Abstract
Chemotropism is an essential response of organisms to external chemical gradients that direct the growth of cells toward the gradient source. Chemotropic responses between single cells have been studied using in vitro gradients of synthetically derived signaling molecules and helped to develop a better understanding of chemotropism in multiple organisms. However, dynamic changes including spatial changes to the gradient as well as fluctuations in levels of cell generated signaling molecules can result in the redirection of chemotropic responses, which can be difficult to model with synthetic peptides and single cells. An experimental system that brings together populations of cells to monitor the population-scale chemotropic responses yet retain single cell spatiotemporal resolution would be useful to further inform on models of chemotropism. Here, we describe a microfluidic platform that can measure the chemotropic response between populations of mating yeast A- and α-cells with spatiotemporal programmability and sensitivity by positioning cell populations side by side in calcium alginate hydrogels along semipermeable membranes with micrometer spatial control. The mating phenotypes of the yeast populations were clearly observed over hours. Three distinct responses were observed depending on the distance between the A- and α-cell populations: the cells either continued to divide, arrest, and develop a stereotypical polarized projection termed a “shmoo” toward the cells of opposite mating type or formed shmoos in random directions. The results from our studies of yeast mating suggest that the biofabricated microfluidic platform can be adopted to study population-scale, spatial-sensitive cell–cell signaling behaviors that would be challenging using conventional approaches.
I. INTRODUCTION
Chemotropism is an essential response of organisms to external chemical gradients that direct the growth of cells toward the gradient source.1 This has been observed in the development of the nervous system in which neurons grow toward a target cell,2 or the growth of a pollen tube toward ovules,3 and the mating of fungi.4 The ability to study how cells respond to gradients of chemical molecules is essential in elucidating signaling mechanisms that are vital for life of all forms. The pheromone-mediated mating response in the budding yeast S. cerevisiae has served as a highly informative model of chemotropism.5,6 Peptides are secreted by yeast of two different mating types and result in polarized growth directed toward a nearby mating partner. However, the gradients are very shallow, and this poses a serious challenge when considering the small size of yeast cells and the clustering of pheromone receptors at the tip of the mating projection (i.e., a shmoo tip).6 Therefore, the length scale for sampling reduces to hundreds of nanometers. Previous studies have relied on observing the mating responses in standard agar-based media using a micro-pipet to supply synthetic pheromone to a specific area and monitoring the formation of mating projections.7 To inform the dynamics and length scale of chemotropic response, new platforms that can generate and manipulate stable gradient as well as positioning cells with spatial resolution are highly desired.8
Microfluidic platforms have proven to be highly effective to precisely manipulate populations of cells and measure complex activities driven by chemical gradients. A variety of microfluidic devices has been developed that can generate in vitro chemical gradients to study diffusion of signaling molecules with a range of different length and time scales9–11 or can perform quantitative analyses of pollen tube guidance.12,13 Microfluidic devices have been used to study the mating response in yeast and provided new insights.14–17 For example, Brett et al. managed to produce and rapidly rotate stable pheromone gradients in a microfluidic device allowing the observation of initial orientation and re-orientation in yeast chemotropism.18 Recently, Jin et al. have developed a programmable static droplet array to analyze cell–cell communication, enabling the distinguishing of gene functions in yeast mutants defective for different aspects of pheromone signaling.19 These studies have largely relied on using synthetically derived pheromones and, therefore, may not be reporting on additional biological parameters that can influence gradient formation. This includes the possibility of variations in pheromone concentration over time. Dynamic changes including spatial changes to the gradient can result in the redirection of shmoos, and this can be difficult to model with synthetic peptides. Furthermore, natural habitats are likely to be heterogeneous containing different species that may influence the chemotropic responses or the chemical gradient. An experimental system in which cells produce the chemical gradient and is flexible enough to bring in other cohabitating species can offer the possibility of constructing chemotropic models that include factors that previous studies with synthetic chemical gradients lack. To date, few studies have been reported that attempt to reproduce the gradients formed between populations of yeast of opposite mating types to mimic what might occur in natural habitats.19 One barrier in performing such experiments is the ability to position populations vs individual yeast cells within the proximity of each other and still support gradient formation.19
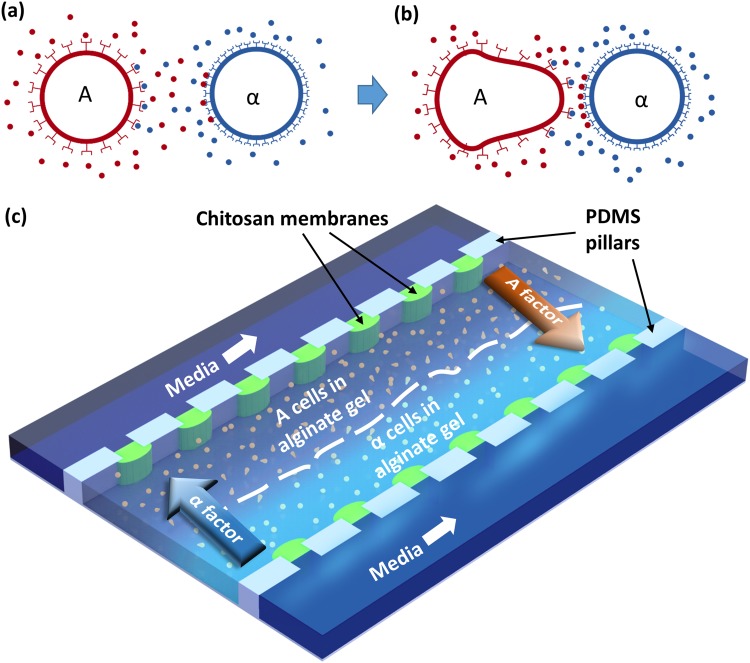
Here, we report on the development of a biofabrication design within a microfluidic device that is capable of precise deposition of populations of mating yeast (A and α) cells [Schemes 1(a) and 1(b)] with spatiotemporal programmability and sensitivity. The observed cell–cell signaling can inform on improving mathematical models of gradient responses. As shown in Scheme 1(c), alginate hydrogels were biofabricated to capture populations of cells side by side in the middle of a three-microchannel device, while a medium was introduced into the two-side microchannels and diffused through the porous chitosan membranes into the middle channel to culture the cells. We have previously reported the diffusion of signaling molecules through alginate hydrogels and nutrients through the chitosan membranes that sustained proper cellular growth and signaling responses.17,20 The unique configuration employed in this study provides several advantages. First, the semi-permeable and freestanding chitosan membranes protected cells from potential shear forces and prevented the microchannels from drying out through gas-permeable polydimethylsiloxane (PDMS) but allowed for the nutrients to diffuse through and the cell morphological changes to occur. Second, the alginate hydrogel matrix was porous enough that pheromone could diffuse in and out of the hydrogel. Finally, this setup allowed for positioning of cell populations to form gradients of pheromones generated by cells rather than requiring synthetic pheromones. In this uniform environment, cells of opposite mating types were fixed in the three-dimensional calcium alginate hydrogel. Cells and the mating response were imaged with live-cell microscopy to monitor morphological changes to gain an insight into the mating response between populations of cells. These results demonstrate that our method can report on chemotropic responses and gradients of naturally generated peptides or chemicals that mediate chemotropism. We envision that this platform can be used to study other forms of population-scale, spatial-sensitive cell–cell signaling behaviors that would be challenging using conventional approaches.
Scheme 1.
Mating yeast chemotropism with biofabrication in microfluidics. (a) Signaling between mating yeast A- and α-cells; (b) chemotropic growth of A-cell toward α-cell resulting in so-called “shmoo” structures, in which A-cells were deleted for BAR1, a gene that encodes for a protease that degrades α-factor to increase their sensitivity to α-cells; and (c) A- and α-cells trapped side by side in alginate hydrogels in a middle microchannel. Biofabricated semi-permeable chitosan membranes separate the middle microchannel from two-side microchannels where media solution is introduced to culture the cells.
II. MATERIALS AND METHODS
A. Materials
Chitosan flakes (85% deacetylated, medium molecular weight), alginate power (extracted from brown algae, medium viscosity), calcium chloride powder, and phosphate buffered saline (PBS) tablets were purchased from Sigma Aldrich. PTFE tubing of 0.022 in. ID and 0.042 in. OD was purchased from Cole-Parmer. Disposable syringes of 1 ml volume were purchased from Becton Dickinson and Company. NE-1000 syringe pumps were purchased from New Era Pump Systems, Inc., and Sylgard 184 and its curing agent were purchased from Ellsworth Adhesives (WI). All yeast reagents were purchased from U.S. Biologicals (TX). The yeast extract peptone dextrose (YPD) solution consists of 1× PBS and 1× YPD (1% yeast extract, 2% peptone, and 1.8 mM tryptophan). The yeast strain is W303 background with the genotype (Mat A bar1 leu2 uras3 his3 trp1).21
PBS solution of pH 7.4 was made with PBS tablets in de-ionized (DI) water. Chitosan solution of 0.5% w/v, pH 5.8 was prepared by dissolving chitosan flakes in DI water, with 1M HCl added dropwise to pH 2 and left by stirring on a stirring plate overnight, followed by dropwise addition of 1M NaOH to adjust the pH to 5.8. The solution was filtered, and DI water was added to bring to the final concentration. Alginate solution of 0.5% w/v was prepared by dissolving sodium alginate powder in DI water, followed by stirring on a stirring plate overnight. DI water was added to the alginate solution to the final concentration. Both solutions were kept in fridge for future use. Before the membrane biofabrication, a small amount of the alginate solution was adjusted to pH 11.5 by adding 1M NaOH solution dropwise. In case that the pH 11.5 alginate solution decreased within days due to the dissolution of carbon dioxide in the air, the pH of the solution was re-adjusted and was only adjusted for a few times before the solution contains too much salt and was disposed, and fresh alginate solution was used.
Yeast were grown in 1× YPD to 2–3 OD600 overnight before pelleting and re-suspended in the 0.5% w/v alginate solution for assembly in microfluidic channels.
B. Device fabrication
The mold for 50-μm-deep microchannels was microfabricated using conventional photolithographic techniques with negative photoresist SU-8 on a 4 in. silicon wafer. Briefly, SU-8 3035 resist was spin coated on the wafer at 2000 rpm in a programmable spin coater (Laurell Technologies), soft baked 15 min at 95 °C on a hot plate, exposed with 200 mJ/cm2 power dose of UV light with a programmable KLOE UV-KUB 2 aligner, baked for 1 min at 65 °C followed by 5 min at 95 °C, developed in SU-8 developer for 8 min, spray rinsed with fresh SU-8 developer, and blown dry. The polydimethylsiloxane (PDMS) microchannels were made with the common soft lithography process.22,23 Briefly, Sylgard 184 and its curing agent were mixed at 10:1 ratio, degassed, poured into the SU-8 molds in a home-made aluminum foil container, and cured at 65 °C on a hot plate for 120 min. The solidified PDMS was then delaminated from the molds and cut into desired pieces. The microchannels were punched for input and output connections. Oxygen plasma (200 mTorr, 10 psi gas source from an oxygen tank, 30 s, medium RF level) was used to bind the punched PDMS channels to methanol-cleaned 1 in. × 3 in. glass glides using a Plasma Cleaner PDC-32G (Harrick Plasma). The bonded device was then put on a hot plate at 150°C and left overnight to restore the hydrophobicity of PDMS before the chitosan membrane biofabrication. The microfluidic device as shown in Fig. 1(a) with a zoom-in view shown in Fig. 1(d) features three microchannels (50 μm in depth and 400 μm in width) connecting with small apertures (50 μm in all three dimensions) between PDMS pillars (100 μm long, 50 μm wide, and 50 μm deep).
FIG. 1.
Positioning yeast cells with biofabrication in microfluidics. (a) A PDMS microfluidic device with multiple sets of microchannel networks, with the zoom-in dash rectangle showing in (b) the biofabricated freestanding and semipermeable chitosan membranes connecting three microchannels; (c) schematic and (d) micrograph of the assembly of yeast cells in the alginate hydrogel; (e) schematic and (f) micrograph of assembled cells after PBS rinsing; (g) cells uniformly distributed in the alginate hydrogel; and (h) cells aggregated in the solution after 10 min without cross-linking the alginate solution into the hydrogel with calcium ions.
C. Chitosan membrane biofabrication
Chitosan membranes were biofabricated following a recently reported procedure by steering air bubbles in PDMS microchannels with an add-on vacuuming layer when needed.22,23 Briefly, the positively charged chitosan solution (pH 5.8) was pumped with a Genie syringe pump (Kent Scientific, CT) into the middle microchannel, while the negative charged alginate solutions (pH 11.5) were pumped into the two-side microchannel, at the flow rate of 0.5 μl/min in all three channels. The flows were stopped after air bubbles were trapped in the apertures due to the natural hydrophobicity of PDMS. To dissipate the air bubbles through the gas-permeable PDMS, an add-on PDMS vacuum layer with clean surface was placed on top clean surface of the device and vacuumed by simply withdrawing a syringe on a syringe pump. Upon the dissipation of air bubbles within 5–8 min, the negatively charged alginate and the positively charged chitosan solutions came into contact and instantaneously formed a thin polyelectrolyte complex membrane (PECM) layer due to electrostatic interactions. The flows were then restarted at 1 μl/min in all three channels. Diffusion of OH− from the alginate side through the PECM into the chitosan side deprotonated chitosan macromolecules and grew a chitosan membrane on each PECM to the desired thickness of 20–30 μm [Fig. 1(d)] before all the microchannels were thoroughly rinsed with PBS. The membranes were freestanding, mechanically robust, and chemically semi-permeable to small molecules,24 which was due to the flow-induced molecular microalignment of the membranes.25
D. Yeast cell assembly along chitosan membranes
Yeast cells were assembled along chitosan membranes with spatiotemporal control following previously reported procedures.23 Diffusion of Ca2+ through the semipermeable chitosan membranes cross-linked alginate and embedded cells in the gel26 as depicted in Fig. 1(b). Briefly, the mixture of alginate (pH 7, 1% w/v) and the target budding yeast α-cells, S. cerevisiae, centrifugingly concentrated to approximately 10 OD600, were introduced into the middle channel [Fig. 1(e)]. Calcium chloride solution (25 mM, pH 7) was introduced into the corresponding side channel. The cell and alginate mixture was then polymerized by the diffusion of Ca2+ from the side microchannel. The width of the hydrogel with embedded cells grew over time, and the growth rate was dependent on the concentration of calcium ions, as reported in a previous study.23 The boundary of the assembled cell–gel was noticeable when the cell–alginate solution was not in complete stop flow, for which Fig. 1(e) was chosen to show that the cells in the solution were under minimum flow. After the cell–gel layer was formed within minutes to the desired thickness, the middle channel was thoroughly rinsed with PBS manually injected into the middle channel [Figs. 1(c) and 1(f)]. Next, the mating A-cells were similarly introduced into the middle channel, assembled upon the first α-cell layer, and occupied the rest of the middle channel. The assembly order of α- and A-cells could be interchanged. Alternatively, the second cell–gel layer could be assembled in the middle channel with calcium chloride solution introduced to the other side microchannel. Therefore, α-cells and A-cells were positioned side by side in the middle channel as shown in Figs. 2(a) and 2(b) for all the experiments.
FIG. 2.
The fate of chemotropism or proliferation of mating yeast cells depended on the distance from the other cell type. (a) Schematic of the A- and α-cells assembled side by side in the alginate hydrogel in the middle channel of the three-channel device. A medium was introduced in the side microchannels to culture the cells over time; (b) relative uniform distribution of A- and α-cells at 1 h after assembly, with the area of A-cells, was separated into four blocks for further analysis in (d) and (e); (c) distribution and morphologies of A- and α-cells at 8 h after assembly, showing chemotropism of A-cells closer to α-cells while continuous proliferation of A-cells away from α-cells; (d) proliferation ratio of A-cells in the four indicated blocks in (b) and (c), showing continuous cell proliferation over 5 h in blocks 1 and 2, while little cell proliferation in block 4. Proliferation over 5 h was not accountable due to cell overlapping; and (e) Projection ratio of A-cells in the same four blocks, showing little or no chemotropism over 8 h for cells in blocks 1 and 2 while increasing chemotropism for cells in block 4.
E. Culturing of yeast mating cells in microfluidics
The 1× YPD medium was introduced into both side microchannels at 1 μl/min flow rate with Genie syringe pumps for the experimental process for 5–8 h. Nutrients were able to diffuse through the semipermeable chitosan membranes from the medium in side microchannels. The mating yeast cells in the middle microchannel were cultured over time at room temperature (22 °C) and monitored under a Ludesco EXI-310 inverted microscope. Images of the cell growth and morphology were taken every half hour. No proliferation or morphological change was observed in the first hour while the cells settled, while the yeast doubling time was slightly slower than the standard 100 min at 30 °C.
F. Image processing, cell counting, and statistics
All the images are bright field images taken with either a Nikon TS100 or a Ludesco EXI-310 inverted microscope. Images were processed using ImageJ (NIH, MD) with particle analysis function and plotted with Microsoft Excel. Only images without overlapping cells that were able to distinguish individual cells were processed for cell counts. The number of A-cells in the four denoted blocks (60 μm per block) in Fig. 2(b) and the controls, α cells close to A (within 60 μm to A-cells) and α cells far from A (farther than 180 μm from A-cells), over the countable 5-h period are shown in Table I. The proliferation ratio was defined as the ratio of cell number at a given hour to that of its initial hour at 60 min. The initial hour was at 60 min when the introduction of media solutions into the side microchannels was started. The morphology change of A-cells was further quantified by computing the projection ratio, the ratio of the long axis dimension of the elongated cells over time compared to the dimension of the corresponding A-cells at the initial hour. The calculated projection ratios over time are shown in Table II. The initial measurement of aspect ratio was not 1.0 probably due to factors including cells undergo budding (thus not perfectly round shape) and measurement errors. The cell proliferation data in Figs. 3 and 4(b) were the ratio of exact cell counts without statistics. The error bars in Fig. 4(c) represent the standard deviations of three measurements of one cell for the case of an individual A-cell or the measurements of five cells for the case of clusters of A-cells.
TABLE I.
Cell numbers and calculated proliferation ratios over time.
| Time (min) | Block 1 (A far from α) |
Block 2 | Block 3 | Block 4 (A close to α) |
α close to A | α far from A | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cell No. | Ratio | Cell No. | Ratio | Cell No. | Ratio | Cell No. | Ratio | Cell No. | Ratio | Cell No. | Ratio | |
| 60 | 23 | 1.0 | 42 | 1.0 | 25 | 1.0 | 17 | 1.0 | 46 | 1.0 | 68 | 1.0 |
| 120 | 30 | 1.3 | 48 | 1.1 | 24 | 1.0 | 17 | 1.0 | 50 | 1.1 | 79 | 1.2 |
| 180 | 42 | 1.8 | 68 | 1.6 | 38 | 1.5 | 21 | 1.2 | 65 | 1.4 | 93 | 1.4 |
| 240 | 55 | 2.4 | 85 | 2.0 | 44 | 1.8 | 23 | 1.4 | 85 | 1.8 | 116 | 1.7 |
| 300 | 74 | 3.2 | 108 | 2.6 | 49 | 2.0 | 29 | 1.7 | 101 | 2.2 | 136 | 2.0 |
TABLE II.
Projection ratio over time.
| Time (min) | Block 1 | Block 2 | Block 3 | Block 4 |
|---|---|---|---|---|
| 60 | 1.10 | 1.09 | 1.03 | 1.22 |
| 240 | 1.13 | 1.10 | 1.34 | 1.13 |
| 420 | 1.14 | 1.17 | 1.67 | 2.44 |
| 480 | 1.21 | 1.23 | 1.97 | 3.25 |
FIG. 3.
Proliferation ratios of A-cells with chemotropism (blue rectangles and black triangles) vs those of α-cells without chemotropism (red diamonds and green circles) as the control experiment. The proliferation of A-cells was spatially sensitive depending on the distance of concerned cells from the mating opposite α-cells, while the proliferation of α-cells was spatially insensitive. The dashed lines are 2nd order polynomial fits. The data points were the ratio of exact cell counts without statistics.
FIG. 4.
Response of one representative individual A-cell vs clusters of A-cells to mating signal. (a) Time lapse images of an individual (indicated with red arrows) and clusters of A-cells near α-cells; (b) the cell proliferation ratios of clusters of A-cells compared with an individual A-cell over a time period of 4.5 h. The data points were the ratio of exact cell counts without statistics; (c) the projection ratio of an individual A-cell compared with that of clusters of A-cells over time. The error bars represent the standard deviations of three measurements of one cell (individual A-cell) or the measurements of five cells (clusters of A-cells).
III. RESULTS AND DISCUSSION
A. Populations of yeast cells assembled with biofabrication in microfluidics
Biofabrication schematically shown in Figs. 1(b) and 1(c) was applied for the assembly of mating type A- and α-cell populations side by side in the middle microchannel as depicted in Scheme 1. We also used mating type A-cells that were deficient for the Bar1 protease to enhance the shmoo response, making it easier to detect signaling events. Inside the PDMS microfluidic device [Fig. 1(a)], freestanding and semi-permeable chitosan membranes were biofabricated in small apertures (50 μm in all three dimensions) connecting the three microchannels of 50 μm in depth [Fig. 1(d)]. The chitosan membrane biofabrication was done with the introduction of chitosan and alginate solutions into the middle and side microchannels, respectively, facilitated with a convenient add-on vacuuming strategy to dissipate naturally trapped air bubbles in the small apertures in hydrophobic PDMS.22 The yeast cells were then cultured over time at room temperature and monitored under microscope with nutrients (1× YPD) diffusing from the two-side microchannels.
Cells assembled in alginate gels remained uniformly distributed over time [Fig. 1(g)], while cells in the alginate solution without cross-linking with calcium ions were subject to the Brownian disturbance and tended to aggregate after 10 min [Fig. 1(h)]. Cells assembled in the alginate hydrogel were easier to monitor for morphological changes over time, while no difference in cell proliferation was observed compared with cells in the free solution. Therefore, directional chemotropic responses can be discerned by assembling cells in alginate hydrogels without being confounded by the Brownian motion.
B. The fate of chemotropism or proliferation of mating yeast was spatially dependent
By capturing cells in biologically compatible alginate hydrogels and positioning the mating yeast A- and α-cell types side by side as depicted in Fig. 2(a), the chemotropic response (cell cycle arrest, shmoo development) of yeast cells to mating pheromone was measured. Nutrients were supplied to both groups of cells with diffusion from flowing media in the side channels. The A-cells were deleted for BAR1, a gene that encodes for a protease that degrades α-factor to increase their sensitivity to α-cells.5,27 While the cell distribution was relatively uniform in the hydrogels at the first hour as shown in Fig. 2(b), the results in Fig. 2(c) show that A-cells in blocks 1 and 2 that are distal to α-cells (far right) continued to proliferate over a period of 8 h. In contrast, cells proximal (in blocks 3 and 4) to α-cells were observed to form shmoo structures and undergo cell cycle arrest, both events are indicative of the mating response to pheromone. In particular, it appeared that all cells in block 4, which were nearest to the α-cells, had formed shmoo structures. It was also observed from Fig. 2(c) that all α-cells proliferated independent of their proximity to A-cells, indicating that α-cells were less sensitive to A-factor as compared with A-cells to α-factor, which is possibly due to the hydrophobic characteristic of A-factor influencing its ability to diffuse long distances in primarily aqueous environment.28 The number cell divisions that A-cells experienced in the four indicated blocks were further quantified by calculating a proliferation ratio, the ratio of the cell numbers over time in each block compared to the cell numbers in the first hours. The results in Fig. 2(d) show that the proliferation ratios increased to 3.2, 2.6, 2.0, and 1.7 for cells in blocks 1, 2, 3, and 4, respectively, over the period of 5 h. After 5 h the density of cells made it difficult to identify individual cells. Therefore, the proliferation ratios were only calculated up to 5 h. These results indicate that the pheromone produced from the α-cells diffused and interacted with the A-cells to induce cell cycle arrest, the initial step in mating.
Furthermore, Fig. 2(c) shows that A-cells in blocks 3 and 4 closer to α-cells started to elongate besides slowing down or completely stopping proliferation over time, the morphological phenotype of chemotropism, while A-cells in blocks 1 and 2 showed no sign of elongation. The morphology change of A-cells was further quantified by computing the projection ratio, the ratio of the long axis dimension of the elongated cells over time compared to the dimension of the corresponding A-cells at the initial hour. The analysis results in Fig. 2(e) show that the projection ratios increased to 1.17, 1.23, 1.97, and 3.25 for cells in blocks 1, 2, 3, and 4, respectively, over the period of 8 h [instead of 5 h for proliferation results in Fig. 2(d)]. This indicates that chemotropism of A-cells was induced by the pheromone produced and diffused from the nearby α-cells, the morphological indication of yeast mating.
Importantly, the cell cycle arrest as well as the morphological elongation of A-cells was spatially dependent on the distance from the α-cells on the scale of tens of micrometers. The closer the A-cells are to the α-cells, presumably the more the pheromone signal, and the more obvious effect on the cell cycle arrest and morphological projections. To further confirm the spatial dependence of the cell cycle arrest in A-cells, the cell proliferation ratios of the α-cells not experiencing chemotropism, both near and far from A-cells, were also analyzed as controls. The proliferation ratios of A-cells, both near and far from α-cells, were plotted in Fig. 3 with 2nd order polynomial fits indicating the corresponding cell growth curves. The results show that the proliferation ratios at 5 h for A-cells far from α-cells and those near α-cells were 3.2 and 1.7, respectively. In other words, the proliferation rate for A-cells far away from α-cells was almost double of that for A-cells close to α-cells. On the contrary, as a control experiment, the proliferation ratio of the α-cells that were near to A-cells compared with those that were far from A-cells were 2.2 and 2.0, respectively, at 5 h. In other words, the proliferation ratios were nearly the same throughout the population of α-cells. Together, these results demonstrate that the proliferation of A-cells was spatially sensitive depending on the distance of concerned cells from the mating opposite α-cells, while the proliferation of α-cells was spatially insensitive.
In summary, we observed that the fate of MAT A-cells (i.e., mating chemotropism or continuous proliferation) depended on their relative proximity to MAT α-cells. The spatial resolution was observed on the scale of tens of micrometers.
C. Cell density influenced the chemotropic responses to mating factor
During the experiments, we observed differences in shmoo formation in A-cells that were well separated from other A-cells compared to A-cells in cell clusters. One representative example is shown in Fig. 4(a), in which an individual A-cell remained undivided over 4.5 h. It elongated dramatically at 2.5 h and elongated much more at 4.5 h. In contrast, A-cells in clusters divided 3.7 times [Fig. 4(b)] during the same period and were approximately the same distance to α-cells as the individual A-cell. At 2.5 h, no elongation was observed for the cells in clusters. At 4.5 h, the A-cells around the edge of clusters also elongated toward the source of α-factor from nearby α-cells. The chemotropism of A-cells in clusters clearly happened at a later stage as compared with that of the individual A-cell.
The projection ratio of the isolated individual A-cell and the average projection ratio of clusters of A-cells were further processed and quantified using ImageJ. The results in Fig. 4(c) show that the projection ratio of the isolated A-cell increased from 1.12 ± 0.02 at 0.5 h to 2.81 ± 0.10 at 2.5 h and finally by 4.5 h, the projection ratio reached 5.57 ± 0.15. In contrast, the average projection ratio of A-cells within clusters minimally increased from 1.07 ± 0.04 at 0.5 h to 1.16 ± 0.17 at 2.5 h and reached a maximum projection ratio of 3.48 ± 0.56 by 4.5 h.
These results reveal that the chemotropic response during mating is influenced by cell density. For individual A-cells, which were predicted to face ample α-pheromone, chemotropism dominated with the shmoo growth directed toward the α-cells. In contrast, clusters of A-cells, which were predicted to face less α-pheromone, initially proliferated but those around the edge of the clusters transitioned from dividing to cell cycle arrest and shmoo formation, presumably due to an increase in the concentration of pheromone generated from nearby α-cells.
D. Directional shmoo projections are gradient-driven or concentration-dependent
There were A-cells displaying characteristic chemotropism, with shmoo tips growing toward the source of α-factor pheromone from α-cells as shown in Fig. 4(a). However, not all A-cells undergoing chemotropism pointed in the same direction as observed in Fig. 2(c). To better understand this, we performed two experiments with different density of A-cells. The results are shown in Fig. 5, where the main difference was that Fig. 5(a) has more A-cells than Fig. 5(b) within the viewing windows. The A-cells in Fig. 5(a) in the three indicated regions showed different levels of chemotropism over the observation time of 4.5 h. This is most obvious in the evolving images at 210 and 270 min. The growth of shmoo morphologies in cells growing in region i was non-directional. In contrast, the projections of cells in region ii all pointed toward the region where α-cells were present, the source of α-factor. In region iii, the most distal from α-cells, A-cells remained mostly round, indicating a lack of chemotropic response. Overall, directional chemotropism was observed in a spatially confined space of tens of micrometers in region ii. When the density of A-cells was reduced to less than half of what was in observed in Fig. 5(a), shmoo projections grew in multiple directions rather than toward the source of mating factor secreted by α-cells as shown in Fig. 5(b).
FIG. 5.
Directional yeast chemotropism was pheromone gradient-dependent. (a) In the case of high density of A-cells near α-cells, the α pheromone signal was consumed by A-cells resulting in sharper gradient, thus the chemotropism of A-cells transited from non-directional (region i) to unidirectional (region ii) to little chemotropism (region iii). This is most obvious in the evolving images at 210 and 270 min. (b) In the case of low density of A-cells near α-cells, the α-factor was saturated around A-cells with little gradient, thus the chemotropism of A-cells remained non-directional.
We hypothesize that the chemotropic response that directs shmoo formation toward the α-pheromone source (i.e., α-cells) occurs at intermediate levels of α-pheromone. The color stripes on the top of images in Figs. 5(a) and 5(b) depict the possible difference in the gradient of α-pheromone in Figs. 5(a) and 5(b). Since there were more A-cells to consume α-pheromone in Fig. 5(a), a sharp gradient of α factors might exist from the border of α/A-cells. As the α-pheromone gradient traversed from regions i–iii, we envision the following: cells in region i might have been saturated with α-pheromone, cells in region ii might have experienced the threshold level of α-pheromone, while cells in region iii might have α-pheromone fallen below the threshold level for chemotropism to occur. For the case in Fig. 5(b), since there were less A-cells to consume the mating α-pheromone, the gradient from the border of α/A-cells toward the right might be very shallow and all the A-cells might be saturated with excessive mating α-pheromone. Therefore, no chemotropism toward the α-cells was observed in Fig. 5(b).
The results suggested that the direction of shmoo projection formation is gradient-driven or concentration-dependent within a spatial resolution of tens of micrometers.
E. Multi-lobed shmoo morphologies
Previous work has shown that A-cells growing in a field of synthetic pheromone will make multiple attempts to form shmoo projections resulting in multi-lobed cells.29 However, it is unclear if these projections (or lobes) would be directed toward the source since cells are not immobilized and can diffuse in multiple directions in these studies. However, the cells in the hydrogels have much more limited motion, which can help one to distinguish if the lobes display directionality. We observed a similar development of multi-lobed cells in highly isolated A-cells over a period of 8 h as shown in Fig. 6(a). The single cell was positioned near the semipermeable chitosan membrane that was about 160 μm away from α-cells. The single A-cells proliferated after 4 h, while initial chemotropism of the mother cell was also observed toward the left. The elongation of the mother cell continued at 5 h, and at 6 h, the daughter cell also elongated in the same direction, while the mother cell developed a second lobe in the lower direction. At 7 h, both mother and daughter cells had produced more than one lobe, and the process continued until 8 h when the experiment was terminated. The multi-lobe cell morphology was not commonly observed using conventional platforms where single cells were not easy to position around a population of opposite mating cells with easy control. Furthermore, because the cells were trapped in the hydrogel, the direction of the shmoo was surely not due to rotation of the cell but most likely due to the response to pheromone. This further confirms the unique advantage of using biofabrication strategy in microfluidics for cell assembly and cellular studies with spatial programmability.
FIG. 6.
Multi-lobed cell morphology under saturated α pheromone signal. (a) Representative multi-lobed morphology of one A-cell (red rectangle) under the configuration that a few A-cells (right) were assembled near to abundant α-cells (left); (b) Time lapse images of the representative A-cell growing into multi-lobed morphology at 3–8 h.
F. Challenges with in vivo chemotropism study
Microfluidic devices have provided new insights into the gradient response14–17 and contributed to improved predictive models. A crucial variable that needs to be included is the potential dynamics of a gradient formed between populations of cells akin to natural environments.30 The biofabricated microfluidic platform we report here brings together populations of mating types A- and α-cells side by side. This is useful to inform on the in vivo chemotropism that accounts for the dynamism which can occur in environments occupied by populations of cells. Our ability to position populations vs individual yeast cells within proximity of each other is straightforward, and the culturing with flow media in the side microchannels separated with porous biopolymer membranes is unique and beneficial for monitoring the chemotropic response of cells over time. One common challenge of studying chemotropism as presented in this report is that the exact amount of the pheromone concentration and gradients cannot be measured directly and dynamically over time. This calls for the long-desired in situ biosensors for measuring signaling molecules within and among microbial species that have drawn much interest in research community. Meanwhile, we believe that the current study provides additional insights into many well-characterized studies of the concentration and gradient-dependent response of mating yeast with synthetically derived pheromones.31
IV. CONCLUSION
We report the chemotropic signaling between populations of mating yeast A- and α-cells with spatiotemporal programmability and sensitivity that were revealed using the biofabrication strategy in a microfluidic device. Biofabricated freestanding and semipermeable chitosan membranes in a microfluidic device provide both the facile assembly of cell populations with micrometer spatial control and the convenient culturing of the assembled cells. The morphological changes of mating yeast cells were imaged with live-cell microscopy to gain an insight into the mating response between cell populations. The mating phenotypes of yeast cells were clearly observed over hours with high spatial resolution and sensitivity. Three distinct responses were observed depending on the distance between A- and α-cell populations: the cells either proliferated, developed a stereotypical polarized projection termed a “shmoo” directed toward the cells of the opposite mating type, or polarized growth that appeared to be non-directional. The demonstrated spatial resolution and sensitivity provide perceptions to further understand how natural pheromone gradients induce a chemotropic response. The observed cell–cell signaling can inform on improving mathematical models of chemotropic responses of yeast populations and potentially chemotropism in other types of cells.
ACKNOWLEDGMENTS
This work was supported in part by the National Science Foundation CAREER Grant No. 1553330 to X.L., the National Institute of Health (NIH) Grant No. 1R15GM129766-01 to X.L. and J.S.C., the Litovitz Family Fund to J.S.C., and the School of Engineering and the School of Arts and Sciences at the Catholic University of America.
Contributor Information
John S. Choy, Email: .
Xiaolong Luo, Email: .
REFERENCES
- 1.Cai H. and Devreotes P. N., “Moving in the right direction: How eukaryotic cells migrate along chemical gradients,” Semin. Cell Dev. Biol. 22(8), 834–841 (2011). 10.1016/j.semcdb.2011.07.020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Tojima T., Hines J. H., Henley J. R., and Kamiguchi H., “Second messengers and membrane trafficking direct and organize growth cone steering,” Nat. Rev. Neurosci. 12(4), 191–203 (2011). 10.1038/nrn2996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Palanivelu R. and Preuss D., “Pollen tube targeting and axon guidance: Parallels in tip growth mechanisms,” Trends Cell Biol. 10(12), 517–524 (2000). 10.1016/S0962-8924(00)01849-3 [DOI] [PubMed] [Google Scholar]
- 4.Ismael A. and Stone D. E., “Yeast chemotropism: A paradigm shift in chemical gradient sensing,” Cell Logist. 7(2), e1314237 (2017). 10.1080/21592799.2017.1314237 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Arkowitz R. A., “Chemical gradients and chemotropism in yeast,” Cold Spring Harb. Perspect. Biol. 1(2), a001958 (2009). 10.1101/cshperspect.a001958 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bardwell L., “A walk-through of the yeast mating pheromone response pathway,” Peptides 26(2), 339–350 (2005). 10.1016/j.peptides.2004.10.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Segall J. E., “Polarization of yeast cells in spatial gradients of alpha mating factor,” Proc. Natl. Acad. Sci. U.S.A. 90(18), 8332–8336 (1993). 10.1073/pnas.90.18.8332 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Marinkovic Z. S., Vulin C., Acman M. et al. , “A microfluidic device for inferring metabolic landscapes in yeast monolayer colonies,” eLife 8, e47951 (2019). 10.7554/eLife.47951 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Somaweera H., Ibraguimov A., and Pappas D., “A review of chemical gradient systems for cell analysis,” Anal. Chim. Acta 907, 7–17 (2016). 10.1016/j.aca.2015.12.008 [DOI] [PubMed] [Google Scholar]
- 10.Lin B. and Levchenko A., “Spatial manipulation with microfluidics,” Front. Bioeng. Biotechnol. 3, 39–39 (2015). 10.3389/fbioe.2015.00039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wang Z., Lee I., Jeon T.-J., and Kim S. M., “Micro-/nanofluidic device for tunable generation of a concentration gradient: Application to Caenorhabditis elegans chemotaxis,” Anal. Bioanal. Chem. 406(11), 2679–2686 (2014). 10.1007/s00216-014-7663-5 [DOI] [PubMed] [Google Scholar]
- 12.Yanagisawa N. and Higashiyama T., “Quantitative assessment of chemotropism in pollen tubes using microslit channel filters,” Biomicrofluidics 12(2), 024113 (2018). 10.1063/1.5023718 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Agudelo C. G., Sanati Nezhad A., Ghanbari M., Naghavi M., Packirisamy M., and Geitmann A., “Tipchip a modular, MEMS-based platform for experimentation and phenotyping of tip-growing cells,” Plant J. 73(6), 1057–1068 (2013). 10.1111/tpj.12093 [DOI] [PubMed] [Google Scholar]
- 14.Brett M. E., DeFlorio R., Stone D. E., and Eddington D. T., “A microfluidic device that forms and redirects pheromone gradients to study chemotropism in yeast,” Lab Chip 12(17), 3127–3134 (2012). 10.1039/c2lc40398f [DOI] [PubMed] [Google Scholar]
- 15.Moore T. I., Chou C. S., Nie Q., Jeon N. L., and Yi T. M., “Robust spatial sensing of mating pheromone gradients by yeast cells,” PLoS One 3(12), e3865 (2008). 10.1371/journal.pone.0003865 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Muller N., Piel M., Calvez V. et al. , “A predictive model for yeast cell polarization in pheromone gradients,” PLoS Comput. Biol. 12(4), e1004795 (2016). 10.1371/journal.pcbi.1004795 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Luo X., Vo T., Jambi F., Pham P., and Choy J. S., “Microfluidic partition with in situ biofabricated semipermeable biopolymer membranes for static gradient generation,” Lab Chip 16(19), 3815–3823 (2016). 10.1039/C6LC00742B [DOI] [PubMed] [Google Scholar]
- 18.Brett M.-E., DeFlorio R., Stone D. E., and Eddington D. T., “A microfluidic device that forms and redirects pheromone gradients to study chemotropism in yeast,” Lab Chip 12(17), 3127–3134 (2012). 10.1039/c2lc40398f [DOI] [PubMed] [Google Scholar]
- 19.Jin S. H., Lee S. S., Lee B., Jeong S. G., Peter M., and Lee C. S., “Programmable static droplet array for the analysis of cell-cell communication in a confined microenvironment,” Anal. Chem. 89(18), 9722–9729 (2017). 10.1021/acs.analchem.7b01462 [DOI] [PubMed] [Google Scholar]
- 20.Luo X., Wu H.-C., Tsao C.-Y. et al. , “Biofabrication of stratified biofilm mimics for observation and control of bacterial signaling,” Biomaterials 33(20), 5136–5143 (2012). 10.1016/j.biomaterials.2012.03.037 [DOI] [PubMed] [Google Scholar]
- 21.Guthrie C. and Fink G. R., “Guide to yeast genetics and molecular biology,” Methods Enzymol. 194, 3–933 (1991). 10.1016/0076-6879(91)94004-V [DOI] [PubMed] [Google Scholar]
- 22.Pham P., Vo T., and Luo X., “Steering air bubbles with an add-on vacuum layer for biopolymer membrane biofabrication in PDMS microfluidics,” Lab Chip 17(2), 248–255 (2017). 10.1039/C6LC01362G [DOI] [PubMed] [Google Scholar]
- 23.Pham P. L. H., Rooholghodos S. A., Choy J. S., and Luo X., “Constructing synthetic ecosystems with biopolymer fluitrodes,” Adv. Biosyst. 2(3), 1700180 (2018). 10.1002/adbi.201700180 [DOI] [Google Scholar]
- 24.Luo X., Wu H.-C., Betz J., Rubloff G. W., and Bentley W. E., “Air bubble-initiated biofabrication of freestanding, semi-permeable biopolymer membranes in PDMS microfluidics,” Biochem. Eng. J. 89, 2–9 (2014). 10.1016/j.bej.2013.12.013 [DOI] [Google Scholar]
- 25.Li K., Correa S. O., Pham P., Raub C. B., and Luo X., “Birefringence of flow-assembled chitosan membranes in microfluidics,” Biofabrication 9(3), 034101 (2017). 10.1088/1758-5090/aa786e [DOI] [PubMed] [Google Scholar]
- 26.Grant G. T., Morris E. R., Rees D. A., Smith P. J. C., and Thom D., “Biological interactions between polysaccharides and divalent cations: The egg-box model,” FEBS Lett. 32(1), 195–198 (1973). 10.1016/0014-5793(73)80770-7 [DOI] [Google Scholar]
- 27.Ciejek E. and Thorner J., “Recovery of S. cerevisiae a cells from G1 arrest by alpha factor pheromone requires endopeptidase action,” Cell 18(3), 623–635 (1979). 10.1016/0092-8674(79)90117-X [DOI] [PubMed] [Google Scholar]
- 28.Marsh L., Neiman A. M., and Herskowitz I., “Signal transduction during pheromone response in yeast,” Annu. Rev. Cell Biol. 7, 699–728 (1991). 10.1146/annurev.cb.07.110191.003411 [DOI] [PubMed] [Google Scholar]
- 29.Diener C., Schreiber G., Giese W., del Rio G., Schroder A., and Klipp E., “Yeast mating and image-based quantification of spatial pattern formation,” PLoS Comput. Biol. 10(6), e1003690 (2014). 10.1371/journal.pcbi.1003690 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Song J., Ryu H., Chung M. et al. , “Microfluidic platform for single cell analysis under dynamic spatial and temporal stimulation,” Biosens. Bioelectron. 104, 58–64 (2018). 10.1016/j.bios.2017.12.038 [DOI] [PubMed] [Google Scholar]
- 31.Moore T. I., Chou C. S., Nie Q., Jeon N. L., and Yi T. M., “Robust spatial sensing of mating pheromone gradients by yeast cells,” PLoS One 3(12), e3865 (2008). 10.1371/journal.pone.0003865 [DOI] [PMC free article] [PubMed] [Google Scholar]