Abstract
Maternal inheritance of mitochondria creates a sex-specific selective sieve through which mitochondrial mutations harmful to males but not females accumulate and contribute to sexual differences in longevity and disease susceptibility. Because eggs and sperm are under disruptive selection, sperm are predicted to be particularly vulnerable to the genetic load generated by maternal inheritance, yet evidence for mitochondrial involvement in male fertility is limited and controversial. Here, we exploit the coexistence of two divergent mitochondrial haplogroups (A and B2) in a Neotropical arachnid to investigate the role of mitochondria in sperm competition. DNA profiling demonstrated that B2-carrying males sired more than three times as many offspring in sperm competition experiments than A males, and this B2 competitive advantage cannot be explained by female mitochondrial haplogroup or male nuclear genetic background. RNA-Seq of testicular tissues implicates differential expression of mitochondrial oxidative phosphorylation (OXPHOS) genes in the B2 competitive advantage, including a 22-fold upregulation of atp8 in B2 males. Previous comparative genomic analyses have revealed functionally significant amino acid substitutions in differentially expressed genes, indicating that the mitochondrial haplogroups differ not only in expression but also in DNA sequence and protein functioning. However, mitochondrial haplogroup had no effect on sperm number or sperm viability, and, when females were mated to a single male, neither male haplogroup, female haplogroup nor the interaction between male/female haplogroup significantly affected female reproductive success. Our findings therefore suggest that mitochondrial effects on male reproduction may often go undetected in noncompetitive contexts and may prove more important in nature than is currently appreciated.
Keywords: atp8, Cordylochernes scorpioides, Differential expression, Mitochondria, OXPHOS, RNA-Seq, Mother’s curse, Sperm competition
1 |. INTRODUCTION
In nearly all animals, mitochondria are inherited exclusively through females (Birky, 2001; Breton & Stewart, 2015). Because males do not transmit mitochondria to offspring, mutations in the mitochondrial genome that are detrimental to males but neutral or advantageous to females cannot respond directly to selection (Frank & Hurst, 1996; Zeh & Zeh, 2005; Hedrick, 2012). The transmission of such pernicious legacies from mothers to sons, known as “mother’s curse” (Gemmel, Metcalf, & Allendorf, 2004), creates a sex-specific selective sieve through which male-harmful mutations can persist in populations (Camus & Dowling, 2018; Connallon, Camus, Morrow, & Dowling, 2018). Mitochondrial mutations have been implicated in a range of pathologies encompassing aging, cancer and degenerative disease, and frequently involve tissues with high metabolic requirements, such as heart, skeletal muscle and brain (Wallace, 2013). Compared to females, males typically exhibit reduced lifespan and greater susceptibility to disease (Wolff & Gemmell, 2013; Ballard & Pichaud, 2014; Austad & Fischer, 2016; Wolff, Pichaud, Camus, Cote, Blier, & Dowling, 2016). In at least some cases, research on humans and model organisms has established mitochondrial mutations as the underlying cause (Camus, Clancy, & Dowling, 2012; Ross, Coppotelli, Hoffer, & Olson., 2014; Milot, Moreau, Gagnon, Cohen, Brais, & Labuda, 2017) of this “fragile male phenomenon” (Zeh & Zeh, 2005). However, the contribution of mitochondria to the sex-specific genetic load in humans and natural populations of other animals remains inadequately investigated and challenging to predict because viscous population structure (Wade & Brandvain, 2009), as well as selection for compensatory nuclear gene expression, can ameliorate the effects of mother’s curse (Rand, Haney, & Fry 2004; Dowling, Friberg, & Lindell, 2008).
At its most fundamental level, sex is defined by the relative size of the gametes. Females produce few, large eggs provisioned with thousands of quiescent mitochondria, whereas males produce many motile sperm fueled by few but metabolically active mitochondria (Zeh & Zeh, 2005; Vaught & Dowling, 2018). Sex-specific selection is therefore likely to be most extreme on mitochondrial function in gametes and gonads, and sperm and testes are predicted to be particularly vulnerable to mother’s curse (Frank & Hurst, 1996; Zeh & Zeh, 2005; Vaught & Dowling, 2018). While early efforts to link disrupted male fertility to mitochondrial variation yielded both supporting evidence (Ruiz-Pesini et al., 2000; Froman & Kirby, 2005; Smith, Turbill, & Suchentrunk, 2010) and contradictory findings (Mossman, Slate, & Birkhead, 2010), it is now clear that mitochondrial mutations can compromise male fertility in a sex-specific manner either directly through altered mitochondrial gene expression (Camus, Wolf, Morrow, & Dowling, 2015; Patel et al., 2016) or indirectly through mitochondrial effects on the expression of a wide range of nuclear genes in testicular tissue (Innocenti, Morrow, & Dowling, 2011). Perhaps because most mutations in both nuclear and mitochondrial genomes are known to be deleterious (Moon & Akey, 2016), empirical studies of mother’s curse have focused on identifying mitochondrial mutations with male-specific effects that are harmful. However, maternal inheritance of mitochondria can also prevent the spread of male-specific beneficial mitochondrial mutations in populations, thereby inhibiting the capacity for the adaptive evolution of sperm (Zeh & Zeh, 2005), a process described as “sister’s curse” (Padua, Zeh, Bonilla, & Zeh, 2014). Support for the sister’s curse hypothesis can be found in the doubly uniparental mitochondrial inheritance systems of some bivalve lineages, in which paternally inherited mitochondria in sperm and male gonadal tissue exhibit accelerated evolution and molecular signatures of positive selection (Zeh & Zeh, 2005; Smietanka, Burzynski, & Wenne, 2010), and in the co-occurrence of divergent mitochondrial haplogroups in a Neotropical arthropod (Padua, Zeh, Bonilla, & Zeh, 2014).
Unlike temperate zone species, tropical species often exhibit high levels of mitochondrial DNA sequence variation across small spatial scales even in the presence of gene flow (Wilcox, Hugg, Zeh, & Zeh, 1997; Strutzenberger, Brehm, & Fiedler, 2011; Stephen, Reynoso, Collett, Hasbun, & Breinholt, 2013). The introgression of divergent mitochondrial haplotypes provides an opportunity to investigate the micro- and macroevolutionary consequences of sexually asymmetric selection acting on mitochondria in natural populations. In assessing the impact of mitochondrial variation on male fertility, it is important to recognize that females in most animal species are polyandrous and produce mixed paternity broods of offspring (Zeh & Zeh, 1996, 1997; Jennions & Petri, 2000). Sperm competitive ability is therefore likely to be a key component of male reproductive success (Parker, 1970). Here, we exploit the coexistence of two divergent mitochondrial haplogroups (A and B2) in the Neotropical pseudoscorpion, Cordylochernes scorpioides, to investigate the role of mitochondrial gene expression in male fertility in a competitive context. Molecular paternity assignment demonstrated that B2-carrying males sired significantly more offspring in sperm competition experiments than A males, and RNA sequencing of testicular tissues implicates differential expression of mitochondrial oxidative phosphorylation (OXPHOS) genes in the B2 competitive advantage, including a more than 20-fold upregulation of atp8 in B2-haplogroup males.
1.1 |. Reproductive biology of the Neotropical pseudoscorpion, Cordylochernes scorpioides
The co-occurrence of two highly divergent mitochondrial haplogroups in central Panamanian populations of the harlequin beetle riding pseudoscorpion, Cordylochernes scorpioides, provides an ideal context for assessing the influence of mitochondrial gene expression on male fertility in both competitive and non-competitive contexts (Zeh et al., 2012; Padua, Zeh, Bonilla, & Zeh, 2014). Cordylochernes scorpioides exhibits a XX-XO male heterogametic sex determination system (Št’áhlavský, Zeh, Zeh & Král, 2009) and, in these pseudoscorpions, mating involves a sequence of stereotypical behaviors in which the male grasps the female while he constructs and deposits a spermatophore on the substrate. The male then maneuvers the female into a position in which the sperm packet directly contacts her genital aperture, and successful sperm transfer is associated with a pronounced abdominal flexure by the female (Zeh, Newcomer, & Zeh 1998). Matings can be interrupted immediately following spermatophore deposition and the sperm packet collected for assessment of sperm number and viability (Zeh, Newcomer, & Zeh 1998). External spermatophore deposition and diagnostic female behavior facilitate unambiguous assessment of female sexual receptivity and success of sperm transfer. Non-invasive monitoring of female reproductive status and embryological development is made possible by the ‘external womb’ mode of viviparity in C. scorpioides, in which females nourish developing embryos in an external, transparent brood sac overlying their genital aperture (Newcomer, Zeh, & Zeh, 1999). In nature, C. scorpioides females produce mixed-paternity broods sired by up to four males (Zeh, Zeh, & Bermingham, 1997), and sperm competitive ability is therefore an important component of male fitness in this pseudoscorpion.
2 |. MATERIALS AND METHODS
2.1 |. Experimental pseudoscorpions
Experimental pseudoscorpions were drawn from a large laboratory population established from 350 C. scorpioides adults and nymphs collected in 2006 and 2008 from six locations spanning a 60 km region in central Panamá (Zeh et al., 2012). In this laboratory population, pseudoscorpions were reared and maintained in individual vials to ensure virginity, and matings were staged to maintain a large number of field-collected matrilines. Each generation, no matings were carried out between full siblings, half siblings or first cousins in order to minimize inbreeding. Within these constraints, pairs for mating were chosen randomly without regard to mitochondrial haplogroup. Because random mating between haplogroups was performed for a minimum of 16 generations, individuals from the two haplogroups were effectively homogenized for nuclear genetic background. To avoid possible confounding effects of the maternally-inherited cellular endosymbiont Wolbachia on mitochondrial gene expression, Wolbachia-specific MLST PCR assays were carried out to confirm that all C. scorpioides matrilines used in the study were uninfected with Wolbachia (for methodology, see Koop, Zeh, Bonilla, & Zeh, 2009).
2.2 |. DNA profiling to assess haplogroup effects on sperm competitive ability
Mitochondrial haplogroup effects on sperm competitive ability were assessed in two-male sperm competition experiments in which initially virgin females were each mated to two males, one carrying A-haplogroup mitochondria and the other B2-haplogroup mitochondria. To complement a previous study in which competing A and B2 haplogroup males were mated exclusively to A-haplogroup females (Padua, Zeh, Bonilla, & Zeh, 2014), only B2-haplogroup females were used in this study. Results from the current study were analyzed separately, and data from the two studies were also pooled to assess possible female haplogroup and male by female haplogroup effects on sperm competitive ability (see below).
Each replication was initiated by placing a virgin female with a virgin male in a 28 mm diameter mating arena under an Olympus SZ6145TR stereomicroscope. Interactions were observed for 45 min or until the female accepted a sperm packet from the male. Mating order was randomized across male haplogroups with a 48 hr time lapse before the procedure was repeated with the second male. Only replicates in which the female unambiguously accepted a sperm packet (see Zeh, Newcomer, & Zeh 1998) from both males were retained for paternity analysis. Following their male B mating, females were maintained in individual vials in a dark incubator at 28.5°C and 80% relative humidity and monitored until giving birth. First-stage nymphs (protonymphs) were then removed from the brood nest, counted and frozen at −80°C, together with the dam and the two putative sires.
DNA was extracted from the dams, putative sires and protonymphs using Invitrogen Chargeswitch gDNA Micro Tissue Kit (ThermoFisher Scientific, Waltham, MA, USA). PCR amplification of alleles at the cCscMS23 minisatellite locus (heterozygosity = 0.99; Zeh, Zeh, & May, 1994; Zeh and Zeh, 2006) was used to assign paternity. For each replication, PCR products from the mother, the two putative sires and an average of 10 offspring were run on a 1.5% agarose gel and stained with ethidium bromide to visualize alleles. Paternity was assigned based on the presence of unique paternal alleles in offspring. The PCR reactions were performed in a 30 μL reaction volume containing ~10 ng of genomic DNA, 3 μL of 10X Advantage Genomic LA Buffer (25 mM MgCl2), 350 mM dNTPs, 750 nM primers, and 1.5U of Advantage Genomic LA Polymerase Mix (Takara Bio USA, Madison, WI, USA). The two-step PCR amplification reaction included a 1-min hot start at 95°C, followed by 34 cycles of 93°C melting for 25 s, and 68°C annealing/extension for 8 min.
Across the two experiments, it was possible to unambiguously assign paternity for 91% of the offspring whose alleles were amplified by PCR (496 of 545 nymphs). In 23 of 29 replications, all paternal alleles were unique and paternity could be assigned for all offspring. In the remaining six replications, the two putative sires shared one of their two alleles and paternity could not be unambiguously assigned for approximately 50% of the offspring. These offspring were excluded from statistical analyses.
DNA sequencing of the mitochondrial nd2 locus from our C. scorpioides laboratory matrilines has established that haplotypes in the A but not the B2 haplogroup possess a ClaI restriction site (Padua, Zeh, Bonilla, & Zeh, 2014). ClaI digested nd2 amplicons were therefore used to confirm the mitochondrial haplotype of all putative sires from successful replicates of the sperm competition experiment. DNA from frozen adults was extracted as described above and PCR was conducted using the nd2 mitochondrial DNA locus forward (5’ – TGTAAGTCTTAAAAYAAAGAAAACC – 3’) and reverse primers (5’ – AAGTCATCGAATAGARACRTTAGC – 3’). PCR reactions were performed, as described above, except that the conditions of the 34 cycles were modified to 93°C for 25 s, 48°C for 50 s, and 68°C for 90 s. Digestion reactions were carried out using 20 μL mixtures of 1X buffer, 1 μg (approximately 12 μL) of PCR product and 1μL of ClaI restriction enzyme. Products were incubated for 60 min at 37°C and the enzyme inactivated at 65°C for 20 min. Products were visualized on 1.5% agarose gels run for approximately 80 volt-hr to determine presence (two fragments) or absence (one fragment) of the restriction site in the nd2 sequence from each male.
2.3 |. RNA-Seq of testicular tissues to assess haplogroup effects on mitochondrial gene expression
Five full-sibling families for each of the two mitochondrial haplogroups were randomly selected for testicular tissue dissection and RNA extraction. To obtain sufficient RNA, dissected testes from four young adult males were pooled for each of the 10 families. Males were frozen in liquid nitrogen, dissected under 20–40X magnification, and the testes surgically removed (see Weygoldt, 1969). Total RNA was purified into small (< 200 nucleotides) and large (> 200 nucleotides) fractions using a PureLink® miRNA Isolation Kit in combination with a PureLink® RNA Mini Kit (ThermoFisher Scientific, Waltham, MA, USA). Purified RNA samples were stored at −80°C for further downstream processing prior to sequencing. RNA quantity and quality were assessed by the Nevada Genomics Center (Reno, NV, USA), using RiboGreen quantitation and Agilent 6000 Nano RNA analysis (Agilent Technologies, Santa Clara, CA, USA), respectively. RNA Integrity Numbers (RI) ranged from 7.90 to 9.60, and RNA quantity from 0.65 μL to 1.52 μL.
Library preparation and next generation RNA sequencing of the 10 large-fraction RNA samples were performed by the Nevada Genomics Center. Prior to sequencing, RiboMinus™ Eukaryote System v2 treatment (ThermoFisher Scientific, Waltham, MA, USA) was used to selectively deplete ribosomal RNAs. Ten barcode-labeled, TruSeq® sequencing libraries were sequenced in multiplex using an Illumina NextSeq 500 and an Illumina Mid-Output flow cell (Illumina, San Diego, CA, USA). Sequencing was paired-end, with the sequencer configured to collect forward and reverse sequences for 76 cycles each, and six cycles for barcode indices. Pre-base-call intensity data, logs and metrics were sent to the Illumina BaseSpace service, and processed via the built-in BaseSpace FASTQ Generation module, which called nucleotide bases, demultiplexed the samples by barcode label, masked adapter sequences, and generated FASTQ-formated sequence files which were downloaded for further processing. Paired-end sequences were further trimmed and filtered to remove sequencing adapters, artifacts, and low-quality bases using Trimmomatic software, version 0.36 (Bolger, Lohse, & Usadel, 2014).
Sequences were aligned to published mitochondrial genomes for the A and B2 haplogroups (Padua, Zeh, Bonilla, & Zeh, 2014). The full mitochondrial genome sequence of each haplogroup was indexed for alignment using `hisat2-build` command within the HISAT2 alignment software package, version 2.1.0 (Kim, Langmead, & Salzberg, 2015). Trimmed read pairs were mapped to their indexed mitochondrial genome with `hisat2` reporting of up to five alignments per read pair, if multiple valid alignments were detected. The resultant sequence alignment maps were then compressed into sorted binary (BAM) files using samtools, version 1.3.1 (Li et al., 2009). The numbers of read pairs aligned to each known mitochondrial gene were calculated using the featureCounts tool within the subread package, version 1.5.1 (Liao, Smyth, & Shi, 2014). Alignments were counted once per pair, summarized as gene loci features, and only read pairs with alignments to one location were counted. For alignments to regions where annotated gene boundaries intersected, counts were assigned to the gene with the largest overlap to the read pair. Feature counts of genes were converted to counts per million (CPM) and further normalized based on gene length.
2.4 |. qPCR to assess whether haplogroups differ in mitochondrial copy number in testicular tissue
For this assay, we isolated testicular tissue and extracted DNA from 34 adult C. scorpioides males, as described above. Comparative quantitative PCR (qPCR) was performed, using an Applied Biosystems 7500 Fast Real-Time PCR System and Power SYBR® Green PCR Master Mix (ThermoFisher Scientific, Waltham, MA, USA). To estimate the number of mitochondria per cell, we amplified a conserved, 142-bp region of the atp6 gene, using the primers 5’-ACTCTTTCAATTCGATTAACAGC-3’ and 5’-TCGCCACTGCAAGTTCGAGAG-3’. As an internal control, we amplified a 113-bp region of a single copy, X-chromosome-linked microsatellite locus, using the primers 5’-GGGAAGACAACGTCGAACAAC 3’ and 5’-CTCCGTCCTTCTCCTTGTCC-3’. Using 10 ng of total DNA as template, qPCR reactions were conducted by the Nevada Genomics Center according to manufacturers’ specifications. For both loci, average critical threshold (CT) and standard deviation values were calculated from three replicates per sample. For each male, comparison of the mean CT values for the two markers was used to estimate the copy number of the mitochondrial gene relative to the nuclear gene.
2.5 |. Statistical analyses
With the exception of the differential gene expression analyses, statistical analyses were performed using SAS/STAT Version 15.1 (SAS, Institute Inc., 2018). For the sperm competition experiment involving only B2 females, the SAS GLIMMIX procedure incorporating a random effect (female identity), a log link function, a Gauss – Hermite Quadrature maximum-likelihood approximation, and a generalized Poisson mixed model for overdispersed count data (SAS Institute Inc., 2018: 3919–3926) was used to assess the effects of male haplogroup and mating order on the number of protonymphs sired by the A and B2 males in each replication. Female identity was included as a random factor in the model to avoid pseudoreplication. To control for slight differences in the numbers of protonymphs assayed for paternity across replications, the total number of protonymphs for which paternity was assigned in each replication was included as a weighting factor in the model. For the pooled data set that incorporated results for both A haplogroup (Padua, Zeh, Bonilla, & Zeh, 2014) and B2 haplogroup females, the SAS GLIMMIX generalized Poisson procedure was again used, in this case to assess the effects of male haplogroup, female haplogroup and mating order on the number of protonymphs sired by the A and B2 males in each replication. To control for differing numbers of protonymphs assayed for paternity in the two experiments, the total number of protonymphs for which paternity was assigned in each replication was included as a weighting factor in the model.
The Bioconductor software package edgeR (Robinson, McCarthy & Smyth, 2010) was used to assess whether the A and B2 haplogroups differentially expressed any of the 13 OXPHOS and two ribosomal RNA genes of the mitochondrial genome. edgeR employs an overdispersed Poisson model to account for biological and technical variability, and empirical Bayes methods to moderate the degree of overdispersion across transcripts (Robinson, McCarthy, & Smyth, 2010). The Bonferroni-Holm method (Holm, 1979) was used to adjust P values for multiple comparisons.
The SAS GLIMMIX procedure incorporating a random effect (male full-sibling family), a Poisson distribution and a log link function was used to assess possible differences in mitochondrial copy number in the testicular tissues of the A and B2 haplotypes. Male full-sibling family was included as a random effect because one to two males were assayed from each of 25 full-sibling families.
3 |. RESULTS
3.1 |. Molecular paternity assignment to assess haplogroup effects on sperm competitive ability
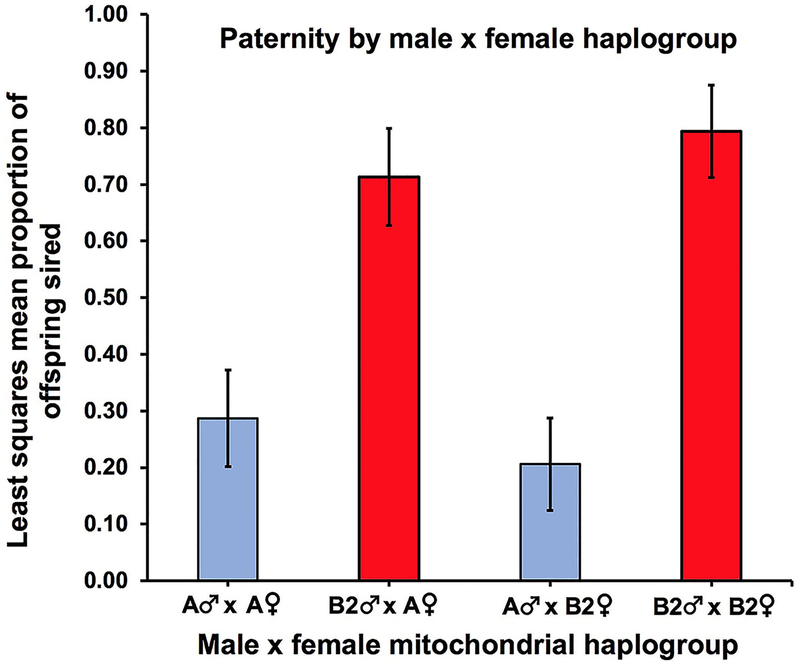
In the sperm competition experiment reported here that incorporated only B2 haplogroup females, there were significant effects of male haplogroup (F1,10 = 15.67, P = 0.0027) and mating order (F1,10 = 6.70, P = 0.0270) on the number of protonymphs sired in each replication. The interaction between male haplogroup and mating order was not significant (F1,10 = 0.58, P = 0.4650). B2 males sired a significantly higher proportion of protonymphs than A males. The least squares mean proportion of offspring sired (± S.E.M) for the B2 haplogroup [LSMB2] = 0.79 ± 0.08 and for A haplogroup [LSMA] = 0.21 ± 0.08).
Pooling the B2 female data with those of our previously published study that used only A haplogroup females (Padua, Zeh, Bonilla, & Zeh, 2014) again yielded a highly significant effect of male haplogroup (F1,26 = 15.96, P = 0.0005). Mating order was also significant (F1,26 = 4.55, P = 0.0426). By contrast, the effects of female haplogroup (F1,26 = 2.54, P = 0.1232), and the interaction between female haplogroup and male haplogroup (F1,26 = 0.51, P = 0.4798) were not significant. For this pooled data set, B2 males again sired a significantly higher proportion of offspring than A males (LSMB2 = 0.74 ± 0.07, LSMA = 0.26 ± 0.07). The LSM proportion of offspring sired by A and B2 males was very similar across the two female haplogroups (Fig. 1).
Fig. 1.
B2 sperm competitive advantage in Cordylochernes scorpioides from central Panamá. Bars represent the least squares mean (LSM) proportion of offspring sired by A versus B2 haplogroup males in two-male sperm competition experiments with either A or B2 females. LSMs for A haplogroup males are indicated in blue and LSMs for B2 haplogroup males in red. Error bars represent standard error of the means.
It should be noted that the 13-month lapse between completion of the A-female sperm competition experiment and initiation of the B2-female sperm competition experiment could have confounded the pooled data analysis due to population and experimental condition effects. However, given that pseudoscorpions in both experiments were derived from the same population that had been maintained in the laboratory for at least 14 generations, population confounding effects seem unlikely. Moreover, such confounding would likely promote male x female haplogroup interaction effects and contribute to between-experiment differences in male haplogroup effects. Contrary to these expectations, the pooled analysis indicated strongly congruent male haplogroup effects across the two experiments and no evidence for male x female haplogroup effects.
3.2 |. RNA-seq of testicular tissues to assess haplogroup effects on mitochondrial gene expression
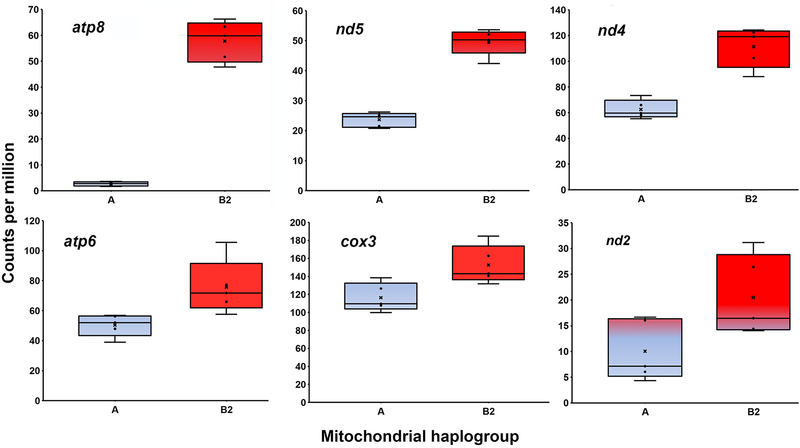
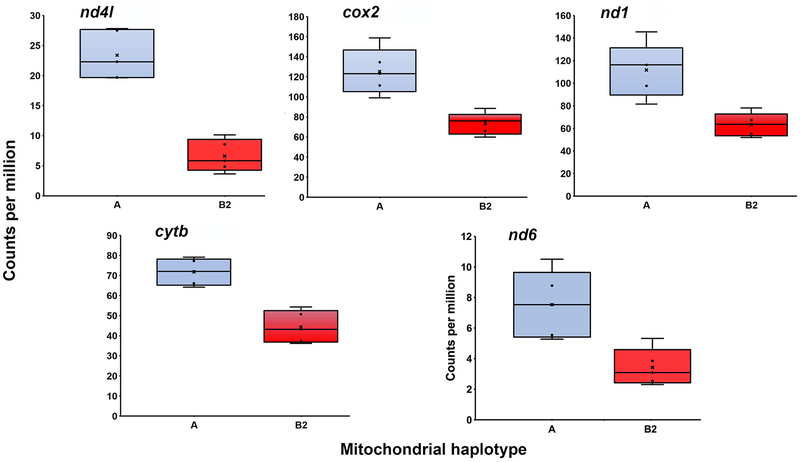
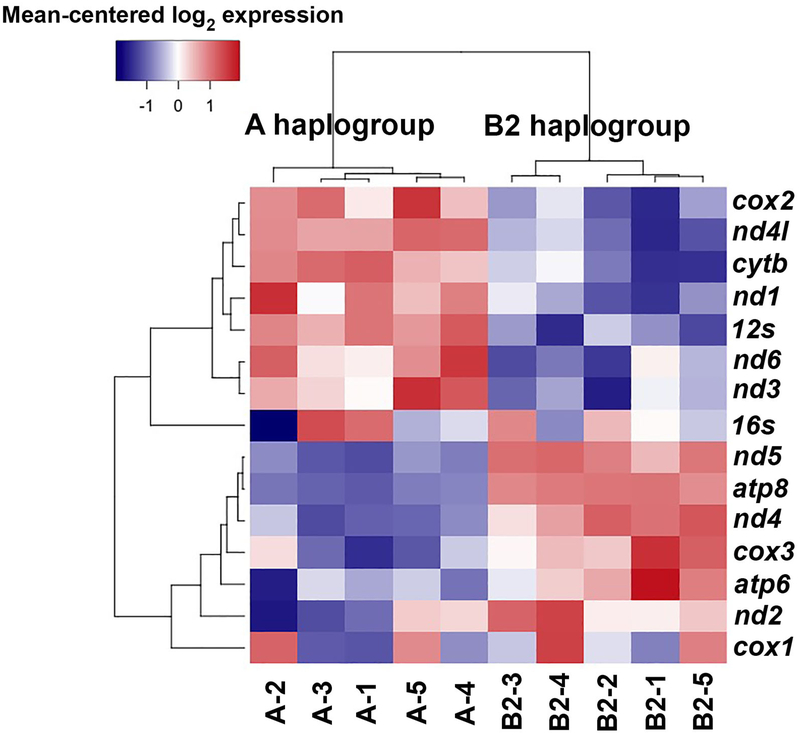
Bonferroni-Holm adjusted p values for haplogroup differential expression of the 13 mitochondrial protein-coding genes and the 2 rRNA genes (Table 1) ranged from 1.41×10−67 (atp8) to 0.9492 (16S). The OXPHOS genes atp8, nd5, nd4, atp6 and nd2 and cox3 were all significantly upregulated in the B2 haplogroup relative to the A haplogroup (Fig. 2), with upregulation values ranging from 1.31 (cox3) to 21.62 (atp8). Genes significantly upregulated in A haplogroup males included 12S, nd4l, cox2, nd1, nd6 and cytb. For these A haplogroup upregulated genes, differential expression was generally less extreme than for B2-upregulated genes (Fig. 3), with upregulation values ranging from 1.62 (cytb) to 3.54 (nd4l). A heatmap of mitochondrial gene expression clearly separates the A and B2 haplogroups into distinct clades (Fig. 4).
Table 1.
Ranking of mitochondrial gene differential expression for the A and B2 haplogroups by level of statistical significance. To account for multiple comparisons, P-values were adjusted using the Bonferroni-Holm (1979) method and are reported as false discovery rates (FDR). The haplogroup means are shown in counts per million.
| Gene | Haplogroup A Mean | Haplogroup B2 Mean | log2 ratio (B2:A) | Ratio of B2:A expression | Significance Rank | FDR |
|---|---|---|---|---|---|---|
| atp8 | 2.67 | 57.76 | 4.39 | 21.62 | 1 | < 0.0001 |
| 12s | 95.97 | 31.75 | −1.57 | 0.33 | 2 | < 0.0001 |
| nd5 | 23.67 | 49.55 | 1.08 | 2.09 | 3 | < 0.0001 |
| nd41 | 23.39 | 6.61 | −1.78 | 0.28 | 4 | < 0.0001 |
| nd4 | 62.50 | 111.33 | 0.85 | 1.78 | 5 | < 0.0001 |
| cox2 | 125.36 | 73.39 | −0.75 | 0.59 | 6 | < 0.0001 |
| nd1 | 111.68 | 63.28 | −0.80 | 0.57 | 7 | < 0.0001 |
| cytb | 71.80 | 44.40 | −0.67 | 0.62 | 8 | < 0.0001 |
| atp6 | 50.38 | 75.67 | 0.60 | 1.50 | 9 | 0.0006 |
| nd2 | 10.04 | 20.50 | 1.04 | 2.04 | 10 | 0.0033 |
| cox3 | 116.40 | 152.63 | 0.41 | 1.31 | 11 | 0.0081 |
| nd6 | 7.53 | 3.43 | −1.09 | 0.46 | 12 | 0.0155 |
| nd3 | 3.95 | 2.09 | −0.87 | 0.53 | 13 | 0.1141 |
| cox1 | 134.02 | 137.55 | 0.06 | 1.03 | 14 | 0.7531 |
| 16s | 172.12 | 168.34 | −0.01 | 0.98 | 15 | 0.9492 |
Fig. 2.
Mitochondrial genes significantly overexpressed in the testicular tissues of B2 haplogroup males. The box and whisker plots compare the levels of gene expression for the mitochondrial genes that were significantly overexpressed by the C. scorpioides B2 haplogroup (red) relative to the A haplogroup (blue).
Fig. 3.
Mitochondrial genes significantly overexpressed by A haplogroup males. The box and whisker plots compare levels of gene expression for the mitochondrial genes that were significantly overexpressed by the C. scorpioides A haplogroup (blue) relative to the B2 haplogroup (red).
Fig. 4.
A heatmap showing mean-centered expression levels of the 15 mitochondrial genes. The A and B2 haplogroup samples cluster into two distinct clades as indicated in the dendrogram.
3. 3 |. qPCR to assess whether haplogroups differ in mitochondrial copy number in testicular tissue
qPCR-based estimates of the number of mitochondrial gene copies per cell in the testicular tissues of A (LSMA = 5.63 ± 0.51) and B2 (LSMB2 = 6.46 ± 1.05) males did not differ significantly (P = 0.4714).
4 |. DISCUSSION
In the study reported here in which B2-haplogroup females were each mated to both an A and a B2 male, DNA profiling demonstrated that B2-haplogroup males sired more than three times as many offspring than A males. In conjunction with the results of an earlier sperm competition experiment involving only A haplogroup females (Padua, Zeh, Bonilla, & Zeh, 2014), our research on C. scorpioides indicates that the B2 sperm competitive advantage is independent of female haplogroup. This intrinsic B2 advantage cannot be attributed to differences in nuclear genetic background. The C. scorpioides individuals used in the two studies were maintained in the laboratory for a minimum of 14 (Padua, Zeh, Bonilla, & Zeh, 2014) or 16 generations (this study) in which they were mated randomly with respect to mitochondrial haplogroup. Consequently, the two haplogroups became effectively homogeneous with respect to nuclear genetic background, greatly increasing the likelihood that any systematic differences between haplogroups in sperm competitive ability and mitochondrial gene expression are causally related to differences in the mitochondrial genome.
Our RNA sequencing of testicular tissues implicates differential expression of 11 mitochondrial oxidative phosphorylation (OXPHOS) genes in the B2 competitive advantage, including a more than 20-fold upregulation of atp8 in B2-haplogroup males. Quantitative PCR demonstrated that differential expression between haplogroups cannot be attributed to differences in mitochondrial copy number. However, mitochondrial haplotype is known to affect nuclear gene expression in male testicular tissue (Innocenti, Morrow, & Dowling, 2011), and it should therefore be acknowledged that a limitation of the current study is the lack of a reference nuclear genome for C. scorpioides. Unfortunately, C. scorpioides appears to possess a large (~2 Gb) and complex genome, and generating a reference genome will require substantial sequencing, assembly and annotation efforts. Phylogenetic analysis of the cox1 gene (Zeh, Zeh & Bonilla. 2003) indicates that the Panamanian A and B clades of C. scorpioides are sister groups that were derived from South American populations of the C. scorpioides species complex (Wilcox, Hugg, Zeh & Zeh, 1997). In the absence of mtDNA gene expression data from South American populations, it is therefore not currently possible to determine the evolutionary polarity of OXPHOS gene expression levels.
Previous comparative genomic analyses (Padua, Zeh, Bonilla, & Zeh, 2014) have revealed functionally significant amino acid substitutions in four of the differentially expressed genes (atp8, nd4, nd4l and nd3), indicating that the mitochondrial haplogroups differ not only in expression but also in DNA sequence and protein functioning. Previous research also demonstrated that the B2 fertility advantage only becomes apparent in a competitive context (Padua, Zeh, Bonilla, & Zeh, 2014). Males carrying the two mitochondrial haplogroups did not differ in sperm number or sperm viability, and, when females were mated to a single male, neither male haplogroup, female haplogroup nor the interaction between male/female haplogroup significantly affected female reproductive success (Padua, Zeh, Bonilla, & Zeh, 2014). To our knowledge, this is the first study to demonstrate intrinsic differences between naturally cooccurring mitochondrial haplogroups in sperm competitive ability linked to the differential expression of mitochondrial genes. The strong but cryptic effects of mtDNA sequence variation and OXPHOS gene expression on male fertility suggest that mitochondrial effects on male reproductive success may frequently go undetected in nature.
While published studies of the evolutionary implications of maternal inheritance of mitochondria have focused on the accumulation of deleterious mutations contributing to male infertility, our research on C. scorpioides is novel in providing evidence that the inability of males to transmit mitochondria can constrain the spread of an advantageous mitochondrial variant in nature. In the wild, C. scorpioides females are polyandrous, and produce mixed-paternity broods sired by up to four males (Zeh, Zeh, & Bermingham, 1997). Sperm competitive ability is therefore likely to be an important component of male reproductive success in this pseudoscorpion. Despite the strong B2 advantage in sperm competition, the B2 haplotype occurs at low frequency (12%) in central Panamanian populations of C. scorpioides (Zeh et al., 2012). The rarity of B2 appears to be the consequence of sister’s curse, that is, the haplogroup’s antagonistic effects on female fitness (Padua, Zeh, Bonilla, & Zeh, 2014). While essentially all C. scorpioides virgin females are sexually receptive, sexual receptivity at second mating has been shown to be significantly lower in females carrying B2 haplogroup mitochondria (Padua, Zeh, Bonilla, & Zeh, 2014). Given the evidence for substantial fitness benefits to polyandry in this pseudoscorpion (Newcomer, Zeh, & Zeh, 1999), the reduced propensity for polyandry by B2 haplogroup females is likely to incur significant costs to female lifetime reproductive success. Ongoing research in our laboratory also suggests that B2 mitochondria exert negative impacts on life history and morphological traits in females, including slower developmental rate and reduced adult body size.
Given the large number of protein-coding mitochondrial genes that were significantly differentially expressed in the two haplogroups (atp6, atp8, nd2, nd4, nd5 and cox3 were upregulated in Haplogroup B2, and nd1, nd4l, cox2 and cytb were upregulated in Haplogroup A), multiple mechanisms are likely to be responsible for the B2 sperm competitive advantage. Established and proposed roles of mitochondrial gene expression in male fertility include meiosis, spermatogenesis, sperm maturation, apoptosis-based quality control, energy for sperm motility, energy for sperm survival and sperm capacitation (Rajender, Rahul, & Mahdi, 2010). Testes are composed of numerous germ and somatic cell types and recent single-cell RNA-Seq studies have demonstrated that mitochondrial gene expression varies between testicular cell types and stages of spermatogenesis. Compared to other testicular cells, the relative abundance of mtDNA transcripts is reduced in differentiating sperm cells, including meiotic spermatocytes, post-meiotic haploid round spermatids, and elongating spermatids, a pattern that likely stems from the reduction in mitochondrial copy number that occurs as sperm mature (Green et al., 2018).
The importance of mitochondrial metabolism in mature sperm remains controversial (Ramalho-Santos, Varum, Amaral, Mota, Sousa, & Amaral, 2009). Sperm clearly require ATP for motility, as well as for the cellular events involved in hyperactivation, capacitation and the acrosome reaction. However, whether the ATP that fuels these activities, especially motility, is provided by OXPHOS or by glycolysis has been debated for decades (Ramalho-Santos, Varum, Amaral, Mota, Sousa, & Amaral, 2009), at least in part because the relative importance of the OXPHOS pathway appears to be species specific (Guo, Gong, He, & Zhao, 2017). Despite uncertainty regarding the role of OXPHOS in ejaculated sperm, multiple lines of evidence indicate that differences in sperm motility result from differences in OXPHOS capacity (Ruiz-Pesini, et al. 2000) and mitochondrial transcript levels in sperm (Jodar, Kalko, Castillo, Ballesca, & Oliva, 2012). In addition, in mice, mitochondrial rRNAs and the mRNAs encoded by several mitochondrial genes have been isolated from sperm midpiece tails (Alcivar, Hake, Millette, Trasler, & Hecht, 1989).
While OXPHOS-based differences in sperm motility remain a plausible explanation for the C. scorpioides B2 sperm competitive advantage, differential A and B2 sperm survival in the female reproductive tract is unlikely to be a contributing factor. As we have shown elsewhere, over their 6 to 12-month lifespan, C. scorpioides females can produce multiple broods from stored A-haplotype sperm derived from a single spermatophore (Newcomer, Zeh, & Zeh, 1999). In fact, in long-lived females, the number of stored sperm (~ 2,000) is only slightly greater than the total number of eggs fertilized (~500). In addition, the time between matings in our sperm competition experiment was 48 hours, and females typically produced broods of embryos within 5 days of mating. It therefore seems unlikely that differential survival of sperm could account for the B2 fertilization advantage.
It is important to acknowledge that our study has not definitively established that haplogroup differences in sperm competitive ability result from differences in levels of OXPHOS gene expression. Haplogroup differences could be due to functionally significant DNA sequence differences in just one or multiple OXPHOS genes (Padua, Zeh, Bonilla, & Zeh, 2014). Nonetheless, the extreme, 22-fold overexpression of atp8 in the testes of B2 males is consistent with this gene playing an important role in the B2 advantage. Embedded in the mitochondrion’s inner membrane, atp8, together with atp6 which was also significantly overexpressed in B2 males, comprises Complex V of the electron transport chain (ETC) (Wallace, 2005). The ETC is responsible for establishing the voltage differential that drives eukaryotic metabolism and generates as much as 90% of cellular energy (Wallace 2013). atp8 and atp6 are directly involved in the final stage processing of ATP from ADP, and are therefore likely to act as rate-limiting enzymes in the production of the cellular energy required for spermatogenesis.
Interestingly, accelerated evolution of the atp8 gene has been detected in the paternally inherited “M” mitochondria of bivalve mollusks (Smietanka, Burzynski, & Wenne, 2010). In several bivalve lineages, including marine mussels, freshwater mussels and marine clams, species exhibit an intriguing pattern of doubly uniparental inheritance (Breton, Beaupre, Stewart, Hoeh, & Blier, 2007; Zouros, 2013) that can at least partially circumvent the asymmetrical selective filter acting on mitochondria (Zeh & Zeh, 2005). In these bivalves, females transmit “F” mitochondria to both daughters and sons, and males also inherit M mitochondria from their fathers. The distribution of these two mitochondrial types in sexually mature males is particularly interesting (Sutherland, Stewart, Kenchington, & Zouros, 1998; Cao, Kenchington, & Zouros, 2004). Female-transmitted mitochondria are predominant in somatic tissues, male-transmitted mitochondria are predominant in testes, and sperm contain only male-transmitted mitochondria. As predicted by sister’s curse, M and F mitochondria diverge rapidly, with evolution most accelerated in the paternally-inherited atp8 gene (Smietanka, Burzynski, & Wenne, 2010; Lubosny, Przylucka, Smietanka, Breton, & Burzynski, 2018). Indeed, paternally inherited atp8 genes have evolved so divergently that they were overlooked in most annotations of bivalve mitochondrial genomes carried out before 2009, and the presence of the gene has only been confirmed by recent comparative mitogenomic studies (Plazzi, Puccio, & Passamonti, 2016; Lubosny, Przylucka, Smietanka, Breton, & Burzynski, 2018). Taken in conjunction with patterns in bivalve atp8 evolution, our findings on the relationship between mitochondrial gene expression and sperm competitive ability in C. scorpioides, suggest that this gene may play a generally important role in male reproductive success.
4.1 |. Conclusions
In an experimental design that controlled for female haplogroup and male nuclear genetic background, our research on the harlequin beetle riding pseudoscorpion provides evidence of a link between differential OXPHOS gene expression and sperm competitive ability in two highly divergent but naturally co-occurring mitochondrial haplogroups. Of particular significance is the identification of the mitochondrial atp8 gene as a potentially key component of sperm competitive ability. In addition, the demonstration that a mitochondrial variant’s beneficial effect is apparent only in a sperm competitive context suggests that mitochondrial impacts on male fertility may be more generally important than is currently recognized.
4.2 |. Data archiving
Reference mitochondrial genome sequences for C. scorpioides A and B2 haplogroups, mitochondrial DNA expression levels, qPCR mtDNA concentration results and male reproductive success data from the two-male sperm competition experiment are available on Dryad (https://datadryad.org/resource/doi:10.5061/dryad.bh19kj8). The fastq files in support of the differential mitochondrial gene expression analyses have been deposited in the NCBI Short Read Archive under accession numbers SRR7162678 - SRR7162697 for the A haplogroup libraries and SRR7956962 - SRR7956966 for the B2 haplogroup libraries.
ACKNOWLEDGMENTS
We thank La Autoridad Nacional del Ambiente for permission to collect pseudoscorpions in Panamá and the Smithsonian Tropical Research Institute for logistical support.
Funding Information
National Science Foundation, Grant/Award Number: IOS-1656670 and National Institutes of Health Award Number GM103440
Footnotes
Publisher's Disclaimer: This article has been accepted for publication and undergone full peer review but has not been through the copyediting, typesetting, pagination and proofreading process, which may lead to differences between this version and the Version of Record. Please cite this article as doi: 10.1111/jeb.13536
CONFLICT OF INTEREST
All authors declare no conflict of interest.
REFERENCES
- Alcivar AA, Hake LE, Millette CF, Trasler JM, & Hecht NB (1989). Mitochondrial gene-expression in male germ-cells of the mouse. Developmental Biology, 135(2), 263–271. 10.1016/0012-1606(89)90178-4 [DOI] [PubMed] [Google Scholar]
- Austad SN, & Fischer KE (2016). Sex Differences in Lifespan. Cell Metabolism, 23(6), 1022–1033. 10.1016/j.cmet.2016.05.019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ballard JWO, & Pichaud N (2014). Mitochondrial DNA: more than an evolutionary bystander. Functional Ecology, 28(1), 218–231. 10.1111/1365-2435.12177 [DOI] [Google Scholar]
- Birky CW (2001). The inheritance of genes in mitochondria and chloroplasts: Laws, mechanisms, and models. Annual Review of Genetics, 35, 125–148. 10.1146/annurev.genet.35.102401.090231 [DOI] [PubMed] [Google Scholar]
- Bolger AM, Lohse M, & Usadel B (2014). Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics, 30(15), 2114–2120. 10.1093/bioinformatics/btu170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Breton S, Beaupre HD, Stewart DT, Hoeh WR, & Blier PU (2007). The unusual system of doubly uniparental inheritance of mtDNA: isn’t one enough? Trends in Genetics, 23(9), 465–474. 10.1016/j.tig.2007.05.011 [DOI] [PubMed] [Google Scholar]
- Breton S, & Stewart DT (2015). Atypical mitochondrial inheritance patterns in eukaryotes. Genome, 58(10), 423–431. 10.1139/gen-2015-0090 [DOI] [PubMed] [Google Scholar]
- Camus MF, Clancy DJ, & Dowling DK (2012). Mitochondria, maternal inheritance, and male aging. Current Biology, 22, 1717–1721. 10.1016/j.cub.2012.07.018 [DOI] [PubMed] [Google Scholar]
- Camus MF, & Dowling DK (2018). Mitochondrial genetic effects on reproductive success: signatures of positive intrasexual, but negative intersexual pleiotropy. Proceedings of the Royal Society B-Biological Sciences, 285(1879). 10.1098/rspb.2018.0187 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camus MF, Wolf JBW, Morrow EH, & Dowling DK (2015). Single nucleotides in the mtDNA sequence modify mitochondrial molecular function and are associated with sex-specific effects on fertility and aging. Current Biology, 25(20), 2717–2722. 10.1016/j.cub.2015.09.012 [DOI] [PubMed] [Google Scholar]
- Cao LQ, Kenchington E, & Zouros E (2004). Differential segregation patterns of sperm mitochondria in embryos of the blue mussel (Mytilus edulis). Genetics, 166(2), 883–894. 10.1534/genetics.166.2.883 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Connallon T, Camus MF, Morrow EH, & Dowling DK (2018). Coadaptation of mitochondrial and nuclear genes, and the cost of mother’s curse. Proceedings of the Royal Society B-Biological Sciences, 285(1871). 10.1098/rspb.2017.2257 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dowling DK, Friberg U, & Lindell J (2008). Evolutionary implications of non-neutral mitochondrial genetic variation. Trends in Ecology & Evolution, 23, 546–554. 10.1111/jeb.12822 [DOI] [PubMed] [Google Scholar]
- Frank SA, & Hurst LD (1996). Mitochondria and male disease. Nature, 383(6597), 224–224. 10.1038/383224a0 [DOI] [PubMed] [Google Scholar]
- Froman DP, & Kirby JD (2005). Sperm mobility: Phenotype in roosters (Gallus domesticus) determined by mitochondrial function. Biology of Reproduction, 72(3), 562–567. 10.1095/biolreprod.104.035113 [DOI] [PubMed] [Google Scholar]
- Gemmell NJ, Metcalf VJ, & Allendorf FW (2004). Mother’s curse: the effect of mtDNA on individual fitness and population viability. Trends in Ecology & Evolution, 19(5), 238–244. 10.1016/j.tree.2004.02.002 [DOI] [PubMed] [Google Scholar]
- Green CD, Ma Q, Manske GL, Shami AN, Zheng X, Marini S, … Hammoud SS (2018). A comprehensive roadmap of murine spermatogenesis defined by single-cell RNASeq. Developmental Cell, 46, 651–667. 10.1016/j.devcel.2018.07.025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo HD, Gong YB, He B, & Zhao RQ (2017). Relationships between mitochondrial DNA content, mitochondrial activity, and boar sperm motility. Theriogenology, 87, 276–283. 10.1016/j.theriogenology.2016.09.005 [DOI] [PubMed] [Google Scholar]
- Hedrick PW (2012). Reversing mother’s curse revisited. Evolution, 66(2), 612–616. 10.1111/j.1558-5646.2011.01465.x [DOI] [PubMed] [Google Scholar]
- Holm S (1979). A simple sequentially rejective multiple test procedure. Scandinavian Journal of Statistics, 6(2), 65–70. [Google Scholar]
- Innocenti P, Morrow EH, & Dowling DK (2011). Experimental evidence supports a sex-specific selective sieve in mitochondrial genome evolution. Science, 332(6031), 845–848. 10.1126/science.1201157 [DOI] [PubMed] [Google Scholar]
- Jennions MD, & Petrie M (2000). Why do females mate multiply? A review of the genetic benefits. Biological Reviews, 75(1), 21–64. 10.1111/j.1469-185X.1999.tb00040.x [DOI] [PubMed] [Google Scholar]
- Jodar M, Kalko S, Castillo J, Ballesca JL, & Oliva R (2012). Differential RNAs in the sperm cells of asthenozoospermic patients. Human Reproduction, 27(5), 1431–1438. 10.1093/humrep/des021 [DOI] [PubMed] [Google Scholar]
- Kim D, Langmead B, & Salzberg SL (2015). HISAT: a fast spliced aligner with low memory requirements. Nature Methods, 12, 357–360. 10.1038/nmeth.3317 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koop JL, Zeh DW, Bonilla MM, & Zeh JA (2009). Reproductive compensation favours male-killing Wolbachia in a live-bearing host. Proceedings of the Royal Society B-Biological Sciences, 276, 4021–4028. 10.1098/rspb.2009.1230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, … Durbin R (2009). The sequence alignment/map format and SAMtools. Bioinformatics, 25(16), 2078–2079. 10.1093/bioinformatics/btp352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao Y, Smyth GK, & Shi W (2014). featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics, 30(7), 923–930. 10.1093/bioinformatics/btt656 [DOI] [PubMed] [Google Scholar]
- Lubosny M, Przylucka A, Smietanka B, Breton S, & Burzynski A (2018). Actively transcribed and expressed atp8 gene in Mytilus edulis mussels. PeerJ, 6: e4897 10.7717/peerj.4897 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milot E, Moreau C, Gagnon A, Cohen AA, Brais B, & Labuda D (2017). Mother’s curse neutralizes natural selection against a human genetic disease over three centuries. Nature Ecology & Evolution, 1(9), 1400–1406. 10.1038/s41559-017-0276-6 [DOI] [PubMed] [Google Scholar]
- Moon S, & Akey JM (2016). A flexible method for estimating the fraction of fitness influencing mutations from large sequencing data sets. Genome Research, 26(6), 834–843. http://www.genome.org/cgi/doi/10.1101/gr.203059.115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mossman JA, Slate J, & Birkhead TR (2010). Mitochondrial haplotype does not affect sperm velocity in the zebra finch Taeniopygia guttata. Journal of Evolutionary Biology, 23(2), 422–432. 10.1111/j.1420-9101.2009.01913.x [DOI] [PubMed] [Google Scholar]
- Newcomer SD, Zeh JA, & Zeh DW (1999). Genetic benefits enhance the reproductive success of polyandrous females. Proceedings of the National Academy of Sciences of the United States of America, 96(18), 10236–10241. 10.1073/pnas.96.18.10236 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Padua MV, Zeh DW, Bonilla MM, & Zeh JA (2014). Sisters’ curse: sexually antagonistic effects constrain the spread of a mitochondrial haplogroup superior in sperm competition. Proceedings of the Royal Society B-Biological Sciences, 281(1797). 10.1098/rspb.2014.1686 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker GA (1970). Sperm competition and its evolutionary consequences in insects. Biological Reviews, 45(4), 525–567. 10.1111/j.1469-185X.1970.tb01176.x [DOI] [Google Scholar]
- Patel MR, Miriyala GK, Littleton AJ, Yang HK, Trinh K, Young JM, … Malik HS (2016). A mitochondrial DNA hypomorph of cytochrome oxidase specifically impairs male fertility in Drosophila melanogaster. Elife, 5: e16923 10.7554/eLife.16923.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plazzi F, Puccio G, & Passamonti M (2016). Comparative large-scale mitogenomics evidences clade-specific evolutionary trends in mitochondrial DNAs of Bivalvia. Genome Biology and Evolution, 8(8), 2544–2564. 10.1093/gbe/evw187 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rajender S, Rahul P, & Mahdi AA (2010). Mitochondria, spermatogenesis and male infertility. Mitochondrion, 10(5), 419–428. 10.1016/j.mito.2010.05.015 [DOI] [PubMed] [Google Scholar]
- Ramalho-Santos J, Varum S, Amaral S, Mota PC, Sousa AP, & Amaral A (2009). Mitochondrial functionality in reproduction: from gonads and gametes to embryos and embryonic stem cells. Human Reproduction Update, 15(5), 553–572. [DOI] [PubMed] [Google Scholar]
- Rand DM, Haney RA, & Fry AJ (2004). Cytonuclear coevolution: the genomics of cooperation. Trends in Ecology & Evolution, 19(12), 645–653. 10.1093/humupd/dmp016 [DOI] [PubMed] [Google Scholar]
- Robinson MD, McCarthy DJ, & Smyth GK (2010). edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics, 26(1), 139–140. 10.1093/bioinformatics/btp616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ross JM, Coppotelli G, Hoffer BJ, & Olson L (2014). Maternally transmitted mitochondrial DNA mutations can reduce lifespan. Scientific Reports, 4, 6569 10.1038/srep06569 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruiz-Pesini E, Lapena AC, Diez-Sanchez C, Perez-Martos A, Montoya J, Alvarez E, … Enriquez JA (2000). Human mtDNA haplogroups associated with high or reduced spermatozoa motility. American Journal of Human Genetics, 67(3), 682–696. 10.1086/303040 [DOI] [PMC free article] [PubMed] [Google Scholar]
- SAS Institute Inc. (2012). SAS/STAT® 14.2; User’s Guide. Cary, NC, USA. [Google Scholar]
- Smietanka B, Burzynski A, & Wenne R (2010). Comparative genomics of marine mussels (Mytilus spp.) gender associated mtDNA: rapidly evolving atp8. Journal of Molecular Evolution, 7sas1(5–6), 385–400. 10.1007/s00239-010-9393-4 [DOI] [PubMed] [Google Scholar]
- Smith S, Turbill C, & Suchentrunk F (2010). Introducing mother’s curse: low male fertility associated with an imported mtDNA haplotype in a captive colony of brown hares. Molecular Ecology, 19(1), 36–43. 10.1111/j.1365-294X.2009.04444.x [DOI] [PubMed] [Google Scholar]
- Stephen CL, Reynoso VH, Collett WS, Hasbun CR, & Breinholt JW (2013). Geographical structure and cryptic lineages within common green iguanas, Iguana iguana. Journal of Biogeography, 40(1), 50–62. [Google Scholar]
- Št’áhlavský F, Zeh JA, Zeh DW, & Král J (2009). Karyotypes of the Neotropical pseudoscorpions Semeiochernes armiger and Cordylochernes scorpioides (Pseudoscorpiones: Chernetidae). The Journal of Arachnology, 37, 287–291. [Google Scholar]
- Strutzenberger P, Brehm G, & Fiedler K (2011). DNA barcoding-based species delimitation increases species count of Eois (Geometridae) moths in a well-studied tropical mountain forest by up to 50%. Insect Science, 18(3), 349–362. 10.1111/j.1744-7917.2010.01366.x [DOI] [Google Scholar]
- Sutherland B, Stewart D, Kenchington ER, & Zouros E (1998). The fate of paternal mitochondrial DNA in developing female mussels, Mytilus edulis: Implications for the mechanism of doubly uniparental inheritance of mitochondrial DNA. Genetics, 148(1), 341–347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vaught RC, & Dowling DK (2018). Maternal inheritance of mitochondria: implications for male fertility? Reproduction, 155(4), R159–R168. 10.1530/REP-17-0600 [DOI] [PubMed] [Google Scholar]
- Wade MJ, & Brandvain Y (2009). Reversing mother’s curse: selection on male mitochondrial fitness effects. Evolution, 63(4), 1084–1089. 10.1111/j.1558-5646.2009.00614.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wallace DC (2005). A mitochondrial paradigm of metabolic and degenerative diseases, aging, and cancer: A dawn for evolutionary medicine. Annual Review of Genetics, 39, 359–407. 10.1146/annurev.genet.39.110304.095751 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wallace DC (2013). Bioenergetics in human evolution and disease: implications for the origins of biological complexity and the missing genetic variation of common diseases. Philosophical Transactions of the Royal Society B-Biological Sciences, 368(1622). 10.1098/rstb.2012.0267 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weygoldt P (1969). The Biology of Pseudoscorpions. Harvard Univ. Press, Cambridge, MA. [Google Scholar]
- Wilcox TP, Hugg L, Zeh JA, & Zeh DW (1997). Mitochondrial DNA sequencing reveals extreme genetic differentiation in a cryptic species complex of neotropical pseudoscorpions. Molecular Phylogenetics and Evolution, 7(2), 208–216. 10.1006/mpev.1996.0388 [DOI] [PubMed] [Google Scholar]
- Wolff JN, & Gemmell NJ (2013). Mitochondria, maternal inheritance, and asymmetric fitness: Why males die younger. Bioessays, 35(2), 93–99. 10.1002/bies.201200141 [DOI] [PubMed] [Google Scholar]
- Wolff JN, Pichaud N, Camus MF, Cote G, Blier PU, & Dowling DK (2016). Evolutionary implications of mitochondrial genetic variation: mitochondrial genetic effects on OXPHOS respiration and mitochondrial quantity change with age and sex in fruit flies. Journal of Evolutionary Biology, 29(4), 736–747. 10.1111/jeb.12822 [DOI] [PubMed] [Google Scholar]
- Zeh DW, Zeh JA, & Bermingham E (1997). Polyandrous, sperm-storing females: Carriers of male genotypes through episodes of adverse selection. Proceedings of the Royal Society B-Biological Sciences, 264(1378), 119–125. 10.1098/rspb.1997.0018 [DOI] [Google Scholar]
- Zeh DW, Zeh JA, & May CA (1994). Charomid cloning vectors meet the pedipalpal chelae - single-locus minisatellite DNA probes for paternity assignment in the harlequin beetle-riding pseudoscorpion. Molecular Ecology, 3(5), 517–522. 10.1111/j.1365-294X.1994.tb00130.x [DOI] [PubMed] [Google Scholar]
- Zeh JA, Bonilla MM, Su EJ, Padua MV, Anderson RV, Kaur D, … Zeh DW (2012). Degrees of disruption: projected temperature increase has catastrophic consequences for reproduction in a tropical ectotherm. Global Change Biology, 18, 1833–1842. 10.1111/j.1365-2486.2012.02640.x [DOI] [Google Scholar]
- Zeh JA, Newcomer SD, & Zeh DW (1998). Polyandrous females discriminate against previous mates. Proceedings of the National Academy of Sciences of the United States of America, 95(23), 13732–13736. 10.1073/pnas.95.23.13732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeh JA, & Zeh DW (1996). The evolution of polyandry I: Intragenomic conflict and genetic incompatibility. Proceedings of the Royal Society B-Biological Sciences, 263(1377), 1711–1717. 10.1098/rspb.1996.0250 [DOI] [Google Scholar]
- Zeh JA, & Zeh DW (1997). The evolution of polyandry II: Post-copulatory defences against genetic incompatibility. Proceedings of the Royal Society B-Biological Sciences, 264(1378), 69–75. 10.1098/rspb.1997.0010 [DOI] [Google Scholar]
- Zeh JA, & Zeh DW (2005). Maternal inheritance, sexual conflict and the maladapted male. Trends in Genetics, 21(5), 281–286. 10.1016/j.tig.2005.03.006 [DOI] [PubMed] [Google Scholar]
- Zeh JA, & Zeh DW (2006). Outbred embryos rescue inbred half-siblings in mixed-paternity broods of live-bearing females. Nature, 439(7073), 201–203. 10.1038/nature04260 [DOI] [PubMed] [Google Scholar]
- Zeh JA, Zeh DW, & Bonilla MM (2003). Phylogeography of the harlequin beetle-riding pseudoscorpion and the rise of the Isthmus of Panamá. Molecular Ecology, 12, 2759–2769. 10.1046/j.1365-294X.2003.01914.x [DOI] [PubMed] [Google Scholar]
- Zouros E (2013). Biparental inheritance through uniparental transmission: the doubly uniparental inheritance (DUI) of mitochondrial DNA. Evolutionary Biology, 40(1), 1–31. https://doi-org.unr.idm.oclc.org/10.1007/s11692-012-9195-2 [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Reference mitochondrial genome sequences for C. scorpioides A and B2 haplogroups, mitochondrial DNA expression levels, qPCR mtDNA concentration results and male reproductive success data from the two-male sperm competition experiment are available on Dryad (https://datadryad.org/resource/doi:10.5061/dryad.bh19kj8). The fastq files in support of the differential mitochondrial gene expression analyses have been deposited in the NCBI Short Read Archive under accession numbers SRR7162678 - SRR7162697 for the A haplogroup libraries and SRR7956962 - SRR7956966 for the B2 haplogroup libraries.