Abstract
To monitor rodent colony health in research facilities, soiled-bedding sentinel (SBS) animals have traditionally been used. SBS can be tested by various methods, which may include serology, PCR analysis, and necropsy. Several pathogens are unreliably detected by using SBS or transmitted poorly through soiled bedding, and collection and evaluation of SBS samples can be time-intensive. Recently, exhaust air dust (EAD) testing through PCR analysis has emerged as an adjunct or replacement method for rodent colony health monitoring. EAD monitoring may provide a more efficient, sensitive, and humane method for monitoring health status. Using both EAD and SBS health monitoring, we evaluated colony health over the course of 1 y in 3 research barrier rooms in which mice were housed exclusively on IVC racks. Three pathogens—Helicobacter spp., Rodentibacter spp. (previously Pasteurella pneumotropica), and murine norovirus (MNV)—were not excluded in 2 of the rooms, and we expected that these mice would test positive with some regularity. EAD monitoring was significantly more sensitive than SBS for detection of the bacterial agents. SBS failed to detect Helicobacter spp. at time points when EAD had 100% detection in the rooms that did not exclude the bacteria. The detection of MNV did not differ between health monitoring systems at any time point. The findings suggest that EAD is especially valuable in detecting bacteria poorly transmitted through soiled bedding. In addition, the corresponding results with MNV detection suggest that EAD surveillance can reliably be implemented as an alternative to SBS monitoring in a facility in which mice are housed exclusively on IVC racks.
Abbreviations: EAD, exhaust air dust; SBS, soiled-bedding sentinel; MB, mouse barrier; MB+, mouse barrier plus; MNV, murine norovirus; LDV, lactate dehydrogenase-elevating virus
Laboratory mice (Mus musculus) have been used in research for at least 300 y36 and are one of the most widely used animal models in biomedical research. Rodent research colonies harbor numerous adventitious agents, which can cause clinical disease as well as interfere with research results. To manage these adventitious agents, barrier housing practices and health monitoring programs have been developed to exclude these pathogens and to efficiently detect them when they are inadvertently introduced into the colony. The traditional form of mouse health monitoring in academic institutions has been soiled bedding sentinels (SBS), but there are limitations to its use.
SBS health monitoring relies on the transmission of the infectious agents from colony animals to sentinel animals, mainly through the transfer of soiled bedding. Sentinel animals are tested at regularly scheduled intervals by using diagnostics such as physical examination, serology, PCR analysis, bacterial culture, necropsy, and histopathology. There are many different methods for implementing an SBS program, and they vary with the caging system in use, sentinel mice used, diagnostics performed, frequency of testing, and how mice are exposed to the soiled bedding from the colony.
SBS mice are exposed to the soiled bedding of research colony mice at each cage change. Typically, a predetermined amount of soiled bedding from each colony cage on the rack is placed in a new clean cage for SBS mice. When open-top caging is used, sentinel cages also have the potential to be exposed to particulates and aerosols from colony mice. With advancements in caging design, such as microisolation cages and IVC, this direct exposure method is unlikely because of the microisolation lid, which provides containment at the cage level. This set-up decreases the exposure potential to what is present in the soiled bedding and therefore decreases the likelihood of transmission of infectious agents to sentinels, especially those agents that transmit by means of aerosols.2,5,8,25
In addition, the effectiveness of an SBS monitoring system is affected by the agents monitored. Some agents are not transmitted through the fecal-oral route, including Sendai virus and Filobacterium rodentium (previously cilia-associated respiratory bacillus), or have low prevalence, such as some host-adapted bacteria and parasites, and are not transmitted reliably or at all through soiled bedding.1,5,8,9,17,23,27,33 Given these known limitations of the use of SBS, other methods of health monitoring have been considered and used to increase the sensitivity of detection of these agents.
In recent years, environmental PCR monitoring has been explored as an adjunct method to health monitoring with SBS.10,12-14,17,19,20 This use has been achieved by using several methods, including swabbing of individual colony cages or either horizontal or vertical exhaust rack plenums. The collection method can provide results for specific cages, a row of cages, or the entire rack, depending on the need. Exhaust debris PCR monitoring of IVC racks has recently been evaluated for its efficacy in detecting several pathogens.2,5,8,17,19,22,25,27-29,39 Most commonly, the studies found that detection through PCR analysis of exhaust debris was a reliable option or a good adjunct to a health monitoring program. Notably, PCR evaluation of exhaust debris was found to be reliable and in some cases superior in detecting mouse hepatitis virus, Sendai virus, Helicobacter spp., Rodentibacter pneumotropicus and heylii (previously Pasteurella pneumotropica),3 pinworms, fur mites, and enteric protozoa.2,8,13,17,19,25,28,29,39 Exhaust air dust surveillance (EAD) is the PCR testing mechanism used by Charles River Laboratories to evaluate exhaust debris or environmental samples. EAD PCR evaluation can be used with both swabs and collection media and is the assay used in this study to evaluate all environmental and exhaust samples. Swabs can be taken from IVC racks, cage tops, inner cage, or other environmental sites of interest.
The current study compares the reliability of EAD surveillance to SBS health monitoring over the course of 1 y in 3 barrier rooms in a facility exclusively using IVC racks from a single vendor. Two of the barrier rooms were known to have a prevalence of murine norovirus (MNV), Helicobacter spp., and Rodentibacter spp. MNV is a nonenveloped RNA virus in the family Caliciviridae that shows few to no clinical signs in immunocompetent as well as many immunodeficient mice.31,36 MNV is mainly shed in the feces and can interfere with immune and enteric studies.31,36 Helicobacter spp. and Rodentibacter spp. are bacteria that also may interfere with immune studies.7,35,36,38 Helicobacter spp. are gram-negative, microaerophilic, curved to spiral-shaped organisms commonly found in the gastrointestinal flora of many mammals and, in mice, are mainly transmitted through the fecal–oral route.7,36,38 Infection in mice may cause typhlocolitis, thus confounding enteric studies.7,35,36,38 Rodentibacter spp. is a gram-negative rod, is widespread in the environment and is most effectively transmitted by direct contact.32,36 Typically, clinical signs are not seen with either agent in immunocompetent mice.7,35,36,38 The prevalence of these pathogens in 2 of the rooms provided an opportunity to compare EAD and SBS health monitoring systems for possible complete replacement of SBS in the facilities. We hypothesized that EAD PCR surveillance would be as sensitive or more than the traditional SBS program in detecting the excluded pathogens in the facility.
Materials and Methods
Husbandry and animal care.
Animals were housed in a vivarium within the Animal Resources Center, which is part of the AAALAC-accredited animal care and use program at the University of Chicago. All animal work was approved by the University of Chicago's IACUC. Three housing rooms were monitored: 2 mouse barrier (MB) rooms and one mouse barrier plus (MB+) room. MB rooms excluded the pathogens noted in Figure 1 except for MNV, Helicobacter spp., and Rodentibacter spp. The MB+ room excluded all the pathogens listed in Figure 1 including the 3 pathogens not excluded from MB rooms. The MB+ room served as a negative control for all pathogens tested. Prior to the study, health monitoring in these rooms was achieved through SBS serology and fecal, fur, and oral swab PCR analysis on a quarterly basis.
Figure 1.
List of pathogens excluded from the Mouse Barrier Plus (MB+) room. All pathogens except those denoted by an asterisk are excluded from Mouse Barrier (MB) rooms. Sentinel serology was performed regarding pathogens in green; PCR analysis of sentinels was performed regarding pathogens in purple. The EAD filter was PCR-tested for all pathogens listed.
The facility houses animals exclusively on Jag 75 Micro-VENT Environmental System IVC racks (Allentown Caging, Allentown, NJ). Animals housed on these racks are assigned to individual investigator protocols, and the number of mice on the racks is completely dependent on the research conducted. Investigators in these rooms have approved protocols spanning a variety of research models including neoplasia, immunology, and enteric biology. Room configuration ranged from 4 to 6 single-sided 70-cage racks and 1 or 2 double-sided 140-cage racks per room for a total of 22 (5 double-sided and 17 single-sided) IVC racks across 3 rooms (Figure 2). There were a total of 16 racks in the MB rooms and 6 racks in the MB+ room. All racks were sanitized in a Basil 9500 Cage and Rack Washer (Steris, Mentor, OH) prior to study initiation. To ensure that the appropriate sanitation temperature is achieved (180 °F), the rack washer runs at the start of each day by using a Temp-Tape 180 (Pharmacal Research Laboratories, Naugatuck, CT) temperature-indicating strip. To confirm MB+ pathogen-negative status prior to the study, each rack was swabbed at each horizontal exhaust plenum (Figure 3 C), and samples were submitted to Charles River Laboratories (Wilmington, MA) for PCR testing for the full panel of excluded pathogens (MB+ pathogen panel) before and after sanitation. Sentinel EAD Collection Media (Allentown) was fit into the exhaust plenum of every clean rack and collected and replaced quarterly for PCR testing (Figure 4). Allentown and Charles River Laboratories collaborated to design the Sentinel EAD Collection Media to maximize collection of exhaust debris from the IVC to be evaluated by using the unique EAD PCR testing method developed by Charles River Laboratories. Placement and collection of media was performed by using clean gloves and a sterile vial for collection. Concurrently, 54 Swiss Webster sentinel mice (female; age, 3 to 4 wk; Taconic Biosciences, Rensselaer, NY) were purchased and housed, 2 per cage, with one cage per rack side. These mice were certified as MB+ pathogen-free by the vendor. In addition, on arrival, feces were pooled and submitted for fecal PCR to screen for MB+ pathogen-free status prior to initiation of experiment. Sentinels were identified by using an ear punch (2 mm) in the right ear (model 1538, The Punch, National Band and Tag Company, Newport, KY) or no ear punch. Sentinel cages were placed on the bottom right corner of each IVC rack or rack side.
Figure 2.
(A) Diagram of room arrangement of both MB rooms. Sentinel cages with 2 mice per cage are indicated with green squares. There were a total of 20 sentinel cages between the 2 rooms. There were a total 16 racks between the 2 rooms: 12 single-sided racks and 4 double-sided racks. (B) Diagram of room arrangement of MB+ room. Sentinel cages with 2 mice per cage are indicated with green squares. There were a total of 7 sentinel cages and 6 total racks: 5 single-sided racks and 1 double-sided rack. BSC, biologic safety cabinet.
Figure 3.
(A) Front side of an IVC rack. Green arrows represent HEPA-filtered air coming in to each individual cage. Red arrows represent air exhausted out of each individual cage. (B) Back side of IVC rack, with representative arrows for the flow of air. Supply and exhaust plenum doors are seen on the left and right sides, respectively. (C) Close-up of open exhaust plenum door. Red arrow represents flow of exhaust air. Representative horizontal plenum is circled in red. The horizontal plenums were swabbed prior to initiation of study to confirm MB+-free status. Black arrow shows the vertical exhaust plenum with Sentinel Collection Media placed.
Figure 4.
(A) Sentinel EAD Collection Media. (B) Placement of Sentinel Collection Media into specifically designed holder to fit into vertical exhaust plenum of IVC rack. (C) Close-up of Sentinel Collection Media placed in vertical exhaust plenum, from above. (D) Collection of Sentinel Collection Media into sterile vial by using clean gloves.
All mice were housed in solid-bottom polycarbonate IVC (19.69 × 30.48 × 16.51 cm; Jag 75 Micro-Barrier, Allentown). All cages, bedding, and enrichment were autoclaved prior to use. Mice were housed on 1/4-in. corncob bedding (no. 7097, Envigo, Indianapolis, IN), received reverse osmosis–purified water without restriction through an automatic watering system (Avidity Science, Waterford, WI), and were fed irradiated diet chow (Teklad 2918, Envigo). Mice were provided cotton nesting squares (NES 3600, Ancare, Bellmore, NY) for enrichment. Animal rooms were maintained on a 12:12-h light:dark cycle with humidity ranging from 30% to 70% and temperatures ranging from 68 to 76 °F (20.0 to 24.4 °C), in compliance with the Guide for the Care and Use of Laboratory Animals.16
Animal cages were changed every 2 wk within a class II type A2 biosafety cabinet (NuAire, Plymouth, MN). All cages on the same rack or rack side were changed before the sentinel cage, and approximately 5 g of soiled bedding from the dirtiest area of each cage was collected by using a 5-finger pinch method and added to the new sentinel cage. At cage change, as each cage entered the biosafety cabinet, the outside was sprayed with Labsan C-Dox (Sanitation Strategies, Holt, MI) prior to removal of the lid. Each mouse was placed in a new clean cage by using atraumatic forceps that had been soaked in Labsan C-Dox. The food was transferred to a new wire bar and topped off. Last, soiled bedding was taken from the old cage and placed in the new clean sentinel cage. The clean cage of mice was returned to the rack, where CaviWipes (Metrex Research, Orange, CA) were used to clean any excess dust build-up from the cage slot. Gloves and sleeves were sprayed with Labsan C-Dox between cages, and gloves were changed between racks. When no colony mice were present on the rack, the sentinel cage was changed every 2 wk in the same manner but without the addition of soiled bedding.
Animal numbers.
In total, 135 Swiss Webster mice were ordered as sentinel animals throughout the 1-y study. In the first quarter, 54 mice were ordered for housing at 2 per rack or rack side on all 22 IVC racks. In addition, 27 new mice were ordered each at the start of the second, third, and fourth quarters, to replace the sentinels that had been collected for testing in the previous quarter.
Sample and data collection.
Every 3 mo, for 1 y (4 time points in total), one sentinel mouse from each single-sided rack or double-sided rack side was collected and euthanized through CO2 asphyxiation at a gas displacement rate of 10% to 30% of the chamber volume per minute. In the first quarter, the sentinel with the right ear punch was collected (housed for 3 mo on the rack), and in subsequent quarters, the older of the 2 animals (housed for 6 mo on the rack) was collected for testing. Cardiac blood collection was performed, and blood was collected on an EZ-Spot card (Charles River Laboratories) for serology of pathogens delineated in green in Figure 1. In addition, pelt and oral swabs as well as feces were collected for PCR testing of pathogens indicated in purple in Figure 1. At the same time interval, samples for EAD collection media were collected, by using clean gloves, from each rack for PCR testing of all MB+ excluded pathogens. The manufacturer recommends leaving EAD collection media in place for 3 mo for optimal dust accumulation. All samples were submitted to Charles River Laboratories for testing.
Each quarter, 27 additional female Swiss Webster mice (age, 3 to 4 wk; Taconic Biosciences, Rensselaer, NY) were ordered; feces were pooled and sent for PCR testing to confirm MB+ status before the previous sentinel on each rack or rack side was replaced. Prior to replacement of the previous sentinel, right ear punches were performed as necessary to identify the new compared with the remaining sentinel.
Since the study was performed in a working barrier, colony census was entirely dependent on individual investigators. To understand the soiled bedding sampling size, at least twice per month, a total census of colony cages on each rack was collected, with the location of cage position noted for every rack on study.
Statistical analysis.
The percentage of racks that tested positive for any organism on the list of pathogens in Figure 1 was compared between EAD and SBS by using the McNemar exact test. This analysis was performed for each pathogen overall and separately for each of the 4 time points. For the overall analysis, conclusions were confirmed by using generalized estimating equations regression models with robust standard errors to account for differences in collection media and sentinel testing multiple time points (data not shown). Analyses were performed by using Stata 15 (StataCorp, College Station, TX). An α level less than 0.01 was considered significant for all analyses.
Results
Confirmation of sanitation and MB+ pathogen-free status.
All racks were confirmed to be sanitized, with no positive results for excluded pathogens from swab samples of rack plenums prior to initiation of the experiment.
All new sentinel animals were purchased as MB+ pathogen-free and were confirmed to be MB+ pathogen-free through fecal PCR testing on arrival.
Census.
Room census was entirely dependent on research investigator usage. On average, most racks were at greater than 50% capacity each quarter (Table 1). Although the overall census indicated that racks were consistently greater than 50% full, some racks occasionally had very few colony cages. In the last quarter, 3 single-sided racks in an MB room housed only the sentinel cage for the entire testing period. The MB+ room tended to have greater shifts in use by investigators but still remained, on average and by rack, to be occupied similarly to the MB rooms. However, one rack in the MB+ room housed only the sentinel cage for most of the study period (about 8 consecutive months).
Table 1.
Rack census per room
| Quarter | ||||
| 1 | 2 | 3 | 4 | |
| MB room 1 | 68.08% | 69.52% | 53.88% | 67.33% |
| MB room 2a | 60.52% | 59.71% | 56.79% | 50.76% |
| MB+ room | 60.59% | 49.46% | 49.29% | 52.07% |
Data are given as the average percentage occupancy per rack.
MB room that housed 3 single-sided racks with sentinel cages only throughout quarter 4.
Health monitoring.
All agents excluded in MB rooms tested negative by both SBS and EAD testing for all racks in all rooms for the entirety of the study. In addition, MNV, Helicobacter spp. and Rodentibacter spp. tested negative in the MB+ room by both SBS and EAD testing throughout the entire year (data not shown).
In both MB rooms, EAD testing was positive for Helicobacter spp. on every rack during all 4 quarters and was significantly more sensitive (P < 0.001) than SBS in detecting Helicobacter spp. (Figure 5). When SBS positives for Helicobacter spp. were further characterized by species, the corresponding EAD species characterization matched the SBS species characterized and identified at least one additional species. H. mastomyrinus was the only species identified by SBS, whereas EAD identified H. mastomyrinus, H. ganmani, H. hepaticus, and H. typhlonius.
Figure 5.
Proportion of IVC racks in MB rooms that tested positive for Helicobacter spp. by EAD or SBS each quarter. EAD identified Helicobacter spp. on every rack during all 4 quarters. EAD was significantly (†, P < 0.01) more sensitive in detection of Helicobacter spp. than SBS.
In both MB rooms, EAD for Rodentibacter spp. was significantly (P < 0.001) more sensitive than SBS (Figure 6). Overall, EAD detected Rodentibacter spp. in 94% of the racks, whereas SBS detected these organisms in 11% of the racks. Any SBS that tested positive had a corresponding positive result on that rack by EAD PCR analysis. Both EAD and SBS detected R. pneumotropicus and R. heylii.
Figure 6.
Proportion of IVC racks in MB rooms that tested positive for Rodentibacter spp. by EAD or SBS each quarter. EAD was significantly (†, P < 0.01) more sensitive in detection of Rodentibacter spp. in each quarter than SBS.
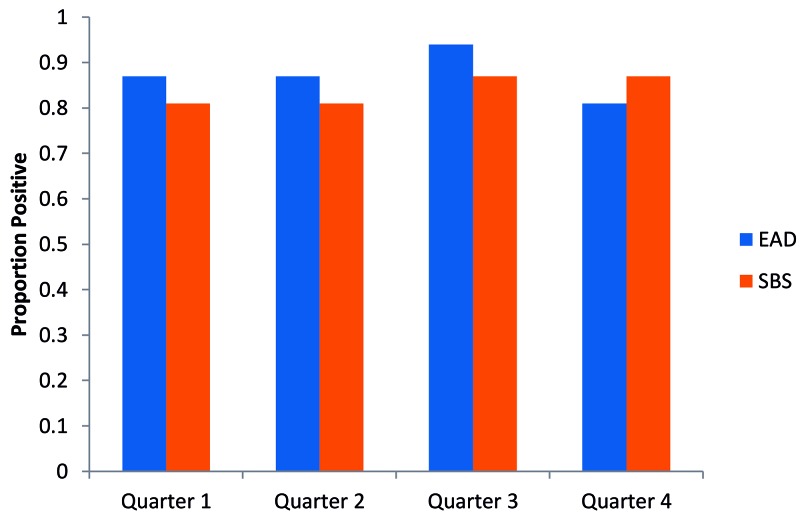
In both MB rooms, MNV detection did not differ significantly between SBS and EAD (Figure 7). All SBS positive results for MNV corresponded with EAD positive results on the same rack, except in one case (Table 2). In quarter 4, SBS tested positive for MNV but EAD tested negative for the same rack. This rack housed only the sentinel cage for the entire 3 mo testing period. There were 3 cases for which EAD tested positive for MNV, but the correlating SBS for that rack tested negative (Table 2).
Figure 7.
Proportion of IVC racks in MB rooms that tested positive for MNV by EAD or SBS each quarter. There was no significant difference (overall P > 0.5) in detection of MNV on each rack between EAD and SBS. Each EAD-positive result from a rack correlated with an SBS-positive result from the same rack.
Table 2.
Detection of MNV
| Quarter 1 | Quarter 2 | Quarter 3 | Quarter 4 | |||||
| EAD | SBS | EAD | SBS | EAD | SBS | EAD | SBS | |
| Rack 1 | + | + | + | + | + | + | + | + |
| Rack 2 | + | + | + | + | + | + | + | + |
| Rack 3 | + | + | + | + | + | + | + | + |
| Rack 4 | + | + | + | + | + | + | + | + |
| Rack 5 | + | + | + | + | + | + | + | + |
| Rack 6 | + | + | + | + | + | + | + | + |
| Rack 7 | + | + | + | + | + | + | + | + |
| Rack 8 | + | + | + | + | + | + | + | + |
| Rack 9a | + | + | + | + | + | + | —c | + |
| Rack 10a | — | — | — | — | + | —b | — | — |
| Rack 11a | — | — | — | — | — | — | — | — |
| Rack 12 | + | + | + | + | + | + | + | + |
| Rack 13 | + | + | + | + | + | + | + | + |
| Rack 14 | + | + | + | + | + | + | + | + |
| Rack 15 | + | + | + | + | + | + | + | + |
| Rack 16 | + | —b | + | —b | + | + | + | + |
+, positive; —, negative
All racks that tested positive for MNV through SBS also tested positive by EAD, except for rack 9 in quarter 4.
Rack that housed only sentinel cages throughout quarter 4.
SBS gave a negative result but EAD tested positive.
EAD gave a negative result but SBS tested positive.
Discussion
In 3 mouse rooms of an established research barrier facility, we compared EAD PCR surveillance of IVC racks with SBS health monitoring over the course of 1 y. Each quarter, 22 racks (5 double-sided and 17 single-sided) were tested for a full panel of MB+ excluded agents through EAD PCR analysis. Concurrently, 27 corresponding SBS were tested each quarter through serology and PCR analysis for MB+ pathogens.
Our hypothesis was confirmed in that EAD testing is more sensitive than SBS in the detection of Helicobacter spp. and Rodentibacter spp. SBS and EAD testing yielded equivalent detection of MNV.
Helicobacter spp. was detected in all 16 MB racks every quarter by EAD but was only detected through SBS on 5 racks in the first quarter and 1 rack in the third quarter. Both Helicobacter spp. and Rodentibacter spp. were tested through PCR assays on either the EAD collection media or swabs or feces from the SBS mice. The collection media accumulates dust particles over the course of 3 mo and has the potential to collect a much higher concentration of each microbe for detection, whereas the sentinel grooms itself and may not shed the microbe at the time of collection, decreasing the likelihood of detection from the SBS samples. Transmission of these bacteria to the sentinels through soiled bedding historically has been poor.8,28,33 Rodentibacter spp., specifically, is described to be viable for only 30 min in bedding, and this characteristic likely plays a role in the organism's poor transmission to SBS.32
In addition, the speciation results for Helicobacter spp. for EAD returned species that have all been described to have been isolated from mice,18,34,36 whereas SBS identified only one of these species, H. mastomyrinus. Other studies have isolated H. mastomyrinus from the liver and cecum of mice34,35 and noted colonization in the liver with potential to cause significant disease in immunocompromised mice.11 Perhaps this species is more often shed once it has colonized an animal and therefore SBS identifies H. mastomyrinus more often than other species. Other Helicobacter spp. may shed more intermittently than H. mastomyrinus and therefore may be more difficult to detect through SBS and more likely to be identified from EAD collection media, which accumulates exhaust dust over the course of 3 mo.
In this study, Helicobacter spp. were evaluated by PCR analysis of SBS feces. Previous studies have shown that PCR analysis of cecal scrapings produces higher detection rates for Helicobacter spp. than fecal PCR.37 This result corroborates the data in our current study, in that, the detection rates after using fecal PCR analysis ranged from 0% to 31% per quarter, compared with the 100% detection rates obtained by using EAD collection media. These results were not compared with cecal scraping PCR analysis, but the evidence suggests that cecal scrapings may be equivalent to EAD testing in the detection of Helicobacter spp.. However, EAD testing would not require the euthanasia of animals, as for cecal scraping evaluation. The described findings for Helicobacter spp. corroborate recent studies28,29 and indicate that EAD is extremely sensitive for the detection of Helicobacter spp. and Rodentibacter spp., even when each rack (on average) was only half occupied.
In a recent study, MNV detection through exhaust air debris was found to be more sensitive than SBS.39 However, our data showed no significant difference in the detection of MNV between EAD and SBS. Indeed, in the fourth quarter testing, one rack in an MB room that tested positive for MNV by SBS serology concurrently tested negative by EAD. This rack housed only one cage of sentinel mice for the entirety of the quarter. Typically a sentinel cage would not remain on an otherwise empty rack, but for the purpose of the study, the cage was left on the rack. Although some investigators have described persistent shedding of MNV,2,21,26 others state that the virus may be shed intermittently.39 The sentinel that we tested for MNV was the older of the 2 sentinels housed on the empty rack for the quarter. When this sentinel was first placed on the rack, in quarter 3, this rack was at 92% capacity (on average) until the last month. This older sentinel may have been infected with MNV and might have seroconverted in the previous quarter, but if shedding was intermittent, in the following quarter, the mouse might not have been shedding the pathogen such that the EAD collection media could pick it up. In addition, the study describing MNV detection as more sensitive through exhaust debris than SBS used ICR mice, whereas we used Swiss Webster mice in the current study. The difference in strains may play a role in the ability of the animals to seroconvert to the virus.4,15,21 The extremely low prevalence on this rack (1.4%) may play a role in detection of MNV through EAD. Low detection rates with low prevalence was seen previously with MNV and mouse parvovirus detection in a study using different IVC systems (Tecniplast and Innovive) than the one we used here (Allentown).2 The low prevalence of MNV in our study likely resulted in minimal generation of contaminated dust, which therefore was insufficient for the collection media to capture.
Although in our current study, the detection of MNV did not differ significantly between EAD and SBS, the results seemed to corroborate each other and support the high prevalence of MNV in MB rooms in our facility. In contrast, MNV was never detected through exhaust air debris when animals were inoculated with the virus and were shedding for the entirety of the experiment.2 In the cited study, the exhaust air debris was collected by using a different type of collection media, placed over a small area on the prefilter, and was collected in small sections as the study progressed. For our study, the collection media was a specifically designed filter to fit at the exhaust plenum, and the entire filter was submitted for testing each quarter. It appears that the collection media, its placement, and its sampling are vital factors in the accumulation of exhaust air dust and subsequent identification of pathogens. The detection of pathogens appears to be more reliable when using a product specifically designed to capture and accumulate exhaust air dust, but differences should be explored further.
Rodents in biomedical research have transitioned to progressively more isolated housing environments from open-top caging to microisolation cages to IVC racks. The increase in isolated environments is making the transmission of previously poorly transmitted pathogens increasingly difficult.2,5,8,25 Some agents, such as many of the rodent enteric viruses, including MNV, can be transmitted in the bedding reliably with adequate dose.33 However, agents that are transmitted poorly through bedding include respiratory pathogens such as Sendai virus and F. rodentium,1,8 fur mites23 and unstable pathogens such as enveloped viruses and Rodentibacter spp.,2,32 are increasingly less likely to be transmitted reliably through bedding. Without the use of open-top caging, respiratory pathogen transmission is rare, and with the increased barrier practices in vivariums housing in microisolation caging and IVC, unstable pathogens have decreased likelihood of being transmitted in bedding. The growing body of literature for evaluation of exhaust dust PCR testing indicates that some of these pathogens that historically have been more difficult to detect can now be identified more readily.2,8,13,17,19,25,28,29,39
Some authors have noted drawbacks to exhaust debris PCR analysis because of crossreactivity to the PCR assay performed and subsequent false positives.22 In addition, others have noted less than 100% sensitivity for the detection of agents such as mouse pinworms, Aspicularis tetraptera, and Syphacia obvelata.13 In addition, due to the high sensitivity of exhaust debris PCR surveillance, the potential for detecting contaminant trace DNA material or inactive agents present in the environment has been noted.27,28 These described drawbacks lead to increased false-positive results, which require costly follow-up tests. In our experience, since implementing EAD health surveillance throughout all facilities at our institution, a total of 4 false positives have been detected over the course of approximately 2 y. The following descriptions are not part of the data collected for the comparison study but do provide some additional insight into the implementation of EAD as a sole method of surveillance. Specifically, of the 4 false positives, one was a reporting error. Each of the other 3 false-positive results was confirmed through follow-up tests. Typically, swabs were taken from the horizontal plenums of the IVC rack, the colony cages, or colony animals. Cage swabs are taken from around the inside walls of the cage, whereas the location for swabbing the colony animals depends on the agent in question. Throughout the course of this study and implementation of EAD in other facilities, we have found that supplemental testing and consultation with the testing laboratory has proven to be an efficient and reliable method for follow-up and health surveillance at our institution.
Beyond the scope of this study and during the implementation of EAD throughout the rest of the campus facilities, we identified 2 pathogens that we would not have otherwise detected by using SBS. EAD surveillance detected a true outbreak of R. affinis in a colony that had never previously tested positive with SBS. With confirmatory swabs testing positive, the colony was quarantined and treated. Follow-up testing with colony swabs showed that the fur mite had been cleared, and the colony was released from quarantine. Detection of R. affinis has been described by using PCR of horizontal plenum swabs17 and filter tops12 where prevalence was known. To reduce future detection of residual DNA, the rack was sanitized, and the EAD collection media was replaced. Racks in the facilities are otherwise sent through the rack washer on an as-needed basis and sanitized, as previously described in the Methods section. Rack sanitation is coordinated when the EAD collection media is collected, so as to ensure adequate accumulation of exhaust dust.
The second unexpected pathogen identified when EAD was implemented in other campus facilities using Allentown IVC racks was lactate dehydrogenase-elevating virus (LDV). LDV is a mouse-specific virus in the Arteriviridae family and is typically rare in laboratory colonies.36 LDV has been found to be a common contaminant of biologic material and can create variables for research.6,24,36 A positive result of a single copy number on EAD collection media was confirmed to be a true positive in consultation with the diagnostic laboratory. The combination of poor transmission, instability in the environment, and lack of clinical signs in the animals make LDV very difficult to detect and unlikely to be transmitted in the soiled bedding. Consequently, SBS on this rack never tested positive for LDV. We were able to contain the virus to the rack and cull all mice that had been inoculated with LDV-contaminated tumor to eradicate the virus. Due to its poor transmission and viability in the environment, as well as the possibility of tumor lines being potentially contaminated, testing for LDV in the facilities has increased to quarterly, and all positives will be considered true until a complete investigation proves otherwise.
Although previous research has evaluated the quality of exhaust dust PCR surveillance,2,5,8,12,17,19,25,27-29,30,39 to our knowledge, our current study is the first to compare immunocompetent SBS and EAD colony surveillance by using sentinel collection media on Allentown IVC racks in a working biomedical research barrier. Other investigators showed similar results from a single 3-mo evaluation using the same collection media and IVC racks; however, sentinel mice were immunocompromised, and MNV status was not evaluated.30 In the current study, MNV—as most of the other viruses— was evaluated through serology; therefore immunocompetent sentinels with the ability to seroconvert were necessary for this screening method. Another group describes similar results to this current report; however, the previous results were obtained by using Tecniplast IVC racks and analyzing gauze pieces that had been attached to the exhaust air prefilter.27,28 Still others describe similar findings to this report regarding bacterial agents; however, SBS were not used in the single 3-mo evaluation period, thereby precluding comparison of surveillance methods.25 Overall, these other studies2,5,8,12,17,19,25,27-29,30,39 evaluate exhaust dust PCR surveillance by using other IVC systems, collection media or mechanisms or collection periods and do not typically directly compare with SBS. Our study compares SBS to EAD surveillance in a practical context by using existing IVC rack systems, barrier prevalences, and manufacturer-recommended collection times for optimal exhaust dust accumulation. Each vivarium is unique, with differing housing systems, exclusion lists, and surveillance programs. The current study provides additional data on exhaust dust surveillance methods, which are instrumental for institutions to evaluate their surveillance programs.
In comparing the 2 health monitoring systems in a mouse barrier supporting daily research with colony mice housed on Allentown IVC racks, it was shown that EAD pathogen surveillance can be implemented to replace SBS health monitoring. This finding has major implications for the 3R principles of animal research. EAD surveillance is not only a refinement that is more sensitive in detecting several pathogens, but it also eliminates the stresses associated with mouse shipment, handling, and sample collection. Using EAD monitoring replaces live sentinel animals in the facilities and greatly reduces the number of animals that otherwise would have been used previously for health monitoring. Since our implementation of EAD monitoring, approximately 1,700 sentinel animals are no longer used in our facilities each year. In addition, the replacement of sentinel animals with EAD monitoring reduces costs 3-fold. The upfront cost of the mice and the cost of maintaining the animals, as well as the time staff spend on transferring soiled bedding and inhouse sample collection and diagnostics are all savings that are gained when implementing EAD surveillance. It is important to evaluate the type of IVC rack system used at each institution and its airflow prior to implementing an exhaust dust PCR surveillance program. Each system is different and therefore optimal locations for diagnostic sampling will differ also. Our study is the first to evaluate EAD testing of sentinel collection media on Allentown IVC racks in a working research barrier using immunocompetent SBS. Thus the findings in this study are specific to the sentinel collection media and the Allentown IVC racks. With the overall sensitivity of EAD proving superior for detection of the bacteria described, the equivalent detection of MNV, the implications for cost savings, and the reduction of animal use in research, EAD surveillance—where possible—is a recommended alternative to SBS in facilities.
Acknowledgments
We thank The University of Chicago Laboratory Animal Medicine Training Program for funding this project. We also thank Ann Schue, Sue Zanelli, and Karin Peterson for assistance with sample collection and Derek Summerville and Curtis Jackson for support with IVC rack and room set up.
This work was supported by The University of Chicago Animal Resources Center. The researchers claim no conflicts of interest.
References
- 1.Artwohl JE, Cera LM, Wright MF, Medina LV, Kim LJ. 1994. The efficacy of a dirty bedding sentinel system for detecting Sendai virus infection in mice: a comparison of clinical signs and seroconversion. Lab Anim Sci 44:73–75. [PubMed] [Google Scholar]
- 2.Bauer BA, Besch-Williford C, Livingston RS, Crim MJ, Riley LK, Myles MH. 2016. Influence of rack design and disease prevalence on detection of rodent pathogens in exhaust debris samples from individually ventilated caging systems. J Am Assoc Lab Anim Sci 55:782–788. [PMC free article] [PubMed] [Google Scholar]
- 3.Benga L, Sager M, Christensen H. 2018. From the [Pasteurella] pneumotropica complex to Rodentibacter spp.: an update on [Pasteurella] pneumotropica. Vet Microbiol 217:121–134. 10.1016/j.vetmic.2018.03.011. [DOI] [PubMed] [Google Scholar]
- 4.Besselsen DG, Wagner AM, Loganbill JK. 2000. Effect of mouse strain and age on detection of mouse parvovirus 1 by use of serologic testing and polymerase chain reaction analysis. Comp Med 50:498–502. [PubMed] [Google Scholar]
- 5.Brielmeier M, Mahabir E, Needham JR, Lengger C, Wilhelm P, Schmidt J. 2006. Microbiological monitoring of laboratory mice and biocontainment in individually ventilated cages: a field study. Lab Anim 40:247–260. 10.1258/002367706777611497. [DOI] [PubMed] [Google Scholar]
- 6.Carlson Scholz JA, Garg R, Compton SR, Allore HG, Zeiss CJ, Uchio EM. 2011. Poliomyelitis in MuLV-infected ICR-SCID Mice after injection of basement membrane matrix contaminated with lactate dehydrogenase-elevating virus. Comp Med 61:404–411. [PMC free article] [PubMed] [Google Scholar]
- 7.Chichlowski M, Hale LP. 2009. Effects of helicobacter infection on research: the case for eradication of helicobacter from rodent research colonies. Comp Med 59:10–17. [PMC free article] [PubMed] [Google Scholar]
- 8.Compton SR, Homberger FR, Paturzo FX, Clark JM. 2004. Efficacy of 3 microbiological monitoring methods in a ventilated cage rack. Comp Med 54:382–392. [PubMed] [Google Scholar]
- 9.de Bruin WC, van de Ven EM, Hooijmans CR. 2016. Efficacy of soiled bedding transfer for transmission of mouse and rat infections to sentinels: a systematic review. PLoS One 11:e0158410 10.1371/journal.pone.0158410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Dole VS, Zaias J, Kyricopoulos-Cleasby DM, Banu LA, Waterman LL, Sanders K, Henderson KS. 2011. Comparison of traditional and PCR methods during screening for and confirmation of Aspiculuris tetraptera in a mouse facility. J Am Assoc Lab Anim Sci 50:904–909. [PMC free article] [PubMed] [Google Scholar]
- 11.Eaton KA, Opp JS, Gray BM, Bergin IL, Young VB. 2010. Ulcerative typhlocolitis associated with helicobacter mastomyrinus in telomerase-deficient mice. Vet Pathol 48:713–725. 10.1177/0300985810383876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gerwin PM, Ricart Arbona RJ, Riedel ER, Henderson KS, Lipman NS. 2017. PCR testing of IVC filter tops as a method for detecting murine pinworms and fur mites. J Am Assoc Lab Anim Sci 56:752–761. [PMC free article] [PubMed] [Google Scholar]
- 13.Gerwin PM, Ricart Arbona RJ, Riedel ER, Lepherd ML, Henderson KS, Lipman NS. 2017. Evaluation of traditional and contemporary methods for detecting Syphacia obvelata and Aspiculuris tetraptera in laboratory mice. J Am Assoc Lab Anim Sci 56:32–41. [PMC free article] [PubMed] [Google Scholar]
- 14.Henderson KS, Perkins CL, Havens RB, Kelly MJ, Francis BC, Dole VS, Shek WR. 2013. Efficacy of direct detection of pathogens in naturally infected mice by using a high-density PCR array. J Am Assoc Lab Anim Sci 52:763–772. [PMC free article] [PubMed] [Google Scholar]
- 15.Henderson KS, Pritchett-Corning KR, Perkins CL, Banu LA, Jennings SM, Francis BC, Shek WR. 2015. A comparison of mouse parvovirus 1 infection in BALB/c and C57BL/6 mice: susceptibility, replication, shedding, and seroconversion. Comp Med 65:5–14. [PMC free article] [PubMed] [Google Scholar]
- 16.Institute for Laboratory Animal Research. 2011. Guide for the care and use of laboratory animals, 8th ed Washington (DC): National Academies Press. [Google Scholar]
- 17.Jensen ES, Allen KP, Henderson KS, Szabo A, Thulin JD. 2013. PCR testing of a ventilated caging system to detect murine fur mites. J Am Assoc Lab Anim Sci 52:28–33. [PMC free article] [PubMed] [Google Scholar]
- 18.Johansson SK, Feinstein RE, Johansson KE, Lindberg AV. 2006. Occurrence of Helicobacter species other than H. hepaticus in laboratory mice and rats in Sweden. Comp Med 56:110–113. [PubMed] [Google Scholar]
- 19.Kapoor P, Hayes YO, Jarrell LT, Bellinger DA, Thomas RD, Lawson GW, Arkema JD, Fletcher CA, Nielsen JN. 2017. Evaluation of anthelmintic resistance and exhaust air dust PCR as a diagnostic tool in mice enzootically infected with Aspiculuris tetraptera. J Am Assoc Lab Anim Sci 56:273–289. [PMC free article] [PubMed] [Google Scholar]
- 20.Karlsson EM, Pearson LM, Kuzma KM, Burkholder TH. 2014. Combined evaluation of commonly used techniques, including PCR, for diagnosis of mouse fur mites. J Am Assoc Lab Anim Sci 53:69–73. [PMC free article] [PubMed] [Google Scholar]
- 21.Kelmenson JA, Pomerleau DP, Griffey S, Zhang W, Karolak MJ, Fahey JR. 2009. Kinetics of transmission, infectivity, and genome stability of 2 novel mouse norovirus isolates in breeding mice. Comp Med 59:27–36. [PMC free article] [PubMed] [Google Scholar]
- 22.Leblanc M, Berry K, Graciano S, Becker B, Reuter JD. 2014. False-positive results after environmental pinworm PCR testing due to rhabditid nematodes in corncob bedding. J Am Assoc Lab Anim Sci 53:717–724. [PMC free article] [PubMed] [Google Scholar]
- 23.Lindstrom KE, Carbone LG, Kellar DE, Mayorga MS, Wilkerson JD. 2011. Soiled bedding sentinels for the detection of fur mites in mice. J Am Assoc Lab Anim Sci 50:54–60. [PMC free article] [PubMed] [Google Scholar]
- 24.Liu H, Bockhorn J, Dalton R, Chang Y-F, Qian D, Zitzow LA, Clarke MF, Greene GL. 2011. Removal of lactate dehydrogenase-elevating virus from human-in-mouse breast tumor xenografts by cell-sorting. J Virol Methods 173:266–270. 10.1016/j.jviromet.2011.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Mahabir E, Durand S, Henderson KS, Hardy P. 2019. Comparison of two prevalent individually ventilated caging systems for detection of murine infectious agents via exhaust air particles. Lab Anim 53:84–88. 10.1177/0023677218785929. [DOI] [PubMed] [Google Scholar]
- 26.Manuel CA, Hsu CC, Riley LK, Livingston RS. 2008. Soiled-bedding sentinel detection of murine norovirus 4. J Am Assoc Lab Anim Sci 47:31–36. [PMC free article] [PubMed] [Google Scholar]
- 27.Miller M, Brielmeier M. 2017. Environmental samples make soiled bedding sentinels dispensable for hygienic monitoring of IVC-reared mouse colonies. Lab Anim 52:233–239. 10.1177/0023677217739329. [DOI] [PubMed] [Google Scholar]
- 28.Miller M, Ritter B, Zorn J, Brielmeier M. 2016. Exhaust air dust monitoring is superior to soiled bedding sentinels for the detection of Pasteurella pneumotropica in individually ventilated cage systems. J Am Assoc Lab Anim Sci 55:775–781. [PMC free article] [PubMed] [Google Scholar]
- 29.Miller M, Ritter B, Zorn J, Brielmeier M. 2016. Exhaust air particle PCR detects Helicobacter hepaticus infections at low prevalence. J Vet Sci Technol 7:1–2. [Google Scholar]
- 30.Niimi K, Maruyama S, Sako N, Miyata K, Yoshimoto T, Bilecki BM, Henderson KS, Takahashi E. 2018. The SentinelTM EADR program can detect more microorganisms than bedding sentine animals. Jpn J Vet Res 66:125–129. 10.14943/jjvr.66.2.125. [DOI] [Google Scholar]
- 31.Perdue KA, Green KY, Copeland M, Barron E, Mandel M, Faucette LJ, Williams EM, Sosnovtsev SV, Elkins WR, Ward JM. 2007. Naturally occurring murine norovirus infection in a large research institution. J Am Assoc Lab Anim Sci 46:39–45. [PubMed] [Google Scholar]
- 32.Scharmann W, Heller A. 2001. Survival and transmissibility of Pasteurella pneumotropica. Lab Anim 35:163–166. 10.1258/0023677011911543. [DOI] [PubMed] [Google Scholar]
- 33.Shek WR, Smith AL, Pritchett-Corning KR. 2015. Microbiological quality control for laboratory rodents and lagomorphs, p 463–510. Chapter 11. In: Fox JG, Anderson LC, Otto GM, Pritchett-Corning KR, Whary MT, Laboratory animal medicine, 3rd ed Boston (MA): Academic Press. [Google Scholar]
- 34.Shen Z, Xu S, Dewhirst FE, Paster BJ, Pena JA, Modlin IM, Kidd M, Fox JG. 2005. A novel enterohepatic Helicobacter species ‘Helicobacter mastomyrinus’ isolated from the liver and intestine of rodents. Helicobacter 10:59–70. 10.1111/j.1523-5378.2005.00292.x. [DOI] [PubMed] [Google Scholar]
- 35.Taylor NS, Xu S, Nambiar P, Dewhirst FE, Fox JG. 2007. Enterohepatic Helicobacter species are prevalent in mice from commercial and academic institutions in Asia, Europe, and North America. J Clin Microbiol 45:2166–2172. 10.1128/JCM.00137-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Whary MT, Baumgarth N, Fox JG, Barthold SW. 2015. Biology and diseases of mice, p 43–149. Chapter 3. In: Fox JG, Anderson LC, Otto GM, Pritchett-Corning KR, Whary MT. Laboratory animal medicine, 3rd ed Boston (MA): Academic Press. [Google Scholar]
- 37.Whary MT, Cline JH, King AE, Hewes KM, Chojnacky D, Salvarrey A, Fox JG. 2000. Monitoring sentinel mice for Helicobacter hepaticus, H. rodentium, and H. bilis infection by use of polymerase chain reaction analysis and serologic testing. Comp Med 50:436–443. [PubMed] [Google Scholar]
- 38.Whary MT, Fox JG. 2004. Natural and experimental Helicobacter infections. Comp Med 54:128–158. [PubMed] [Google Scholar]
- 39.Zorn J, Ritter B, Miller M, Kraus M, Northrup E, Brielmeier M. 2016. Murine norovirus detection in the exhaust air of IVCs is more sensitive than serological analysis of soiled bedding sentinels. Lab Anim 51:301–310. 10.1177/0023677216661586. [DOI] [PubMed] [Google Scholar]