Abstract
Background.
In kidney transplant recipients, gastrointestinal (GI) pathogens in feces are only evaluated during diarrheal episodes. Little is known about the prevalence of GI pathogens in asymptomatic individuals in this population.
Methods.
We recruited 142 kidney transplant recipients who provided a non-diarrheal fecal sample within the first 10 days after transplantation. The specimens were evaluated for GI pathogens using the BioFire® FilmArray® GI Panel (BioFire® Diagnostics, LLC; Salt Lake City, UT), which tests for 22 pathogens. The fecal microbiome was also characterized using 16S rRNA gene sequencing of the V4-V5 hypervariable region. We evaluated whether detection of Clostridioides difficile and other GI pathogens was associated with post-transplant diarrhea within the first 3 months after transplantation.
Results.
Among the 142 subjects, a potential pathogen was detected in 43 (30%) using the GI Panel. The most common organisms detected were C. difficile (n=24, 17%), enteropathogenic Escherichia coli (n=8, 6%), and norovirus (n=5, 4%). Detection of a pathogen on the GI panel or detection of C. difficile alone was not associated with future post-transplant diarrhea (P > 0.05). The estimated number of gut bacterial species was significantly lower in subjects colonized with C. difficile than those not colonized with a GI pathogen (P = 0.01)
Conclusion.
Colonization with GI pathogens, particularly C. difficile, is common at the time of kidney transplantation but does not predict subsequent diarrhea. Detection of C. difficile carriage was associated with decreased microbial diversity and may be a biomarker of gut dysbiosis.
Keywords: C. difficile, Colonization, Gastrointestinal Panel, Kidney Transplantation, Microbiome, Diarrhea
Introduction
Post-transplant diarrhea is a common complication after kidney transplantation, affecting approximately 20% of patients in the first year after transplantation 1. The rate attributable to infectious etiologies in this population is variable, ranging from 25 to 80%, with norovirus and Clostridioides difficile being most common 2,3. In another study focused on solid organ transplant recipients, predominately kidney transplant recipients, 25 to 40% of etiologies were identified 3. Norovirus has been reported to account for approximately 25% of post-transplant diarrhea cases and C. difficile infection occurs in 2 to 7% of kidney transplant recipients 4–6.
No study to date has evaluated the frequency of asymptomatic gastrointestinal (GI) pathogen carriage in kidney transplant recipients or the impact of colonization on the risk of future development of diarrhea. If colonization frequently precedes clinically significant infection, then strategies of screening for colonization followed by treatment of colonized patients may warrant investigation to prevent post-transplant diarrhea. Understanding the prevalence of asymptomatic colonization with each diarrheal pathogen in this population may also assist clinicians with interpreting the significance of positive test results in patients with other apparent causes of diarrhea. Furthermore, detection of a high prevalence of highly transmissible GI pathogens, such as norovirus, may have important infection control implications.
In this study, we evaluated whether detection of C. difficile and other GI pathogens was associated with post-transplant diarrhea in the first 3 months after transplantation and whether the carriage state was associated with decreased gut abundance of commensal bacteria. We utilized a multiplex PCR assay for GI pathogens (BioFire® FilmArray® GI Panel; BioFire Diagnostics, LLC; Salt Lake City, UT) which allows for the simultaneous detection of 22 GI pathogens, including C. difficile and 12 other bacteria, 5 viruses, and 4 protozoa, and evaluated whether detection of GI pathogens is associated with future development of post-transplant diarrhea. We also characterized the gut microbiota in these 142 subjects and determined whether detection of C. difficile and other GI pathogens on the GI Panel is associated with gut microbiota dysbiosis.
Patients and Methods
Kidney Transplant Cohort and Specimen Collection
Between August 2015 and November 2016, 280 kidney transplant recipients were recruited at the time of kidney transplantation for fecal specimen collection at NewYork Presbyterian Hospital – Weill Cornell Medical Center and 153 subjects provided a fecal specimen during the first 10 days after transplantation. Eleven subjects were excluded for having diarrhea during this collection, for having invalid BioFire® FilmArray® GI Panel result (an invalid result was returned initially and after repeat testing), or for having insufficient material for GI panel testing. Among the 142 subjects, 3 subjects had dual organ transplantation (2 kidney-pancreas transplantations and 1 liver-kidney transplantation). As an inpatient, subjects were given a Fisherbrand™ commode specimen collection kit (Fisher Scientific, Hampton, NH) to collect the specimen and the specimen was stored at 4°C. As an outpatient, subjects self-collected the fecal specimens with the commode specimen collection kit and stored the specimen with ice packs or in a refrigerator. Among the 142 fecal specimens, 126 (89%) were collected in hospital and 16 (11%) were collected at home. In addition to the 142 initial fecal specimens, we also included 10 diarrheal specimens that were collected from unique subjects within the cohort to complement analyses. Specimens were aliquoted into approximately 200 milligram aliquots and stored at −80°C for further analysis. The time from production of the fecal specimen to storage at −80°C was within 24 hours for 105 (74%), within 48 hours for 34 (24%), and within 72 hours for 3 (2%). Demographic and transplant-related data, including development of diarrhea, antibiotic prophylaxis (including preoperative antibiotic prophylaxis as well as Pneumocystis jirovecii prophylaxis), and diagnoses of GI pathogen infections were collected for all subjects. The Institutional Review Board at Weill Cornell Medicine (Protocol number: 1207012703) approved this study and each participant provided written informed consent.
Gastrointestinal Panel Testing
An aliquot of a frozen fecal specimen was thawed and combined with Para-Pak C&S media (Meridian Bioscience, Inc., Cincinnati, OH) in a ratio of 1 mg of feces to 3 μL of Para-Pak C&S 7. The mixture was rigorously vortexed to resuspend the feces. The aliquot was analyzed using the BioFire® FilmArray® GI Panel (v2.1) (BioFire Diagnostics, LLC) per the manufacturer’s instructions. This panel utilizes a multiplexed PCR approach to simultaneously detect nucleic acids from the following pathogens: Campylobacter species (C. jejuni, C. coli, and C. upsaliensis), Clostridioides (formerly Clostridium) difficile (toxin A/B), Plesiomonas shigelloides, Salmonella, Yersinia enterocolitica, Vibrio species (V. parahaemolyticus, V. vulnificus, and V. cholerae), enteroaggregative Escherichia coli (EAEC), enteropathogenic E. coli (EPEC), enterotoxigenic E. coli (ETEC) lt/st, shiga-like toxin-producing E. coli (STEC) stx1/stx2, including E. coli 0157, Shigella/enteroinvasive E. coli (EIEC), Cryptosporidium, Cyclospora cayetanensis, Entamoeba histolytica, Giardia lamblia, adenovirus F40/41, astrovirus, norovirus GI/GII, rotavirus A, and sapovirus (I, II, IV and V) 8.
Each Para-Pak C&S media vial used to resuspend fecal specimens was tested on the GI Panel to ensure that the Para-Pak C&S media was not contaminated with GI pathogen nucleic acids. All media tested negative for the analytes detected by the BioFire® FilmArray® GI Panel. Each shipment of BioFire® FilmArray® GI Panels was tested with quality control material (FilmArray® GI Control Panel M238, Molecular Maine Quality Controls, Inc., Saco, ME) that spans the 22 pathogens covered by the BioFire® FilmArray® GI Panel. All quality control material tested as expected.
C. difficile toxin A and B immunoassay was performed using the Immunocard® Toxins A&B test (Meridian Biosciences, Inc) per the manufacturer’s protocol. An external positive and negative control was performed each day of testing.
Diarrhea Classification
Post-transplant diarrhea was defined as a complaint of diarrhea and by the World Health Organization classification of 3 or more unusually loose/liquid bowel movements per day 9. Workup for infectious etiologies for post-transplant diarrhea by the clinician was not performed in all reported cases. Potential standard of care testing included the Xpert® C. difficile/Epi PCR assay (Cepheid, Inc., Sunnyvale, CA), bacterial fecal culture, examination for ova and parasites, norovirus RT-PCR (Viracor Eurofins Clinical Diagnostics, Lee’s Summit, MO), and/or BioFire® FilmArray® GI Panel testing (which was initiated in-house at our institution in May 2016).
DNA Extraction for 16S rRNA Gene Sequencing
DNA extraction has been previously described10. Briefly, an aliquot of a fecal specimen was suspended in a solution containing 500 μL of extraction buffer (200 mM Tris-HCl [pH 8.0]; 200 mM NaCl; 20 mM EDTA), 500 μL of phenol:chloroform:isoamyl alcohol (25:24:1), 200 μL of 20% (w/v) sodium dodecyl sulphate, and 500 μL of 0.1-mm-diameter zirconia/silica beads (BioSpec Products, Inc., Bartlesville, OK). Mechanical disruption with a bead beater (BioSpec Products, Inc.) was performed for 2 minutes. Two rounds of phenol:chloroform:isoamyl alcohol extraction were performed, and DNA was precipitated with ethanol and resuspended in 50 μL of 1 × TE (10 mM Tris-HCl [pH 8.0]; 1 mM EDTA) buffer with 100μg/mL RNase. Additional purification was performed using the QIAamp mini spin columns according to the manufacturer’s protocol (Qiagen, Inc., Germantown, MD).
16S rRNA Gene Amplification, Sequencing, and Bioinformatics Analyses
Duplicate 50 μL PCR reactions were performed, each containing 50 ng of purified DNA, 0.2 mM dNTPs, 2.5 U Platinum Taq DNA polymerase, 1.5 mM MgCl2, 2.5 μL of 10 × PCR buffer, and 0.5 μM of each primer designed to amplify the V4-V5: 563F (5’-nnnnnnnn-NNNNNNNNNNNN-AYTGGGYDTAAAGNG-3’) and 926R (5′- nnnnnnnn-NNNNNNNNNNNN-CCGTCAATTYHTTTRAGT-3′). A unique 12-base Golay barcode (denoted by the “N”s) precedes primers for sample identification 11, and 1 to 8 additional nucleotides (denoted by the “n”s) precedes the barcode to offset primer sequencing. Conditions for cycling were 94°C for 3 minutes, followed by 27 cycles of 94°C for 50 seconds, 51°C for 30 seconds, and 72°C for 1 minute with 72°C for 5 min for the elongation step. For each specimen, replicate PCR products were pooled and were purified using the Qiaquick PCR Purification Kit (Qiagen, Inc.). Illumina barcodes and adaptors were ligated on using the Illumina TruSeq Sample Preparation protocol. The products were sequenced on an Illumina Miseq platform (Illumina, Inc., San Diego, CA) (paired end 250 × 250 base pair) using Illumina recommended procedures.
The 16S rRNA gene paired-end reads were merged and demultiplexed. The reads were analyzing with the UPARSE 12 pipeline to: (a) perform error filtering using maximum expected error 13, (b) group sequences into operational taxonomic units (OTUs) (97% distance-based similarity), and (c) identify and remove potential chimeric sequences. Taxonomic assignment was performed using a Python script incorporating nucleotide BLAST 14, with NCBI RefSeq 15 as a reference training set with a minimum E-value threshold of 1 × 10−10 for assignments. Microbial diversity and estimated number of bacterial species were estimated using Shannon diversity index and Chao1 index, respectively.
Statistical Analyses
The distribution of continuous variables between two groups was compared using the Wilcoxon rank sum test and between three groups using the Kruskal-Wallis test. Analyses involving multiple comparisons utilized the Benjamini-Hochberg correction. The distribution of categorical variables was compared using the Fisher’s exact test. All analyses were performed using R 3.3.1 and RStudio 0.99.902.
Results
Kidney Transplant Cohort and Post-Transplant Diarrhea
One hundred and fifty-three kidney transplant recipients provided a fecal specimen at the time of transplantation which refers to a fecal specimen within the first 10 days after kidney transplantation. Eleven subjects were excluded: 6 had specimens were associated with diarrhea at the time of collection, 2 had specimens that failed BioFire® FilmArray® GI Panel testing (generating an invalid result after testing aliquots twice), and 3 had specimens that had insufficient material for testing, leaving 142 non-diarrheal fecal specimens from unique subjects in the final cohort.
The median time from transplant to fecal specimen collection in the final cohort was 3 days with an interquartile range of 2 to 4 days. Among the 142 kidney transplant recipients, 47 (33%) developed post-transplant diarrhea in the first 3 months after transplantation. Of note, 5 subjects did not reach the 3 month time point after transplantation either because of death or were lost to follow up. Among these 5 subjects, 2 developed post-transplant diarrhea and 3 did not have post-transplant diarrhea and were grouped as if they never developed post-transplant diarrhea. Demographic and transplant characteristics are shown in Table 1. Female gender (P = 0.01), deceased donor transplantation (P = 0.01), and delayed graft function (P = 0.002) were significantly associated with the Diarrhea Group. Because clinical use of the GI Panel was not routinely performed during the study, we were unable to assess all infectious etiologies using the same methodology. However, C. difficile associated diarrhea was analyzed throughout the duration of the study using the Xpert® C. difficile/Epi PCR assay.
Table 1.
Characteristics of the Diarrhea Group and the No Diarrhea Group.
| Characteristic | Diarrhea Group†
(N=47) |
No Diarrhea Group† (N=95) |
P value‡ |
|---|---|---|---|
| N (%) / Median (IQR) | N (%) / Median (IQR) |
||
| Age | 53 (42–64) | 56 (46–66) | 0.50 |
| Female Gender | 27 (57%) | 32 (34%) | 0.01 |
| African American | 15 (32%) | 22 (23%) | 0.31 |
| Hispanic | 9 (19%) | 10 (11%) | 0.19 |
| History of Diabetes Mellitus | 14 (30%) | 30 (32%) | 0.99 |
| Deceased Donor Transplantation* | 20 (43%) | 19 (20%) | 0.01 |
| Delayed Graft Function | 13 (28%) | 7 (7%) | 0.002 |
| Anti-thymocyte Globulin Induction | 32 (68%) | 75 (79%) | 0.21 |
| Cefazolin Preoperative Prophylaxis (2 g) § | 39 (83%) | 78 (82%) | 0.99 |
| Trimethoprim/Sulfamethoxazole SS or DS Prophylaxis‖ | 45 (96%) | 91 (96%) | 0.99 |
| Mycophenolate Mofetil Dosage of 2000 mg / day# | 26 (55%) | 38 (40%) | 0.11 |
| Prednisone Maintenance Therapy (2.5 to 20 mg / day) | 16 (34%) | 23 (24%) | 0.24 |
Abbreviations:
IQR, interquartile range
SS, single-strength (trimethoprim 80 mg / sulfamethoxazole 400 mg)
DS, double-strength (trimethoprim 160 mg / sulfamethoxazole 800 mg)
Among the 142 subjects, 47 experienced post-transplant diarrhea within the first three months of transplantation and the remaining 95 participants remained diarrhea free during the 3 months following transplantation.
P value was calculated using Fisher’s exact test for categorical values and Wilcoxon rank sum test for continuous values.
Two deceased donor transplants were simultaneous kidney-pancreas transplants in the Diarrhea Group and one deceased donor transplant was a simultaneous liver-kidney transplant in the Diarrhea Group.
Cefazolin 4 g was utilized in 12 subjects and 3 g in 2 subjects. The remaining preoperative prophylaxis included mostly vancomycin 1 g.
The 1 liver-kidney transplant recipient received DS prophylaxis. The remaining prophylaxis included dapsone 100 mg / day and atovaquone 1500 mg / day.
The remaining antimetabolite prophylaxis included mostly mycophenolate mofetil dosage of 1000 mg/day
Prevalence of Gastrointestinal Pathogens in Asymptomatic Kidney Transplant Recipients
Forty-three (30%) of the 142 kidney transplant recipients were found to be colonized with at least one diarrheal pathogen at the time of transplantation using the GI Panel and 5 (4%) had ≥2 pathogens detected. Colonization with C. difficile was observed in 24 subjects; EPEC (n = 8 subjects); norovirus (n = 5); Y. enterocolitica (n=4); C. jejuni (n = 2); STEC (n = 1); Cryptosporidium (n = 1); ETEC (n = 1); EAEC (n = 1); Giardia lamblia (n = 1). Of the 24 subjects whose stool specimens were positive for C. difficile by the FilmArray® GI Panel, only 4 (17%) had detectable C. difficile toxins A&B by immunoassay. Demographic and transplant characteristics were grouped by BioFire® FilmArray® GI Panel status and there were no significant differences in characteristics among the groups (Table 2).
Table 2.
Characteristics in the C. difficile Positive Group, Other GI Panel Positive Group, and the GI Panel Negative Group.
| 5 | GI Panel | C. difficile | Other GI Panel | ||
|---|---|---|---|---|---|
| Negative Group | Positive Group | Positive Group | |||
| Characteristics | (N=99) | (N=24) | (N=19) | P value† | |
| N (%) / Median (IQR) | N (%) / Median (IQR) | N (%) / Median (IQR) | |||
| Age | 57 (47–65) | 48 (37–61) | 55 (46–69) | 0.16 | |
| Female Gender | 41 (41%) | 13 (54%) | 5 (26%) | 0.21 | |
| African American | 25 (25%) | 7 (29%) | 5 (26%) | 0.96 | |
| Hispanic | 13 (13%) | 5 (21%) | 1 (5%) | 0.32 | |
| History of Diabetes Mellitus | 30 (30%) | 9 (38%) | 5 (26%) | 0.75 | |
| Deceased Donor Transplantation‡ | 30 (30%) | 8 (33%) | 1 (5%) | 0.0505 | |
| Delayed Graft Function | 17 (17%) | 3 (13%) | 0 (0%) | 0.13 | |
| Anti-thymocyte Globulin Induction | 77 (78%) | 17 (71%) | 13 (68%) | 0.52 | |
| Cefazolin Preoperative Prophylaxis (2 g)* | 85 (86%) | 16 (67%) | 16 (84%) | 0.11 | |
| Trimethoprim/ Sulfamethoxazole SS or DS Prophylaxis§ | 96 (97%) | 23 (96%) | 17 (89%) | 0.24 | |
| Mycophenolate Mofetil Dosage of 2000 mg‖ | 42 (42%) | 12 (50%) | 10 (53%) | 0.61 | |
| Prednisone Maintenance Therapy (2.5 to 20 mg) | 25 (25%) | 8 (33%) | 6 (32%) | 0.64 | |
Abbreviations:
IQR, Interquartile Range
SS, single-strength (trimethoprim 80 mg / sulfamethoxazole 400 mg)
DS, double-strength (trimethoprim 160 mg / sulfamethoxazole 800 mg)
P value was calculated using Fisher’s exact test for categorical values and Kruskal Wallis test for continuous values.
Two deceased donor transplants were simultaneous kidney-pancreas transplants in the Diarrhea Group and one deceased donor transplant was simultaneous liver-kidney transplant in the Diarrhea Group.
Cefazolin 4 g was utilized in 12 subjects and 3 g in 2 subjects. The remaining preoperative prophylaxis included mostly vancomycin 1 g.
The 1 liver-kidney transplant recipient received DS prophylaxis. The remaining prophylaxis included dapsone 100 mg / day and atovaquone 1500 mg / day.
The remaining antimetabolite prophylaxis included mostly mycophenolate mofetil dosage of 1000 mg/day
Colonization with Gastrointestinal Pathogens and Development of Post-Transplant Diarrhea
We compared the incidence of diarrhea in the first 3 months after transplantation between the group that detected any pathogen on the GI BioFire® FilmArray® Panel (GI Panel Positive Group, n=43) and the group with no pathogens detected (GI Panel Negative Group, n=99). The incidence of diarrhea in the GI Panel Positive Group was not significantly different than that from the GI Panel Negative Group (28% versus 35%, P = 0.44). The GI Panel Positive Group was further divided into subjects whose fecal specimens were positive for C. difficile (C. difficile Positive Group) (n=24) and subjects whose fecal specimens were positive for a pathogen other than C. difficile (Other GI Panel Positive Group) (n=19). There was no increase in the incidence of diarrhea in the first 3 months after transplantation between the C. difficile Positive Group and the GI Panel Negative Group (42% versus 35%, P = 0.64). The incidence of diarrhea was significantly lower in the Other GI Panel Positive Group than that in the GI Panel Negative Group (11% versus 35%, P= 0.03). Two (8%) of the 24 subjects colonized with C. difficile at the time of transplantation developed diarrhea associated with C. difficile during the first 3 months after transplantation, compared to 6 (6%) of the 99 subjects from the GI Panel Negative Group (P = 0.65).
We also evaluated whether a positive result for a given GI pathogen on the FilmArray® GI Panel at the time of transplantation led to development of diarrhea by the same pathogen. Among the 43 subjects with a positive GI panel result, 31 subjects did not have post-transplant diarrhea and thus were not tested clinically. Among the remaining 12 subjects who had post-transplant diarrhea, only 2 were tested clinically with the FilmArray® GI Panel. However, we had 10 diarrheal specimens banked for 6 of the 12 subjects with post-transplant diarrhea. As such, we performed testing on these additional 10 specimens as part of this study. For these 6 subjects for whom we had archived fecal specimens, at the time of transplantation, 3 subjects had a positive C. difficile result; 1 subject was positive for both C. difficile and Campylobacter; 1 subject had a positive norovirus result; and 1 subject had a positive EAEC result. Three of the 6 subjects had the same GI pathogen detected at the time of transplantation and at time of diarrhea. One subject who had a positive C. difficile result at the time of transplantation had a subsequent C. difficile positive result determined using the FilmArray® GI Panel; another subject who had a positive C. difficile result at the time of transplantation had a subsequent positive C. difficile result determined using the clinical Xpert® C. difficile/Epi PCR assay; and the third subject who had a positive norovirus result at the time of transplantation had a subsequent positive norovirus result determined using the FilmArray® GI Panel.
Microbial Diversity and Beta Diversity in the Fecal Specimens
Shannon diversity index was not significantly different between the GI Panel Positive Group and the GI Panel Negative Group (median 3.1 versus median 3.2, respectively, P = 0.46) (Fig. 1A), between the C. difficile Positive Group and the GI Panel Negative Group (median 2.9 versus median 3.2, respectively, P = 0.20) (Fig. 1B), or between the Other GI Panel Positive Group and the GI Panel Negative Group (median 3.2 versus 3.2, respectively, P = 0.80) (Fig. 1C). However, the Chao1 (estimated bacterial species) was significantly lower in the GI Panel Positive Group than in the GI Panel Negative Group (median 193 versus median 234, respectively, P= 0.03) (Fig. 1D). This decrease was observed only in the C. difficile Positive Group, where the number of bacterial species was lower than observed in the GI Panel Negative Group (median 164 versus median 234, respectively, P= 0.01) (Fig. 1E). The number of bacterial species was not significantly different in the Other GI Panel Positive Group compared to the GI Panel Negative Group (median 218 versus median 234, respectively, P= 0.42) (Fig. 1F).
Figure 1. Microbial Diversity by GI Panel Status.
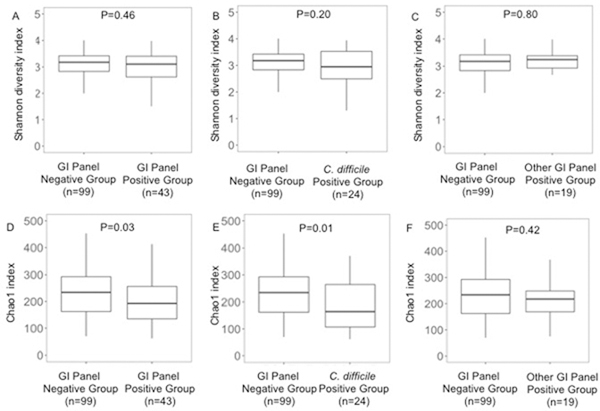
Box and whisker plots are plotted with the median, 25% and 75% representing the edges of the boxplot; and 1.5 times the median representing the whiskers of the boxplot. P values were calculated using the Wilcoxon rank sum test. Panel A. Box and whisker plot of Shannon diversity index in the fecal specimens from the GI Panel Negative Group and the GI Panel Positive Group. Panel B. Box and whisker plot of Shannon diversity index in the fecal specimens from the GI Panel Negative Group and the C. difficile Positive Group. Panel C. Box and whisker plot of Shannon diversity index in the fecal specimens from the GI Panel Negative Group and the Other GI Panel Positive Group. Panel D. Box and whisker plot of Chao1 in the fecal specimens from the GI Panel Negative Group and the GI Panel Positive Group. Panel E. Box and whisker plot of Chao1 in the fecal specimens from the GI Panel Negative Group and the C. difficile Positive Group. Panel F. Box and whisker plot of Chao1 in the fecal specimens from the GI Panel Negative Group and the Other GI Panel Positive Group.
Discussion
This study provides insight into the detection of GI pathogens at the time of kidney transplantation as well as the microbiota status of those individuals colonized with GI pathogens. To the best of our knowledge, this is the first study to utilize a multiplex PCR approach to assess for common GI pathogen colonization in non-diarrheal fecal specimens in kidney transplant recipients at the time of kidney transplantation. We report a significant proportion of kidney transplant recipients have asymptomatic detection of GI pathogens, particularly C. difficile, at the time of transplantation and that kidney transplant recipients with C. difficile carriage have decreased microbial diversity as measured by Chao1 index.
In this study, we did not find a significant relationship between detection of a GI pathogen at the time of transplant and development of post-transplant diarrhea. Because many of the transplant recipients in this study predated the introduction of the FilmArray® GI Panel at our institution for routine clinical use, we were not always able to assess whether colonized subjects developed clinically significant diarrhea due to their colonizing organism after transplantation. Nonetheless, we were able to assess those subjects with C. difficile diarrhea and did not find a significant association between C. difficile detection at the time of transplantation and subsequent development of C. difficile diarrhea. However, we had only 8 cases of C. difficile associated diarrhea. Interestingly, the Other GI Panel Positive Group had a significantly lower incidence of post-transplant diarrhea than the GI Panel Negative Group. The Other GI Panel Positive Group had a lower percentage of women, of deceased donor transplantations, and of delayed graft function; all factors that were positively associated with post-transplant diarrhea (Table 1). Therefore, this unexpected negative association between the Other GI Panel Positive Group and lower incidence of diarrhea might be explained by one of the variables listed above.
Our data suggest that colonization with GI pathogens is common but is not associated with diarrhea development, which highlights the importance of judiciously testing kidney transplant recipients with post-transplant diarrhea for infectious etiologies. Importantly, only a minority of subjects who were colonized with a GI pathogen at the time of transplantation developed post-transplant diarrhea, and of those that did, very few subjects had the same pathogen at time of transplantation and at time of post-transplant diarrhea. In addition, we found that C. difficile toxins A&B were not detected by the toxin immunoassay in the majority of cases. Given the established view that C. difficile infection is a toxin-mediated disease16, these data suggest that the majority of subjects did not have active C. difficile infection at the time of transplantation. Furthermore, the high prevalence of asymptomatic colonization with these GI pathogens also raises the possibility of patient-to-patient transmission on an inpatient transplant unit or in an outpatient transplant clinic and thus raises the importance of adherence to hand hygiene and standard infection prevention precautions among providers of transplant recipients. However, at least in the case of C. difficile diarrhea, Eyre and colleagues reported that transmission of C. difficile from symptomatic patients only accounted for approximately a third of cases, suggesting many cases emerge from genetically diverse sources.17
Our study has several limitations. We used frozen rather than fresh feces to assess for the presence of nucleic acids of various GI pathogens and for the presence of C. difficile toxins A and B. Although our samples underwent only one freeze-thaw cycle, it is possible that the prevalence of detection could have been higher using fresh fecal specimens. In addition, only a fraction of post-transplant diarrheal episodes had any specimen collected and thus fecal diagnostic testing performed as part of routine clinical care. Therefore the infectious etiologies of diarrhea within the first three months after transplant, may be underestimated. Also, the GI Panel was not used clinically for part of the study. Thus, the full impact of asymptomatic colonization with GI pathogens on the risk of developing post-transplant diarrhea due to the colonizing organism could not be assessed. Nevertheless, we observed that a minority of subjects with asymptomatic colonization with GI pathogens developed post-transplant diarrhea and few subjects with asymptomatic colonization developed post-transplant diarrhea with the same GI pathogen detected at the time of transplantation. However, we were limited by the number of cases evaluated. While we found C. difficile colonization was not associated with future development of C. difficile associated diarrhea, we were limited due to the paucity of C. difficile diarrhea cases. Furthermore, we also employed a different C. difficile detection methodology for clinical use (Xpert® C. difficile/Epi) and assessment of C. difficile colonization status in asymptomatic individuals (FilmArray® GI Panel), which may explain the discordance. Another limitation is that we did not recover or type C. difficile isolates and were unable to assess potential C. difficile transmission rates in our study. Also, pre-transplant fecal specimens were not analyzed and use of surgical preoperative prophylaxis or P. jiroveci prophylaxis may have influenced the microbiota post-transplantation although these variables were not significantly different between the Diarrhea Group and no Diarrhea Group. Finally, we retrospectively reviewed the charts of the subjects so we do not have a complete catalogue of the episodes of post-transplant diarrhea, which may have been underestimated in our study.
Despite these limitations, the current study is the first to evaluate the prevalence of C. difficile and other diarrheal pathogens at the time of kidney transplantation in a large cohort. Our study reports that a significant portion of kidney transplant recipients have C. difficile carriage and that the gut microbiome in these asymptomatic individuals have an altered microbiota with decreased microbial diversity as measured by the Chao1 index. Future studies investigating C. difficile detection in a larger cohort of kidney transplant recipients are warranted to further assess whether C. difficile carriage and its associated gut dysbiosis leads to an increased risk for C. difficile diarrhea or other adverse outcomes.
Acknowledgments
We thank Dr. Carey-Ann Burnham for her review of the manuscript. We thank BioFire® Diagnostics, LLC for providing an investigator-initiated research grant and the National Institutes of Allergy and Infectious Diseases for their research support via NIH grant K23 AI 124464 (J.R.L.) and NIH grant R37 AI 051652 (M.S.).
Funding
This research work was supported, in part, by an investigator-initiated research grant from BioFire Diagnostics, LLC; and by K23 AI 124464 from the National Institute of Allergy and Infectious Diseases to J.R.L and R37 AI 051652 from the National Institute of Allergy and Infectious Diseases to M.S.
L.F.W. receives research support from BioFire Diagnostics, LLC; M.J.S. receives research support from BioFire Diagnostics, LLC and Merck. E.G.P. has received speaker honoraria from Bristol Myers Squibb, Celgene, Seres Therapeutics, MedImmune, Novartis and Ferring Pharmaceuticals and is an inventor on patent application # WPO2015179437A1, entitled “Methods and compositions for reducing Clostridium difficile infection” and #WO2017091753A1, entitled “Methods and compositions for reducing vancomycin-resistant enterococci infection or colonization” and holds patents that receive royalties from Seres Therapeutics, Inc. J.R.L. receives research support from BioFire Diagnostics, LLC.
Abbreviations
- EAEC
enteroaggregative Escherichia coli
- EPEC
enteropathogenic E. coli
- ETEC
enterotoxigenic E. coli
- GI
gastrointestinal
- PCR
polymerase chain reaction
- SS
single-stranded
- STEC
shiga-like toxin-producing E. coli stx1/stx2 including E. coli 0157
Footnotes
Disclosures
References
- 1.Mycophenolate mofetil in renal transplantation: 3-year results from the placebo-controlled trial. European Mycophenolate Mofetil Cooperative Study Group. Transplantation. 1999;68(3):391–396. [PubMed] [Google Scholar]
- 2.Arslan H, Inci EK, Azap OK, Karakayali H, Torgay A, Haberal M. Etiologic agents of diarrhea in solid organ recipients. Transpl Infect Dis. 2007;9(4):270–275. [DOI] [PubMed] [Google Scholar]
- 3.Echenique IA, Penugonda S, Stosor V, Ison MG, Angarone MP Diagnostic yields in solid organ transplant recipients admitted with diarrhea. Clin Infect Dis. 2015;60(5):729–737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Paudel S, Zacharioudakis IM, Zervou FN, Ziakas PD, Mylonakis E. Prevalence of Clostridium difficile infection among solid organ transplant recipients: a meta-analysis of published studies. PLoS One. 2015;10(4):e0124483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ramos A, Ortiz J, Asensio A, et al. Risk Factors for Clostridium Difficile Diarrhea in Patients With Solid Organ Transplantation. Prog Transplant. 2016;26(3):231–237. [DOI] [PubMed] [Google Scholar]
- 6.Spinner ML, Stephany BR, Cerrato PM, Lam SW, Neuner EA, Patel KS Risk factors associated with Clostridium difficile infection in kidney transplant recipients. Transpl Infect Dis. 2018;20(4):e12918. [DOI] [PubMed] [Google Scholar]
- 7.Lee JR, Magruder M, Zhang L, et al. Gut microbiota dysbiosis and diarrhea in kidney transplant recipients. Am J Transplant. 2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Buss SN, Leber A, Chapin K, et al. Multicenter evaluation of the BioFire FilmArray gastrointestinal panel for etiologic diagnosis of infectious gastroenteritis. J Clin Microbiol. 2015;53(3):915–925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Organization WH. The treatment of diarrhoea: a manual for physicians and other senior health workers. 2005. [Google Scholar]
- 10.Lee JR, Muthukumar T, Dadhania D, et al. Gut microbial community structure and complications after kidney transplantation: a pilot study. Transplantation. 2014;98(7):697–705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Caporaso JG, Lauber CL, Walters WA, et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 2012;6(8):1621–1624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Edgar RC UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods. 2013;10(10):996–998. [DOI] [PubMed] [Google Scholar]
- 13.Edgar RC, Flyvbjerg H. Error filtering, pair assembly and error correction for next-generation sequencing reads. Bioinformatics. 2015;31(21):3476–3482. [DOI] [PubMed] [Google Scholar]
- 14.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ Basic local alignment search tool. J Mol Biol. 1990;215(3):403–410. [DOI] [PubMed] [Google Scholar]
- 15.Tatusova T, Ciufo S, Fedorov B, O’Neill K, Tolstoy I. RefSeq microbial genomes database: new representation and annotation strategy. Nucleic Acids Res. 2014;42(Database issue):D553–559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Polage CR, Gyorke CE, Kennedy MA, et al. Overdiagnosis of Clostridium difficile Infection in the Molecular Test Era. JAMA Intern Med. 2015;175(11):1792–1801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Eyre DW, Cule ML, Wilson DJ, et al. Diverse sources of C. difficile infection identified on whole-genome sequencing. N Engl J Med. 2013;369(13):1195–1205. [DOI] [PMC free article] [PubMed] [Google Scholar]


