Abstract
Tuberculosis is an infectious disease of global concern. Members of the diazaquinomycin (DAQ) class of natural products have shown potent and selective activity against drug-resistant Mycobacterium tuberculosis. However, poor solubility has prevented further development of this compound class. Understanding DAQ biosynthesis may provide a viable route for the generation of derivatives with improved properties. We have sequenced the genomes of two actinomycete bacteria that produce distinct DAQ derivatives. While software tools for automated biosynthetic gene cluster (BGC) prediction failed to detect DAQ BGCs, comparative genomics using MAUVE alignment led to the identification of putative BGCs in the marine Streptomyces sp. F001 and in the freshwater Micromonospora sp. B006. Deletion of the identified daq BGC in strain B006 using CRISPR-Cas9 genome editing abolished DAQ production, providing experimental evidence for BGC assignment. A complete model for DAQ biosynthesis is proposed based on the genes identified. Insufficient knowledge of natural product biosynthesis is one of the major challenges of productive, genome mining approaches. The results reported here fill a gap in knowledge regarding the genetic basis for the biosynthesis of DAQ antibiotics. Moreover, identification of the daq BGC shall enable future generations of improved derivatives using biosynthetic methods.
Graphical Abstract

Tuberculosis (TB) is a global health threat. In 2016, the World Health Organization reported 10.4 million new TB cases and approximately 1.7 million deaths worldwide, with over 95% of them occurring in low- and middle-income countries.1 Approximately 20% of the reported TB cases are estimated to be resistant to at least one first- and second-line anti-TB drug.1 The increasing number of multidrug-resistant TB cases highlights the importance and clinical need for continued anti-TB drug discovery and development efforts. Members of the diazaquinomycin (DAQ) class of diaza-anthraquinone antibiotics (Figure 1), have shown potent and selective inhibitory activity against Mycobacterium tuberculosis (MIC of ~0.1 μM against strain H37Rv).2 The in vitro activity profile of DAQ A (1) was equivalent to clinical anti-TB drugs and the potency was maintained against a panel of drug-resistant M. tuberculosis strains.2
Figure 1.
Structures of diazaquinomycin (DAQ) natural products and related compounds. The bacterial and environmental source is indicated below each structure.
DAQ A (1) was first isolated from soil Streptomyces sp. OM-704 in 1982.3 It was reported then to exhibit weak antibiotic activity.3 The structures of DAQ A and its reduced derivative DAQ B (2) were published the following year.4 To date, seven additional derivatives have been isolated, DAQs C (3) and D (4) from soil Streptomyces sp. GW48/1497,5 DAQs E (5), F (6), and G (7) from the marine-derived Streptomyces sp. F001,6 and DAQs H (8) and J (9) from the freshwater-derived Micromonospora spp. B0262 and B0067 (Figure 1). Although DAQs have been known for over three decades, their potent and selective anti-TB activity was only recently discovered after a screening campaign of freshwater actinomycete extracts for anti-TB hit identification.2
In 1985, Murata and colleagues reported that DAQs target mammalian and bacterial thymidylate synthase.8 More recent data indicated that at least in M. tuberculosis this is however not the case.2 Despite extensive efforts, the anti-TB mechanism of action of DAQs remains unknown.9 Transcription profiling experiments and the central quinone substructure of DAQs suggest that cell damage in M. tuberculosis may be caused by extensive redox chemistry.9 DAQs show structural similarity to nybomycin (10),10, 11 deoxynybomycin (11),10, 11 SCH 538415 (12),12 and deoxynyboquinone (13) (Figure 1).10, 13 Nybomycin was originally isolated from the terrestrial Streptomyces sp. A717 and showed anti-phage and antimicrobial activity.14 It was re-discovered in 2015 as an anti-TB agent that targets both the active and dormant states of M. tuberculosis.15 Deoxynybomycin (11), originally described as a degradation product of 10,10 was later found to be produced by a soil Streptomyces hyalinum strain and to have more potent activity than 10.16, 17 SCH 538415 (12) was isolated in 2003 from an unidentified bacterial microbe during a mechanism-based drug discovery campaign aimed at identifying inhibitors of bacterial acyl carrier protein synthetase,12 and again three years later from a riverbank soil-derived Streptomyces sp.18 It was reported to possess anticancer and moderate antibacterial activity.12, 18 A nearly identical compound, 13, was originally synthesized during studies of the antibiotic 1010 and was more recently found to be a natural product produced by a deep-sea actinomycete, Pseudonocardia sp. SCSIO 01299.19, 20 Deoxynyboquinone (13) has been shown to be reductively activated by the enzyme NAD(P)H:quinone oxidoreductase 1, and to undergo redox cycling that induces cancer cell death through a reactive oxygen species (ROS)-based mechanism.13, 21
DAQs suffer from poor water solubility, presumably due to their planar structures leading to intermolecular pi-stacking interactions, and the hydrophobicity of the aliphatic side chains, as has been reported for 13 and 10.10, 13 This has prevented extensive animal studies thus far. While total synthesis routes to DAQs have been described,22–25 modifications to the alkyl chains remain a challenge, and analogs of improved pharmaceutical properties have yet to be reported. We are interested in understanding the molecular basis for differential DAQ biosynthesis in Streptomyces sp. F001 and Micromonospora sp. B006, and in applying this knowledge towards the generation of DAQ derivatives with improved properties. For instance, enzymes involved in side ring biosynthesis could potentially be harnessed to introduce functional groups that may lead to improved solubility.
The structural similarity of 1 – 13 indicates that they share a similar biosynthetic origin. Precursor labeling studies performed in the 1970s with the nybomycin (10) producer defined acetate as the source of the side ring carbons of 10, while the central nucleus was shown to originate from the shikimate pathway.26, 27 Herein we report the identification of daq biosynthetic gene clusters (BGCs) in the marine-derived Streptomyces sp. F001 and the freshwater-derived Micromonospora sp. B006 by comparative genomics.
We recently sequenced the genome of Micromonospora sp. strain B006, the producer of 8 and 9.28 Automated genome mining tools were unsuccessful in detecting a putative daq BGC. Incomplete knowledge of natural product biosynthesis is one of the major challenges of productive, genome mining approaches that try to connect natural products to BGCs. To overcome this challenge, we report analysis of an additional genome, that of Streptomyces sp. F001 as the producer of 1 and 5 – 7, and the identification of putative daq BGCs by comparative genomics. Moreover, deletion of the identified genes in strain B006 using CRISPR-Cas9 genome editing provided evidence for gene cluster assignment. Based on the genes identified we propose a model for diazaquinomycin biosynthesis.
RESULTS AND DISCUSSION
Comparative Genomics Leads to the Identification of a Putative DAQ Gene Cluster.
The genome of the marine actinomycete bacterium Streptomyces sp. F001 was sequenced with short-read Illumina and long-read Pacific Biosciences technologies.29 Bioinformatic analyses revealed 24 secondary metabolite BGCs (Tables S1 to S26, and Supplementary Results). However, automated genome mining techniques, such as antiSMASH 4.0, failed to identify putative daq BGCs in strains F001 or B006, suggesting that biosynthesis of DAQs does not conform to generally known biosynthetic pathways for secondary metabolites or that their biosynthesis resembles primary metabolism. We anticipated that the latter is the case given that involvement of the shikimate pathway and fatty acid biosynthesis is suggested based on precursor labeling studies with 10.26, 27
In order to identify genes putatively involved in DAQ biosynthesis, the genome of Streptomyces sp. strain F00129 was compared with that of Micromonospora sp. strain B00628 using progressive MAUVE alignment.30 The genome of the well-studied and non-producer strain Streptomyces coelicolor A3(2)31 was added to the alignment to exclude common genes from primary metabolism. We were then able to identify an approximately 19-kb region that was shared between strains F001 and B006 (Figure 2). Interestingly, in Micromonospora sp. strain B006 the identified region lacked four genes putatively encoding a 3-deoxy-D-arabinoheptulosonic acid 7-phosphate (DAHP) synthase (daqJ), an anthranilate synthase (daqG), an isochorismatase (daqH) and a 2,3-dihydro-2,3-dihydroxybenzoate dehydrogenase (daql). Homologues of these four genes were identified within the previously described diazepinomicin BGC using BLAST searches.28 The daq gene cluster is located in the direct vicinity of the diazepinomicin BGC in strain B006 (Figure 2). Strain F001 does not contain a diazepinomicin BGC.
Figure 2.
Diazaquinomycin BGCs from Streptomyces sp. F001 and Micromonospora sp. B006. Homologous genes are connected by grey areas. Genes are color-coded by proposed function as shown.
Open reading frames (ORFs) within the daq BGCs were annotated and analyzed for sequence similarity to known proteins with the aim of assigning their putative function in the biosynthesis of DAQs (Table 1, and Table S27). The putative daq BGCs identified in strains F001 and B006 did not only differ in gene organization but also in gene content (Figure 2 and Table 1). For instance, the BGC in strain F001 contained two genes, daqL and daqO, encoding putative NAD-dependent epimerases while only one, which showed higher similarity with daqO, was found in strain B006. Furthermore, the cluster in B006 contained an additional gene, daqV, which encodes a putative transcriptional regulator, and which is absent in strain F001. Additionally, a second gene encoding a putative β-ketoacyl-ACP synthase III (daqU) was found at one of the borders of the daq BGC in strain F001 but not in strain B006.
Table 1.
Predicted Function of ORFs in the daq BGCs.
| Gene | Locus Tag |
Protein Putative Function(s) |
Identity/ Similarity (%)* |
|
|---|---|---|---|---|
| F001 | B006 | |||
| daqA | StrepF001_25980 | MicB006_2935 | NADPH:quinone reductase | 59/79 |
| daqB | StrepF001_25975 | MicB006_2937 | cyclase | 57/76 |
| daqC | StrepF001_25970 | MicB006_2938 | long-chain acyl-CoA synthetase | 69/86 |
| daqD | StrepF001_25965 | MicB006_2939 | FAD-dependent oxidoreductase/3-hydroxybenzoate 6-monooxygenase | 60/81 |
| daqE | StrepF001_25960 | MicB006_2940 | hypothetical protein with FAD-NAD(P)-binding domain | 69/82 |
| daqF | StrepF001_25955 | MicB006_2941 | lyase family protein | 65/84 |
| daqG | StrepF001_25950 | MicB006_2906 | anthranilate synthase | 57/77 |
| daqH | StrepF001_25945 | MicB006_2898 | isochorismatase | 58/75 |
| daqI | StrepF001_25940 | MicB006_2899 | 2,3-dihydro-2,3-dihydroxybenzoate dehydrogenase | 62/85 |
| daqJ | StrepF001_25935 | MicB006_2892 | DAHP synthase | 48/71 |
| daqK | StrepF001_25930 | MicB006_2930 | transposase | 43/68 |
| daqL | StrepF001_25925 | - | NAD-dependent epimerase/dehydratase | - |
| daqM | StrepF001_25920 | MicB006_2928 | TetR family transcriptional regulator | 59/83 |
| daqN | StrepF001_25915 | MicB006_2931 | ketoacyl-ACP synthase III | 65/88 |
| daqO | StrepF001_25910 | MicB006_2929 | NAD-dependent epimerase/SDR family oxidoreductase | 65/85 |
| daqP | StrepF001_25905 | MicB006_2927 | WrbA, a NADPH:quinone oxidoreductase | 74/88 |
| daqQ | StrepF001_25900 | MicB006_2926 | dehydratase | 52/76 |
| daqR | StrepF001_25895 | MicB006_2932 | decarboxylase | 62/83 |
| daqS | StrepF001_25890 | MicB006_2933 | N-acetyltransferase | 54/79 |
| daqT | StrepF001_25885 | MicB006_2934 | N-acetyltransferase | 57/81 |
| daqU | StrepF001_25880 | - | ketoacyl-ACP synthase III | - |
| daqV | - | MicB006_2936 | PucR family transcriptional regulator | - |
The identity/similarity percentages shown are between proteins from F001 and B006.
More detailed information, including top BLAST hits, can be found in Table S27.
Homologous daq BGCs are Present in Other Sequenced Strains.
We identified homologous daq BGCs in Streptomyces sp. AS58, Streptomyces sp. yr375, Micromonospora sp. TSRI0369, and Micromonospora sp. M42. While the putative daq BGCs identified in the Streptomyces strains F001 and yr375 are identical, the cluster found in strain AS58 differs in gene organization and by the absence of the putative NADPH:quinone oxidoreductase gene daqA (Figure S2A). The daq BGCs found in Micromonospora sp. strains TSRI0369 and M42 are identical to that in strain B006 (Figure S2B), and they are also located in the direct vicinity of a diazepinomicin gene cluster. Genes up- and downstream of the putative daq BGC are different in strains F001 and yr375 as well as in strains B006 and M42, indicating that the boundaries of the BGCs are as shown in Figure 2. Furthermore, the daq BGCs in all three Streptomyces strains contain a second putative β-ketoacyl-ACP synthase III gene (daqU) suggesting that DaqU is indeed part of the BGC and that it is important to the biosynthesis of DAQs produced by Streptomyces sp. F001 and potentially to the other Streptomyces strains, although they haven’t been shown to be DAQ producers. In addition, the BGC of 10 (Figure 1) from Streptomyces albus subsp. chlorinus NRRL B-24108 was recently published (Figure S2C).32 A comparison of the daq and nyb BGCs is presented in Table S28. The genes daqA to daqH, daqJ, and daqL to daqU share sequence identity with nyb genes. No homologue was found for daqI encoding a putative 2,3-dihydro-2,3-dihydroxybenzoate dehydrogenase. Furthermore, the daq BGC contains two genes, daqS and daqT, encoding N-acetyltransferases, while only one was found in the nyb BGC. Finally, the nybomycin BGC encodes one gene, nybM, which shares sequence similarity to the putative β-ketoacyl-ACP synthase III genes daqN/daqU.
Deletion of the daq BGC Abolishes DAQ Production.
We recently established a reverse genetics system for Micromonospora sp. B006 and showed that gene disruption could be performed in this strain using classical homologous recombination via a single crossover.28 By contrast, Streptomyces sp. F001 turned out to be more recalcitrant to genetic engineering. We therefore chose to probe whether the identified region is indeed responsible for the biosynthesis of DAQs by gene cluster deletion in Micromonospora sp. B006. This time, we opted for CRISPR-Cas9 for genome editing. Protospacer sequences were selected within the genes at the start and end of the cluster (MicB006_2924 and MicB006_2941, respectively; Figure 3A). The genome editing design ensured that the diazepinomicin BGC remained intact. A 2-kb editing template was constructed from 1-kb sequences immediately flanking the daq cluster. Following conjugation, five exconjugants were evaluated by PCR using primers that anneal outside of the 1-kb homology arms (Figure 3A). The 3.5-kb band indicative of gene cluster deletion was observed for exconjugants 1, 2, 4, and 5, while no band was observed when wild-type (as expected) and exconjugant 3 genomic DNA were used as templates (Figure 3B). Similarly, PCR amplification with a primer annealing within the deleted region only produced a 2-kb band from wild type and exconjugant 3 (Figure 3C). These results confirmed four of the five tested exconjugants as Adaq mutants. This is only the second report of using CRISPR-Cas9 for genome editing in Micromonospora.33
Figure 3.
Deletion of the putative daq biosynthetic gene cluster abolishes DAQ production in Micromonospora sp. B006. (A) Design of CRISPR-Cas9 genome editing. (B) PCR using primers oJB172/174; expected fragment lengths are 21.5 kb (wt), and 3.5 kb (mutants). (C) PCR using primers oJB140/174; expected fragment lengths are 2 kb (wt), and no band (mutants). (D) HPLC chromatograms, extracted at 280 nm, highlighting the m/z values for DAQ peaks, along with their overlaid UV spectra shown in full scale. The m/z values for DAQ peaks were obtained by UHPLC-QTOFMS (Figure S3). wt, wild type.
The four confirmed Δdaq mutants and the wild-type strain were grown in A1 liquid medium with filtered Lake Michigan water and the MeOH extracts were analyzed by HPLC-PDA and UHPLC-HRESIQTOFMS (Figure 3D and Figure S3). From this analysis, four DAQs were observed from HPLC-PDA analysis of the extract obtained from the wild-type strain. Analogs 8 and 9 were confirmed by comparing HRMS, MS2 fragments, and retention times in UHPLC-MS/MS with purified DAQ H and DAQ J samples isolated from previous studies (Figure S4).2 The remaining two DAQ analogs are putatively new, however they are minor components of the fermentation extract and insufficient material precluded their full structural characterization. Despite this, based on characteristic DAQ MS2 fragmentation data, chemical structures are proposed in Figure S4. The deletion of the identified ~19-kb region in Micromonospora sp. B006 abolished the production of DAQs (Figure 3D), providing experimental evidence for gene cluster assignment.
Biosynthesis Hypothesis.
Based on the genes present in the daq BGC, a working biosynthesis hypothesis was formulated (Figures 4 to 6 and Figure S5). The proposed model is in accordance with previous precursor labeling studies which showed incorporation of labeled D-glucose, pyruvate and D-erythrose consistent with the central ring carbons of the related compound 10 (Figure 1) originating from a shikimate-type pathway.27 Moreover, labeling studies established acetate as the source of the side ring carbons of 10.26
Figure 4.
Proposed pathway for biosynthesis of the diazaquinomycin central core. The first four steps involving DaqJ, DaqG, DaqH and DaqI and leading to 3-hydroxy-anthranilic acid are according to the proposal for diazepinomicin biosynthesis by McAlpine et al.34 See text for further details.
Figure 6.
Proposed final steps in diazaquinomycin biosynthesis.
According to the hypothesis presented in Figure 4, biosynthesis of DAQ’s central ring would start with DAHP synthase DaqJ. DAHP synthase catalyzes the first committed step of the shikimate pathway, that is, the condensation of the glycolysis pathway intermediate phosphoenolpyruvate and the pentose phosphate pathway intermediate D-erythrose-4-phosphate. Strict feedback regulation of housekeeping DAHP synthase ensures appropriate flux through the shikimate pathway for aromatic amino acid biosynthesis. DAHP synthase is known to be regulated at the gene and enzyme levels. For instance, allosteric regulation by end products phenylalanine, tyrosine and tryptophan leads to loss in enzyme activity35, 36 As such, BGCs for compounds that rely on the shikimate pathway usually contain a gene encoding a DAHP synthase isoenzyme that can bypass feedback regulation by aromatic amino acids.37–39
In Micromonospora sp. B006, daqJ is present in the diazepinomicin BGC (Figure 2). In addition to daqJ, three other genes are shared between the daq and the diazepinomicin BGCs, anthranilate synthase daqG, isochorismatase daqH, and 2,3-dihydro-2,3-dihydroxybenzoate dehydrogenase daqI. According to the proposal by McAlpine et al.34 the corresponding proteins ORF33, ORF19 and ORF27 of Micromonospora sp. DPJ12, respectively, catalyze the conversion of chorismic acid to 3-hydroxyanthranilic acid as shown in Figure 4. 3-hydroxyanthranilic acid is then activated by adenylation, followed by condensation with aminohydroxy[1,4]benzoquinone to form the dibenzodiazepinone core, which after prenylation results in diazepinomicin (Figure S5).34,40
We propose that 3-hydroxyanthranilic acid is the branching point between DAQ and diazepinomicin biosynthesis as shown in Figure S5. 3-hydroxyanthranilic acid would be converted to the 2,6-diaminoquinone central ring of DAQs in a series of steps. For instance, the daq BGC contains a gene, daqD, that shows 85% sequence identity to 3-hydroxybenzoate 6-monooxygenases.41 Accordingly, we propose that DaqD catalyzes hydroxylation at C-6 of 3-hydroxyanthranilic acid. Further, although DaqF shows sequence similarity to enzymes putatively annotated as an adenylosuccinate lyase family protein or a 3-carboxy-cis, cis-muconate cycloisomerase, it also contains a lyase I domain. The class I lyase superfamily includes phenylalanine, tyrosine and histidine ammonia lyases, which catalyze the reversible addition of ammonia to a double bond (cinnamic acid, p-hydroxycinnamic acid and urocanic acid, respectively).42–44 Thus, we tentatively propose that DaqF catalyzes amination at C-4, which followed by re-aromatization, possibly involving DaqE and/or DaqO, may lead to 2,4-diamino-3,6-dihydroxybenzoic acid. DaqE contains an uncharacterized FAD-NAD(P) binding domain at its N-terminus and a conserved domain of unknown function (DUF4175) at its C-terminus. DaqO contains a NAD binding domain and appears to belong to the short-chain dehydrogenase/reductase superfamily. A PDB search with DaqO identified an imine reductase as a potential homolog.45
Next, DaqR shows a metallo-dependent hydrolase fold and sequence similarity to proteins putatively annotated as amidohydrolases. In addition, DaqR shows 40% sequence identity to an enzyme from Burkholderia cepacia that has been recently shown to be a novel member of the amidohydrolase 2 family that catalyzes the nonoxidative decarboxylation of 2-hydroxy-1-naphthoic acid.46 Thus, we propose that DaqR catalyzes decarboxylation of 2,4-diamino-3,6-dihydroxybenzoic acid to form the 2,6-diaminobenzene-1,4-diol that is later incorporated as the central quinone core of DAQs.
Regarding the biosynthesis of the side rings, the gene daqC encodes a putative long-chain acyl-CoA synthetase that is proposed to be responsible for activation of acyl substrates (Figure 5). Actinomycetes are known to biosynthesize both straight and branched-chain fatty acids.47 It is conceivable that differential preference of DaqCF001 and DaqCB006 for straight and branched-chain fatty acids, respectively, may help explain the different DAQ analogs produced by each strain. The substrate preferences of enzymes downstream in the pathway may play a role as well. Next, it is interesting to note that the daq BGC in strain B006 contains only one β-ketoacyl-ACP synthase III (KASIM) gene, daqN, whereas two KASIII genes, daqN and daqU are found in strain F001. DaqN and DaqU would catalyze the Claisen condensation of an acyl-CoA unit with malonyl-or methylmalonyl-ACP. Based on the observation that DAQ congeners produced by strain B006 lack methyl groups at C-3 and C-6 and that DAQ congeners produced by strain F001 contain a methyl group at C-3 and may contain a methyl group at C-6, we hypothesize that DaqN prefers malonyl-ACP as a substrate, whereas DaqU prefers methylmalonyl-ACP (Figure 5). Another point to note is that there are no ACP genes present in the daq BGC. Given that KASIII enzymes are known to accept ACP-bound substrates,48, 49 (methyl)malonyl-ACP from primary metabolism (fatty acid biosynthesis) may be co-opted for DAQ biosynthesis. Alternatively, it is also conceivable that CoA-activated substrates may be used instead.50
Figure 5.
Proposed pathway for biosynthesis of diazaquinomycin’s side rings. Proposed model to explain the molecular basis for the distinct DAQ derivatives produced by Micromonospora sp. B006 and Streptomyces sp. F001. We hypothesize that DaqNB006 preferentially accepts malonyl-ACP (or malonyl-CoA) and a branched-chain fatty acyl-CoA or acetyl-CoA monomer as substrates, whereas DaqNF001 preferentially accepts malonyl-ACP (or malonyl-CoA) and a straight-chain fatty acyl-CoA monomer. DaqU, which is present only in strain F001 would preferentially accept methylmalonyl-ACP (or methylmalonyl-CoA). CoA, coenzyme A. ACP, acyl carrier protein.
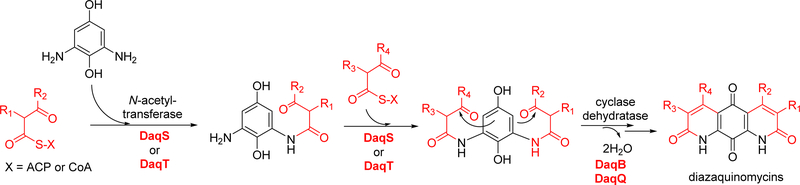
The resulting β-ketoacyl units would then be condensed with the primary amines of 2,6-diaminohydroquinone as catalyzed by the activity of two putative N-acetyltransferases DaqS and DaqT (Figure 6). Finally, the putative polyketide cyclase DaqB would catalyze cyclization, followed by dehydration, presumably catalyzed by DaqQ which appears to belong to the nuclear transport factor 2 family of proteins that includes dehydratases. Reactions presumably catalyzed by DaqB and DaqQ would be the enzymatic equivalents of double Knorr quinoline synthetic routes reported for DAQs.22, 24, 51
Regulatory and Unknown Genes.
In addition to structural genes, the daq BGC contains a TetR-family transcriptional regulator gene, daqM, in both actinomycetes and a putative PucR-like transcriptional regulator in the BGC from strain B006 only, daqV. Moreover, both BGCs contain two genes that show sequence similarity to quinone oxidoreductases. DaqP shows sequence similarity to the type IV, two-electron, FMN-dependent NAD(P)H:quinone oxidoreductase WrbA, which has been shown to be implicated in oxidative stress defense.52–54 For instance, deletion of WrbA-like proteins in Candida albicans lend them more sensitive to benzoquinone.55 DaqA shows sequence similarity to the zeta-crystallin type of one-electron quinone oxidoreductases. This type of quinone oxidoreductase has also been proposed to play a role in quinone detoxification. In addition, binding of human and yeast zeta-crystallin to mRNA has also been reported with a possible role in gene expression.56–58 The roles of DaqA and DaqP remain to be demonstrated.
Conclusions.
Members of the diazaquinomycin class of natural products have shown potent and selective inhibitory activity against a panel of drug-resistant M. tuberculosis strains, a pathogen of global concern.2 The genome of the marine-derived actinomycete bacterium Streptomyces sp. F001, a producer of DAQ A (1) and DAQ E-G (5–7),6 was analyzed using automated software tools, revealing 24 BGCs, none of which could be assigned to DAQs (Table S1). The genome of Streptomyces sp. F001 was then compared with that of the Lake Michigan-derived Micromonospora sp. B006,28 a producer of DAQ H (8) and J (9).7 An approximately 19kb region that is shared between both strains was identified as a putative daq BGC (Figure 2). We then provided experimental evidence, through BGC deletion mediated by CRISPR-Cas9, that the identified BGC in strain B006 is indeed responsible for the biosynthesis of DAQs (Figure 3). Based on the genes identified, we proposed a working hypothesis for DAQ biosynthesis (Figures 4 to 6).
Genome mining approaches rely on current knowledge of secondary metabolite biosynthesis and on BGC databases such as the Minimum Information about a Biosynthetic Gene cluster (MIBiG) repository.59, 60 The fact that DAQ biosynthesis resembles primary metabolism (shikimate pathway and fatty acid biosynthesis) may explain why automated software tools were unsuccessful in detecting daq BGCs in the genomes of strains F001 and B006. The daq BGCs identified here through comparative genomics will allow not only elucidation of DAQ biosynthesis for basic and applied purposes, but will also facilitate detection of related BGCs in the future.
MATERIAL AND METHODS
General Experimental Procedures.
All chemicals were acquired from Sigma-Aldrich, Alfa Aesar, VWR, and Fisher Scientific. Solvents were of HPLC grade or higher. Restrictions enzymes were purchased from New England Biolabs. Oligonucleotide primers were synthesized by Sigma-Aldrich. Molecular biology procedures were carried out according to the manufactures’ instructions (New England Biolabs, Thermo Fisher Scientific, Qiagen, and Zymo Research).
Strains and Cultivation Conditions.
Streptomyces sp. F001 was isolated from a sediment sample from Raja Ampat, Birds Head, Papua, Indonesia (0°33’54.72“S, 130°40’35.04”E) in March 2011.29 The strain was routinely cultivated on A1 medium (33.3 g instant ocean sea salt (Marineland), 10 g starch, 4 g yeast extract, 2 g peptone, 1 g calcium carbonate, 100 mg potassium bromide, and 40 mg iron sulfate per liter A1 medium, and 20 g agar for solid medium) at 30 °C.
Micromonospora sp. strain B006 was isolated from a Lake Michigan sediment sample collected by PONAR at a depth of 56 m, from approx. 16.5 miles off the coast of north Milwaukee, WI, USA (43°13’27.0“N, 87°34’12.0”W) on August 23rd, 2010. For genomic DNA isolation and the preparation of frozen stocks, the wild-type strain and the obtained exconjugants were cultivated in TSB medium (3% tryptic soy broth) for three days at 30 °C and 200 rpm. The cultures were inoculated with a loopful of cell material from five-day old pure cultures grown on A1 medium plates (0.5 L of filtered Lake Michigan water, 0.5 L DI H2O, 10 g starch, 4 g yeast extract, 2 g peptone, 1 g calcium carbonate, 100 mg potassium bromide, and 40 mg iron sulfate per liter A1 medium).2 Frozen stocks were prepared with mycelium from three-day old ISP2 (0.4% yeast extract, 1% malt extract, 0.4% dextrose, pH 7.3) liquid cultures by adding glycerol to 20% [v/v] final concentration followed by storage at −80 °C.
E. coli strain ET12567/pUZ8002 was cultivated in LB medium supplemented with the appropriate antibiotics. The following antibiotics were used as selection markers: apramycin (final concentration: 50 μg/mL), kanamycin (50 μg/mL), chloramphenicol (25 μg/mL), and nalidixic acid (25 μg/mL).
Bioinformatics Analysis.
The genome of strain F001 was sequenced using Illumina and Pacific Biosciences (PacBio) technologies.29 CDS were detected using RAST.61–63 Ribosomal RNA genes and transfer RNA were detected by using RAST, the RNAmmer 1.2 software64 and tRNAscan-SE 2.065. Biosynthetic gene clusters and possible encoded compounds were roughly predicted with antiSMASH66 and BAGEL467, and further annotated using BLAST68 followed by manual curation. A multi-locus species tree was generated using autoMLST in placement mode (http://automlst.ziemertlab.com).69 The genome of strain B006 was sequenced using Illumina and Oxford Nanopore technologies and analyzed as previously reported.28 Multiple whole genome sequence alignment of Streptomyces sp. F001 (GenBank accession: QZWF00000000; SRA accession: PRJNA483497), Micromonospora sp. B006 (GenBank accession: CP030865), and Streptomyces coelicolor A3(2) (GenBank accession: GCA_000203835.1) was performed using progressive MAUVE (version 2.4.0) with default settings.30 The daq BGCs were deposited in MIBiG under accession codes BGC0001848 (for strain B006) and BGC0001850 (for strain F001).
Deletion of the Diazaquinomycin BGC in Micromonospora sp. Strain B006.
For multiplex gene deletion in Micromonospora sp. B006, two sgRNA cassettes targeting two different positions outside the putative DAQ gene cluster were designed (Figure 3). The spacer sequences were identified using the CRISPR tool in GeneiousR9 (version 9.1.8). A synthetic DNA fragment was designed (5’-GTC TTC TGC CGC GTA CTC CAG ATA GCG TTT TAG AGC TAG AAA TAG CAA GTT AAA ATA AGG CTA GTC CGT TAT CAA CTT GAA AAA GTG GCA CCG AGT CGG TGC TAA ACC GAT ACA ATT AAA GGC TCC TTT TGG AGC CTT TTT TTG CTG CTC CTT CGG TCG GAC GTG CGT CTA CGG GCA CCT TAC CGC AGC CGT CGG CTG TGC GAC ACG GAC GGA TCG GGC GAA CTG GCC GAT GCT GGG AGA AGC GCG CTG CTG TAC GGC GCG CAC CGG GTG CGG AGC CCC TCG GCG AGC GGT GTG AAA CTT CTG TGA ATG GCC TGT TCG GTT GCT TTT TTT ATA CGG CTG CCA GAT AAG GCT TGC AGC ATC TGG GCG GCT ACC GCT ATG ATC GGG GCG TTC CTG CAA TTC TTA GTG CGA GTA TCT GAA AGG GGA TAC GCA TCG GGG CGA CCA GGA GGG GGT TTT AGA GCT AGA AAT AGC AAG TTA AAA TAA GGC TAG TCC GTT ATC AAC TTG AAA AAG TGG CAC CGA GTC GGT GCT TTT TAC TCC ATC TGG ATT TGT TCA GAA CGC TCG GTT GCC GCC GGG CGT TTT TTA TCT AGA-3’) and ordered from GenScript. The two restriction sites BbsI and XbaI (underlined) were used to clone the synthetic DNA fragment into the same sites of the vector pCRISPomyces-2,70 yielding plasmid pJB026EL.
Primer pairs oJB104/oJB105 and oJB106/oJB107 were used to amplify two 1-kb homology arms from genomic DNA isolated from strain B006 (Table S29). Genomic DNA was isolated from Micromonospora sp. B006 using the GenElute Bacterial Genomic DNA Kit (Sigma-Aldrich). The 50 μL PCR reactions consisted of 0.2 mM of each dNTP, 3% DMSO, 0.25 μm of each primer, and 1 U/μL Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific) in HF reaction buffer supplied with the enzyme. Thermocycling parameters were initial denaturation for 2 min at 98 °C; amplification: 30 cycles (98 °C for 10 s, 69 °C for 30 s, 72 °C for 30 s); and terminal hold for 5 min at 72 °C. The two obtained PCR fragments were ligated by overlap extension (OE) PCR using primers oJB104 and oJB107 (Table S29). The 50-μL reaction consisted of the two purified DNA fragments in equimolar amounts, 0.2 mM of each dNTP, 3% DMSO, and 1 U/μL Phusion High-Fidelity DNA Polymerase in HF reaction buffer supplied with the enzyme. Thermocycling conditions were initial denaturation: 60 s at 98 °C; amplification: 3 cycles (98 °C for 10 s, 72 °C for 30 s, 72 °C for 60 s). Subsequently, 0.25 mM of each primer were added to the reaction. Thermocycling parameters were initial denaturation for 60 s at 98 °C; amplification: 30 cycles (98 °C for 10 s, 69.8 °C for 30 s, 72 °C for 60 s); and terminal hold for 5 min at 72 °C. The primers oJB104 and oJB107 introduced the restriction site Xbal into the 2 kb PCR product, which was cloned into the same site of plasmid pJB026EL to generate plasmid pJB027EL.
Subsequently, plasmid pJB027EL was transferred into Micromonospora sp. strain B006 by conjugation from E. coli ET12567/pUZ8002 as described earlier.28 Apramycin-resistant colonies were streaked on A1 medium plates containing 50 μg/mL apramycin and 25 μg/mL nalidixic acid until pure cultures were obtained. Genomic DNA was isolated using the GenElute Bacterial Genomic DNA Kit (Sigma-Aldrich). Gene cluster deletion was confirmed by PCR with primer pairs oJB140/oJB174 and oJB172/oJB174 (Table S29). The 20 μL reactions for primer pair oJB140/174 contained 0.2 mM of each dNTP, 3% DMSO, 0.25 μM of each primer, and 1.25 U DreamTaq DNA Polymerase (Thermo Fisher Scientific) in reaction buffer supplied with the enzyme. Thermocycling parameters were initial denaturation for 2 min at 95 °C; amplification: 30 cycles (95 °C for 30 s, 57.1 °C for 30 s, 72 °C for 60 s); and terminal hold for 5 min at 72 °C. The 25μL reactions with primers oJB172 and oJB174 consisted of 0.2 mM of each dNTP, 1× Q5 High GC Enhancer, 0.5 μM of each primer, and 0.02 U/μL Q5 High-Fidelity DNA Polymerase (New England Biolabs) in Q5 reaction buffer supplied with the enzyme. The following thermal cycling conditions were used: 30 s at 98 °C; 30 cycles of 98 °C for 10 s; and 72 °C for 75 s; and a terminal hold at 72 °C for 2 min.
Fermentation and Metabolite Analysis.
For DAQ production, 50 mL TSB liquid medium was inoculated with 200 μL of a frozen stock of Micromonospora sp. strain B006 wild-type strain and mutants, respectively. The seed cultures were incubated for three days at 30 °C and 200 rpm. Subsequently, 5% [v/v] of these seed cultures were used to inoculate 50 mL A1 liquid medium in filtered Lake Michigan water2 in a 250 mL Erlenmeyer flask. The cultures were incubated at 30 °C and 200 rpm. After 7 days, 5% conditioned Diaion HP-20 resin was added to the production cultures. The HP-20 resin was conditioned prior to use by soaking in MeOH and then rinsing thoroughly with distilled H2O. The production cultures were incubated for another 24 h at 30 °C and 200 rpm. Subsequently, the cultures were harvested by centrifugation. The supernatant was decanted and the cell/resin pellet was extracted three times with 20 mL MeOH each. After removing the solvent under reduced pressure, the extracts were dissolved in MeOH for HPLC analysis.
HPLC analysis was performed on an Agilent 1260 Infinity system equipped with a Kinetex C18 column (150 × 4.6 mm, 5 μm particle size, 100 Å pore size, Phenomenex). Solvent A was 0.1% [v/v] trifluoroacetic acid (TFA) in H2O, and solvent B was CH3CN. Method: isocratic flow of 50% B for 2 min; linear gradient from 50% to 100% B from 2–10 min; isocratic flow of 100% B for 5 min. The detection wave length range was 200 – 600 nm; chromatograms were analyzed at λ = 280 nm.
MS analyses were performed on a Bruker impact II Q-TOF (quadrupole time-of-flight) mass spectrometer (Thermo Electron Corporation) in positive mode and a Shimadzu Nexera X2 UHPLC, equipped with a Kinetex C18 column (50 × 2.1 nm, 1.7 μm particle size, 100 Å pore size, Phenomenex), at a flow rate of 0.5 mL/min. Solvent A was 0.1% [v/v] formic acid in H2O, and solvent B was 0.1% [v/v] formic acid in CH3CN. The gradient was: initial hold at 5% B for 0.1 min, linear gradient from 5% to 100% B within 6 min, and held for 2 min. The detection mass range was 50 Da to 1,500 Da (positive mode).
Supplementary Material
ACKNOWLEDGEMENTS
We thank G. Pauli (University of Illinois at Chicago) and N. Ziemert (University of Tübingen) for access to LC/MS instrumentation, and suggestions regarding MAUVE alignments, respectively. We also thank H. Zhao (University of Illinois at Urbana-Champaign) for pCRISPomyces-2 (via Addgene, plasmid #61737). Financial support for this work was provided by the National Center for Advancing Translational Sciences, National Institutes of Health (NIH), under grant KL2TR002002 (to ASE), and by startup funds from the Department of Medicinal Chemistry and Pharmacognosy and the Center for Biomolecular Sciences, University of Illinois at Chicago (to ASE). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Footnotes
ASSOCIATED CONTENT
Supporting Information
The Supporting Information is available free of charge in the ACS Publications website at DOI:10.1021/acs.jnatprod.8b01028.
Biosynthetic gene clusters (BGCs) identified in Streptomyces sp. F001 and Protein BLAST statistics of those BGCs (Supplementary Results and Tables S1 to S26), top BLAST hits of ORFs identified in daq clusters (Table S27), comparison of the nybomycin BGC with the ORFs identified in the daq BGCs of Streptomyces sp. F001 and Micromonospora sp. B006, respectively (Table S28), list of oligonucleotide primers used (Table S29), multi-locus species tree (Figure S1), comparison of daq BGCs in strains F001 and B006 with homologous clusters in other actinomycete strains as well as the nybomycin BGC (Figure S2), MS and MS/MS data for 8 and 9 as well as the uncharacterized DAQs (Figures S3 and S4), and proposed model for biosynthesis of diazaquinomycin and diazepinomicin in strain B006 (Figure S5). (PDF)
REFERENCES
- (1).Tiberi S; du Plessis N; Walzl G; Vjecha MJ; Rao M; Ntoumi F; Mfinanga S; Kapata N; Mwaba P; McHugh TD; Ippolito G; Migliori GB; Maeurer MJ; Zumla A. Lancet Infect. Dis 2018, 18, e183–e198. [DOI] [PubMed] [Google Scholar]
- (2).Mullowney MW; Hwang CH; Newsome AG; Wei X; Tanouye U; Wan B; Carlson S; Barranis NJ; O hAinmhire E; Chen WL; Krishnamoorthy K; White J; Blair R; Lee H; Burdette JE; Rathod PK; Parish T; Cho S; Franzblau SG; Murphy BT ACS Infect. Dis 2015, 1, 168–174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (3).Omura S; Iwai Y; Hinotozawa K; Tanaka H; Takahashi Y; Nakagawa AJ Antibiot 1982, 35, 1425–1429. [DOI] [PubMed] [Google Scholar]
- (4).Omura S; Nakagawa A; Aoyama H; Hinotozawa K; Sano H. Tetrahedron Lett 1983, 12, 3643–3646. [Google Scholar]
- (5).Maskey RP; Grün-Wollny I; Laatsch H Nat. Prod. Res 2005, 19, 137–142. [DOI] [PubMed] [Google Scholar]
- (6).Mullowney MW; O hAinmhire E; Shaikh A; Wei X; Tanouye U; Santarsiero BD; Burdette JE; Murphy BT Mar. Drugs 2014, 12, 3574–3586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (7).Shaikh AF; Elfeki M; Landolfa S; Tanouye U; Green SJ; Murphy BT Nat. Prod. Sci 2015, 21, 261–267. [Google Scholar]
- (8).Murata M; Miyasaka T; Tanaka H; Omura SJ Antibiot 1985, 38, 1025–1033. [DOI] [PubMed] [Google Scholar]
- (9).Mullowney MW Antibiotics from Aquatic-Derived Actinomycete Bacteria that Inhibit M. tuberculosis. Ph.D. Dissertation, University of Illinois at Chicago, Chicago, IL, 2016. [Google Scholar]
- (10).Rinehart KL Jr.; Renfroe HB J. Am. Chem. Soc 1961, 83, 3729–3731. [Google Scholar]
- (11).Rinehart KL Jr.; Leadbetter G; Larson RA; Forbis RM J. Am. Chem. Soc 1970, 92, 6994–6995. [DOI] [PubMed] [Google Scholar]
- (12).Chu M; Mierzwa R; Xu L; Yang S-W; He L; Patel M; Stafford J; Macinga D; Black T; Chan T-M; Gullo V. Bioorganic Med. Chem. Lett 2003, 13, 3827–3829. [DOI] [PubMed] [Google Scholar]
- (13).Bair JS; Palchaudhuri R; Hergenrother PJ J. Am. Chem. Soc 2010, 132, 5469–5478. [DOI] [PubMed] [Google Scholar]
- (14).Strelitz F; Flon H; Asheshov IN Proc. Natl. Acad. Sci. U.S.A 1955, 41, 620–624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (15).Arai M; Kamiya K; Pruksakorn P; Sumii Y; Kotoku N; Joubert JP; Moodley P; Han C; Shin D; Kobayashi M. Bioorg. Med. Chem. Lett 2015, 23, 3534–3541. [DOI] [PubMed] [Google Scholar]
- (16).Naganawa H; Wakashiro T; Yagi A; Kondo S; Takita TJ Antibiot 1970, 23, 365–368. [DOI] [PubMed] [Google Scholar]
- (17).Hiramatsu K; Igarashi M; Morimoto Y; Baba T; Umekita M; Akamatsu Y. Int. J. Antimicrob. Agents 2012, 39, 478–485. [DOI] [PubMed] [Google Scholar]
- (18).Pettit GR; Du J; Pettit RK; Richert LA; Hogan F; Mukku VJRV; Hoard MS J. Nat. Prod 2006, 69, 804–806. [DOI] [PubMed] [Google Scholar]
- (19).Li S; Tian X; Niu S; Zhang W; Chen Y; Zhang H; Yang X; Zhang W; Li W; Zhang S; Ju J; Zhang C. Mai’. Drugs 2011, 9, 1428–1439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (20).Tian XP; Long LJ; Li SM; Zhang J; Xu Y; He J; Li J; Wang FZ; Li WJ; Zhang CS; Zhang S. Int. J. Syst. Evol. Microbiol 2013, 63, 893–899. [DOI] [PubMed] [Google Scholar]
- (21).Parkinson EI; Bair JS; Cismesia M; Hergenrother PJ ACS Chem. Biol 2013, 8, 2173–2183. [DOI] [PubMed] [Google Scholar]
- (22).Kelly TR; Field JA; Li Q. Tetrahedron Lett 1988, 29, 3545–3546. [Google Scholar]
- (23).Pérez JM; López-Alvarado P; Avendano C; Menéndez JC Tetrahedron Lett 1998, 39, 673–676. [Google Scholar]
- (24).Prior A; Sun D. Synthesis 2018, 50, 859–871. [Google Scholar]
- (25).Pérez JM; López-Alvarado P; Pascual-Alfonso E; Avendano C; Ménendez JC Tetrahedron 2000, 56, 4574–4583. [Google Scholar]
- (26).Knoell WM; Huxtable RJ; Rinehart KL Jr. J. Am. Chem. Soc 1973, 95, 2703–2704. [DOI] [PubMed] [Google Scholar]
- (27).Nadzan AM; Rinehart KL Jr. J. Am. Chem. Soc 1976, 98, 5012–5014. [DOI] [PubMed] [Google Scholar]
- (28).Braesel J; Crnkovic CM; Kunstman KJ; Green SJ; Maienschein-Cline M; Orjala J; Murphy BT; Eustaquio AS J. Nat. Prod 2018, 81, 2057–2068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (29).Braesel J; Clark CM; Kunstman KJ; Green SJ; Maienschein-Cline M; Murphy BT; Eustaquio AS submitted to Microbiology Resource Announcement 2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (30).Darling AE; Mau B; Perna NT PLoS One 2010, 5, e11147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (31).Bentley SD; Chater KF; Cerdeno-Tarraga AM; Challis GL; Thomson NR; James KD; Harris DE; Quail MA; Kieser H; Harper D; Bateman A; Brown S; Chandra G; Chen CW; Collins M; Cronin A; Fraser A; Goble A; Hidalgo J; Hornsby T; Howarth S; Huang CH; Kieser T; Larke L; Murphy L; Oliver K; O’Neil S; Rabbinowitsch E; Rajandream MA; Rutherford K; Rutter S; Seeger K; Saunders D; Sharp S; Squares R; Squares S; Taylor K; Warren T; Wietzorrek A; Woodward J; Barrell BG; Parkhill J; Hopwood DA Nature 2002, 417, 141–147. [DOI] [PubMed] [Google Scholar]
- (32).Estevez MR; Myronovskyi M; Gummerlich N; Nadmid S; Luzhetskyy A. Mar. Drugs. 2018, 16, 435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (33).Cohen DR; Townsend CA Nat. Chem 2018, 10, 231–236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (34).McAlpine JB; Banskota AH; Charan RD; Schlingmann G; Zazopoulos E; Piraee M; Janso J; Bernan VS; Aouidate M; Farnet CM; Feng X; Zhao Z; Carter GT J. Nat. Prod 2008, 71, 1585–1590. [DOI] [PubMed] [Google Scholar]
- (35).Webby CJ; Jiao W; Hutton RD; Blackmore NJ; Baker HM; Baker EN; Jameson GB; Parker EJ J. Biol. Chem 2010, 285, 30567–30576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (36).Blackmore NJ; Reichau S; Jiao W; Hutton RD; Baker EN; Jameson GB; Parker EJ J. Mol. Biol 2013, 425, 1582–1592. [DOI] [PubMed] [Google Scholar]
- (37).Pierson LSI; Gaffney T; Lam S; Gong F. FEMS Microbiol. Lett 1995, 134, 299–307. [DOI] [PubMed] [Google Scholar]
- (38).August PR; Tang L; Yoon y. J.; Ning S; Müller R; Yu TW; Taylor M; Hoffmann D; Kim CG; Zhang X; Hutchinson CR; Floss HG Chem. Biol 1998, 5, 69–79. [DOI] [PubMed] [Google Scholar]
- (39).Kloosterman H; Hessels GI; Vrijbloed JW; Euverink GJ; Dijkhuizen L. Microbiology 2003, 149, 3321–3330. [DOI] [PubMed] [Google Scholar]
- (40).Bonitz T; Zubeil F; Grond S; Heide L. PLoS One 2013, 8, e85707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (41).Montersino S; Orru R; Barendregt A; Westphal AH; van Duijn E; Mattevi A; van Berkel WJ J. Biol. Chem 2013, 288, 26235–26245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (42).Poppe L; Retey J. Angew. Chem. Int. Ed. Engl 2005, 44, 3668–3688. [DOI] [PubMed] [Google Scholar]
- (43).Watts KT; Mijts BN; Lee PC; Manning AJ; Schmidt-Dannert C. Chem. Biol 2006, 13, 1317–1326. [DOI] [PubMed] [Google Scholar]
- (44).MacDonald MJ; D’Cunha GB Biochem. Cell Biol 2007, 85, 273–282. [DOI] [PubMed] [Google Scholar]
- (45).Aleku GA; Man H; France SP; Leipold F; Hussain S; Toca-Gonzalez L; Marchington R; Hart S; Turkenburg JP; Grogan G; Turner NJ ACS Catalysis 2016, 6, 3880–3889. [Google Scholar]
- (46).Pal Chowdhury P; Basu S; Dutta A; Dutta TK J. Bacteriol 2016, 198, 1755–1763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (47).Wallace KK; Zhao B; McArthur HAI; Reynolds KA FEMS Microbiol. Lett 1995, 131, 227–234. [DOI] [PubMed] [Google Scholar]
- (48).Tsay JT; Oh W; Larson TJ; Jackowski S; Rock CO J. Biol. Chem 1992, 267, 6807–6814. [PubMed] [Google Scholar]
- (49).Lai CY; Cronan JE J. Biol. Chem 2003, 278, 51494–51503. [DOI] [PubMed] [Google Scholar]
- (50).Okamura E; Tomita T; Sawa R; Nishiyama M; Kuzuyama T. Proc. Natl. Acad. Sci. U.S.A 2010, 107, 11265–11270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (51).Prior AM; Sun D. RSC Advances 2019, 9, 1759–1771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (52).Kishko I; Harish B; Zayats V; Reha D; Tenner B; Beri D; Gustavsson T; Ettrich R; Carey J. PLoS One 2012, 7, e43902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (53).Kishko I; Carey J; Reha D; Brynda J; Winkler R; Harish B; Guerra R; Ettrichova O; Kukacka Z; Sheryemyetyeva O; Novak P; Kuty M; Kuta Smatanova I; Ettrich R; Lapkouski M. Acta Crystallogr. D. Biol. Crystallogr 2013, 69, 1748–1757. [DOI] [PubMed] [Google Scholar]
- (54).Green LK; La Flamme AC; Ackerley DF J. Microbiol 2014, 52, 771–777. [DOI] [PubMed] [Google Scholar]
- (55).Li L; Naseem S; Sharma S; Konopka JB PLoS Pathog 2015, 11, e1005147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (56).Maruyama A; Kumagai Y; Morikawa K; Taguchi K; Hayashi H; Ohta T. Microbiology 2003, 149, 389–398. [DOI] [PubMed] [Google Scholar]
- (57).Porte S; Crosas E; Yakovtseva E; Biosca JA; Farres J; Fernandez MR; Pares X. Chem. Biol. Interact 2009, 178, 288–294. [DOI] [PubMed] [Google Scholar]
- (58).Crosas E; Sumoy L; Gonzalez E; Diaz M; Bartolome S; Farres J; Pares X; Biosca JA; Fernandez MR FEBS J 2015, 282, 1953–1964. [DOI] [PubMed] [Google Scholar]
- (59).Medema MH; Kottmann R; Yilmaz P; Cummings M; Biggins JB; Blin K; de Bruijn I; Chooi YH; Claesen J; Coates RC; Cruz-Morales P; Duddela S; Dusterhus S; Edwards DJ; Fewer DP; Garg N; Geiger C; Gomez-Escribano JP; Greule A; Hadjithomas M; Haines AS; Helfrich EJ; Hillwig ML; Ishida K; Jones AC; Jones CS; Jungmann K; Kegler C; Kim HU; Kotter P; Krug D; Masschelein J; Melnik AV; Mantovani SM; Monroe EA; Moore M; Moss N; Nutzmann HW; Pan G; Pati A; Petras D; Reen FJ; Rosconi F; Rui Z; Tian Z; Tobias NJ; Tsunematsu Y; Wiemann P; Wyckoff E; Yan X; Yim G; Yu F; Xie Y; Aigle B; Apel AK; Balibar CJ; Balskus EP; Barona-Gomez F; Bechthold A; Bode HB; Borriss R; Brady SF; Brakhage AA; Caffrey P; Cheng YQ; Clardy J; Cox RJ; De Mot R; Donadio S; Donia MS; van der Donk WA; Dorrestein PC; Doyle S; Driessen AJ; Ehling-Schulz M; Entian KD; Fischbach MA; Gerwick L; Gerwick WH; Gross H; Gust B; Hertweck C; Hofte M; Jensen SE; Ju J; Katz L; Kaysser L; Klassen JL; Keller NP; Kormanec J; Kuipers OP; Kuzuyama T; Kyrpides NC; Kwon HJ; Lautru S; Lavigne R; Lee CY; Linquan B; Liu X; Liu W; Luzhetskyy A; Mahmud T; Mast Y; Mendez C; Metsa-Ketela M; Micklefield J; Mitchell DA; Moore BS; Moreira LM; Muller R; Neilan BA; Nett M; Nielsen J; O’Gara F; Oikawa H; Osbourn A; Osburne MS; Ostash B; Payne SM; Pernodet JL; Petricek M; Piel J; Ploux O; Raaijmakers JM; Salas JA; Schmitt EK; Scott B; Seipke RF; Shen B; Sherman DH; Sivonen K; Smanski MJ; Sosio M; Stegmann E; Sussmuth RD; Tahlan K; Thomas CM; Tang Y; Truman AW; Viaud M; Walton JD; Walsh CT; Weber T; van Wezel GP; Wilkinson B; Willey JM; Wohlleben W; Wright GD; Ziemert N; Zhang C; Zotchev SB; Breitling R; Takano E; Glöckner FO Nat. Chem. Biol 2015, 11, 625–631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (60).Epstein SC; Charkoudian LK; Medema MH Stand. Genomic Sci 2018, 13, 16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (61).Aziz RK; Bartels D; Best AA; DeJongh M; Disz T; Edwards RA; Formsma K; Gerdes S; Glass EM; Kubal M; Meyer F; Olsen GJ; Olson R; Osterman AL; Overbeek RA; McNeil LK; Paarmann D; Paczian T; Parrello B; Pusch GD; Reich C; Stevens R; Vassieva O; Vonstein V; Wilke A; Zagnitko O. BMC Genomics 2008, 9, 75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (62).Overbeek R; Olson R; Pusch GD; Olsen GJ; Davis JJ; Disz T; Edwards RA; Gerdes S; Parrello B; Shukla M; Vonstein V; Wattam AR; Xia F; Stevens R. Nucleic Acids Res 2014, 42, D206–214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (63).Brettin T; Davis JJ; Disz T; Edwards RA; Gerdes S; Olsen GJ; Olson R; Overbeek R; Parrello B; Pusch GD; Shukla M; Thomason JA 3rd; Stevens R; Vonstein V; Wattam AR; Xia F. Sci. Rep 2015, 5, 8365. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (64).Lagesen K; Hallin P; Rodland EA; Staerfeldt HH; Rognes T; Ussery DW Nucleic Acids Res 2007, 35, 3100–3108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (65).Lowe TM; Chan PP Nucleic Acids Res 2016, 44, W54–57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (66).Blin K; Wolf T; Chevrette MG; Lu X; Schwalen CJ; Kautsar SA; Suarez Duran HG; de Los Santos ELC; Kim HU; Nave M; Dickschat JS; Mitchell DA; Shelest E; Breitling R; Takano E; Lee SY; Weber T; Medema MH Nucleic Acids Res 2017, 45, W36–W41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (67).van Heel AJ; de Jong A; Song C; Viel JH; Kok J; Kuipers OP Nucleic Acids Res 2018, 46, W278–W281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (68).Altschul SF; Gish W; Miller W; Myers EW; Lipman DJ J. Mol. Biol 1990, 215, 403–410. [DOI] [PubMed] [Google Scholar]
- (69).Nguyen LT; Schmidt HA; von Haeseler A; Minh BQ Mol. Biol. Evol 2015, 32, 268–274. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (70).Cobb RE; Wang YJ; Zhao HM ACS Synth. Biol 2015, 4, 723–728. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.