Abstract
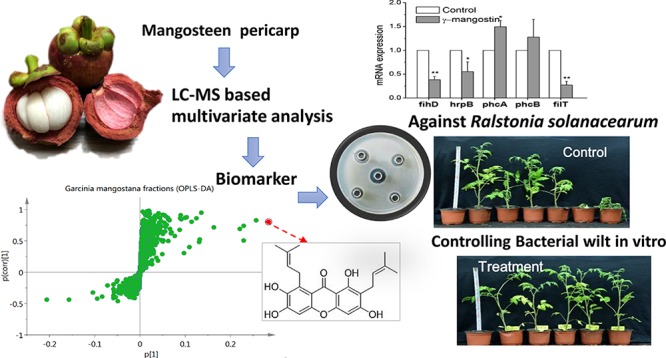
Bacterial wilt caused by Ralstonia solanacearum is one of the most destructive bacterial diseases in agriculture. There is no effective control method, although chemical pesticides are used to prevent this disease, but they may lead to serious problems of environmental pollution. Natural products from plants can be rich and environmentally friendly sources for a broad spectrum biological control of bacteria. This study focuses on the pericarp of mangosteen (Garcinia mangostana) using bioactivity-guided analysis of different fractions and liquid chromatography–mass spectrometry combined with multivariate analysis to determine markers of active fractions. Six prenyl xanthones, including two new xanthones, garcimangosxanthones H and I, were isolated and identified by NMR and HRESIMS. The biomarker γ-mangostin displayed significant activity against the phytopathogen R. solanacearum with an IC50 of 34.7 ± 1.5 μg/mL; γ-mangostin affected the bacterial morphology at a concentration of 16.0 μg/mL as seen with a scanning electron microscope image, and it significantly repressed the virulence-associated genes HrpB, FihD, and PilT of R. solanacearum. γ-Mangostin also reduced the symptoms of bacterial wilt disease effectively that is caused by R. solanacearum in tomato and tobacco seedlings in vitro. These results suggested that the use of γ-mangostin from the mangosteen pericarp against R. solanacearum may be used as a natural bacteriostatic agent in agriculture.
Introduction
Ralstonia solanacearum is an extremely virulent soil-borne phytopathogenic bacterium with a broad geographical distribution, extensive host range, persistence, and high viability in soil. This soil-borne bacterium causes bacterial wilt disease in more than 200 plant species (50 families), including key crops like tomato, peanut, potato, pepper, tobacco, banana, and eggplant,1 and this has resulted in serious economic losses in agriculture. For example, R. solanacearum can destroy potato crops, leading to an estimated $1 billion losses annually worldwide.2 The management of bacterial wilt with chemicals, biological agents, and cultural methods has been used to protect crops for decades.3 Even though various management strategies have been developed to control this disease, an efficient and environmentally friendly method is still lacking.4
Phytochemical biopesticides have great promise because they are often less toxic, less persistent, more environmentally friendly, and are nontarget organisms.5 Scientists have focused on plant-derived natural products that attempt to exploit and develop new biopesticides to overcome bacterial wilt. For instance, screening the activity of 52 tropical medicinal plants has shown that 25% (13 species) inhibited the growth of R. solanacearum.6 The isolated natural products included essential oils and monoterpenes,7,8 lansiumamide B,9 methyl gallate,10 protocatechualdehyde,11 hydroxycomarins,12 and flavonoids13 from different species that inhibited the survival of the soil-borne pathogen R. solanacearum and revealed potential for controlling bacterial wilt.
Mangosteen (Garcinia mangostana) is grown in Southeast Asia; it is a well-known tropical fruit and termed as the “queen of fruits” for its unique sweet–sour taste.14 The pericarp has been used in traditional medicine in Southeast Asia for the treatment of conditions likely caused by bacteria such as skin infections, diarrhea, abdominal pain, dysentery, and infected wounds.15,16 Phytochemical investigations revealed that the pericarp of mangosteen is rich in isoprenylated xanthones, which have a wide range of reported biological functions from antioxidant to anti-inflammatory, but for the purposes of this study, the most germane reports are its antimicrobial activity.17 Mangosteen has been used in various food products and animal feed supplementation, and mangosteen-based products may benefit for engineering and biomedical industries.18 However, the use of the fruit pericarp (peel) waste in crops and agriculture have received limited attention.
As part of our ongoing research on biological control agents from medicinal plants in South China, we have screened >150 medicinal plants against R. solanacearum in vitro, and the mangosteen pericarp was one of the most active. The objective of this study was to identify natural metabolites from the mangosteen pericarp with antimicrobial activity against R. solanacearum and thereby identifying a way to exploit this waste stream as a useful biological control for bacterial wilt.
Results and Discussion
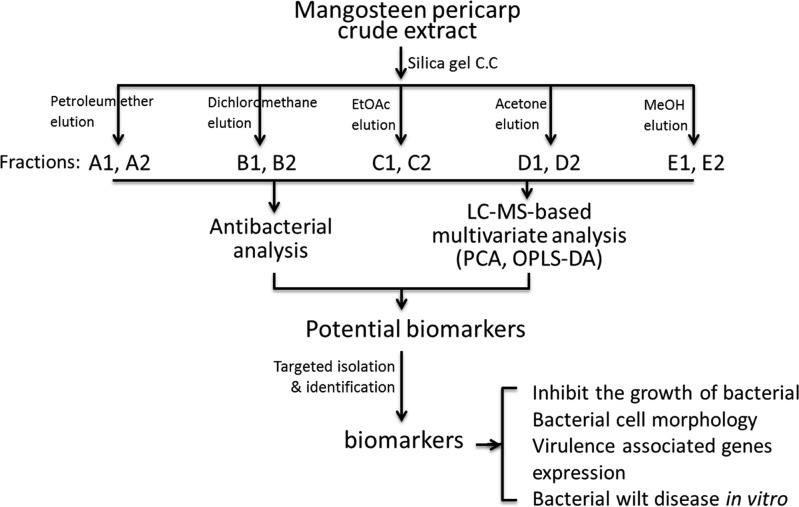
An ethanol extract of the mangosteen pericarp was separated over a silica gel column by eluting with petroleum ether, dichloromethane, ethyl acetate, acetone, and methanol sequentially to obtain 10 fractions. Each of the 10 fractions was then analyzed by liquid chromatography–mass spectrometry (LC–MS) and antibacterial assays. The results of bioactivity were used to construct multivariate analysis to help identify underlying biomarkers, which were further isolated and identified (Figure 1).
Figure 1.
Strategy applied to identify antibacterial metabolites from the mangosteen pericarp.
Antibacterial Activity of Screened Crude Extracts against R. solanacearum
The crude extract of the mangosteen pericarp showed significant effect on the growth of R. solanacearum using an agar disk dilution method (Figure S1A). In this study, the antibacterial metabolites from the pericarp were identified. First, the mangosteen pericarp was extracted with ethanol and separated over a silica gel column with petroleum ether, dichloromethane, ethyl acetate, acetone, and methanol sequentially, and this resulted in 10 crude fractions (A1, A2, B1..., E2). Each of the 10 fractions was tested for antibacterial activity under the same conditions. Only the dichloromethane fractions (Fr. B1 and B2) showed significant antibacterial activity (Figure S1B), and therefore, B1 and B2 were selected for metabolomics analysis and compound isolation.
LC–MS-Based Multivariate Analysis of Potential Antibacterial Markers
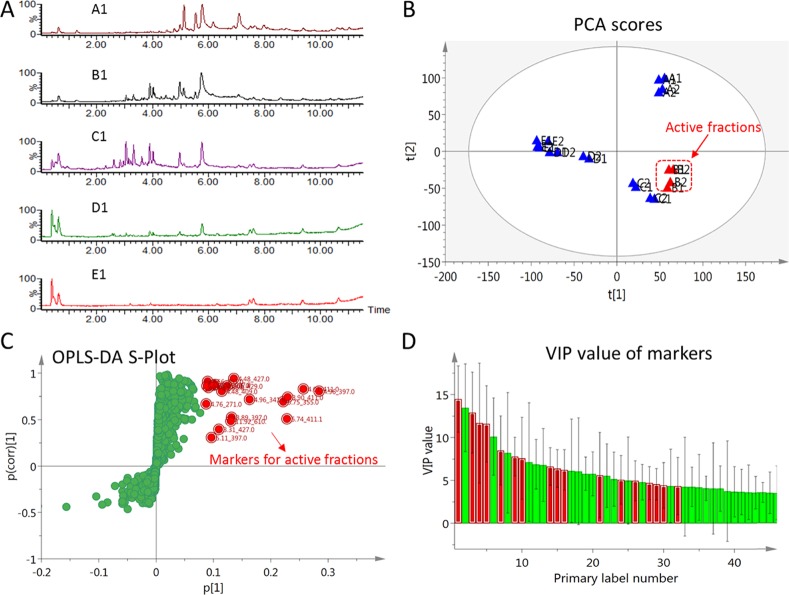
To investigate the specific antibacterial (R. solanacearum) compounds from fractions B1 and B2, bioactivity-guided isolation of antibacterial compounds coupled with LC–MS-based multivariate analysis was used in this study to determine biomarkers effectively. LC–MS analysis was optimized for all 10 fractions, as shown in Figure 2A. The chemical constituents of five selected fractions were different but share a common peak at 5.80 min. Positive mode was selected because more peaks were detected, and their relative abundance was higher compared to negative mode. For example, in negative mode, most of the peaks visible in positive mode from 1.0–4.0 and 7.3–12.0 min were missing (Figure S2). Therefore, the positive raw data was used and included retention time, exact mass (3858 ions in total), and ion intensity as variables in the multivariate analysis. Using a nontargeted PCA score plot, fractions B1–B2 and C1–C2 were clustered into one group, thus indicating that they share common metabolites (Figure 2B). However, among the 10 fractions, only dichloromethane fractions (Fr. B1 and B2) displayed significant antibacterial activity against R. solanacearum.
Figure 2.
UPLC-MS combined with multivariate analysis to determine markers from the most active fractions of the mangosteen pericarp extract. (A) Total ion chromatograms (TIC) of different fractions isolated by silica gel chromatography (positive mode). A1, B1..., E1 represented the TIC of selected fractions A1, B1..., E1. (B) Untargeted PCA analysis of all the fractions, R2X[1] = 0.22 and R2X[2] = 0.15, drawn with Hotelling’s 95% confidence ellipse; the red highlight shows the active fractions. A1, A2, B1..., E2 indicated the fractions A1, A2.., E2, as shown in Figure 1. (C) S-plots based on OPLS-DA analysis to determine molecular ions of markers for the active fractions (inactive group, −1; active group, 1). R2Y = 0.98 and Q2 = 0.85. Red dots indicate the priority potential biomarkers in active groups. (D) Variable importance in the projection (VIP) obtained from the OPLS-DA model for the most discrimination among active fractions from inactive fractions. Red bar graphs correspond to the selected markers in OPLS-DA.
OPLS-DA with the S-plot model was used to separate trends between active and inactive samples or clarify different samples, leading to select potential biologically active compounds.19 We compared two groups using OPLS-DA: fractions B1 and B2 were active, and fractions A1, A2, C1, C2, D1, D2, E1, and E2 were inactive. The constructed OPLS-DA model performed well based on the R2Y = 0.98 and Q2 = 0.85 (Figure 2C). The OPLS-DA model was validated by permutation test 200 times, and Q2 intercepts the y axis below zero (−0.282), indicating that the model is not overfitting (Figure S3). For the active fractions, the highlighted points in the red box at the top of the S-plot (Figure 2C) may correlate with biological activity. Therefore, 17 ions were selected as markers for potential antibacterial metabolites in fractions B1 and B2 (Table 1). The variable importance for the projection (VIP) >1 generally identifies a variable’s importance to the model, which helped to identify significant metabolites that contributed most to group separation in the OPLS-DA.20 The higher values of VIP suggested which active markers in the active fraction had priority potential (Figure 2D). Several markers had the same molecular weights, and it was therefore challenging to separate and isolate these isomers. Because the exact mass is not possible to measure with the TQD-MS used in this study, mass ions and retention times provided important information to identify target markers.
Table 1. Markers Information for the Active Fractions Were Found Using the S-Plot Model of Supervised Orthogonal Partial Least Squares Discriminate Analysis.
| marker no. | retention time(min) | mass [M + H]+(m/z) | VIP valuea |
|---|---|---|---|
| 1 | 4.96 | 397.07 | 14.48 |
| 2 | 4.01 | 411.09 | 12.91 |
| 3 | 3.90 | 411.08 | 11.68 |
| 4 | 5.75 | 355.05 | 11.60 |
| 5 | 4.96 | 341.05 | 8.49 |
| 6 | 5.11 | 397.07 | 7.79 |
| 7 | 3.89 | 397.07 | 7.58 |
| 8 | 4.48 | 427.07 | 6.63 |
| 9 | 3.31 | 427.06 | 6.39 |
| 10 | 4.06 | 429.08 | 6.24 |
| 11 | 4.48 | 409.06 | 5.61 |
| 12 | 4.30 | 427.07 | 5.08 |
| 13 | 5.02 | 465.09 | 4.97 |
| 14 | 4.76 | 271.05 | 4.70 |
| 15 | 4.02 | 355.05 | 4.56 |
| 16 | 4.56 | 397.07 | 4.42 |
| 17 | 4.99 | 411.08 | 4.34 |
Variable importance for the projection (VIP) was obtained from OPLS-DA with a threshold of 1.0. The VIP plot is sorted from high to low and shows confidence intervals for the VIP values, normally at the 95% level.
Identification of Possible Biomarkers
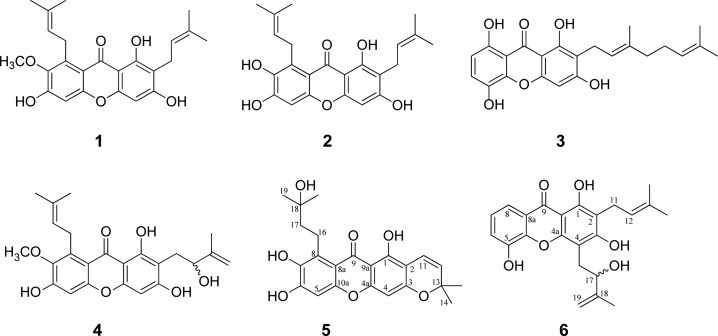
LC–MS-based multivariate analysis provided 17 candidate marker ions. These candidate antibacterial metabolites were isolated by sequential chromatographies that included a silica gel column, Sephedax LH-20 chromatography, and semipreparative HPLC to yield six compounds. Their structures were identified using nuclear magnetic resonance (NMR), UV, and HRESIMS spectroscopy as four known compounds α-mangostin (1),21 γ-mangostin (2),21 smeathxanthone A (3),22 and mangostenol (4),23 and compounds 5 and 6 were characterized as new xanthones (Figure 3).
Figure 3.
Isolated compounds 1 to 6 from the mangosteen pericarp (1; α-mangostin; 2, γ-mangostin; 3, smeathxanthone A; 4, mangostenol; 5, garcimangosxanthone H; 6, garcimangosxanthone I).
Compound 5, a yellow amorphous powder with a molecular ion at m/z 413.1608 [M + H]+, corresponds to a molecular formula of C23H24O7, with an index of hydrogen deficiency of 12. The 1H NMR spectrum (Table 2) for 5 showed the presence of four methyl groups [δ 1.42 (6H, s) and 1.20 (6H, s)], one chelated phenolic hydroxyl group (δ 14.18), two methylene protons [δ 1.57 (2H, m) and 3.30 (2H, dd, J = 9.7, 6.8 Hz)], one cis-olefinic proton [δ 6.60 (1H, d, J = 10.0 Hz), 5.72 (1H, d, J = 10.0 Hz)], and two aromatic singlets (δ 7.10 and 7.04). In the 13C NMR spectrum, there were signals for two methylene groups at δ 21.9 (C-16) and δ 43.6 (C-17), two methyl carbons at δ 29.1 (C-19 and C-20), and a quaternary carbon connected to a hydroxyl group at δ 69.3 (C-18); these data suggested that 5 contains a 3-hydroxy-3-methylbutyl moiety. Other signal sets included a 2,2-dimethychromene ring at δ 114.8 (C-11), 127.8 (C-12), 77.9 (C-13), and 27.9 (C-14 and C-15). All these moieties and the NMR chemical shift of 5 are similar to those of garcimangosxanthone E,24 except for the absence of a signal for the aromatic methyoxy group. The structure assignment of 5 was supported further by the HMBC spectrum (Figure 4). The 3-hydroxy-3-methylbutyl moiety was attached to the C-8 carbon based on the HMBC corrections from H-16 to C-7, C-8, and C-8a. The HMBC correlations between Me-15/C-12, C-13, and C-14, H-12/C-2, H-11/C-1, C-3, and C-13 deduced the 2,2-dimethylchromene ring that was attached at C-2 and C-3. Other key HMBC and 1H-1H COSY are shown in Figure 4. Accordingly, the structure of 5 was determined as 1,6,7-trihydroxy-8-(3-hydroxy-3-methylbutyl)-6′,6′-dimethylpyrano[2′,3′:3,2] xanthone, a new trioxyengated xanthone named as garcimangosxanthone H.
Table 2. Nuclear Magnetic Resonance (NMR) Spectroscopic Data (600 MHz, DMSO) for 5 and 6.
| 5 | 6 | |||
|---|---|---|---|---|
| position | δC | δH (J in Hz) | δC | δH (J in Hz) |
| 1 | 158.9 | 14.18, s | 158.0 | |
| 2 | 103.4 | 110.6 | ||
| 3 | 157.1 | 162.4 | ||
| 4 | 93.6 | 6.32, s | 104.3 | |
| 4a | 155.6 | 152.6 | ||
| 5 | 100.0 | 6.73, s | 147.6 | |
| 6 | 152.1 | 11.18, s | 120.6 | 7.28, dd (7.8, 1.3) |
| 7 | 141.0 | 8.69, s | 124.0 | 7.24, t (7.8) |
| 8 | 129.9 | 114.6 | 7.55, dd (7.8, 1.3) | |
| 8a | 110.1 | 120.3 | ||
| 9 | 181.7 | 180.8 | ||
| 9a | 102.9 | 102.2 | ||
| 10a | 152.8 | 146.5 | ||
| 11 | 114.8 | 6.60, d (10.0) | 21.4 | 3.28, d (7.0) |
| 12 | 127.8 | 5.72, d (10.0) | 122.4 | 5.18, d (7.4) |
| 13 | 77.9 | 130.8 | ||
| 14 | 29.1 | 1.20, s | 25.6 | 1.61, s |
| 15 | 29.1 | 1.20, s | 17.9 | 1.67, s |
| 16 | 43.6 | 1.57, m | 29.2 | 3.21, dd (14.7, 3.2), 3.03, dd (14.8, 7.7) |
| 17 | 21.9 | 3.30, dd (9.7, 6.8) | 74.6 | 4.32, m |
| 18 | 69.3 | 1.3, s | 144.9 | |
| 19 | 27.9 | 1.42, s | 109.8 | 4.86, s; 4.72, s |
| 20 | 27.9 | 1.42, s | 18.5 | 1.81, s |
Figure 4.

Selected key COSY and HMBC correlations of the two new xanthones.
Compound 6, a yellow amorphous, colorless powder with a molecular formula of C23H24O6, is based on HR-ESI-MS (m/z 397.1659, [H + M]+) and 12° of unsaturation. The 1H and 13C NMR spectra (Table 2) contained one chelated hydroxyl group at δ 13.21. The aromatic region showed a clear ABX pattern, which was revealed by resonances at δ 7.28 (1H, dd, J = 7.8, 1.3 Hz), 7.24 (1H, t, J = 7.8 Hz), and δ 7.28 (1H, dd, J = 7.8, 1.3 Hz). A 2-hydroxy-3-methylbut-3-enyl group was established based on its characteristic pattern [δ 4.86 and 4.72 (1H, each, s), 4.32 (1H, m), 3.21(1H, dd, J = 14.7, 3.2), 3.03 (1H, dd, J = 14.8, 7.7), and 1.81(3H, s)] and one prenyl group [δ 3.28 (2H, d, J = 7.0, 1.3 Hz), 5.18 (1H, d, J = 7.4 Hz), 1.61 (3H, s), and 1.67 (3H, s)]. The 13C NMR and DEPT spectroscopic data disclosed the presence of 23 carbon atoms, the characteristic carbonyl at δ 180.9, two benzene rings, and 2-hydroxy-3-methylbut-3-enyl and prenyl groups, which indicated that 6 was a trioxygenated xanthone derivative.
The characteristic 6, 7, and 8 protons in the B ring was confirmed further by the 1H-1H COSY (Figure 4). The HMBC showed that H-8 correlated with C-9 (δ 180.9), which indicated that C-5 was substituted by a phenolic group. In the HMBC spectrum, the chelated hydroxyl proton correlated to three aromatic carbons at δ 110.6 (C-2), δ 158.0 (C-1), and δ 102.2 (C-9a). Furthermore, H-11 correlated with C-1, C-2, and C-3 of the A ring, which indicated that the prenyl group was attached to the C-2 carbon (Figure 4). Moreover, the key HMBC correlation (Figure 4) between H-16 and two oxygenated aromatic carbons at δ 162.4 (C-3) and δ 152.6 (C-4a) confirmed that the 2-hydroxy-3-methylbut-3-enyl group was assigned to C-4. Therefore, compound 6 was identified as 1,3,5-trihydroxy-2-(3-methylbut-2-enyl)-4-(2-hydroxy-3-methylbut-3-enyl) xanthone, a new xanthone isolated from a natural source named as garcimangosxanthone I.
Antibacterial Analysis of Markers
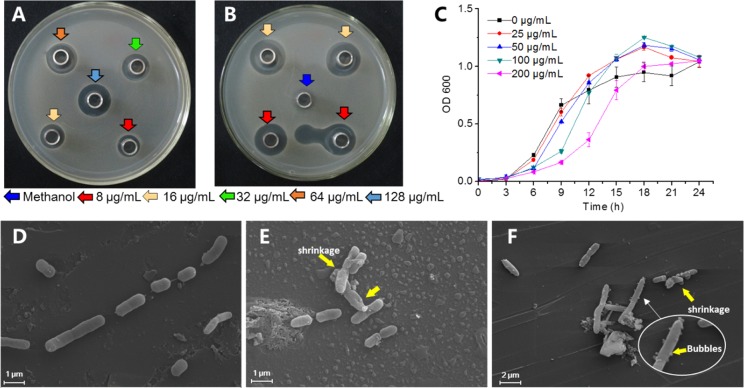
Six isolated compounds (1–6) were tested for their antibacterial activity against R. solanacearum. Only γ-mangostin (2) showed a significant effect on the growth of R. solanacearum in an agar medium (Figure 5A,B), α-mangostin (1) displayed weak antibacterial activity, and other compounds were inactive. The VIP value of 2 (14.48) was the highest in the OPLS-DA model (Table 1), which was the most active compound in the active fractions.
Figure 5.
Biomarker γ-mangostin inhibited the growth of R. solanacearum. (A) Antibacterial studied using the Oxford cup method on the agar medium at different concentrations of γ-mangostin. (B) Different concentrations of streptomycin sulfate served as positive control against R. solanacearum; methanol used as a negative control. (C) Effect of γ-mangostin on the growth of R. solanacearum in a liquid medium. (D–F) SEM image of R. solanacearum cells treated with γ-mangostin. Panel (D), untreated control; panels (E) and (F), treated with γ-mangostin at final concentrations of 16 and 128 μg/mL, respectively. Yellow arrows show the changes in the cells.
We conducted further research on the effect of γ-mangostin (2) on the growth of R. solanacearum. The inhibitory rate of γ-mangostin using a treatment of 8 μg/mL was 37.5%, and it showed a relatively high effect with an IC50 value of 34.7 ± 1.5 μg/mL (Figure S4). This indicates that the γ-mangostin in the pericarp can contribute significantly to the antibacterial activity against R. solanacearum. In a liquid medium, a low concentration of γ-mangostin did not affect the growth curve of pathogenic bacteria (Figure 5C), but when the bacterium was treated with higher concentrations (100 and 200 μg/mL), it delayed the growth in the logarithmic phase. In addition, morphological changes in R. solanacearum treated with γ-mangostin were observed using a scanning electron microscope. Some cells were deformed, and the cell surface showed different degrees of shrinkage under a treatment with 16 μg/mL γ-mangostin, although the cell surface was still smooth (Figure 5E). In treatments with high concentrations at 128 μg/mL, cell surfaces were rough, irregular, exhibited significant shrinkage, and they were accompanied by many bubble-like protrusions (Figure 5F), which suggested that γ-mangostin inhibited the growth of R. solanacearum and caused morphological changes.
Biomarker Represses the Virulence Associated Genes of R. solanacearum
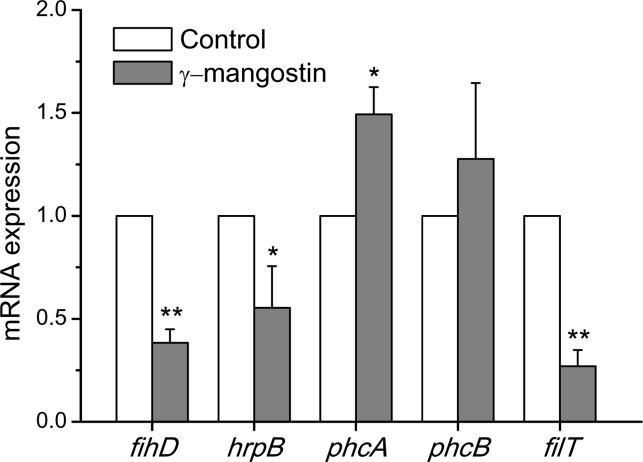
Biomarker γ-mangostin affected the pathogenesis of R. solanacearum, which was supported further by analyses of the expression of five virulence-associated genes (hrpB, phcA, phcB, fihD, and pilT). Quantitative RT-PCR showed that γ-mangostin significantly repressed the expression of hrpB, fihD, and pilT with a treatment of 64 μg/mL, but in contrast, genes PhcA and PhcB showed enhanced expression (Figure 6). The virulence and pathogenicity in R. solanacearum is controlled by a complex regulatory network, although two determinants that include a phenotype conversion (Phc) system and a hrp-encoded Type III secretion system (TTSS) responded to environmental stimuli.25 The regulatory gene hrpB controlled TTSS in diseases caused by R. solanacearum,26 but its expression was negatively regulated by phcA.27 Our data indicated that an increase in phcA and phcB led to the repression of hrpB genes under the treatment by γ-mangostin. Swimming motility played a key role in the prophase infection by R. solanacearum. Two important genes (fihD and pilT) that were responsible for motility were repressed significantly that they affected the swimming motility of R. solanacearum. The process of pathogenicity in R. solanacearum is controlled by a completed regulation system and involves the expression of multiple genes in response to the environment. γ-Mangostin affected the virulence-associated genes of R. solanacearum, but this requires more experiments for clarification.
Figure 6.
Quantitative RT-PCR analyses of virulence-associated genes of R. solanacearum treated with γ-mangostin. R. solanacearum was treated with DMSO for the control and with 64 μg/mL γ-mangostin. *P < 0.05, **P < 0.001 compared with the control group in one-way ANOVA.
Efficacious Control of Ralstonia Wilt Disease
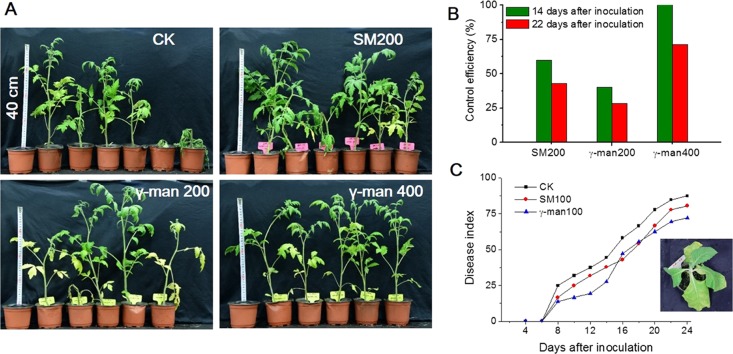
The bioactive compound γ-mangostin was evaluated for antibacterial activity in vivo caused by R. solanacearum in both tomato and tobacco seedlings. In the tomato pot experiment, some of the plants in the negative control group (CK) wilted completely after 14 days of post-inoculation, although treatment with γ-mangostin and streptomycin sulfate (SM) reduced Ralstonia wilt disease (Figure 7A). In particular, no leaves wilted under a high concentration of γ-mangostin (400 mg/L). The control efficiency of γ-mangostin was evaluated at 14 and 22 days after inoculation with R. solanacearum (Figure 7B). γ-Mangostin control efficiency was lower than streptomycin sulfate (SM) at the same treatment concentration. However, a higher concentration at 400 mg/L reduced bacterial wilt disease effectively, and the control efficiency was 71.4% at 22 days after treatment. These results suggest that the effectiveness of γ-mangostin against tomato bacterial wilt related closely with the concentration that was applied. In addition, the leaves of tobacco seedlings inoculated with R. solanacearum wilted 8 days after inoculation (Figure 7C). The trends in the disease index for γ-mangostin and streptomycin sulfate were similar, and the emergence of wilting symptoms was delayed as the disease index decreased compared with the untreated control (CK). The disease index of γ-mangostin was lower than that of streptomycin sulfate before 14 days. These results suggested that γ-mangostin reduced bacterial wilt in tomato and tobacco seedlings, and it might control plant bacterial wilt in plants generally.
Figure 7.
Effect of bioactive compound from the mangosteen pericarp on tomato and tobacco seedlings inoculated with R. solanacearum. (A) Photographs of tomato seedlings after 14 days of inoculation. (B) Control efficiency was assessed 14 and 22 days after inoculation, and it was calculated as the percentage of plants that wilted for one experiment (CK, untreated control; SM200, indicated 200 mg/L streptomycin sulfate; γ-man200 and γ-man400, represented seedlings treated with γ-mangostin of 200 and 400 mg/L, respectively). (C) Progress of Ralstonia wilt disease on tobacco plants (SM100, 100 mg/L streptomycin sulfate, γ-man100, and 100 mg/L γ-mangostin). Each data point represents the mean percentage of leaves that wilted for two independent experiments. Each experiment included 36 plants.
Conclusions
The pericarps of mangosteen underwent bioactivity-guided analysis of different fractions. Liquid chromatography–mass spectrometry combined with multivariate analysis was used to effectively identify antibacterial fractions in the active fractions. Six prenyl xanthones, including γ-mangostin and two new xanthones, were isolated and identified. Among the six tested compounds, γ-mangostin showed the best antibacterial activity against the phytopathogen R. solanacearum in vitro. The effect of γ-mangostin on selected genes in R. solanacearum was evaluated for the first time. γ-Mangostin controlled bacterial wilt effectively, and it might have the potential to be developed as a natural bactericide to control plant bacterial wilt in the future.
Experimental Section
General Experimental Procedures
NMR spectra were recorded by a Bruker AVANCE-600 (600 MHz) instrument (Bruker Biospin, Zurich, Switzerland). UV spectra were recorded by an Evolution 300 UV–vis spectrometer (Thermo Fisher Scientific). UPLC-TQD-MS was operated using an Acquity UPLC system (Waters Corporation, Milford, MA, USA) coupled with a MS (Xevo TQD, Waters MS Technologies, Manchester, UK). HRESIMS spectra were obtained from UPLC-QTOF-MS (Agilent Inc., Santa Clara, CA, USA). HPLC separations used the Agilent 1100 HPLC equipped with a UV detector with a semipreparative column (Zorbax 300 SB-C18 column, 9.4 mm × 25 cm, 4 μm). Sephadex LH-20 (25–100 μm) was purchased from Pharmacia Fine Chemicals (Piscataway, NJ), and HPLC-MS grade acetonitrile, water, and formic acid were purchased from J. T. Baker (Phillipsburg, NJ, USA). Thin-layer chromatography (TLC) silica gel plates and a silica gel of 200–300 mesh were obtained from Qingdao Haiyang Chemical Co. Ltd. (P. R. China), and all reagents were analytical grade (Guangzhou Chemical Reagent Factory, P. R. China).
Bacterial Strains and Culture Conditions
The R. solanacearum strain was obtained from the Laboratory of Crop Ecology (South China Agricultural University) from a previous study.28 The strain was cultured on a nutrient agar (SMSA) medium (peptone, 10.0 g; casein enzymolysis, 1.0 g; glucose, 10.0 g; agar, 18.0 g; deionized water, 1000 mL; pH 7.0). Single colonies were transferred to nutrient broth (NB; liquid medium). The flasks were shaken at 180 r/min at 30 °C for 24 h and then stored at −80 °C before use.
Extraction and Isolation
Fresh mangosteen (G. mangostana) was purchased from the Guangzhou fruit market in July 2015 (Guangdong Province, southern China). The dried pericarp of mangosteen (0.7 kg) was grounded into a powder and extracted with a 95% ethanol soak for 36 h. The filter solution was concentrated under reduced pressure at 40 °C to yield the crude extract (280 g). Then, 255 g of extract was chromatographed using silica gel column chromatography and eluted by petroleum ether, dichloromethane, ethyl acetate, acetone, and methanol (2000 mL, each), which produced 10 combined fractions (Fr. A1, Fr. A2, Fr. B1, Fr. B2, ..., Fr. E1, and Fr. E2). Each fraction was then analyzed by LC–MS and antibacterial assays before preparative-scale isolation. The active fractions Fr. B1 and B2 were combined and labeled as Fr. B (65 g), and it was chromatographed using silica gel column chromatography with a gradient system of petroleum-acetone (100–10 → 0:100) to produce seven fractions (Fr. B1 → Fr. B7). Fr. B5 was isolated using Sephadax LH-20 (MeOH-dichloromethane, v/v = 1:1) to produce eight subfractions (Fr. B5–1 → Fr. B5–8). Fr. B5–5 was chromatographed using Sephadex LH-20 column chromatography to produce compounds 1 (25.5 mg) and 2 (40.2 mg). Fr. B5–6 was purified subsequently by reversed-phase preparative HPLC and gradient eluted MeOH/H2O to obtain compounds 4 (12.0 mg) and 6 (10.5 mg). Fr. B5–8 was chromatographed repeatedly over reversed-phase HPLC and eluted with a step gradient of MeCN/H2O, which yielded compounds 5 (4.5 mg) and 3 (9.0 mg). The spectroscopic data for new compounds are as follows:
Garcimangosxanthone H (5): yellow amorphous powder; 1H and 13C NMR (DMSO, 600 MHz) data, see Table 2; UV (MeOH) λ max (log ε): 243 (4.35), 260 (4.23), and 289 (4.22) nm; IR(KBr) υmax 2965, 2933, 1655, and 1621 cm–1; and HRESIMS m/z 413.1608 [M + H]+ (calculated for C23H25O7, 413.1600).
Garcimangosxanthone I (6): yellow amorphous powder; 1H and 13C NMR (DMSO, 600 MHz) data, see Table 2; UV (MeOH) λ max (log ε): 238 (4.40), 267 (0.77), 319(4.21), and 380 (3.50) nm; IR(KBr) υmax 3423, 2941, 2843, 1652, 1560, and 1381 cm–1; and HRESIMS m/z 397.1659 [M + H]+ (calculated for C23H25O6, 397.1651).
Antibacterial Activity of Fractions/Compounds from the Mangosteen Pericarp
Antibacterial activity of crude extracts or fractions against R. solanacearum was detected by the agar diffusion method with minor modifications.9 Nutrient agar (SMSA) was melted and cooled at room temperature to 35 °C, and the overnight-cultured, bacterial suspension was inoculated with SMSA to obtain bacteria-containing media with a concentration of R. solanacearum of OD600 = 0.1 (≈108 cfu/mL), mixed well, and poured into 9 cm diameter Petri dishes (15 mL each). Oxford cups were placed into plates, and 150 μL of different concentrations of samples was pipetted into the cups. All samples were performed three times, and methanol was used as a negative control. The diameters of inhibition zones were measured after samples were cultured for 48 h at 30 °C.
Antibacterial assay of compounds were performed in 96-well microtiter plates by a previously described method.29 Briefly, 50 μL of cultured bacterial suspension (OD600 = 0.2) was added to 50 mL of NB medium containing γ-mangostin at final concentrations ranging from 3.13 to 100 mg/L. The final volume is 100 μL in each well. The negative controls were treated with 1% methanol, and the dilutions were prepared with six replicates. The inoculated plates were incubated at 30 °C for 24 h after shaking at 280 rpm for 20 min on a shaker table. The absorbance data process at 600 nm was measured by a microplate reader (BioTek, Epoch2, USA). Inhibition percent was calculated using the following equation: % inhibition = (absorbancecontrol – absorbancesample)/(absorbancecontrol) × 100. The antibacterial activity was displayed as IC50.
The Growth Curve of R. solanacearum
A solution of γ-mangostin (200 mg/L) was added to 30 mL of NB medium at an aseptic workstation to obtain phytochemical-containing media with test concentrations of 10, 25, 50, 100, and 200 mg/L. A final concentration of DMSO of 0.5% was used as the solvent control. Then, 100 μL of bacterial suspension (OD600 = 1.0) that had been cultured for 24 h was inoculated with the phytochemical-containing medium, which was cultured at 30 °C with shaking at 180 r/min. The OD600 values of the samples were measured every 3 h for 24 h using an ultraviolet spectrophotometer. The experiment was repeated three times.
Observations of Cell Morphology with Scanning Electron Microscopy (SEM)
After 24 h, 100 μL of cultured bacterial suspension (OD600 = 1.0) was added to 30 mL of NB medium containing γ-mangostin at final concentrations of 16 and 128 mg/L, respectively. The mixtures were shaken at 180 r/min at 30 °C for 24 h. A mixture (1.0 mL) with OD600 = 1.6 was centrifuged for 8 min at 5000 r/min, and the supernatant was removed. Then, the mixture was rinsed with 0.1 M PBS buffer, and it was centrifuged three times to obtain the sediment. A 500 μL 2.5% glutaraldehyde was added to the sediment and mixed completely. A small amount of bacteria droplets was pipetted onto the coverslip, spread gently, and stored in a refrigerator at 4 °C for 12 h. The bacteria sample was washed with 0.1 M PBS buffer three times, fixed with 1% osmic acid in a hood for 30 min, and then washed with 0.1 M PBS buffer again. Next, different concentrations of ethanol were used to dehydrate the sample gradually (10 min per concentration). The morphology of the bacteria was observed under a scanning electron microscope after drying overnight in an oven.
Quantitative Real-Time PCR Assays
Total RNA was extracted from the collected cells of R. solanacearum using the RNAiso Plus reagent, according to the manufacturer’s instructions (Takara Bio Inc., Shanghai, China). RNA degradation and contamination were checked on 1% agarose gels, and RNA concentration and purity were monitored using a Thermo Scientific NanoDrop One spectrophotometer (Thermo Fisher Scientific, Inc., USA). cDNA was synthesized from 0.8 μg of total RNA using the PrimeScript II 1st Strand cDNA Synthesis Kit (Takara Bio Inc., Shanghai, China).
All quantitative real-time PCR (qRT-PCR) analyses were performed on the ABI 7500 Manager (Life Technologies Holdings Pte. Ltd., Singapore) in a 15 μL reaction system, which consisted of 5 μL of 3× SYBR Green qPCR mix (Takara Bio Inc., Shanghai, China), 5 μL of diluted cDNA, 0.3 μL of 50 × Rox, 0.4 mM each primer, and 3.5 μL of ddH2O. The amplification protocols were as follows: 2 min at 95 °C, followed by 40 cycles of 95 °C for 15 s, 60 °C for 45 s, and 72 °C for 30 s. After that, a melting curve from 60 to 95 °C was applied to test the specificity and consistency of the PCR products. Normalized gene expression was calculated by the Bio-Rad CFX Manager 3.0 software using the Δ Δ Cq method.
LC–MS-Based Multivariate Analysis
Comparative analyses of different fractions were performed on an ACQUITY UPLC system coupled with a triple-quadrupole Xevo TQD mass spectrometer (Waters MS Technologies, Manchester, UK). An ACQUITY UPLC BEH C18 column (2.1 mm × 50 mm, 1.7 μm) was employed, and the column temperature was maintained at 40 °C. Gradient elution with acetonitrile that contained 0.1% formic acid (A) and water that contained 0.1% formic acid (B) was performed as follows: 0–0.5 min, 20% B; 0.5–2.5 min, 20–65% B; 2.5–5.0 min, 65–70% B; 5.0–7.5 min, 70–80% B; 7.5–11.0 min, 80–95% B; and 11.0–13.0 min, 95% B; the column was reconditioned at 20% B for 2 min to prepare for the next injection. The flow rate was set at 0.3 mL/min. The auto-sampler was conditioned at 25 °C, and the injection volume of the solution was 2 μL for analysis. Mass spectrometric detection was performed on a Xevo TQD equipped with an electrospray ionization source (ESI). The capillary voltage was set to 3.5 kV, and the source temperature was maintained at 150 °C. The collision gas was Ar, N2 gas was used as desolvation at 400 °C, cone gas was at a flow rate of 750 L/h, and cone gas was set to 50 L/h. Both positive and negative data were collected by MS mode using a scan time of 0.5 s.
UPLC-MS full scan data were processed as described previously.19 Briefly, data preprocessing that included peak peaking, alignment, peak integration, and correction of retention time for all raw data was processed using MarkerLynx. The parameters used included a retention time range of 0.5–11.5 min, a mass range of 100–1000 Da, and a mass tolerance of 50 mDa. The intensity threshold (counts) for collection parameters was set at 1000, mass window was set at 0.05, retention time tolerance was set at 0.20, noise elimination level was set at 6.00, and isotopic peaks were excluded for analysis. The SIMCA 14.0 software (Umetrics, Umea, Sweden) was used for untargeted principal component analysis (PCA) and for supervised orthogonal partial least squares discriminate analysis (OPLS-DA) with Pareto scaling (Par) to normalization. To guard against model overfitting, the OPLS-DA model was validated with a 200 times permutation test. VIP scores and S-Plot of the OPLS-DA model were extracted for selected markers.
Pot Experiment
The control of bacterial wilt in tomato and tobacco seedlings was evaluated after they were inoculated with the bioactive compounds of R. solanacearum using the pot experiment performed with minor modification.29 Tomato seedlings were grown for 3 weeks in a controlled chamber and then transferred to vinyl pots with a diameter of 10 cm (one plant each) until the 5–6 leaf stage of plants. γ-Mangostin was prepared at final concentrations of 200 and 400 mg/L that contained 1% methanol in water. The phytochemical solution (20 mL) was poured into each tomato pot. After 3 h, potted plants were inoculated with 20 mL of R. solanacearum suspension (OD600 = 0.1) by pouring over the wounded root. The second treatment with γ-mangostin solutions was conducted 5 days after the first treatment. Plants treated with streptomycin sulfate (200 mg/L) were used as a bactericide control, and 1% methanol in water served as a negative control (CK). Each treatment consisted of nine plants. Bacterial wilt incidence (WI) was assessed at 14 and 22 days. Wilt incidence was calculated as the percentage of plants, which were completely wilted. Control efficacy was calculated as the following equation:10 control efficacy (%) = 100 × (WI of control – WI of treatment)/ WI of treatment.
The experiment of controlling tobacco bacterial wilt was conducted as in a previous study.30 Briefly, 30 mL of γ-mangostin (100 mg/L, with 1% methanol) was added to pots that contained one tobacco plant at the 6–7 leaf stage. After 12 h, potted plants were inoculated with 20 mL of a R. solanacearum suspension (OD600 = 0.1) by pouring over the root (noninjured inoculation). Streptomycin sulfate (100 mg/L) was used as the bactericide control, and 1% methanol in water was used as a negative control (CK). Each treatment consisted of two replicates, and each replicate contained nine plants. All plants in both experiments were maintained in a greenhouse at a temperature of 30 ± 2 °C, a relative humidity of 70–80%, and with a light cycle of 14 h of light and 10 h of dark. The occurrence of diseased tobacco seedlings was recorded every 2 days. Disease index was recorded accordingly as previously described29 and modified into a scale of 0–4: 0, no symptoms; 1, one leaf partially wilted; 2, one to two leaves wilted; 3, two to three leaves wilted; and 4, four or more leaves wilted or dead. The disease index was calculated as follows: disease index = [Σ(number of diseased plants × number of disease plants)/(total number of plants × representative value of the highest grade)] × 100.
Statistical Analysis
Data were present as means plus standard deviations. The significance of the treatments was analyzed by one-way ANOVA (P < 0.05). The IC50 values were calculated using GraphPad Prism 6.0.
Acknowledgments
This work was supported by the National Key Research and Development Program of China (2017YFD0201300), the Natural Science Foundation of Guangdong Province of China (2017A030310283), the National Natural Science Foundation of China (31800283), and the Science and Technology Planning Project of Guangdong Province, China (2019B030301007). We would like to thank Thomas A. Gavin, Professor Emeritus, Cornell University and Edward J. Kennelly, Professor, Lehman College, City University of New York, for helping us edit this paper.
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsomega.9b02746.
Supplementary figures and spectroscopic data including HRESIMS, 1D, and 2D NMR spectra of compounds 5 and 6 (PDF)
Author Contributions
Y.Z., B.T., and Q.Z performed the experiments. P.L. analyzed the NMR spectra and multivariate analysis. J.W., J.Z., and Z.C. contributed the reagents/materials/analysis tools. J.Y. and L.S. conceived and designed the experiments, and P. L. and J. Y. wrote the manuscript.
The authors declare no competing financial interest.
Supplementary Material
References
- Nahar K.; Matsumoto I.; Taguchi F.; Inagaki Y.; Yamamoto M.; Toyoda K.; Shiraishi T.; Ichinose Y.; Mukaihara T. Ralstonia solanacearum type III secretion system effector Rip36 induces a hypersensitive response in the nonhost wild eggplant Solanum torvum. Mol. Plant Pathol. 2014, 15, 297–303. 10.1111/mpp.12079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elphinstone J. G.The current bacterial wilt situation: a global overview. In Bacterial wilt the disease & the Ralstonia Solanacearum species complex; Allen C., Prior P., Hayward A. C., Eds.; APS Press: St. Paul, MN, USA, 2005, 9–28. [Google Scholar]
- Yuliar; Nion Y. A.; Toyota K. Recent trends in control methods for bacterial wilt diseases caused by Ralstonia solanacearum. Microbes Environ. 2015, 30, 1–11. 10.1264/jsme2.ME14144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang G.; Wei Z.; Xu J.; Chen H.; Zhang Y.; She X.; Macho A. P.; Ding W.; Liao B. Bacterial wilt in China: History, current status, and future perspectives. Front. Plant Sci. 2017, 8, 1549. 10.3389/fpls.2017.01549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walia S.; Saha S.; Tripathi V.; Sharma K. K. Phytochemical biopesticides: Some recent developments. Phytochem. Rev. 2017, 16, 989–1007. 10.1007/s11101-017-9512-6. [DOI] [Google Scholar]
- Jiang W. T.; Dai H. F.; Liu S. B.; Mei W. L. Activity screening of fifty-two species of tropical officinals against tobacco green blight (Ralstonia solanacearum) in vitro. J. Microbiol. 2011, 31, 15–18. [Google Scholar]
- Pradhanang P. M.; Momol M. T.; Olson S. M.; Jones J. B. Effects of plant essential oils on Ralstonia solanacearum population density and bacterial wilt incidence in tomato. Plant Dis. 2003, 87, 423–427. 10.1094/PDIS.2003.87.4.423. [DOI] [PubMed] [Google Scholar]
- Mohamed A. A.; Behiry S. I.; Younes H. A.; Ashmawy N. A.; Salem M. Z. M.; Márquez-Molina O.; Barbabosa-Pilego A. Antibacterial activity of three essential oils and some monoterpenes against Ralstonia solanacearum phylotype II isolated from potato. Microb. Pathog. 2019, 135, 103604. 10.1016/j.micpath.2019.103604. [DOI] [PubMed] [Google Scholar]
- Li L.; Feng X.; Tang M.; Hao W.; Han Y.; Zhang G.; Wan S. Antibacterial activity of Lansiumamide B to tobacco bacterial wilt ( Ralstonia solanacearum ). Microbiol. Res. 2014, 169, 522–526. 10.1016/j.micres.2013.12.003. [DOI] [PubMed] [Google Scholar]
- Yuan G.-Q.; Li Q.-Q.; Qin J.; Ye Y.-F.; Lin W. Isolation of methyl gallate from Toxicodendron sylvestre and its effect on tomato bacterial wilt. Plant Dis. 2012, 96, 1143–1147. 10.1094/PDIS-03-11-0150-RE. [DOI] [PubMed] [Google Scholar]
- Li S.; Yu Y.; Chen J.; Guo B.; Yang L.; Ding W. Evaluation of the antibacterial effects and mechanism of action of protocatechualdehyde against Ralstonia solanacearum. Molecules 2016, 21, 754. 10.3390/molecules21060754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang L.; Ding W.; Xu Y.; Wu D.; Li S.; Chen J.; Guo B. New insights into the antibacterial activity of hydroxycoumarins against Ralstonia solanacearum. Molecules 2016, 21, 468. 10.3390/molecules21040468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao X.; Mei W.; Gong M.; Zuo W.; Bai H.; Dai H. Antibacterial activity of the flavonoids from Dalbergia odorifera on Ralstonia solanacearum. Molecules 2011, 16, 9775–9782. 10.3390/molecules16129775. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aizat W. M.; Jamil I. N.; Ahmad-Hashim F. H.; Noor N. M. Recent updates on metabolite composition and medicinal benefits of mangosteen plant. PeerJ 2019, 7, e6324 10.7717/peerj.6324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Obolskiy D.; Pischel I.; Siriwatanametanon N.; Heinrich M. Garcinia mangostana L.: a phytochemical and pharmacological review. Phytother. Res. 2009, 23, 1047–1065. 10.1002/ptr.2730. [DOI] [PubMed] [Google Scholar]
- Pedraza-Chaverri J.; Cárdenas-Rodríguez N.; Orozco-Ibarra M.; Pérez-Rojas J. M. Medicinal properties of mangosteen (Garcinia mangostana). Food Chem. Toxicol. 2008, 46, 3227–3239. 10.1016/j.fct.2008.07.024. [DOI] [PubMed] [Google Scholar]
- Ovalle-Magallanes B.; Eugenio-Pérez D.; Pedraza-Chaverri J. Medicinal properties of mangosteen (Garcinia mangostana L.): A comprehensive update. Food Chem. Toxicol. 2017, 109, 102–122. 10.1016/j.fct.2017.08.021. [DOI] [PubMed] [Google Scholar]
- Aizat W. M.; Ahmad-Hashim F. H.; Syed Jaafar S. N. Valorization of mangosteen, ″The Queen of Fruits,″ and new advances in postharvest and in food and engineering applications: A review. J. Adv. Res. 2019, 20, 61–70. 10.1016/j.jare.2019.05.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li P.; Yue G. G. L.; Kwok H. F.; Long C. L.; Lau C. B. S.; Kennelly E. J. Using ultra-performance liquid chromatography quadrupole time of flight mass spectrometry-based chemometrics for the identification of anti-angiogenic biflavonoids from edible Garcinia species. J. Agric. Food Chem. 2017, 65, 8348–8355. 10.1021/acs.jafc.7b02867. [DOI] [PubMed] [Google Scholar]
- Zhao Y.; Zhao Y. Y.; Du Y.; Kang J. S. Characterization and classification of three common Bambusoideae species in Korea by an HPLC-based analytical platform coupled with multivariate statistical analysis. Ind. Crops Prod. 2019, 130, 389–397. 10.1016/j.indcrop.2018.12.096. [DOI] [Google Scholar]
- Suksamrarn S.; Suwannapoch N.; Phakhodee W.; Thanuhiranlert J.; Ratananukul P.; Chimnoi N.; Suksamrarn A. Antimycobacterial activity of prenylated xanthones from the fruits of Garcinia mangostana. Chem. Pharm. Bull. 2003, 51, 857–859. 10.1248/cpb.51.857. [DOI] [PubMed] [Google Scholar]
- Komguem J.; Meli A. L.; Manfouo R. N.; Lontsi D.; Ngounou F. N.; Kuete V.; Kamdem H. W.; Tane P.; Ngadjui B. T.; Sondengam B. L.; Connolly J. D. Xanthones from Garcinia smeathmannii (Oliver) and their antimicrobial activity. Phytochemistry 2005, 66, 1713–1717. 10.1016/j.phytochem.2005.03.010. [DOI] [PubMed] [Google Scholar]
- Suksamrarn S.; Suwannapoch N.; Ratananukul P.; Aroonlerk N.; Suksamrarn A. Xanthones from the green fruit hulls of Garcinia mangostana. J. Nat. Prod. 2002, 65, 761–763. 10.1021/np010566g. [DOI] [PubMed] [Google Scholar]
- Zhou X.; Huang R.; Hao J.; Huang H.; Fu M.; Xu Z.; Zhou Y.; Li X.-E.; Qiu S. X.; Wang B. Two new prenylated xanthones from the pericarp of Garcinia mangostana (Mangosteen). Helv. Chim. Acta 2011, 94, 2092–2098. 10.1002/hlca.201100157. [DOI] [Google Scholar]
- Schell M. A. Control of virulence and pathogenicity genes of Ralstonia solanacearum by an elaborate sensory network. Annu. Rev. Phytopathol. 2000, 38, 263–292. 10.1146/annurev.phyto.38.1.263. [DOI] [PubMed] [Google Scholar]
- Cunnac S.; Occhialini A.; Barberis P.; Boucher C.; Genin S. Inventory and functional analysis of the large Hrp regulon in Ralstonia solanacearum: identification of novel effector proteins translocated to plant host cells through the type III secretion system. Mol. Microbiol. 2004, 53, 115–128. 10.1111/j.1365-2958.2004.04118.x. [DOI] [PubMed] [Google Scholar]
- Genin S.; Brito B.; Denny T. P.; Boucher C. Control of theRalstonia solanacearumType III secretion system (Hrp) genes by the global virulence regulator PhcAControl of the Ralstonia solanacearum Type III secretion system (Hrp) genes by the global virulence regulator PhcA. FEBS Lett. 2005, 579, 2077–2081. 10.1016/j.febslet.2005.02.058. [DOI] [PubMed] [Google Scholar]
- Chen Y.; Liu M.; Wang L.; Lin W.; Fan X.; Cai K. Proteomic characterization of silicon-mediated resistance against Ralstonia solanacearum in tomato. Plant Soil 2015, 387, 425–440. 10.1007/s11104-014-2293-4. [DOI] [Google Scholar]
- Le Dang Q.; Shin T. S.; Park M. S.; Choi G. J.; Jang K. S.; Kim I. S.; Kim J.-C. Antimicrobial activities of novel mannosyl lipids isolated from the biocontrol fungus Simplicillium lamellicola BCP against phytopathogenic bacteria. J. Agric. Food Chem. 2014, 62, 3363–3370. 10.1021/jf500361e. [DOI] [PubMed] [Google Scholar]
- Chen J.; Yu Y.; Li S.; Ding W. Resveratrol and coumarin: Novel agricultural antibacterial agent against Ralstonia solanacearum in vitro and in vivo. Molecules 2016, 21, 1501. 10.3390/molecules21111501. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.