Abstract
High throughput T cell receptor (TCR) sequencing allows interrogation of the human TCR repertoire, potentially connecting TCR sequences to antigenic targets. Unlike the highly polymorphic major histocompatibility complex proteins, monomorphic antigen presenting molecules such as MR1, CD1d and CD1b present antigens to T cells with species-wide TCR motifs. CD1b tetramer studies and a survey of the 27 published CD1b-restricted TCRs demonstrated a TCR motif in humans defined by the TCR β chain variable gene 4–1 (TRBV4–1) region. Unexpectedly, TRBV4–1 was involved in recognition of CD1b regardless of the chemical class of the carried lipid. Crystal structures of two CD1b-specific TRBV4–1+ TCRs show that germline-encoded residues in CDR1 and CDR3 regions of TRBV4–1 encoded sequences interact with each other and consolidate the surface of the TCR. Mutational studies identified a key positively charged residue in TRBV4–1 and a key negatively charged residue in CD1b that is shared with CD1c, which is also recognized by TRBV4–1 TCRs. These data show that one TCR V region can mediate a mechanism of recognition of two related monomorphic antigen presenting molecules that does not rely of a defined lipid antigen.
Introduction
Major histocompatibility complex (MHC) encoded antigen-presenting molecules, and the structurally related CD1 and MR1 molecules all present antigens to αβ T cells. In humans, the CD1 family consists of four cell surface-expressed antigen presenting molecules, CD1a, CD1b, CD1c, CD1d, which present lipids to T cells (1). Whereas MHC, CD1, and MR1 proteins all display chemically diverse antigens, a major difference among them involves genetic diversity. Whereas the human MHC locus shows the highest rates of polymorphism among genomes, polymorphisms in the coding regions of CD1 and MR1 are rare. The few single nucleotide polymorphisms in CD1 genes are not known to control antigen presentation, suggesting that CD1 proteins are functionally equivalent among most or all humans (2, 3). These marked differences in the inter-individual variability of MHC, MR1 and CD1 antigen-presenting molecules, together with their distinct structural features, potentially translate into differing patterns of αβ T cell receptors (TCRs) controlled by these systems.
Based on more than 100 solved MHC-peptide-TCR structures (4–7), docking of TCRs on MHC is well understood in general terms. First, the variable (V) domains of the α chain and β chain are positioned over the α2 helix and α1 helix of MHC-I, respectively. Second, variable (V) gene-encoded complementarity determining region (CDR) 1 or CDR2 frequently interact with the outer α-helices of MHC molecules. Despite notable exceptions (8, 9), this general structural model might predict that certain MHC types or allomorphs preferentially interact with certain V gene-encoded segments, and that MHC haplotype biases the selected TCR repertoire in ways that can be specifically detected by sequencing of the TCR repertoire. However, there is only limited data available to support this concept, and the existing data only account for a minor association between MHC haplotype and TCR gene bias (10). The alternative hypothesis is that population-based genetic diversity in the MHC system, the diversity of the bound peptides for any allomorph, and the fact that each MHC molecule or allomorph can interact with many TCR V segments, all contribute to extreme TCR repertoire diversity, such that connections between individual MHC-peptide complexes and TCR gene segments or sequences is presently undecipherable. Discerning which of these two views is correct is not only a fundamental question for immunologists, but is of potential practical importance, given rapid advances in high throughput TCR sequencing and emerging interest in immunodiagnosis based on TCR sequence bias in peripheral blood T cells (11, 12).
Considering the entire human population, the human leukocyte antigen (HLA) complex system uses > 10,000 allomorphs to present peptides (13), but the human CD1 system uses essentially four monomorphic proteins to present antigens, and the human MR1 system uses only one. Thus, CD1 and MR1 can be thought of as genetically simple cases to test questions relating to connection of individual antigens with TCR gene bias. In the CD1 and MR1 systems clear examples of TCR bias, which extends broadly or universally to all humans have been identified. The most widely known example is type I Natural Killer T (NKT) cells, which recognize CD1d-α-galactosylceramide complexes. In human type I NKT cells, TRAV10 genes rearrange to TRAJ18 to form identical or almost identical TCR α chains that pair with moderately diverse TCR β chains. A more recently discovered example is the GEM TCR that uses TRAV1–2-TRAJ9 genes with nearly identical TCR α chains to recognize CD1b bound to mycobacterial glucose monomycolate antigens (GMM) (14, 15). Similarly, the monomorphic MR1 displays vitamin B metabolites known as (2-oxopropylideneamino)-6-D-ribitylaminouracil (5-OP-RU) to activate human MAIT TCRs with invariant TRAV1–2-TRAJ33 α chains (16), and coevolution between MR1 and TRAV1–2 has been suggested (17).
In these three examples, the invariant TCR α chains pair with moderately diverse TCR β chains to form ‘semi-invariant’ TCRs. Within one individual and among unrelated individuals, variations of these invariant TCR α chains are limited to a few nucleotide substitutions, which is referred to as type 3 bias (18). These three examples clearly establish linkages between named antigens and defined TCR motifs, but whether TCR bias dominates in other parts of the human CD1-reactive repertoire remains unknown. Human CD1a, CD1b and CD1c proteins also activate T cells with diverse TCR patterns (19) and type II NKT cells with TCRs that express diverse TCR are well described for humans and mice (20–25).
Based on six clones, we previously identified a human T cell type known as LDN5-like T cells, which are named after the prototype clone, LDN5 (26). We tentatively defined LDN5-like T cells by specificity for CD1b and mycobacterial GMM, as well as the expression of TRBV4–1 or TRAV17-utilizing TCRs (27). As contrasted with type I NKT cells, MAIT cells and GEM T cells, LDN5-like T cells show lower affinity binding to their antigenic target and much less stringent TCR sequence conservation (14, 15, 27): none of their TCR chains are invariant. Here we used CD1b tetramers with defined, but chemically diverse lipid antigens to demonstrate a connection between CD1b recognition and TRBV4–1 expression in clones, lines and polyclonal T cells from unrelated donors demonstrating a broad relationship between a TCR variable region and its protein target. Unexpectedly, these studies show that TRBV4–1 expression correlates with CD1b specificity, but not with lipid antigen specificity. Thus, LDN5-like cells are just one example of what is actually a broader pattern of TRBV4–1 usage by CD1b-specific T cells. Finally, together with the identification of TRBV4–1 enrichment in the human CD1c-reactive T cell repertoire (28), these data spurred a crystallographic study of two CD1b-specific TRBV4–1+ TCRs and mutational analysis of TCR and CD1b. These studies provide insights into how genome-encoded parts of the TRBV4–1+ TCR could potentially mediate recognition across two distinct monomorphic antigen-presenting molecules.
Material and methods
Protein production, antigen loading, and tetramerization
For tetramers, 20 μg CD1b monomers obtained from the National Institute Health (NIH) tetramer facility were loaded with 32 μg PG (Avanti lipids), GMM (Bill and Melinda Gates Foundation Lipid Bank) or synthetic BbGL-II (a gift from J. Kubler-Kielb, NIH). Lipids were dissolved in citrate buffer pH 4.5 with 0.5% 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonate (Sigma) and incubated overnight at 37°C. After incubation the pH was neutralized by adding Tris pH 8.5. As negative control CD1b monomers were treated in the same way without adding exogenous lipids lipid, so they carry endogenous (CD1b-endo) lipids from the expression system. CD1b mutants and wild type control for experiments using these mutants, were produced in HEK293 S GnTI− (American Type Culture Collection) and purified via HisTrap Metal chelating-Ni2+ affinity chromatography and size exclusion. For SPR, purified CD1b was loaded with a molar excess of C36 GMM (Bill and Melinda Gates Foundation lipid bank) over a 16 h incubation period at 20°C and purified by anion exchange chromatography. Recombinant PG10 and clone 2 TCRs were cloned into the pET30a vector and expressed in Escherichia coli BL21(DE3) as insoluble inclusion bodies, refolded in 20 mM Tris-HCl (pH 8.0), 5M urea, 400 mM L-arginine, 0.5 mM Na-EDTA, 5 mM reduced glutathione, and 0.5 mM oxidized glutathione for 3 days at 273 K, and purified.
Crystallization and structure determination of PG10 and clone 2 TCRs
Crystals of the PG10 and clone 2 TCRs were grown by hanging drop vapor diffusion, with a protein-mother liquor drop ratio of 1:1, at a protein concentration of 5 mg mL−1 in 10 mM Tris-HCl (pH 8.0), 150 mM NaCl, with a crystallization condition of 22 % (v/v) PEG 3350, and 0.2 M ammonium sulphate. Crystals were soaked in a cryoprotectant of mother liquor comprising 10 % glycerol or ethylene glycol and flash frozen in liquid nitrogen. Data were collected at the Australian Synchrotron at the MX2 Beamline (29) for both TCRs. Data were processed using the iMosflm software and scaled using Aimless as part of the CCP4i Program Suite (30). Crystal structures were solved by molecular replacement using PHASER (31). Crystal structures of the PG90 TCR (PDB accession code: 5WJO, https://www.rcsb.org/structure/5WJO) and GEM42 TCR (PDB accession code: 4G8F, https://www.rcsb.org/structure/4G8F) were used as models for solving PG10 and clone 2 respectively. Manual adjustment of solved models was conducted in the Coot graphics program (32), followed by maximum-likelihood refinement using Phenix-refine (31). All molecular representations were generated in PyMOL. Core r.m.s.d. values were calculated using Coot (32), with alignments of the TCRs that were generated against the TCR constant domains.
Surface Plasmon Resonance of clone 2 and CD1b-GMM
SPR analysis of CD1b-GMM and clone 2 TCR WT and mutants was conducted on the BIAcore 3000 instrument at 20°C in 10 mM Tris-HCl (pH 8.0), 150 mM NaCl, and 1 % (w/v) bovine serum albumin. WT and mutant CD1b-GMM expressing a BirA tag were biotinylated and non-covalently coupled to a streptavidin (SA) chip. Purified refolded clone 2 in solution was passed over CD1b as the analyte at 5 μL/min. All experiments were conducted as two independent experiments in duplicates. Data analysis and visualization was conducted using Graphpad prism 7.0, using the 1:1 Langmuir binding model.
Human subjects
PBMCs were isolated from venous blood from donors in Lima, Peru, which were recruited under oversight from the Institutional Committee of Ethics in Research (CIEI) of the Peruvian Institutes of Health, the Institutional Review Board (IRB) of the Harvard Faculty of Medicine, and the Partners Healthcare IRB. Peruvian patients provided oral and written informed consent in Spanish and met study criteria for lack of prior tuberculosis infection defined as negative Quantiferon test result and no clinical evidence of active tuberculosis. Separately, random blood bank donor-derived PBMCs were obtained from the Brigham and Women’s Hospital Specimen Bank in Boston, USA or donated by an asymptomatic tuberculin skin test-positive subject with no clinical or radiographic evidence of active tuberculosis, at the Massachusetts General Hospital blood bank in Boston, USA.
T cell lines
The previously published primary T cell lines A25Salmonella, which contains the clones PG10 and PG90 (33), LDN5 (26), BC24A, BC24B, and BC24C (34) were grown by stimulation with 30 ng/mL anti-CD3 antibody and 25 × 106 irradiated PBMC and 5 × 106 irradiated Epstein-Barr virus-transformed B cells, and 1 ng/mL IL-2, which was added on day 2 of the culture. To study tetramer binding to the GEM42 TCR, we used a TCR-transduced 5KC-78.3.20 hybridoma (14, 15). To generate primary TRBV4–1+ and TRBV4–1− T cell lines, PBMC from random blood bank donor D43 were stained with anti-CD3 (555342, BD) and anti-TRBV4–1 (IM2287, Beckman Coulter). Cells were sorted on a BD FACSAria (BD Biosciences), and lymphocytes were selected based on forward scatter and side scatter. 1×106 CD3+TRBV4–1+ and 1×106 CD3+TRBV4–1− cells were sorted. After 2 weeks of stimulation as described above both cells were stained with anti-CD3 and anti-TRBV4–1 antibodies to check purity.
Single cell TCR sequencing
To each well of a Vapor-Lock (Qiagen)-coated 96 well plate (Eppendorf) a mixture of 0.5 μl 5x reaction buffer, 0.5 μl reverse transcriptase (Iscript, Bio-Rad), and 1.25 μl H2O was added per well, with a final concentration of 0.1 % 4-(1,1,3,3-tetramethylbutyl)phenyl-polyethylene glycol (Triton X-100). Single cells were sorted into individual wells in this 96 well plate using a FACSAria cell sorter (BD biosciences). The plate was centrifuged at 3000 rotations per minute at 4°C for 10 min. For cDNA synthesis, the plate was incubated at 25°C for 5 min, followed by 42°C for 30 min, and 80°C for 5 min. TCR transcripts were amplified in two subsequent, nested polymerase chain reactions (PCRs). The primary reaction consisted of 2.5 μl of the cDNA synthesis reaction mixture as a template, 0.75 units of Thermus aquaticus (Taq) polymerase (Denville), 2.5 μl 10x PCR buffer (Denville), 0.5 μl 10 mM dNTPs, 2.5 pMol of each external TRAV and TRBV primer, 10 pMol antisense TRAC, and 10 pMol antisense TRBC primer as described by Wang (35) in a total volume of 25 μl. The following PCR conditions were used: 95°C for 2 min, 35 cycles of (95°C for 20 sec, 50°C for 20 sec, 72°C for 45 sec), followed by 1 cycle of 72°C for 7 min. This reaction mixture was used as a template in two separate secondary PCR reactions. The mixtures are identical to the primary PCR except that in one reaction the internal TRAV and TRAC primers were used and in the other reaction the internal TRBV and TRBC primers as described by Wang (35). The following PCR conditions were used for the secondary PCR: 95°C for 2 min, 35 cycles of (95°C for 20 sec, 56°C for 20 sec, 72°C for 45 sec), followed by 1 cycle of 72°C for 7 min. PCR products were analyzed by gel electrophoresis and α and β chain PCR products that resulted from the same single cell-containing well were sequenced by Sanger sequencing.
Flow Cytometry
T cells (3×106) and tetramers (0.2 μg per 50 μl staining volume) were incubated for 10 min at room temperature in phosphate buffered saline (PBS) with 1% bovine serum albumin (BSA). Subsequently anti-CD3 (clone SK7, BD Biosciences) antibody was added an incubated for 10 min at room temperature. Subsequently, additional antibodies (anti-TRBV4–1, IM2287, Beckman Coulter) were added and incubated for 20 min at 4°C. Cells were washed with PBS with 1% BSA and analyzed on an LSRFortessa (BD Biosciences) flow cytometer. For display in a matrix, percentage positive cell or mean fluorescence intensity of the positive cells were normalized to Z score and a heat map was created using the heatmap.2 function of R (https://cran.r-project.org/web/packages/gplots/index.html).
Statistics
In Figure 3C, significance was calculated by a paired Wilcoxon ranked sums test. Significance of the data in Fig. 1A was calculated using a Kruskal-Wallis test.
Figure 3: TRBV4–1+ T cells are enriched among CD1b tetramer+ T cells.
(A) PBMC from three blood donors were stimulated with autologous monocyte-derived dendritic cells and mycolic acid (MA) for 18 days. The resulting cells were stained with CD1b-MA tetramers and an anti-CD3 followed by sorting of tetramer+ and tetramer− cells as shown in Supplemental Figure 1D and subjected to high throughput TCR sequencing. The percentages of TRBV gene usage are shown. (B) Using the approach described above, we reanalyzed a publicly available dataset (48) of CD1b-glucose monomycolate (GMM) tetramer+ and tetramer− cells from four donors. (C) Summary of TRBV4–1 percentage among tetramer+ and tetramer− cells of the three donors shown in A and four donors shown in B.
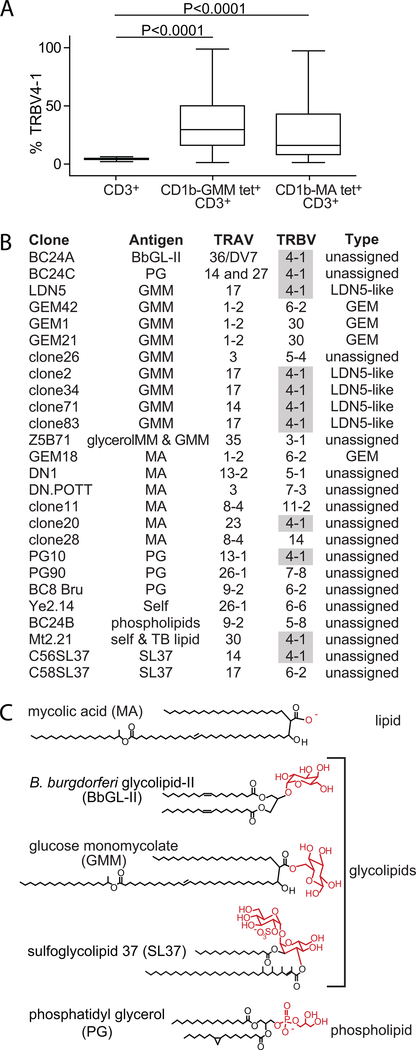
Figure 1. TRBV4–1+ CD1b-restricted clones recognize a wide variety of antigens.
(A) PBMC from 49 healthy donors were pre-gated for CD3 expression and CD1b-GMM tetramer or CD1b-MA tetramer binding (Supplemental Fig. 1A). Within the tetramer− or tetramer+ gates, the percentage TRBV4–1-expressing cells was determined. Box: median with interquartile range; whiskers: minimum to maximum value. (B) Previously published CD1b-restricted TCRs are shown according to variable (TRAV, TRBV) genes and the antigen they recognize: LDN5 (26); GEM1, GEM18, GEM21,GEM42 (27); clone 2, clone26, clone34, clone71, clone83 (27); Z5B71 (36); DN1 (37); DN.POTT (19, 38); PG10, PG90, BC8 Bru (33); YE2.14 (39); MT2.21 (40); C56SL37, C58SL37 (41); clone 11, clone 20, clone 28 (42), BC24A, BC24B, and BC24C (34). BC24C expresses two different α chains. (C) Lipid antigens illustrate the structural diversity of molecules, with polar head groups indicated in red, recognized by TRBV4–1 TCRs.
Results
TRBV4–1 TCRs recognize CD1b bound to diverse lipid antigens
A prior study of 5 clones (LDN5, clone 2, clone 71, clone 34, clone 83) defined LDN5-like T cells as expressing a candidate TCR motif encoded by TRBV4–1 or TRAV17, which recognized mycobacterial GMM presented by CD1b (27). However, these clones were isolated from only four blood donors, and the distribution of LDN5-like cells among the broader human population was not determined. We designed a study in a cohort of 49 healthy donors from Lima, Peru, that lacked positive antigen-recall tests for tuberculosis, but were likely vaccinated with BCG according to the Peruvian national vaccination program. Besides CD1b-GMM and CD1b-mycolic acid (MA)-tetramers, an antibody against TRBV4–1 was included in the cytometry panel to enable the identification of LDN5-like T cells (CD3+, CD1b-GMM tetramer+, TRBV4–1+). An antibody against TRAV17 might have improved identification of LDN5-like T cells but such an antibody is unavailable. We found that TRBV4–1 usage among CD1b-GMM tetramer+ T cells (mean 35 %) was ~10-fold higher than among the total T cell population (3.2 %) (P < 0.0001). Thus, initial results from 5 clones were confirmed in a large study of polyclonal T cells among unrelated donors demonstrating the existence LDN5-like cells as a defined, trackable T cell type in humans (Supplemental Fig. 1A and Fig. 1A).
Whereas these and prior results were generated using GMM antigen, we also observed TRBV4–1 usage among CD1b-MA tetramer+ T cells (mean 26 %) was also highly enriched above the total T cell population (P < 0.0001). This finding was unexpected because free mycolic acid (MA) lacks the glucose that has previously been demonstrated to be crucial for the recognition of GMM by LDN5 (26). This finding prompted the alignment of all available complete TCR sequences from a large panel of 26 previously published, functionally confirmed CD1b-restricted T cell clones. (19, 26, 27, 33, 34, 36–42). Similar to frequencies of TRBV-1 expression in CD1b tetramer positive T cells, eleven (42 %) of CD1b reactive clones expressed TRBV4–1 (Fig. 1B). Excluding TCRs that conform to the other known CD1b-reactive TCR motif (GEM T cells), 11 of 22 TCRs (50 %) expressed TRBV4–1. These rates are well in excess of TRBV4–1 expression among randomly chosen T cells, as measured by TCR variable region specific antibodies (~1 percent) or by high throughput TCR sequencing (0.5 – 4%) (43–45).
However, prior studies had not tested or considered candidate TCR motifs for antigens other than the mycobacterial glycolipid GMM. Overall, this search identified at least four additional structurally distinct lipid antigens presented by CD1b to TRAV4–1 encoded TCRs: Borrelia burgdorferi glycolipid II (BbGL-II), sulfoglycolipid 37 (SL37), phosphatidylglycerol (PG), and MA. Although most GMM-specific TRBV4–1+ clones known as LDN5-like T cells co-express TRAV17 (27), none of the TRBV4–1+ clones that recognize these four other lipid antigens express TRAV17. These lipids are not structurally related to one another or to GMM with regard to lipid tails, carbohydrate groups or inorganic sulfate or phosphate groups (Fig. 1C). In particular, the lipid head groups that protrude from the CD1b cleft (34, 46, 47) and function as TCR epitopes are comprised of a hexose sugar, a dihexose sugar, a phosphoglycerol unit, or a small negatively charged headgroup with no carbohydrate, respectively (Fig. 1C). These data confirmed and extended the relationship between TRBV4–1 and CD1b reactivity but raised new questions about the specificity of TCRs for lipids carried by CD1b. In particular, could TRBV4–1 sequences mediate recognition of CD1b while ignoring the antigen carried by CD1b, while other parts of the TCR are available for antigen recognition?
CD1b-specific T cells among polyclonal T cell lines
Because these T cell clones were derived with diverse methods and from genetically unrelated donors, we carried out controlled experiments in which TRBV4–1+ and TRBV4–1− cells were sorted from the peripheral blood mononuclear cells (PBMC) of the same blood bank donor 43 (D43) and cultivated in parallel, under equivalent conditions. Using a monoclonal antibody that binds to TCRs that use TRBV4–1, equal numbers of CD3+TRBV4–1+ and CD3+TRBV4–1− T cells were sorted (Supplemental Fig. 1B) and expanded by stimulation with anti-CD3 antibody, after which the uniform expression or absence of TRBV4–1 by the cell lines was confirmed (Fig. 2A). Both T cell lines were stained with CD1b tetramers that were mock treated and so contained diverse cell-derived endogenous lipids (CD1b-endo), or were loaded with PG or GMM as examples of structurally divergent phospholipid or glycolipid antigens. In the TRBV4–1+ polyclonal T cell line there was an increased frequency of cells that bind to CD1b-PG or CD1b-GMM compared to the TRBV4–1− population (Fig. 2B). Thus, TRBV4–1+ is more likely to confer CD1b-reactivity than the combined action of the other Vβ segments. Further the CD1b reactivity is not limited to CD1b bound to one specific antigen. In fact, PG and GMM are very different, as structural studies of TCR-CD1b-lipid complexes show that they use distinct glucose versus phosphoglycerol units on the TCR-facing surface of CD1b (14, 46).
Figure 2: CD1b recognition by TRBV4–1+ T cells.
(A) Cell lines sorted from blood bank donor D43 based on expression or absence of TRBV4–1 (Supplemental Figure 1B), were tested for TRBV4–1 expression. (B) Both cell lines were stained with CD1b-PG or CD1b-GMM tetramers, or mock loaded CD1b tetramers carrying diverse endogenous lipids (CD1b-endo). (C) TCR sequences were obtained by single cell TCR sequencing of CD1b-GMM tetramer+ and CD1b-PG tetramer+ cells are shown with germline (grey) and non-germline (white) residues encoded by the indicated variable and joining region genes. After pre-gating using anti-CD3 and anti-TRBV4–1 TCR antibodies (D), TRBV4–1+ and TRBV4–1− T cells from PBMC from three random blood bank donors were analyzed for binding of CD1b-PG and CD1b-GMM tetramers directly ex vivo (E). Equal numbers of cells are shown in each plot. All acquired TRBV4–1− cells are shown in Supplemental Figure 1B (top).
Confirming TRBV4–1 expression, single cell TCR sequencing demonstrated that among the CD1b-PG binding cells were several single cells that expressed identical pairs of α and β chains composed of TRAV8–2-TRAJ38 and TRBV4–1-TRBJ1–6. Likewise, among cells that bind to CD1b-GMM were several cells that expressed identical pairs of α and β chains composed of TRAV13–2-TRAJ32 and TRBV4–1-TRBJ1–4 (Fig. 2C). Even though in healthy blood bank-derived donors the frequency of GMM- and PG-specific cells is typically below or at the detection limit of ~10−4, pre-enrichment for TRBV4–1 apparently enriches for CD1b-specific cells to a level that they can now be detected.
CD1b-specific T cells among TRBV4–1+ T cells ex vivo
To bypass artifacts related to T cell culture and outgrowth, we next analyzed polyclonal T cells directly ex vivo. We used fresh PBMC from three random blood bank donors (D6, D7, D48) and co-stained them with anti-CD3, anti-TRBV4–1 and CD1b tetramers treated with the phospholipid PG or the glycolipid GMM, to measure the rate of staining with tetramers in the TRBV4–1+ T cell gate (Fig. 2D–E). As negative controls, we measured staining with mock treated CD1a tetramers (Supplemental Fig. 1C) and CD1b tetramer staining in the TRBV4–1− gate (Fig. 2D–E). In all three donors, we detected a higher percentage of cells that stain with CD1b-PG or CD1b-GMM tetramer in the TRBV4–1+ population compared to the TRBV4–1− population (Fig. 2E) or any T cell population stained with CD1a tetramers (Supplemental Fig. 1C). Most tetramer+ T cells stained with both tetramers, which suggests that they represent a broadly cross-reactive population that has been described previously (34). The lack of T cells that were single positive for the CD1b-GMM tetramer is most likely due to extremely low frequencies of CD1b-GMM-positive T cells among blood bank donors in Boston, which are most likely not exposed to BCG vaccine or M. tuberculosis. Overall, enrichment of CD1b tetramer+ cells among TRBV4–1+ T cells was confirmed directly ex vivo. Thus, polyclonal T cells from genetically unrelated donors provided a new and strong linkage between TRBV4–1 TCR expression and CD1b recognition.
High throughput sequencing of TCR rearrangements in CD1b tetramer-positive and -negative T cells
Next, we used high throughput TCR sequencing (Adaptive Biotechnologies, Seattle, USA) to measure the TRBV gene usage in functional TCR rearrangements among T cells that were sorted into CD1b tetramer+-enriched and CD1b tetramer− populations (Fig. 3). We chose MA as the ligand for CD1b tetramers based on its antigenic properties in the CD1 system (19, 37, 42), including its stimulation of polyclonal (Fig. 1A) and clonal (Fig. 1B) TRBV4–1+ T cells. In Boston, we obtained PBMC from two blood bank donors (CX and C63) and one person who was latently infected with M. tuberculosis (C58). T cells were stimulated once with autologous monocyte-derived dendritic cells and MA and cultured for 17 days prior to sorting into CD1b-MA tetramer+ and CD1b-MA tetramer− populations (Supplemental Fig. 1D). The junctional regions of their TCRs were sequenced in high throughput and their V genes assigned, showing that tetramer+ cells used TRBV4–1 at markedly increased rates for donors CX and C58 (Fig. 3A, green arrows). In donor C63, the post-sort analysis demonstrated high contamination with tetramer− cells (92.6 %) in the ‘tetramer+’ cells (Supplemental Fig. 1D), so a sort failure explained the highly similar profiles for TRBV genes in this patient.
To increase the number of known antigens and donors analyzed, we took advantage of a publicly available dataset, which used a similar approach based on CD1b-GMM tetramer sorting for high throughput TCR sequencing (48). Similar to CD1b-MA tetramer results, all four patients analyzed with CD1b-GMM tetramers showed marked enrichment of TRBV4–1 among tetramer+-enriched cells as compared to tetramer− cells (Fig. 3B). Overall, TRBV4–1 use among CD1b tetramer+-enriched polyclonal T cells was higher in all six subjects studied and the result was statistically significantly (p = 0.016) (Fig. 3C). Further, because MA and GMM are chemically distinct antigens, and TCRs specific for one do not typically cross-react with the other (14, 15, 26, 27, 49), preferred expansion of TRBV4–1+ cells by both antigens suggests that TRBV4–1 is driving CD1b recognition rather than lipid antigen recognition, in agreement with the clonal T cell analyses (Fig. 1).
Detectable TCR conservation is limited to the V gene
TRBV4–1 encodes all CDR1β and CDR2β amino acids and the first four amino acids of CDR3β. The C-terminal part of CDR3β is encoded by non-germline encoded N nucleotides, D segments and part of the J segment. To develop hypotheses about which TCR β chain residues could interact with CD1b we examined the known CD1b-reactive, TRBV4–1+ β chain sequences in more detail. Among 12 TRBV4–1+ TCRs that recognize CD1b in combination differing lipid, glycolipid or phospholipid antigens, we observed no enrichment for any single TRBJ segment or a preference for TRBJ segments that belong to the TRBC1 or TRBC2 group (Fig. 4A). Further, we could not discern CDR3β common sequence patterns beyond the TRBV4–1-encoded residues at positions 104–107 (CASS). This is in sharp contrast with MAIT, NKT, and GEM TCRs which have TCR α chains that are nearly identical in length and sequence, including CDR3α regions, and are formed by identical V and J segments. Therefore, further studies investigated the hypothesis that TRBV4–1 encoded residues might mediate recognition of CD1b.
Figure 4: Structural analysis of TRBV4–1+ TCRs.
CD1b-specific TRBV4–1+ TCR conservation is limited to the germline encoded TRBV gene. (A) The schematic shows the role of TRB locus genes in encoding residues in the CDR1, CDR2 and CDR3 regions. CDRβ regions of new and previously sequenced TRBV4–1+ and TRBV4–1− CD1b-specific clones were aligned. BbGL-II is 1,2-di-oleyl-a-galactopyranosyl-sn-glycerol; SL37 is synthetic di-acylated sulfoglycolipid analog (67). (B) Upper: Side view of clone 2, PG10, PG90 and GEM42 TCRs, with α-chains (grey), TRBV4–1 β-chains (cyan), and other β chains (green and pink) highlighted. Lower: Bottom-up view of TCR interface surface electrostatic potential. (C) In comparison, top-down view of the CD1b interface surface electrostatic potential are shown (right), with CD1b presenting GMM (brown, upper), and PG (blue, lower). Potential contours are shown on a scale from + 5.0 (positive charge, blue) to – 5.0 kBT e−1 (negative charge, red); white indicates value close to 0 kBT e−1 (neutral charge). (D) Overlay image shows CDR regions of clone 2 (green) and PG10 (Blue) TCRs. (E) Key interactions in the TRBV4–1+ CDRβ regions are shown, including positions of H29β and R30β on the CDR2β region of clone 2 (green, left) and PG10 (blue, right) TCRs. Amino acid residues involved in contacts are represented as sticks, with hydrogen bonds represented as black dashes. Nitrogen, oxygen, and phosphate are represented in blue, red, and orange respectively, and color coding of CDR regions are highlighted in the legend.
Structural analysis of conserved TRBV4–1 TCRs
First, we focused on the residues encoded by TRBV4–1 that encode CDR1β (MGHRA), CDR2β (YSYEKL) and the first four residues of CDR3β (CASS) (Fig. 4A). For a detailed understanding of where these residues are positioned within αβ TCR heterodimers, we cloned two TRBV4–1+ CD1b-reactive TCRs (clone 2 and PG10 TCR), expressed them as heterodimeric proteins, and determined their structures to 1.8 Å and 2.5 Å resolution, respectively (Supplemental Table 1, Fig. 4B). The clone 2 TCR recognizes CD1b-GMM (27) and so represents an LDN5-like TCR. The PG10 TCR recognizes CD1b-phosphatidylglycerol, so represents the non-GMM specific TRBV4–1+ TCR motif identified here (33). These were compared to two existing structures of TRBV4–1− TCRs: a GEM TCR (GEM42) and a phosphatidylglycerol-CD1b-reactive TCR (Fig. 4B).
Structural comparisons between the TRBV4–1+ and TRBV4–1− TCRs reveals extensive electropositive regions at the antigen recognition site. Compared to PG90 and GEM42 TCRs, the TRBV4–1+ TCRs exhibit greater electropositivity in a region located predominantly between the CDR3 loops, extending towards the CDR2β loop (Fig 4B, blue). This electropositive feature correlates with the electronegative surface of the CD1b antigen-binding cleft (Fig. 4C, red). As previously determined co-crystal structures of PG90-CD1b-PG and GEM42-CD1b-GMM demonstrate selective co-recognition between an electropositive TCR interface and electronegative CD1b (14, 34), a broadly similar mode of docking for the TRBV4–1+ TCRs onto CD1b might potentially operate.
Despite the different lipid antigens recognized, the overall positioning of the Cα carbon backbone, the amino acid residue side chain position in the CDR1β loop, and the germline-encoded CASS region of the CDR3β loop were highly conserved between the two TRBV4–1+ TCRs, with root mean squared deviation (r.m.s.d.) values of 0.2 Å and 0.1 Å, respectively (Fig. 4D). The CDR2β loop of the clone 2 TCR is involved in crystal contacts, and as such, aligned less well to the PG10 TCR CDR2β loop (r.m.s.d. value of 1.3 Å). Notably, the position of H29β in the CDR1β loop is highly conserved in both TCRs, being wedged between the germline encoded S106β and P108β residues of the CDR3β region (Fig. 4E). Here, H29β appears to play a stabilizing role that anchors the germline-encoded N-terminal end of the CDR3β loop via hydrogen bond formation with S106β. In turn, the main chain carbonyl group of H29β is stabilized via hydrogen bonds by the CDR2β residues S57β or Y58β (Fig. 4E). Of interest, a similar interaction is observed in the two non-TRBV4–1 TCRs, PG90 and GEM42. Despite using TRBV7–8 and TRBV6–2 respectively, H29β is also encoded in the CDR1β region (Fig. 4A), and stabilizes the germline encoded CDR3β region. By analogy to the known docking modes of PG90 and GEM42 TCRs (Supplemental Fig. 2), this inter-CDR1β-CDR3β region is positioned distally to the CD1b-lipid-TCR interface, where it acts to stabilize the CDR3β region to allow for direct lipid headgroup contact. In the absence of a trimolecular TRBV4–1+ TCR-CD1b-lipid structure it is unclear whether H29β plays a similar role when the TCR uses TRBV4–1.
While H29β is conserved in these four TCRs, R30β is present only in the two TRBV4–1+ TCRs (Fig 4A). Here, R30β was solvent exposed and adopted two distinct conformations in the Clone 2 and PG10 TCRs (Fig 4D). In the PG10 TCR structure, R30β hydrogen bonds with Y58β, where the residue contributes towards the electropositive interface surface of the PG10 TCR. In the Clone 2 TCR structure, R30β is orientated away from the interface in a manner not related to crystal contacts (Fig 4D), and as such, the interface surface is less electropositive (Fig 4B). Due to the electrostatic complementary nature of the TCR and CD1b interface, and the positioning of R30β towards the interface surface in the PG10 TCR structure, this residue might be involved in contacts between CD1b and TRBV4–1.
Mapping of essential residues in the CD1b-clone 2 TCR interaction
These crystal structures guided additional alanine substitution experiments to test which TRBV4–1 CDR residues are crucial for recognition of CD1b-GMM by the clone 2 TCR. Mutations along the CDR1β and CDR2β regions were generated, but recombinant protein production was reduced dramatically with CDR2β mutants. The side chains of these residues are solvent exposed, so changes of this type are known to affect overall protein solubility and integrity. To bypass this technical limitation, we mutated CDR1β amino acid H29β, R30β, and a framework region 1 residue, T16, as a negative control. Surface plasmon resonance (SPR) experiments were performed to measure steady state binding affinities between these TCR mutants and GMM-loaded CD1b monomers coupled to a sensor chip. The KD of wild type clone 2 TCR and the T16A mutant were comparable: 6.9 ± 1.0 μM and 6.1 ± 0.6 μM respectively (Fig. 5A). However, both mutations in the CDR1β region showed a marked decrease in CD1b-GMM binding affinity, resulting in a KD of >200 μM (Fig. 5A). This indicates that the H29β and R30β are critical TRBV4–1+ residues for mediating high affinity TCR interactions with CD1b-GMM. Based on the locations of the residues in CDR1 loop, such effects are likely mediated via an indirect (H29β) or direct (R30β) impact on TRBV4–1 docking onto CD1b.
Figure 5. Mutational analysis of the TCR-CD1b interaction.
(A) CD1b-reactive clone 2 TCR was expressed as a heterodimer encoded by wild type sequences or subjected to β chain point mutation with alanine substitutions in the TRBV4–1 encoded region at positions 29 or 30. Binding to GMM-loaded CD1b complexes was measured using surface plasmon resonance to generate steady state affinities, with binding curves (upper) and equilibrium curves (lower) shown here. (B) The steady state affinities of the wild type clone 2 TCR for wild type and mutant CD1b proteins loaded with GMM was determined. Equilibrium curves for CD1b-E80A, CD1b-Y151A, and CD1b-Y154A showed no observable binding. Error bars represent mean + standard error of the mean (SEM). (C) Surface representation of the CD1b-GMM surface (white), with residues, when mutated to alanine, demonstrate less than a 3-fold decrease in affinity (yellow), 3–5-fold decrease in affinity (orange), and greater than a 5-fold decrease (red), upon binding against the clone 2 TCR. Positions of residues E80, Y151, I154, and T157 are indicated.
Next, we carried out alanine scanning mutation across the surface of CD1b to functionally evaluate the CD1b-clone 2 TCR interaction via SPR (Fig. 5B). As the goal was to specifically assess the TCR-CD1b interaction interface, we did not mutate sites that function as interdomain tethers to control antigen entry into the cleft (D60, E62) (50). Instead, we studied ten alanine point mutants on the outer surface of CD1b, which are located on the TCR facing aspect of the α1 (E65A, I69A, V72A, R79A, E80A) and α2 (Y151A, I154A, T157A, R159A, I160A) helices, as determined from prior crystal structures (14, 51). Only E80A (non-binding), Y151A (non-binding), I154A (non-binding), and T157A (38.9 + 8.3), mutants demonstrated a significant reduction in steady state affinities, resulting in significantly reduced or abrogated Clone 2 TCR binding onto CD1b-GMM (Fig. 5B).
Consistent with this interpretation, most human CD1 group 1-reactive TCRs, including all three of the known CD1b-binding αβ TCRs (14, 34, 46) bind CD1 such that the TCR α-chain is positioned over the A’-roof, distant from the hot spot established here. The established hot spot that significantly affects Clone 2 TCR binding resides at the F’ portal and involves E80 and Y151 (Fig. 5C). As observed with the GEM42 and PG90 TCRs, this site is critical for TCR β chain interaction with CD1b (14, 46).
Conserved functional hotspots for all TRBV4–1+ TCRs
Next, we wanted to determine if any CD1b residues are important in the interaction with the broader spectrum of CD1b-reactive TCRs. First, we assembled a panel of TRBV4–1+ (LDN5, PG10, D43, BC24C) and TRBV4–1− antigen-specific, CD1b reactive clones (BC24B, PG90, GEM42). These clones were evaluated for staining with human CD1b tetramers that were treated with the relevant glycolipid (GMM) or phospholipid (PG) antigen and CD1a tetramers as a negative control. At the same time 13 alanine substituted CD1b proteins (K61A, E65A, E68A, I69A, V72A, R79A, E80A, Q150A, Y151A, Q152A, E156A, I160A, E164A) were assembled into tetramers, mock or antigen treated, and then tested on all clones to map the functional interaction sites on CD1b. Finally, the clone BC24B was known to show autoreactive response to mock-loaded CD1b and PG-treated CD1b, so it was tested with all mutant tetramers prepared in mock and PG loading conditions. This tetramer-based, biophysical approach was chosen because it emphasized clonal TCR binding to defined combinations of CD1b and lipid antigen. The use of staining instead of cytokine release eliminated clone to clone differences in activation outcomes downstream of TCR ligation. Further, as compared to cellular assays, this approach minimized the effects of antigen presenting cell-derived lipid antigens or differential processing of PG or GMM antigens. Analysis of staining of LDN5 and GEM42 by CD1b-GMM tetramers (Fig. 6A) and PG10 and BC24B by CD1b-PG tetramers (Fig. 6B) illustrate key outcomes, which when combined with results from the additional four staining patterns (Supplemental Fig. 3A–D), generated a matrix of staining results (Fig. 6C).
Figure 6: The effect of mutations in CD1b on interaction with T cells.
(A) T cell clones LDN5 and GEM42 were tested for binding of GMM-loaded CD1b tetramers with the indicated point mutations. (B) T cell clones PG10 and BC24B were tested for binding of PG-loaded CD1b tetramers with the indicated point mutations. (C) In addition, four additional T cell clone-CD1b-antigen combinations were tested (Supplemental Fig. 3A–D) and percentages of tetramer positive cells or MFI (Supplemental Fig. 3E) were Z score normalized per cell line and shown as a heatmap. The negative control (“neg”) is mock-loaded wild type CD1a tetramer and the positive control (“pos”) is the indicated antigen-loaded wild type tetramer.
No tetramer stained every T cell clone, and in no case did CD1a tetramers stain T cells. These two findings largely rule out non-specific interaction of tetramers with lymphocytes. Every CD1b tetramer-antigen combination stained at least one T cell line, demonstrating that no mutant tetramer failed to fold or load lipid. In all cases, dependence on the glycolipid or phospholipid matched the known antigen specificity of the clone, consistent with the interactions being mediated by TCRs interacting with combinations of CD1b and antigen. As measured by the percentage of cells staining above background (Fig. 6C) or mean fluorescence intensity (Supplemental Fig. 3E), the matrix staining conditions demonstrated clear, CD1b position-dependent changes in staining and linked these to the antigen and TCR type used.
Except for the K61A mutation, which had little effect compared to wild type CD1b, most mutations had a different effect on tetramer binding, depending on which T cell line was tested. Thus, TCR interactions with CD1b were not precisely conserved, especially for the diverse TRBV4–1− TCRs (Fig. 6C). However, there was a striking commonality among all four TRBV4–1+ T cells tested: mutation of the E80A on CD1b abolished recognition. This outcome was true regardless of whether GMM or PG was carried. The E80A CD1b tetramers were not non-functional in a general way based on positive results with other clones, and both PG and GMM were loaded, as assessed by bright staining BC24B and GEM42 (Figs. 6A, C). These patterns demonstrate that E80 in CD1b is an important residue for interaction with TRBV4–1 encoded TCRs, establishing a functional hot spot for TRBV4–1+ TCRs that acts independently of the antigen loaded.
Discussion
T cells recognizing monomorphic antigen presenting molecules represent an important exception to the general idea that specific TCR sequences cannot be linked to the molecular identity of the antigenic targets recognized. Through study of polyclonal T cells from genetically unrelated donors, our new data show that CD1b tetramer+ cells are enriched among TRBV4–1+ cells in vitro and ex vivo. Conversely, among CD1b-specific clones and polyclonal CD1b tetramer+ T cells, TCR sequencing demonstrated markedly increased use of TRBV4–1. Thus, bidirectional evidence supports linkage between TRBV4–1 expression and CD1b recognition. CD1b-restricted T cells expressing TRBV4–1+ TCRs represent a population of which LDN5-like T cells form a subset that is defined by recognition of GMM.
MAIT cells, type I NKT cells and GEM T cells express TCRs with invariant α chains and mediate highly specific responses to 5-OPRU, α-GalCer and GMM, respectively. CD1b-specific T cells described here differ from the known TCR-defined T cell types in several ways. First, TCR sequence motifs are found in the TCR β not the TCR α chains. Second, the TCRs show a lower degree (type 1 bias) of conservation that is restricted to the TRBV4–1 gene segment, rather than multiple segments or N-region additions. Thirdly, and most surprisingly, even though each individual TRBV4–1+ TCR that we studied here is specific for a lipid antigen, the TRBV4–1 motif biases towards CD1b recognition without regard to the antigen carried. A recent study independently established a link between TRBV4–1 and CD1c recognition (28). The broad antigen response pattern for TRBV4–1+ T cells is, to our knowledge, unprecedented in the sense that one germline encoded part of a TCR acts without regard to the antigen carried and likely reacts across two types of antigen presenting molecules. For MAIT cells, NKT cells and GEM T cells, the recognition mechanism is straightforward and well proven: the particular residues encoded by TRAV and TRAJ regions of each of these invariant TCR α chains make extensive physical contact with MR1, CD1d or CD1b and protruding epitopes on the carried lipid or metabolite antigens (14, 52–54). Our data show that the shared TCR features are found in the germline residues of TRBV4–1, but not in N-region or TRBJ residues that dominate the CDR3 loop. Therefore, a straightforward structural explanation for these functional responses would be that residues encoded by TRBV4–1 recognize some shared epitope on CD1b and CD1c that does not involve the carried lipid.
While direct proof of the molecular mechanism requires ternary TRBV4–1 TCR-CD1b crystal structures, much existing evidence points toward specific roles of TRBV4–1 sequences in TCR stabilization and binding to anionic surfaces on CD1b and CD1c. First, TCR crystal structures of clone 2 and PG10, as well as mutational scanning, pinpointed roles of two TRBV4–1 encoded residues, H29β and R30β. H29, which forms a hydrogen bond between the CDR1 and CDR3 loops, is located distally from the TCR surface so not expected to directly contact CD1b or antigen, but could stabilize the general internal structure of TCRβ. However, this intrachain, interloop interaction is common among non-TRBV4–1 TCRs, so does not likely account for their CD1-biasing nature. Instead, the positively charged residue R30β, which is found only in TRBV4–1, TRBV5–1, and TRBV10–3, is well positioned to contact CD1b. Also published mutational mapping independently implicates R30 of TRBV5–1 as crucial for TCR interaction with CD1b (55) and R30β of TRBV4–1 is crucial for the interaction with CD1c (28).
On the CD1 side, our mutational mapping for CD1b implicated E80, an anionic residue as essential for binding of for all five TRBV4–1+ TCRs tested here. E80 is necessary for a TRBV5–1+ and one other TRBV4–1+ CD1b-specific TCR response (40). E80 is also found in CD1c, but not in CD1a and CD1d, so its presence correlates with the two isoforms recognized by TRBV4–1 TCRs (56). Although no TRBV4–1 TCR has been solved in contact with CD1b, most αβ TCR-lipid-CD1 structures and all three ternary CD1b structures solved to date, show that CDR1β and CDR2β loops that carry R30 are positioned near the right margin of the CD1 platform, where E80 is also located (14, 34, 46). Overall, these data support a potential scenario in which the conserved positively charged residue R30 could bind CD1b near the negatively charged E80, although other anionic residues are present on the CD1b surface. Finally, CD1B and CD1C genes encoding E80 are widely present among mammals (57–63). No CD1B or CD1C (64) orthologs are present in mice, and mice lack a TRBV4–1 ortholog (65, 66). Therefore, analogous to the simultaneous presence or absence of MR1 and TRAV1–2 orthologs among mammals (17), a comparable co-evolutionary relationship might exist between TRBV4–1 and CD1B or CD1C orthologs. These data, and the availability of human tetramers made from non-polymorphic antigen presenting molecules, now provide new avenues to discovering hidden TCR patterns in the complex human TCR repertoire.
Supplementary Material
Key points.
TCR β variable gene 4–1 (TRBV4–1) is overrepresented among CD1b-specific T cells
The association between TRBV4–1 usage and CD1b is independent of lipid antigen
CD1b residue E80 is essential for binding of TRBV4–1-utilizing TCRs
Acknowledgments
This work was supported by the Australian Research Council and National Health and Medical Research Council, the National Institutes of Health (AI049313, AR048632, AI111224), and Netherlands Organization for Scientific Research (NWO). SG is a NHMRC Senior Research Fellow. JR is supported by an ARC Laureate Fellowship.
Footnotes
Competing interests
The authors declare that they have no competing interests.
References
- 1.Salio M, Silk JD, Jones EY, and Cerundolo V. 2014. Biology of CD1- and MR1-restricted T cells. Annu. Rev. Immunol 32:323–366. [DOI] [PubMed] [Google Scholar]
- 2.Han M, Hannick LI, DiBrino M, and Robinson MA. 1999. Polymorphism of human CD1 genes. Tissue Antigens 54:122–127. [DOI] [PubMed] [Google Scholar]
- 3.Van Rhijn I, and Moody DB. 2015. Donor Unrestricted T Cells: A Shared Human T Cell Response. J. Immunol 195:1927–1932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Rossjohn J, Gras S, Miles JJ, Turner SJ, Godfrey DI, and McCluskey J. 2015. T cell antigen receptor recognition of antigen-presenting molecules. Annu. Rev. Immunol 33:169–200. [DOI] [PubMed] [Google Scholar]
- 5.Garcia KC, Degano M, Stanfield RL, Brunmark A, Jackson MR, Peterson PA, Teyton L, and Wilson IA. 1996. An alphabeta T cell receptor structure at 2.5 A and its orientation in the TCR-MHC complex. Science 274:209–219. [DOI] [PubMed] [Google Scholar]
- 6.Garboczi DN, Utz U, Ghosh P, Seth A, Kim J, VanTienhoven EA, Biddison WE, and Wiley DC. 1996. Assembly, specific binding, and crystallization of a human TCR-alphabeta with an antigenic Tax peptide from human T lymphotropic virus type 1 and the class I MHC molecule HLA-A2. J. Immunol 157:5403–5410. [PubMed] [Google Scholar]
- 7.La Gruta NL, Gras S, Daley SR, Thomas PG, and Rossjohn J. 2018. Understanding the drivers of MHC restriction of T cell receptors. Nat. Rev. Immunol 18:467–478. [DOI] [PubMed] [Google Scholar]
- 8.Gras S, Chadderton J, Del Campo CM, Farenc C, Wiede F, Josephs TM, Sng XYX, Mirams M, Watson KA, Tiganis T, Quinn KM, Rossjohn J, and La Gruta NL. 2016. Reversed T Cell Receptor Docking on a Major Histocompatibility Class I Complex Limits Involvement in the Immune Response. Immunity 45:749–760. [DOI] [PubMed] [Google Scholar]
- 9.Beringer DX, Kleijwegt FS, Wiede F, van der Slik AR, Loh KL, Petersen J, Dudek NL, Duinkerken G, Laban S, Joosten A, Vivian JP, Chen Z, Uldrich AP, Godfrey DI, McCluskey J, Price DA, Radford KJ, Purcell AW, Nikolic T, Reid HH, Tiganis T, Roep BO, and Rossjohn J. 2015. T cell receptor reversed polarity recognition of a self-antigen major histocompatibility complex. Nat. Immunol 16:1153–1161. [DOI] [PubMed] [Google Scholar]
- 10.Sharon E, Sibener LV, Battle A, Fraser HB, Garcia KC, and Pritchard JK. 2016. Genetic variation in MHC proteins is associated with T cell receptor expression biases. Nat. Genet 48:995–1002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Dash P, Fiore-Gartland AJ, Hertz T, Wang GC, Sharma S, Souquette A, Crawford JC, Clemens EB, Nguyen THO, Kedzierska K, La Gruta NL, Bradley P, and Thomas PG. 2017. Quantifiable predictive features define epitope-specific T cell receptor repertoires. Nature 547:89–93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Glanville J, Huang H, Nau A, Hatton O, Wagar LE, Rubelt F, Ji X, Han A, Krams SM, Pettus C, Haas N, Arlehamn CSL, Sette A, Boyd SD, Scriba TJ, Martinez OM, and Davis MM. 2017. Identifying specificity groups in the T cell receptor repertoire. Nature 547:94–98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Robinson J, Halliwell JA, Hayhurst JD, Flicek P, Parham P, and Marsh SG. 2015. The IPD and IMGT/HLA database: allele variant databases. Nucleic Acids Res 43:D423–431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gras S, Van Rhijn I, Shahine A, Cheng TY, Bhati M, Tan LL, Halim H, Tuttle KD, Gapin L, Le Nours J, Moody DB, and Rossjohn J. 2016. T cell receptor recognition of CD1b presenting a mycobacterial glycolipid. Nat Commun 7:13257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Van Rhijn I, Kasmar A, de Jong A, Gras S, Bhati M, Doorenspleet ME, de Vries N, Godfrey DI, Altman JD, de Jager W, Rossjohn J, and Moody DB. 2013. A conserved human T cell population targets mycobacterial antigens presented by CD1b. Nat. Immunol 14:706–713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Corbett AJ, Eckle SB, Birkinshaw RW, Liu L, Patel O, Mahony J, Chen Z, Reantragoon R, Meehan B, Cao H, Williamson NA, Strugnell RA, Van Sinderen D, Mak JY, Fairlie DP, Kjer-Nielsen L, Rossjohn J, and McCluskey J. 2014. T-cell activation by transitory neo-antigens derived from distinct microbial pathways. Nature 509:361–365. [DOI] [PubMed] [Google Scholar]
- 17.Boudinot P, Mondot S, Jouneau L, Teyton L, Lefranc MP, and Lantz O. 2016. Restricting nonclassical MHC genes coevolve with TRAV genes used by innate-like T cells in mammals. Proc. Natl. Acad. Sci. U. S. A [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Turner SJ, Doherty PC, McCluskey J, and Rossjohn J. 2006. Structural determinants of T-cell receptor bias in immunity. Nat. Rev. Immunol 6:883–894. [DOI] [PubMed] [Google Scholar]
- 19.Grant EP, Degano M, Rosat JP, Stenger S, Modlin RL, Wilson IA, Porcelli SA, and Brenner MB. 1999. Molecular recognition of lipid antigens by T cell receptors. J. Exp. Med 189:195–205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cardell S, Tangri S, Chan S, Kronenberg M, Benoist C, and Mathis D. 1995. CD1-restricted CD4+ T cells in major histocompatibility complex class II-deficient mice. J. Exp. Med 182:993–1004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Behar SM, Podrebarac TA, Roy CJ, Wang CR, and Brenner MB. 1999. Diverse TCRs recognize murine CD1. J. Immunol 162:161–167. [PubMed] [Google Scholar]
- 22.Makowska A, Kawano T, Taniguchi M, and Cardell S. 2000. Differences in the ligand specificity between CD1d-restricted T cells with limited and diverse T-cell receptor repertoire. Scand.J.Immunol 52:71–79. [DOI] [PubMed] [Google Scholar]
- 23.Jahng A, Maricic I, Aguilera C, Cardell S, Halder RC, and Kumar V. 2004. Prevention of autoimmunity by targeting a distinct, noninvariant CD1d-reactive T cell population reactive to sulfatide. J. Exp. Med:947–957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Van Rhijn I, Young DC, Im JS, Levery SB, Illarionov PA, Besra GS, Porcelli SA, Gumperz J, Cheng TY, and Moody DB. 2004. CD1d-restricted T cell activation by nonlipidic small molecules. Proc. Natl. Acad. Sci. U. S. A 101:13578–13583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Chiu YH, Jayawardena J, Weiss A, Lee D, Park SH, Dautry-Varsat A, and Bendelac A. 1999. Distinct subsets of CD1d-restricted T cells recognize self-antigens loaded in different cellular compartments. J. Exp. Med 189:103–110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Moody DB, Reinhold BB, Guy MR, Beckman EM, Frederique DE, Furlong ST, Ye S, Reinhold VN, Sieling PA, Modlin RL, Besra GS, and Porcelli SA. 1997. Structural requirements for glycolipid antigen recognition by CD1b-restricted T cells. Science 278:283–286. [DOI] [PubMed] [Google Scholar]
- 27.Van Rhijn I, Gherardin NA, Kasmar A, de Jager W, Pellicci DG, Kostenko L, Tan LL, Bhati M, Gras S, Godfrey DI, Rossjohn J, and Moody DB. 2014. TCR Bias and Affinity Define Two Compartments of the CD1b-Glycolipid-Specific T Cell Repertoire. J. Immunol 193:5338–5344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Guo T, Koo MY, Kagoya Y, Anczurowski M, Wang CH, Saso K, Butler MO, and Hirano N. 2018. A Subset of Human Autoreactive CD1c-Restricted T Cells Preferentially Expresses TRBV4–1(+) TCRs. J. Immunol 200:500–511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Aragao D, Aishima J, Cherukuvada H, Clarken R, Clift M, Cowieson NP, Ericsson DJ, Gee CL, Macedo S, Mudie N, Panjikar S, Price JR, Riboldi-Tunnicliffe A, Rostan R, Williamson R, and Caradoc-Davies TT. 2018. MX2: a high-flux undulator microfocus beamline serving both the chemical and macromolecular crystallography communities at the Australian Synchrotron. J Synchrotron Radiat 25:885–891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AG, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, and Wilson KS. 2011. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr 67:235–242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Adams PD, Afonine PV, Bunkoczi G, Chen VB, Davis IW, Echols N, Headd JJ, Hung LW, Kapral GJ, Grosse-Kunstleve RW, McCoy AJ, Moriarty NW, Oeffner R, Read RJ, Richardson DC, Richardson JS, Terwilliger TC, and Zwart PH. 2010. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr 66:213–221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Emsley P, Lohkamp B, Scott WG, and Cowtan K. 2010. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr 66:486–501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Van Rhijn I, van Berlo T, Hilmenyuk T, Cheng TY, Wolf BJ, Tatituri RV, Uldrich AP, Napolitani G, Cerundolo V, Altman JD, Willemsen P, Huang S, Rossjohn J, Besra GS, Brenner MB, Godfrey DI, and Moody DB. 2016. Human autoreactive T cells recognize CD1b and phospholipids. Proc. Natl. Acad. Sci. U. S. A 113:380–385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Shahine A, Reinink P, Reijneveld JF, Gras S, Holzheimer M, Cheng TY, Minnaard AJ, Altman JD, Lenz S, Prandi J, Kubler-Kielb J, Moody DB, Rossjohn J, and Van Rhijn I. 2019. A T-cell receptor escape channel allows broad T-cell response to CD1b and membrane phospholipids. Nat Commun 10:56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Wang GC, Dash P, McCullers JA, Doherty PC, and Thomas PG. 2012. T cell receptor alphabeta diversity inversely correlates with pathogen-specific antibody levels in human cytomegalovirus infection. Science translational medicine 4:128ra142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Layre E, Collmann A, Bastian M, Mariotti S, Czaplicki J, Prandi J, Mori L, Stenger S, De Libero G, Puzo G, and Gilleron M. 2009. Mycolic acids constitute a scaffold for mycobacterial lipid antigens stimulating CD1-restricted T cells. Chem. Biol 16:82–92. [DOI] [PubMed] [Google Scholar]
- 37.Beckman EM, Porcelli SA, Morita CT, Behar SM, Furlong ST, and Brenner MB. 1994. Recognition of a lipid antigen by CD1-restricted alpha beta+ T cells. Nature 372:691–694. [DOI] [PubMed] [Google Scholar]
- 38.Stenger S, Mazzaccaro RJ, Uyemura K, Cho S, Barnes PF, Rosat JP, Sette A, Brenner MB, Porcelli SA, Bloom BR, and Modlin RL. 1997. Differential effects of cytolytic T cell subsets on intracellular infection. Science 276:1684–1687. [DOI] [PubMed] [Google Scholar]
- 39.Vincent MS, Leslie DS, Gumperz JE, Xiong X, Grant EP, and Brenner MB. 2002. CD1-dependent dendritic cell instruction. Nat. Immunol 3:1163–1168. [DOI] [PubMed] [Google Scholar]
- 40.Spada FM, Grant EP, Peters PJ, Sugita M, Melian A, Leslie DS, Lee HK, van Donselaar E, Hanson DA, Krensky AM, Majdic O, Porcelli SA, Morita CT, and Brenner MB. 2000. Self-recognition of CD1 by gamma/delta T cells: implications for innate immunity. J. Exp. Med. %20;191:937–948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.James CA, Yu KKQ, Gilleron M, Prandi J, Yedulla VR, Moleda ZZ, Diamanti E, Khan M, Aggarwal VK, Reijneveld JF, Reinink P, Lenz S, Emerson RO, Scriba TJ, Souter MNT, Godfrey DI, Pellicci DG, Moody DB, Minnaard AJ, Seshadri C, and Van Rhijn I. 2018. CD1b Tetramers Identify T Cells that Recognize Natural and Synthetic Diacylated Sulfoglycolipids from Mycobacterium tuberculosis. Cell chemical biology 25:392–402 e314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Van Rhijn I, Iwany SK, Fodran P, Cheng TY, Gapin L, Minnaard AJ, and Moody DB. 2017. CD1b-mycolic acid tetramers demonstrate T-cell fine specificity for mycobacterial lipid tails. Eur. J. Immunol 47:1525–1534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Emerson R, Sherwood A, Desmarais C, Malhotra S, Phippard D, and Robins H. 2013. Estimating the ratio of CD4+ to CD8+ T cells using high-throughput sequence data. J. Immunol. Methods 391:14–21. [DOI] [PubMed] [Google Scholar]
- 44.Freeman JD, Warren RL, Webb JR, Nelson BH, and Holt RA. 2009. Profiling the T-cell receptor beta-chain repertoire by massively parallel sequencing. Genome Res. 19:1817–1824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Warren RL, Freeman JD, Zeng T, Choe G, Munro S, Moore R, Webb JR, and Holt RA. 2011. Exhaustive T-cell repertoire sequencing of human peripheral blood samples reveals signatures of antigen selection and a directly measured repertoire size of at least 1 million clonotypes. Genome Res. 21:790–797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Shahine A, Van Rhijn I, Cheng TY, Iwany S, Gras S, Moody DB, and Rossjohn J. 2017. A molecular basis of human T cell receptor autoreactivity toward self-phospholipids. Science immunology 2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Garcia-Alles LF, Collmann A, Versluis C, Lindner B, Guiard J, Maveyraud L, Huc E, Im JS, Sansano S, Brando T, Julien S, Prandi J, Gilleron M, Porcelli SA, de la Salle H, Heck AJ, Mori L, Puzo G, Mourey L, and De Libero G. 2011. Structural reorganization of the antigen-binding groove of human CD1b for presentation of mycobacterial sulfoglycolipids. Proc. Natl. Acad. Sci. U. S. A 108:17755–17760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.DeWitt WS, Yu KKQ, Wilburn DB, Sherwood A, Vignali M, Day CL, Scriba TJ, Robins HS, Swanson WJ, Emerson RO, Bradley PH, and Seshadri C. 2018. A Diverse Lipid Antigen-Specific TCR Repertoire Is Clonally Expanded during Active Tuberculosis. J. Immunol 201:888–896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Moody DB, Guy MR, Grant E, Cheng TY, Brenner MB, Besra GS, and Porcelli SA. 2000. CD1b-mediated T cell recognition of a glycolipid antigen generated from mycobacterial lipid and host carbohydrate during infection. J. Exp. Med 192:965–976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Gadola SD, Zaccai NR, Harlos K, Shepherd D, Castro-Palomino JC, Ritter G, Schmidt RR, Jones EY, and Cerundolo V. 2002. Structure of human CD1b with bound ligands at 2.3 A, a maze for alkyl chains. Nat.Immunol 3:721–726. [DOI] [PubMed] [Google Scholar]
- 51.Relloso M, Cheng TY, Im JS, Parisini E, Roura-Mir C, DeBono C, Zajonc DM, Murga LF, Ondrechen MJ, Wilson IA, Porcelli SA, and Moody DB. 2008. pH-dependent interdomain tethers of CD1b regulate its antigen capture. Immunity 28:774–786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Patel O, Kjer-Nielsen L, Le Nours J, Eckle SB, Birkinshaw R, Beddoe T, Corbett AJ, Liu L, Miles JJ, Meehan B, Reantragoon R, Sandoval-Romero ML, Sullivan LC, Brooks AG, Chen Z, Fairlie DP, McCluskey J, and Rossjohn J. 2013. Recognition of vitamin B metabolites by mucosal-associated invariant T cells. Nat Commun 4:2142. [DOI] [PubMed] [Google Scholar]
- 53.Lopez-Sagaseta J, Dulberger CL, Crooks JE, Parks CD, Luoma AM, McFedries A, Van Rhijn I, Saghatelian A, and Adams EJ. 2013. The molecular basis for Mucosal-Associated Invariant T cell recognition of MR1 proteins. Proc. Natl. Acad. Sci. U. S. A 110:E1771–1778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Borg NA, Wun KS, Kjer-Nielsen L, Wilce MC, Pellicci DG, Koh R, Besra GS, Bharadwaj M, Godfrey DI, McCluskey J, and Rossjohn J. 2007. CD1d-lipid-antigen recognition by the semi-invariant NKT T-cell receptor. Nature 448:44–49. [DOI] [PubMed] [Google Scholar]
- 55.Grant EP, Beckman EM, Behar SM, Degano M, Frederique D, Besra GS, Wilson IA, Porcelli SA, Furlong ST, and Brenner MB. 2002. Fine specificity of TCR complementarity-determining region residues and lipid antigen hydrophilic moieties in the recognition of a CD1-lipid complex. J. Immunol 168:3933–3940. [DOI] [PubMed] [Google Scholar]
- 56.Wun KS, Reijneveld JF, Cheng TY, Ladell K, Uldrich AP, Le Nours J, Miners KL, McLaren JE, Grant EJ, Haigh OL, Watkins TS, Suliman S, Iwany S, Jimenez J, Calderon R, Tamara KL, Leon SR, Murray MB, Mayfield JA, Altman JD, Purcell AW, Miles JJ, Godfrey DI, Gras S, Price DA, Van Rhijn I, Moody DB, and Rossjohn J. 2018. T cell autoreactivity directed toward CD1c itself rather than toward carried self lipids. Nat. Immunol 19:397–406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Dascher CC, Hiromatsu K, Naylor JW, Brauer PP, Brown KA, Storey JR, Behar SM, Kawasaki ES, Porcelli SA, Brenner MB, and LeClair KP. 1999. Conservation of a CD1 multigene family in the guinea pig. J. Immunol 163:5478–5488. [PubMed] [Google Scholar]
- 58.Dossa RG, Alperin DC, Hines MT, and Hines SA. 2014. The equine CD1 gene family is the largest and most diverse yet identified. Immunogenetics 66:33–42. [DOI] [PubMed] [Google Scholar]
- 59.Eguchi-Ogawa T, Morozumi T, Tanaka M, Shinkai H, Okumura N, Suzuki K, Awata T, and Uenishi H. 2007. Analysis of the genomic structure of the porcine CD1 gene cluster. Genomics 89:248–261. [DOI] [PubMed] [Google Scholar]
- 60.Hayes SM, and Knight KL. 2001. Group 1 CD1 genes in rabbit. J. Immunol 166:403–410. [DOI] [PubMed] [Google Scholar]
- 61.Looringh van Beeck FA, Zajonc DM, Moore PF, Schlotter YM, Broere F, Rutten VP, Willemse T, and Van Rhijn I. 2008. Two canine CD1a proteins are differentially expressed in skin. Immunogenetics 60:315–324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Reinink P, and Van Rhijn I. 2016. Mammalian CD1 and MR1 genes. Immunogenetics 68:515–523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Van Rhijn I, Koets AP, Im JS, Piebes D, Reddington F, Besra GS, Porcelli SA, van Eden W, and Rutten VP. 2006. The bovine CD1 family contains group 1 CD1 proteins, but no functional CD1d. J. Immunol 176:4888–4893. [DOI] [PubMed] [Google Scholar]
- 64.Bradbury A, Belt KT, Neri TM, Milstein C, and Calabi F. 1988. Mouse CD1 is distinct from and co-exists with TL in the same thymus. EMBO J. 7:3081–3086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Bosc N, and Lefranc MP. 2000. The mouse (Mus musculus) T cell receptor beta variable (TRBV), diversity (TRBD) and joining (TRBJ) genes. Exp. Clin. Immunogenet 17:216–228. [DOI] [PubMed] [Google Scholar]
- 66.Koop BF, and Hood L. 1994. Striking sequence similarity over almost 100 kilobases of human and mouse T-cell receptor DNA. Nat. Genet 7:48–53. [DOI] [PubMed] [Google Scholar]
- 67.Larrouy-Maumus G, Layre E, Clark S, Prandi J, Rayner E, Lepore M, de Libero G, Williams A, Puzo G, and Gilleron M. 2017. Protective efficacy of a lipid antigen vaccine in a guinea pig model of tuberculosis. Vaccine 35:1395–1402. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.