Abstract
The interior of eukaryotic cells is mysterious. How do the large communities of macromolecular machines interact with each other? How do the structures and positions of these nanoscopic entities respond to new stimuli? Questions like these can now be answered with the help of a method called electron cryotomography (cryo-ET). Cryo-ET will ultimately reveal the inner workings of a cell at the protein, secondary structure, and perhaps even side-chain levels. Combined with genetic or pharmacological perturbation, cryo-ET will allow us to answer previously unimaginable questions, such as how structure, biochemistry, and forces are related in situ. Because it bridges structural biology and cell biology, cryo-ET is indispensable for structural cell biology—the study of the 3-D macromolecular structure of cells. Here we discuss some of the key ideas, strategies, auxiliary techniques, and innovations that an aspiring structural cell biologist will consider when planning to ask bold questions.
INTRODUCTION
Electron cryomicroscopy (cryo-EM) is a family of methods that is used to study biological structure, from atoms to cells. To preserve a biological sample’s molecular details, the cryo-EM sample is kept free of the chemical fixation, dehydration, and heavy-metal staining that are commonly used in traditional EM. Furthermore, the sample is frozen so quickly that water molecules are immobilized in an amorphous state, which keeps the biological material “hydrated.” Once frozen, the sample must be kept colder than −135°C before and during image acquisition in a transmission electron cryomicroscope. Because biological matter is damaged rapidly by the microscope’s electron beam, cryo-EM imaging is done using a very limited electron dose. The resulting low-dose images are noisy and therefore require careful processing and interpretation. These stringent requirements ensure that the cryo-EM data represent biological structures in a minimally perturbed, lifelike state.
Two popular forms of cryo-EM are “single-particle” analysis (SPA) and electron cryotomography (cryo-ET). Structural biologists have used both these approaches to study macromolecular complexes, herein called complexes for brevity. SPA has revolutionized the structure determination of purified complexes. Because this method can routinely produce 3-D density maps in which protein side chains are resolved, SPA is a popular replacement for X-ray crystallography studies of complexes larger than ∼100 kDa. In contrast, cryo-ET is a method that generates modest ∼40 Å resolution 3-D density maps of complexes, organelles, and even cells. If multiple copies of a complex are identified, they can be analyzed by subtomogram averaging (STA). STA involves aligning and averaging multiple copies of the same complex, thereby increasing the signal-to-noise ratio and suppressing the missing-wedge artifacts (detailed later), resulting in higher-resolution maps (Asano et al., 2016; Oikonomou and Jensen, 2017; Hutchings and Zanetti, 2018; Schur, 2019).
This Perspective is written for cell biologists who are interested in learning and applying cryo-ET to their favorite eukaryotic systems. To cover a broad range of ideas, we have skipped some technical details, which are found in other literature and online training resources (Henderson, 1995; Koster et al., 1997; Frank, 2006a,b; Dubochet, 2007; Penczek, 2010; Shen and Iwasa, 2018; Vos and Jensen, 2018; Rodenburg, 2019). We discuss some nonintuitive principles of cryo-ET and our expectations from current and future technologies. We also review some studies that use artificially thin samples like cryosections and cryolamellae because such samples vastly expand the range of questions that can be asked. Finally, we explore how cryo-ET can be used to study large cellular machines that defy conventional approaches.
TOMOGRAPHY AND TOMOGRAMS
Because heavy-metal stains are not used, cryo-EM image features come from the atoms within proteins, nucleic acids, and other biological molecules. Cryo-EM images are approximations of 2-D “projections” of 3-D objects, meaning that the pixel values are proportional to the mass summed along parallel linear paths through the sample (Figure 1A). Real-world projections deviate from ideal projections because they are influenced by the transfer functions of the microscope and the camera system. Projection images encode information about the interior of the object, making them fundamentally different from shadows (Figure 1B). The pixel values, called densities are usually represented in grayscale. Note that the term “densities” is contextual and can also mean 3-D pixel (voxel) intensity values or groups of pixels (or voxels) that belong to the same complex.
FIGURE 1:
The principles of cryo-EM and cryo-ET. (A) The example object is “NUS” in block letters, centered within a thin spherical shell. The object’s projection image (bounded by a trapezoid) is shown below the Cartesian coordinates symbol. Electrons (e−, white arrow) travel along the z-axis. To show the object’s orientation, the front of the shell has been removed. (B) Cryo-EM images are projections, not shadows. Density profiles (ρ) along the dashed lines (x-axis) are shown in the plots below. (C) In a cryo-ET experiment, the sample is rotated around the tilt axis (y-axis in this example), typically over a tilt range from −60° to +60°. Five possible tilted orientations are shown. (D) A tilt series consists of the set of cryo-EM images recorded over this tilt range. Each image corresponds to the object, as oriented directly above in C. The image gray levels are proportional to the samples’ projected mass along the z-axis. Together, the images encode the information needed for a 3-D reconstruction. Supplemental Movie S1 shows the full tilt series. (E) After alignment, the images (bounded by trapezoids) are oriented according to their corresponding tilt angles and then “back projected” to generate a 3-D density map called a cryotomogram. The limited tilt range of (±60°) results in missing-wedge artifacts, which manifest as image distortions. These distortions include the triangular features at 6 and 12 o’clock, the spokelike feature protruding from the bottom of the U, and the poorly defined lower portion of the letter S.
Movie S1.
Understanding tilt series This movie shows a structure, “NUS” enclosed within a spherical shell, and its projection images, corresponding to Figure 1. Contains an audio narration and subtitles.
A cryo-ET raw data set is a series of images, typically recorded over a tilt range from −60° to +60° (±60°), with tilt increments ranging from 1° to 4°. This data set has discrete angular sampling and is called a tilt series (Figure 1, C and D, and Supplemental Movie S1). Tilt-series images are mutually aligned to a common 3-D reference model. This alignment is done using image features from the cell itself or with the aid of fiducial markers, such as gold colloids that were added prior to sample freezing. The aligned images are then typically combined by an algorithm called “back projection” to produce the cryotomogram, a 3-D representation of the original sample (Figure 1E). Note that there are other cryotomogram reconstruction algorithms (Frank, 2006a).
If we ignore radiation-damage effects, the reconstruction of an isotropically resolved 3-D volume of a sample requires images of samples tilted up to ±90°. The typical usable tilt range in cryo-ET, however, is approximately ±60°. This limitation stems from cryo-EM samples’ slab-like geometry, which increases the electron-optical path length by 1/cosine(tilt angle). Furthermore, most cryo-EM samples, commercial cryo sample holders, and microscopes are not compatible with tilts much higher than 70°. Exotic cylindrical samples such as bacterial cells loaded into a sub-micron-diameter carbon nanopipette do not have this limitation (Palmer and Lowe, 2014). Most cryotomograms are therefore missing approximately 33% of a complete tilt-series data set, which causes the cryotomogram to appear distorted relative to the original structure (Figure 1, A and E).
When visualized in Fourier space, the missing cryo-ET data appear like a pair of empty wedges that intersect along a line that runs parallel to the tilt axis (Figure 2A). This missing wedge limits the resolution along the electron-optical “Z” axis and results in elongated and smeared density-map features. We illustrate these distortions using the Dam1C/DASH outer-kinetochore complex and the nucleosome, which we have simulated with 60° of data missing (Figure 2, B and C). Supplemental Movies S2 and S3 illustrate the bizarre appearance of cryotomograms when these complexes are rotated relative to a stationary missing wedge. Fortunately, many subtomogram-analysis programs have routines that account for the missing wedge during the alignment process. Within the cell, most known complexes are randomly oriented. When multiple copies of the complex are aligned, their respective missing wedges will be randomly oriented. The Fourier transform of the resulting average has the missing data filled in, meaning that the reconstruction no longer has missing-wedge artifacts.
FIGURE 2:

The missing wedge influences cryotomograms in unintuitive ways. The left subpanels correspond to a “complete” data set (180° tilt range) while the right subpanels correspond to an incomplete data set, with a 60° wedge of data missing. The tilt axis is parallel to the y-axis, and the electron beam is parallel to the z-axis. (A) Volumetric representations of cryo-ET data in Fourier space; kx, ky, and kz denote the Fourier-space coordinate system, which is aligned to the real-space coordinate system shown in the other panels. The right panel shows the two missing wedges, bounded by dashed lines. (B) Outer-kinetochore Dam1C/DASH ring densities simulated at 12 Å resolution. The rings in the bottom panel are rotated 90° around the x-axis, relative to an unchanged missing wedge. From PDB 6CFZ (Jenni and Harrison, 2018). (C) Nucleosome densities, also simulated at 12 Å resolution and shown in two orientations like in B. From PDB 1KX5 (Davey et al., 2002). Supplemental Movies S2 and S3 show how the Dam1C/DASH ring and the nucleosome are distorted in other orientations relative to the missing wedge. Note that real cryotomograms are affected by additional factors not modeled here. These factors include defocus, radiation damage, and discrete angular sampling.
Movie S2.
Dam1C/DASH and missing-wedge effects This movie shows how the Dam1C/DASH outer-kinetochore ring complex's appearance is influenced by missing-wedge effects, corresponding to Figure 2. Contains an audio narration and subtitles.
Movie S3.
The nucleosome and missing-wedge effects This movie shows how the nucleosome's appearance is influenced by missing-wedge effects, corresponding to Figure 2. Contains an audio narration and subtitles.
How to learn cryo-ET
There are three major skill sets to cryo-ET. The first skill set is sample preparation, meaning the operation of rapid-freezing machines. Training in the two principal techniques of rapid freezing—plunge freezing and high-pressure freezing (Dahl and Staehelin, 1989; Dobro et al., 2010)—is typically offered by cryo-EM facility staff. The second skill set is the operation of cryo-EM hardware and software. Some cryo-EM facilities also offer hands-on training in cryomicroscope operation. The third skill set is image processing, which unlike the other two, can be self-taught with the assistance of online tutorials and “walk-throughs.” Recently, some academic organizations have started offering on-site courses in one or more of these critical skill sets. Training workshops have also been offered by cryo-EM centers such as Diamond eBIC (Diamond, 2019), NeCEN (Netherlands Centre for Electron Nanoscopy, www.necen.nl, 2019), and NYSBC (New York Structural Biology Center, https://nysbc.org, 2019). These intensive hands-on courses are usually announced on the cryo-EM mailing lists CCP-EM (CCP-EM, 2019) and 3DEM (Perkins, 2019) and the IMOD list (Mastronarde, 2019) and also on social media such as the Twitter platform.
New professionally produced web resources such as em-learning.com and cryoem101.org are making theoretical training more accessible (Shen and Iwasa, 2018; Vos and Jensen, 2018). In addition, the University of Colorado provides a step-by-step tomogram-reconstruction tutorial using their popular IMOD package (Mastronarde, 1997; O’Toole, 2018). This tutorial includes a tilt series of a negatively stained plastic section of a Chlamydomonas cell. The IMOD tutorial can be done in 1 day and is a gateway to the analysis of cryo-ET data, which are more challenging. Cellular cryo-ET data are accessible from the EMPIAR database (Iudin et al., 2016) and from the Jensen lab’s ETDB-Caltech database (Ortega et al., 2019). Furthermore, we have deposited our surplus data in EMPIAR-10227 (Gan et al., 2019). Surplus data are tomograms or positions within tomograms (and their corresponding tilt series) that contain cellular structures not presented or analyzed for a research paper. Resources like these enable new practitioners to gain proficiency in the essential steps of cryo-ET image processing from the comfort of their office.
How to decide on the suitability and feasibility of cellular cryo-ET
The three most popular techniques of cellular imaging are light microscopy, traditional EM, and cryo-ET. Each of these approaches has advantages (Table 1). The biggest advantage of cryo-ET is that it can provide structural information, which we define as the 3-D arrangement of image densities that correspond to the proteins, nucleic acids, and other biological material within the cell. Furthermore, cryo-ET is essential if the study requires 3-D localization at the nanometer level or if the complex being studied is either invisible or distorted in traditional and immuno-EM data. Superresolution light microscopy can in principle achieve 3-D localization at the nanometer level, but this positional information only applies to the fluorophore, which in the case of fluorescent proteins is displaced from the complex by a few nanometers. Note that many structural cell biology projects require two or more forms of imaging.
TABLE 1.
Comparison of popular cell biology microscopy methods
| Method | ||||
|---|---|---|---|---|
| Information | Example | LM | EM | Cryo-ET |
| Cytology | Architecture of mitochondrial network | + | + | |
| Location | Ribosome assembly factor in nucleoli | + | + | |
| Distance | Diameter of a membrane tubule | + | + | + |
| Volume | Enlargement of lipid bodies | + | + | + |
| Counts | Nucleosomes per unit volume | + | + | + |
| Positions | Ribosome redistribution under stress | + | + | + |
| Structure | 3-D structure of kinetochore subunits | + | ||
| Heterogeneity | Oligomerization states of proteasomes | + | ||
| Higher-order structure | Path and orientation of sequential nucleosomes and ribosomes; mammalian kinetochores | + | ||
LM, conventional and superresolution light microscopy; EM, traditional, immuno, and serial EM. Note that there are exceptions to these broad guidelines and that superresolution light microscopy may eventually be able to answer some of the questions that are now exclusive to cryo-ET.
The next questions concern feasibility. First, can the sample be cooled to a frozen-hydrated state by plunge freezing? Samples that are smaller than approximately 10 µm can be frozen without the formation of crystalline ice when plunged into a cryogen like liquid ethane or liquid propane-ethane mixture (Tivol et al., 2008). To increase the chances of complete vitrification (the absence of crystalline ice), the plunge-frozen sample should be as thin as possible. Small cells like yeast are amenable to plunge freezing. Larger plunge-frozen cells have vitreous ice in their thinner extremities and crystalline ice in their thicker parts. Samples up to nearly 1 mm thick are high-pressure-frozen in the presence of 2000 bars of pressure to ensure the ice is vitreous throughout their entire volume. The upper limit of such “thick” samples include cell pastes, tissues, and small animals, which are sometimes high-pressure-frozen in the presence of a cryoprotectant (Harapin et al., 2015; Schaffer et al., 2019).
Next, one needs to ask, what complex needs to be observed? Can these complexes be seen in a cellular cryotomogram? How abundant must these complexes be to answer the project’s main question? Are the complexes large and monolithic enough to be unambiguously identified? These questions do not have simple answers, so we will make some calculations using a few well-studied complexes. As a cellular sample, we will consider a 100-nm-thick cryosection cut through the center of an early-interphase yeast nucleus, which has a ∼2 µm diameter. In this example, the total nuclear volume is ∼4.2 µm3 and the cryotomogram contains ∼0.3 µm3, or approximately 7% of the nucleus.
Our first example is the nucleosome, a 200-kDa complex shaped like a cylinder, 10 nm wide and 6 nm thick. The nucleosome is currently one the smallest complexes that can be detected in a cellular cryotomogram. Yeast cells have ∼12 megabases of genomic DNA and a ∼160-base-pair-long nucleosome repeat length, meaning that there are ∼75,000 nucleosomes per nucleus. We therefore expect a yeast nucleus cryosection to contain ∼5000 nucleosomes per cryotomogram, making nucleosomes amenable to identification by a purification in silico approach. Our second example is the 66-MDa nuclear pore complex (Rout and Blobel, 1993), which is more than 96 nm wide and 35 nm thick (Yang et al., 1998). There are fewer than 100 nuclear pore complexes per yeast cell, so it is not surprising that cryotomograms (in our experience) rarely contain more than one complete complex. Nuclear pore complexes can nevertheless be easily located and then identified because they are conspicuously positioned at sites of inner- and outer-nuclear membrane fusion. Both nucleosomes and nuclear pore complexes have a largely random nuclear distribution, meaning that cryotomograms that contain a piece of the nucleus usually contain these two complexes.
Rare complexes whose structures are poorly characterized in situ are more challenging to study. One such example is the replication origin, which hosts numerous interesting macromolecular complexes. Each yeast cell contains ∼300 replication origins (Nieduszynski et al., 2007). We estimate that a yeast cryosection cryotomogram should contain ∼20 origins. Let’s assume that these origins can be identified (a different challenge discussed later) and that the replication subassembly of interest is conformationally and constitutionally homogeneous. Then 50 cryotomograms will provide ∼1000 origin-containing subtomograms, which is enough to generate an average at better than 30 Å resolution. Extremely rare complexes (averaging fewer than 1 per cryotomogram) that have nonrandom locations, such as kinetochores and microtubule-organizing centers, require alternative strategies that are discussed in later sections.
WORKFLOWS AND STRATEGIES
Preparation of thin frozen-hydrated cell samples
Cryo-ET samples typically are thinner than ∼500 nm, but may be thicker or much thinner depending on the resolution requirements and the microscope hardware (Grimm et al., 1998). Cryotomograms of ∼500-nm-thick samples have enough resolution to reveal the positions of membranes and large complexes such as ribosomes. STA generally requires samples thinner than 200 nm. Thin cellular samples can be prepared by cryomicrotomy or cryo-focused-ion-beam (cryo-FIB) milling, which produce cryosections and cryolamellae, respectively. Cryosections are typically ∼50–200 nm thick and are cut from a block of frozen-hydrated cells (or tissue) by a diamond knife cooled to ≤ −140°C. Cryomicrotomy was once considered feasible only for those with extremely steady hands and years of training. Advanced micromanipulation techniques have now made cryomicrotomy easier and more reproducible (Ladinsky et al., 2006; Studer et al., 2014). FIB milling is routinely used in materials science and has been painstakingly adapted for frozen-hydrated life science sample preparation (Marko et al., 2006; Hayles et al., 2007; Rigort et al., 2010; Villa et al., 2013; Mahamid et al., 2015; Medeiros et al., 2018). In a cryo-FIB milling experiment, the operator uses a beam of gallium ions to remove a frozen-hydrated cell mass to produce a thin plank-like sample suspended between two unmilled positions.
Cryomicrotomy has a few advantages over cryo-FIB milling. First, the instrument and the consumables are an order of magnitude cheaper. Second, cryomicrotomy can deal with tissues and small animals. Third, cryomicrotomy can generate cryo-ET samples faster than cryo-FIB milling. While cryomicrotomy is a more established technique, cryo-FIB milling has several advantages that are making it the more popular choice of cellular cryo-ET students. First, cryo-FIB milling does not cause compression and cutting artifacts like in cryomicrotomy (Dubochet et al., 2007). Cryo-FIB milling is therefore less likely to damage biological structures. Second, cryo-FIB milling is done within the vacuum of a computer-controlled scanning electron microscope, meaning that it suffers less contamination from room humidity, is less dependent on dexterity, and has better potential for automation. Cryomicrotomy and cryo-FIB milling can be combined to generate compression-free cryolamellae from macroscopic samples (Rigort et al., 2010; Zhang et al., 2019). Further advances in sample freezing (Ravelli et al., 2019) and lamellae preparation (Schaffer et al., 2019; Wolff et al., 2019) will greatly increase the diversity of milled cellular samples that can be studied by cryo-ET.
Searching for targets of interest
Cryo-ET imaging experiments begin with a search for grid positions that contain the cell(s) or complex(es) of interest. This search is done at low magnification to maximize the field of view and at ultra-low dose to minimize radiation damage prior to tilt-series collection. If the complex of interest is abundant, the microscopist can image cells at random. Rarer complexes can first be localized by fluorescence light cryomicroscopy. These positions are then imaged by cryo-EM (correlative light and electron cryomicroscopy, cryo-CLEM). In a cryo-CLEM experiment, the complex is tagged with a protein or molecule that is fluorescent at temperatures colder than −135°C (Hampton et al., 2017). Some fluorescent proteins are photoactivatable at these cryogenic temperatures, meaning that superresolution light cryomicroscopy is possible (Chang et al., 2014; Dahlberg et al., 2018; Tuijtel et al., 2019). Cryo-CLEM can facilitate the identification of complexes for which no SPA or crystal structure exists, as shown in a study of a bacterial compartmentalization complex (Schlimpert et al., 2012). Recently, cryo-CLEM of a membrane-associated GFP-fusion protein enabled the classification of yeast plasma-membrane domains called eisosomes (Bharat et al., 2018). Cryo-CLEM was also used to sample different positions associated with the actin-wave phenomenon in Dictyostelium cells (Jasnin et al., 2019). Cryo-CLEM will facilitate even more localizations (and possibly identification of the complex; see below) once superresolution light cryomicroscopy extends to 3-D and the workflow becomes routine.
Data collection
Modern electron microscopes are highly reliable and automated, driven by state-of-the-art software (Mastronarde, 2005; Suloway et al., 2009; Zheng et al., 2009). Hardware advances have also kept pace, enabling more structural details to be extracted from cellular samples. One major advance is the complementary metal–oxide semiconductor–based direct detector, which operates in “movie mode” and thereby enables partial correction of beam-induced motion blur (Brilot et al., 2012). The direct detectors with the fastest readouts make it possible to record data in which each imaging electron is registered. Such electron-counted data have the highest signal-to-noise ratios, which are evident in comparisons of the cameras’ detective quantum efficiencies (Ruskin et al., 2013). A second major advance is the phase plate, a device that increases the phase difference between the scattered and the unscattered electrons (Danev et al., 2014). The phase plate increases the low-resolution contrast, which makes complexes easier to detect. New developments in laser phase plates could make this technology more user-friendly and higher throughput than possible with the current carbon film-based phase plates (Schwartz et al., 2019). Cryosections and cryolamellae are thick by cryo-EM standards, meaning that they scatter more electrons inelastically. Commercially available cryomicroscopes cannot focus these lower-energy electrons at the image plane, leading to worse chromatic aberration. To deal with this problem, cryo-ET imaging should be done with an energy filter, which can remove the inelastically scattered electrons.
Data processing
Tomogram reconstruction is a multistep process, including motion correction, contrast transfer-function correction, image alignment, low-pass filtering, 3-D reconstruction, and denoising. These steps are all automated to some degree in the popular packages IMOD (Mastronarde, 1997) and EMAN2 (Chen et al., 2019). Automated tilt-series alignment without fiducials is also being addressed by software such as Protomo (Noble and Stagg, 2015). Following cryotomogram reconstruction, subcellular image features can be annotated either manually or semiautomatically using new machine-learning approaches (Chen et al., 2017; Zeng et al., 2018). Known complexes can be annotated semiautomatically by template matching (Heumann, 2016; Castaño-Díez, 2017). These annotations (also called segmentations) are the starting point for quantitative analysis. Note that cryo-ET data are commonly stored as MRC-formatted files (Cheng et al., 2015). Cryo-ET practitioners should write format-interconversion scripts if they want to adapt analysis programs from other microscopy fields.
Identification of complexes
Cryotomograms are noisy, making the identification of complexes (or their subunits) a major challenge in cryo-ET of cells. Large, abundant complexes can be identified by classification and STA, which produce density maps that resemble known SPA or crystal structures. The best-studied examples are ribosomes and proteasomes (Mahamid et al., 2016; Albert et al., 2017). In contrast, the identification of rare complexes is facilitated with the knowledge of their structure and intracellular location. For example, we identified Dam1C/DASH outer-kinetochore complexes based on their localization to the plus ends of kinetochore microtubules, their absence from cytoplasmic microtubules, and their unmistakable structural similarity to recombinant complexes (Ng et al., 2019). To identify subunits within a complex, each subunit can be either deleted or fused with a protein tag and then analyzed by STA (Oda and Kikkawa, 2013; Chang et al., 2016). Density-map subtraction of the mutant from wild-type densities will then reveal either a “positive” or a “negative” density feature that corresponds to the position of the deleted or tagged subunit, respectively.
Ideally, all copies of a complex should be identifiable without the need of STA or CLEM. This not-yet-invented technology is known colloquially as the “GFP of cryo-EM” and has several requirements. The tag must be a metal nanoparticle, which will be visible as a dense punctum. This tag must also be synthesized in situ without cell permeabilization. Finally, the GFP of cryo-EM must be small and inert enough that it does not compromise the function of the protein of interest. An example of such tags was recently demonstrated in fixed, plastic-embedded cells, which are compatible with in situ gold nanoparticle synthesis (Jiang et al., 2019). Some microorganisms can naturally synthesize metal nanoparticles (Komeili et al., 2006; Scheffel et al., 2006), meaning that the required chemistry is theoretically possible inside living cells. Reproducible, inducible nanoparticle synthesis in living cells will require ideas from inorganic chemistry and materials science. In short, there are exciting opportunities for breakthroughs in cryo-EM in situ labeling.
STA, classification, and analysis
Owing to radiation damage, the signal-to-noise ratio of cryotomograms beyond ∼40 Å resolution is very low. If multiple conformationally identical copies of a complex exist, they can be averaged together by STA, yielding a new map that has a higher signal-to-noise ratio. If the complex is conformationally heterogeneous, then some of the conformational states can be “purified in silico” by 2-D and 3-D classification. The combination of STA and classification bridges cell biology and structural biology and is being actively developed (Nicastro et al., 2006; Heymann et al., 2008; Bharat et al., 2015; Galaz-Montoya et al., 2015; Castaño-Díez, 2017; Himes and Zhang, 2018). Work done on HIV capsid subunits showed that subtomogram averages can exceed 4 Å resolution (Schur et al., 2016; Himes and Zhang, 2018). At these resolutions, amino acids with bulky side chains are recognizable and can thereafter be used as landmarks for building and/or refining an atomic model. Lower-resolution subtomogram averages are also valuable because they can be docked with atomic models from crystallography, NMR, or SPA.
Once a SPA or STA experiment produces an averaged structure, the relative translations and rotations between all contributing particles within an image become known. In SPA studies, these alignment data are rarely used for biological interpretations because in most ideal SPA samples, the complexes have random positions and orientations. In cell cryotomograms, however, the 3-D relationships between multiple complexes can encode biological information. To recover this information, the subtomogram average is copied and then remapped into each individual complex’s position and orientation within the cell (Forster et al., 2010). This method has revealed the interactions and distributions of complexes such as ribosomes (Mahamid et al., 2016) and proteasomes (Albert et al., 2017) and higher-order structures of polymers such as nucleosome chains (Cai et al., 2018a).
Expecting the unexpected
Cellular cryo-ET data are a rich source of biological surprises. In the recent study of fission yeast cytokinesis, cryo-CLEM clearly detected myosin at the division septum (Swulius et al., 2018). However, the cryotomograms of the myosin-positive positions did not reveal any structures with the size and morphology expected of myofilaments. This finding raises questions about how the actomyosin machinery pinches the two daughters apart. Our study of the Dam1C/DASH complex also revealed unexpected phenotypes, such as the absence of evidence of a globular inner-kinetochore complex coaxial with the kinetochore microtubule and the absence of contacts between Dam1C/DASH and the curved microtubule tips (Ng et al., 2019). Our data also showed that most Dam1C/DASH rings are incomplete, consistent with an earlier calibrated-fluorescence microscopy study (Dhatchinamoorthy et al., 2017). These findings challenged existing models of how the outer kinetochore anchors chromosomes to the plus ends of depolymerizing microtubules for chromosome segregation. As cryo-ET image quality improves, even more unexpected discoveries will be made in situ.
Studying function in situ
Because we know so little about the nanometer-level organization of eukaryotic cells, cryo-ET studies are often labeled as “descriptive”—they reveal new phenotypes and they inspire new hypotheses. To use cryo-ET as a more hypothesis-driven tool, one needs to perturb a cellular process and then characterize the phenotypic changes. Genetic perturbations in the form of deletions and temperature-sensitive mutants are available in model organisms. Such mutants may have unexpected phenotypes arising from adaptation to these mutations. To mitigate these potential artifacts, future studies will need to use conditional rapid mislocalization or knockdown technologies, such as anchor-away and the auxin-inducible degron (Haruki et al., 2008; Nishimura et al., 2009).
STRUCTURAL INSIGHTS FROM THE DEPTHS
Cryo-EM has come a long way since its birth four decades ago (Frank, 1975; Taylor and Glaeser, 1976; Dubochet and McDowall, 1981; Adrian et al., 1984; Henderson et al., 1986). Cryo-EM studies of eukaryotic cells were historically challenging because cells first had to be cryosectioned. Furthermore, the hardware and software needed for automation were still nascent at the turn of the century (Koster et al., 1997). Cryo-ET of eukaryotic cells only became feasible in 2002 (Hsieh et al., 2002; Medalia et al., 2002). Owing to the challenges of cryomicrotomy, early cryo-ET studies of eukaryotes were done primarily with plunge-frozen samples. These samples include picoplankton, which are the size of bacteria (Henderson et al., 2007), and cell parts thinner than 1 µm (Nicastro et al., 2005; Koning et al., 2008).
The eukaryotic chromatin field has been a major beneficiary of cryo-EM. Eukaryotic genes are packaged into chains of nucleosomes, each consisting of eight histone proteins that are wrapped by ∼147 base pairs of DNA (Kornberg, 1974; Luger et al., 1997). Sequential nucleosomes are thought to fold into higher-order structures that can either occlude or expose DNA sequences to the nucleoplasm. DNA accessibility controls important cellular functions such as transcription and replication, so the central question in the field is: what is the higher-order structure of chromatin (Maeshima et al., 2019)? The dominant model of higher-order structure was the 30-nm fiber, which portrayed nucleosome chains compacted into long helical fibers that have crystalline order. This model was conceived based on electron microscopy images of negatively stained chromatin fragments and X-ray scattering data of oriented chromatin fragments (Finch and Klug, 1976; Widom and Klug, 1985). The 30-nm fiber was not seriously challenged until cryo-EM of cells made it possible to visualize chromatin in situ, without traditional-EM-associated artifacts.
The first major challenge to the 30-nm-fiber model came from cryo-EM studies of sectioned CHO cells (Figure 3A). These pioneering analyses did not reveal any evidence that supported the 30-nm-fiber model in situ (McDowall, 1984; McDowall et al., 1986). Nevertheless, this model persisted, in part because X-ray scattering of purified chromosomes showed diffraction features corresponding to regular 30-nm structure; these diffraction features have since been shown to be artifacts from ribosomes that copurified with and coated the chromosomes (Nishino et al., 2012). Furthermore, the data from early cryo-EM studies of cellular cryosections were limited by “1980’s state-of-the-art technology,” including the microscope (poor beam stability) and image-recording media (film) (Chang et al., 1983; McDowall et al., 1983). The absence of evidence for abundant ordered helical mammalian chromatin was reproduced in HeLa cells using newer microscope hardware and newer image-processing techniques (Eltsov et al., 2008).
FIGURE 3:
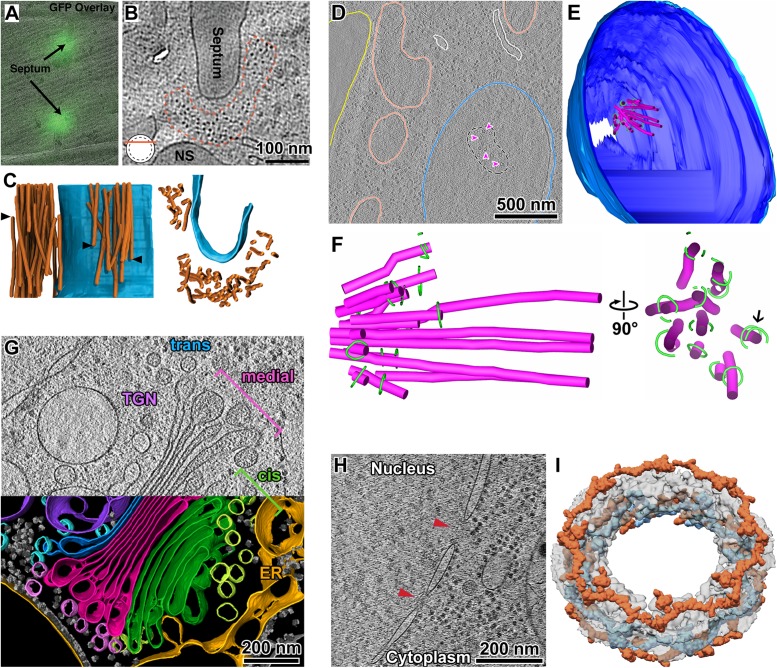
Structural cell biology of chromatin. (A) An early example projection image of a cryosectioned CHO cell (McDowall, 1984). Key cytological features include the nucleus (N), chromatin (Ch), nucleolus (Nu), and a nuclear pore complex (P). Cryomicrotomy artifacts such as knife marks (KM) and chatter (W) are also indicated. Image courtesy of A. McDowall. (B) Cryotomographic slice of a Saccharomyces cerevisiae nucleus, imaged with “defocus” phase contrast. The boxed region is enlarged twofold and shown in the top-right panel. This position is also rendered as isosurfaces in the bottom-right panel. Adapted from Chen et al. (2016). (C) Cryotomographic slice of a Schizosaccharomyces pombe nucleus imaged with “Volta” phase contrast. The bottom-left subpanel shows a threefold enlarged view of a representative nucleosome (white arrow) and a larger multi-MDa complex (megacomplex) in the top-left corner. Both complexes are from the position boxed in the main panel. The bottom-right subpanels are cartoons of the organization of nucleosomes and megacomplexes in G2 and prometaphase (PM) nuclei. Adapted from Cai et al. (2018b). (D) Cryotomographic slice of an embryonic Drosophila neuronal nucleus. Heterochromatin is marked out with white dashed lines and the nuclear pore complex (NPC) is labeled. Adapted from Eltsov et al. (2018). (E) Nucleosome remapping in a HeLa cell nucleus. Left: cryotomographic slice of a HeLa cell nucleus. The regions with higher local nucleosome concentrations are marked out with purple dashed lines. The white arrow indicates a nuclear pore complex. Right: 3-D annotation of the same cryotomogram after nucleosome remapping. (F) Arrangement and organization of di- and trinucleosomes in the HeLa cell nucleus. The two nucleosome conformational classes that were “purified” in silico are shaded magenta and blue. E and F were adapted from Cai et al. (2018a).
Using cryo-ET, we have shown that chromatin does not pack into any periodic arrays or monolithic masses inside picoplankton (Gan et al., 2013), budding yeast (Chen et al., 2016) (Figure 3B), and fission yeast cells (Cai et al., 2018b) (Figure 3C). Strangely, chromatin is more compact when released intact from cells, without any divalent-cation chelators added, than when confined within nuclei (Cai et al., 2018c). Our studies, considered together with a recent cryo-ET study of cryosectioned insect cells (Eltsov et al., 2018) (Figure 3D), firmly support the notion that chromatin’s irregular structure in situ is conserved. How, then, do nucleosome chains fold up in situ? In the interphase HeLa cell nuclear periphery, we found that asymmetric nucleosomes follow paths consistent with irregular zigzags (Cai et al., 2018a) (Figure 3, E and F). Much more work is needed to explore the extent of chromatin folding in both transcriptionally active and silenced chromatin.
Cryo-ET studies of transient or rare cellular complexes are more feasible if they can be enriched by either pharmacological or genetic perturbation. For example, an arrest-and-release strategy was used to enrich for fission-yeast cells that have actively constricting actomyosin rings (Swulius et al., 2018). The division septa were then localized by cryo-CLEM (Figure 4A). In these dividing cells, dense bundles of actin filaments were located peripheral to—but not in contact with—the division septum (Figure 4, B and C). We have used both drugs and cell-cycle mutants to arrest picoplankton and budding yeast, respectively, in metaphase. In picoplankton, we found that the unusual spindle has fewer microtubules than chromosomes (Gan et al., 2011) while in budding yeast, we found that the outer-kinetochore Dam1C/DASH complex rarely (if ever) contacts the curved tips of the kinetochore microtubules (Figure 4, D–F) (Ng et al., 2019). The vast repertoire of well-characterized yeast mutants will bring even more genetic precision to future studies.
FIGURE 4:
Cryo-ET analysis of model organisms. (A) Correlation of a fluorescence light cryomicroscopy image of a S. pombe cell cryosection and the cryo-EM projection image of the same position. The fluorescence signal comes from Rlc1-GFP, which marks myosin at the septum’s leading edge. (B) Cryotomographic slice of S. pombe, taken near the contractile actomyosin ring densities (outlined with red dots). (C) Two views (left and right) of a 3-D annotation of an actomyosin ring (orange) and septum (blue). The arrowheads denote the termini of some actomyosin ring densities. A–C were adapted from Swulius et al. (2018). (D) Cryotomographic slice of a metaphase S. cerevisiae cell. Cell membrane (yellow); mitochondria (salmon); nuclear membrane (blue); spindle microtubules (magenta arrowheads). (E) 3-D annotation of seven serial cryotomograms. Outer-nuclear membrane (light blue); inner nuclear membrane (dark blue); spindle microtubules (magenta); Dam1C/DASH complex (green). (F) Enlarged orthogonal views of the 3-D annotation of the metaphase S. cerevisiae spindle from E. D–F were adapted from Ng et al. (2019). (G) Cryotomographic slice of a C. reinhardtii Golgi body (top half) and its corresponding 3-D annotation (bottom half). Portions of the trans-Golgi network (TGN) and endoplasmic reticulum (ER) were captured in this cryotomogram. Adapted from Bykov et al. (2017). (H) Cryotomographic slice of the nuclear periphery of C. reinhardtii. Red arrowheads indicate nuclear pore complexes. (I) A subtomogram average of the C. reinhardtii nuclear pore complex. The Y-complexes (orange and light blue) make up the bulk of the cytoplasmic and nuclear rings of the nuclear pore. H and I were adapted from Mosalaganti et al. (2018).
The model alga Chlamydomonas reinhardtii is convenient for structural studies of its flagella, which can be isolated intact and then analyzed at high resolution (Nicastro et al., 2006; Bui et al., 2008). Recently, cryo-ET studies of cryo-FIB-milled C. reinhardtii have uncovered new details about its cytoplasmic and nucleoplasmic organization. These include the presence of periodic intracisternal protein arrays in the Golgi body (Engel et al., 2015), the nuclear-pore-proximal distribution of proteasomes in the nucleus (Albert et al., 2017), and the distribution of rubisco in the chloroplast (Freeman Rosenzweig et al., 2017). C. reinhardtii has an extensive vesicular-transport cytology (Figure 4G), making it amenable to studies of COP I cargo-sorting vesicles (Bykov et al., 2017). These cells also have easy-to-find nuclear pore complexes, which facilitated the high-resolution STA analysis (Mosalaganti et al., 2018) (Figure 4H).
Reproducible cryomicrotomy and cryo-FIB milling have empowered cell biologists to write rich stories in unconventional contexts. For example, cryo-ET of tissues has shown that desmosomal proteins are arranged in a 2-D lattice in human skin (Al-Amoudi et al., 2007, 2011). This group further devised a clever cryomicrotomy workflow to obtain orthogonal views along the two principal axes of rod cells from mouse retina (Gunkel et al., 2015), showing unambiguously that the G protein–coupled receptor rhodopsin forms ordered arrays. Thin proteins called “tricalbins” have very recently been observed bridging the plasma membrane and endoplasmic reticulum in budding yeast (Collado et al., 2019; Hoffmann et al., 2019). Two-dimensional classification of these tricalbin densities has even revealed their putative globular domains (Hoffmann et al., 2019).
Cryo-ET studies are also revealing the details of cell death and pathogenesis. For example, overexpression of the Bax, a Bcl-2 protein, causes extensive damage to the mitochondria of HeLa cells (Figure 5, A–D). These rare membrane-rupture events were captured with the help of cryo-CLEM (Ader et al., 2019). Cryo-CLEM, combined with integrative modeling (Lasker et al., 2009), were used to propose a model in which a mutant form of LRRK2 kinase decorates microtubules as a right-handed double-helix (Watanabe et al., 2019). Host–pathogen interactions were explored in a study of how a herpes virus maturation intermediate transits the nuclear envelope (Hagen et al., 2015) and how bacteria escape from phagosomes and then divide within their amoeba host (Böck et al., 2017) (Figure 5E). Amyloids were observed within neurons in studies that cleverly used GFP for cryo-CLEM and as a cryo-ET density tag (Bauerlein et al., 2017; Guo et al., 2018). Because yeast can also have amyloid inclusion bodies (Saibil et al., 2012) (Figure 5, F–H), future comparative studies can potentially identify conserved in situ protein-aggregation conditions. Cryo-ET has also been used to study cellular stress, such as in the recent discovery that both starved and mitotically arrested HeLa cells have a high incidence of crystalline lipid droplets (Mahamid et al., 2019).
FIGURE 5:
Structural cell biology of cellular pathology. (A) Fluorescence light cryomicrograph of a cryosection of HeLa cells. The green fluorescence signals correspond to GFP-Bax. (B) Correlation of the fluorescence cryomicroscopy signals from A with a low-magnification cryo-EM projection image of the same cell. (C) Cryotomographic slice corresponding to the white square in B. GFP-Bax cluster densities are annotated in red. (D) Details of a rupturing mitochondrion from another cell. The red and yellow arrowheads, respectively, indicate the inner and outer membranes. A–D were adapted from Ader et al. (2019). (E) Cryotomographic slice showing the bacterium Amoebophilus asiaticus in the cytoplasm of an infected amoeba cell. An array of type 6 secretion structures (T6S array) is indicated within the cytoplasm of the bacterium. Adapted from Böck et al. (2017). (F, G) Projection image and cryotomographic slice, respectively, of cryosections of S. cerevisiae cells that have large prion structures. F and G show parts of a “dot” and a “ring” prion structure, respectively. (H) Model of arrangement of Sup35 fiber arrays (purple) and cross-bridges (pink). F–H were adapted from Saibil et al. (2012).
FASTER, FEWER, FLOPPIER, MESSIER
Many interesting complexes are extremely challenging to study in situ. Such complexes are too rare, or too transient, or have both conformational and constitutional heterogeneity, or are not monolithic, or a combination of the above. Nevertheless, these “biochemist’s nightmare” properties often underlie their function and mechanisms. Examples of such challenging complexes—if they can be called complexes at all—include signalosomes, focal adhesions, neuron synapses, membrane-membrane contact sites, chromatin, and kinetochores. To make such complexes easier to study, cryo-ET needs to be much faster. New tomography hardware and imaging workflows will make tilt-series collection ten times faster (Chreifi et al., 2019; Eisenstein et al., 2019), potentially without sacrificing resolution. Other steps in the workflow will need to speed up: cryolamellae preparation, cryo-CLEM localization, and reconstruction. Recently, researchers have indeed reported advanced automation in cryo-FIB (Buckley et al., 2019; Zachs et al., 2019) and the integration of cryo-FIB milling and light cryomicroscopy in a single instrument (Gorelick et al., 2019). Image processing and analysis will also become faster as researchers exploit the massive parallelism of consumer graphics cards and multicore processors (Kimanius et al., 2016; Castaño-Díez, 2017; Himes and Zhang, 2018; Zivanov et al., 2018).
How would one use high-throughput cryo-ET to answer the most challenging questions? Consider the kinetochore—a large multifunctional complex central to chromosome segregation. Kinetochores connect chromosomes to the spindle apparatus, detect and correct erroneous attachments, delay mitosis until all chromosomes are correctly attached to the spindle apparatus, and couple spindle-microtubule depolymerization to poleward-directed chromosome motion. Kinetochores have dimensions in the tens to hundreds of nanometers and contain multiple elongated subassemblies (Jenni et al., 2017). Recently, many high-resolution cryo-EM structures of reconstituted kinetochore subcomplexes were published (Pentakota et al., 2017; Chittori et al., 2018; Jenni and Harrison, 2018; Leber et al., 2018; Yan et al., 2018; Zhang et al., 2018; Hinshaw et al., 2019; Hinshaw and Harrison, 2019; Yan et al., 2019). Yet, very little progress on the kinetochore’s in situ structure(s) has been made. Using a combination of serial cryo-ET of cryosections, with kinetochore microtubules as macromolecular signposts, we recently characterized the Dam1C/DASH complex—just one piece of the yeast outer kinetochore (Ng et al., 2019). The in situ structure of the rest of the kinetochore remains unknown.
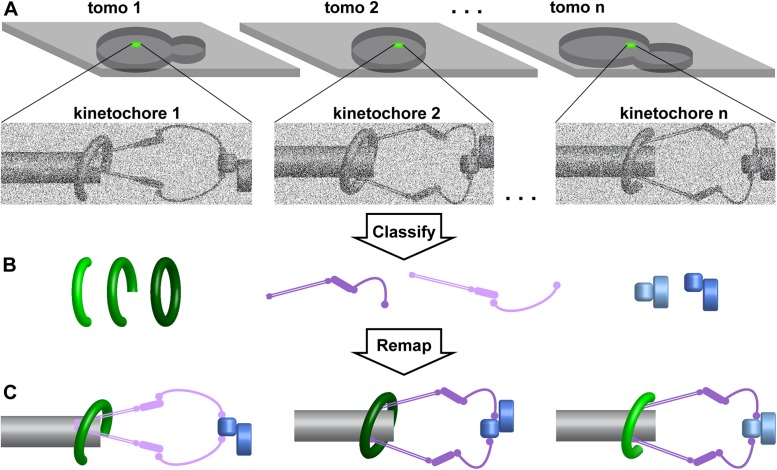
We now speculate on how high-throughput cryo-ET, combined with a “divide, conquer, and unite” subtomogram-analysis strategy, will reveal kinetochore structures in situ. In this exercise, we wishfully assume that the just-discussed microscopy bottlenecks will be solved by automation and improved workflows. Our starting point is a collection of thousands of cryotomograms of budding yeast cryolamellae, each containing a few kinetochores that were automatically localized by cryo-CLEM. The first step is to extract subtomograms that overlap with the light cryomicroscopy signals (Figure 6A). Identification of the kinetochore subassemblies will be guided by our knowledge of their relative positions (Joglekar et al., 2009; Aravamudhan et al., 2014). Small subtomograms that contain these subassemblies will next be classified into their various conformational states (Figure 6B). We imagine that some kinetochore subassemblies have only a few translational and rotational degrees of freedom relative to kinetochore’s center of mass, analogous to side-chain rotamers in protein structures. In contrast, there are also some flexible, elongated subassemblies such as the coiled–coil-rich Ndc80 complex, which gives the kinetochore its gossamer nature. Flexible subassemblies like Ndc80 are expected to have many conformations and would therefore produce the lowest-resolution models. Next, each kinetochore will be reassembled in silico by remapping the subtomogram averages of each subcomponent (Figure 6C). Kinetochores pieced together this way will have many components remapped with subnanometer accuracy. Remapped kinetochores from spindles with and without tension will reveal a wealth of information. We will learn which complexes undergo the largest conformational changes, what conformational changes are sensitive to spindle pulling forces, how checkpoint complexes dock in mitotically arrested cells, if detached kinetochores have the same conformation as tensionless kinetochores, and if anaphase kinetochores attach to the spindle the same way as metaphase ones. These in situ kinetochore structures will stimulate even more (presently unimaginable) questions.
FIGURE 6:
Structural cell biology of floppy complexes. This schematic shows how a divide, conquer, and unite strategy can reveal kinetochore structures in situ. (A) Cartoons of cryotomograms (thin gray slabs; tomo1, tomo2, etc.) of thinned yeast (rounded gray bodies). In this example, the kinetochores are first localized by fluorescence cryomicroscopy (green signal). Cryo-CLEM greatly facilitates the identification of subtomograms that contain kinetochores. (B) Alignment and classification of smaller subtomograms that contain kinetochore subassemblies. Owing to the flexibility and conformational heterogeneity of the kinetochores, each subassembly must be windowed and then tracked throughout the alignment and classification process. Subassemblies that have similar conformations are aligned and averaged, producing a higher signal-to-noise ratio “class average.” In this schematic, the outer kinetochore, inner kinetochore, and centromere-associated complexes are colored green, violet, and blue, respectively. (C) These class averages can then be rotated and translated to the orientations and coordinates of each copy in their in situ context at the tips of kinetochore microtubules (gray). This remapping approach can deal with “floppy” complexes as long as some of the subassemblies are monolithic.
Supplementary Material
Acknowledgments
We thank Eunyoung Chae, Min Wu, Tony Kanchanawong, Shu Fen Tan, and Alasdair McDowall for discussions and the anonymous reviewers for their constructive feedback. The images and movies were created with Bsoft, IMOD, UCSF Chimera, avconv, and Adobe Creative Cloud ( Mastronarde, 1997; Pettersen et al., 2004; Heymann and Belnap, 2007; Libav_team, 2018). All panels in Figures 3 – 5 were adapted with permission from the authors and publishers. This work was supported by Singapore Ministry of Education T1 R-154-000-A49-114 and MOE2018-T2-2-146.
Abbreviations used:
- cryo-CLEM
correlative light and electron cryomicroscopy
- cryo-EM
electron cryomicroscopy/cryo-electron microscopy
- cryo-ET
electron cryotomography/cryo-electron tomography
- cryo-FIB
cryo focused-ion beam
- SPA
single-particle analysis
- STA
subtomogram averaging.
Footnotes
REFERENCES
- Ader NR, Hoffmann PC, Ganeva I, Borgeaud AC, Wang C, Youle RJ, Kukulski W. (2019). Molecular and topological reorganizations in mitochondrial architecture interplay during Bax-mediated steps of apoptosis. Elife , e40712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adrian M, Dubochet J, Lepault J, McDowall AW. (1984). Cryo-electron microscopy of viruses. Nature , 32–36. [DOI] [PubMed] [Google Scholar]
- Al-Amoudi A, Castano-Diez D, Devos DP, Russell RB, Johnson GT, Frangakis AS. (2011). The three-dimensional molecular structure of the desmosomal plaque. Proc Natl Acad Sci USA , 6480–6485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al-Amoudi A, Diez DC, Betts MJ, Frangakis AS. (2007). The molecular architecture of cadherins in native epidermal desmosomes. Nature , 832–837. [DOI] [PubMed] [Google Scholar]
- Albert S, Schaffer M, Beck F, Mosalaganti S, Asano S, Thomas HF, Plitzko JM, Beck M, Baumeister W, Engel BD. (2017). Proteasomes tether to two distinct sites at the nuclear pore complex. Proc Natl Acad Sci USA , 13726–13731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aravamudhan P, Felzer-Kim I, Gurunathan K, Joglekar AP. (2014). Assembling the protein architecture of the budding yeast kinetochore-microtubule attachment using FRET. Curr Biol , 1437–1446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asano S, Engel BD, Baumeister W. (2016). In situ cryo-electron tomography: a post-reductionist approach to structural biology. J Mol Biol , 332–343. [DOI] [PubMed] [Google Scholar]
- Bauerlein FJB, Saha I, Mishra A, Kalemanov M, Martinez-Sanchez A, Klein R, Dudanova I, Hipp MS, Hartl FU, Baumeister W, Fernandez-Busnadiego R. (2017). In situ architecture and cellular interactions of polyQ inclusions. Cell , 179–187.e110. [DOI] [PubMed] [Google Scholar]
- Bharat TAM, Hoffmann PC, Kukulski W. (2018). Correlative microscopy of vitreous sections provides insights into BAR-domain organization in situ. Structure , 879–886.e873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bharat TA, Russo CJ, Lowe J, Passmore LA, Scheres SH. (2015). Advances in single-particle electron cryomicroscopy structure determination applied to sub-tomogram averaging. Structure , 1743–1753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Böck D, Medeiros JM, Tsao HF, Penz T, Weiss GL, Aistleitner K, Horn M, Pilhofer M. (2017). In situ architecture, function, and evolution of a contractile injection system. Science , 713–717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brilot AF, Chen JZ, Cheng A, Pan J, Harrison SC, Potter CS, Carragher B, Henderson R, Grigorieff N. (2012). Beam-induced motion of vitrified specimen on holey carbon film. J Struct Biol , 630–637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buckley G, Gervinskas G, Venugopal H, Taveneau C, Whisstock JC, de Marco A. (2019). Automated cryo-lamella preparation for high-throughput in-situ structural biology. bioRxiv, 797506. [DOI] [PubMed] [Google Scholar]
- Bui KH, Sakakibara H, Movassagh T, Oiwa K, Ishikawa T. (2008). Molecular architecture of inner dynein arms in situ in Chlamydomonas reinhardtii flagella. J Cell Biol , 923–932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bykov YS, Schaffer M, Dodonova SO, Albert S, Plitzko JM, Baumeister W, Engel BD, Briggs JA. (2017). The structure of the COPI coat determined within the cell. Elife , e32493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai S, Böck D, Pilhofer M, Gan L. (2018a). The in situ structures of mono-, di-, and trinucleosomes in human heterochromatin. Mol Biol Cell , 2450–2457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai S, Chen C, Tan ZY, Huang Y, Shi J, Gan L. (2018b). Cryo-ET reveals the macromolecular reorganization of S. pombe mitotic chromosomes in vivo. Proc Natl Acad Sci USA , 10977–10982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai S, Song Y, Chen C, Shi J, Gan L. (2018c). Natural chromatin is heterogeneous and self-associates in vitro. Mol Biol Cell , 1652–1663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castaño-Díez D. (2017). The Dynamo package for tomography and subtomogram averaging: components for MATLAB, GPU computing and EC2 Amazon Web Services. Acta Crystallogr D Struct Biol , 478–487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- CCP-EM (2019). CCP-EM discussion list, www.jiscmail.ac.uk/CCPEM.
- Chang YW, Chen S, Tocheva EI, Treuner-Lange A, Lobach S, Sogaard-Andersen L, Jensen GJ. (2014). Correlated cryogenic photoactivated localization microscopy and cryo-electron tomography. Nat Methods , 737–739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang J-J, McDowall AW, Lepault J, Freeman R, Walter CA, Dubochet J. (1983). Freezing, sectioning and observation artefacts of frozen hydrated sections for electron microscopy. J Microsc , 109–123. [DOI] [PubMed] [Google Scholar]
- Chang YW, Rettberg LA, Treuner-Lange A, Iwasa J, Sogaard-Andersen L, Jensen GJ. (2016). Architecture of the type IVa pilus machine. Science , aad2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen M, Bell JM, Shi X, Sun SY, Wang Z, Ludtke SJ. (2019). A complete data processing workflow for cryo-ET and subtomogram averaging. Nat Methods , 1161–1168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen M, Dai W, Sun SY, Jonasch D, He CY, Schmid MF, Chiu W, Ludtke SJ. (2017). Convolutional neural networks for automated annotation of cellular cryo-electron tomograms. Nat Methods , 983–985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen C, Lim HH, Shi J, Tamura S, Maeshima K, Surana U, Gan L. (2016). Budding yeast chromatin is dispersed in a crowded nucleoplasm in vivo. Mol Biol Cell , 3357–3368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheng A, Henderson R, Mastronarde D, Ludtke SJ, Schoenmakers RH, Short J, Marabini R, Dallakyan S, Agard D, Winn M. (2015). MRC2014: Extensions to the MRC format header for electron cryo-microscopy and tomography. J Struct Biol , 146–150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chittori S, Hong J, Saunders H, Feng H, Ghirlando R, Kelly AE, Bai Y, Subramaniam S. (2018). Structural mechanisms of centromeric nucleosome recognition by the kinetochore protein CENP-N. Science , 339–343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chreifi G, Chen S, Metskas LA, Kaplan M, Jensen GJ. (2019). Rapid tilt-series acquisition for electron cryotomography. J Struct Biol , 163–169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collado J, Kalemanov M, Campelo F, Bourgoint C, Thomas F, Loewith R, Martínez-Sánchez A, Baumeister W, Stefan CJ, Fernández-Busnadiego R. (2019). Tricalbin-mediated contact sites control ER curvature to maintain plasma membrane integrity. Dev Cell , 476–487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dahl R, Staehelin LA. (1989). High-pressure freezing for the preservation of biological structure: theory and practice. J Electron Microsc Tech , 165–174. [DOI] [PubMed] [Google Scholar]
- Dahlberg PD, Sartor AM, Wang J, Saurabh S, Shapiro L, Moerner WE. (2018). Identification of PAmKate as a red photoactivatable fluorescent protein for cryogenic super-resolution imaging. J Am Chem Soc , 12310–12313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Danev R, Buijsse B, Khoshouei M, Plitzko JM, Baumeister W. (2014). Volta potential phase plate for in-focus phase contrast transmission electron microscopy. Proc Natl Acad Sci USA , 15635–15640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davey CA, Sargent DF, Luger K, Maeder AW, Richmond TJ. (2002). Solvent mediated interactions in the structure of the nucleosome core particle at 1.9 a resolution. J Mol Biol , 1097–1113. [DOI] [PubMed] [Google Scholar]
- Dhatchinamoorthy K, Shivaraju M, Lange JJ, Rubinstein B, Unruh JR, Slaughter BD, Gerton JL. (2017). Structural plasticity of the living kinetochore. J Cell Biol , 3551–3570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diamond (2019). eBIC: Electron Bio-Imaging Centre, www.diamond.ac.uk/Instruments/Biological-Cryo-Imaging/eBIC.html.
- Dobro MJ, Melanson LA, Jensen GJ, McDowall AW. (2010). Plunge freezing for electron cryomicroscopy. Methods Enzymol , 63–82. [DOI] [PubMed] [Google Scholar]
- Dubochet J. (2007). The physics of rapid cooling and its implications for cryoimmobilization of cells. Methods Cell Biol , 7–21. [DOI] [PubMed] [Google Scholar]
- Dubochet J, McDowall AW. (1981). Vitrification of pure water for electron microscopy. J Microsc , RP3–RP4. [Google Scholar]
- Dubochet J, Zuber B, Eltsov M, Bouchet-Marquis C, Al-Amoudi A, Livolant F. (2007). How to “read” a vitreous section. Methods Cell Biol , 385–406. [DOI] [PubMed] [Google Scholar]
- Eisenstein F, Danev R, Pilhofer M. (2019). Improved applicability and robustness of fast cryo-electron tomography data acquisition. J Struct Biol , 107–114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eltsov M, Grewe D, Lemercier N, Frangakis A, Livolant F, Leforestier A. (2018). Nucleosome conformational variability in solution and in interphase nuclei evidenced by cryo-electron microscopy of vitreous sections. Nucleic Acids Res , 9189–9200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eltsov M, MacLellan KM, Maeshima K, Frangakis AS, Dubochet J. (2008). Analysis of cryo-electron microscopy images does not support the existence of 30-nm chromatin fibers in mitotic chromosomes in situ. Proc Natl Acad Sci , 19732–19737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engel BD, Schaffer M, Albert S, Asano S, Plitzko JM, Baumeister W. (2015). In situ structural analysis of Golgi intracisternal protein arrays. Proc Natl Acad Sci USA , 11264–11269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finch JT, Klug A. (1976). Solenoidal model for superstructure in chromatin. Proc Natl Acad Sci USA , 1897–1901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forster F, Han BG, Beck M. (2010). Visual proteomics. Methods Enzymol , 215–243. [DOI] [PubMed] [Google Scholar]
- Frank J. (1975). Averaging of low exposure electron micrographs of non-periodic objects. Ultramicroscopy , 159–162. [DOI] [PubMed] [Google Scholar]
- Frank J. (2006a). Electron Tomography: Methods for Three-Dimensional Visualization of Structures in the Cell, New York: Springer. [Google Scholar]
- Frank J. (2006b). Three-Dimensional Electron Microscopy of Macromolecular Assemblies: Visualization of Biological Molecules in Their Native State, New York: Oxford University Press. [Google Scholar]
- Freeman Rosenzweig ES, Xu B, Kuhn Cuellar L, Martinez-Sanchez A, Schaffer M, Strauss M, Cartwright HN, Ronceray P, Plitzko JM, Forster F, et al (2017). The Eukaryotic CO2-concentrating organelle is liquid-like and exhibits dynamic reorganization. Cell , 148–162.e119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galaz-Montoya JG, Flanagan J, Schmid MF, Ludtke SJ. (2015). Single particle tomography in EMAN2. J Struct Biol , 279–290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gan L, Ladinsky MS, Jensen GJ. (2011). Organization of the smallest eukaryotic spindle. Curr Biol , 1578–1583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gan L, Ladinsky MS, Jensen GJ. (2013). Chromatin in a marine picoeukaryote is a disordered assemblage of nucleosomes. Chromosoma , 377–386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gan L, Ng CT, Chen C, Cai S. (2019). A collection of yeast cellular electron cryotomography data. GigaScience , giz077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorelick S, Buckley G, Gervinskas G, Johnson TK, Handley A, Caggiano MP, Whisstock JC, Pocock R, de Marco A. (2019). PIE-scope, integrated cryo-correlative light and FIB/SEM microscopy. Elife , e45919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grimm R, Singh H, Rachel R, Typke D, Zillig W, Baumeister W. (1998). Electron tomography of ice-embedded prokaryotic cells. Biophys J , 1031–1042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gunkel M, Schoneberg J, Alkhaldi W, Irsen S, Noe F, Kaupp UB, Al-Amoudi A. (2015). Higher-order architecture of rhodopsin in intact photoreceptors and its implication for phototransduction kinetics. Structure , 628–638. [DOI] [PubMed] [Google Scholar]
- Guo Q, Lehmer C, Martinez-Sanchez A, Rudack T, Beck F, Hartmann H, Perez-Berlanga M, Frottin F, Hipp MS, Hartl FU, et al (2018). In situ structure of neuronal C9orf72 Poly-GA aggregates reveals proteasome recruitment. Cell , 696–705.e612. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hagen C, Dent KC, Zeev-Ben-Mordehai T, Grange M, Bosse JB, Whittle C, Klupp BG, Siebert CA, Vasishtan D, Bauerlein FJ, et al (2015). Structural basis of vesicle formation at the inner nuclear membrane. Cell , 1692–1701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hampton CM, Strauss JD, Ke Z, Dillard RS, Hammonds JE, Alonas E, Desai TM, Marin M, Storms RE, Leon F, et al (2017). Correlated fluorescence microscopy and cryo-electron tomography of virus-infected or transfected mammalian cells. Nat Protoc , 150–167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harapin J, Bormel M, Sapra KT, Brunner D, Kaech A, Medalia O. (2015). Structural analysis of multicellular organisms with cryo-electron tomography. Nat Methods , 634–636. [DOI] [PubMed] [Google Scholar]
- Haruki H, Nishikawa J, Laemmli UK. (2008). The anchor-away technique: rapid, conditional establishment of yeast mutant phenotypes. Mol Cell , 925–932. [DOI] [PubMed] [Google Scholar]
- Hayles MF, Stokes DJ, Phifer D, Findlay KC. (2007). A technique for improved focused ion beam milling of cryo-prepared life science specimens. J Microsc , 263–269. [DOI] [PubMed] [Google Scholar]
- Henderson GP, Gan L, Jensen GJ. (2007). 3-D ultrastructure of O. tauri: electron cryotomography of an entire eukaryotic cell. PLoS One , e749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson R. (1995). The potential and limitations of neutrons, electrons and X-rays for atomic resolution microscopy of unstained biological molecules. Q Rev Biophys , 171–193. [DOI] [PubMed] [Google Scholar]
- Henderson R, Baldwin JM, Downing KH, Lepault J, Zemlin F. (1986). Structure of purple membrane from halobacterium halobium: recording, measurement and evaluation of electron micrographs at 3.5 Å resolution. Ultramicroscopy , 147–178. [Google Scholar]
- Heumann JM. (2016). PEET: University of Colorado Boulder, http://bio3d.colorado.edu/PEET.
- Heymann JB, Belnap DM. (2007). Bsoft: image processing and molecular modeling for electron microscopy. J Struct Biol , 3–18. [DOI] [PubMed] [Google Scholar]
- Heymann JB, Cardone G, Winkler DC, Steven AC. (2008). Computational resources for cryo-electron tomography in Bsoft. J Struct Biol , 232–242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Himes BA, Zhang P. (2018). emClarity: software for high-resolution cryo-electron tomography and subtomogram averaging. Nat Methods , 955–961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinshaw SM, Dates AN, Harrison SC. (2019). The structure of the yeast Ctf3 complex. Elife , e48215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinshaw SM, Harrison SC. (2019). The structure of the Ctf19c/CCAN from budding yeast. Elife , e44239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffmann PC, Bharat TAM, Wozny MR, Boulanger J, Miller EA, Kukulski W. (2019). Tricalbins contribute to cellular lipid flux and form curved ER-PM contacts that are bridged by rod-shaped structures. Dev Cell , 488–502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsieh CE, Marko M, Frank J, Mannella CA. (2002). Electron tomographic analysis of frozen-hydrated tissue sections. J Struct Biol , 63–73. [DOI] [PubMed] [Google Scholar]
- Hutchings J, Zanetti G. (2018). Fine details in complex environments: the power of cryo-electron tomography. Biochem Soc Trans , 807–816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iudin A, Korir PK, Salavert-Torres J, Kleywegt GJ, Patwardhan A. (2016). EMPIAR: a public archive for raw electron microscopy image data. Nat Methods , 387–388. [DOI] [PubMed] [Google Scholar]
- Jasnin M, Beck F, Ecke M, Fukuda Y, Martinez-Sanchez A, Baumeister W, Gerisch G. (2019). The architecture of traveling actin waves revealed by cryo-electron tomography. Structure , 1211–1223 e1215. [DOI] [PubMed] [Google Scholar]
- Jenni S, Dimitrova YN, Valverde R, Hinshaw SM, Harrison SC. (2017). Molecular structures of yeast kinetochore subcomplexes and their roles in chromosome segregation. Cold Spring Harb Symp Quant Biol , 83–89. [DOI] [PubMed] [Google Scholar]
- Jenni S, Harrison SC. (2018). Structure of the DASH/Dam1 complex shows its role at the yeast kinetochore-microtubule interface. Science , 552–558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang Z, Jin X, Li Y, Liu S, Liu X-M, Wang Y-Y, Zhao P, Cai X, Liu Y, Tang Y, et al (2019). Direct synthesis of EM-visible gold nanoparticles on genetically encoded tags for single-molecule visualization in cells. bioRxiv, 520999. [Google Scholar]
- Joglekar AP, Bloom K, Salmon ED. (2009). In vivo protein architecture of the eukaryotic kinetochore with nanometer scale accuracy. Curr Biol , 694–699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimanius D, Forsberg BO, Scheres SH, Lindahl E. (2016). Accelerated cryo-EM structure determination with parallelisation using GPUs in RELION-2. Elife , e18722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komeili A, Li Z, Newman DK, Jensen GJ. (2006). Magnetosomes are cell membrane invaginations organized by the actin-like protein MamK. Science , 242–245. [DOI] [PubMed] [Google Scholar]
- Koning RI, Zovko S, Barcena M, Oostergetel GT, Koerten HK, Galjart N, Koster AJ, Mieke Mommaas A. (2008). Cryo electron tomography of vitrified fibroblasts: microtubule plus ends in situ. J Struct Biol , 459–468. [DOI] [PubMed] [Google Scholar]
- Kornberg RD. (1974). Chromatin structure: a repeating unit of histones and DNA. Science , 868–871. [DOI] [PubMed] [Google Scholar]
- Koster AJ, Grimm R, Typke D, Hegerl R, Stoschek A, Walz J, Baumeister W. (1997). Perspectives of molecular and cellular electron tomography. J Struct Biol , 276–308. [DOI] [PubMed] [Google Scholar]
- Ladinsky MS, Pierson JM, McIntosh JR. (2006). Vitreous cryo-sectioning of cells facilitated by a micromanipulator. J Microsc , 129–134. [DOI] [PubMed] [Google Scholar]
- Lasker K, Topf M, Sali A, Wolfson HJ. (2009). Inferential optimization for simultaneous fitting of multiple components into a CryoEM map of their assembly. J Mol Biol , 180–194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leber V, Nans A, Singleton MR. (2018). Structural basis for assembly of the CBF3 kinetochore complex. EMBO J , 269–281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Libav_team (2018). avconv, https://libav.org.
- Luger K, Mader AW, Richmond RK, Sargent DF, Richmond TJ. (1997). Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature , 251–260. [DOI] [PubMed] [Google Scholar]
- Maeshima K, Ide S, Babokhov M. (2019). Dynamic chromatin organization without the 30-nm fiber. Curr Opin Cell Biol , 95–104. [DOI] [PubMed] [Google Scholar]
- Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Cuellar LK, Forster F, Hyman AA, Plitzko JM, Baumeister W. (2016). Visualizing the molecular sociology at the HeLa cell nuclear periphery. Science , 969–972. [DOI] [PubMed] [Google Scholar]
- Mahamid J, Schampers R, Persoon H, Hyman AA, Baumeister W, Plitzko JM. (2015). A focused ion beam milling and lift-out approach for site-specific preparation of frozen-hydrated lamellas from multicellular organisms. J Struct Biol , 262–269. [DOI] [PubMed] [Google Scholar]
- Mahamid J, Tegunov D, Maiser A, Arnold J, Leonhardt H, Plitzko JM, Baumeister W. (2019). Liquid-crystalline phase transitions in lipid droplets are related to cellular states and specific organelle association. Proc Natl Acad Sci USA , 16866–16871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marko M, Hsieh C, Moberlychan W, Mannella CA, Frank J. (2006). Focused ion beam milling of vitreous water: prospects for an alternative to cryo-ultramicrotomy of frozen-hydrated biological samples. J Microsc , 42–47. [DOI] [PubMed] [Google Scholar]
- Mastronarde DN. (1997). Dual-axis tomography: an approach with alignment methods that preserve resolution. J Struct Biol , 343–352. [DOI] [PubMed] [Google Scholar]
- Mastronarde DN. (2005). Automated electron microscope tomography using robust prediction of specimen movements. J Struct Biol , 36–51. [DOI] [PubMed] [Google Scholar]
- Mastronarde DN. (2019). IMOD Discussion Group, https://bio3d.colorado.edu/imod/joinlist.html.
- McDowall A. (1984). Ultracryotomy: An Investigation of the Cryotechnical Problems Involved in the Preparation of Frozen-Hydrated Cells and Tissues for High Resolution Electron Microscopy. PhD Thesis. Paris: Universite Pierre et Marie Curie (Sorbonne University).
- McDowall AW, Chang JJ, Freeman R, Lepault J, Walter CA, Dubochet J. (1983). Electron microscopy of frozen hydrated sections of vitreous ice and vitrified biological samples. J Microsc , 1–9. [DOI] [PubMed] [Google Scholar]
- McDowall AW, Smith JM, Dubochet J. (1986). Cryo-electron microscopy of vitrified chromosomes in situ. EMBO J , 1395–1402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Medalia O, Weber I, Frangakis AS, Nicastro D, Gerisch G, Baumeister W. (2002). Macromolecular architecture in eukaryotic cells visualized by cryoelectron tomography. Science , 1209–1213. [DOI] [PubMed] [Google Scholar]
- Medeiros JM, Bock D, Weiss GL, Kooger R, Wepf RA, Pilhofer M. (2018). Robust workflow and instrumentation for cryo-focused ion beam milling of samples for electron cryotomography. Ultramicroscopy , 1–11. [DOI] [PubMed] [Google Scholar]
- Mosalaganti S, Kosinski J, Albert S, Schaffer M, Strenkert D, Salome PA, Merchant SS, Plitzko JM, Baumeister W, Engel BD, Beck M. (2018). In situ architecture of the algal nuclear pore complex. Nat Commun , 2361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ng CT, Deng L, Chen C, Lim HH, Shi J, Surana U, Gan L. (2019). Electron cryotomography analysis of Dam1C/DASH at the kinetochore-spindle interface in situ. J Cell Biol , 455–473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicastro D, McIntosh JR, Baumeister W. (2005). 3D structure of eukaryotic flagella in a quiescent state revealed by cryo-electron tomography. Proc Natl Acad Sci USA , 15889–15894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicastro D, Schwartz C, Pierson J, Gaudette R, Porter ME, McIntosh JR. (2006). The molecular architecture of axonemes revealed by cryoelectron tomography. Science , 944–948. [DOI] [PubMed] [Google Scholar]
- Nieduszynski CA, Hiraga S, Ak P, Benham CJ, Donaldson AD. (2007). OriDB: a DNA replication origin database. Nucleic Acids Res , D40–D46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishimura K, Fukagawa T, Takisawa H, Kakimoto T, Kanemaki M. (2009). An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat Methods , 917–922. [DOI] [PubMed] [Google Scholar]
- Nishino Y, Eltsov M, Joti Y, Ito K, Takata H, Takahashi Y, Hihara S, Frangakis AS, Imamoto N, Ishikawa T, Maeshima K. (2012). Human mitotic chromosomes consist predominantly of irregularly folded nucleosome fibres without a 30-nm chromatin structure. EMBO J , 1644–1653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noble AJ, Stagg SM. (2015). Automated batch fiducial-less tilt-series alignment in Appion using Protomo. J Struct Biol , 270–278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oda T, Kikkawa M. (2013). Novel structural labeling method using cryo-electron tomography and biotin-streptavidin system. J Struct Biol , 305–311. [DOI] [PubMed] [Google Scholar]
- Oikonomou CM, Jensen GJ. (2017). Cellular electron cryotomography: toward structural biology in situ. Annu Rev Biochem , 873–896. [DOI] [PubMed] [Google Scholar]
- Ortega DR, Oikonomou CM, Ding HJ, Rees-Lee P, Alexandria, Jensen GJ. (2019). ETDB-Caltech: A blockchain-based distributed public database for electron tomography. PLoS One , e0215531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Toole E. (2018). ETomo Tutorial for IMOD Version 4.9: University of Colorado, Boulder, https://bio3d.colorado.edu/imod/doc/etomoTutorial.html. [Google Scholar]
- Palmer CM, Lowe J. (2014). A cylindrical specimen holder for electron cryo-tomography. Ultramicroscopy , 20–29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Penczek PA. (2010). Image restoration in cryo-electron microscopy. Methods Enzymol , 35–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pentakota S, Zhou K, Smith C, Maffini S, Petrovic A, Morgan GP, Weir JR, Vetter IR, Musacchio A, Luger K. (2017). Decoding the centromeric nucleosome through CENP-N. Elife , e33442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perkins G. (2019). 3DEM mailing list, http://3dem.ucsd.edu/mailinglist.shtm.
- Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE. (2004). UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem , 1605–1612. [DOI] [PubMed] [Google Scholar]
- Ravelli RBG, Nijpels FJT, Henderikx RJM, Weissenberger G, Thewessem S, Gijsbers A, Beulen BWAMM, López-Iglesias C, Peters PJ. (2019). Automated cryo-EM sample preparation by pin-printing and jet vitrification. bioRxiv, 651208. [Google Scholar]
- Rigort A, Bauerlein FJ, Leis A, Gruska M, Hoffmann C, Laugks T, Bohm U, Eibauer M, Gnaegi H, Baumeister W, Plitzko JM. (2010). Micromachining tools and correlative approaches for cellular cryo-electron tomography. J Struct Biol , 169–179. [DOI] [PubMed] [Google Scholar]
- Rodenburg J. (2019). Tutorials in Transmission Electron Microscopy, www.rodenburg.org/index.html.
- Rout MP, Blobel G. (1993). Isolation of the yeast nuclear pore complex. J Cell Biol , 771–783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruskin RS, Yu Z, Grigorieff N. (2013). Quantitative characterization of electron detectors for transmission electron microscopy. J Struct Biol , 385–393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saibil HR, Seybert A, Habermann A, Winkler J, Eltsov M, Perkovic M, Castano-Diez D, Scheffer MP, Haselmann U, Chlanda P, et al (2012). Heritable yeast prions have a highly organized three-dimensional architecture with interfiber structures. Proc Natl Acad Sci USA , 14906–14911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaffer M, Pfeffer S, Mahamid J, Kleindiek S, Laugks T, Albert S, Engel BD, Rummel A, Smith AJ, Baumeister W, Plitzko JM. (2019). A cryo-FIB lift-out technique enables molecular-resolution cryo-ET within native Caenorhabditis elegans tissue. Nat Methods , 757–762. [DOI] [PubMed] [Google Scholar]
- Scheffel A, Gruska M, Faivre D, Linaroudis A, Plitzko JM, Schuler D. (2006). An acidic protein aligns magnetosomes along a filamentous structure in magnetotactic bacteria. Nature , 110–114. [DOI] [PubMed] [Google Scholar]
- Schlimpert S, Klein EA, Briegel A, Hughes V, Kahnt J, Bolte K, Maier UG, Brun YV, Jensen GJ, Gitai Z, Thanbichler M. (2012). General protein diffusion barriers create compartments within bacterial cells. Cell , 1270–1282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schur FK. (2019). Toward high-resolution in situ structural biology with cryo-electron tomography and subtomogram averaging. Curr Opin Struct Biol , 1–9. [DOI] [PubMed] [Google Scholar]
- Schur FK, Obr M, Hagen WJ, Wan W, Jakobi AJ, Kirkpatrick JM, Sachse C, Krausslich HG, Briggs JA. (2016). An atomic model of HIV-1 capsid-SP1 reveals structures regulating assembly and maturation. Science , 506–508. [DOI] [PubMed] [Google Scholar]
- Schwartz O, Axelrod JJ, Campbell SL, Turnbaugh C, Glaeser RM, Muller H. (2019). Laser phase plate for transmission electron microscopy. Nat Methods , 1016–1020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen PS, Iwasa JH. (2018). CryoEM 101, https://cryoem101.org.
- Studer D, Klein A, Iacovache I, Gnaegi H, Zuber B. (2014). A new tool based on two micromanipulators facilitates the handling of ultrathin cryosection ribbons. J Struct Biol , 125–128. [DOI] [PubMed] [Google Scholar]
- Suloway C, Shi J, Cheng A, Pulokas J, Carragher B, Potter CS, Zheng SQ, Agard DA, Jensen GJ. (2009). Fully automated, sequential tilt-series acquisition with Leginon. J Struct Biol , 11–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swulius MT, Nguyen LT, Ladinsky MS, Ortega DR, Aich S, Mishra M, Jensen GJ. (2018). Structure of the fission yeast actomyosin ring during constriction. Proc Natl Acad Sci USA , E1455–E1464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor KA, Glaeser RM. (1976). Electron microscopy of frozen hydrated biological specimens. J Ultrastruct Res , 448–456. [DOI] [PubMed] [Google Scholar]
- Tivol WF, Briegel A, Jensen GJ. (2008). An improved cryogen for plunge freezing. Microsc Microanal , 375–379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tuijtel MW, Koster AJ, Jakobs S, Faas FGA, Sharp TH. (2019). Correlative cryo super-resolution light and electron microscopy on mammalian cells using fluorescent proteins. Sci Rep , 1369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villa E, Schaffer M, Plitzko JM, Baumeister W. (2013). Opening windows into the cell: focused-ion-beam milling for cryo-electron tomography. Curr Opin Struct Biol , 771–777. [DOI] [PubMed] [Google Scholar]
- Vos MR, Jensen GJ. (2018). Getting Started in Cryo-EM online course, https://em-learning.com.
- Watanabe R, Buschauer R, Böhning J, Audagnotto M, Lasker K, Lu TW, Boassa D, Taylor S, Villa E. (2019). The in situ structure of Parkinson’s disease-linked LRRK2. bioRxiv, 837203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Widom J, Klug A. (1985). Structure of the 300A chromatin filament: X-ray diffraction from oriented samples. Cell , 207–213. [DOI] [PubMed] [Google Scholar]
- Wolff G, Limpens R, Zheng S, Snijder EJ, Agard DA, Koster AJ, Barcena M. (2019). Mind the gap: Micro-expansion joints drastically decrease the bending of FIB-milled cryo-lamellae. J Struct Biol, www.ncbi.nlm.nih.gov/pubmed/31536774. [DOI] [PubMed] [Google Scholar]
- Yan K, Yang J, Zhang Z, McLaughlin SH, Chang L, Fasci D, Ehrenhofer-Murray AE, Heck AJR, Barford D. (2019). Structure of the inner kinetochore CCAN complex assembled onto a centromeric nucleosome. Nature , 278–282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan K, Zhang Z, Yang J, McLaughlin SH, Barford D. (2018). Architecture of the CBF3-centromere complex of the budding yeast kinetochore. Nat Struct Mol Biol , 1103–1110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Q, Rout MP, Akey CW. (1998). Three-dimensional architecture of the isolated yeast nuclear pore complex: functional and evolutionary implications. Mol Cell , 223–234. [DOI] [PubMed] [Google Scholar]
- Zachs T, Medeiros JM, Schertel A, Weiss GL, Hugener J, Pilhofer M. (2019). Fully automated, sequential focused ion beam milling for cryo-electron tomography. bioRxiv, 797514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng X, Leung MR, Zeev-Ben-Mordehai T, Xu M. (2018). A convolutional autoencoder approach for mining features in cellular electron cryo-tomograms and weakly supervised coarse segmentation. J Struct Biol , 150–160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang J, Zhang D, Sun L, Ji G, Huang X, Niu T, Sun F. (2019). VHUT-cryo-FIB, a method to fabricate frozen-hydrated lamella of tissue specimen for in situ cryo-electron tomography. bioRxiv, 727149. [DOI] [PubMed] [Google Scholar]
- Zhang W, Lukoyanova N, Miah S, Lucas J, Vaughan CK. (2018). Insights into centromere DNA bending revealed by the Cryo-EM structure of the core centromere binding factor 3 with Ndc10. Cell Rep , 744–754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng SQ, Matsuda A, Braunfeld MB, Sedat JW, Agard DA. (2009). Dual-axis target mapping and automated sequential acquisition of dual-axis EM tomographic data. J Struct Biol , 323–331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zivanov J, Nakane T, Forsberg BO, Kimanius D, Hagen WJ, Lindahl E, Scheres SH. (2018). New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife , e42166. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.