Abstract
Background
Telocytes (TCs) have the capacity of cell–cell communication with adjacent cells within the tissue, contributing to tissue repair and recovery from injury. The present study aims at investigating the molecular mechanisms by which the TGFβ1-ITGB1-PI3K signal pathways regulate TC cycle and proliferation.
Methods
Gene expression of integrin (ITG) family were measured in mouse primary TCs to compare with other cells. TC proliferation, movement, cell cycle, and PI3K isoform protein genes were assayed in ITGB1-negative or positive mouse lung TCs treated with the inhibition of PI3Kp110α, PI3Kα/δ, PKCβ, or GSK3, followed by TGFβ1 treatment.
Results
We found the characters and interactions of ITG or PKC family member networks in primary mouse lung TCs, different from other cells in the lung tissue. The deletion of ITGB1 changed TCs sensitivity to treatment with multifunctional cytokines or signal pathway inhibitors. The compensatory mechanisms occur among TGFβ1-induced PI3Kp110α, PI3Kα/δ, PKCβ, or GSK3 when ITGB1 gene was deleted, leading to alterations of TC cell cycle and proliferation. Of those PI3K isoform protein genes, mRNA expression of PIK3CG altered with ITGB1-negative TC cycle and proliferation.
Conclusion
TCs have strong capacity of proliferation through the compensatory signaling mechanisms and contribute to the development of drug resistance due to alterations of TC sensitivity.
Keywords: Telocytes, Lung, PI3K, TGFβ, ITGB1
Introduction
Telocytes (TCs) play an important role in cell–cell communication with adjacent cells within the tissue via direct patterns or indirect ways, such as cellular junctions, production of multi-factors and extracellular vesicles [1, 2]. TCs maintain local tissue homeostasis and cooperate with stem cells for organ repair and regeneration in diseases, inflammation, and fibrosis [3, 4]. However, mechanisms by which bio-behaviors of TCs are regulated are still unclear. Our recent study demonstrated that the transforming growth factor beta-1 (TGFβ1) interacted with phosphoinositide 3-kinase (PI3K) and regulated the proliferation and cell cycle phases of TCs [5]. Class IA PI3K is a heterodimer composed with two types of subunits, including one regulatory subunit and three catalytic subunits (p110α, p110β, p110δ). PI3K isoform proteins, e.g. PI3Kα/δ/β, PI3K/mTOR, and PI3K p110δ, contribute to the interactions and network functions of genes or proteins during TCs proliferation, especially PI3Kα/δ/β. TGFβ1 up-regulated the expression of PIK3CA coding p110α and PIK3CB coding p110β, while down-regulated the expression of PIK3CD coding p110δ and PIK3CG coding p110-γ in lung TCs [6]. PI3K p110α is involved in tumor growth, hypoxia, metastasis, or cell communication by increasing the tight junction formation [7] and the activity of glycogen synthase kinase-3 beta (GSK-3β) to promote cyclin D1 expression [8]. The present study furthermore investigates potential mechanisms of the interaction between TGFβ1 and PI3K isoforms in the regulation of TCs bio-behaviors.
PI3K/protein kinase B AKT/GSK3β signaling pathway-activated cell proliferation depends upon the alternations of TGFβ signaling by binding to integrins (ITG) [9–11]. TCs have the strong capacity of proliferation and of cell–cell communication with adjacent cells within the tissue, contributing to tissue repair and recovery from injury [6, 12]. The present study aims at investigating the molecular mechanisms by which the TGFβ1- integrin beta1 (ITGB1)-PI3K signal pathways regulate TCs cycle and proliferation. Gene expression profiles and special network characteristics of ITG family members were investigated among murine pulmonary TCs on days 5 (TC 5) and 10 (TC 10), fibroblasts, mesenchymal stem cells, alveolar type II cells (ATII), airway basal cells, proximal airway cells (PACs), CD8+ T cells come from bronchial lymph nodes (CD8 T BL), and CD8+ T cells from lung (CD8 T LL), respectively, like other genes [13]. Mouse lung TC Line was applied for investigating the patterns of PI3K catalytic isoform proteins or GSK3 and the regulation of TGF-β1 in TCs bio-behaviors were defined in mouse lung TCs [6]. We furthermore demonstrated effects of ITGB1 in PI3K catalytic isoform proteins or GSK3β-regulated mRNA expression of PI3K isoforms and defined the interactions among ITGB1, PI3K, and GSK3β in TCs bio-behaviors.
Materials and methods
Framework of the current study
We first analyzed the special network characteristics of ITG family molecules in primary lung TCs harvested from mice, as compared with alveolar type II cells, mesenchymal stem cells, airway epithelial cells, lymphocytes, and fibroblasts. After then mouse lung TCs was applied for investigating the patterns of PI3K catalytic isoform proteins (e.g. PI3K/p110α, PI3Kα/δ), Protein Kinase Cβ (PKCβ), or GSK3 in TCs proliferation, movement, differentiation and death. TGF-β1-regulated PI3K catalytic isoform proteins activity in TCs proliferation were validated in TCs with or without ITGB1. Effects of ITGB1 in PI3K catalytic isoform proteins or GSK3-regulated mRNA expression of PI3K isoforms were evaluated to define the interactions among ITGB1, PI3K, and GSK3.
Network and molecular interactions of PKC family, GSK family or ITG family
Genomic probes were used for gene expression profiles analysis. 23,861 probes were contained in gene expression profiles of murine lung TCs, MSCs and Fbs, and 45,101 probes were included in ATII, ABCs, PACs, CD8 T-BL, and CD8 T-LL from GEO originally. Differentially expressed genes between two samples were identified through fold change filtering as reported previously [13]. Network and molecular interactions of PKC family, GSK family or ITG family were analyzed and figured according to the previous publication [14]. To reconstruct and show the state of gene network corresponding to each sample, we used differential network models (DEN) [14, 15] with the sample-specific network measurements [16]. The R package networkD3 (https://cran.r-project.org/web/packages/networkD3/index.html) was used to visualize the network structure of each sample, where the network nodes represent genes and network edges represent significant samples-specific edges or gene associations.
Cell culture
This study was approved by the Fudan University Ethical Committee for animal experiments and mice were provided by Animal Facility in Biomedical Research Center of Zhongshan Hospital, Fudan University. Female BABL/c mice aged 6–8 weeks were used and the isolation of lung primary TCs were practiced as previous study [11]. After identification, lung primary TCs were transfected with SV40 large and small T antigen to constructed TCs. The telopodes were recorded and the expression of ckit, CD34, vimentin and PDGFR-α were detected for the identification of TCs maintaining the specific morphology and markers expression of TCs from generation 5 to generation 50 [6]. TCs were cultured in Dulbecco’s modified Eagle’s medium/F12 (DMEM/F12, GIBCO; Thermo Fisher Scientific, Inc.) supplemented with 10% fetal calf serum (FBS; GIBCO; Thermo Fisher Scientific, Inc.), 100 UI/ml penicillin, 0.1 mg/ml streptomycin (Sigma-Aldrich, St. Louis, USA), and cultured in a incubator, with 5% CO2 in air, at 37 °C. Cells were passaged when cell density were 60–70%. The cells were washed with PBS, digest with 0.25% trypsin, collection in new tubes and centrifuged at 400 × g for 5 min at room temperature. Then cells were seeded in new culture flasks.
Lentivirus construction and infection
Lentivirus particles containing the ITGB 1 shRNA sequence were constructed. The oligo (listed 5′–3′) includes ITGB1 shRNA forward: 5′-CCACAGAAGTTTACATTAA-3′and reverse: 5′- TTAATGTAAAGTTCTGTGG-3′; negative control forward: 5′-TTCTCCGAACGTGTCACGT-3′ and reverse: 5′-ACGAGACACGTTCGGAGAA-3′. TCs were seeded on 6-well plates with a density of 104 cells/well and cultured for 24 h at 37 °C and replaced in fresh medium. Two μl of shRNA or negative control was added into each well and incubated for 12 h at 37 °C. Puromycin were added for screening. The cells were harvested for Real-time Quantitative PCR Detecting System (qPCR) analysis to determine the ITGB 1 mRNA expression level after being cultured for 96 h at 37 °C. ITGB1 knocked down TC cells were mentioned as TCITGB1− and TCs with ITGB1 were named as TCITGB1+.
Immunofluorescent staining
Triple immunofluorescent staining for CD34/Vimentin/PDGFRα was used as previously reported [17, 18]. In brief, TCs were cultured on glass bottom cell culture dishes with 20 mm diameter glass (NEST, Nanjing, China) and were fixed in 4% paraformaldehyde containing 0.05% Triton-X-100 for 20 min. Then washed the dishes three times wash with 1 × PBS and blocked in 5% Bovine serum albumin (BSA) for 1 h. After incubated overnight at 4 °C with mouse anti-CD34 antibody, goat anti-vimentin antibody or rat anti- PDGFRα antibody (1:200 dilution; Abcam, Cambridge, UK) diluted in 1% bovine serum albumin (BSA) in PBS, the dishes were washing in PBS for three times. Then, dishes were incubated with APC conjugated anti-mouse secondary antibodies, PE conjugated anti-rat secondary antibodies and FITC conjugated anti-goat secondary antibodies (1:200 dilution; Jackson ImmunoResearch, USA). The nuclear were marked by DAPI according to the manufacture (KeyGEN BioTECH, Nanjing, China). Cells were observed and recorded under Olympus FV3000 Confocal Laser Scanning Microscope (DSS Imagetech Pvt. Ltd, New Delhi, India).
Measurement of cell proliferation
TCITGB1+ or TCITGB1− were digested and cultured in 96-well plates with the density of 5 × 103 cells/well, 3–6 wells per group, followed by treatment with TGFβ1 at 0.5, 5.0, or 50 ng/ml, respectively, for 24 h to characterize TGFβ1-induced TCs proliferation. In order to evaluate roles of PI3K catalytic isoform proteins, 5 μM HS173 (PI3K p110α inhibitor, SelleckChem Co., Houston, USA), 5 μM GDC0941(PI3Kα/δ inhibitor, SelleckChem), 0.25 μM Enzastaurin (PKCβ inhibitor, SelleckChem) or 0.5 μM SB216763 (a potent and selective GSK-3 inhibitor for both GSK-3α and GSK-3β, SelleckChem)on TCITGB1+ or TCITGB1− with or without TGFβ1 for 48 h. Ten microliters of CCK-8 reagents (Dojindo Molecular Technologies, Inc., Maryland, USA) was added to every well and incubated for 0.5 h at 37° C, with 5% CO2. We determined the absorbance at 450 nm using SpectraMax M5 Microplate Reader (Molecular Devices Instruments Inc., Sunnyvale, California, USA).
RNA extraction and PCR
TC cells were seeded in 24-well plates with a density of 104 cells/well and cultured for 24 h at 37 °C. Cells were starving for 12–24 h before stimulation. 0.05 ng/ml, 0.5 ng/ml or 5 ng/ml TGFβ1 for 24 h or 48 h were used for stimulation, and the PI3K inhibitors were added 2 h before TGFβ1 treatment. Cells were washed thrice with cold PBS, and total RNA was isolated and transcribed into single-stranded cDNA using the 1st Strand cDNA Synthesis Kit (AMV, Roche Molecular Systems, Inc., Branchburg, USA) for Reverse Transcription-Polymerase Chain Reaction (RT-PCR) following the recommendations of the manufacturer. cDNA was synthesized from 1 µg of total RNA using PrimeScript® RT reagent Kit (Takara Bio Inc., Shiga, Japan). PCR was performed with 1 µl of cDNA using GoTaq polymerase (Promega Corporation, Madison, USA) for 25 cycles with specific primers for genes ITGB1 forward primer: GTCTTGGAACGGATTTGATGA and reverse: TTTGCTGGGGTTGTGCTAAT; GAPDH forward: CGGAGTCAACGGATTTGGTCGTAT and reverse: AGCCTTCTCCATGGTGGTGAAGAC. PCR reaction products were resolved through a 0.8% agarose gel in 1 × TAE and stained with Gelred (Biotium Inc., Newark, USA).
Detection of cell bio-behaviors
TCs were treated with 0.05 μM, 0.5 μM or 5 μM HS173 (PI3K/p110α inhibitor), GDC0941(PI3Kα/δ inhibitor), or 0.5 μM SB216763 (a potent and selective GSK-3 inhibitor for both GSK-3α and GSK-3β) or 0.025 μM, 0.25 μM or 2.5 μM Enzastaurin (PKCβ inhibitor)with or without TGFβ1 for 48 h. The bio-behaviors of TCs were recorded and analyzed using a Cell-IQ cell culturing platform (Chip-Man Technologies, Tampere, Finland), equipped with a phase-contrast microscope (Nikon CFI Achromat phase contrast objective with 10 magnification) and a camera [12]. These bio-behaviors, including total cell number, cell morphology, and cell movement, can be monitored and recorded as time-lapse data by this Cell-IQ system uses machine vision technology. Images were captured at about 30 min intervals for 1 week. Analysis was carried out with a freely distributed Image software (McMaster Biophotonics Facility, Hamilton, ON), using the Manual Tracking plugin created by Fabrice Cordelie´res (Institute Curie, Orsay, France).
Cell cycle assay
PI staining was used for cell cycle analysis of primary TCs as described in manufacturer. In brief, cells were collected and fixed in 75% ethanol at 4 °C for overnight. After centrifuged and washed, 0.5 ml PI/RNase Staining Buffer (BD Pharmingen, NJ, USA) was added to each tube for 15 min at room temperature. Samples were examined with a fluorescence-activated cell sorting flow cytometer, BD FACS Aria II (Becton, Dickinson and Company, NJ, USA) and DNA histograms were analyzed with Flowjo 7.6.1 software. Each test was repeated in triplicate.
Statistics
Data were analyzed using SPSS Statistics 20 (IBM, Chicago, USA). Statistical differences between two groups were compared by t test. Statistical differences among more than two groups were determined using ANOVA. Each point corresponds to the mean ± S.E.M. and p < 0.05 was considered significant.
Results
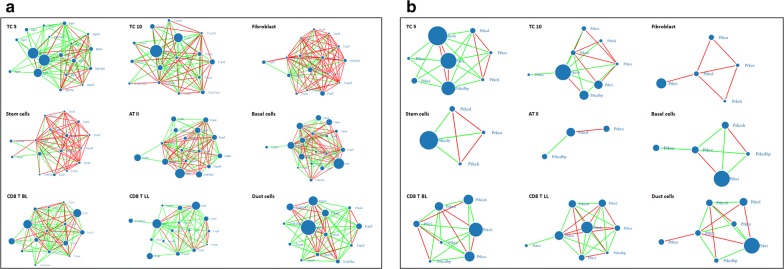
Figure 1 demonstrated the specificity of PKC, GSK3 and ITG family in TCs, network characters and molecular interactions of PKC family genes, GSK3 family genes or ITG family genes. Network elements and interactions of ITG subunit genes in mouse lung primary TCs were changed from day 5 to day 10 after the culture, and had positive communication (Fig. 1a). The network characters and interactions of PKC genes in TCs were different from those in fibroblasts, mesenchymal stem cells, ATII, airway basal cells, PACs, and varied significantly between TCs on days 5 and 10 (Fig. 1b). The network characters of GSK subunit genes had negative communication on day 5, and became positive links on day 10 (Additional file 1: Figure S1).
Fig. 1.
The characters of ITGB1, and PKC networks and interactions in mouse lung primary TCs cultured for 5 days (TC 5) and 10 days (TC 10) were compared with other tissue cells, e.g. fibroblasts, mesenchymal stem cells (Stem cells), alveolar type II cells (ATII), airway basal cells (Basal cells), proximal airway cells (Duct cells), CD8+ T cells come from bronchial lymph nodes (CD8 T BL), and CD8+ T cells from lung (CD8 T LL). a Network characters and molecular interactions of ITG family genes; b Network characters and molecular interactions of PKC family genes and its receptor family genes
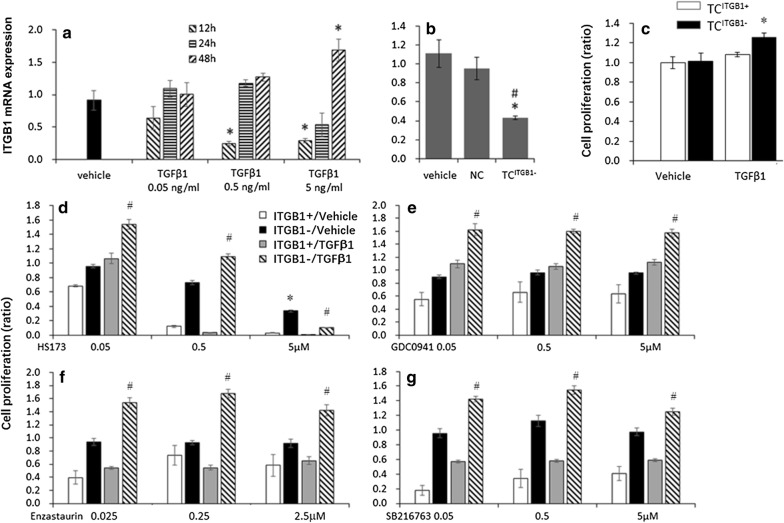
Mouse lung TCs were successfully isolated and constructed as a cell line as our previous reported [6]. These TCs have special characteristics including relatively small cell body and very long and thin Tps with lots of dilations (Fig. 4a). To further confirm that the cells were TCs, triple immunofluorescent staining for CD34/PDGFR-α/vimentin was used. We found that these cells were triple positive for CD34/PDGFR-α/vimentin (Fig. 2), indicating that these cells were TCs. To understand potential regulations between those factors in TCs, we firstly investigated effects of external TGFβ1 at different concentrations and time points on TCs with ITGB1 (TCITGB1+) and selected the concentration of TGFβ1 at 5 ng/ml for 48 h as the condition of further studies (Fig. 3a). ITGB1 knocked down TC cells (TCITGB1−) were generated by lentivirus construction and infection with ITGB1 shRNA and evaluated by expression of ITGB1 gene (Fig. 3b). TGFβ1 could induced the increased cell proliferations in TCITGB1− cells at 48 h, as compared to TCITGB1+ cells (Fig. 3c). HS173 inhibited cell proliferation significantly in a dose-dependent pattern in TCITGB1+ cells with or without TGFβ1 administration and had higher responses about 98% inhibitory rate at 5 μM, as compared to TCITGB1− cells (Fig. 3d). The proliferation of TGFβ1-pretreated TCITGB1− cells was higher than vehicle-pretreated ones after HS173 administration, especially at doses of 0.05 and 0.5 μM. The cell proliferation reduced from the low dose of GDC0941 (0.05 μM, Fig. 3e), Enzastaurin (0.025 μM, Fig. 3f), or SB216763 (0.05 μM, Fig. 3g), rather than in TCITGB1− cells. TGFβ1-pretreated TCITGB1− cells had significantly higher cell proliferation than vehicle-pretreated TCITGB1− cells, even higher than TCITGB1+ cells.
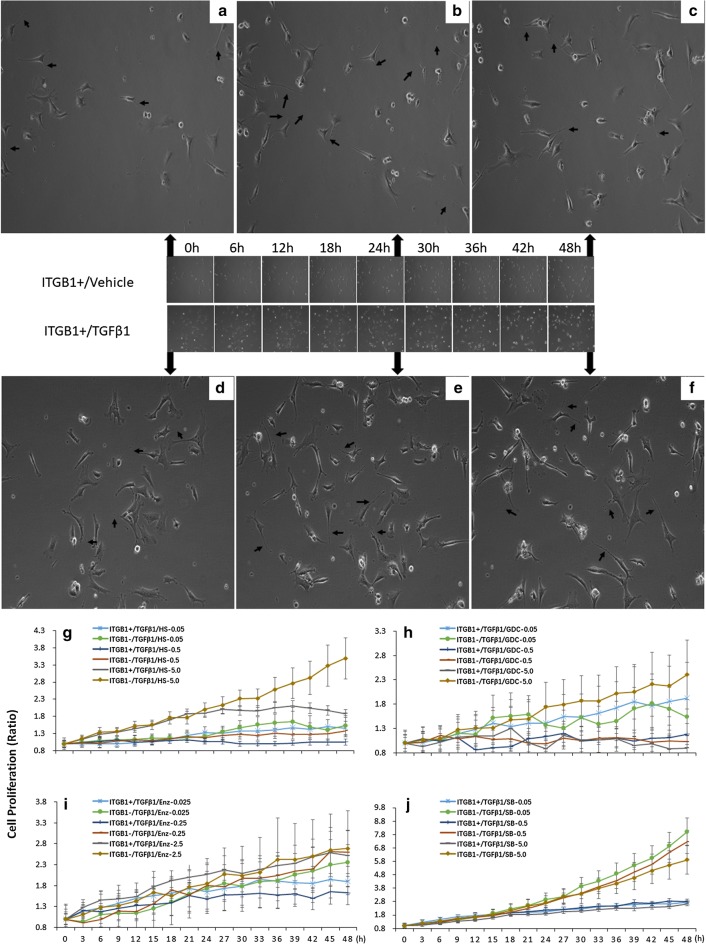
Fig. 4.
The effects of ITGB1on the proliferation curve of TC ITGB1+ treated with PI3K inhibitors and TGFβ1. a–c Representative photos of TCs cultured for 0 h, 24 h and 48 h captured by celliq, respectively. d–f representative photos of TCs treated within TGFβ1 for 0 h, 24 h and 48 h, respectively. Arrows show typical TCs with long and thin telopodes with dilations. Original magnification ×100. g Analysis of TC ITGB1+ or TCITGB1− proliferation with the treatment of HS173 and TGFβ1. h Analysis of TC ITGB1+ or TCITGB1− proliferation with the treatment of GDC0941 and TGFβ1. i Analysis of TC ITGB1+ or TCITGB1− proliferation with the treatment of Enzastaurin and TGFβ1. j Analysis of TC ITGB1+ or TCITGB1− proliferation with the treatment of SB216763 and TGFβ1, n = 6–8, **stand for p values less than 0.05, as compared with TC ITGB1+ treated with TGFβ1
Fig. 2.
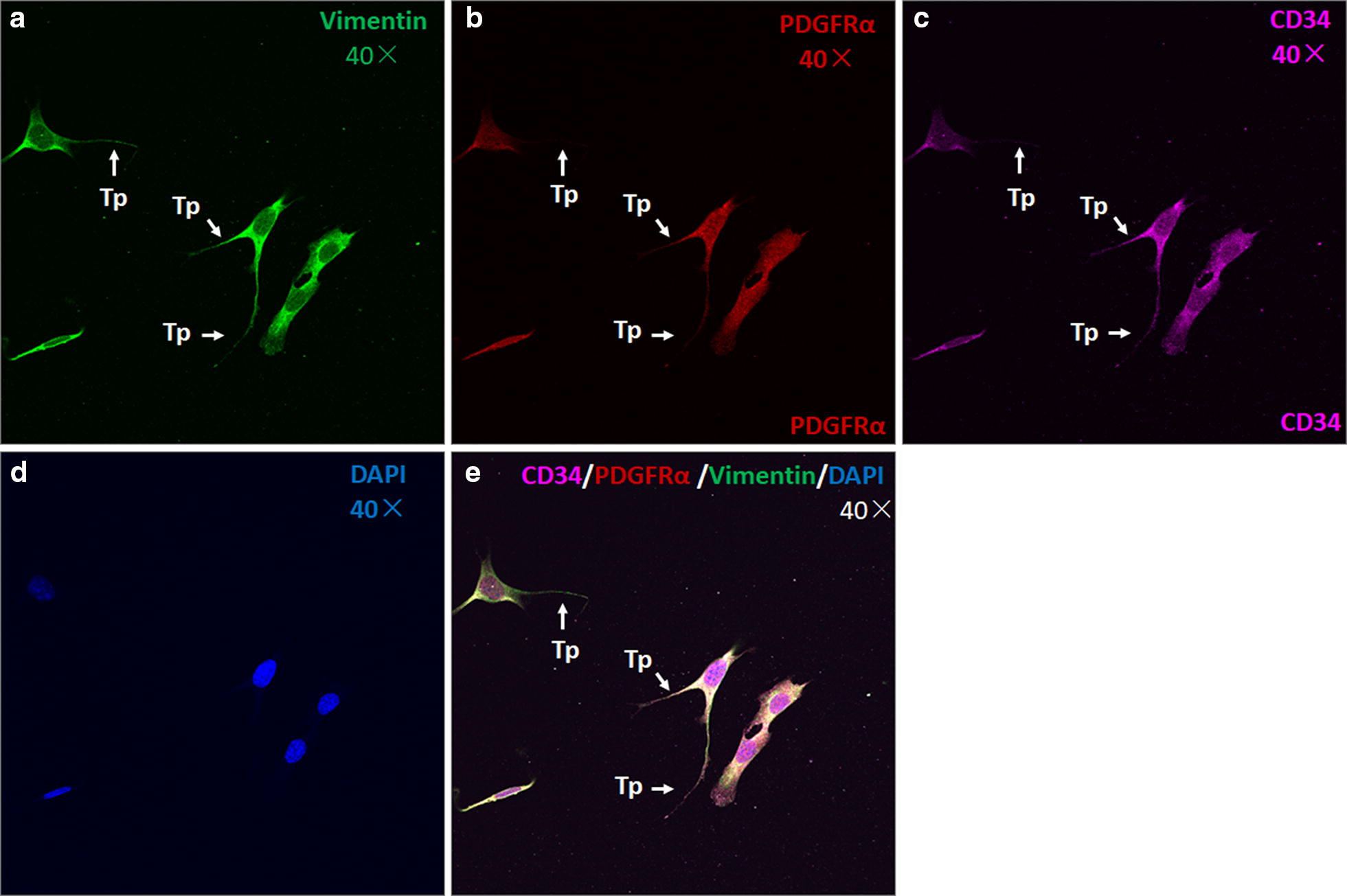
TCs are CD34/PDGFR-α/vimentin triple positive in vitro. Triple immunofluorescence labelling for vimentin (a, green), PDGFR-α (b, red) and CD34 (c, purple) with DAPI (d, blue) counterstain for nuclei. TCs are CD34, PDGFR-α and vimentin positive (e). Arrows show typical TCs with long and thin telopodes (Tp) with dilations. Original magnification ×400
Fig. 3.
The effects of ITGB1 on cell proliferation of TCs treated with PI3K/p110α, PI3Kα/δ, PKCβ, or GSK3 inhibitors. a Expression of ITGB1 mRNA in TCs 12, 24, and 48 h after treatment with TGFβ1 at 0.05, 0.5, or 5 ng/ml, respectively. b Expression of ITGB1 mRNA in TCs treated with vehicle, non-specific sequencing (NC), or ITGB1 siRNA (TCITGB1−), * stand for p values less than 0.05, as compared with TCs; # stand for p values less than 0.05, respectively, as compared with NC. c Proliferation of TCs treated with vehicle (TCITGB1+) or ITGB1 siRNA (TCITGB1−), *stand for p values less than 0.05, as compared with TCITGB1+ treated with TGFβ1. d Proliferation of TGFβ1 or vehicle-treated TCITGB1+ or TCITGB1− 24 h after treatment with HS173 (PI3Kp110α inhibitor) at 0.05, 0.5 or 5 μM. e Proliferation of TGFβ1 or vehicle-treated TCITGB1+ or TCITGB1− after treatment with GDC0941 (PI3Kα/δ inhibitor) at 0.05, 0.5 or 5 μM. f Proliferation of TGFβ1 or vehicle-treated TCITGB1+ or TCITGB1− after treatment with Enzastaurin (PKCβ inhibitor) at 0.025, 0.25 or 2.5 μM. g Proliferation of TGFβ1 or vehicle-treated TCITGB1+ or TCITGB1− after treatment with SB216763 (GSK3 inhibitor). n = 3–6, *stand for p values less than 0.05, respectively, as compared with TCITGB1+; # stand for p values less than 0.05, respectively, as compared with TCITGB1+ treated with TGFβ1
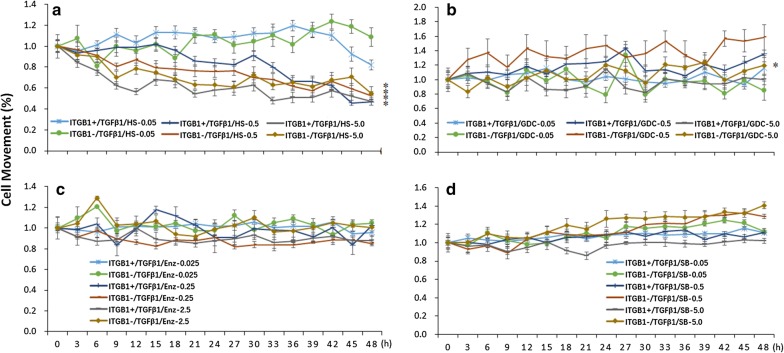
We furthermore investigated dynamic sensitivities of TCs with or without ITGB1 gene to TGFβ1 and found that TGFβ1 reduced dynamic proliferation of TCs as compared to vehicle, and TCITGB1− cells had lower proliferation than TCITGB1+ cells during TGFβ1 challenge (Additional file 2: Figure S2A). The morphology of TC ITGB1+ treat with or without TGFβ1 for 24 h and 48 h were shown with Tps in Fig. 4a–f. TCITGB1− cells had higher levels of cell proliferation in co-existence of TGFβ1 with HS173 at different doses with a dose-dependent pattern (Fig. 4g), in TGFβ1 with GDC0941 at 5.0 μM (Fig. 4h), in TGFβ1 with Enzastaurin at 0.25 μM (Fig. 4i), or in TGFβ1 with SB216763 at three doses (Fig. 4j). Of those TGFβ1-treated TCs, cell proliferation was lower after the administration of GDC0941 at 0.5 μM than at 0.05 μM. There was no significant difference of the dynamic movement capacity among TCs and TCITGB1− cells with or without TGFβ1 challenge (Additional file 2: Figure S2B). HS173 at 0.5 and 5.0 μM reduced the movement of TCITGB1+ or TCITGB1− cells in co-existence of TGFβ1 (Fig. 5a). GDC0941 at 5.0 μM increased the movement of TCITGB1+ or TCITGB1− cells, between which TCITGB1− cells movement was higher during TGFβ1 challenge (Fig. 5b). Cell movement of TCs with or without ITGB1 gene did not show significantly different during the co-existence of TGFβ1 with Enzastaurin (Fig. 5c) or SB216763 (Fig. 5d), although TCITGB1− cells treated with TGFβ1 and SB216763 showed slightly high movement.
Fig. 5.
The effects of ITGB1on the movement curve of TCs treated with PI3K inhibitors and TGFβ1. a Analysis of TC ITGB1+ or TCITGB1− movement with the treatment of HS173 and TGFβ1. b Analysis of TCITGB1+ or TCITGB1− movement with the treatment of GDC0941 and TGFβ1. c Analysis of TCITGB1+ or TCITGB1− movement with the treatment of Enzastaurin and TGFβ1. d Analysis of TCITGB1+ or TCITGB1− movement with the treatment of SB216763 and TGFβ1, n = 6–8, *stand for p values less than 0.05, as compared with TC ITGB1+
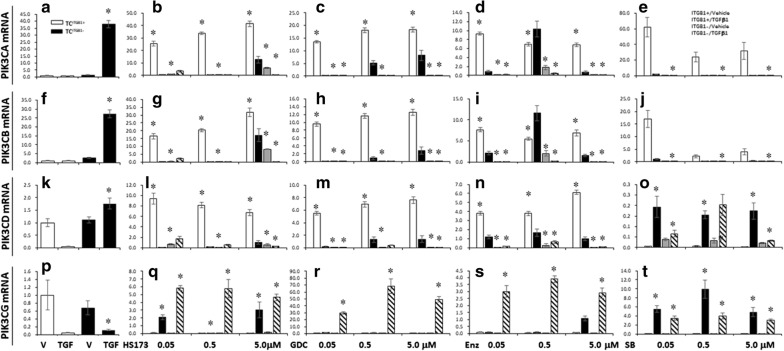
In order to define roles of ITGB1 in the expression of PIK3 class 1 catalytic genes, PIK3CA (coding PI3K p110α), PIK3CB (coding PI3K p110β), PIK3CD (coding PI3K p110δ), and PIK3CG genes (coding PI3K p110γ) were monitored after TCs were co-treated with TGFβ1 and pan-PI3K, PI3K p110α, PKC, GSK3β inhibitors. We found that TGFβ1 increased expression of PIK3CA (Fig. 6a), PIK3CB (Fig. 6f), PIK3CD (Fig. 6k), PIK3CG (Fig. 6p) in TCITGB1− cells, respectively. mRNA levels of PIK3CA, PIK3CB, and PIK3CD were significantly higher in TCITGB1+ treated with HS173 (Fig. 6b, g, l, q), GDC0941 (Fig. 6c, h, m, r), or Enzastaurin (Fig. 6d, i, n, s), as compared to other groups (p < 0.05 or less, respectively). We also noticed that mRNA levels of PIK3CA, PIK3CB, and PIK3CD were lower in TCITGB1− cells treated with HS173 at 5 μM as compared to the corresponding TCITGB1+ with vehicle, and in TGFβ1-treated cells after HS173 at 5.0 μM, GDC0941 at 0.5 and 5.0 μM, Enzastaurin at all doses, or SB216763 as compared to vehicle-treated cells. mRNA expression of PIK3CA and PIK3CB had similar patterns of changes. mRNA of PIK3CD was lower in TCITGB1− treated with SB216763 (Fig. 6o). mRNA of PIK3CG was higher in TCITGB1− treated with SB216763 (Fig. 6o) in TCITGB1− with HS173 (Fig. 6q), GDC0941(Fig. 6r), Enzastaurin (Fig. 6s), and SB216763(Fig. 6t). An increase in cell differentiation was noted in TCITGB1− treated with TGFβ1 and HS173 (Additional file 3: Figure S3), while the cell death number was significantly lower in TCITGB1− (vs TCITGB1+) treated with TGFβ1 and HS173 (Additional file 4: Figure S4A) or with TGFβ1 and Enzastaurin (Additional file 4: Figure S4C), but significantly higher in TCITGB1− (vs TCITGB1+) treated with TGFβ1 and SB216763 (Additional file 1: Figure S4D). Relevant morphological data was recorded and shown in Figure 4 A-F and Additional file 5: Figure S5.
Fig. 6.
Analysis of mRNA levels of PIK3CA, PIK3CB, PIK3CG, or PIK3CD in TCITGB1+or TCITGB1− treated with PI3K inhibitors and/or stimulated with TGFβ1. a–e mRNA levels of PIK3CA, b–j mRNA levels of PIK3CB,k–o mRNA levels of PIK3CD, p–t mRNA levels of PIK3CG monitored after TCITGB1+or TCITGB1− co-treated with TGFβ1 and PI3Kp110α, PI3Kα/δ, PKCβ, GSK3 inhibitors, respectively, n = 3–6, *stand for p values less than 0.05, as compared with TC ITGB1+
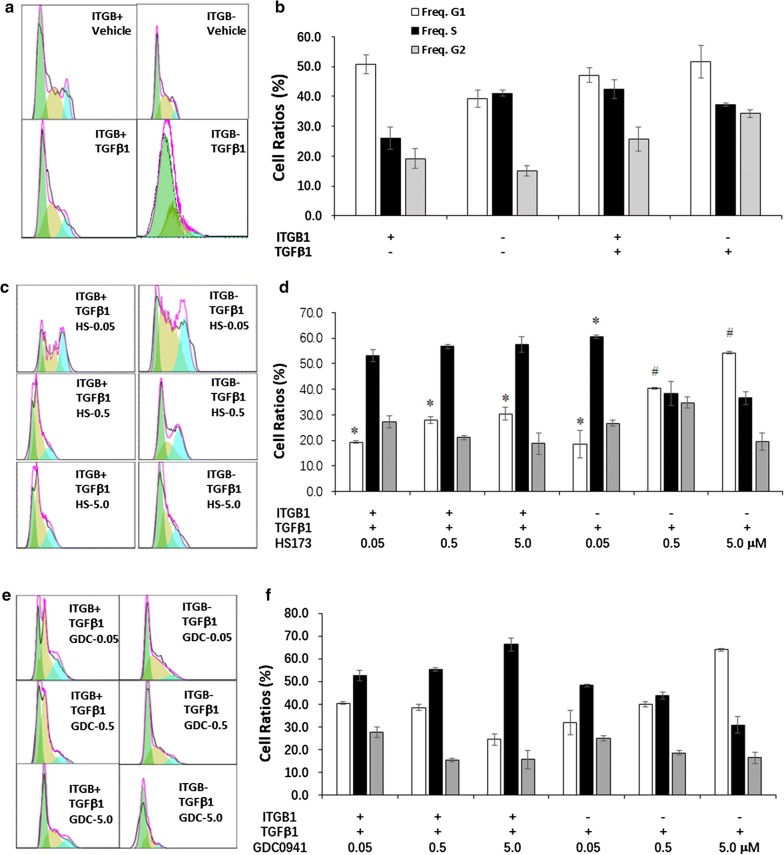
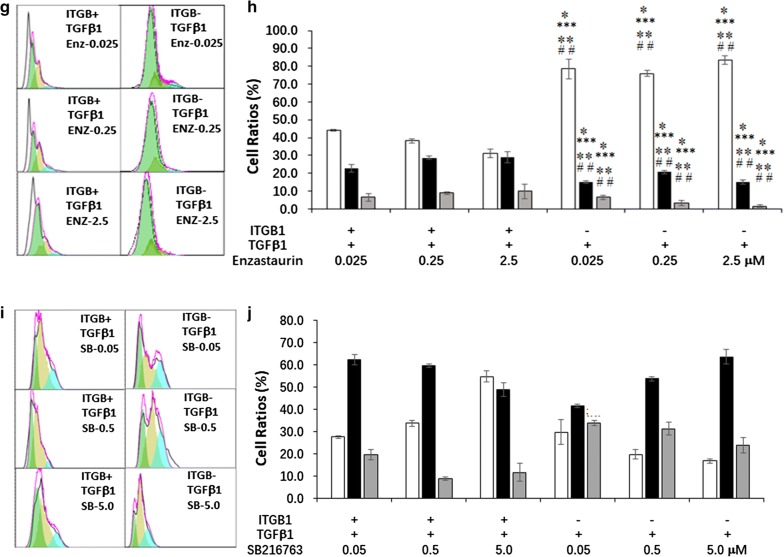
We further analyzed the cell cycle of TC ITGB1+ and TCsITGB1− to find out the mechanisms in ITGB1 mediated TCs proliferation (Fig. 7a, b) alteration and found that TGFβ1-treated TC ITGB1+ or TCITGB1− number in G1 phase increased in a dose-dependent patter of HS173, while still was lower than after TGFβ1 and vehicle, except for HS173 at 5.0 μM (Fig. 7c, d). TCITGB1− number in G1 was significantly higher than TC ITGB1+ after co-treated with TGFβ1 and HS173. HS173 at 0.05 μM decreased the ratios of G1 phase and induced S phase arrest in TCITGB1− compared with TC ITGB1+ which decreased in G1 and increased in S phase after treated with TGFβ1 and HS173(Fig. 7c, d). TC ITGB1+ decreased while TCITGB1− increased in G2 phase with a dose-dependent pattern after TGFβ1 and GDC0941 (Fig. 7e, f). In opposite, TCITGB1− decreased while TCs increased in S and G2 phases. TCITGB1− number increased in G1 phase and deceased in S and G2 phases after treatment with Enzastaurin, as compared with TC ITGB1+ or TCITGB1− with or without TGFβ1 (Fig. 7g, h). TC ITGB1+ increased in G1 phase and TCITGB1− decreased in G1 and G2 phases gradually with increased doses of SB216763 after treated with TGFβ1 and SB216763 (Fig. 7i, j), where the number of TCITGB1− in G2 phage was higher than TC ITGB1+.
Fig. 7.
The effects of ITGB1 on cell cycle of TCs treated with PI3K inhibitors and TGFβ1. a–d Cell cycle analysis of HS173 (PI3K/p110α inhibitor) in TGFβ1-induced TCITGB1+ by flowcytometry. e–f. Cell cycle analysis of GDC0941 (pan-PI3K inhibitor) in TGFβ1-induced TCITGB1+ by flowcytometry. g–h Cell cycle analysis of Enzastaurin (PI3K/PKC inhibitor) in TGFβ1-induced TCITGB1+ by flowcytometry. i–j Cell cycle analysis of SB216763 (GSK-3 inhibitor) in TGFβ1-induced TCITGB1+ by flowcytometry, n = 3–6, * stand for p values less than 0.05, as compared with TGFβ1-induced TC ITGB1+, # stand for p values less than 0.05, as TCITGB1− treated with PI3K inhibitors and TGFβ1compared with TC ITGB1+ treated with PI3K inhibitors and TGFβ1. ** stand for p values less than 0.05, as compared with TC ITGB1+, ## stand for p values less than 0.05, as compared with TC ITGB1−, ***stand for p values less than 0.05, as compared with TGFβ1-induced TC ITGB1+
Discussion
TCs connect with themselves or other cells through three-dimensional networks of TCs-specific telopodes and cell proliferation to maintain the cell–cell communication and microenvironmental stability and accelerate the process of tissue repair [19]. PI3K and TGF family networks in TCs differed from other cells which appear in the lung tissue and PI3K-TGF signal pathways play the critical role in the regulation of TCs proliferation and expression of PI3K isoform genes [5]. The present study furthermore investigates roles of ITGB1 in PI3K-TGF signal pathways-regulated TCs bio-behaviors. Different integrin subunits binding to the same ligand can trigger different signal transduction, and the expression patterns of integrins on cell surface is the key to determine the response of cells to microenvironments to immune and inflammatory regulation [19–21]. Although ITGB1 on alveolar epithelium was found to alleviate epithelial injury and remodeling in acute respiratory distress syndrome [22], profiles and functions of ITGB1 and their family members in lung TCs remain unclear due to the limited source of preliminary lung TCs. The present study firstly reported characters of ITGB family member networks and interactions in the primary mouse lung TCs, highly dependent upon cell types and functions. Of elements in TCs networks and interactions, ITGB4 declined and ITGB1 increased rapidly on day 10. We found that the interaction between PI3K and ITGB1 could complement the mechanisms of TCs proliferation and ITGB1 regulated TC sensitivity to PI3K isoform protein inhibition or PKC inhibition.
ITGB1 plays critical roles in the maintenance of cell proliferation and movement, evidenced by the finding that the over-expression of ITGB1 genes could increase cancer cell proliferation and metastasis which was declined by deletion of ITGB1 gene [23]. TGFβ activation depends upon the cytoplasmic domain integrity of ITGB subunit which connects with actin cytoskeleton or the extracellular environment, e.g. specific binding of ITGαvβ6 and αvβ8 to the arm domain of Pro-TGFβ1 [24]. Our data suggest that ITGB1 play the decisive role in TC survival and proliferation and change TC sensitivity to stimulators or inhibitors. The deletion of ITGB1 increased the sensitivity of TCs to PI3Kp/110α, PI3Kα/δ, PKCβ, or GSK3 inhibitors, indicating that ITGB1 contributes to multiple signal pathways and the interaction between ITGB1 and TGFβ regulates cell fate and proliferation through intracellular PI3K/PKC/GSK pathways. For example, the binding of ITGαvβ6 with TGFβ1 could regulate the production of inflammatory mediators through the activation of HMGB1 in injured epithelial cells, accelerating lung tissue repair and curing [25]. We noticed that the dose of TGFβ1 which did not increase TCITGB1+ proliferation could accelerate TCITGB1− proliferation, and such effects of TGFβ1 became more obvious when PI3K/PKC/GSK pathways were down-regulated in TCITGB1− cells, especially GSK3α and β signal pathways.
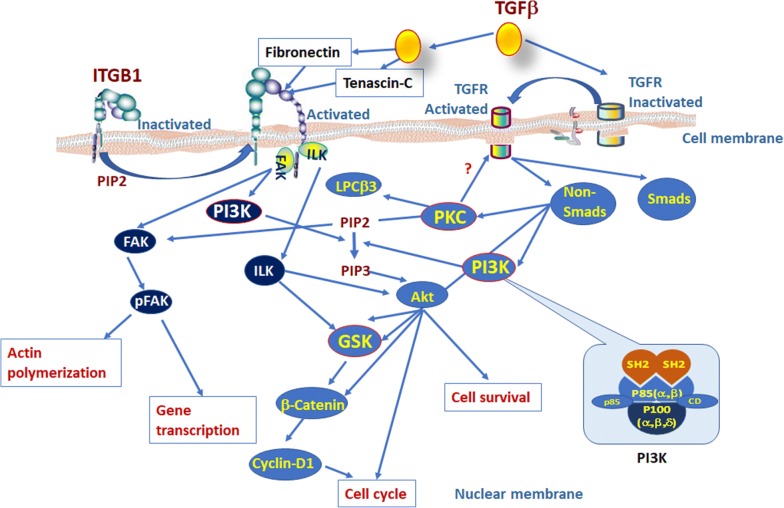
The present study demonstrates that intracellular ITGB1-TGFβ1-PI3K signal pathways in TCs have the strong compensatory capacity to maintain cell survival and proliferation. It was evidenced by the finding that TGFβ1 increased proliferation increased in TCs with combined inhibition of ITGB1 and one of PI3K/p110α, PI3Kα/δ, PKCβ, or GSK3β signaling. We propose that the hyper-sensitivity of TCITGB1− with either inhibitor may result from the compensatory mechanisms of other signal pathway function or crosstalk as in other condition [26–28]. The interaction of ITGB-PI3K plays important roles in the maintenance of normal lung cell hemostasis and function and the response to inflammation, though it is unclear in lung TCs. For example, ITGB1 protein could regulate the activation of NFκB and the expression of pro-inflammatory cytokines in alveolar macrophage [20], control intracellular transcription, accelerate cell cycle progression, movement, survival, metabolism, and DNA damage [29]. TGF-β regulates cell fate and plasticity as well as multifunctional cellular responses via mothers against decapentaplegic homolog (SMAD) or non-SMAD pathways or ITGB1 pathway band with fibronectins and tenascin-C PI3K/AKT pathway, as explained in Fig. 7. The conversion from oleate-activated transcription factor (PIP2) to plasma membrane intrinsic protein 3 (PIP3) regulated by PI3K isoform proteins is the central process of the interaction between Akt and GSK or between PKC and Akt directly or indirectly [30–32].
It is also possible that PI3K inhibitors could inhibit the fibrinogen/tenascin-C binding to ITGαIIbβ3 by inducing GTPase-Rap1 or the PI3K p110β and PI3K p110γ directly as downstream integrins by inducing activation of G protein-coupled receptors [30]. Figure 7 demonstrates that integrins regulate the activity of PI3K signaling pathway and cause the accumulation of PIP3. ITGα2β1 was found to activate PI3K p110β, RasGRP2, or GTP binding to Rap1b, to amplify the activation of ITGα2β1signals and forming regulatory feedback loops around the cell membrane [31]. The interaction between ITGB-PI3K could influence the selective bind to ITGαvβ3 receptor to regulate cell cycle, cell proliferation and cell migration [32]. Our results demonstrate that ITGB1 plays the significant role in the regulation of TC proliferation, since the inhibition of PI3Kp110α, PI3Kα/δ, or PKCβ caused the increase in S phase of TCs with ITGB1, while mainly in G1 phase in TCs without ITGB1, different from the inhibition of GSK3β. We hypothesize that PIK3CG gene and protein p110-γ may play the regulatory roles in cell cycle responses to TGF and those inhibitors in the absence of ITGB1. PI3K/Akt plays important roles in the molecular mechanism of cell cycle and survival through the signals of Akt/PKC/nuclear factor kappa-B (NF-κB) and GSK-3 [33] and regulates GSK-3β-dominated downstream survival signaling pathway of serine/threonine kinase [34]. Our results indicate that both TGF-β-band type II and type I serine and threonine kinase receptors (TβRII and TβRI) and ITGB1 can activate PI3K signal pathway either via the switch of the heterotetrameric receptor complex from inactivated to activated TβRII and TβRI or via the switch of the integrin-mediated focal adhesion kinase signaling from inactivated to activated ITGB1. Both switches are required for the activation of class IA PI3K mediated by TGFβ1 [35] and also result in the activation of PI3Kp110α, PI3Kα/δ, PKCβ, or GSK3 (Fig. 8). Molecules and networks in the TGFβ/ITGB1/PI3K pathways can be considered as TC-specific functional biomarkers and therapeutic targets like others [36–40].
Fig. 8.
The explanation of TGF-β regulates cell fate and plasticity as well as multifunctional cellular responses via SMADor non-SMAD pathways or ITGB1 pathway band with fibronectins and tenascin-C PI3K/AKT pathway. The molecules with red circles were key signaling pathways in our current study [30–34]
In conclusion, we found the characters and interactions of ITG or PKC family member networks in primary mouse lung TCs and defined their specificity and dynamic alternations, different from other cells which often appear in the lung tissue. The deletion of ITGB1 changed TCs sensitivity to treatment with multifunctional cytokines or signal pathway inhibitors. The compensatory mechanisms may occur among TGFβ1-induced PI3Kp110α, PI3Kα/δ, PKCβ, or GSK3 when ITGB1 gene was deleted, leading to alterations of TCs cell cycle and proliferation. Of those PI3K isoform protein genes, mRNA expression of PIK3CG altered with ITGB1-negative TC cycle and proliferation. Such compensatory mechanism can provide insights for understanding molecular mechanisms by which TCs have strong capacity of proliferation and repair and of cell–cell communication and for investigating the potential roles of TCs in the development of drug resistance.
Supplementary information
Additional file 1: Figure S1. The characters of GSK networks and interactions in mouse lung primary TCs cultured for 5 days (TC 5) and 10 days (TC 10) were compared with other tissue cells, e.g. airway basal cells (Basal cells), proximal airway cells (Duct cells).
Additional file 2: Figure S2. The effects of ITGB1 on the cell proliferation and movement curves of TCs treated with TGFβ1. A. Analysis of TC ITGB1+ or TCITGB1− cell proliferation curves with the treatment of TGFβ1. B. Analysis of TC ITGB1+ or TCITGB1− cell movement curves with the treatment of TGFβ1, n = 6–8, *stand for p values less than 0.05, as compared with TC ITGB1+; # stand for p values less than 0.05, respectively, as compared with NC.
Additional file 3: Figure S3. The effects of ITGB1on the cell differentiation curve of TCs treated with TGFβ1 and PI3Kp110α, PI3Kα/δ, PKCβ, GSK3 inhibitors, respectively, n = 6–8.
Additional file 4: Figure S4. The effects of ITGB1on the cell death curve of TCs treated with TGFβ1 and PI3Kp110α, PI3Kα/δ, PKCβ, GSK3 inhibitors, respectively, n = 6–8, *stand for p values less than 0.05, as compared with TC ITGB1+ treated with TGFβ1 and PI3K inhibitors.
Additional file 5: Figure S5. Cell bio-behaviors of TC ITGB1+ or TCITGB1− treated with TGFβ1 and PI3Kp110α, PI3Kα/δ, PKCβ, GSK3 inhibitors, respectively, n= 6–8.
Acknowledgements
The work was supported by Zhongshan Distinguished Professor Grant (XDW), The National Nature Science Foundation of China (91230204, 81270099, 81320108001, 81270131, 81300010, 81700008, 81873409), The Shanghai Committee of Science and Technology (12JC1402200, 12431900207, 11410708600, 14431905100), Operation funding of Shanghai Institute of Clinical Bioinformatics, Ministry of Education for Academic Special Science and Research Foundation for Ph.D. Education (20130071110043), and National Key Research and Development Program (2016YFC0902400, 2017YFSF090207, 2017YFC0909500).
Abbreviations
- TCs
telocytes
- Tps
telopodes
- SV40
simian vacuolating virus 40 small and large T antigen
- TGF-β
transforming growth factor-β
- SPSS
Statistical Package for the Social Sciences
- ANOVA
one‑way analysis of variance
- PI3K
phosphatidylinositol 3′ –kinase
- GSK-3
glycogen synthasc kinase-3
- mTOR
Mammalian target of rapamycin
- GSK-3
glycogen synthase kinase-3
- ITG
integrins
- ITGB1
integrin beta1
- PKC
protein kinase C
- DEN
differential network model
- CDK
cyclin-dependent kinases
- TCITGB1−
TCs knocked out of ITGB1
- TC 5
mouse lung primary TCs cultured for 5 days
- TC 10
mouse lung primary TCs cultured for 10 days
- Fbs
fibroblasts
- MSCs
mesenchymal stem cells
- ATII
alveolar type II cells
- ABCs
airway basal cells
- PACs
proximal airway cells
- CD8+ T-BL
CD8+ T cells come from bronchial lymph nodes
- CD8+ T-LL
CD8+ T cells come from lung
- ARDS
acute respiratory distress syndrome
- PIP2
oleate-activated transcription factor
- PIP3
plasma membrane intrinsic protein 3
- SMAD
mothers against decapentaplegic homolog
- NF-κB
nuclear factor kappa-B
Authors’ contributions
DS designed the study and completed the experimental process, literature search, and generation of figures. DS and XW wrote and edited the manuscript. DS, LT, NYH, LW and TZ completed generation of figures. All authors reviewed the manuscript. All authors read and approved the final manuscript.
Funding
This work was supported by Grants from the National Natural Science Foundation of China (81700008, 81873409).
Availability of data and materials
Not applicable.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Dongli Song, Li Tang and Jianan Huang contribute to this article equally as the first author
Supplementary information
Supplementary information accompanies this paper at 10.1186/s12967-019-02181-2.
References
- 1.Song D, Yang D, Powell CA, et al. Cell–cell communication: old mystery and new opportunity. Cell Biol Toxicol. 2019;35(2):89–93. doi: 10.1007/s10565-019-09470-y. [DOI] [PubMed] [Google Scholar]
- 2.Nawaz M, Shah N, Zanetti BR, et al. Extracellular vesicles and matrix remodeling enzymes: the emerging roles in extracellular matrix remodeling, progression of diseases and tissue repair. Cells. 2018;7(10):E167. doi: 10.3390/cells7100167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Marini M, Ibba-Manneschi L, Manetti M. Cardiac telocyte-derived exosomes and their possible implications in cardiovascular pathophysiology. Adv Exp Med Biol. 2017;998:237–254. doi: 10.1007/978-981-10-4397-0_16. [DOI] [PubMed] [Google Scholar]
- 4.Shoshkes-Carmel M, Wang YJ, Wangensteen KJ, et al. Subepithelial telocytes are an important source of Wnts that supports intestinal crypts. Nature. 2018;557(7704):242–246. doi: 10.1038/s41586-018-0084-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Song D, Tang L, Wang L, et al. Roles of TGFbeta1 in the expression of phosphoinositide 3-kinase isoform genes and sensitivity and response of lung telocytes to PI3K inhibitors. Cell Biol Toxicol. 2019 doi: 10.1007/s10565-019-09487-3. [DOI] [PubMed] [Google Scholar]
- 6.Song D, Xu M, Qi R, et al. Influence of gene modification in biological behaviors and responses of mouse lung telocytes to inflammation. J Transl Med. 2019;17(1):158. doi: 10.1186/s12967-019-1870-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kim SJ, Jung KH, Son MK, et al. Tumor vessel normalization by the PI3K inhibitor HS-173 enhances drug delivery. Cancer Lett. 2017;403:339–353. doi: 10.1016/j.canlet.2017.06.035. [DOI] [PubMed] [Google Scholar]
- 8.Ren C, Chen X, Du N, et al. Low-intensity pulsed ultrasound promotes Schwann cell viability and proliferation via the GSK-3beta/beta-catenin signaling pathway. Int J Biol Sci. 2018;14(5):497–507. doi: 10.7150/ijbs.22409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhang H, Su Y, Wang J, et al. Ginsenoside Rb1 promotes the growth of mink hair follicle via PI3K/AKT/GSK-3beta signaling pathway. Life Sci. 2019;229:210–218. doi: 10.1016/j.lfs.2019.05.033. [DOI] [PubMed] [Google Scholar]
- 10.Agrawal B. New therapeutic targets for cancer: the interplay between immune and metabolic checkpoints and gut microbiota. Clin Transl Med. 2019;8(1):23. doi: 10.1186/s40169-019-0241-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ghosh D, Nandi S, Bhattacharjee S. Combination therapy to checkmate Glioblastoma: clinical challenges and advances. Clin Transl Med. 2018;7(1):33. doi: 10.1186/s40169-018-0211-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Rusu MC, Cretoiu D, Vrapciu AD, et al. Telocytes of the human adult trigeminal ganglion. Cell Biol Toxicol. 2016;32(3):199–207. doi: 10.1007/s10565-016-9328-y. [DOI] [PubMed] [Google Scholar]
- 13.Zhu Y, Zheng M, Song D, et al. Global comparison of chromosome X genes of pulmonary telocytes with mesenchymal stem cells, fibroblasts, alveolar type II cells, airway epithelial cells, and lymphocytes. J Transl Med. 2015;13:318. doi: 10.1186/s12967-015-0669-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zeng T, Zhang W, Yu X, et al. Big-data-based edge biomarkers: study on dynamical drug sensitivity and resistance in individuals. Brief Bioinform. 2016;17(4):576–592. doi: 10.1093/bib/bbv078. [DOI] [PubMed] [Google Scholar]
- 15.Yu X, Zeng T, Wang X, et al. Unravelling personalized dysfunctional gene network of complex diseases based on differential network model. J Transl Med. 2015;13:189. doi: 10.1186/s12967-015-0546-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang W, Zeng T, Liu X, et al. Diagnosing phenotypes of single-sample individuals by edge biomarkers. J Mol Cell Biol. 2015;7(3):231–241. doi: 10.1093/jmcb/mjv025. [DOI] [PubMed] [Google Scholar]
- 17.Mou Y, Wang Y, Li J, et al. Immunohistochemical characterization and functional identification of mammary gland telocytes in the self-assembly of reconstituted breast cancer tissue in vitro. J Cell Mol Med. 2013;17(1):65–75. doi: 10.1111/j.1582-4934.2012.01646.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Zhou Q, Wei L, Zhong C, et al. Cardiac telocytes are double positive for CD34/PDGFR-alpha. J Cell Mol Med. 2015;19(8):2036–2042. doi: 10.1111/jcmm.12615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Wang X, Cretoiu D. Telocytes 2016: Connecting Cells.
- 20.Zhang M, Yan X, Liu W, et al. Endothelial semaphorin 7A promotes seawater aspiration-induced acute lung injury through plexin C1 and beta1 integrin. Mol Med Rep. 2017;16(4):4215–4221. doi: 10.3892/mmr.2017.7097. [DOI] [PubMed] [Google Scholar]
- 21.Kadel D, Zhang Y, Sun HR, et al. Current perspectives of cancer-associated fibroblast in therapeutic resistance: potential mechanism and future strategy. Cell Biol Toxicol. 2019;35(5):407–421. doi: 10.1007/s10565-019-09461-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shi L, Dong N, Ji D, et al. Lipopolysaccharide-induced CCN1 production enhances interleukin-6 secretion in bronchial epithelial cells. Cell Biol Toxicol. 2018;34(1):39–49. doi: 10.1007/s10565-017-9401-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Guo L, Sun C, Xu S, et al. Knockdown of long non-coding RNA linc-ITGB1 inhibits cancer stemness and epithelial-mesenchymal transition by reducing the expression of Snail in non-small cell lung cancer. Thorac Cancer. 2019;10(2):128–136. doi: 10.1111/1759-7714.12911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Dong X, Zhao B, Iacob RE, et al. Force interacts with macromolecular structure in activation of TGF-beta. Nature. 2017;542(7639):55–59. doi: 10.1038/nature21035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pittet JF, Koh H, Fang X, et al. HMGB1 accelerates alveolar epithelial repair via an IL-1beta- and alphavbeta6 integrin-dependent activation of TGF-beta1. PLoS ONE. 2013;8(5):e63907. doi: 10.1371/journal.pone.0063907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Li D, Lo W, Rudloff U. Merging perspectives: genotype-directed molecular therapy for hereditary diffuse gastric cancer (HDGC) and E-cadherin-EGFR crosstalk. Clin Transl Med. 2018;7(1):7. doi: 10.1186/s40169-018-0184-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Truong L, Zheng YM, Song T, et al. Potential important roles and signaling mechanisms of YPEL4 in pulmonary diseases. Clin Transl Med. 2018;7(1):16. doi: 10.1186/s40169-018-0194-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Li Q, Qi F, Meng X, et al. Mst1 regulates colorectal cancer stress response via inhibiting Bnip3-related mitophagy by activation of JNK/p53 pathway. Cell Biol Toxicol. 2018;34(4):263–277. doi: 10.1007/s10565-017-9417-6. [DOI] [PubMed] [Google Scholar]
- 29.Gao L, Zhao M, Li P, et al. Glycogen synthase kinase 3 (GSK3)-inhibitor SB216763 promotes the conversion of human umbilical cord mesenchymal stem cells into neural precursors in adherent culture. Hum Cell. 2017;30(1):11–22. doi: 10.1007/s13577-016-0146-6. [DOI] [PubMed] [Google Scholar]
- 30.Laurent PA, Severin S, Hechler B, et al. Platelet PI3Kbeta and GSK3 regulate thrombus stability at a high shear rate. Blood. 2015;125(5):881–888. doi: 10.1182/blood-2014-07-588335. [DOI] [PubMed] [Google Scholar]
- 31.Manganaro D, Consonni A, Guidetti GF, et al. Activation of phosphatidylinositol 3-kinase beta by the platelet collagen receptors integrin alpha2beta1 and GPVI: the role of Pyk2 and c-Cbl. Biochim Biophys Acta. 2015;1853(8):1879–1888. doi: 10.1016/j.bbamcr.2015.05.004. [DOI] [PubMed] [Google Scholar]
- 32.Cen B, Wei Y, Huang W, et al. An efficient bivalent Cyclic RGD-PIK3CB siRNA conjugate for specific targeted therapy against glioblastoma in vitro and in vivo. Mol Ther Nucleic Acids. 2018;13:220–232. doi: 10.1016/j.omtn.2018.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.He X, Zhang L, Queme LF, et al. A histone deacetylase 3-dependent pathway delimits peripheral myelin growth and functional regeneration. Nat Med. 2018;24(3):338–351. doi: 10.1038/nm.4483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Russo I, Femmino S, Barale C, et al. Cardioprotective properties of human platelets are lost in uncontrolled diabetes mellitus: a study in isolated rat hearts. Front Physiol. 2018;9:875. doi: 10.3389/fphys.2018.00875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Han S, Ding S, Miao X, et al. TGF-beta1 expression in regulatory NK1.1(−)CD4(+)NKG2D(+) T cells dependents on the PI3K-p85alpha/JNK, NF-kappaB and STAT3 pathways. Am J Cancer Res. 2018;8(3):489–501. [PMC free article] [PubMed] [Google Scholar]
- 36.Wang X. Clinical trans-omics: an integration of clinical phenomes with molecular multiomics. Cell Biol Toxicol. 2018;34(3):163–166. doi: 10.1007/s10565-018-9431-3. [DOI] [PubMed] [Google Scholar]
- 37.Liu X, Wu J. History, applications, and challenges of immune repertoire research. Cell Biol Toxicol. 2018;34(6):441–457. doi: 10.1007/s10565-018-9426-0. [DOI] [PubMed] [Google Scholar]
- 38.Yang X, Mao X, Ding X, et al. miR-146a down-regulation alleviates H2O2-induced cytotoxicity of PC12 cells by regulating MCL1/JAK/STAT pathway: miR-146a down-regulation relieves H2O2-induced PC12 cells cytotoxicity by MCL1/JAK/STAT. Cell Biol Toxicol. 2018;34(6):479–489. doi: 10.1007/s10565-018-9424-2. [DOI] [PubMed] [Google Scholar]
- 39.Yan F, Zhao H, Zeng Y. Lipidomics: a promising cancer biomarker. Clin Transl Med. 2018;7(1):21. doi: 10.1186/s40169-018-0199-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhang Y. Cell toxicity mechanism and biomarker. Clin Transl Med. 2018;7(1):34. doi: 10.1186/s40169-018-0212-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Figure S1. The characters of GSK networks and interactions in mouse lung primary TCs cultured for 5 days (TC 5) and 10 days (TC 10) were compared with other tissue cells, e.g. airway basal cells (Basal cells), proximal airway cells (Duct cells).
Additional file 2: Figure S2. The effects of ITGB1 on the cell proliferation and movement curves of TCs treated with TGFβ1. A. Analysis of TC ITGB1+ or TCITGB1− cell proliferation curves with the treatment of TGFβ1. B. Analysis of TC ITGB1+ or TCITGB1− cell movement curves with the treatment of TGFβ1, n = 6–8, *stand for p values less than 0.05, as compared with TC ITGB1+; # stand for p values less than 0.05, respectively, as compared with NC.
Additional file 3: Figure S3. The effects of ITGB1on the cell differentiation curve of TCs treated with TGFβ1 and PI3Kp110α, PI3Kα/δ, PKCβ, GSK3 inhibitors, respectively, n = 6–8.
Additional file 4: Figure S4. The effects of ITGB1on the cell death curve of TCs treated with TGFβ1 and PI3Kp110α, PI3Kα/δ, PKCβ, GSK3 inhibitors, respectively, n = 6–8, *stand for p values less than 0.05, as compared with TC ITGB1+ treated with TGFβ1 and PI3K inhibitors.
Additional file 5: Figure S5. Cell bio-behaviors of TC ITGB1+ or TCITGB1− treated with TGFβ1 and PI3Kp110α, PI3Kα/δ, PKCβ, GSK3 inhibitors, respectively, n= 6–8.
Data Availability Statement
Not applicable.