herrmann S, Hess J, Högberg N, Johansson T, Khouja H-R, LaButti K, Lahrmann U, Levasseur A, Lindquist EA, Lipzen A, Marmeisse R, Martino E, Murat C, Ngan CY, Nehls U, Plett JM, Pringle A, Ohm RA, Perotto S, Peter M, Riley R, Rineau R, Ruytinx J, Salamov A, Shah F, Sun H, Tarkka M, Tritt A, Veneault-Fourrey C, Zuccaro A, Mycorrhizal Genomics Initiative Consortium, Tunlin A, Grigoriev IV, Hibbett DS, Martin F, 2015. Convergent losses of decay mechanisms and rapid turnover of symbiosis genes in mycorrhizal mutualists. Nature Genetics. 47, 410–415. [DOI] [PubMed] [Google Scholar]
- Kumar S, Jones M, Koutsovoulos G, Clarke M, Blaxter M, 2013. Blobology: exploring raw genome data for contaminants, symbionts, and parasites using taxon-annotated GC-coverage plots. Front. Genet 4, 1–12. 10.3389/fgene.2013.00237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lafferty KD, Kurtis AM, 2002. Trophic strategies, animal diversity and body size. Trends Ecol. Evol 17, 507–513. 10.1016/S0169-5346(02)02615-0. [DOI] [Google Scholar]
- Lewis PO, 2001. A likelihood approach to estimating phylogeny from discrete morphological character data. Syst. Biol 50, 913–925. [DOI] [PubMed] [Google Scholar]
- Luo H, Mo MH, Huang XW, Li X, Zhand KQ, 2004. Coprinus somatus: a basidiomycete fungus forms novel spiny structures and infects nematodes. Mycologia 96, 1218–1225. [PubMed] [Google Scholar]
- Maddison WP, Maddison DR, 2006. Stochar: A package of Mesquite modules for stochastic models of character evolution. Version 1.1.
- Maddison WP, Maddison DR, 2018. Mesquite: A modular system for evolutionary analysis. Version 3.40. http://mesquiteproject.org.
- McInerney J, 1998. GCUA: general codon usage analysis. Bioinformatics 14 (4),372–373. 10.1093/bioinformatics/14.4.372. [DOI] [PubMed] [Google Scholar]
- Michel R, Steid M, Köhsler M, Walochnik J, 2015. Acaulopage tetraceros Dreschler 1935 (Zoopagales): cultivation, prey pattern, and molecular characterization. J. Endocytobiosis Cell Res 26, 76–82. [Google Scholar]
- Mikhailov KV, Simdyanov TG, Aleoshin VV, 2016. Genomic survey of a hyperparasitic microsporidian Amphiamblys sp. (Metchnikovellidae). Genome Biol. Evol 9, 454–467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirarab S, Reaz R, Bayzid Md., S., Zimmermann T, Swenson MS, Warnow T, 2014. ASTRAL: Genome-scale coalescent-based species tree. Bioinformatics 30, i541–i548. 10.1093/bioinformatics/btu462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molles MC Jr., 2008. Ecology: Concepts and Applications. McGraw-Hill, Boston,Massachusetts, pp. 604. [Google Scholar]
- Parra G, Bradnam K, Korf I, 2007. CEGMA: a pipeline to accurate annotate core genes in eukaryotic genomes. Bioinformatics 9, 1061–1067. 10.1093/bioinformatics/btm071. [DOI] [PubMed] [Google Scholar]
- Pati A, Heath LS, Kyrpides NC, Ivanova N, 2011. ClaMS: A classifier for metagenomic sequences. Standards Genomic Sci. 5, 248–253. https://doi.org10.4056/sigs.2075298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pinard R, de Winter A, Sarkis GJ, Gerstein MB, Tartaro KR, Plant RN, Egholm M, Rothberg JM, Leamon JH, 2006. Assessment of whole genome amplification-induced bias through high-throughput, massively parallel whole genome sequencing. BMC Genomics 7, 216 10.1186/1471-2164-7-216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prowse GA, 1954. Sommerstorffia spinose and Zoophagus insidians predaceous on rotifers, and Rozellaopsis inflata, the endoparasite of Zoophagus. Transactions Br. Mycol. Soc 37, 134–150. [Google Scholar]
- Quandt CA, Kohler A, Hesse CN, Sharpton TJ, Martin R, Spatafora JW, 2015. Metagenome sequence of Elaphomyces granulatus from sporocarp tissue reveals Ascomycota ectomycorrhizal fingerprints of genome expansion and a Proteobacteriarich microbiome. Environ. Microbiol 17, 2952–2968. [DOI] [PubMed] [Google Scholar]
- Quandt CA, Patterson, Spatafora J, 2018. Harnessing the power of the phylogenomics to disentangle the directionality and signatures of interkingdom host jumping in the parasitic fungal genus Tolypocladium. Mycologia 110, 104–117. 10.1080/00275514.2018.1442618. [DOI] [PubMed] [Google Scholar]
- Raffel TR, Martin LB, Rohr JR, 2008. Parasites as predators: unifying natural enemy ecology. Trends Ecol. Evol 23, 610–618. [DOI] [PubMed] [Google Scholar]
- Reyes E, Nadot A, von Balthazar M, Schönenberger J, Sauqute H, 2018. Testing the impact of morphological rate heterogeneity on ancestral state reconstruction of five floral traits in angiosperms. Sci. Rep 9473 10.1038/s41598-018-27750-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saikawa M, Baba N, Aoki Y, 1994. Electron microscopy of amoeba-capturing Dactylella tylopaga, showing the morphology of a basidiomycete. Mycologia 86, 474–477. [Google Scholar]
- Salichos L, Stamatakis A, Rokas A, 2014. Novel information theory-based measures for quantifying incongruence among phylogenetic trees. Mol. Biol. Evol 31, 1261–1271. 10.1093/molbev/msu061. [DOI] [PubMed] [Google Scholar]
- Schluter D, Price T, Mooers AØ, Ludwig D, 1997. Likelihood of ancestor states in adaptive radiation. Evolution 51, 1699–1711. [DOI] [PubMed] [Google Scholar]
- Shaffer JP, U’Ren J, Gallery RE, Baltrus DA, Arnold AE, 2017. An endohyphal bacterium (Chitinophaga, Bacteroidetes) alters carbon source use by Fusarium keratoplasticum (F. solani Species complex, Nectriaceae). Frontiers Microbiol. 8, 350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spatafora JW, Owensby CA, Douhan GW, Boehm EWA, Schoch CL, 2012. Phylogenetic placement of the ectomycorrhizal genus Cenococcumin Gloniaceae (Dothideomycetes). Mycologia 104 (758), 765. [DOI] [PubMed] [Google Scholar]
- Spatafora JW, Chang Y, Benny GL, Lazarus K, Smith ME, Berbee ML, Bonito G, Corradi N, Grigoriev I, Gryganski A, James TY, O’Donnell K, Roberson RW, Taylor TN, Uehling J, Vilgalys R, White MM, 2016. A phylum-level phylogenetic classification of zygomycete fungi based on genome-scale data. Mycologia 108, 1028–1046. 10.3852/16042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stamatakis A, 2006. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analysis with thousands of taxa and mixed models. Bioinfomatics. 22, 2688–2690. [DOI] [PubMed] [Google Scholar]
- Stamatakis A, Hoover P, Rougemount J, 2008. A rapid algorithm for the RAxML web servers. SystBiol. 57, 758–771. [DOI] [PubMed] [Google Scholar]
- Stanke M, Waack S, 2003. Gene prediction with a hidden-markov model and a new intron submodel. Bioinformatics 19 ii215–ii225. [DOI] [PubMed] [Google Scholar]
- Tanabe Y, O’Donnell K, Saikawa M, Sugiyama J, 2000. Molecular phylogeny of parasitic Zygomycota (Dimargaritales, Zoopagales) based on nuclear small subunit ribosomal DNA sequences. Mol. Phylogenet. Evol 16, 253–262. [DOI] [PubMed] [Google Scholar]
- Tedersoo L, May TW, Smith ME, 2010. Ectomycorrhizal lifestyle in fungi: global diversity, distribution and evolution of phylogenetic lineages. Mycorrhiza 20, 217–263. [DOI] [PubMed] [Google Scholar]
- Thorn RG, Barron GL, 1984. Carnivorous mushrooms. Science 224, 76–78. [DOI] [PubMed] [Google Scholar]
- Thorn RG, Moncalvo J-M, Reddy CA, Vilgalys R, 2000. Phylogenetic analyses and the distribution of nematophagy support a monophyletic Pleurotaceae within the polyphyletic pleurotoid-lentinoid fungi. Mycologia 92, 241–252. [Google Scholar]
- Weisburg WG, Barns SM, Pelletier DA, Lane DJ, 1991. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol 173 (2), 697–703. 10.1128/jb.173.2.697-703.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- White MM, James TY, O’Donnell K, Cafaro MJ, Tanabe Y, Sugiyama J, 2006. Phylogeny of the Zygomycota based on nuclear ribosomal sequence data. Mycologia 98, 872–884. [DOI] [PubMed] [Google Scholar]
- Woyke T, Doud DFR, Schluz F, 2017. The trajectory of single-cell sequencing. Nat. Methods 14, 1045. [DOI] [PubMed] [Google Scholar]
- Yang E, Xu L, Yang Y, Zhang X, Xiang M, Wang C, An Z, Liu X, 2012. Origin and evolution of carnivorism in the Ascomycota (Fungi). Proc. Natl. Acad. Sci 109, 10960–10965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang C, Sayyari E, Mirarab S, 2017. ASTRAL-III: Increased scalability and impacts of contracting low support branches. In: Comparative Genomics: 15th International Workshop. RECOMB CG. 10.1007/978-3-319-67979-2_4. [DOI] [Google Scholar]
- Zhou X, Lutteropp S, Czech L, Stamatakis A, von Looz M, Rokas A, 2017. Quartet-based computations of internode certainty provide accurate and robust measures of phylogenetic incongruence. bioRxiv. 10.1101/168526. [DOI] [PubMed] [Google Scholar]
- Zhou X, Shen XX, Hittinger CT, Rokas A, 2018. Evaluating fast maximum likelihood-based phylogenetic programs using empirical phylogenomic data sets. Mol. Biol. Evol 35, 486–503. 10.1093/molbev/msx302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zouhar M, Douda O, Nováková J, Doudová E, Mazáková J, Wenzlová J, Ryšánek P, Renčo, 2013. First report about the trapping activity of Stopharia rugosoannulata acanthocytes for northern root knot nematode. Helminthologia. 50, 127–131. 10.2478/s11687-013-0120-8. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Assemblies and raw reads were deposited in NCBI’s databases under Bioproject PRJNA451036. Scripts and representative output files are available on GitHub (https://github.com/wjdavis90/ZyGoLife/tree/master/phylogenomics). The concatenated alignment, ML tree, ASTRAL tree, and trophic character matrix are deposited in TreeBase (http://purl.org/phylo/treebase/phylows/study/TB2:S22698).