Abstract
The prognosis of pancreatic ductal adenocarcinoma (PDAC) remains poor even among patients with the same Tumor-Node-Metastasis stage. Thus, it is necessary to identify biomarkers that can accurately predict outcomes. There is accumulating evidence suggesting that microRNA (miR) expression influences overall survival (OS) time in patients with PDAC, via the regulation of tumor suppressor genes and oncogene expression. Specifically, miR-608 expression is hypothesized to regulate PDAC progression via the downregulation of bromodomain-containing protein 4 (BRD4) expression and the promotion of cell apoptosis. The present study aimed to investigate this theory. Thus, whole genome expression microarray analysis was performed on three patient samples with OS time >30 months, and compared with three samples with <12 months, in order to identify differentially expressed miRNAs (DEMs), via EdgeR analysis. A total of 591 DEMs were identified that exhibited a fold change >1, including 390 upregulated and 201 downregulated genes. Subsequently, 10 DEMs were identified using quantitative PCR in a different population of 68 tissues, collected from patients with PDAC. Notably, a high level of miRNA-608 expression was associated with longer OS times (P<0.05). Bioinformatics analysis was then performed to predict the molecular mechanism underlying the regulation of cell apoptosis by miRNA-608, and a dual-luciferase assay determined that overexpression of mimics in the Panc-1 and Bxpc-3 pancreatic cancer cell lines increased levels of apoptosis compared with the control. Additionally, high miRNA-608 expression decreased the protein level of BRD4. A luciferase reporting assay was used to elucidate whether miRNA-608 may directly inhibit the expression of BRD4 by binding to the 3′-untranslated region of its mRNA in the same cell lines. A subsequent rescue experiment indicated that the upregulation of BRD4 may reverse the apoptosis-promoting effect induced by miRNA-608. In summary, the present study revealed that miRNA-608 promotes apoptosis in PDAC via the negative regulation of BRD4. The results of the present study provide a theoretical basis that may improve the prediction of prognosis in patients with PDAC, and also indicate an opportunity to develop individualized treatment and investigate novel therapeutics that target these mechanisms.
Keywords: pancreatic ductal adenocarcinoma, microRNA-608, bromodomain-containing protein 4, apoptosis
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is a highly malignant tumor of the digestive system, which exhibits a high-mortality rate (1). Advances in surgical and chemotherapeutic techniques have improved the treatment and overall survival (OS) rates of the majority of patients with gastrointestinal tumors; however, these advances have not yet conferred improvement on patients with PDAC, who currently exhibit a 5-year relative OS rate of 8%, in the USA (2). The early diagnosis of PDAC is rare and, consequently, numerous patients are in the advanced stages upon primary diagnosis, which results in fewer surgical options and an increase in chemotherapeutic resistance (2). Currently used clinical biomarkers, such as carbohydrate antigen 19-9 and carcinoembryonic antigen, are not able to accurately diagnose early disease or predict the chemotherapeutic response/OS times in patients with PDAC (3). Therefore, the identification of novel biomarkers for PDAC progression and treatment evaluation represents a shared objective for both clinical and laboratory staff, which may confer great benefit on patient outcomes.
MicroRNAs (miRNAs) are a form of endogenous non-coding RNAs that consist of 20–24 nucleotides and serve a post-transcriptional regulatory role, via binding to the 3′-untranslated region (3′-UTR) of mRNAs (4). It has been reported that >1,000 miRNAs are encoded by the mammalian genome (5), with 30% of the human genome (~5,300 genes) targeted by miRNAs (6). Abnormal expression of miRNAs has been revealed in numerous types of cancer and is implicated in various physiological and pathological processes of tumor cells, such as proliferation, differentiation, apoptosis, metabolism, metastasis and cell signaling (7). As a result of significant advances in bioinformatics and microarray techniques, human genome sequencing represents a novel technique for the identification of the genes that influence cancer progression. Gene expression microarray analysis has been used to discover multiple molecular signatures associated with PDAC progression (8,9). A plethora of differentially expressed miRNAs have been discovered by comparing the gene expression profiles between cancer tissues and adjacent tissues of European and American populations (10). However, due to differences in the genetic backgrounds and living environments between the aforementioned populations and Asian populations, the findings may not be transferable. Therefore, more experiments need to be conducted on Asian populations in order to identify miRNAs that are highly specific and sensitive to PDAC, allowing for the accurate predictions of diagnosis and prognosis.
The aim of the present study was to identify miRNAs that are able to predict the prognosis of patients with PDAC. miRNA expression levels were detected in cancerous tissues from six patients with different prognoses of PDAC via microarray analysis, and the top 10 miRNAs with the largest differences in fold change were screened and verified in an independent group of 68 patients. It was revealed that a low expression level of miRNA-608 was associated with poor prognosis. Furthermore, overexpression of miRNA-608 in pancreatic cancer cell lines (Panc-1 and Bxpc-3) significantly increased apoptosis and decreased the protein level of bromodomain-containing protein (BRD)-4 (no difference was observed in transcription). Finally, luciferase reporter gene and rescue experiments were conducted and confirmed that miRNA-608 promotes apoptosis in PDAC tissues via the negative regulation of BRD4. The current results provide a theoretical basis for a novel technique to predict the prognosis of PDAC patients, which may result in improved individualized treatment and the identification of new drug targets.
Materials and methods
Gene expression microarray analysis
A total of six patients with PDAC who had received a surgical resection at Tianjin Medical University Cancer Institute & Hospital (TMUCIH; Tianjin, China) between January 2008 and December 2015 were recruited for the present study, and gene expression dataset was collected (clinicopathological data of the patients are detailed in Table I). The six samples were divided into two groups, dependent on prognosis. The long-survival group included three patients with OS times >30 months, and the short-survival group consisted of three patients with OS times <12 months. The inclusion criteria were as follows: i) Patients who underwent surgery from the same surgeon in Tianjin Medical University Cancer Institute & Hospital; ii) patients who had a pathological diagnosis of PDAC; iii) patients who had received an R0 resection; iv) patients who had not received preoperative chemoradiotherapy; and v) patients who exhibited no serious postoperative complications. The exclusion criteria were as follows: i) Patients who died with 7–12 days following surgery; ii) patients with severe postoperative complications requiring secondary surgery or ICU treatment; and iii) patients whose death was not related to tumor.
Table I.
Clinicopathological characteristics of the six patients.
| Long survival group | Short survival group | |||||
|---|---|---|---|---|---|---|
| Patient no. | 1 | 2 | 3 | 4 | 5 | 6 |
| Sex | Male | Female | Male | Male | Male | Male |
| Age, years | 48 | 36 | 65 | 72 | 69 | 60 |
| Differentation state | Medium | Poor | High | High | Medium | Poor |
| TNM | T4N0M0 | T1N1M0 | T2N0M0 | T1bN0M0 | T3N1M0 | T1bN1M0 |
| AJCC Stage | III | IIB | I | IIA | IIB | III |
| OS, months | 30 | 39 | 33 | 9 | 7 | 12 |
| RFS, months | 18 | 20 | 33 | 5 | 1 | 11 |
OS, overall survival; RFS, recurrence free survival; TNM, Tumor-Node-Metastasis; AJCC, American Joint Committee on Cancer.
Total RNA was extracted from cancer and paracancerous tissues using TRIzol® (Invitrogen; Thermo Fisher Scientific, Inc.)/chloroform and purified with Agencourt Ampure magnetic beads (cat, no, APN 000132; Beckman Coulter, Inc.). Sample preparation for the microarray processing was carried out according to the protocol detailed in the GeneChip® WT PLUS Reagent kit (Thermo Fisher Scientific, Inc.). Briefly, a total of 0.5 µg RNA was subjected to two rounds of cDNA synthesis. Following fragmentation, the second-cycle single-stranded (ss)cDNA was labeled with biotin using terminal deoxynucleotidyl transferase (TdT). Subsequently the sample was hybridized using the Affymetrix Human Gene 1.0ST Array (Thermo Fisher Scientific, Inc.) for 16–18 h at 45°C. Following hybridization, the microarrays were washed and stained with streptavidin phycoerythrin on the Affymetrix Fluidics Station 450. Microarrays were scanned using the Affymetrix® GeneChip Command Console (AGCC; Thermo Fisher Scientific, Inc.), which was installed in GeneChip® Scanner 3000 7G (https://www.thermofisher.com/order/catalog/product/00-0213). The data were analyzed using the Robust Multichip Analysis (RMA) algorithm in the Partek® Genomics Suite 6.6(http://cgs.hku.hk/portal/index.php/software-for-microarray-analysis/partek-genome-suite-with-partek-pathway), the default analysis settings were used, and global scaling was used as the normalization method. The values presented are the log2 RMA signal intensity.
Clinicopathological characteristics and follow-up of study population
A total of 68 patients with pancreatic cancer, who had received a surgical resection at Tianjin Medical University Cancer Institute & Hospital (TMUCIH; Tianjin, China) between January 2008 and December 2015, were selected for use in the present study. All patients had received a confirmation of pancreatic cancer diagnosis from a pathologist in the Department of Pathology, TMUCIH. A total of 37 males and 31 females (median age, 61 years; range, 36–80 years) were included in the present study. Of the included patients, five were diagnosed with well-differentiated tumors (5/68; 7.4%), 55 with moderately-differentiated tumors (55/68; 80.9%) and eight with poorly-differentiated tumors (8/68; 11.8%), and a total of 17 patients (17/68; 25.0%) presented with lymphatic invasion. Furthermore, 21/68 (30.9%), 30/68 (44.1%), 15/68 (22.1%) and 2/68 patients (2.9%) were diagnosed at stages I, IIA, IIB and III, respectively, according to the 7th American Joint Committee on Cancer Tumor-Node-Metastasis classification system (11). The median OS time was 18.4 months and the 2-year OS rate was 35.2%. The clinicopathological characteristics of the 68 patients with PDAC are detailed in Table II. The use of all human tissues in the present study was reviewed and approved by the Ethics Committee of TMUCIH, and written informed consent was provided by all participants.
Table II.
Clinicopathological characteristics and OS times of 68 patients with pancreatic cancer.
| Characteristic | n | % |
|---|---|---|
| Sex | ||
| Male | 37 | 54.4 |
| Female | 31 | 45.6 |
| Age | ||
| ≤60 years | 31 | 45.6 |
| >60 years | 37 | 54.4 |
| Differentiation | ||
| Low | 8 | 11.8 |
| Medium | 55 | 80.9 |
| High | 5 | 7.3 |
| Tumor size | ||
| ≤4 cm | 40 | 58.8 |
| >4 cm | 28 | 41.2 |
| Vascular invasion | ||
| Present | 13 | 19.1 |
| Absent | 55 | 80.9 |
| T stage | ||
| T1+T2 | 28 | 41.2 |
| T3 | 40 | 58.8 |
| N stage | ||
| N0 | 51 | 75.0 |
| N1 | 17 | 25.0 |
| AJCC stage | ||
| I | 21 | 30.9 |
| IIA | 30 | 44.1 |
| IIB | 15 | 22.1 |
| III | 2 | 2.9 |
| OS | ||
| ≤12 months | 23 | 33.8 |
| >12 months | 45 | 66.2 |
OS, overall survival; AJCC, American Joint Committee on Cancer; T, tumor; N, node.
RNA samples and quantitative (q) PCR
Genomic RNA samples from 68 patients were collected from the TMUCIH tumor tissue bank. cDNA was synthesized from 0.5 µg of total RNA using the miRcute Plus miRNA First-Strand cDNA Kit (Tiangen Biotech Co., Ltd.) according to the manufacturer's instructions. Candidate genes and U6 were amplified using qPCR in a fluorescence reader QuantStudio 5 Real-Time PCR system (Applied Biosystems). Each sample was run in a miRcute Plus miRNA qPCR kit (Tiangen Biotech Co., Ltd.). The thermocycling conditions were as follows: Initial enzyme activation at 95°C for 15 min, followed by 40 cycles of denaturing at 94°C for 20 sec, and then annealing/extension at 60°C for 34 sec. mRNA levels were quantified using the 2−ΔΔCq method and normalized to U6 (12). The sequences of the primers used in the present study are presented in Table III.
Table III.
Sequences designed for the validation of candidate miR levels and U6 using qPCR.
| Primer | Sequence (5′-3′) |
|---|---|
| miR-221 | CGAGCTACATTGTCTGCTGGGTTTC |
| miR-492 | AGGACCTGCGGGACAAGATTCTT |
| miR-192 | GCGCTGACCTATGAATTGACAGCC |
| miR-573 | GCGCTGAAGTGATGTGTAACTGATCAG |
| miR-222 | GCGTGTCAGTTTGTCAAATACCCCA |
| miR-106b | GCGTAAAGTGCTGACAGTGCAGAT |
| miR-409 | AGGTTACCCGAGCAACTTTGCAT |
| miR-21 | GGCGCTAGCTTATCAGACTGATGTTG |
| miR-186 | GCCCAAAGAATTCTCCTTTTGGGCT |
| miR-608 | AGGGGTGGTGTTGGGACAGC |
| U6-Forward | GCGCGTCGTGAAGCGTTC |
| U6-Reverse | GTGCAGGGTCCGAGGT |
miR, microRNA.
Cell transfection
The pancreatic cancer cell lines Panc-1 and Bxpc-3 (American Type Culture Collection; ATCC) were transfected with miR-608 mimics (50 nM) and their respective negative controls (NC; non-targeting sequence; 50 nM), which were acquired from Guangzhou RiboBio Co., Ltd. The sequence of miR-608 mimic was as follow: 5′-AGGGGUGGUGUUGGGACAGCUCCGU-3′. Opti-MEM® I and Lipofectamine 3000 reagent (both from Thermo Fisher Scientific, Inc.) were used for transfection. The cells were collected 24 h after transfection.
Flow cytometry
Flow cytometry was performed using a FITC Annexin V Apoptosis Detection kit I (Becton, Dickinson and Company). Panc-1 and Bxpc-3 cells (with or without miR-608 mimic transfection) were seeded in 6-well plates at a density of ~4×105 cells/well. After overnight incubation at 37°C and 5% CO2, the cells were collected, washed twice with cold PBS, resuspended in 500 µl 1× binding buffer at a concentration of 1×106 cells/ml, and incubated with 5 µl V-fluorescein isothiocyanate (FITC) and 5 µl propidium iodide (Thermo Fisher Scientific, Inc.). The samples were collected via centrifugation at 800 rpm/min and 25°C and incubated for 15 min at 25°C in the dark. Flow cytometry was performed within 1 h, using a FACScan flow cytometer (version V10.0; BD Biosciences).
The prediction of miR-608 target by bioinformatics analyses
The potential target genes of miR-608 were predicted by using TargetScan (http://www.targetscan.org/), miRDB (http://mirdb.org/) and miRanda (http://miranda.org.uk/). Venn diagram was drawn usinkg R Project (X64; version 3.2.5) to obtain genes that appeared simultaneously on the three websites. Subsequently RNAhybrid (https://bibiserv.cebitec.uni-bielefeld.de/rnahybrid/) was used to evaluate the binding capacity of miR-608 to its target.
Western blotting
Total protein was extracted from Bxpc-3 and Panc-1 cell lines using RIPA buffer supplemented with protease and phosphatase inhibitors (Roche Diagnostics; cat. no. 04906837001). Total protein was quantified using a BCA protein assay kit. Proteins (20 µg) were separated via SDS-PAGE (10–15% gel). The separated proteins were transferred to nitrocellulose membranes, which were then blocked with 5% non-fat milk in Tris-buffered saline-Tween-20 (TBST) overnight at 4°C. The membranes were incubated with primary antibodies against BRD4 (1:500; Abcam; cat. no. ab244221) and GAPDH (1:1,000; Cell Signaling Technology, Inc.; cat. no. 8884). The proteins were then incubated with secondary antibodies conjugated with horseradish peroxidase (OriGene Technologies, Inc.; cat. no. DS-0002) for 1 h at room temperature. The immunoreactive protein bands were visualized using enhanced chemiluminescence (ECL kit; Santa Cruz Biotechnology) and scanned using Amersham Imager 600 (GE Healthcare). Western blotting experiments were performed three times.
Dual-luciferase assay
A dual-luciferase reporter assay was performed in order to determine the effect of miR-608 on the BRD4 3′ untranslated region (3′UTR) and the functional binding sites in the BRD4 3′UTR. BRD4-3′UTR-Wild type (Wt) and BRD4-3′UTR-Mutant (Mut; all from Shanghai GeneChem Co., Ltd.)] were produced by sub-cloning the full-length 3′UTR fragments of the BRD4 gene and its mutant into the miR-608 binding sites using a pmirGlo Dual-Luciferase miRNA target expression vector (Promega Corporation). 293T (ATCC), Panc-1 and Bxpc-3 cells were seeded into 96-well plates (5×103 cells/well). Following incubation for 24 h at 37°C, the cells were co-transfected with either BRD4-3′UTR-Wt, BRD4-3′UTR-Mut, miR-608 or miR-608 NC. The cells were divided into three groups as follows: miR-608 negative control (NC) + BRD4-3′UTR-Wt; miR-608 + BRD4-3′UTR-Wt; and miR-608 + BRD4-3′UTR-Mut. Following transfection for 24 h, luciferase assays were performed using the Dual-Luciferase Reporter Assay system (Promega Corporation), and the firefly luciferase activity was normalized to Renilla luciferase activity.
BRD4 rescue experiment
The BRD4 plasmid was purchased from Shanghai Genechem Co., Ltd. and short hairpin (sh) BRD4 was purchased from Guangzhou RiboBio Co., Ltd. The experiment included four groups: miR-608 mimics + NC plasmid; miR-608 mimics + BRD4 plasmid; NC mimics + siBRD4 plasmid; and NC mimics + NC plasmid. Each group of mimics and BRD4 were co-transfected into Panc-1 and Bxpc-3 cells. The cells were harvested 24 h after transfection and were analyzed via flow cytometry. The sequence of shBRD4 was as follows: 5′-GATCCGCCTGGAGATGACATAGTCTTATTCAAGAGATAAGACTATGTCATCTCCAGGTTTTTTC-3′
Statistical analysis
Statistical analysis was performed using SPSS software (version 16.0; SPSS, Inc.). Receiver operating characteristic (ROC) curves were constructed and the area under the curve (AUC) was calculated in order to estimate the OS times for the selected candidate genes. The difference between two groups was assessed using the Student's t-test and one-way ANOVA followed by Bonferroni post hos test was used for comparisons between multiple groups. Kaplan-Meier analysis was utilized to generate survival curves and these were compared using the log-rank test. P<0.05 was considered to indicate a statistically significant difference.
Results
Screening for differentially expressed miRNAs (DEMs) between patients with pancreatic cancer exhibiting different prognoses using a gene expression microarray
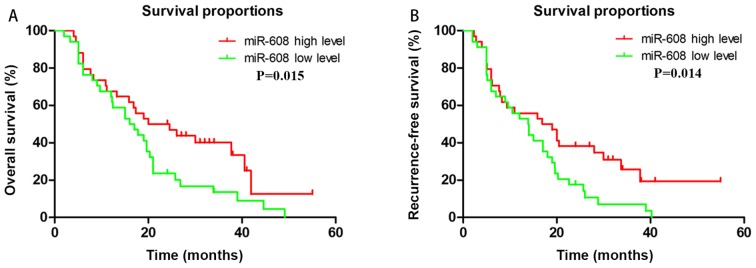
In order to identify the potential prognostic markers correlated with OS time, a microarray analysis was conducted on samples from six patients with PDAC. Compared with the short-survival group (OS, <12 months), 390 miRNAs (65.9%) were upregulated and 201 (34.1%) were downregulated (|fold change| >1) in the long-survival group (OS, >30 months; data not shown). Combined with our preliminary study (13) and associated literature retrieved from PubMed, 10 miRNAs (miRNA-21, −106b, −186, −192, −221, −222, −409, −492, −573 and −608) were selected to investigate their association with OS in a population of 68 patients with PDAC. The expression levels of these 10 miRNAs were detected using qPCR, and a Kaplan-Meier plot was constructed in order to analyze the association between the miRNA expression levels and OS. It was revealed that only miRNA-608 expression was able to accurately predict patient outcomes. The results demonstrated that high expression of miRNA-608 was associated with improved OS (P=0.015) and RFS (P=0.014; Fig. 1).
Figure 1.
Association between miRNA-608 expression and the prognosis of patients with pancreatic cancer. Expression level of miRNA-608 was determined using quantitative PCR, and patients were divided into a high expression group and low expression group (cut-off is median) based on the median expression value. Survival curve was generated using the Kaplan-Meier method. High expression levels of miRNA-608 suggested a longer prognosis in both (A) overall survival (P=0.015) and (B) recurrence-free survival (P=0.014). miRNA, microRNA.
miRNA-608 induces apoptosis in pancreatic cancer cell lines
miRNA-608 was associated with the prognosis of patients with PDAC. However, the mechanism behind the association between miR-608 and pancreatic cancer was unclear. miRNAs post-transcriptionally regulate their target genes, thus, 3 online bioinformatics tools were utilized to estimate potential target genes; TargetScan, miRDB and miRanda websites yielded 5,267, 916 and 2,840 possible miR-608 targets, respectively. R Project was then used to obtain 638 genes from all three bioinformatics tools. Subsequently, the online bioinformatics tool RNAhybrid indicated that there were high scoring binding sites from miRNA-608 in the 3′-UTR of BRD4 mRNA. It was previously reported that BRD4 serves a role in the modulation of the transcription of certain essential genes associated with apoptosis, including c-Myc and BCL2 (14). These data suggested that miRNA-608 was primarily associated with ‘apoptosis’ and ‘proliferation’ via the targeting of BRD4.
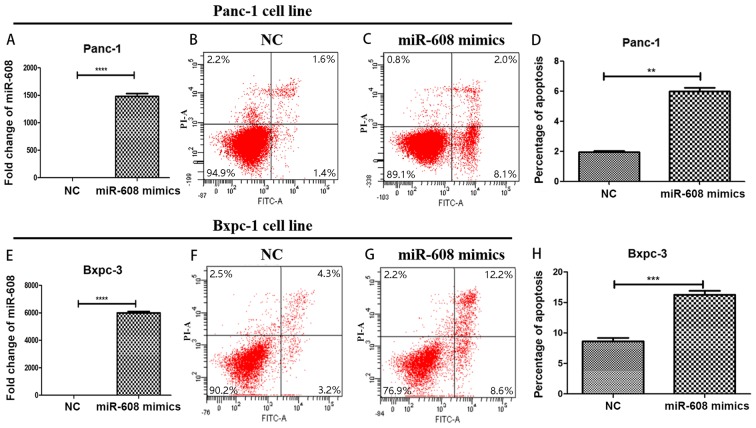
In order to verify the results predicted using bioinformatics, miRNA-608 was overexpressed in two pancreatic cancer cell lines: Panc-1 and Bxpc-3, using mimics. qPCR was used to confirm the transfection success and was compared with the NC group. The expression level of miRNA-608 in the mimic group was significantly higher. The fold changes were increased by ~1,500 and ~6,000 times in Panc-1 and Bxpc-3, respectively (P<0.0001). Flow cytometry was then performed in order to detect the function of miRNA-608, and it was revealed that a high level of miRNA-608 expression promoted apoptosis in both cell lines used (Fig. 2).
Figure 2.
miRNA-608 induces apoptosis in pancreatic cancer cell lines. (A and E) miRNA-608 was overexpressed in the (A) Panc-1 and (E) Bxpc-3 cell lines, and apoptosis was detected using flow cytometry. Transfection was confirmed in both the Panc-1 and Bxpc-3 cell lines. (B, C, F and G) Compared with the NC group, the (B and F) miRNA-608 mimic group had a significantly higher level of apoptosis in both Panc-1 and Bxpc-3 cell lines. (C and G) The statistical results of flow cytometry in the Panc-1 cell line. (D and H) Compared with the NC group, the miRNA-608 mimics group had a significantly higher level of apoptosis in both Bxpc-3 and panc-1 cell lines. Statistical results of flow cytometry in the Panc-1 and Bxpc-3 cell lines. **P<0.01, ***P<0.001 and ****P<0.0001. miRNA, microRNA; NC, negative control.
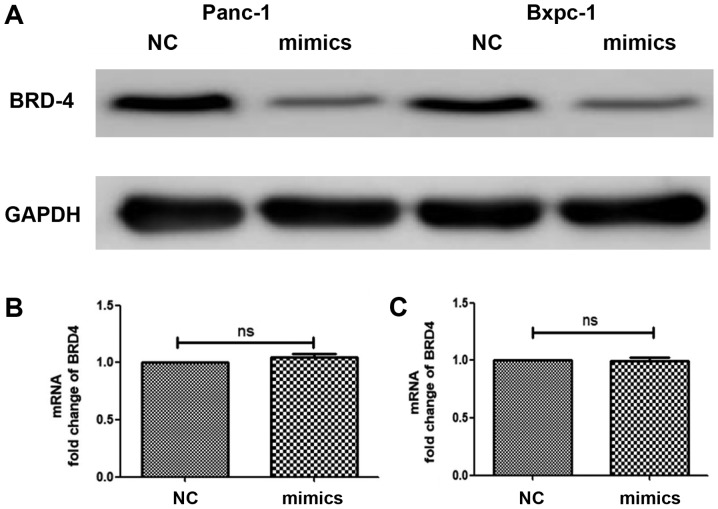
Furthermore, mimics were used to increase miRNA-608 expression and cells were collected in order to detect the expression level of BRD4 in both transcription and translation processes, using qPCR and western blot analyses, respectively. It was revealed that the protein levels of BRD4 in Panc-1 and Bxpc-3 were significantly decreased following overexpression of miRNA-608, but there was no change in the BRD4 mRNA level (Fig. 3).
Figure 3.
miRNA-608 regulates BRD4 expression. Overexpression of miRNA-608 was achieved using mimics in the Panc-1 and Bxpc-3 cell lines. BRD4 expression was analyzed at the mRNA and protein levels using quantitative PCR and western blotting, respectively. (A) High-expression level of miRNA-608 decreases BRD4 expression at the protein level. (B and C) There were no significant differences in the mRNA level of BRD4 by changing miRNA-608 expression. miRNA, microRNA; NC, negative control; BRD4, bromodomain-containing protein 4.
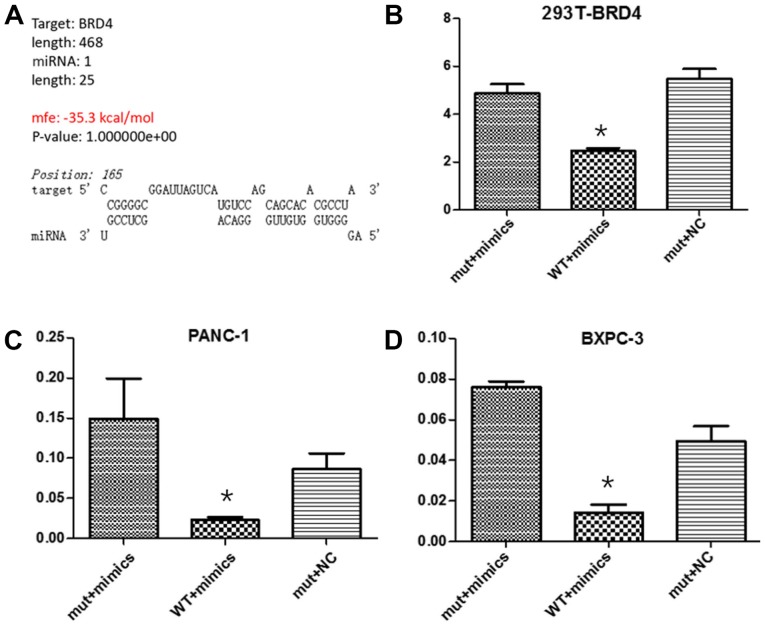
In order to investigate miRNA-608 regulation of BRD4 after transcription, the fragment of the BRD4 3′-UTR containing the predicted wild-type miRNA-608 binding site was incorporated into the downstream region of firefly luciferase, in the Dual-Luciferase miRNA Target Expression Vector, and a vector with the corresponding mutated binding sites was simultaneously constructed. The relative luciferase activity in 293T, Panc-1 and Bxpc-3 cells co-transfected with miRNA-608 mimic and a Wt vector was inhibited, while in cells transfected with the miR-608 mimic mutant vector, luciferase activity was unaffected (Fig. 4). The results of the present study supports the hypothesis that miRNA-608 may directly inhibit the expression of BRD4, via binding to the 3′-UTR of its mRNA.
Figure 4.
miR-608 decreases the BRD4 protein level by directly targeting the 3′-UTR. A dual-luciferase reporting assay was performed to detect that miRNA-608 may directly inhibit the expression of BRD4 by binding to the 3′-UTR of its mRNA in Panc-1 and Bxpc-3 cell lines. (A) Online biological information predicted the binding site and capacity between miRNA-608 and BRD4. The relative luciferase activity in (B) 293T cells, (C) Panc-1 cells and (D) Bxpc-3 cells, co-transfected with miR-608 mimics and the WT vector, were significantly inhibited, while in cells simultaneously transfected with the Mut type vector and miR-608 mimics or NC mimics, luciferase activity was unaffected. *P<0.05. miRNA, microRNA; NC, negative control; WT, wild type; Mut, mutant; BRD4, bromodomain-containing protein 4; 3′-UTR, 3′-untranslated region.
Suppression of BRD4 serves a crucial role in miRNA-608-induced apoptosis
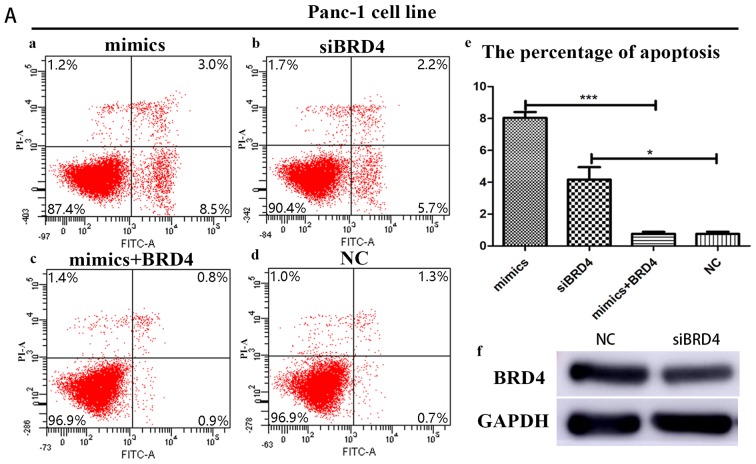
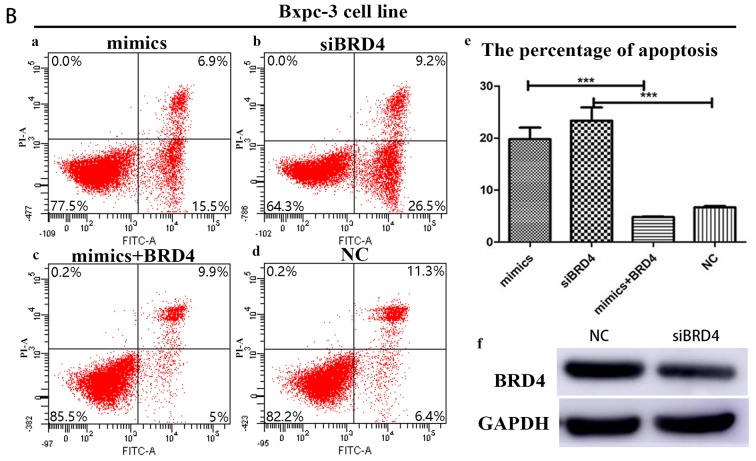
In order to further confirm the influence of BRD4 on miRNA-608-induced apoptosis, Panc-1 and Bxpc-3 cells were transfected with siBRD4 in order to specifically knock-down the expression of BRD4. Transfection with BRD4 increased apoptosis in both cell lines, a result consistent with the effects of miRNA-608 overexpression, using mimics. Finally, Panc-1 and Bxpc-3 cells were transfected with both the miRNA-608 and BRD4 plasmid, and upregulation of BRD4 decreased the miRNA-608-mediated induction of apoptosis (Fig. 5).
Figure 5.
(Ab and Bb) Suppression of BRD4 influences miR-608-induced apoptosis. (Aa-f) In Panc-1 cell line, (Ad and Bd) compared with the control (NC group), (Aa) overexpression of miR-608 or (Ab) suppression of BRD4 separately as mimics or siBRD4 groups may significantly increase the level of apoptosis. (Ac) Simultaneously increasing the levels of miRNA-608 and BRD4 mRNA may significantly decrease the level of apoptosis induced by the overexpression of miRNA-608. (Ae) There is the statistical result of the percentage of apoptosis in different level of miR-608 and BRD4 expression. (Af) It also shows the knock down result of siBRD4 in protein level. (Ba-f) In the Bxpc-3 cell line, similar results are obtained. (Ba) Overexpression of miR-608 (mimics group) or (Bb) suppression of BRD4 (siBRD4 group) significantly increase the level of apoptosis than that in (Bd) control (NC group). (Bc) Overexpression of BRD4 may subtract the increased apoptosis induced by miR-608. (Be and f) There are the percentage of apoptosis and BRD4 knock down efficiency. *P<0.05 and ***P<0.001. miRNA, microRNA. siBRD4, small interfering bromodomain-containing protein 4.
Discussion
The present study demonstrated that miRNA-608 is associated with the prognosis of patients with PDAC; a high level of miRNA-608 expression was associated with a longer OS time. In addition, miRNA-608 exhibited a low expression level in cancerous tissues compared with adjacent paracancerous tissues, and the overexpression of miRNA-608 promoted the apoptosis of pancreatic cancer cells. Furthermore, the results of the dual-luciferase reporter assay and bioinformatics database analysis revealed that miRNA-608 targets the BRD4 gene, which results in the downregulation of BRD4 expression. Finally, using a rescue assay, it was determined that miRNA-608 serves a tumor-suppressive role via the targeting of BRD4.
Pancreatic cancer is highly malignant and has a low 5-year OS rate, which often results in patient mortality only several months post-diagnosis (1). However, in TMUCIH, it was observed that in patients at the same pathological stage, certain patients exhibited significantly longer OS times compared with others. The present study aimed to elucidate the causal factor behind this difference, improving the understanding of disease progression and potentially conferring improvements on the outcome of patients with PDAC.
Numerous studies have revealed the capacity of miRNAs to serve as promising therapeutic targets, and to be significant diagnostic or prognostic biomarkers in the treatment of malignant tumors (15–17). In addition, miRNA-608 has been widely reported to be a tumor suppressor gene and is downregulated in multiple types of malignant tumor, including glioma, bladder, colon, liver and lung cancer (18–22). However, the function of miRNA-608 in pancreatic cancer progression is yet to be determined.
In the present study, tissue samples were collected from six patients with similar clinicopathological characteristics, but exhibiting significantly different prognostic outcomes. The samples were analyzed using a whole-genome expression microarray and a total of 10 miRNAs, with significant fold-changes between patient groups, were selected for further analyses. A limitation of the present study was that the microarray was conducted on a small sample population size (n=6), and that the groups did not have an even sex distribution. Thus, future studies should be conducted on a larger population size, with an even sex distribution.
The 10 pre-identified DEMs were verified using another population comprising 68 patients with PDAC. qPCR analysis revealed that miRNA-608 was associated with OS time. In addition, miRNA-608 expression level was significantly decreased in cancerous tissues compared with paracancerous tissues (68 vs. 11 tissues; P<0.05; data not shown). However, the mechanism behind the effect of miRNA-608 on pancreatic cancer is yet to be determined. Therefore, overexpression of miRNA-608 was induced in Panc-1 and Bxpc-3 cells in order to investigate the biological function of miRNA-608, and it was revealed that the rate of apoptosis was increased in the miRNA-608-overexpression group. Furthermore, it was revealed that miR-608 also inhibited the proliferation of pancreatic cells, but the association was not significant. Furthermore, there was no significant association between cell cycle stage and expression level of miR-608. Consequently, the results concerning proliferation and cell cycle are not shown. Subsequent experiments determined that miRNA-608 may promote apoptosis in pancreatic cancer cells via the targeting of the bromodomain and extra-terminal (BET) family protein, BRD4. BRD4 contains two tandem bromodomains that recognize acetylated-lysine residues in nucleosome histones, facilitating the recruitment of transcriptional proteins to chromatin (23). Furthermore, BRD4 may downregulate MYC Proto-Oncogene, BHLH transcription factor (24), which is a well-characterized apoptotic gene.
Certain studies have suggested that miRNAs regulate gene expression by serving as molecular sponges, resulting in mRNA degradation. This indicates that miRNA-mRNA interactions with high degrees of complementarity result in mRNA degradation (25). Despite this, it has been hypothesized that miRNA-mediated inhibition of expression may occur following the initiation of translation, but not transcription. miRNA has been revealed to decrease the rate of translation, and ribosomes are able to prematurely terminate the translation of miRNA-targeted mRNAs. Following miRNA-mediated mRNA translation, miRNA is often rapidly hydrolyzed; however, the full extent of the influence of miRNAs on protein expression levels, after the initiation of translation, is yet to be elucidated (26). Notably, in the present study, overexpression of miRNA-608 (via transfection with mimics in pancreatic cancer cell lines) caused the protein level of BRD4 to decrease significantly; however, no change in mRNA level was observed following qPCR analysis.
In conclusion, the present study revealed that miRNA-608 is associated with OS times in patients with pancreatic cancer, and that a high-expression level of miRNA-608 is associated with longer OS times. It was also discovered that miRNA-608 may promote apoptosis in pancreatic cancer cells via the targeting of the BET family protein, BRD4. The results indicate that miRNA-608 may represent a potential target for the treatment of human pancreatic cancer.
Acknowledgements
Not applicable.
Funding
This study was supported by the Key Projects of Tianjin Science and Technology Support Program (grant no. 13ZCZCSY20300).
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Authors' contributions
ML performed the experiment and analyzed data. TL interpreted the data. WM collected the samples and patients' data. XW interpreted the clinical and survival analysis. GZ interpreted the data, wrote and revised the manuscript. All authors read and approved the final version of the manuscript.
Ethics approval and consent to participate
This study was approved by the Ethics Committee of Tianjin Medical University Cancer Institute and Hospital. All patients or their family members were fully informed and consented to this study.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
References
- 1.Hidalgo M. Pancreatic cancer. N Engl J Med. 2010;362:1605–1617. doi: 10.1056/NEJMra0901557. [DOI] [PubMed] [Google Scholar]
- 2.Siegel RL, Miller KD, Jemal A. Cancer Statistics, 2017. CA Cancer J Clin. 2017;67:7–30. doi: 10.3322/caac.21387. [DOI] [PubMed] [Google Scholar]
- 3.Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64:9–29. doi: 10.3322/caac.21208. [DOI] [PubMed] [Google Scholar]
- 4.Bartel DP. MicroRNAs: Target recognition and regulatory functions. Cell. 2009;136:215–233. doi: 10.1016/j.cell.2009.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Berezikov E, van Tetering G, Verheul M, van de Belt J, van Laake L, Vos J, Verloop R, van de Wetering M, Guryev V, Takada S, et al. Many novel mammalian microRNA candidates identified by extensive cloning and RAKE analysis. Genome Res. 2006;16:1289–1298. doi: 10.1101/gr.5159906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120:15–20. doi: 10.1016/j.cell.2004.12.035. [DOI] [PubMed] [Google Scholar]
- 7.Hata A, Lieberman J. Dysregulation of microRNA biogenesis and gene silencing in cancer. Sci Signal. 2015;8:re3. doi: 10.1126/scisignal.2005825. [DOI] [PubMed] [Google Scholar]
- 8.Stratford JK, Bentrem DJ, Anderson JM, Fan C, Volmar KA, Marron JS, Routh ED, Caskey LS, Samuel JC, Der CJ, et al. A six-gene signature predicts survival of patients with localized pancreatic ductal adenocarcinoma. PLoS Med. 2010;7:e1000307. doi: 10.1371/journal.pmed.1000307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Collisson EA, Sadanandam A, Olson P, Gibb WJ, Truitt M, Gu S, Cooc J, Weinkle J, Kim GE, Jakkula L, et al. Subtypes of pancreatic ductal adenocarcinoma and their differing responses to therapy. Nat Med. 2011;17:500–503. doi: 10.1038/nm.2344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Donahue TR, Tran LM, Hill R, Li Y, Kovochich A, Calvopina JH, Patel SG, Wu N, Hindoyan A, Farrell JJ, et al. Integrative survival-based molecular profiling of human pancreatic cancer. Clin Cancer Res. 2012;18:1352–1363. doi: 10.1158/1078-0432.CCR-11-1539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.AJCC 7th Edition Cancer Staging Manual. https://cancerstaging.org/references-tools/deskreferences/Documents/AJCC%207th%20Ed%20Cancer%20Staging%20Manual.pdf. [Apr 10;2019 ];
- 12.Li D, Liu H, Li Y, Yang M, Qu C, Zhang Y, Liu Y, Zhang X. Identification of suitable endogenous control genes for quantitative RT-PCR analysis of miRNA in bovine solid tissues. Mol Biol Rep. 2014;41:6475–6480. doi: 10.1007/s11033-014-3530-x. [DOI] [PubMed] [Google Scholar]
- 13.Ma W, Li T, Wu S, Li J, Wang X, Li H. LOX and ACSL5 as potential relapse markers for pancreatic cancer patients. Cancer Biol Ther. 2019;20:787–798. doi: 10.1080/15384047.2018.1564565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jung M, Gelato KA, Fernández-Montalván A, Siegel S, Haendler B. Targeting BET bromodomains for cancer treatment. Epigenomics. 2015;7:487–501. doi: 10.2217/epi.14.91. [DOI] [PubMed] [Google Scholar]
- 15.Sahraei M, Chaube B, Liu Y, Sun J, Kaplan A, Price NL, Ding W, Oyaghire S, García-Milian R, Mehta S, et al. Suppressing miR-21 activity in tumor-associated macrophages promotes an antitumor immune response. J Clin Invest. 2019;129:5518–5536. doi: 10.1172/JCI127125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Juang V, Chang CH, Wang CS, Wang HE, Lo YL. pH-Responsive PEG-shedding and targeting peptide-modified nanoparticles for dual-delivery of irinotecan and microRNA to enhance tumor-specific therapy. Small. 2019;15:e1903296. doi: 10.1002/smll.201903296. [DOI] [PubMed] [Google Scholar]
- 17.Basu A, Jiang X, Negrini M, Haldar S. MicroRNA-mediated regulation of pancreatic cancer cell proliferation. Oncol Lett. 2010;1:565–568. doi: 10.3892/ol_00000100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Liang Z, Wang X, Xu X, et al. MicroRNA-608 inhibits proliferation of bladder cancer via AKT/FOXO3a signaling pathway. Mol Cancer. 2017;16:96. doi: 10.1186/s12943-017-0664-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Yang H, Li Q, Niu J, Li B, Jiang D, Wan Z, Yang Q, Jiang F, Wei P, Bai S. microRNA-342-5p and miR-608 inhibit colon cancer tumorigenesis by targeting NAA10. Oncotarget. 2016;7:2709–2720. doi: 10.18632/oncotarget.6458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wang Z, Xue Y, Wang P, Zhu J, Ma J. MiR-608 inhibits the migration and invasion of glioma stem cells by targeting macrophage migration inhibitory factor. Oncol Rep. 2016;35:2733–2742. doi: 10.3892/or.2016.4652. [DOI] [PubMed] [Google Scholar]
- 21.Wang K, Liang Q, Wei L, Zhang W, Zhu P. MicroRNA-608 acts as a prognostic marker and inhibits the cell proliferation in hepatocellular carcinoma by macrophage migration inhibitory factor. Tumour Biol. 2016;37:3823–3830. doi: 10.1007/s13277-015-4213-5. [DOI] [PubMed] [Google Scholar]
- 22.Othman N, Nagoor NH. miR-608 regulates apoptosis in human lung adenocarcinoma via regulation of AKT2. Int J Oncol. 2017;51:1757–1764. doi: 10.3892/ijo.2017.4174. [DOI] [PubMed] [Google Scholar]
- 23.Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith WB, Fedorov O, Morse EM, Keates T, Hickman TT, Felletar I, et al. Selective inhibition of BET bromodomains. Nature. 2010;468:1067–1073. doi: 10.1038/nature09504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Delmore JE, Issa GC, Lemieux ME, et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell. 2011;146:904–917. doi: 10.1016/j.cell.2011.08.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Lu TX, Rothenberg ME. MicroRNA. J Allergy Clin Immunol. 2018;141:1202–1207. doi: 10.1016/j.jaci.2017.08.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Seggerson K, Tang L, Moss EG. Two genetic circuits repress the Caenorhabditis elegans heterochronic gene lin-28 after translation initiation. Dev Biol. 2002;243:215–225. doi: 10.1006/dbio.2001.0563. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.