Abstract
The growing concern with the environment is making important for livestock producers to focus on selection for efficiency-related traits, which is a challenge for commercial cattle herds due to the lack of pedigree information. To explore a cost-effective opportunity for genomic evaluations of commercial herds, this study compared the accuracy of bulls’ genomic estimated breeding values (GEBV) using different pooled genotype strategies. We used ten replicates of previously simulated genomic and phenotypic data for one low (t1) and one moderate (t2) heritability trait of 200 sires and 2,200 progeny. Sire’s GEBV were calculated using a univariate mixed model, with a hybrid genomic relationship matrix (h-GRM) relating sires to: 1) 1,100 pools of 2 animals; 2) 440 pools of 5 animals; 3) 220 pools of 10 animals; 4) 110 pools of 20 animals; 5) 88 pools of 25 animals; 6) 44 pools of 50 animals; and 7) 22 pools of 100 animals. Pooling criteria were: at random, grouped sorting by t1, grouped sorting by t2, and grouped sorting by a combination of t1 and t2. The same criteria were used to select 110, 220, 440, and 1,100 individual genotypes for GEBV calculation to compare GEBV accuracy using the same number of individual genotypes and pools. Although the best accuracy was achieved for a given trait when pools were grouped based on that same trait (t1: 0.50–0.56, t2: 0.66–0.77), pooling by one trait impacted negatively on the accuracy of GEBV for the other trait (t1: 0.25–0.46, t2: 0.29–0.71). Therefore, the combined measure may be a feasible alternative to use the same pools to calculate GEBVs for both traits (t1: 0.45–0.57, t2: 0.62–0.76). Pools of 10 individuals were identified as representing a good compromise between loss of accuracy (~10%–15%) and cost savings (~90%) from genotype assays. In addition, we demonstrated that in more than 90% of the simulations, pools present higher sires’ GEBV accuracy than individual genotypes when the number of genotype assays is limited (i.e., 110 or 220) and animals are assigned to pools based on phenotype. Pools assigned at random presented the poorest results (t1: 0.07–0.45, t2: 0.14–0.70). In conclusion, pooling by phenotype is the best approach to implementing genomic evaluation using commercial herd data, particularly when pools of 10 individuals are evaluated. While combining phenotypes seems a promising strategy to allow more flexibility to the estimates made using pools, more studies are necessary in this regard.
Keywords: beef cattle, DNA pooling, genomic selection, hybrid genomic relationship matrix
Introduction
Farming practices around the world are changing due to concerns with the environmental impact of human activities and global demand for protein sources, presenting both opportunities and challenges for the livestock sector (Webb and Buratini, 2016). While beef cattle herds are responsible for 65% of livestock greenhouse gas emissions, there is considerable genetic variation in efficiency-related traits, such as feed efficiency, fertility, and adaptation to climate, which represents a significant opportunity to select superior animals to reduce the environmental impact without compromising productivity (Gerber et al., 2013; Hayes et al., 2013). The challenge is that, due to the extensive nature of the production system and the simultaneous use of multiple sires, particularly in tropical conditions, there is a lack of pedigree information so that the large number of phenotypic measures routinely collected in commercial supply chains are not accessible for breeding plans to achieve genetic improvement. The use of genetically superior herd bulls is often the only way to achieve genetic progress in commercial herds and strategies to enable the identification of those animals are crucial to assure that, given the opportunity, the next generation will perform to their genetic potential, toward a more sustainable production system.
DNA technology is being increasingly applied to the livestock breeding sector as a tool to estimate genomic breeding values (GEBV) with no requirement for pedigree information, and thus, has the potential to revolutionize the sector by informing management and selection decisions (Hayes et al., 2013). However, obtaining individual genotypes to assess sires’ performance at a commercial level still represents a substantial expense. For more than 30 years, human genetics has been using DNA pooling as a cost-effective strategy for screening molecular markers (Arnheim et al., 1985). More recently, new methodologies have been developed to explore this alternative for genomic evaluation of livestock species using single nucleotide polymorphism (SNP) assays (Henshall et al., 2012; Reverter et al., 2014). Reverter et al. (2016) proposed the use of a hybrid genomic relationship matrix to provide a link between pooled genotyping data from commercial cow herds and individual DNA genotypes from bulls in the nucleus herds, which was shown to be a useful genotyping strategy for cow fertility in tropical cattle. This strategy has also been shown to be suitable for dag score in sheep (Bell et al., 2017), demonstrating its viability for the selection for efficiency-related traits that are often expensive to measure accurately. Although a promising approach to inform sire’s GEBV using information from commercial herds, pooling strategies imply a loss of prediction accuracy. Therefore, assessing the impact of different pooling schemes on prediction accuracy while considering genotyping costs is fundamental to support the implementation of such strategies at a commercial level.
The objective of this study was to compare the accuracy of sires’ GEBV based on different pool sizes and pooling criteria of progeny genotypes, for a lowly and a moderately heritable trait, using simulated data.
Materials and Methods
Simulated Data
The data used to perform the analysis proposed in this study was previously simulated by Karaman et al. (2018). Briefly, genotype data for the first five bovine chromosomes (11,154 SNPs) was simulated for one generation (G1), based on 50K haplotype information of 2,200 Danish Holstein animals (G0). Data were simulated across 10 random replicates and for each one, 200 and 2,000 animals of G0 were randomly chosen as male and females, respectively. Animals from G0 were randomly mated at a rate of 1:10 and one mating was replicated for each sire, resulting in 11 progeny per sire to retain the population size at 2,200 for G1. Recombination on each chromosome was simulated as previously described (Karaman et al., 2018) and mutation was not considered. Phenotypic values were simulated to represent a low (0.1) and a moderate (0.4) heritability trait (referred as trait 1 and trait 2, respectively), with a genetic correlation of 0.48 considering all animals. For each replicate, 200 SNPs were randomly assigned to QTL and they were kept in the final data set of SNPs. This does not influence the comparison between the relative merit of each approach tested in this study. Genotypes and phenotypes for both traits were simulated for all animals over 10 replicates, and all reported results were averaged between those replicates.
Estimation of Genomic Breeding Values
Several approaches were used regarding progeny genotypic data (G1) used to calculate GEBV for the 200 sires of G0 (Figure 1) and they were all based on the following methodology, with slight modifications to accommodate pooled genotypes. Firstly, all individual genotypes of the 200 sires and 2,200 progeny were used to generate a genomic relationship matrix (GRM) using the first method of VanRaden (2008). For calculation of GEBV, the following univariate mixed model was used:
Figure 1.
Experimental design. Three dots indicate analyses were performed as described immediately above.
| (1) |
where y is the vector of phenotypes (trait 1 or trait 2), 1 is a vector of ones, µ is the overall mean, Z is the incidence matrix relating random additive effects in u with observations in y, and e is the vector of random residual effects. Genomic best linear unbiased prediction (GBLUP) was performed in Qxpak5 (Pérez-Enciso and Misztal, 2011) to obtain sires’ GEBV from the solutions to additive effects in u. It was assumed that , and , where and are the additive genetic and residual variances, respectively, and G is the GRM. The accuracy of sires’ GEBVs were calculated as the correlation between GEBV and simulated true breeding values (BV) for each sire. As an alternative, sires with the top 100 GEBV were compared to sires with the top 100 true BV (henceforward referred to as TOP100), to estimate the proportion of concordance between the two ranks. This analysis was performed to generate reference values for comparison to pooled genotyping strategies.
Pooled Genotyping Strategies
Seven different pool sizes were tested, in which data for all 2,200 progeny were combined into pools as follows: 1) 1,100 pools of 2 animals (PS2); 2) 440 pools of 5 animals (PS5); 3) 220 pools of 10 animals (PS10); 4) 110 pools of 20 animals (PS20); 5) 88 pools of 25 animals (PS25); 6) 44 pools of 50 animals (PS50); and 7) 22 pools of 100 animals (PS100). Animals were assigned to pools based on four pooling criteria: at random, grouped sorting by trait 1, grouped sorting by trait 2, and grouped sorting by a combination of trait 1 and 2 (henceforward referred as “combo”) which consisted of the mean of trait 1 Z-score plus trait 2 Z-score for all the animals (Figure 1). Phenotypes for each pool were calculated as the average phenotype of animals in that pool.
To combine individual genotypes into pools of different sizes, the frequency of B allele was computed for all SNPs in each pool. Then, genotypes were determined similarly to the scheme proposed by Reverter et al. (2016), except that instead of defining a hard frequency threshold to assign AA (0), AB (1) or BB (2) genotypes, pooled genotypes were defined as follows: 1) if the B-allele frequency ≤0.17, then SNP genotype = 0; 2) if the B-allele frequency >0.25 and ≤0.75, then SNP genotype = 1; 3) if the B-allele frequency >0.82, then SNP genotype = 2; 4) if the B-allele frequency >0.17 and ≤0.25 or >0.75 and ≤0.82, then a “flipping coin” function assigned SNP genotype to 0 or 1 and 1 or 2, respectively. Based on our experience with genotypic data from different species, we believe this methodology reflects better what occurs in reality than using the hard thresholds originally proposed.
Considering for each analysis all pools were the same size (i.e., had the same number of animals represented), the hybrid GRM (h-GRM) proposed by Reverter et al. (2016) relating all 200 individually genotyped sires and different numbers of pools were computed in a normal fashion using the first method of VanRaden (2008), so that three blocks were created: one relating only sires (individual genotypes), one relating only progeny (pools) and one relating sires to progeny (individual genotypes to pools) (Figure 2). Sires’ GEBV, accuracy and TOP100 were also calculated as described for individually genotyped progeny, except that the entire h-GRM was used as relationship matrix, and in equation 1, y now represents the vector of phenotypes for pools. Analyses performed with pools assigned at random were run 100 times to account for the expected variation in GEBV since each random run assembled different pools of animals. For the analysis of pools assigned by phenotype, although for each run the same animals are always grouped together, the flipping coin function used to generate the pooled genotype introduces a variation to GEBV, therefore, those analyses were also performed 100 times. Reported results were calculated by first computing the average within runs and then the average across the 10 replicates. The retained accuracy (%) was calculated by comparing the accuracy obtained for each pool size and pooling criteria to the maximum possible accuracy for each trait, estimated using individual genotypes.
Figure 2.
Heatmap of a hybrid relationship matrix of 200 individually genotyped sires and 220 pools of 10 animals each. Highlighted the difference between the three blocks relating only sires (individual genotypes), only progeny (pools) and sires to progeny (individual genotypes to pools). Black color represents values close to zero and intensity of red color increase as values increase.
Finally, we asked the question: given a limited number of possible genotyping assays, would it be better to genotype individual animals or pools? We then repeated the estimations for the 200 sires using the same number of individually genotyped progeny as the numbers of pools we previously tested, that is, 110, 220, 440, and 1,100. Animals were selected at random or as extremes of trait 1, trait 2, or combo (Figure 1). Analyses considering randomly selected animals were performed 100 times and averaged as previously described. To access the significance of the results we computed how many times, out of 100, pools performed better than individual genotypes according to the accuracy of GEBV.
Results and Discussion
The results of accuracies and TOP100 GEBV of trait 1 (low heritability) and trait 2 (moderate heritability) for the six pool sizes (PS2, PS5, PS10, PS20, PS25, PS50, PS100) and four pooling criteria (at random, sorted by trait 1, sorted by trait 2, and sorted by combo) analyzed are summarized in Figure 3 and Supplementary Table 1. Notably, TOP100 behaves very similarly to the accuracy, as both parameters measure the ability of predictions to reflect reality. The TOP100 measure is not meant to indicate the top sires for selection, in which case the top 10% would be more appropriate. Here, as a measure of accuracy, the TOP100 is better to sample the variation around expectation while keeping the same proportions of results compared to the top 10%.
Figure 3.
Mean accuracy of sires’ genomic estimated breeding value (GEBV) (A) and number of sires captured as top 100 GEBV that are also top 100 true breeding value (B) for a lowly (trait 1) and a moderately (trait 2) heritable trait, using pooled genotypes with different numbers of progeny chosen using four criteria: random (byRandom), grouped sorting by trait 1 (byT1), grouped sorting by trait 2 (byT2) or grouped sorting by a combination of traits (byCombo). Dots represent the average of 100 analyses within each pool size, pooling criteria and phenotype for each of the 10 replicates, except for pool size 1 that represents results when using individually genotyped progeny.
The best result for a given trait was achieved when pools were grouped based on that same trait. Indeed, pooling animals by phenotype has been suggested as the design of choice (Henshall et al., 2012) and a variety of phenotypes have been used to pool genotypes of sheep, beef and dairy cattle, and aquaculture species for genomic evaluations (Sonesson et al., 2010; Strillacci et al., 2014; Keele et al., 2015; Reverter et al., 2016; Bell et al., 2017). However, when considering two traits, our data shows that pooling by one trait impacted negatively on the accuracy of GEBV’s for the second trait, even when the latter trait is moderately heritable. This negative impact was not as detrimental as was observed when we assigned animals to pools at random, probably because of the positive genetic correlation that exists between the two traits. If there was no correlation between the traits, one can speculate accuracies for the second trait would be closer to those observed with randomly assigned pools. To the best of our knowledge, no study has addressed situations where pools are used to evaluate more than one trait simultaneously.
The combination of traits based on Z-scores turned out being a good strategy to estimate GEBV for both traits, particularly for smaller pools. The combination of phenotypes based on Z-scores as opposed to ranks preserves the spreading of the data and seems to be a good way to combine phenotypes, although different scenarios still need to be tested regarding how many phenotypes can be combined and how much their respective heritabilities influence the results. In general, the drop in accuracy seems to be more pronounced for pools based on phenotype until PS20 and then remains relatively constant. Still, when we compare accuracies between individually genotyped progeny (i.e., the best accuracy possible) and 22 pools of 100 animals (the most extreme scenario), trait 1 present an accuracy drop of 0.07 and 0.12 when progeny are grouped sorting by trait 1 and by combo, respectively, and trait 2 present an accuracy drop of 0.15 and 0.19 when progeny are grouped sorting by trait 2 and by combo, respectively. Considering that genotyping 22 pools of 100 animals (22 genotype assays) represent 1% the cost of genotyping 2,200 individual animals, that loss of accuracy seems relatively small. While larger pool sizes have the potential to dramatically decrease the cost of genomic evaluations, it remains necessary to consider the total number of pools required to achieve a satisfactory accuracy level, therefore, a balance must be drawn.
Figure 4 shows the increase in cost savings with genotype assays as pool size increases in comparison to the decrease in the accuracy retained (relative to the maximum possible accuracy) and their point of intersection. With PS10 it is possible to maintain close to 90% of the maximum accuracy for both traits and save 90% of the cost of genotyping all the progeny (i.e., 220 genotype assays needed instead of 2,200 for individual genotypes), which is a good compromise, especially considering the possibility of using a combined measure that enables genomic estimates for two traits using the same pools (Supplementary Table 2). Indeed, Dominik et al. (2018) observed increased accuracy and dollar response for domestic Australian Angus selection index using commercial records from DNA pooling, indicating pools of 10 progeny as a cost-effective strategy. Still, estimating the costs involved in generating genotyping data is not a straightforward task as, for instance, unit prices could decrease with the increase in total number of genotype assay. Here, to avoid the uncertainty of the real cost of individual genotype assays, we simply considered that by requiring half the number of genotype assays one would save 50% with genotyping cost, which in our view is a fair approximation. It is worth mentioning that our estimation of cost savings is only related to the genotype assay, the costs of collecting individual phenotypes, tissue samples, and extracting DNA would remain the same independently of genotyping individual animals or pools. Therefore, using routinely collected phenotypes and incorporating fast tissue sampling protocols to this routine would be the best way to make genomic analysis viable for beef cattle at commercial level.
Figure 4.
Comparison between the percentage of genotype cost savings and the percentage of sires’ genomic estimated breeding value (GEBV) accuracy retained (AR) by each pool size (relative to the accuracy using individual genotypes) for trait 1, when pools are grouped sorting by itself (AR_T1byT1) or by combo (AR_T1byCombo), and for trait 2, when pools are grouped sorting by itself (AR_T2byT2) or by combo (AR_T2byCombo). The dashed line indicates the pool size which the percentage of genotyping cost savings meet the GEBV accuracy retained.
The drop of accuracy as the pool size increases is more dramatic when animals are randomly assigned to pools (Figure 3). Random pooling is, in principle, a good strategy as it enables the use of the resulting pools for genomic predictions of any recorded phenotype, and has been applied before in aquaculture species for population studies (Johnston et al., 2013; Henshall et al., 2014). However, the randomness is responsible for a variation in GEBV accuracy relative to pool size (Figure 5). As pool size gets smaller, so does the variation in the accuracy of sires GEBV for randomly assigned pools, and the mean accuracy increases. For the moderately heritable trait (trait 2), small pool sizes (i.e., PS2 and PS5) present mean accuracy comparable to the accuracy of the lowly heritable trait (trait 1) when that trait is calculated based on individually genotyped progeny. Nevertheless, the decrease in accuracy of sires GEBV as the pool size increases is more evident in trait 2. While results demonstrate that pooling based on phenotype performs better than randomized pooling, differences in observed accuracy of GEBVs are highly influenced by the heritability of the trait. Randomly selected PS5 performs better for trait 2 (0.55) than for trait 1 when animals are assigned to pools based on the same trait (0.53, Supplementary Table 1).
Figure 5.
Accuracy of sires’ genomic estimated breeding value (GEBV) and number of sires captured as top 100 GEBV that are also top 100 true breeding value (n = 200) for a lowly (trait 1—A and B, respectively) and a moderately (trait 2—C and D, respectively) heritable trait, using pooled genotypes with different numbers of randomly chosen progeny (n = 2,200; PS + number of animals in each pool). Distributions represent 100 analyses within each pool size and phenotype for one of the 10 replicates. Dashed lines indicate means, the continuous red line indicates results when using individually genotyped progeny and the continuous gray line indicate the limit around which results are completely random.
In real situations and due to budget constraints, it is important to consider the affordability of the testing, potentially limiting the number of genotype assays which can be undertaken. Based on the comparison between the same number of individually genotyped samples and genotyped pools (Figure 6, Supplementary Table 3), Table 1 shows pools perform better than individual genotypes more than 90% of the time when the number of genotype assays is small (i.e., 110 or 220) and provided 1) GEBV for trait 1 are estimated using progeny pools grouped sorting by trait 1; 2) GEBV for trait 2 are estimated using progeny pools grouped sorting by trait 2; or 3) GEBV for traits 1 and 2 are estimated using progeny pools grouped sorting by combo. On the other hand, when it is possible to genotype a large number of samples (i.e., 440 or 1,100), pooled genotypes do not significantly increase the accuracy of predictions. Genotyping randomly selected individual genotypes versus pools chosen at random do not result in appreciable differences in the accuracy of predictions, in fact, in some cases individual genotypes seem to perform slightly better. Taken together, if there’s a reason for which animals need to be randomly selected for genotyping, individual genotypes are preferable to pools, independently of the number of individuals.
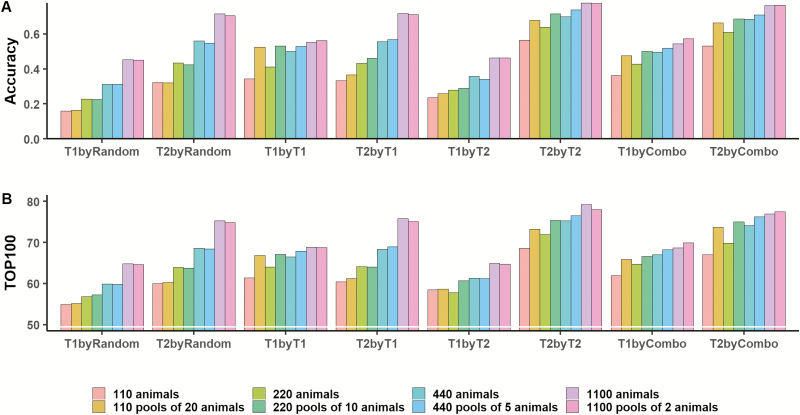
Figure 6.
Mean accuracy of sires’ genomic estimated breeding value (GEBV) (A) and number of sires captured as top 100 GEBV that are also top 100 true breeding value (B) for a lowly (trait 1) and a moderately (trait 2) heritable phenotype, using 110, 220, 440, or 1,100 individually genotyped progeny or all 2,200 progeny grouped in pools of 20, 10, 5, or 2 animals. Animals were assigned to pools using four criteria: random (byRandom), grouped sorting by trait 1 (byT1), grouped sorting by trait 2 (byT2) or grouped sorting by a combination of traits (byCombo). Individually genotyped animals were selected at random or as extremes of phenotypes. For pools, bars represent the average of 100 analysis within each trait, pooling criteria and phenotype for all 10 replicates. For individual animals, bars represent the average within replicates.
Table 1.
Number of times, out of 100 replicates, that pools lead to better accuracy of sires GEBV than individual genotypes
| Pooling criteria | Trait | 110 individual genotypes vs. 110 pools of 20 animals | 220 individual genotypes vs. 220 pools of 10 animals | 440 individual genotypes vs. 440 pools of 5 animals | 1,100 individual genotypes vs. 1,100 pools of 2 animals |
|---|---|---|---|---|---|
| Random | Trait 1 | 52.0 | 49.4 | 50.9 | 50.0 |
| Trait 2 | 50.1 | 45.5 | 42.4 | 38.7 | |
| Trait 1 | Trait 1 | 100.0 | 100.0 | 81.2 | 53.2 |
| Trait 2 | 50.7 | 63.5 | 60.0 | 45.5 | |
| Trait 2 | Trait 1 | 42.1 | 54.0 | 28.2 | 42.1 |
| Trait 2 | 99.6 | 100.0 | 89.5 | 41.5 | |
| Combo | Trait 1 | 95.2 | 98.8 | 63.8 | 77.3 |
| Trait 2 | 100.0 | 90.6 | 82.1 | 55.6 |
The same “pool vs. individual genotypes” comparison shows that when pooling is based on phenotypes, 110 pools of 20 animals each perform about the same as 220 individually genotyped progeny (Figure 6, Supplementary Table 3). Likewise, 220 pools of 10 animals perform about the same as 440 individually genotyped progeny. That implies not only pools perform better than individual genotypes when the number of genotype assay is limited, provided animals are assigned to pools based on phenotype, but also that pools perform better using half the number of genotypes and therefore, costs for genotyping are halved. However, this benefit would be somewhat offset by the persisting need to collect DNA samples from all individual animals for pooling, as previously discussed.
The better performance of pools when a limited number of genotypes are available can be explained by the number of sires represented by the progeny used for genomic evaluation (Table 2). Out of the 200 sires evaluated by all proposed strategies, only around 84 are represented when 110 individual genotypes are used, while when using 110 pools of 20 animals for evaluation all sires are represented, as a result of all progeny being represented within the pools (the average number of sires represented in each pool can be found in Supplementary Table 4). A similar scenario occurs when 220 individually genotyped progeny are evaluated, with around 136 of the 200 sires represented in comparison to all sires again being represented in the 220 pools of 10 animals. As the number of sires represented by individually genotyped progeny increases, there are smaller differences in the accuracy of GEBV in comparison to pools. This suggests that pooled genotyping strategies are particularly useful to enable genomic evaluation of bulls from commercial herds when resources are only available for genotyping a fraction of the herd. By increasing the number of animals in each pool, and therefore decreasing the number of necessary genotypes, resources could be made available to individually genotype young bulls which are sons of the progeny tested sires, so they could be included in GEBV analysis. This would enable early selection decisions to be made on those animals and consequent reduction of generation interval (Dominik et al., 2018). Likewise, there could be an opportunity of accessing GEBVs for females in nucleus herds, especially for complex and difficult-to-measure traits, such as feed efficiency or marbling score, which could contribute to the overall genetic progress. Still, depending on the size of the herd, genotyping all animals can be impractical even by DNA pooling. In that case, pooling only animals ranked as extremes for a trait or a combined trait could be an alternative, which has been successfully applied to a number of association studies and to genomic selection in marine fish (Yang et al., 2015; Dong et al., 2016; Bjørnland et al., 2018).
Table 2.
Number of sires represented considering individually genotyped progeny (the total number of sires is 200)
| Pooling criteria | 110 animals | 220 animals | 440 animals | 1,100 animals |
|---|---|---|---|---|
| Random | 86.38 | 137.04 | 182.75 | 199.91 |
| Trait 1 | 84.70 | 135.40 | 181.90 | 199.90 |
| Trait 2 | 82.60 | 135.20 | 182.30 | 199.70 |
| Combo | 83.70 | 135.20 | 183.00 | 199.90 |
Comparing pooling criteria with respect to the variation in the number of sires represented by the same number of individually genotyped progeny, our simulated data does not model the presence of particularly superior sires for the traits, otherwise there would be a significant decrease in the number of sires represented when animals are assigned to pools by phenotype in comparison to animals assigned randomly. That could be the case for real data when one, or a few, particularly outstanding bulls for a given trait would cause their progeny to be overrepresented in the GEBV analysis, leading to even lower accuracies when compared with pooled genotypes. It is also often the case that different sires have different number of progeny, which could lead to similar impact. This simulation study is limited in its ability to represent all the different scenarios encountered in real data, where population structure and the density of SNP assay are important features impacting the accuracy of predictions. Nevertheless, our results on simulated data are a great incentive in the quest of making high throughput technologies applicable at commercial level.
Conclusions
The comparison between different strategies for estimation of genomic breeding values of bulls using progeny pooled genotypes suggests that when limited resources are available for genotyping, which is generally the case for large commercial herds, the strategy of genotyping by pools is particularly appealing, especially considering the possibility to estimate breeding values of bulls that otherwise would not be evaluated. In practice, while these results highlight the significant potential for genetic gain at the commercial level using genomic pooling strategies which are cost-effective, it also represents a great challenge for breeding programs which are currently not set up to deal with this kind of data, requiring that new methodologies are developed. If the availability of samples with phenotypic information is not a constraint, then there is no doubt creating pools based on phenotypes leads to the best accuracy levels for sires’ GEBV. Pool sizes of around 10 individuals are suggested as a good compromise between loss of accuracy and cost savings. Indeed, the high accuracies obtained based on the combination of phenotypes presents as an opportunity to estimate breeding values for more than one trait using the same genotyped pools. More studies are necessary to better evaluate the combination of phenotypes in terms of numbers and heritabilities, and the cost-benefit for important traits, by comparing the economic benefits realized through genetic gains to the cost associated with testing using the pooled genotyping approach.
Supplementary Material
Footnotes
The authors declare no conflict of interest and thank Dr Brad Hine and Dr Yutao Li for reviewing the article. Additional information and data are available upon request.
Literature Cited
- Arnheim N., Strange C., and Erlich H.. . 1985. Use of pooled DNA samples to detect linkage disequilibrium of polymorphic restriction fragments and human disease: studies of the HLA class II loci. Proc. Natl Acad. Sci. U. S. A. 82:6970–6974. doi: 10.1073/pnas.82.20.6970 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bell A. M., Henshall J. M., Porto-Neto L. R., Dominik S., McCulloch R., Kijas J., and Lehnert S. A.. . 2017. Estimating the genetic merit of sires by using pooled DNA from progeny of undetermined pedigree. Genet. Sel. Evol. 49:28. doi: 10.1186/s12711-017-0303-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bjørnland T., Bye A., Ryeng E., Wisløff U., and Langaas M.. . 2018. Powerful extreme phenotype sampling designs and score tests for genetic association studies. Stat. Med. 37:4234–4251. doi: 10.1002/sim.7914 [DOI] [PubMed] [Google Scholar]
- Dominik S., Porto-Neto L. R., Reverter A., and Lehnert S.. . 2018. Commercial records from DNA pooling can augment classical information sources in an Australian Angus selection index to increase accuracy. In: Proceedings of the World Congress on Genetics Applied to Livestock Production, vol. Genetic Gain - Genotyping & Phenotyping Strategies. p. 94. [Google Scholar]
- Dong L., Xiao S., Chen J., Wan L., and Wang Z.. . 2016. Genomic selection using extreme phenotypes and pre-selection of SNPs in large yellow croaker (Larimichthys crocea). Mar. Biotechnol. (NY). 18:575–583. doi: 10.1007/s10126-016-9718-4 [DOI] [PubMed] [Google Scholar]
- Gerber P. J., Steinfeld H., Henderson B., Mottet A., Opio C., Dijkman J., Falcucci A., and Tempio G.. . 2013. Tackling climate change through livestock—a global assessment of emissions and mitigation opportunities.Rome: Available from http://www.fao.org/docrep/018/i3437e/i3437e.pdf [DOI] [PubMed] [Google Scholar]
- Hayes B. J., Lewin H. A., and Goddard M. E.. . 2013. The future of livestock breeding: genomic selection for efficiency, reduced emissions intensity, and adaptation. Trends Genet. 29:206–214. doi: 10.1016/j.tig.2012.11.009 [DOI] [PubMed] [Google Scholar]
- Henshall J. M., Dierens L., and Sellars M. J.. . 2014. Quantitative analysis of low-density SNP data for parentage assignment and estimation of family contributions to pooled samples. Genet. Sel. Evol. 46:51. doi: 10.1186/s12711-014-0051-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henshall J. M., Hawken R. J., Dominik S., and Barendse W.. . 2012. Estimating the effect of SNP genotype on quantitative traits from pooled DNA samples. Genet. Sel. Evol. 44:12. doi: 10.1186/1297-9686-44-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston S. E., Lindqvist M., Niemelä E., Orell P., Erkinaro J., Kent M. P., Lien S., Vähä J. P., Vasemägi A., and Primmer C. R.. . 2013. Fish scales and SNP chips: SNP genotyping and allele frequency estimation in individual and pooled DNA from historical samples of Atlantic salmon (Salmo salar). BMC Genomics 14:439. doi: 10.1186/1471-2164-14-439 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karaman E., Lund M. S., Anche M. T., Janss L., and Su G.. . 2018. Genomic prediction using multi-trait weighted GBLUP accounting for heterogeneous variances and covariances across the genome. G3 (Bethesda). 8:3549–3558. doi: 10.1534/g3.118.200673 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keele J. W., Kuehn L. A., McDaneld T. G., Tait R. G., Jones S. A., Smith T. P., Shackelford S. D., King D. A., Wheeler T. L., Lindholm-Perry A. K., . et al. 2015. Genomewide association study of lung lesions in cattle using sample pooling. J. Anim. Sci. 93:956–964. doi: 10.2527/jas.2014-8492 [DOI] [PubMed] [Google Scholar]
- Pérez-Enciso M., and Misztal I.. . 2011. Qxpak.5: old mixed model solutions for new genomics problems. BMC Bioinformatics. 12:202. doi: 10.1186/1471-2105-12-202 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reverter A., Henshall J. M., McCulloch R., Sasazaki S., Hawken R., and Lehnert S. A.. . 2014. Numerical analysis of intensity signals resulting from genotyping pooled DNA samples in beef cattle and broiler chicken. J. Anim. Sci. 92:1874–1885. doi: 10.2527/jas.2013-7133 [DOI] [PubMed] [Google Scholar]
- Reverter A., Porto-Neto L. R., Fortes M. R., McCulloch R., Lyons R. E., Moore S., Nicol D., Henshall J., and Lehnert S. A.. . 2016. Genomic analyses of tropical beef cattle fertility based on genotyping pools of Brahman cows with unknown pedigree. J. Anim. Sci. 94:4096–4108. doi: 10.2527/jas.2016-0675 [DOI] [PubMed] [Google Scholar]
- Sonesson A. K., Meuwissen T. H., and Goddard M. E.. . 2010. The use of communal rearing of families and DNA pooling in aquaculture genomic selection schemes. Genet. Sel. Evol. 42:41. doi: 10.1186/1297-9686-42-41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strillacci M. G., Frigo E., Canavesi F., Ungar Y., Schiavini F., Zaniboni L., Reghenzani L., Cozzi M. C., Samoré A. B., Kashi Y., . et al. 2014. Quantitative trait loci mapping for conjugated linoleic acid, vaccenic acid and ∆(9)-desaturase in Italian Brown Swiss dairy cattle using selective DNA pooling. Anim. Genet. 45:485–499. doi: 10.1111/age.12174 [DOI] [PubMed] [Google Scholar]
- VanRaden P. M. 2008. Efficient methods to compute genomic predictions. J. Dairy Sci. 91:4414–4423. doi: 10.3168/jds.2007-0980 [DOI] [PubMed] [Google Scholar]
- Webb R., and Buratini J.. . 2016. Global challenges for the 21st century: the role and strategy of the agri-food sector. Anim. Reprod. 13:133–142. doi: 10.21451/1984-3143-ar882 [DOI] [Google Scholar]
- Yang J., Jiang H., Yeh C. T., Yu J., Jeddeloh J. A., Nettleton D., and Schnable P. S.. . 2015. Extreme-phenotype genome-wide association study (XP-GWAS): a method for identifying trait-associated variants by sequencing pools of individuals selected from a diversity panel. Plant J. 84:587–596. doi: 10.1111/tpj.13029 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.