Abstract
Bhendi yellow vein mosaic disease (BYVMD) and Okra enation leaf curl disease (OELCuD) are common diseases of okra/bhendi [Abelmoschus esculentus (L.) Moench] affecting both pod yield and quality in the Indian subcontinent. BYVMD is caused by the infection of a begomovirus and associated betasatellite. In this study, we have made an attempt to investigate the diversity of begomoviral and the satellite sequences in okra samples showing BYVMD and OELCuD, by using a rapid PCR-based approach on 46 samples collected from 23 locations of Southern and Western India. We have also analyzed nine RCA-generated full-length begomoviral clones, some generated from the above samples displaying BYVMD and some OELCuD. By the PCR approach, we find the presence of begomovirus okra enation leaf curl virus (OELCuV) in most samples, irrespective of the disease being displayed (BYVMD or OELCuD). The nine apparently full-length sequences also show high identities with OELCuV and show instances of both intra-specific as well as intra-strainal recombination. We have also analyzed the begomoviral sequences associated with BYVMD and OELCuD from publicly available nucleotide sequence databases and show much higher sequence diversity amongst BYVMV, as compared to OELCuV. This is the first study which comprehensively demonstrates the presence of OELCuV in okra samples showing BYVMD and those showing OELCuD.
Electronic supplementary material
The online version of this article (10.1007/s13337-019-00551-4) contains supplementary material, which is available to authorized users.
Keywords: Begomovirus, Okra, Betasatellite, Alphasatellite, Diversity, PCR, Recombination
Introduction
Geminiviruses are plant-infecting viruses with single-stranded (ss) DNA genomes. Geminiviruses are a major threat to crop production in the tropical and sub-tropical regions of the world. The family Geminiviridae has been classified recently to comprise nine genera: Becurtovirus, Begomovirus, Capulavirus, Curtovirus, Eragrovirus, Grablovirus, Mastrevirus, Topocuvirus and Turncurtovirus [1]. Begomoviruses can have bipartite DNA (having two molecules, DNA-A and DNA-B) or monopartite DNA (having a single DNA molecule, resembling DNA-A [2]). The majority of the monopartite begomoviruses are found to be associated with a class of ssDNA satellites known as betasatellites (Genus; Betasatellite and Family; Tolecusatellitidae [3]; previously known as DNA β [4]).
Okra (Abelmoschus esculentus, family Malvaceae), which is commonly known as Bhendi in India, is an important vegetable grown in major parts of the country. One of the major production constraints for okra is yellow vein mosaic disease/bhendi yellow vein mosaic disease (YVMD/BYVMD), first reported in 1924 from Bombay Presidency [5], and subsequently, the causal agent reported to be a virus sixteen years later [6]. In recent years, okra enation leaf curl disease (OELCuD), another disease of okra, where the plant shows upward leaf curling, petiole bending, leathery leaves and enations in the underside of leaves, has also appeared sporadically in India [7]. Both BYVMD and OELCuD are now considered to be spreading rapidly in areas under okra cultivation in the entire Indian subcontinent [8, 9]. BYVMD is caused by a complex of a monopartite begomovirus, belonging to the species Bhendi yellow vein mosaic virus (BYVMV) and its cognate betasatellite, Bhendi yellow vein mosaic betasatellite (BYVMB) [10], demonstrated elegantly by using infectious clones of the corresponding viral DNA (10). In addition to the above, the association of a new begomoviral species, Bhendi yellow vein Delhi virus (BYVDV) from okra plants exhibiting YVMD from a location near Delhi, in Northern India, has been reported [11]. Another study [12] described a new monopartite begomovirus species Bhendi yellow vein Bhubhaneswar virus from eastern part of India. Subsequently, the association of Cotton leaf curl Bangalore virus (CLCuBaV) with naturally-infected okra plants exhibiting yellow vein and leaf curl symptoms was reported [13], which was different from typical symptoms of OELCuD. Recently, there has been a report of the association of an alphasatellite with OELCuD [14]. Later, the association of BYVMV and the DNA-B component of another bipartite begomovirus, belonging to the species Tomato leaf curl New Delhi virus (ToLCNDV) from BYVMD-affected okra plants was reported [15] from three widely separated geographical locations in India. A begomoviral DNA sequence resembling the species Mesta yellow vein mosaic virus (MeYVMV), which has been reported earlier from a fibre crop Mesta (kenaf, Hibiscus cannabinus L.) in different parts of India [16–20], has also been found associated with BYVMD [21].
Several sequences belonging to the begomoviral species Okra enation leaf curl virus (OELCuV) have been deposited in publicly accessible databases and approved by the International Committee for Taxonomy of Viruses (ICTV), which are associated with okra plants exhibiting OELCuD, including some reported recently [22]. However, in the absence of infectious clones capable of producing OELCuD symptoms in okra, the causative agents of OELCuD are, as yet, unclear. None of the cloned viral and satellite DNAs, mentioned above, have been shown to be infectious on okra, except BYVMV, along with BYVMB [10].
This study was conducted to gather additional information on the geminiviruses and satellite sequences present in naturally-infected okra plants displaying symptoms of BYVMD and OELCuD, collected from 46 samples, taken from 23 locations of Southern and Western India. Using a simple PCR procedure, we report the detection of only OELCuV from a majority of the above samples. Sequence analysis of nine full-length clones obtained from some of the above samples showed them to be variants of OELCuV, displaying evidence of extensive recombination events. We also analyzed the variability of sequences of BYVMV and OELCuV retrieved from databases.
Materials and methods
Sequence analysis of okra-infecting begomoviruses deposited in the databases
Thirteen BYVMV and twelve OELCuV nucleotide sequences each were retrieved from GenBank. Only those sequences were chosen that were enlisted by ICTV. Using the ClustalW algorithm in Sequence Demarcation Tool version 1.2 [23] percentage pair-wise identity of the representative sequences from the database was generated. To analyze diversity in open reading frames (ORFs) in BYVMV and OELCuV, protein sequence of each ORF was compared with corresponding protein sequence of infectious BYVMV clone and one representative OELCuV. BYVMV and OELCuV sequences were retrieved from GenBank and pairwise identity of sequence analysis was calculated using algorithm BlastP.
Sample collection and genomic DNA isolation
Leaf samples of okra plants showing typical symptoms of BYVMD were collected from 9 field locations of Gujarat (near Anand, Baroda, Bhavnagar, Gandhinagar, Himmatnagar, Navsari and Tapi), 10 of Maharashtra (near Amravati, Aurangabad, Buldhana, Jalgaon, Jalna, Nagpur, Nashik, Pune, Satara, and Wardha), and 3 of Tamil Nadu (near Theni, Tirunelveli and Tuticorin) in the winter season of the year 2014. A map of Gujarat, Maharashtra and Tamil Nadu with the origins of all isolated BYVMD-affected samples is shown in Fig. 1. The number of the samples collected was 42. In addition, four samples showing symptoms of OELCuD were collected from Jalgaon, Maharashtra in the summer of 2017, making a total of 46 samples (Table 2). Isolation of total genomic DNA from the samples was performed by employing CTAB-Mucilage-free method [24].
Fig. 1.
Geographic origins of okra samples. Map of Gujarat, Maharashtra and Tamil Nadu showing the geographic origins of the okra samples with begomovirus like symptoms from which partial sequences were cloned. The stars indicate 7 field locations of Gujarat (near Anand, Baroda, Bhavnagar, Gandhinagar, Himmatnagar, Navsari and Tapi), 10 field locations of Maharashtra (near Amravati, Aurangabad, Buldhana, Jalgaon, Jalna, Nagpur, Nashik, Pune, Satara, and Wardha), and 3 field locations of Tamil Nadu (near Theni, Tirunelveli and Tuticorin)
Table 2.
Locations of sites for the collection of okra samples for PCR amplification to detect the presence of BYVMV, OELCuV and satellites (alphasatellites and betasatellites)
| S.no | State | District | PCR amplification using specific primers for: | |||
|---|---|---|---|---|---|---|
| OELcV | BYVMV | Betasatellite | Alphasatellite | |||
| 1 | Maharashtra | Nagpur | + | − | + | + |
| + | − | − | − | |||
| 2 | Wardha | + | − | + | − | |
| + | − | + | + | |||
| 3 | Amravati | + | − | − | − | |
| + | − | − | + | |||
| 4 | Buldhana | + | − | − | − | |
| + | − | + | + | |||
| 5 | Jalna | + | − | + | − | |
| + | − | + | − | |||
| 6 | Aurangabad | + | − | + | + | |
| + | − | + | − | |||
| 7 | Nashik | + | − | + | − | |
| + | − | + | − | |||
| 8 | Jalgaon-1 | + | − | − | − | |
| 9 | Satara | + | − | + | + | |
| + | − | − | + | |||
| 10 | Pune | + | − | + | − | |
| + | − | − | − | |||
| 11 | Gujarat | Gandhinagar | + | − | − | − |
| + | − | − | − | |||
| 12 | Himmatnagar | + | − | + | − | |
| + | − | − | − | |||
| 13 | Baroda | + | − | − | − | |
| + | − | + | − | |||
| 14 | Anand | + | − | − | + | |
| Kantharia | + | − | − | − | ||
| 15 | Anand | + | − | + | + | |
| Sisva | + | − | − | − | ||
| 16 | Bhavnagar | + | − | − | + | |
| 17 | Navsari | + | − | − | − | |
| Valsad | + | − | − | + | ||
| 18 | Navsari | + | − | + | + | |
| + | − | − | − | |||
| 19 | Tapi | + | − | + | + | |
| + | − | − | + | |||
| 20 | Tamil Nadu | Tirenelveli | + | + | + | − |
| + | − | − | − | |||
| 21 | Tuticorin | + | − | + | − | |
| + | − | − | − | |||
| 22 | Theni | + | − | − | − | |
| + | + | + | − | |||
| 23 | Maharashtra | Jalgaon-2 | + | − | N/A | N/A |
| Jalgaon-3 | + | − | N/A | N/A | ||
| Jalgaon-4 | + | − | N/A | N/A | ||
| Jalgaon-5 | + | − | N/A | N/A | ||
Two samples were collected from each site, except Sr. no. 8 and 16, from where one sample each were collected and Sr. no. 23, from where four samples were collected. Samples from locations 1–22 displayed BYVMD and samples from location 23 displayed OELCuD
N/A indicates not analyzed, Sr. no. 1–22 were collected in the year 2014 and Sr. no. 23 was collected in the year 2017
Polymerase chain reaction (PCR), Rolling circle amplification (RCA), cloning and sequencing
For detecting BYVMV, OELCuV and satellites (alphasatellites and betasatellites (BYVMB), reported to be associated with BYVMD and/or OELCuD [10, 14], four pairs of primers were designed. List of all the primers with the sequence, position and amplicon size for each is given in Supplementary Table 1. A gradient PCR was performed to confirm the optimal annealing temperature, for the amplification of the template DNAs. PCR was performed in 10 µl reaction mixes containing 1 µl of template DNA (genomic DNA), 0.4 units of Phusion™ High fidelity DNA polymerase (Thermo Scientific), 0.2 mM dNTPs, 2.5 pmol of each forward and reverse primers, 2 µl of 5X Phusion GC Buffer and 0.1 µl of 100% DMSO. The amplification was carried out using a ProFlex Base PCR thermal cycler (Applied Biosystems, UK). Amplification programmes consisted of one initial cycle of denaturation at 95 °C for 5 min, annealing at (56–62 °C) for 30 s and final extension at 72 °C for 7 min. Thirty cycles of denaturation at 95 °C for 1 min, annealing (56–62 °C) for 30 s and extension at 72 °C for 30 s, were then performed finishing the amplification programme with 7 min at 72 °C. Amplified products were electrophoresed through 1.2% (w/v) agarose gels in 0.5 XTris Boric acid EDTA (TBE) electrophoresis buffer, and visualised under UV. However, an annealing temperature of 58 °C was eventually chosen for better yield in the PCR assays.
With the aim of amplifying and cloning full-length sequences of the known okra-infecting begomoviruses in BYVMD-affected leaf samples, three pairs of abutting primers were designed, based on the coat protein (CP) sequence of BYVMV for the amplification of full-length viral DNA (BYVMV-F; BYVMV-R, SspI-F; SspI-R, EcoRI-F; EcoRI-R.). All the primer sets were designed using Gene Runner (v 3.01). PCR amplification was carried out using these three primer sets, details of which are given in Supplementary Table 2. PCR was performed in 15 µl reaction mix containing 1 µl of template DNA (10X Genomic DNA), 0.4 units of iTaq DNA polymerase (iNtRON), 0.5 mM dNTPs, 2.5 pmol of each forward and reverse primers, 1.5 µl of 10X Buffer. The amplification was carried out using a ProFlex Base PCR thermal cycler (Applied Biosystems, UK). Amplification programmes consisted of one initial cycle of denaturation at 95 °C for 5 min, annealing at (48–52 °C) for 30 s, extension at 72 °C for 1 min 30 s. Thirty cycles of denaturation at 94 °C for 30 s, annealing (48–52 °C) for 30 s and extension at 72 °C for 1 min 30 s, were then performed finishing the amplification programme with 7 min at 72 °C. Amplified products were electrophoresed through 1.2% (w/v) agarose gels in 0.5 XTris Boric acid EDTA (TBE) electrophoresis buffer, and visualised under UV. The PCR amplified product was eluted, A-tailed, ethanol precipitated and ligated in pTZ57R/T vector (InsTAclone PCR cloning kit, Thermo Fisher Scientific) and 42 cloned approximately 2.7 kb DNA fragments were subjected to end sequencing using M13F primer at DNA sequencing facility of the University of Delhi South Campus, New Delhi, India. The obtained sequences were then subjected to Blastn analysis [25] against NCBI nucleotide database. Three clones were fully sequenced (Supplementary Table 3).
Full-length genomes of resident begomoviruses from six random samples (4 displaying OELCuD and 2 BYVMD) were amplified by using the sequence-independent process of Rolling Circle Amplification (RCA) [26] using TempliPhi DNA amplification kit (GE Healthcare, USA) according to manufacturer’s instructions. RCA products were digested with the unique cutter restriction enzyme KpnI (NEB, USA), which had single recognition sites in the BYVMV, MeYVMV and OELCuV DNA sequences available in the database. The purified DNA fragments (ca 1.4 and 2.7 kb) were cloned into pGreen0029 vector and transformed into E. coli DH5α cells. The DNA of one clone from each sample was sequenced by automated sequencer at UDSC, New Delhi (Supplementary Table 3).
Sequence analysis
Partial sequences from different field locations of Gujarat, Maharashtra and Tamil Nadu were analyzed using Blastn search program (https://blast.ncbi.nlm.nih.gov/Blast.cgi). ORF finder was used for ORF identification (https://www.ncbi.nlm.nih.gov/orffinder/). The ExPASy proteomic server tool was used to translate set of protein encoding genes (https://web.expasy.org/translate/). The sequences were submitted to GenBank. Multiple sequence alignments and phylogenetic trees were drawn using MEGA software version 6 [27]. Pair wise alignment was done using SDT version 1.2 [23]. For recombination analysis full length sequences of the begomoviruses and associated satellite DNA of the present study were analyzed using Recombination Detection Program (RDP) version 4.0 [28]. It uses six different algorithms/automated methods namely RDP, GENECONV, BootScan, Chimera, SiScan and 3Seq with highest acceptable probability value P = 0.05.
Data availability
The partial sequences generated in this study are provided with accession numbers MK962353, MK962354, MK962355, MK962356, MK962357, MK962358, MK962359, MK962360, MK962361, MK962362, MK962363, MK962364, MK962365, MK962366, MK962367, MK962368, MK962369, MK962370, MK962371, MK962372, MK962373 from NCBI while the full length sequences generated in this study are available in the GenBank under the accession numbers MK069433, MK069434, MK069435, MK069436, MK069437, MK084765, MK084766; MK084767, MK084768, MK145076, MK145077, MK145078 and MK145075.
Results
Variability of nucleotide sequences from okra-infecting begomoviruses from databases
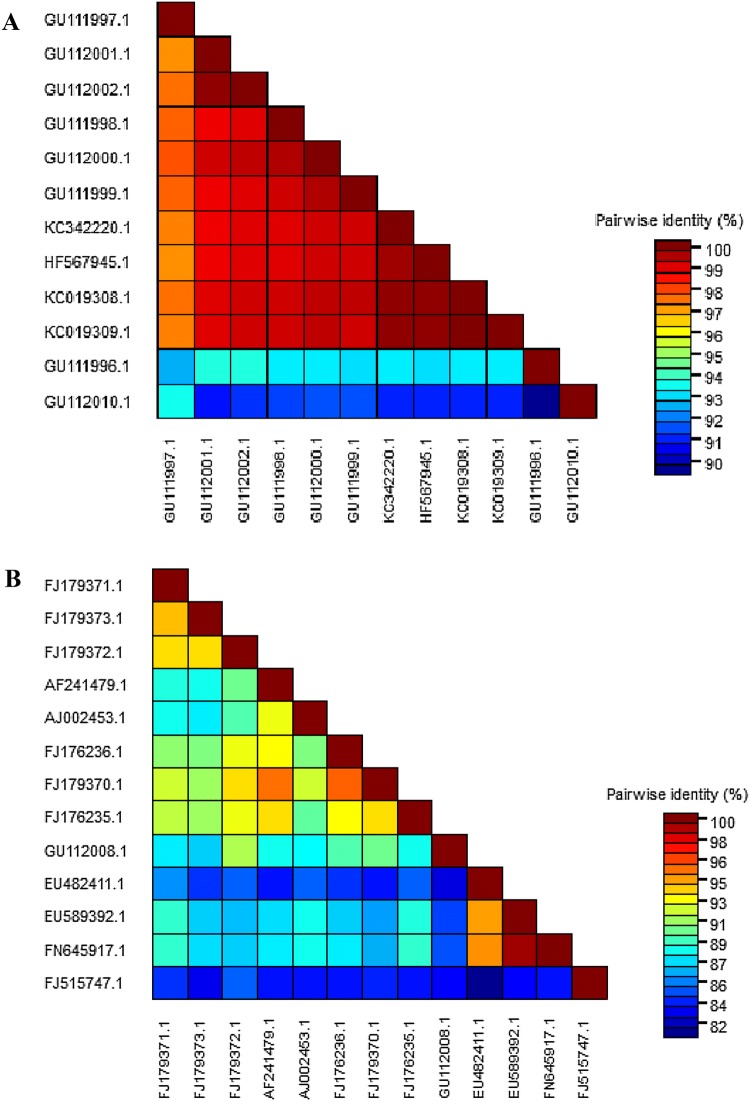
When begomoviral sequences from okra already deposited in the database were compared, it was noticed that within BYVMV-like group, which included thirteen sequences of species belonging to BYVMV, the overall nucleotide sequence identity varied from 91 to 98%. Within the OELCuV-like group, consisting of twelve sequences approved by ICTV, the nucleotide sequence variability was much less, from 97 to 99% identities (Fig. 2). Although the sequences had been deposited from samples collected from regions widely separated in geographical distance from each other, no geographical clustering of sequences could be noticed. For example, FJ179373, a clone from the eastern state of West Bengal showed about 93% identity with FJ179372, a clone from the southern state of Tamil Nadu and with FJ179371, a clone from the northern region (Chandigarh), but only about 86% identity with another clone FJ176235, from Tamil Nadu. The clones FJ176235 and FJ179372, both from Tamil Nadu, collected a few hundred kilometers (kms) from each other, shared 93% identity with each other, about the same as shared by clones from West Bengal, at least 2000 km away. When the amino acid sequences of the six ORFs of nineteen selected sequences were compared to that of the only infectious clone of BYVMV (Table 1), the ORF V2 was seen to be the most conserved, whereas the ORF C4 had the least conserved residues.
Fig. 2.
Sequence Demarcation Tool based pairwise sequence comparisons. Colour coded pairwise color-code similarity matrix generated using thirteen full length OELCuV (a) and full length BYVMV (b) nucleotide sequences using Clustalw algorithm. Each colored cell represents a share identity score between 2 sequences (one indicated horizontally to the left and also the alternative vertically at the bottom). A colored key indicates the percentage similarity correspondence between pairwise identities. a GU111997.1 Okra enation leaf curl virus [India:Sonipat EL14:2006] segment DNA-A; GU112001.1 Okra enation leaf curl virus [India:Munthal EL39:2006] segment DNA-A; GU112002.1 Okra enation leaf curl virus [India:Munthal EL38:2006] segment DNA-A; GU111998.1 Okra enation leaf curl virus [India:Munthal EL37:2006] segment DNA-A; GU112000.1 Okra enation leaf curl virus [India:Munthal EL32:2006] segment DNA-A; GU111999.1 Okra enation leaf curl virus [India:Munthal EL41:2006] segment DNA-A; KC342220.1 Okra enation leaf curl virus isolate Surat; HF567945.1 Okra enation leaf curl virus, isolate Pakistan:cotton Burewala:2011; KC019308.1 Okra enation leaf curl virus isolate India:Gandhinagar:2012 segment DNA-A, complete sequence;KC019309.1 Okra enation leaf curl virus isolate India:Gandhinagar:2012 segment DNA-A; GU111996.1 Okra enation leaf curl virus [India:Sonipat EL10:2006] segment DNA-A; GU112010.1 Okra leaf curl India virus [India:Sonipat EL14A:2006] segment DNA-A, complete sequence. b FJ179371.1 Bhendi yellow vein mosaic virus isolate OY177 segment DNA-A; FJ179373.1 Bhendi yellow vein mosaic virus isolate OY-W1 segment DNA-A; FJ179372.1 Bhendi yellow vein mosaic virus isolate OYCO4 segment DNA-A; AF241479.1 Bhendi yellow vein mosaic virus-[Madurai] segment A; AJ002453.1 Bhendi yellow vein mosaic virus-[301] Pakistan strain segment A; FJ176236.1 Bhendi yellow vein mosaic virus [2005:Pandaralli:India]; FJ179370.1 Bhendi yellow vein mosaic virus isolate OY-T segment DNA-A; FJ176235.1 Bhendi yellow vein mosaic virus [2005:Thadagan:India] segment DNA-A; GU112008.1 Bhendi yellow vein India virus [India:Karnal OY80A:2005] segment DNA-A; EU482411.1 Bhendi yellow vein mosaic virus isolate Nol 101 segment DNA A; EU589392.1 Bhendi yellow vein mosaic virus isolate NOL751 segment DNA A; FN645917.1 Bhendi yellow vein mosaic virus segment A; FJ515747.1 Bhendi yellow vein mosaic virus [2004:New Delhi] isolate OY131 segment DNA-A (colour figure online)
Table 1.
Pairwise protein similarity of different ORFs
| Virus name | C1 | C2 | C3 | C4 | C5 | V1 | V2 |
|---|---|---|---|---|---|---|---|
| (a) BYVMV when compared with ORFs of Bhendi yellow vein mosaic virus-[Madurai] | |||||||
| BYVMV-[Madurai] | 100 | 100 | 100 | 100 | 100 | 100 | 100 |
| BYVMV isolate OYCO4 segment | 87 | 94 | 95 | 73 | 81 | 87 | 94 |
| BYVMV [2004:ND] OY131 segment | 87 | 98 | 90 | 71 | 84 | 87 | 99 |
| BYVMV [IN:Tirupathi OY98:2005] | 88 | 99 | 96 | 73 | 89 | 88 | 96 |
| BYVMV [IN:Srinivaspur OYSK2:2006] | 81 | 96 | 89 | 49 | 69 | 81 | 92 |
| BYVMV [IN:Thadagan:OY156:2006] | 80 | 83 | 92 | 49 | 90 | 80 | 95 |
| BYVMV [IN:Hyderabad:OY65:2005] | 75 | 95 | 95 | 49 | 88 | 75 | 95 |
| BYVMV [IN:Trichy:OY115B:2005] | 77 | 95 | 86 | 49 | 81 | 77 | 94 |
| BYVMV [IN:Sonipat:OY96:2005] | 79 | 94 | 88 | 54 | 44 | 79 | 98 |
| BYVMV [IN:Karnal:OY79A:2005] | 90 | 98 | 90 | 72 | 87 | 90 | 97 |
| BYVMV [IN:Karnal:OY80B:2005] | 88 | 97 | 95 | 71 | 81 | 88 | 95 |
| BYVMV [IN:Delhi:OY133:2006] | 89 | 95 | 95 | 71 | 90 | 90 | 97 |
| BYVMV [IN:Coimbator:OYCO4:2005] | 87 | 97 | 95 | 73 | 81 | 87 | 94 |
| BYVMV isolate OY77B | 87 | 94 | 98 | 73 | 85 | 87 | 99 |
| BYVMV isolate OY79B | 88 | 99 | 96 | 70 | 89 | 88 | 98 |
| BYVMV isolate OY173 | 91 | 98 | 96 | 73 | 90 | 91 | 97 |
| BYVMV isolate OY138B | 91 | 97 | 97 | 73 | 90 | 91 | 99 |
| BYVMV isolate OYDWR1 | 88 | 99 | 95 | 67 | 88 | 88 | 98 |
| BYVMV isolate OY118 | 91 | 98 | 97 | 74 | 88 | 91 | 99 |
| BYVMV isolate 126B | 93 | 99 | 96 | 79 | 89 | 93 | 94 |
| (b) OELCuV when compared with ORFs of Okra enation leaf curl virus [India:Sonipat EL10:2006] | |||||||
| OELCuV [IN:Son EL10:2006] | 100 | 100 | 100 | 100 | 100 | 100 | |
| OELCuV [IN:Son EL14:2006] | 87 | 85 | 92 | 62 | 97 | 100 | |
| OELCuV [IN:Mun EL37:2006] | 89 | 87 | 93 | 62 | 98 | 100 | |
| OELCuV [IN:Mun EL41:2006] | 89 | 86 | 93 | 62 | 96 | 98 | |
| OELCuV [IN:Mun EL32:2006] | 90 | 85 | 92 | 63 | 98 | 98 | |
| OELCuV [IN:Mun EL39:2006] | 90 | 85 | 92 | 63 | 100 | 100 | |
| OELCuV [IN:Mun EL38:2006] | 89 | 86 | 93 | 62 | 100 | 100 | |
| OELCuV [IN:Son EL14A:2006] | 89 | 97 | 96 | 63 | 85 | 98 | |
| OELCuV, isolate Pakistan:cotton-Burewala:2011 | 90 | 85 | 93 | 63 | 94 | 99 | |
| OELCuV isolate IN:Gandhinagar:2012 | 90 | 87 | 93 | 63 | 94 | 99 | |
| OELCuV isolate IN:Gandhinagar:2012 | 90 | 87 | 93 | 63 | 93 | 99 | |
| OELCuV isolate Surat | 89 | 87 | 93 | 63 | 94 | 99 | |
PCR-based detection of begomoviruses and satellites
To detect the presence of BYVMV, OELCuV, BYVMB and alphasatellite from 42 BYVMD affected okra leaf samples and 4 affected by OELCuD, PCR were performed using specific primers. These primers were designed based on the full-length sequences of BYVMV (AF241479), OELCuV (MK084766), alphasatellite (NC_018574) and BYVMB (KJ462076) available from publicly-accessible databases. At an annealing temperature range of 54–62 °C, a 470 bp DNA fragment, expected of an amplification product of BYVMV and a 327 bp DNA fragment expected from OELCuV were amplified (Supplementary Fig. 1a, b). At an annealing temperature range of 56–66 °C, a 467 bp DNA fragment, indicative of an alphasatellite and a 345 bp DNA fragment indicative of the presence of BYVMB were amplified (Supplementary Fig. 1c). However, an annealing temperature of 58 °C was eventually chosen for better yield in the gradient PCR assays. An optimized PCR assay for the amplification of BYVMV and OELCuV was performed, which amplified a single intense band of an expected amplicon size, about 327 bp indicating the presence of OELCuV from 42 BYVMD-affected samples and 4 OELCuD-affected samples (Supplementary Fig. 1, Table 2). Two samples, one from Tirunelveli and another from Theni, both from Tamil Nadu, showing symptoms of BYVMD amplified both BYVMV- and OELCuV-specific bands, indicating the possible presence of both the viruses in these samples. PCR assay to detect the presence of BYVMB and alphasatellite indicated the presence of the former in 20 and the latter in 14 samples (Table 2). Interestingly, 8 samples showing symptoms of BYVMD indicated the presence of both alphasatellite as well as BYVMB.
Full-length clones representing DNA of begomoviruses and betasatellites
With the aim of amplifying and cloning full-length sequences of any of the known okra-infecting begomoviruses in BYVMD-affected leaf samples, three pairs of abutting primers were used for the amplification of full-length viral DNA. However, despite repeated efforts to get a full-length fragment from all the 42 infected samples from BYVMV-F and BYVMV-R set of primers, the complete 2.7 kb fragment of begomoviruses was PCR amplified only by using the other two sets of primers based on a conserved region present in the full-length nucleotide sequences available in NCBI nucleotide database. A total of 6 apparently full-length viral DNA collected from Tamil Nadu were amplified using EcoRI-F/EcoRI-R primer set, while 36 full-length viral DNA collected from regions of Maharashtra and Gujarat were amplified using SspI-F/SspI-R primer set. All 42 apparently full-length clones (2.7 kb) were cloned in TA vector. Preliminary Blastn analysis of partial sequences (700–850 bp) obtained from end-sequencing using M13F primer from 4 locations of Gujarat (Anand, Baroda, Bhavnagar and Tapi), 9 of Maharashtra (Amravati, Aurangabad, Buldhana, Jalgaon, Jalna, Nagpur, Nashik, Satara and Wardha) and 3 of Tamil Nadu (Theni, Tirunelveli and Tuticorin) showed 99.85% identity with OELCuV (KX710156.1). Begomoviral partial sequences from Wardha region of Maharashtra showed highest sequence identity of 99.85% OELCuV (KX710156.1). Out of the 42 BYVMD-affected leaf samples, three full-length sequences of begomoviruses were obtained using primer walking facility at DNA sequencing facility at Department of Biochemistry, University of Delhi South Campus (Supplementary Table 3).
Figure 3 shows the pairwise alignment of thirteen partial sequences described here, representing C-terminal portions of the coat protein gene and the corresponding sequences of other begomoviral sequences associated with okra, based on information available in the literature. All the thirteen partial sequence showed 99–100% pairwise identity to the sequences of OELCuV (KC019308.1) and MeYVMV (KJ462074.1). With BYVMV (AF241479.1) the pairwise identity was 70–75%.
Fig. 3.
Sequence Demarcation Tool based pairwise sequence comparisons between partial sequences of coat protein genes of nine full-length sequences generated in this study and six corresponding sequences taken from database. Colour coded pairwise color-code similarity matrix generated using C-terminal region of the coat protein gene of two full length sequences each of BYVMV (AF241479, EU482411), OELCuV (KC019308, GU111996) and MeYVMV (KJ462074, EF373060) with the corresponding region of thirteen partial sequences using Clustalw algorithm. Each colored cell represents a share identity score between 2 sequences (one indicated horizontally to the left and also the alternative vertically at the bottom). A colored key indicates the percentage similarity correspondence between pairwise identities (colour figure online)
Nine complete sequences of begomovirus (three clones were amplified using PCR, while six were amplified using RCA, Supplementary Table 3) and four complete sequences of betasatellites isolated from BYVMD-affected and OELCuD-affected okra were cloned, sequenced and submitted in GenBank. All nine complete sequences of begomovirus either isolated from BYVMD-affected okra (Nagpur, MK069433; Himmatnagar, MK069434; Jalna, MK069435; Aurangabad, MK069436 and Anand, MK069437) or OELCuD-affected okra (Jalgaon, MK084765; MK084766; MK084767 and MK084768) shared maximum identity 98% with OELCuV (GU111997). Two sequences from Jalgaon (Maharashtra) region viz. MK084766 and MK084767 which were isolated from OELCuD-affected samples shared maximum identity 97% and 98% identities with OELCuV-Sonipat (GU111997) and OELCuV-Gandhinagar (KC019309) isolates, respectively (Fig. 4). As per the species demarcation threshold accepted for geminiviruses (91%) [29], all the nine sequences determined in this study can be described as isolates of OELCuV. BYVMV sequences could not be obtained. However, the betasatellite sequences which are associated with BYVMD (Nashik, MK145076; MK145077; MK145078) and OELCuD (Jalgaon, MK145075) revealed maximum identity 99% and 98% with BYVMB and with the species belonging to Okra enation leaf curl betasatellite (OELCuB), respectively, greater than the species demarcation threshold for betasatellites (78%) [30], indicating them to be isolates of BYVMB and OELCuB, respectively [30].
Fig. 4.
Sequence Demarcation Tool based pairwise sequence comparisons. Colour-coded pairwise identity matrix generated from (A to I) 33 begomovirus genomes (Okra enation leaf curl virus and Bhendi yellow vein mosaic virus). Each colored cell represents a share identity score between 2 sequences (one indicated horizontally to the left and also the alternative vertically at the bottom). A colored key indicates the correspondence between pairwise identities and also the colors displayed within the matrix. The sequences generated by this study are indicated by red boxes (colour figure online)
Phylogenetic analysis revealed that the sequences generated in the present study grouped with other OELCuV isolates reported from India and Pakistan, but not those of BYVMV, which grouped separately (Fig. 5). Phylogenetic dendogram showed that the two begomovirus sequences of Jalgaon isolate (MK084766 and MK084767) identified from OELCuD, clustering together with other isolates of OELCuV, while the remaining seven sequences either isolated from BYVMD or OELCuD grouped with other isolates of OELCuD in a separate clade (Fig. 5a). However, the four betasatellite sequences were closely related to BYVMB and OELCuB previously reported from India (Fig. 5b). The betasatellite sequences (MK145076, MK145077 and MK145078), which were associated with BYVMD clustered with BYMVB and the betasatellite sequence associated with OELCuD, grouped with other isolates of OELCuB in a separate clade (Fig. 5b). The DNA fragments amplified using primers specific for alphasatellites have not been analyzed yet.
Fig. 5.
a Phylogenetic dendogram showing the relationship of Okra enation leaf curl virus (OELCuV, this study) with other isolates of OELCuV. b Phylogenetic dendogram showing the relationship of Bhendi yellow vein mosaic betasatellites (BYVMB) and Okra enation leaf curl betasatellite (OELCuB) associated with OELCuV (this study) with other betasatellites associated with Bhendi yellow vein mosaic virus and OELCuV. Red squares indicate sequences obtained from samples exhibiting OELCuD and green triangles from BYVMD (colour figure online)
Detection of recombination
Out of nine complete sequences of begomoviruses, seven showed recombination (inter-species and intra-species) and six showed more than one recombinant breakpoints. Four complete sequences of OELCuV viz. MK069434, MK069435, MK084766 and MK084767 showed inter-species recombination of BYVMV from India and Pakistan isolates (Supplementary Table 4). Two complete sequences namely MK084765 and MK084768 showed both inter-species and inter-strainal recombination (Supplementary Table 4). Similarly, OELCuB (MK145075) showed inter-species and inter-strain recombination of BYVMB (KJ462080) and OELCuB (KC019310) respectively. The recombination events are shown in Fig. 6.
Fig. 6.
Recombination events detected within Okra enation leaf curl virus (a) and Okra enation leaf curl betasatellite (b) using RDP4
An isolate of OELCuV reported in this study (MK069433) showed one recombination breakpoint at the position 2731-624 (IR, V1, V2 and C5) and was contributed by minor parents (OELCuV-IN, GU111998). The recombinant event in this sample was detected by seven methods (Supplementary Tables 4). Two recombinant breakpoints were detected in isolates OELCuV Himmatnagar (MK069434) and OELCuV Jalgaon 2 isolate (MK084766). One recombination breakpoint in both the isolate was signified BYVMV-IN (GU112050) as a common minor parental sequence and differed only in length of their spanning regions. BYVMV-IN (GU112050) contributed in V1, C5, C3 and C2 (696-1260) regions of OELCuV Himmatnagar isolate MK069434, and in V1, C5, C3, C2 and C1 regions (623-1985) of OELCuV (MK084766). Second breakpoint in OELCuV Himmatnagar isolate (MK069434) was contributed by BYVMV-IN (FN645917) as a minor parent in the region of C3, C2, C1, C4 and IR (1261-2740). While in the sample OELCuV (MK084766) second recombination breakpoint occurred at C1, C4 and IR region (2431-2682) and this fragment was donated by BYVMV-IN (FJ176236) (Supplementary Table 4). OELCuV Jalna isolate (MK069435) also showed two recombination break points at nucleotide region 1262-1625 (C3, C2 and C1) and 2740-1261 (IR, V2, V1, C5, C3 and C2) and these sequences were donated by BYVMV-PK (AJ002453) and BYVMV-IN (JQ326263), respectively.
The clones MK084767, MK084765 and MK084768 showed 3, 4 and 5 recombination breakpoints respectively. Recombination breakpoints for MK084767 were detected in 1476-1976 (C2 and C1), 1986-153 (C1, C4, IR and V2) and 2032-2587 (C1 and C4) regions and these fragments were donated by BYVMV-IN (JQ326263), BYVMV-PK (AJ002453) and BYVMV-IN (FR694925) respectively. Four recombination breakpoints were detected in MK084765 in 8-622 (IR, V2, V1 and C5), 1301-1952 (C3, C2 and C1), 2256-2558 (C1 and C4) and 2653-2730 (IR) sequence coordinates and these fragments were donated by OELCuV-IN (GU111996), BYVMV-PK (AJ002453), BYVMV-IN (GU112050) and BYVMV-IN (GU112007) respectively. In MK084768, 5 recombinant breakpoints were detected in the nucleotide coordinates 623-1985 (V1, C5, C3 and C2), 1452-1786 (C3, C2 and C1), 2732-622 (IR, V2, V1 and C5), 993-1451 (V1, C5, C3 and C2) and 2348-2562 (C1 and C4), and these fragments were contributed by OELCuV-IN (KC019309), BYVMV-TH (JX678967), OELCuV-IN (GU111998), BYVMV-PK (AJ002451) and BYVMV-IN (FJ176236) respectively. The recombination events detected in all the complete sequences of OELCuV of this study except MK084765 (2256-2558) and MK084768 (993-1451) were supported by more than 5 methods in RDP (Supplementary Table 4). OELCuB (MK145075) exhibited two recombination breakpoints in the nucleotide regions 127-708 (βC1) and 713-917 (A-rich) and these fragments were granted by OELCuB (KC019310) and BYVMB (KJ462080) respectively (Supplementary Table 4).
The positions of the predicted ORFs in the nine full-length OELCuV sequences generated in this study are presented in Supplementary Table 5. The percent amino acid identities of two of the full-length clones generated in this study with those of a few okra-infecting begomoviruses deposited in the database are presented in Supplementary Table 6.
Discussion
BYVMD, the first plant viral disease reported from India, and studied extensively for the past 95 years, has been shown to be associated with a large number of begomoviruses, many of which have not been examined in sufficient detail. Apart from the seminal work which showed the appearance of the symptoms of BYVMD upon agroinoculation with cloned monopartite virus BYVMV and the betasatellite BYVMB [10], the associations of various other reported viral species, some of them bipartite, with this disease [11, 12, 31], have not been fully analyzed. Some of the above might have been detected because of multiple infections at the field and their roles in the disease development need to be examined carefully. Hence, apart from BYVMV and BYVMB, there is no evidence of any other begomoviral or betasatellite species to be the causative agent for BYVMD.
Analysis of full-length assembled sequences resembling begomoviruses from 89 diseased okra samples, representing 39 widely separated locations from India [15], indicated that a majority (86 out of 89) resembled the BYVMV variant (BYVMV-IN-Mah-NOL751, accession no. EU589392) or BYVMV (accession no. AF241479), the infectious clone reported earlier [10]. Sequences resembling BYVMV were also found in five BYVMD-affected samples, from which full-length sequences were amplified [32]. By that logic, our PCR-based assay to detect BYVMV should have amplified it in most samples. However, upon using the specific primers, we saw amplification in only two samples out of 46 tested, despite 42 of them showing BYVMD (Table 2). The primers designed to amplify BYVMV also had the potential to amplify BYVMaV-IN-Mah-NOL751, based on its sequence. The predominance of OELCuV in the samples analyzed was also reflected in the 13 partial DNA sequences analyzed from the full-length clones generated in this study (Fig. 3), which all showed high sequence identity with OELCuV sequences. Also to be noted was the high sequence identity between the DNAs, some of which were isolated from samples taken from widely separated geographical locations. Although the above sequences were generated using the error-prone Taq DNA polymerase, as a preliminary step in the analysis, the sequence identity was remarkably high. Hence, we conclude that only OELCuV-type viruses were present in BYVMD- and OELCuD-affected okra samples analyzed in this study.
There could be several reasons for the lack of detection of BYVMV sequences from most samples in this report, compared to the earlier report [15], the most likely reason could be a change in the population of okra-infecting begomoviruses in recent years. The earlier report was based on samples collected in the years 2005–2006, whereas, this study used samples collected in the year 2014. A more in-depth study, possibly involving Deep Sequencing would be needed to firmly establish the nature of the viral sequences present in BYVMD- and OELCuD-affected okra in India and the neighboring countries.
Our analysis of BYVMV and OELCuV sequences retrieved from the databases showed that the former was significantly more variable compared to the latter (Fig. 2). The wider geographical areas and the longer time scale of the collected samples from which the sequences were deposited for BYVMV, compared to OELCuV could explain this finding. Another significant conclusion, which could be drawn from the analysis of sequences taken from the database, was the lack of geographical clustering, especially in OELCuV sequences, a conclusion also supported by the analysis of partial sequences of the full-length clones generated in this study (Fig. 3). Recently, the association of OELCuV with OELCuD at high frequencies in okra fields has also been demonstrated [22, 32]. In the absence of an infectious cloned OELCuV DNA, the exact role of OELCuV in causing OELCuD is still awaited.
The elegant demonstration of appearance of BYVMD in okra upon agroinoculation with BYVMV and BYVMB together [10] indicated that BYVMB was necessary for symptom production of the disease. Surprisingly, we find in this study that only 20 samples out of 46 amplified a product suggesting the presence of BYVMB (Table 2). This finding needs to be further confirmed by more rigorous study of the satellites and the associated begomoviruses and their roles in the two diseases.
Two earlier reports [14, 32] present evidence of extensive recombination amongst the begomoviruses infecting okra in India. The nine sequences of full-length clones of OELCuV generated by us support the same conclusion (Fig. 6 and Supplementary Table 4).
When the amino acid identities of the seven ORFs between two of the clones generated in this study and five clones each representing BYVMV sequences from Indian subcontinent and Thailand and OELCuV from India were compared, the most conserved ORF was seen to be C1, and the least conserved was C4, which, generally agreed with the pattern in most begomoviruses (Supplementary Table 6). However, the absence of C5 from some BYVMV and OELCuV sequences was puzzling. In our analysis of BYVMV and OELCuV sequences deposited in the database, V2 was the most conserved, followed by C2 in BYVMV, whereas it was V2, followed by V1 for OELCuV (Table 1a, b).
In conclusion, we show, using a rapid PCR-based approach, viruses resembling OELCuV are associated with okra plants exhibiting symptoms of BYVMD as well as those affected with OELCuD. BYVMB-like sequences were present only in some of the above samples. Our analysis of 9 full-length OELCuV sequences supported the above conclusion and found evidence of both intra-specific as well as inter-specific recombination.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgements
This work was funded by BIPP project no. BT/BIPP0574/18/11 by the Department of Biotechnology, Government of India as a collaborative project between University of Delhi South Campus and Nirmal Seeds Pvt. Ltd., Pachora. ID would like to acknowledge the DU-DST-PURSE Grant and the DST-FIST infrastructure grant. MN and RK are thankful to University Grants Commission, New Delhi and Department of Biotechnology, Government of India, New Delhi respectively for Research Fellowships. ZAK is thankful to Science & Engineering Research Board (DST, Government of India) for financial support as National Post Doctoral Fellowship (PDF/2016/003514).
Author contributions
This study was conceived by ID and JCR, samples were collected by SPK and VMP, samples were processed by MN and ZAK, cloning, sequence analyses and recombination analyses were carried out by MN, ZAK and RK, data analysis was carried out by MN, ZAK, RK and ID, the manuscript was written by MN, ZAK and ID, funding was arranged by JCR and ID and all the authors approved the final version of the manuscript.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Adams MJ, Lefkowitz EJ, King AMQ, Harrach B, Harrison RL, Knowles NJ, Kropinski AM, Krupovic M, Kuhn JH, Mushegian AR, Nibert A, Sabanadzovic S, Sanfaçon H, Siddell SG, Simmonds P, Varsani A, Zerbini FM, Gorbalenya AE, Davison AJ. Changes to taxonomy and the international code of virus classification and nomenclature ratified by the International Committee on Taxonomy of Viruses (2017) Arch Virol. 2017;162:2505–2538. doi: 10.1007/s00705-017-3358-5. [DOI] [PubMed] [Google Scholar]
- 2.Fondong VN. Geminivirus protein structure and function. Mol Plant Pathol. 2013;14(6):635–649. doi: 10.1111/mpp.12032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Lozano G, Trenado HP, Olivé EF, Chirinos D, Pouey FG, Briddon RW, Castillo JN. Characterization of non-coding DNA satellites associated with sweepoviruses (Genus Begomovirus, Geminiviridae)—definition of a distinct class of begomovirus-associated satellites. Front Microbiol. 2016;7:162. doi: 10.3389/fmicb.2016.00162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Briddon RW, Stanley J. Subviral agents associated with plant single-stranded DNA viruses. Virology. 2006;344:198–210. doi: 10.1016/j.virol.2005.09.042. [DOI] [PubMed] [Google Scholar]
- 5.Kulkarni GS. Mosaic and other related disease of crops in the Bombay presidency. Poona Agric Coll Mag. 1924;16:6–12. [Google Scholar]
- 6.Uppal BN, Varma PM, Capoor SP. Yellow mosaic of Bhendi. Curr Sci. 1940;9:2278. [Google Scholar]
- 7.Singh SJ, Dutta OP. Enation leaf curl of okra-a new virus disease. Indian J Virol. 1986;2:114–117. [Google Scholar]
- 8.Singh SJ. Etiology and epidemiology of whitefly-transmitted virus diseases of okra in India. Plant Dis Res. 1990;5:64–70. [Google Scholar]
- 9.Biswas KK, Palchoudhury S, Shukla P, Godara S, Balram N, Bhattacharyya UK, Makeshkumar T. DNA-A sequences of whitefly transmitted begomovirus infecting okra (Abelmoschus esculentus) in India are extensively diverse. Indian Phytopathol. 2018;71:249–256. doi: 10.1007/s42360-018-0042-y. [DOI] [Google Scholar]
- 10.Jose J, Usha R. Bhendi yellow vein Mosaic disease in india is caused by association of a DNA β satellite with a begomovirus. Virology. 2003;305:310–317. doi: 10.1006/viro.2002.1768. [DOI] [PubMed] [Google Scholar]
- 11.Venkataravanappa V, Reddy CNL, Jalali S, Krishna Reddy MK. Molecular characterization of distinct bipartite begomovirus infecting bhendi (Abelmoschus esculentus L.) in India. Virus Genes. 2012;44:522–535. doi: 10.1007/s11262-012-0732-y. [DOI] [PubMed] [Google Scholar]
- 12.Venkataravanappa V, Reddy CNL, Jalali S, Reddy MK. Molecular characterization of a new species of begomovirus associated with yellow vein mosaic of bhendi (Okra) in Bhubaneswar, India. Eur J Plant Pathol. 2013;136:811–822. doi: 10.1007/s10658-013-0209-4. [DOI] [Google Scholar]
- 13.Venkataravanappa V, Reddy CNL, Devraju A, Jalali S, Reddy MK. Association of a recombinant Cotton leaf curl Bangalore virus with yellow vein and leaf curl disease of okra in India. Ind. J. Virol. 2013;24:188–198. doi: 10.1007/s13337-013-0141-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chandran SA, Packialakshmi RM, Subhalakshmi K, Prakash C, Poovannan K, Prabu AN, Gopal P, Usha R. First report of an alphasatellite associated with Okra enation leaf curl virus. Virus Genes. 2013;46:585–587. doi: 10.1007/s11262-013-0898-y. [DOI] [PubMed] [Google Scholar]
- 15.Venkataravanappa V, Reddy CNL, Jalali S, Reddy MK. Association of tomato leaf curl New Delhi virus DNA-B with bhendi yellow vein mosaic virus in okra showing yellow vein mosaic disease symptoms. Acta Virol. 2015;59:125–139. doi: 10.4149/av_2015_02_125. [DOI] [PubMed] [Google Scholar]
- 16.Chatterjee A, Roy A, Padmalatha KV, Malathi VG, Ghosh SK. Occurrence of a Begomovirus with yellow vein mosaic disease of mesta (Hibiscus cannabinus and Hibiscus sabdariffa) Australas Plant Pathol. 2005;34:609–610. doi: 10.1071/AP05062. [DOI] [Google Scholar]
- 17.Chatterjee A, Ghosh SK. A new monopartite begomovirus isolated from Hibiscus cannabinus L. in India. Arch Virol. 2007;2007(152):2113–2118. doi: 10.1007/s00705-007-1029-7. [DOI] [PubMed] [Google Scholar]
- 18.Das S, Roy A, Ghosh R, Paul S, Acharyya S, Ghosh SK. Sequence variability and phylogenetic relationship of betasatellite isolates associated with yellow vein mosaic disease of mesta in India. Virus Genes. 2008;37:414–424. doi: 10.1007/s11262-008-0287-0. [DOI] [PubMed] [Google Scholar]
- 19.Das S, Ghosh R, Paul S, Roy A, Ghosh SK. Complete nucleotide sequence of a monopartite begomovirus associated with yellow vein mosaic disease of mesta from north India. Arch Virol. 2008;153:1791–1796. doi: 10.1007/s00705-008-0181-z. [DOI] [PubMed] [Google Scholar]
- 20.Roy A, Acharyya S, Das S, Ghosh R, Paul S, Srivastava RK, Ghosh SK. Distribution, epidemiology and molecular variability of the begomovirus complexes associated with yellow vein mosaic disease of mesta in India. Virus Genes. 2009;141(237):246. doi: 10.1016/j.virusres.2008.11.022. [DOI] [PubMed] [Google Scholar]
- 21.Rishishwar R, Mazumdar B, Dasgupta I. Diverse and recombinant begomoviruses and various satellites are associated with Bhendi yellow vein mosaic disease of okra in India. J Plant Biochem Biotechnol. 2015;24(4):470–475. doi: 10.1007/s13562-015-0305-y. [DOI] [Google Scholar]
- 22.Kumar R, Esakky R, Palicherla SR. The emergence of okra enation leaf curl virus—an important begomovirus infecting okra in several states across India. Arch Phytopathol Plant Prot. 2019;52:234–238. doi: 10.1080/03235408.2019.1606383. [DOI] [Google Scholar]
- 23.Muhire BM, Varsani A, Martin DP. SDT: a virus classification tool based on pairwise sequence alignment and identity calculation. PLoS ONE. 2014;9:e108277. doi: 10.1371/journal.pone.0108277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ghosh R, Paul S, Ghosh SK, Roy A. An improved method of DNA isolation suitable for DNA- based detection of begomoviruses from jute and other mucilaginous plants. J Virol Methods. 2009;159:34–39. doi: 10.1016/j.jviromet.2009.02.020. [DOI] [PubMed] [Google Scholar]
- 25.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. JMB. 1990;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 26.Haible D, Kober S, Jeske H. Rolling circle amplification revolutionizes diagonis and genomics of geminiviruses. JMC. 2006;135(1):9–16. doi: 10.1016/j.jviromet.2006.01.017. [DOI] [PubMed] [Google Scholar]
- 27.Tamura K, Stecher G, Peterson D, Filipski A, Kumar S. MEGA6: molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol. 2013;30(12):2725–2729. doi: 10.1093/molbev/mst197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Martin DP, Lemey P, Lott M, Moulton V, Posada D, Lefeuvre P. RDP3: a flexible and fast computer program for analyzing recombination. Bioinformatics. 2010;26:2462–2463. doi: 10.1093/bioinformatics/btq467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Brown JK, Zerbini FM, Navas-Castillo J, Moriones E, Ramos-Sobrinho R, Silva JC, Fiallo-Olivé E, Briddon RW, Hernández-Zepeda C, Idris A, Malathi VG, Martin DP, Rivera-Bustamante R, Ueda S, Varsani A. Revision of Begomovirus taxonomy based on pairwise sequence comparisons. Arch Virol. 2015;160:1593–1619. doi: 10.1007/s00705-015-2398-y. [DOI] [PubMed] [Google Scholar]
- 30.Briddon RW, Brown JK, Moriones E, Stanley J, Zerbini M, Zhou X, Fauquet CM. Recommendations for the classification and nomenclature of the DNA-b satellites of begomoviruses. Arch Virol. 2008;153:763–781. doi: 10.1007/s00705-007-0013-6. [DOI] [PubMed] [Google Scholar]
- 31.Venkataravanappa V, Reddy CNL, Devaraju A, Jalali S, Reddy MK. Association of a recombinant Cotton leaf curl Bangalore virus with yellow vein and leaf curl disease of okra in India. Indian J Virol. 2013;24(2):188–198. doi: 10.1007/s13337-013-0141-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kumar RV, Prasanna HC, Singh AK, Ragunathan D. Garg GK and Chakraborty S (2017) Molecular genetic analysis and evolution of begomoviruses and betasatellites causing yellow mosaic disease of bhendi. Virus Genes. 2017;53:275–285. doi: 10.1007/s11262-016-1414-y. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The partial sequences generated in this study are provided with accession numbers MK962353, MK962354, MK962355, MK962356, MK962357, MK962358, MK962359, MK962360, MK962361, MK962362, MK962363, MK962364, MK962365, MK962366, MK962367, MK962368, MK962369, MK962370, MK962371, MK962372, MK962373 from NCBI while the full length sequences generated in this study are available in the GenBank under the accession numbers MK069433, MK069434, MK069435, MK069436, MK069437, MK084765, MK084766; MK084767, MK084768, MK145076, MK145077, MK145078 and MK145075.